The Cystine Knot Is Responsible for the Exceptional Stability of the Insecticidal Spider Toxin ω-Hexatoxin-Hv1a
Abstract
:1. Introduction
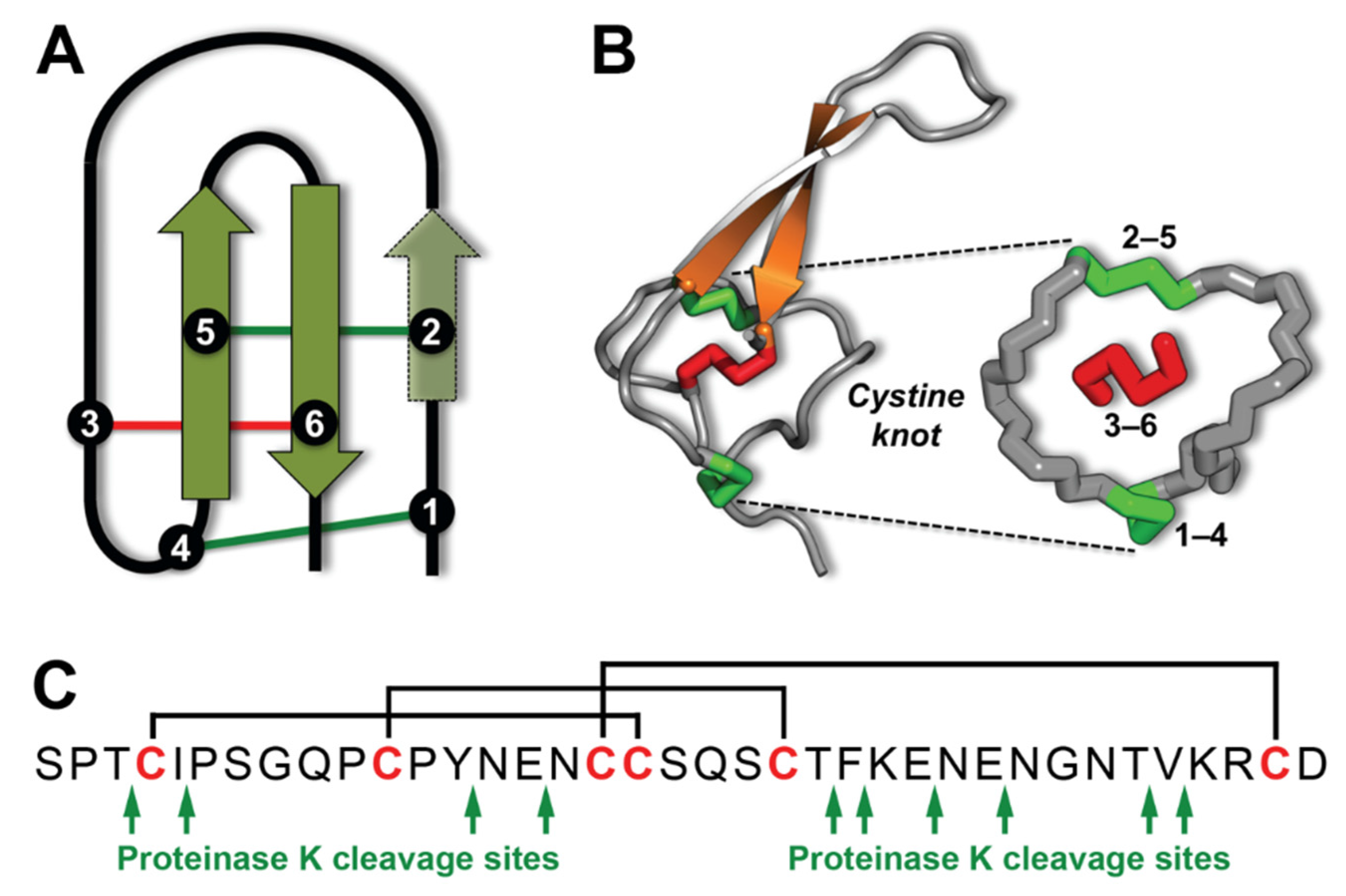
2. Results and Discussion
2.1. Thermal Stability
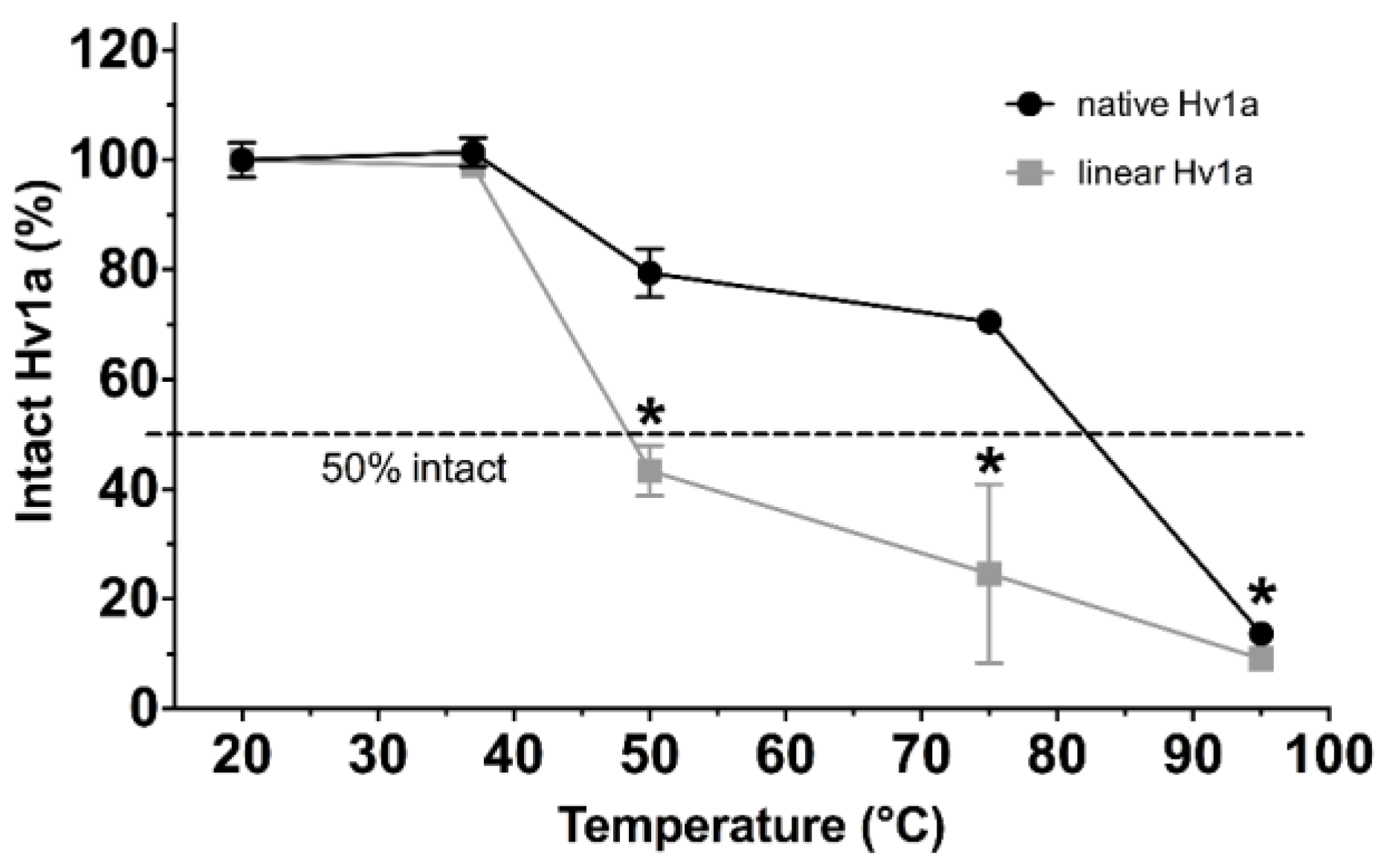
2.2. pH Stability
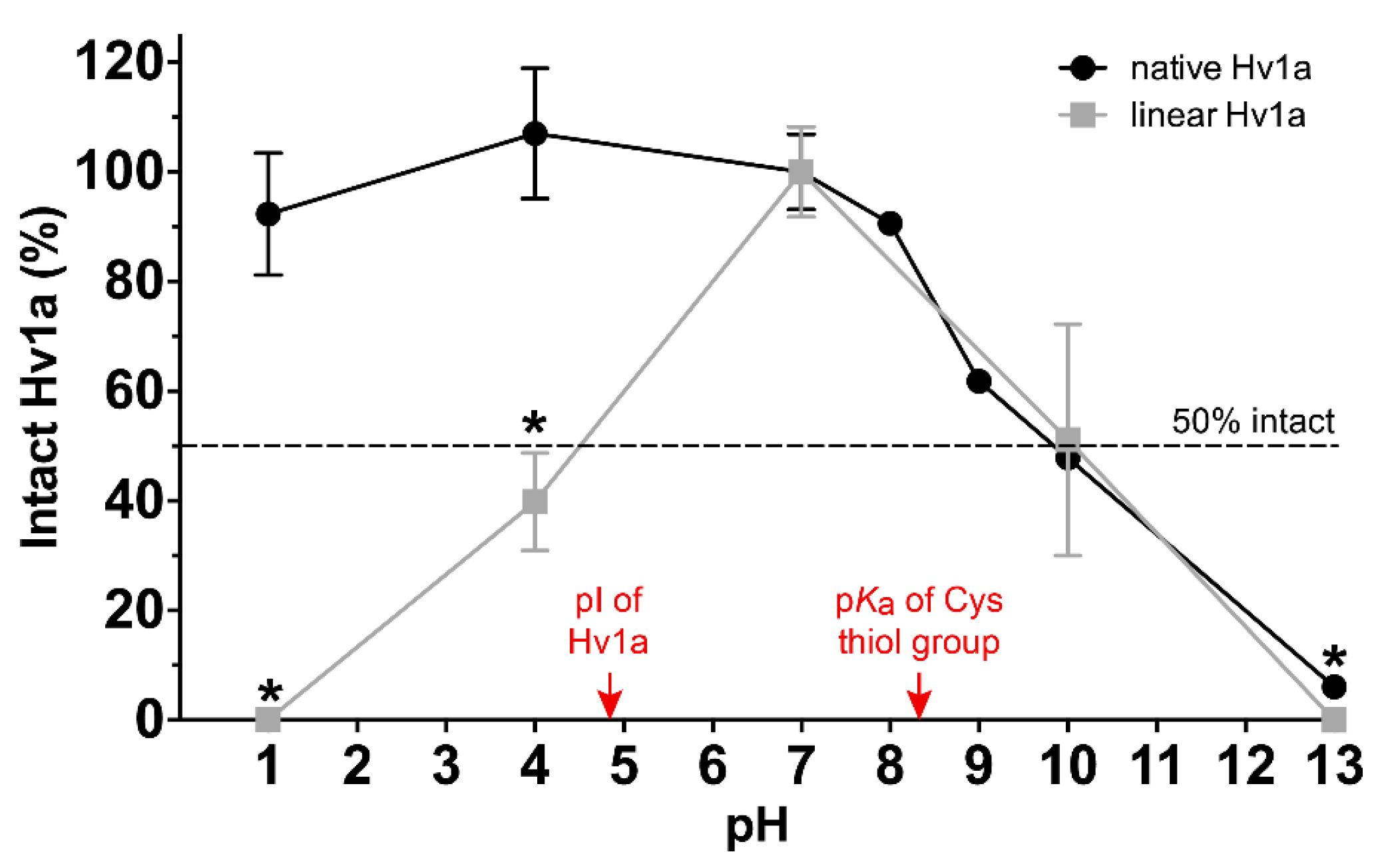
2.3. Chemical Stability
2.4. Proteolytic Stability
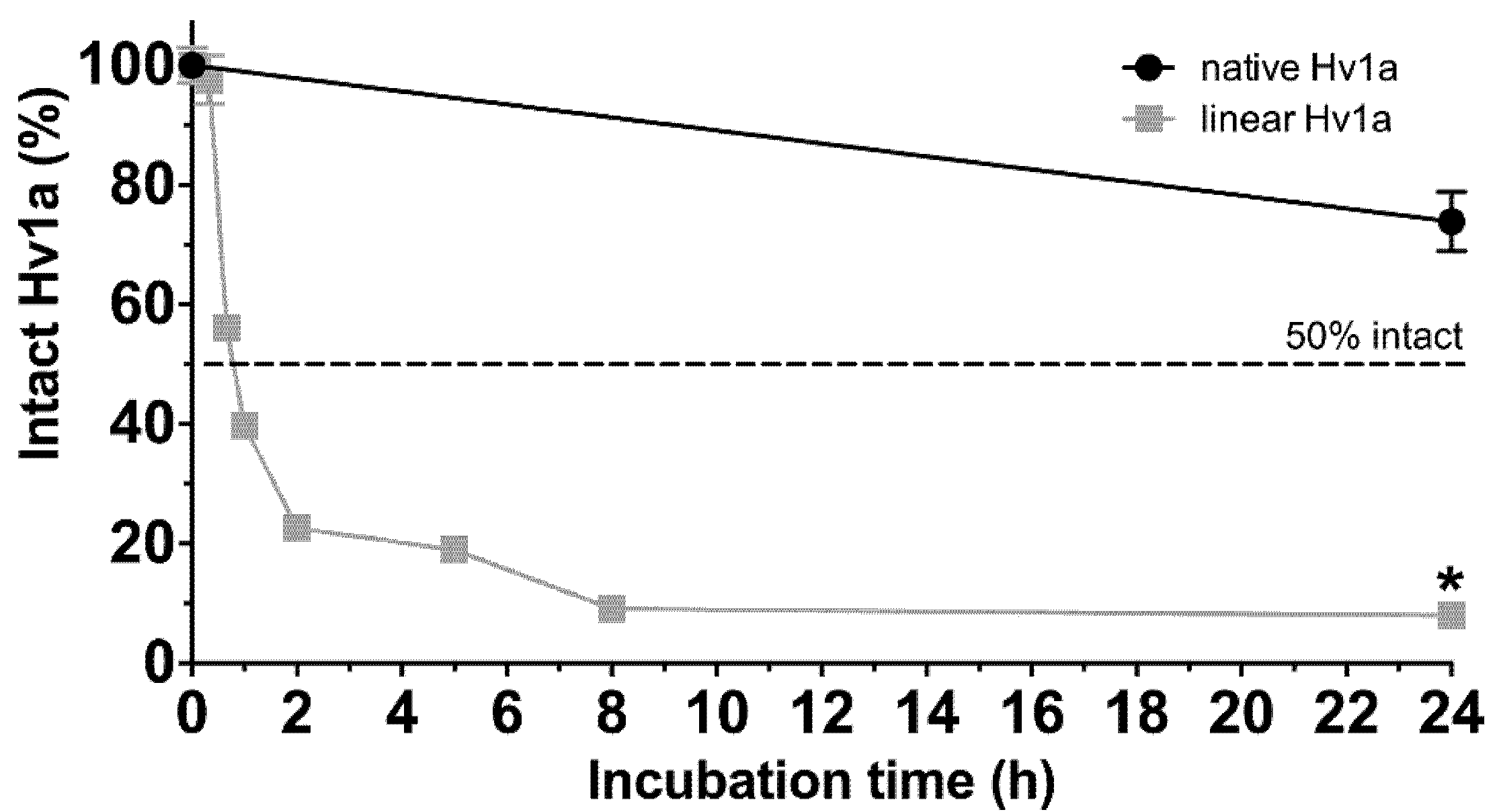
2.5. Stability in Insect Hemolymph

2.6. Stability in Human Plasma
2.7. Comparison of Hv1a Stability with Other ICK Peptides
3. Experimental Section
3.1. Chemicals
3.2. Sample Treatment
3.3. Thermal Stability
3.4. pH Stability
3.5. Chemical Stability
3.6. Proteolytic Stability
3.7. Stability in Insect Hemolymph
3.8. Stability in Human Plasma
3.9. Statistical Analyses
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Pallaghy, P.K.; Nielsen, K.J.; Craik, D.J.; Norton, R.S. A common structural motif incorporating a cystine knot and a triple-stranded β-sheet in toxic and inhibitory polypeptides. Protein Sci. 1994, 3, 1833–1839. [Google Scholar] [CrossRef] [PubMed]
- Norton, R.S.; Pallaghy, P.K. The cystine knot structure of ion channel toxins and related polypeptides. Toxicon 1998, 36, 1573–1583. [Google Scholar] [CrossRef]
- King, G.F.; Tedford, H.W.; Maggio, F. Structure and function of insecticidal neurotoxins from Australian funnel-web spiders. J. Toxicol.-Toxin Rev. 2002, 21, 359–389. [Google Scholar] [CrossRef]
- Saez, N.J.; Senff, S.; Jensen, J.E.; Er, S.Y.; Herzig, V.; Rash, L.D.; King, G.F. Spider-venom peptides as therapeutics. Toxins 2010, 2, 2851–2871. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, J.I.; Smith, R.; O’Donoghue, S.I.; Nilges, M.; Connor, M.; Howden, M.E.; Christie, M.J.; King, G.F. The structure of a novel insecticidal neurotoxin, ω-atracotoxin-HV1, from the venom of an Australian funnel web spider. Nat. Struct. Biol. 1997, 4, 559–566. [Google Scholar] [CrossRef] [PubMed]
- PeptideCutter. Available online: http://web.expasy.org/peptide_cutter (accessed on 23 October 2015).
- Craik, D.J.; Daly, N.L.; Waine, C. The cystine knot motif in toxins and implications for drug design. Toxicon 2001, 39, 43–60. [Google Scholar] [CrossRef]
- King, G.F.; Hardy, M.C. Spider-venom peptides: Structure, pharmacology, and potential for control of insect pests. Annu. Rev. Entomol. 2013, 58, 475–496. [Google Scholar] [CrossRef] [PubMed]
- Colgrave, M.L.; Craik, D.J. Thermal, chemical, and enzymatic stability of the cyclotide kalata B1: The importance of the cyclic cystine knot. Biochemistry 2004, 43, 5965–5975. [Google Scholar] [CrossRef] [PubMed]
- Tedford, H.W.; Gilles, N.; Ménez, A.; Doering, C.J.; Zamponi, G.W.; King, G.F. Scanning mutagenesis of ω-atracotoxin-Hv1a reveals a spatially restricted epitope that confers selective activity against invertebrate calcium channels. J. Biol. Chem. 2004, 279, 44133–44140. [Google Scholar] [CrossRef] [PubMed]
- Tedford, H.W.; Sollod, B.L.; Maggio, F.; King, G.F. Australian funnel-web spiders: Master insecticide chemists. Toxicon 2004, 43, 601–618. [Google Scholar] [CrossRef] [PubMed]
- Windley, M.J.; Herzig, V.; Dziemborowicz, S.A.; Hardy, M.C.; King, G.F.; Nicholson, G.M. Spider-venom peptides as bioinsecticides. Toxins 2012, 4, 191–227. [Google Scholar] [CrossRef] [PubMed]
- Nakasu, E.Y.; Williamson, S.M.; Edwards, M.G.; Fitches, E.C.; Gatehouse, J.A.; Wright, G.A.; Gatehouse, A.M. Novel biopesticide based on a spider venom peptide shows no adverse effects on honeybees. Proc. R. Soc. B. 2014, 281, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Bonning, B.C.; Pal, N.; Liu, S.; Wang, Z.; Sivakumar, S.; Dixon, P.M.; King, G.F.; Miller, W.A. Toxin delivery by the coat protein of an aphid-vectored plant virus provides plant resistance to aphids. Nat. Biotechnol. 2014, 32, 102–105. [Google Scholar] [CrossRef] [PubMed]
- Bloomquist, J.R. Mode of action of atracotoxin at central and peripheral synapses of insects. Invertebr. Neurosci. 2003, 5, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Manning, M.C.; Chou, D.K.; Murphy, B.M.; Payne, R.W.; Katayama, D.S. Stability of protein pharmaceuticals: An update. Pharm. Res. 2010, 27, 544–575. [Google Scholar] [CrossRef] [PubMed]
- Furman, J.L.; Chiu, M.; Hunter, M.J. Early engineering approaches to improve peptide developability and manufacturability. AAPS J. 2015, 17, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Robinson, N.E. Protein deamidation. Proc. Natl. Acad. Sci. USA 2002, 99, 5283–5288. [Google Scholar] [CrossRef] [PubMed]
- Harrison, J.F. Insect acid-base physiology. Annu. Rev. Entomol. 2001, 46, 221–250. [Google Scholar] [CrossRef] [PubMed]
- Nation, J.L. Insect Physiology and Biochemistry, 2nd ed.; CRC Press: Boca Raton, FL, USA, 2008. [Google Scholar]
- Berenbaum, M.R. Adaptive significance of midgut pH in larval Lepidoptera. Am. Nat. 1980, 115, 138–146. [Google Scholar] [CrossRef]
- Pentzold, S.; Zagrobelny, M.; Roelsgaard, P.S.; Moller, B.L.; Bak, S. The multiple strategies of an insect herbivore to overcome plant cyanogenic glucoside defence. PLoS One 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.A.; Zafar, Y.; Briddon, R.W.; Malik, K.A.; Mukhtar, Z. Spider venom toxin protects plants from insect attack. Transgenic Res. 2006, 15, 349–357. [Google Scholar] [CrossRef] [PubMed]
- Shah, A.D.; Ahmed, M.; Mukhtar, Z.; Khan, S.A.; Habib, I.; Malik, Z.A.; Mansoor, S.; Saeed, N.A. Spider toxin (Hvt) gene cloned under phloem specific RSs1 and RolC promoters provides resistance against American bollworm (Heliothis armigera). Biotechnol. Lett. 2011, 33, 1457–1463. [Google Scholar] [CrossRef] [PubMed]
- Omar, A.; Ali Chatha, K. National Institute for Biotechnology and Genetic Engineering (NIBGE): Genetically modified spider cotton. Asian J. Manag. Cases 2012, 9, 33–58. [Google Scholar] [CrossRef]
- Foy, C.L.; Pritchard, D.W. Pesticide Formulation and Adjuvant Technology; CRC Press: Boca Raton, FL, USA, 1996. [Google Scholar]
- Strauss, W.M. UNIT 2.2 Preparation of genomic DNA from mammalian tissue. Curr. Protoc. Mol. Biol. 2001. [Google Scholar] [CrossRef]
- Chapman, R.F. The Insects: Structure and Function, 5th ed.; Cambridge University Press: Cambridge, UK, 1998. [Google Scholar]
- Kanost, M.R. Hemolymph. In Encylopedia of Insects, 2nd ed.; Resh, V.H., Cardé, R.T., Eds.; Academic Press: Burlington, MA, USA, 2009; pp. 446–448. [Google Scholar]
- Brown, S.E.; Patton, R.L.; Zerillo, R.T.; Douglas, S.M.; Breillatt, J.P.; Mazzone, H.M. Comparative properties of hemolymph of the Gypsy moth and the European pine sawfly. J. N.Y. Entomol. Soc. 1977, 85, 36–42. [Google Scholar]
- Wang, Z.; Haunerland, N.H. Storage protein uptake in Helicoverpa zea. purification of the very high density lipoprotein receptor from perivisceral fat body. J. Biol. Chem. 1993, 268, 16673–16678. [Google Scholar] [PubMed]
- Hardy, M.C.; Daly, N.L.; Mobli, M.; Morales, R.A.; King, G.F. Isolation of an orally active insecticidal toxin from the venom of an Australian tarantula. PLoS One 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Armishaw, C.J.; Daly, N.L.; Nevin, S.T.; Adams, D.J.; Craik, D.J.; Alewood, P.F. α-Selenoconotoxins, a new class of potent α7 neuronal nicotinic receptor antagonists. J. Biol. Chem. 2006, 281, 14136–14143. [Google Scholar] [CrossRef] [PubMed]
- Gracy, J.; Le-Nguyen, D.; Gelly, J.C.; Kaas, Q.; Heitz, A.; Chiche, L. KNOTTIN: The knottin or inhibitor cystine knot scaffold in 2007. Nucleic Acids Res. 2008, 36, D314–D319. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Su, M.; Hamann, M.T.; Bowling, J.J.; Kim, H.S.; Jung, J.H. Solution structure of a sponge-derived cystine knot peptide and its notable stability. J. Nat. Prod. 2014, 77, 304–310. [Google Scholar] [CrossRef] [PubMed]
- Fujitani, N.; Kouno, T.; Nakahara, T.; Takaya, K.; Osaki, T.; Kawabata, S.; Mizuguchi, M.; Aizawa, T.; Demura, M.; Nishimura, S.; et al. The solution structure of horseshoe crab antimicrobial peptide tachystatin B with an inhibitory cystine-knot motif. J. Pept. Sci. 2007, 13, 269–279. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, A.A.; Salceda, E.; Garateix, A.G.; Zaharenko, A.J.; Peigneur, S.; López, O.; Pons, T.; Richardson, M.; Díaz, M.; Hernández, Y.; et al. A novel sea anemone peptide that inhibits acid-sensing ion channels. Peptides 2014, 53, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Olivera, B.M.; Hillyard, D.R.; Marsh, M.; Yoshikami, D. Combinatorial peptide libraries in drug design: Lessons from venomous cone snails. Trends Biotechnol. 1995, 13, 422–426. [Google Scholar] [CrossRef]
- Sollod, B.L.; Wilson, D.; Zhaxybayeva, O.; Gogarten, J.P.; Drinkwater, R.; King, G.F. Were arachnids the first to use combinatorial peptide libraries? Peptides 2005, 26, 131–139. [Google Scholar] [CrossRef] [PubMed]
- Ackerman, S.E.; Currier, N.V.; Bergen, J.M.; Cochran, J.R. Cystine-knot peptides: Emerging tools for cancer imaging and therapy. Expert Rev. Proteomics 2014, 11, 561–572. [Google Scholar] [CrossRef] [PubMed]
- Werle, M.; Schmitz, T.; Huang, H.L.; Wentzel, A.; Kolmar, H.; Bernkop-Schnürch, A. The potential of cystine-knot microproteins as novel pharmacophoric scaffolds in oral peptide drug delivery. J. Drug Target 2006, 14, 137–146. [Google Scholar] [CrossRef] [PubMed]
- Werle, M.; Kafedjiiski, K.; Kolmar, H.; Bernkop-Schnürch, A. Evaluation and improvement of the properties of the novel cystine-knot microprotein McoEeTI for oral administration. Int. J. Pharm. 2007, 332, 72–79. [Google Scholar] [CrossRef] [PubMed]
- Wiener, S. Observations on the venom of the Sydney funnel-web spider (Atrax robustus). Med. J. Aust. 1961, 48, 693–699. [Google Scholar] [PubMed]
- Sheumack, D.D.; Claassens, R.; Howden, M.E.H.; Whitley, N.M. Complete amino acid sequence of of a new type of lethal neurotoxin from the venom of the funnel-web spider Atrax robustus. FEBS Lett. 1985, 181, 154–156. [Google Scholar] [CrossRef]
- Fletcher, J.I.; Chapman, B.E.; Mackay, J.P.; Howden, M.E.H.; King, G.F. The structure of versutoxin (δ-atracotoxin-Hv1) provides insights into the binding of site 3 neurotoxins to the voltage-gated sodium channel. Structure 1997, 5, 1525–1535. [Google Scholar] [CrossRef]
- Pallaghy, P.K.; Alewood, D.; Alewood, P.F.; Norton, R.S. Solution structure of robustoxin, the lethal neurotoxin from the funnel-web spider Atrax robustus. FEBS Lett. 1997, 419, 191–196. [Google Scholar] [CrossRef]
- Nicholson, G.M.; Little, M.J.; Tyler, M.; Narahashi, T. Selective alteration of sodium channel gating by Australian funnel-web spider toxins. Toxicon 1996, 34, 1443–1453. [Google Scholar] [CrossRef]
- Hale, J.E.; Butler, J.P.; Gelfanova, V.; You, J.S.; Knierman, M.D. A simplified procedure for the reduction and alkylation of cysteine residues in proteins prior to proteolytic digestion and mass spectral analysis. Anal. Biochem. 2004, 333, 174–181. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Herzig, V.; King, G.F. The Cystine Knot Is Responsible for the Exceptional Stability of the Insecticidal Spider Toxin ω-Hexatoxin-Hv1a. Toxins 2015, 7, 4366-4380. https://doi.org/10.3390/toxins7104366
Herzig V, King GF. The Cystine Knot Is Responsible for the Exceptional Stability of the Insecticidal Spider Toxin ω-Hexatoxin-Hv1a. Toxins. 2015; 7(10):4366-4380. https://doi.org/10.3390/toxins7104366
Chicago/Turabian StyleHerzig, Volker, and Glenn F. King. 2015. "The Cystine Knot Is Responsible for the Exceptional Stability of the Insecticidal Spider Toxin ω-Hexatoxin-Hv1a" Toxins 7, no. 10: 4366-4380. https://doi.org/10.3390/toxins7104366
APA StyleHerzig, V., & King, G. F. (2015). The Cystine Knot Is Responsible for the Exceptional Stability of the Insecticidal Spider Toxin ω-Hexatoxin-Hv1a. Toxins, 7(10), 4366-4380. https://doi.org/10.3390/toxins7104366






