Variation in Type A Trichothecene Production and Trichothecene Biosynthetic Genes in Fusarium goolgardi from Natural Ecosystems of Australia
Abstract
:1. Introduction
2. Results
2.1. Mycotoxin Analysis

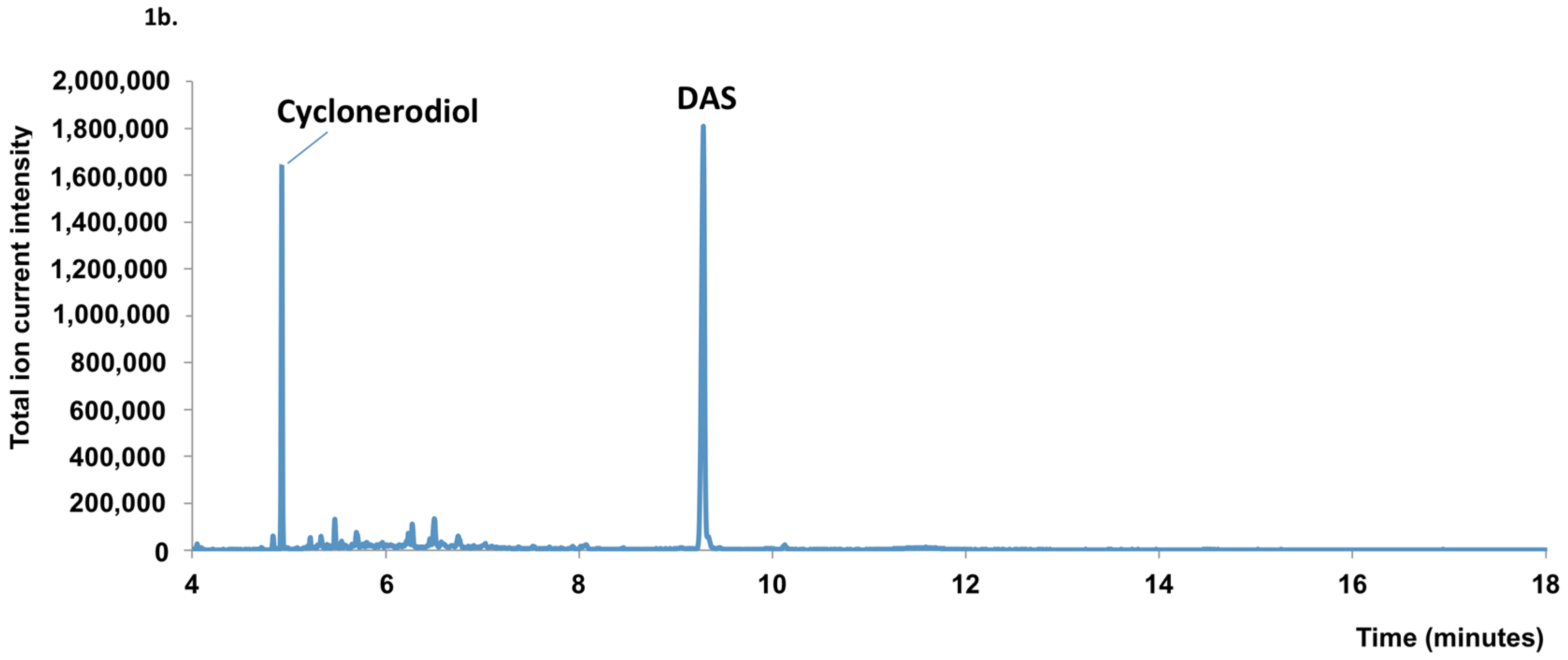
2.2. Sequence Analysis
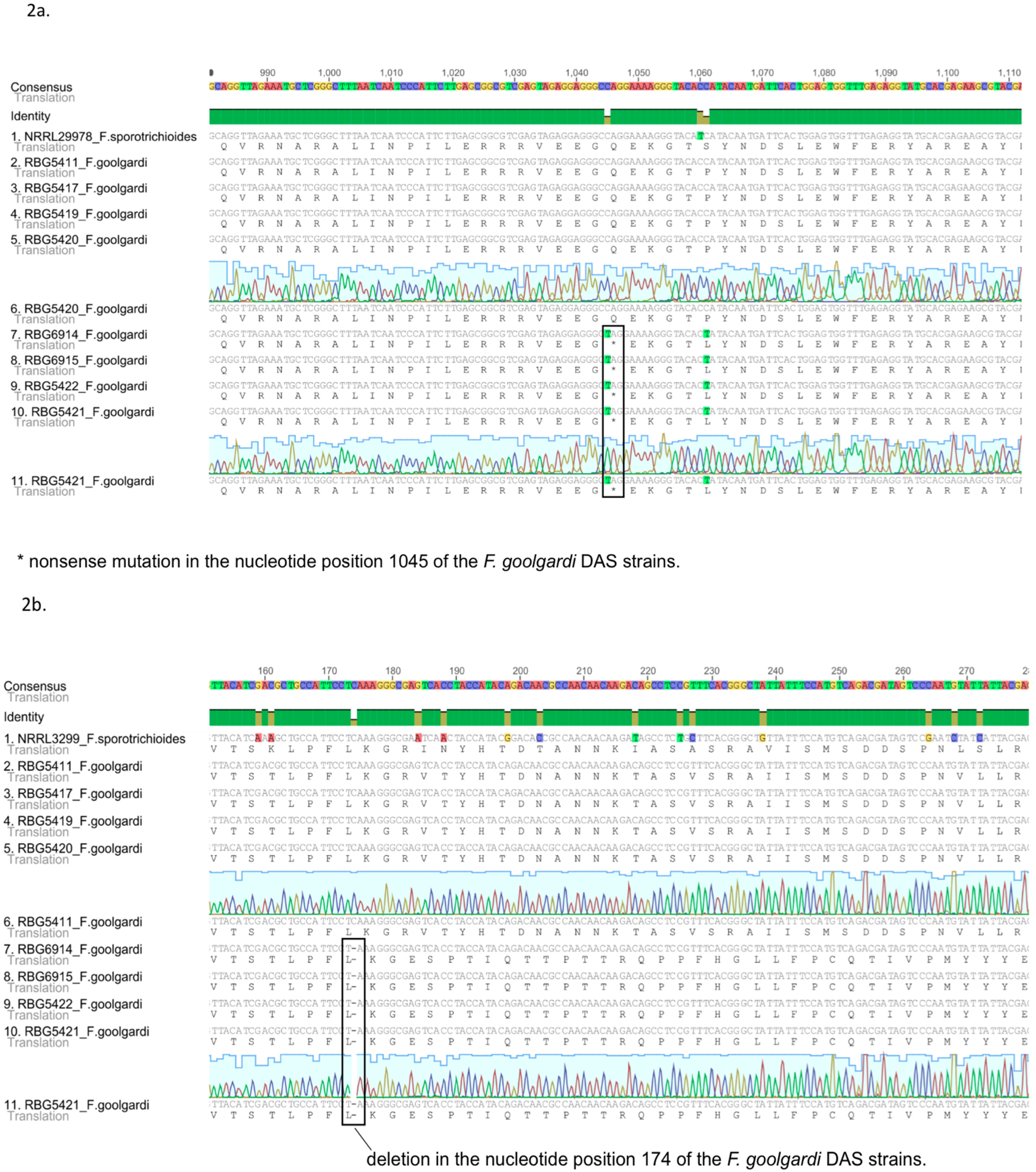
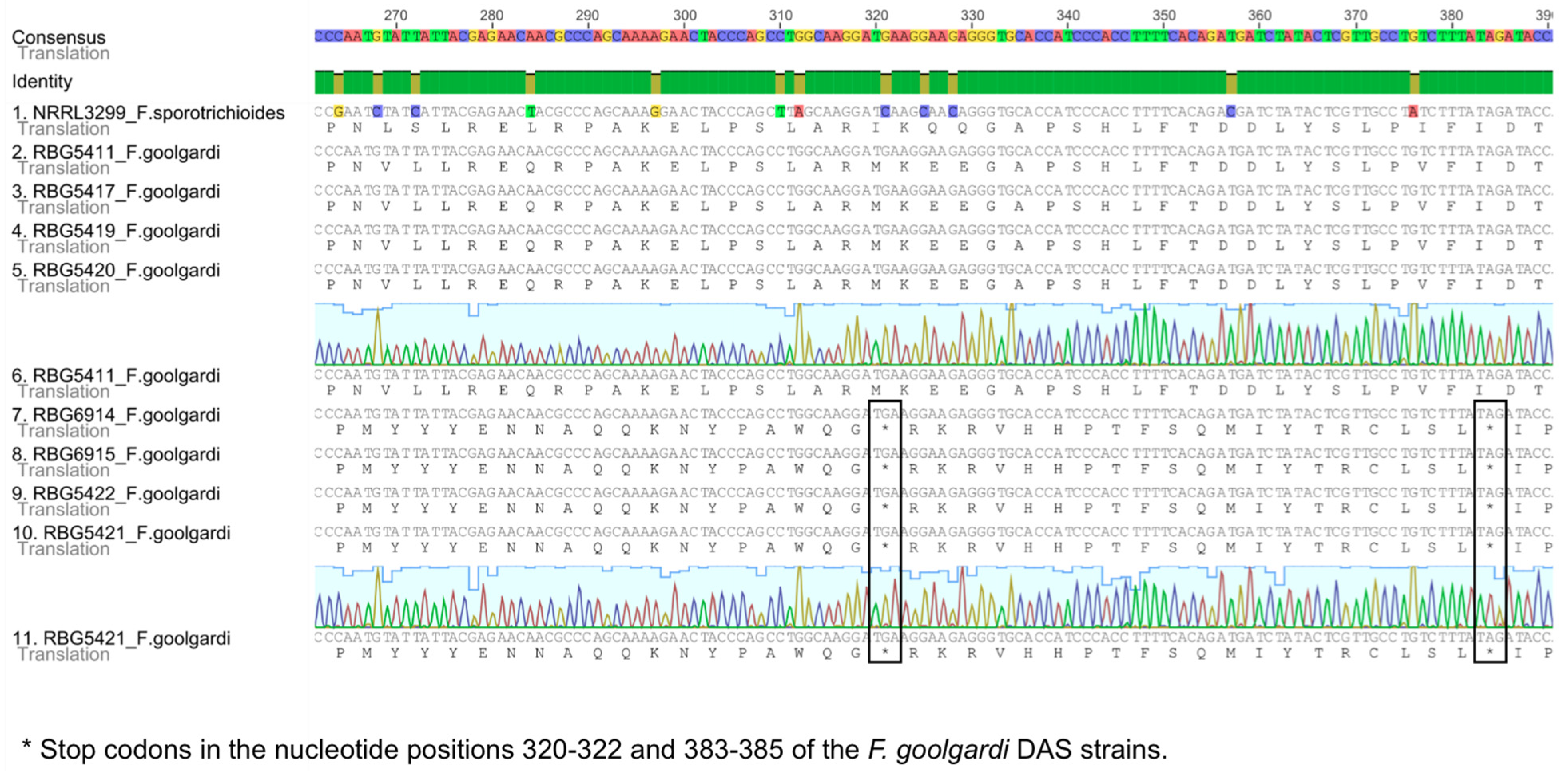
2.3. Phylogenetic Analyses
2.3.1. RNA Polymerase II Largest (RPB1) and Second Largest (RPB2) Subunits
2.3.2. TRI Gene Cluster
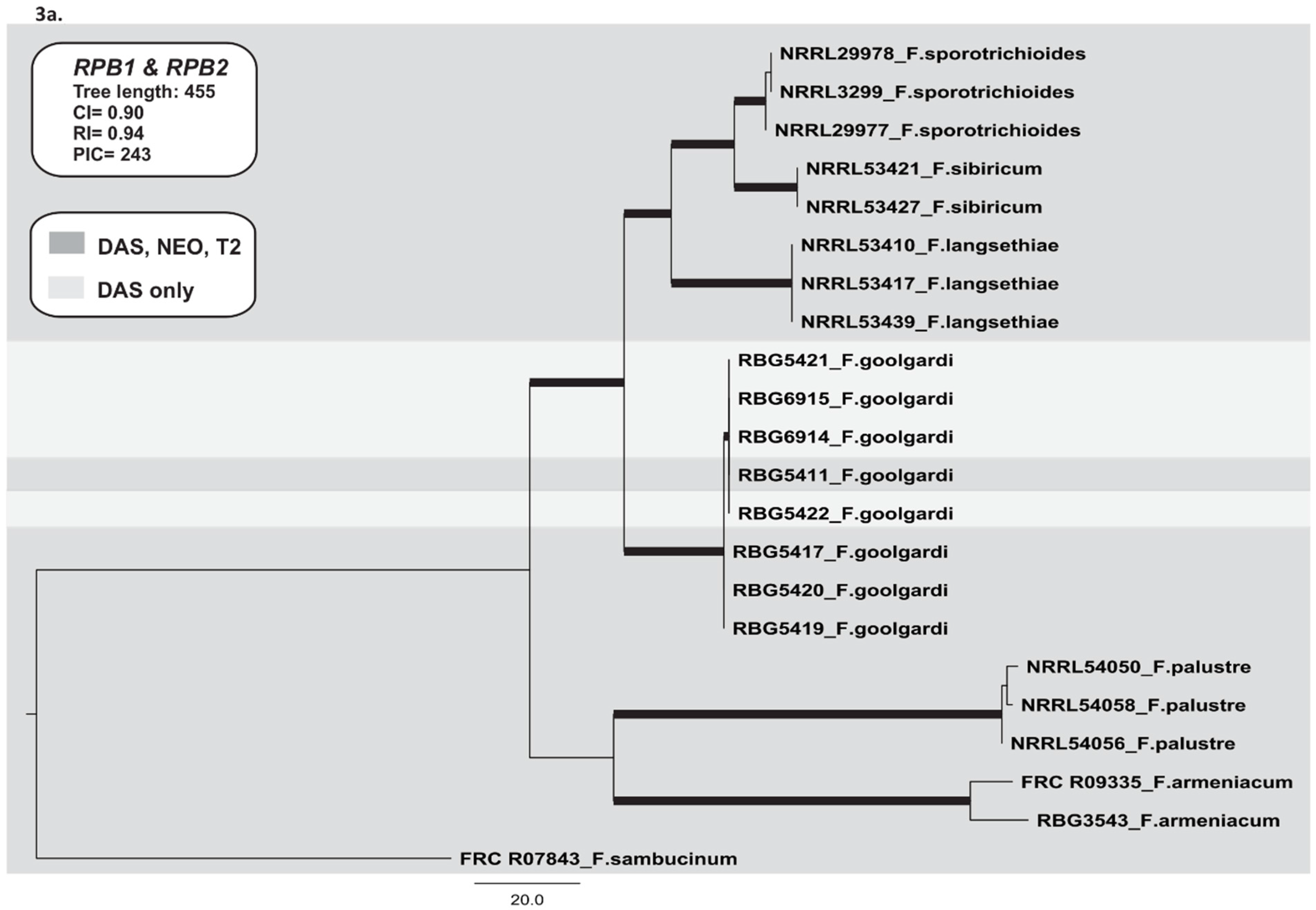
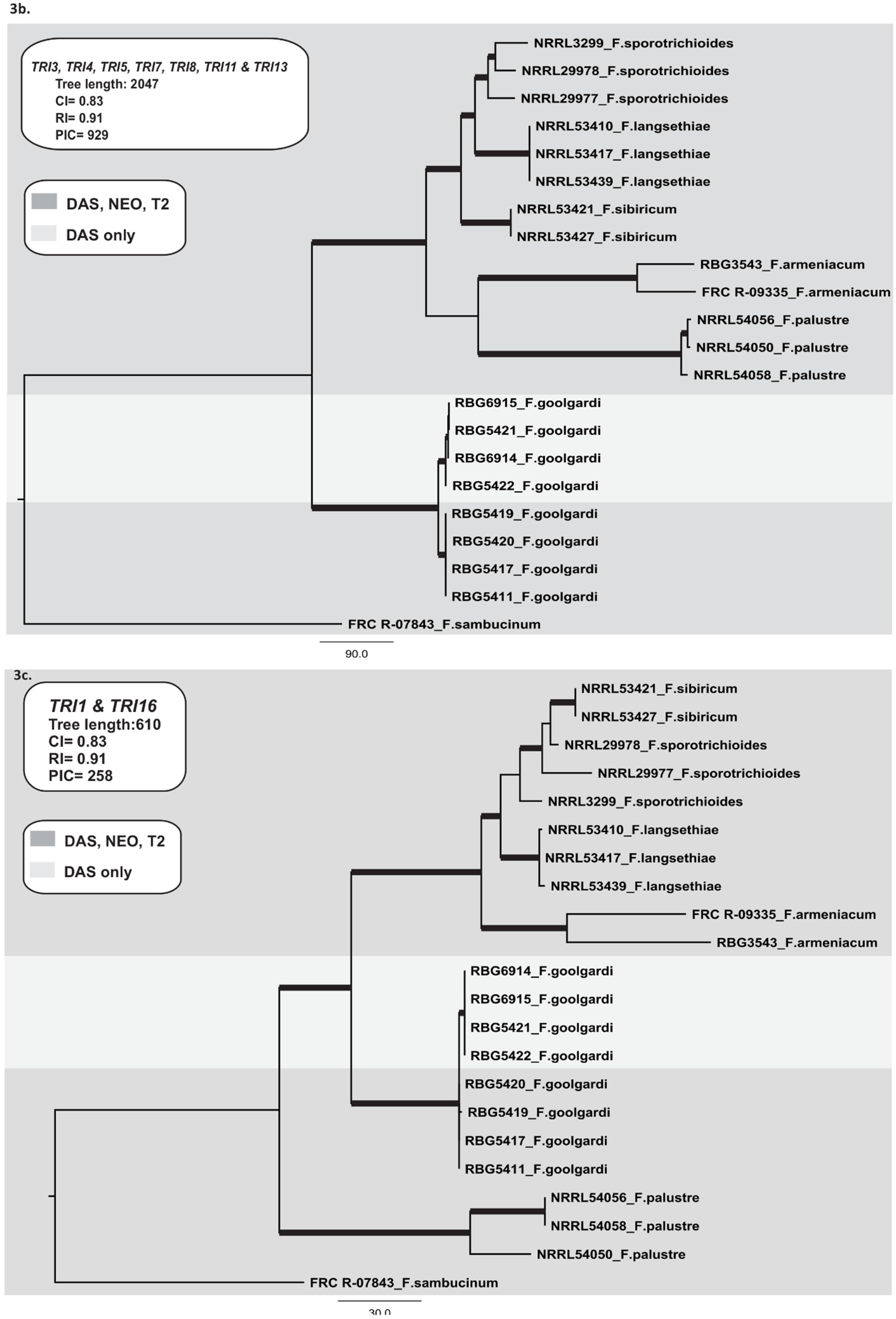
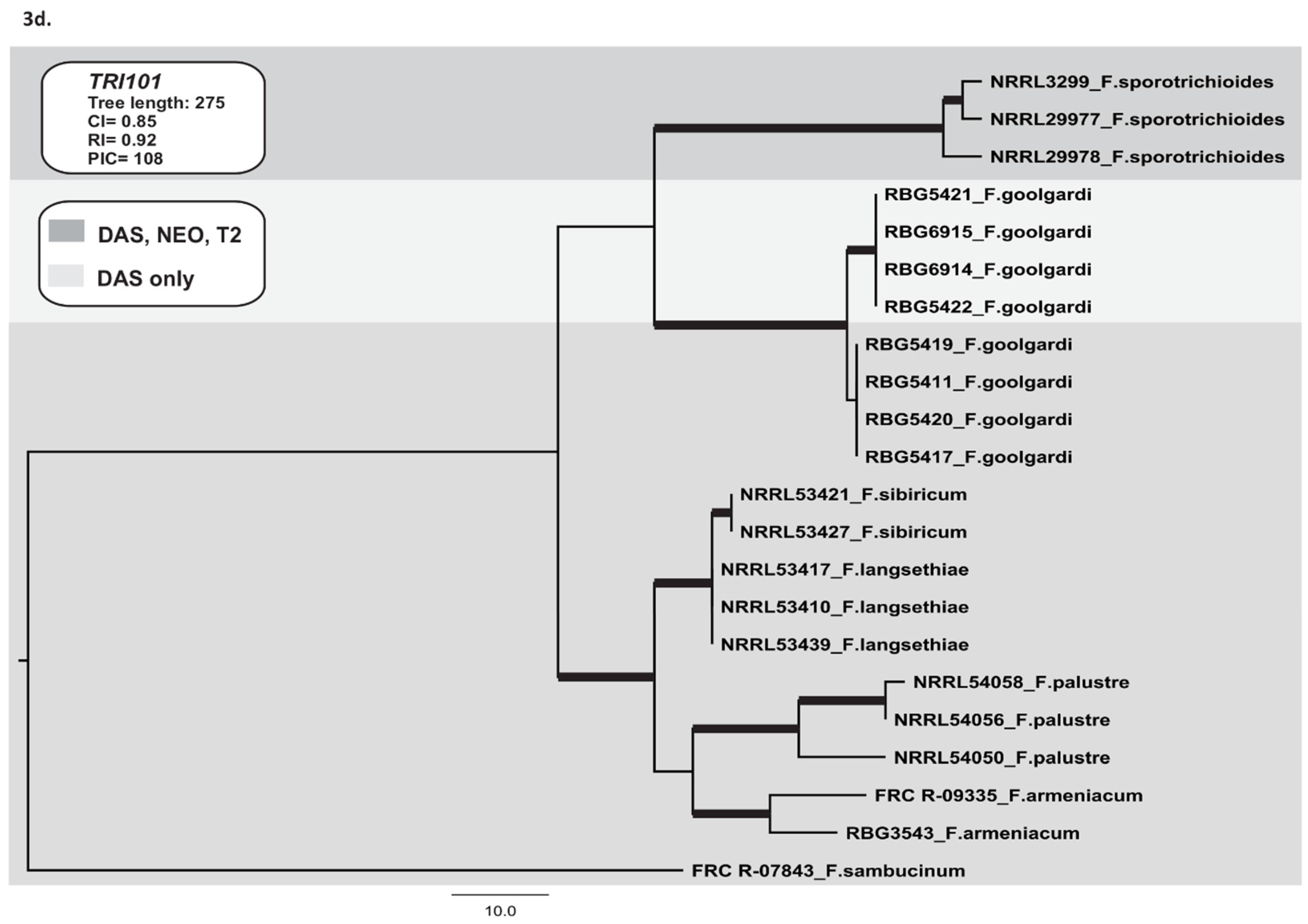
2.3.3. TRI1 and TRI16
2.3.4. TRI101
3. Discussion
4. Experimental Section
4.1. Fusarium Isolates
4.2. Mycotoxin Analysis
4.3. Locus Selection
4.4. DNA Extraction, PCR Amplification and Sequence Analysis
4.5. Phylogenetic Analyses
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wang, B.; Brubaker, C.L.; Burdon, J.J. Fusarium species and Fusarium wilt pathogens associated with native Gossypium populations in Australia. Mycol. Res. 2004, 108, 35–44. [Google Scholar] [CrossRef] [PubMed]
- Bentley, A.R.; Petrovic, T.; Griffiths, S.P.; Burgess, L.W.; Summerell, B.A. Crop pathogens and other Fusarium species associated with Austrostipa aristiglumis. Australas. Plant Pathol. 2007, 36, 434–438. [Google Scholar] [CrossRef]
- Petrovic, T.; Burgess, L.W.; Cowie, I.; Warren, R.A.; Harvey, P.R. Diversity and fertility of Fusarium sacchari from wild rice (Oryza australiensis) in Northern Australia, and pathogenicity tests with wild rice, rice, sorghum and maize. Eur. J. Plant Pathol. 2013, 136, 773–788. [Google Scholar] [CrossRef]
- Laurence, M.H.; Walsh, J.L.; Shuttleworth, L.A.; Robinson, D.M.; Johansen, R.M.; Petrovic, T.; Vu, T.T.H.; Burgess, L.W.; Summerell, B.A.; Liew, E.C.Y. Six novel species of Fusarium from natural ecosystems in Australia. Fungal Divers. 2015. [Google Scholar] [CrossRef]
- Elmer, W.H.; Marra, R.E. New species of Fusarium associated with dieback of Spartina alterniflora in Atlantic salt marshes. Mycologia 2011, 103, 806–819. [Google Scholar] [CrossRef] [PubMed]
- Eudes, F.; Comeau, A.; Rioux, S.; Collin, J. Impact of trichothecenes on Fusarium head blight (Fusarium graminearum) development in spring wheat (Triticum aestivum). Can. J. Plant Pathol. 2001, 23, 318–322. [Google Scholar] [CrossRef]
- Desjardins, A.E.; Hohn, T.M. Mycotoxins in plant pathogenesis. Mol. Plant Microbe Interact. 1997, 10, 147–152. [Google Scholar] [CrossRef]
- Cuzick, A.; Urban, M.; Hammond-Kosack, K. Fusarium graminearum gene deletion mutants map1 and tri5 reveal similarities and differences in the pathogenicity requirements to cause disease on Arabidopsis and wheat floral tissue. New Phytol. 2008, 177, 990–1000. [Google Scholar] [CrossRef] [PubMed]
- Proctor, R.H.; McCormick, S.P.; Alexander, N.J.; Desjardins, A.E. Evidence that a secondary metabolic biosynthetic gene cluster has grown by gene relocation during evolution of the filamentous fungus Fusarium. Mol. Microbiol. 2009, 74, 1128–1142. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M.; Tokai, T.; Takahashi-Ando, N.; Ohsato, S.; Fujimura, M. Molecular and genetic studies of Fusarium trichothecene biosynthesis: Pathways, genes, and evolution. Biosci. Biotechnol. Biochem. 2007, 71, 2105–2123. [Google Scholar] [CrossRef] [PubMed]
- McCormick, S.P.; Stanley, A.M.; Stover, N.A.; Alexander, N.J. Trichothecenes: From simple to complex mycotoxins. Toxins 2011, 3, 802–814. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Jing, L.; Yuan, H.; Peng, S. T-2 toxin induces apoptosis in ovarian granulosa cells of rats through reactive oxygen species-mediated mitochondrial pathway. Toxicol. Lett. 2011, 202, 168–177. [Google Scholar] [CrossRef] [PubMed]
- Torp, M.; Langseth, W. Production of T-2 toxin by a Fusarium resembling Fusarium poae. Mycopathologia 1999, 147, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Rocha, O.; Ansari, K.; Doohan, F.M. Effects of trichothecene mycotoxins on eukaryotic cells: A review. Food Addit. Contam. 2015, 22, 369–378. [Google Scholar] [CrossRef] [PubMed]
- Masuda, D.; Ishida, M.; Yamaguchi, K.; Yamaguchi, I.; Kimura, M.; Nishiuchi, T. Phytotoxic effects of trichothecenes on the growth and morphology of Arabidopsis thaliana. J. Exp. Bot. 2007, 58, 1617–1626. [Google Scholar] [CrossRef] [PubMed]
- Desjardins, A.E. Fusarium Mycotoxins: Chemistry, Genetics and Biology; APS Press: St. Paul, MN, USA, 2006; pp. 33–53. [Google Scholar]
- O’Donnell, K.; Kistler, H.C.; Tacke, B.K.; Casper, H.H. Gene genealogies reveal global phylogeographic structure and reproductive isolation among lineages of Fusarium graminearum, the fungus causing wheat scab. Proc. Natl. Acad. Sci. USA 2000, 97, 7905–7910. [Google Scholar] [CrossRef] [PubMed]
- Ward, T.J.; Bielawski, J.P.; Kistler, H.C.; Sullivan, E.; O’Donnell, K. Ancestral polymorphism and adaptive evolution in the trichothecene mycotoxin gene cluster of phytopathogenic Fusarium. Proc. Natl. Acad. Sci. USA 2002, 99, 9278–9283. [Google Scholar] [CrossRef] [PubMed]
- Aoki, T.; Ward, T.J.; Kistler, H.C.; O’Donnell, K. Systematics, phylogeny and trichothecene mycotoxin potential of Fusarium Head Blight cereal pathogens. Mycotoxins 2012, 62, 91–102. [Google Scholar] [CrossRef]
- O’Donnell, K.; Rooney, A.P.; Proctor, R.H.; Brown, D.W.; McCormick, S.P.; Ward, T.J.; Frandsen, R.J.N.; Lysøe, E.; Rehner, S.A.; Aoki, T.; et al. Phylogenetic analyses of RPB1 and RPB2 support a middle Cretaceous origin for a clade comprising all agriculturally and medically important fusaria. Fungal Genet. Biol. 2013, 52, 20–31. [Google Scholar] [CrossRef] [PubMed]
- Marasas, W.F.O.; Yagen, B.; Sydenham, E.W.; Combrinck, S.; Thiel, P.G. Comparative yields of T-2 toxin and related trichothecenes from five toxicologically important strains of Fusarium sporotrichioides. Appl. Environ. Microbiol. 1987, 53, 693–696. [Google Scholar] [PubMed]
- Langseth, W.; Bernhoft, A.; Rundberget, T.; Kosiak, B.; Gareis, M. Mycotoxin production and cytotoxicity of Fusarium strains isolated from Norwegian cereals. Mycopathologia 1999, 144, 103–113. [Google Scholar] [CrossRef]
- Thrane, U.; Adler, A.; Clasen, P.E.; Galvano, F.; Langseth, W.; Lew, H.; Logrieco, A.; Nielsen, K.F.; Ritieni, A. Diversity in metabolite production by Fusarium langsethiae, Fusarium poae, and Fusarium sporotrichioides. Int. J. Food Microbiol. 2004, 95, 257–266. [Google Scholar] [CrossRef] [PubMed]
- Yli-Mattila, T.; Ward, T.J.; O’Donnell, K.; Proctor, R.H.; Burkin, A.A.; Kononenko, G.P.; Gavrilova, O.P.; Aoki, T.; McCormick, S.P.; Gagkaeva, T.Y. Fusarium sibiricum sp. nov, a novel type A trichothecene-producing Fusarium from northern Asia closely related to F. sporotrichioides and F. langsethiae. Int. J. Food Microbiol. 2011, 147, 58–68. [Google Scholar] [CrossRef] [PubMed]
- Beremand, M.N.; Desjardins, A.E. Trichothecene biosynthesis in Gibberella pulicaris: Inheritance of C-8 hydroxylation. J. Ind. Microbiol. 1988, 3, 167–174. [Google Scholar] [CrossRef]
- Brown, D.W.; Proctor, R.H.; Dyer, R.B.; Plattner, R.D. Characterization of a Fusarium 2-gene cluster involved in trichothecene C-8 modification. J. Agric. Food Chem. 2003, 51, 7936–7944. [Google Scholar] [CrossRef] [PubMed]
- Meek, I.B.; Peplow, A.W.; Ake, C.; Phillips, T.D.; Beremand, M.N. Tri1 encodes the cytochrome P450 monooxygenase for C-8 hydroxylation during trichothecene biosynthesis in Fusarium sporotrichioides and resides upstream of another new Tri gene. Appl. Environ. Microbiol. 2003, 69, 1607–1613. [Google Scholar] [CrossRef] [PubMed]
- Peplow, A.W.; Meek, I.B.; Wiles, M.C.; Phillips, T.D.; Beremand, M.N. Tri16 is required for esterification of position C-8 during trichothecene mycotoxin production by Fusarium sporotrichioides. Appl. Environ. Microbiol. 2003, 69, 5935–5940. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.J.; Geiser, D.M.; Proctor, R.H.; Rooney, A.P.; O’Donnell, K.; Trail, F.; Gardiner, D.M.; Manners, M.; Kazan, K. Fusarium pathogenomics. Ann. Rev. Microbiol. 2013, 67, 399–416. [Google Scholar] [CrossRef] [PubMed]
- Proctor, R.H.; Hohn, T.M.; McCormick, S.P. Reduced virulence of Gibberella zeae caused by disruption of a trichothecene toxin biosynthetic gene. Mol. Plant Microbe Interact. 1995, 8, 593–601. [Google Scholar] [CrossRef] [PubMed]
- Bai, G.H.; Desjardins, A.E.; Plattner, R.D. Deoxynivalenol-nonproducing Fusarium graminearum causes initial infection but does not cause disease spread in wheat spikes. Mycopathologia 2002, 153, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Woloshuk, C.P.; Shim, W.B. Aflatoxins, fumonisins and trichothecenes: A convergence of knowledge. FEMS Microbiol. Rev. 2013, 37, 94–109. [Google Scholar] [CrossRef] [PubMed]
- Hogenhout, S.A.; van der Hoorn, R.A.L.; Terauchi, R.; Kamoun, S. Emerging concepts in effector biology of plant-associated organisms. Mol. Plant Microbe Interact. 2009, 22, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Idnurm, A.; Howlett, B. Pathogenicity genes of phytopathogenic fungi. Mol. Plant Pathol. 2001, 2, 241–255. [Google Scholar] [CrossRef] [PubMed]
- McCormick, S.P.; Alexander, N.J. Fusarium Tri8 encodes a trichothecene C-3 esterase. Appl. Environ. Microbiol. 2002, 68, 2959–2964. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, K.; Sarver, B.A.J.; Brandt, M.; Chang, D.C.; Noble-Wang, J.; Park, B.J.; Sutton, D.A.; Benjamin, L.; Lindsley, M.; Padhye, A.; et al. Phylogenetic diversity and microsphere array-based genotyping of human pathogenic fusaria, including from the multistate contact lens-associated U.S. keratitis outbreaks of 2005 and 2006. J. Clin. Microbiol. 2007, 45, 2235–2248. [Google Scholar] [PubMed]
- O’Donnell, K.; Sutton, D.A.; Rinaldi, M.G.; Sarver, B.A.; Balajee, S.A.; Schroers, H.J.; Summerbell, R.C.; Robert, V.A.; Crous, P.W.; Zhang, N.; et al. Internet-accessible DNA sequence database for identifying fusaria from human and animal infections. J. Clin. Microbiol. 2010, 48, 3708–3718. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The CLUSTALX windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucl. Acids Res. 1997, 24, 4876–4882. [Google Scholar] [CrossRef]
- Drummond, A.J.; Ashton, B.; Buxton, S.; Cheung, M.; Cooper, A.; Duran, C.; Field, M.; Heled, J.; Kearse, M.; Markowitz, S.; et al. Geneious v5.4. 2011. Available online: http://www.geneious.com/ (accessed on 10 January 2015).
- National Centre for Biotechnology Information. Available online: http://blast.ncbi.nlm.nih.gov/Blast.cgi (accessed on 10 January 2015).
- Shimodaira, H.; Hasegawa, M. Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Mol. Biol. Evol. 1999, 16, 1114–1116. [Google Scholar] [CrossRef]
- Swofford, D.L. PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods); Sinauer Associates: Sunderland, MA, USA, 2002; pp. 24–206. [Google Scholar]
- Posada, D. jModelTest: Phylogenetic model averaging. Mol. Biol. Evol. 2008, 25, 1253–1256. [Google Scholar] [CrossRef] [PubMed]
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef] [PubMed]
- Rambaut, A. FigTree. 2013. Available online: http://tree.bio.ed.ac.uk/software/figtree/ (accessed on 10 January 2015).
- Planet, P.J. Tree disagreement: Measuring and testing incongruence in phylogenies. J. Biomed. Inform. 2006, 39, 86–102. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rocha, L.O.; Laurence, M.H.; Proctor, R.H.; McCormick, S.P.; Summerell, B.A.; Liew, E.C.Y. Variation in Type A Trichothecene Production and Trichothecene Biosynthetic Genes in Fusarium goolgardi from Natural Ecosystems of Australia. Toxins 2015, 7, 4577-4594. https://doi.org/10.3390/toxins7114577
Rocha LO, Laurence MH, Proctor RH, McCormick SP, Summerell BA, Liew ECY. Variation in Type A Trichothecene Production and Trichothecene Biosynthetic Genes in Fusarium goolgardi from Natural Ecosystems of Australia. Toxins. 2015; 7(11):4577-4594. https://doi.org/10.3390/toxins7114577
Chicago/Turabian StyleRocha, Liliana O., Matthew H. Laurence, Robert H. Proctor, Susan P. McCormick, Brett A. Summerell, and Edward C. Y. Liew. 2015. "Variation in Type A Trichothecene Production and Trichothecene Biosynthetic Genes in Fusarium goolgardi from Natural Ecosystems of Australia" Toxins 7, no. 11: 4577-4594. https://doi.org/10.3390/toxins7114577
APA StyleRocha, L. O., Laurence, M. H., Proctor, R. H., McCormick, S. P., Summerell, B. A., & Liew, E. C. Y. (2015). Variation in Type A Trichothecene Production and Trichothecene Biosynthetic Genes in Fusarium goolgardi from Natural Ecosystems of Australia. Toxins, 7(11), 4577-4594. https://doi.org/10.3390/toxins7114577