Fossilized Venom: The Unusually Conserved Venom Profiles of Heloderma Species (Beaded Lizards and Gila Monsters)
Abstract
:1. Introduction
| Protein Type/ Toxin Class | Toxic Action | Uniprot Accession #(s) |
|---|---|---|
| CRiSP (cysteine rich secretory protein) | Paralysis of peripheral smooth muscle and induction of hypothermia through blockage of various channels including ryanodine and L-type calcium channels. | Q91055 |
| Exendin | Induces hypotension via relaxation of cardiac smooth muscle. | C6EVG1, C6EVG2, P04203, P04204, P20394, P26349 |
| Helofensin | Lethal toxin that inhibits direct electrical stimulation of the isolated hemi-diaphragm. | C6EVG6, D2X5W3, D2X5W4, Q7LZ31 |
| Kallikrein | Increase of vascular permeability, production of hypotension, stimulation of inflammation in addition to cleavage of fibrinogen. | P43685, C6EVG4, C6EVG5 |
| B-type Natriuretic peptide/helokinestatin precursor | Natriuretic peptides produce hypotension through the relaxation of aortic smooth muscle. The helokinestatin peptides are antagonists of bradykinin at the B2 bradykinin receptor. | C6EVG7, D7FB56, D7FB57, E8ZCG5 |
| Phospholipase A2 (Type III) | Inhibition of platelet aggregation via the epinephrine-induced pathway. | C6EVG9, C6EVH0 |
2. Results and Discussion

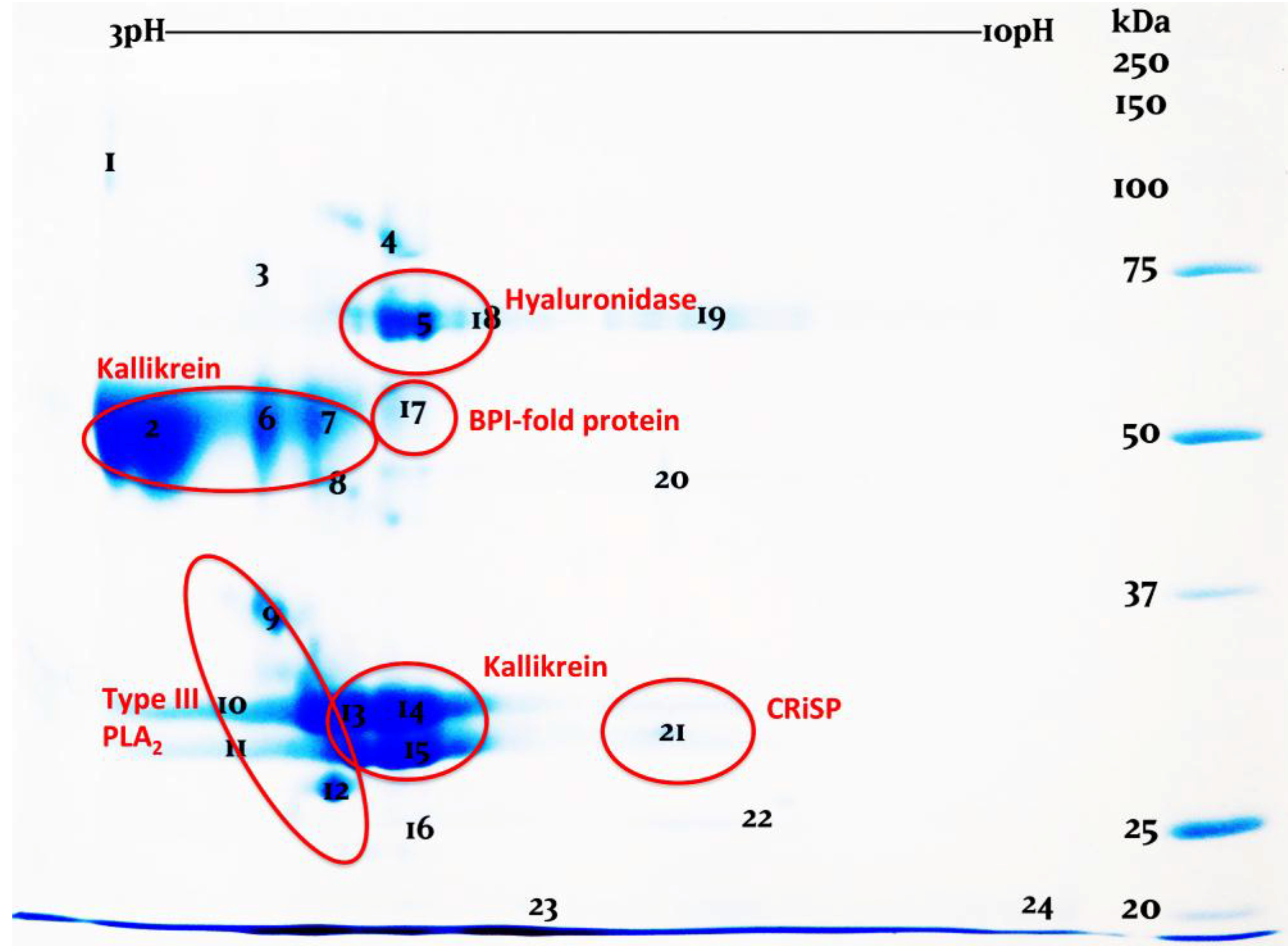

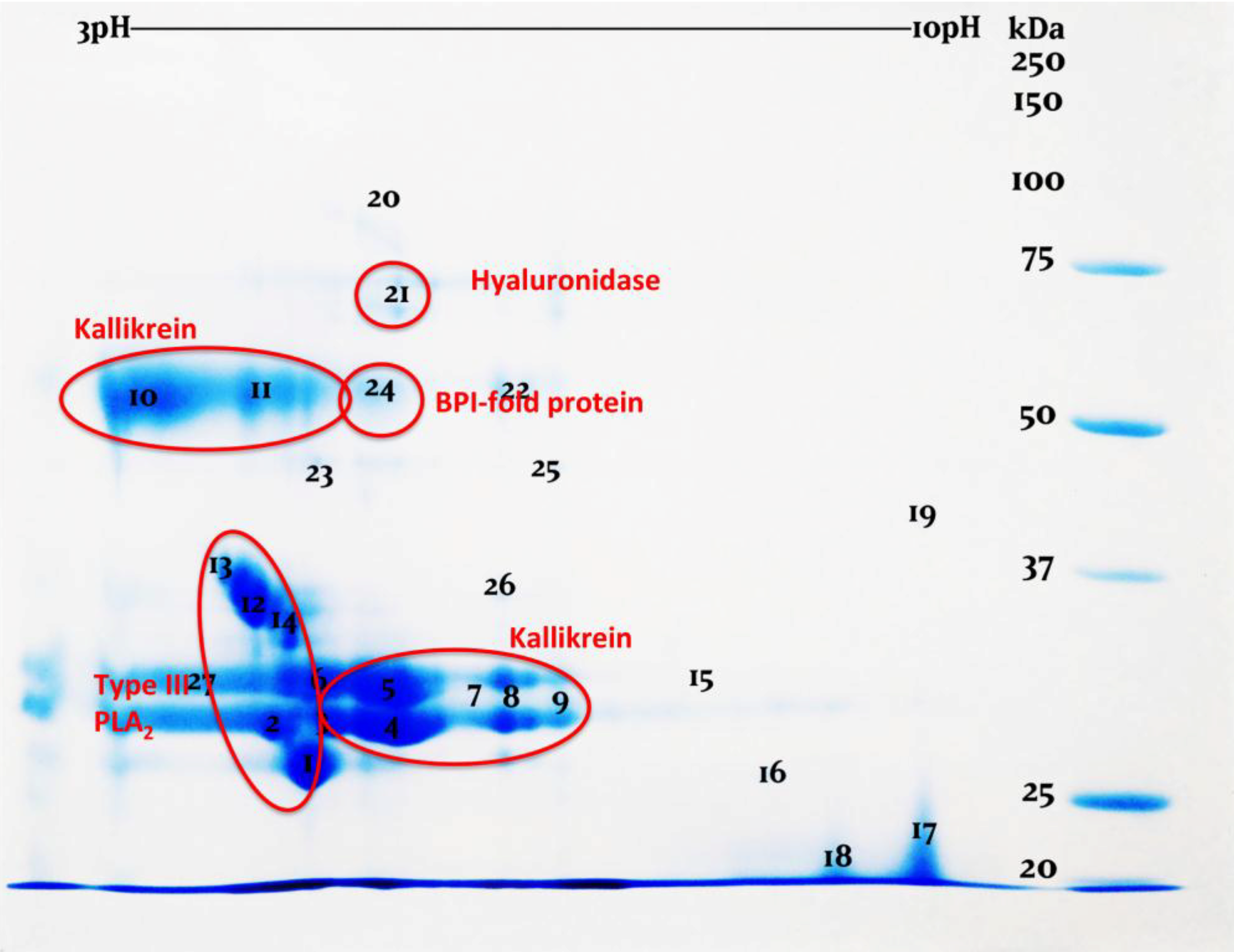
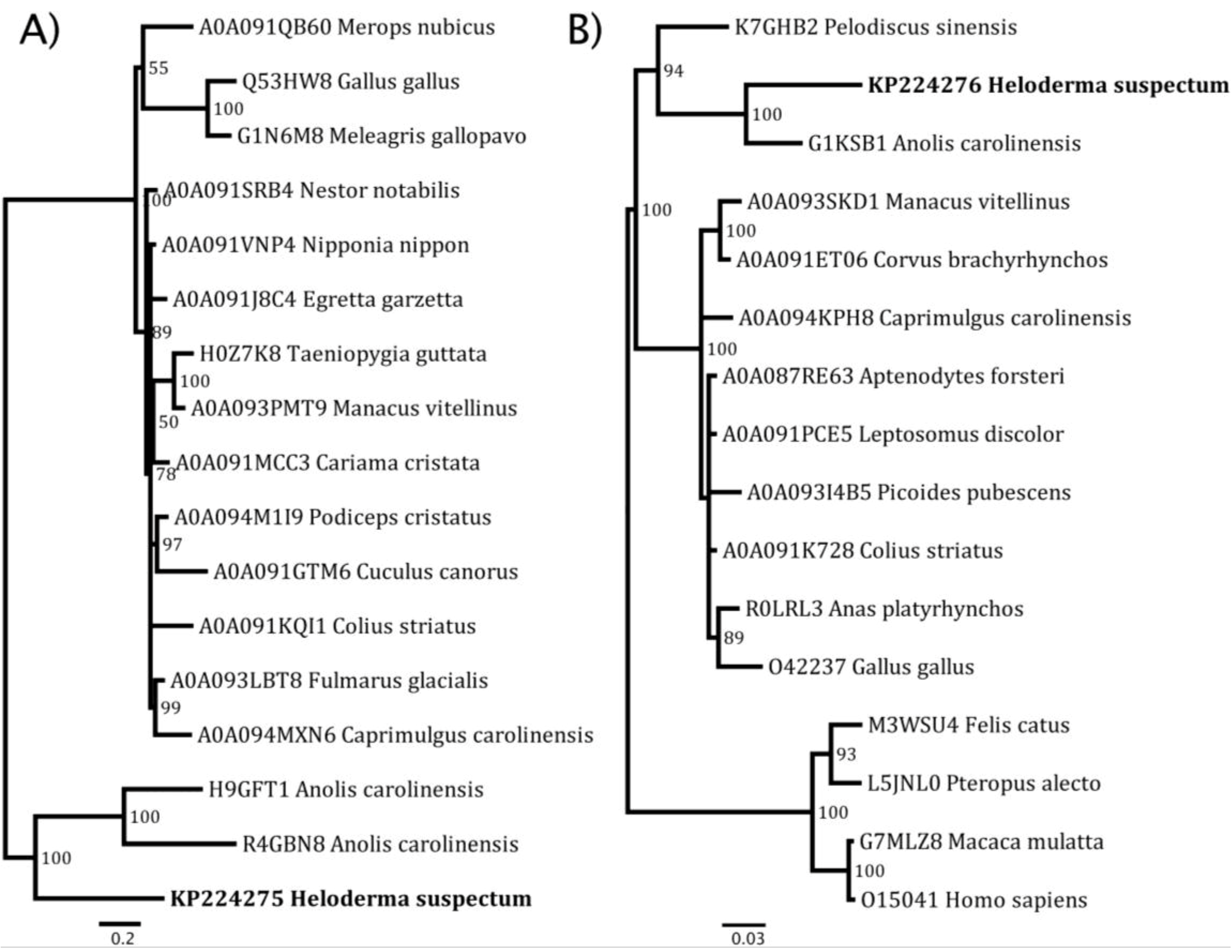

| Toxin Type | Sequence pairs | Estimates |
|---|---|---|
| Kallikrein | EU790962.1 (H. suspectum) vs. HM437246.1 (H. horridum) | dN: 0.180; dS: 0.225; dN/dS: 0.80 |
| EU790963.1 (H. suspectum) vs. HM437246.1 (H. horridum) | dN: 0.242; dS: 0.450; dN/dS: 0.53 | |
| EU790962.1 (H. suspectum) vs. EU790963.1 (H. suspectum) | dN: 0.081; dS: 0.173; dN/dS: 0.47 | |
| Average | dN: 0.167; dS: 0.282; dN/dS:  | |
| CRiSP | EU790958.1 (H. suspectum) vs. U13619.1 (H. horridum) | dN: 0.011; dS: 0.022; dN/dS:  |
| Helofensin | GQ918270.1 (H. suspectum) vs. EU790964.1 (H. suspectum) | dN: 0.030; dS: 0.036; dN/dS: 0.84 |
| GQ918271.1 (H. suspectum) vs. EU790964.1 (H. suspectum) | dN: 0.052; dS: 0.065; dN/dS: 0.80 | |
| GQ918271.1 (H. suspectum) vs. GQ918270.1 (H. suspectum) | dN: 0.020; dS: 0.027; dN/dS: 0.74 | |
| Average | dN: 0.034; dS: 0.042; dN/dS:  |
3. Experimental Section
3.1. Venom Collection
3.2. Shotgun Sequencing
3.3. One-Dimensional Gel Electrophoresis
3.4. Two-Dimensional Gel Electrophoresis
3.5. LC-MS/MS
3.6. Pairwise-Estimation of dN/dS
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Douglas, M.E.; Douglas, M.R.; Schuett, G.W.; Beck, D.D.; Sullivan, B.K. Conservation phylogenetics of helodermatid lizards using multiple molecular markers and a supertree approach. Mol. Phylogenet. Evol. 2010, 55, 153–167. [Google Scholar] [PubMed]
- Reiserer, R.S.; Schuett, G.W.; Beck, D.D. Taxonomic reassessment and conservation status of the beaded lizard, Heloderma horridum (Squamata: Helodermatidae). Amphib. Reptile Conserv. 2013, 7, 74–96. [Google Scholar]
- Pianka, E.R.; King, D.R. Varanoid Lizards of the World; Pianka, E., King, D., Eds.; Indiana University Press: Bloomington, IN, USA, 2004. [Google Scholar]
- Beck, D.D. Biology of Gila Monsters and Beaded Lizards; University of California Press: Oakland, CA, USA, 2005; p. 212. [Google Scholar]
- Venom Evolution Lab, School of Biological Sciences, the University of Queensland, St. Lucia, Queensland. Private observation. 2014.
- Bogert, C.M.; del Campo, R.M. The gila monster and its allies. The relationships, habits, and behavior of the lizards of the family Helodermatidae. Bull. Am. Mus. Natl. Hist. 1956, 109, 1–238. [Google Scholar]
- Bouabboud, C.F.; Kardassakis, D.G. Acute myocardial-infarction following a gila monster (Heloderma suspectum cinctum) bite. West. J. Med. 1988, 148, 577–579. [Google Scholar]
- Cantrell, F.L. Envenomation by the Mexican beaded lizard: A case report. J. Toxicol.-Clin. Toxicol. 2003, 41, 241–244. [Google Scholar] [CrossRef] [PubMed]
- Hooker, K.R.; Caravati, E.M. Gila monster envenomation. Ann. Emerg. Med. 1994, 24, 731–735. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.F. Gila monster envenomation. Ann. Emerg. Med. 1995, 25, 720. [Google Scholar] [PubMed]
- Strimple, P.D.; Tomassoni, A.J.; Otten, E.J.; Bahner, D. Report on envenomation by a Gila monster (Heloderma suspectum) with a discussion of venom apparatus, clinical findings, and treatment. Wilderness Environ. Med. 1997, 8, 111–116. [Google Scholar] [CrossRef] [PubMed]
- Drucker, D.J.; Nauck, M.A. The incretin system: Glucagon-like peptide-1 receptor agonists and dipeptidyl peptidase-4 inhibitors in type 2 diabetes. Lancet 2006, 368, 1696–1705. [Google Scholar] [CrossRef] [PubMed]
- Angulo, Y.; Escolano, J.; Lomonte, B.; Gutierrez, J.M.; Sanz, L.; Calvete, J.J. Snake venomics of Central American pitvipers: Clues for rationalizing the distinct envenomation profiles of Atropoides nummifer and Atropoides picadoi. J. Proteome Res. 2008, 7, 708–719. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Juarez, P.; Sanz, L. Snake venomics. Strategy and applications. J. Mass Spectrom. 2007, 42, 1405–1414. [Google Scholar] [CrossRef]
- Fry, B.G.; Wickramaratana, J.C.; Lemme, S.; Beuve, A.; Garbers, D.; Hodgson, W.C.; Alewood, P. Novel natriuretic peptides from the venom of the inland taipan (Oxyuranus microlepidotus): Isolation, chemical and biological characterisation. Biochem. Biophys. Res. Commun. 2005, 327, 1011–1015. [Google Scholar] [CrossRef]
- Fry, B.G.; Wickramaratna, J.C.; Hodgson, W.C.; Alewood, P.F.; Kini, R.M.; Ho, H.; Wuster, W. Electrospray liquid chromatography/mass spectrometry fingerprinting of Acanthophis (death adder) venoms: Taxonomic and toxinological implications. Rapid. Commun. Mass Spectrom. 2002, 16, 600–608. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G.; Wuster, W.; Ramjan, S.F.R.; Jackson, T.; Martelli, P.; Kini, R.M. Analysis of Colubroidea snake venoms by liquid chromatography with mass spectrometry: Evolutionary and toxinological implications. Rapid Commun. Mass Spectrom. 2003, 17, 2047–2062. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, J.M.; Sanz, L.; Escolano, J.; Fernandez, J.; Lomonte, B.; Angulo, Y.; Rucavado, A.; Warrell, D.A.; Calvete, J.J. Snake venomics of the Lesser Antillean pit vipers Bothrops caribbaeus and Bothrops lanceolatus: Correlation with toxicological activities and immunoreactivity of a heterologous antivenom. J. Proteome Res. 2008, 7, 4396–4408. [Google Scholar] [CrossRef] [PubMed]
- Lomonte, B.; Escolano, J.; Fernandez, J.; Sanz, L.; Angulo, Y.; Gutierrez, J.M.; Calvete, J.J. Snake venomics and antivenomics of the arboreal neotropical pitvipers Bothriechis lateralis and Bothriechis schlegelii. J. Proteome Res. 2008, 7, 2445–2457. [Google Scholar] [CrossRef] [PubMed]
- Mackessy, S.P. Evolutionary trends in venom composition in the western rattlesnakes (Crotalus viridis sensu lato): Toxicity vs. tenderizers. Toxicon 2010, 55, 1463–1474. [Google Scholar] [CrossRef]
- Salazar, A.M.; Guerrero, B.; Cantu, B.; Cantu, E.; Rodríguez-Acosta, A.; Pérez, J.C.; Galán, J.A.; Tao, A.; Sánchez, E.E. Venom variation in hemostasis of the southern Pacific rattlesnake (Crotalus oreganus helleri): Isolation of hellerase. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2009, 149, 307–316. [Google Scholar] [CrossRef]
- Sanz, L.; Escolano, J.; Ferretti, M.; Biscoglio, M.J.; Rivera, E.; Crescenti, E.J.; Angulo, Y.; Lomonte, B.; Gutierrez, J.M.; Calvete, J.J. Snake venomics of the South and Central American Bushmasters. Comparison of the toxin composition of Lachesis muta gathered from proteomic versus transcriptomic analysis. J. Proteomics 2008, 71, 46–60. [Google Scholar]
- Sanz, L.; Gibbs, H.L.; Mackessy, S.P.; Calvete, J.J. Venom proteomes of closely related Sistrurus rattlesnakes with divergent diets. J. Proteome Res. 2006, 5, 2098–2112. [Google Scholar] [CrossRef] [PubMed]
- Tashima, A.K.; Sanz, L.; Camargo, A.C.; Serrano, S.M.; Calvete, J.J. Snake venomics of the Brazilian pitvipers Bothrops cotiara and Bothrops fonsecai Identification of taxonomy markers. J. Proteomics 2008, 71, 473–485. [Google Scholar]
- Wagstaff, S.C.; Sanz, L.; Juarez, P.; Harrison, R.A.; Calvete, J.J. Combined snake venomics and venom gland transcriptomic analysis of the ocellated carpet viper, Echis ocellatus. J. Proteomics 2009, 71, 609–623. [Google Scholar] [CrossRef] [PubMed]
- Boldrini-Franca, J.; Correa-Netto, C.; Silva, M.M.; Rodrigues, R.S.; de la Torre, P.; Perez, A.; Soares, A.M.; Zingali, R.B.; Nogueira, R.A.; Rodrigues, V.M.; et al. Snake venomics and antivenomics of Crotalus durissus subspecies from Brazil: Assessment of geographic variation and its implication on snakebite management. J. Proteomics 2010, 73, 1758–1776. [Google Scholar] [CrossRef] [PubMed]
- Castro, E.N.; Lomonte, B.; del Carmen Gutiérrez, M.; Alagón, A.; Gutiérrez, J.M. Intraspecies variation in the venom of the rattlesnake Crotalus simus from Mexico: Different expression of crotoxin results in highly variable toxicity in the venoms of three subspecies. J. Proteomics 2013, 87, 103–121. [Google Scholar] [CrossRef] [PubMed]
- Daltry, J.C.; Wuster, W.; Thorpe, R.S. Diet and snake venom evolution. Nature 1996, 379, 537–540. [Google Scholar] [CrossRef] [PubMed]
- Forstner, M.; Hilsenbeck, R.; Scudday, J. Geographic variation in whole venom profiles from the mottled rock rattlesnake (Crotalus lepidus lepidus) in Texas. J. Herpetol. 1997, 31, 277–287. [Google Scholar] [CrossRef]
- French, W.J.; Hayes, W.K.; Bush, S.P.; Cardwell, M.D.; Bader, J.O.; Rael, E.D. Mojave toxin in venom of Crotalus helleri (Southern Pacific Rattlesnake): Molecular and geographic characterization. Toxicon 2004, 44, 781–791. [Google Scholar] [CrossRef] [PubMed]
- Sunagar, K.; Undheim, E.A.; Scheib, H.; Gren, E.C.; Cochran, C.; Person, C.E.; Koludarov, I.; Kelln, W.; Hayes, W.K.; King, G.F.; et al. Intraspecific venom variation in the medically significant Southern Pacific Rattlesnake (Crotalus oreganushelleri): Biodiscovery, clinical and evolutionary implications. J. Proteomics 2014, 99, 68–83. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Fasoli, E.; Sanz, L.; Boschetti, E.; Righetti, P.G. Exploring the venom proteome of the western diamondback rattlesnake, Crotalus atrox, via snake venomics and combinatorial peptide ligand library approaches. J. Proteome Res. 2009, 8, 3055–3067. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Lozano, J.L.; de Sousa, M.V.; Ricart, C.A.; Chavez-Olortegui, C.; Flores Sanchez, E.; Muniz, E.G.; Buhrnheim, P.F.; Morhy, L. Ontogenetic variation of metalloproteinases and plasma coagulant activity in venoms of wild Bothrops atrox specimens from Amazonian rain forest. Toxicon 2002, 40, 997–1006. [Google Scholar] [CrossRef] [PubMed]
- Mackessy, S.P. Venom ontogeny in the Pacific rattlesnakes Crotalus viridis helleri and C. v. oreganus. Copeia 1988, 1988, 92–101. [Google Scholar] [CrossRef]
- Brust, A.; Sunagar, K.; Undheim, E.A.; Vetter, I.; Yang, D.C.; Casewell, N.R.; Jackson, T.N.; Koludarov, I.; Alewood, P.F.; Hodgson, W.C.; et al. Differential evolution and neofunctionalization of snake venom metalloprotease domains. Mol. Cell. Proteomics 2013, 12, 651–663. [Google Scholar] [CrossRef] [PubMed]
- Casewell, N.R.; Wuster, W.; Vonk, F.J.; Harrison, R.A.; Fry, B.G. Complex cocktails: The evolutionary novelty of venoms. Trends Ecol. Evol. 2013, 28, 219–229. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G.; Scheib, H.; van der Weerd, L.; Young, B.; McNaughtan, J.; Ramjan, S.F.; Vidal, N.; Poelmann, R.E.; Norman, J.A. Evolution of an arsenal: Structural and functional diversification of the venom system in the advanced snakes (Caenophidia). Mol. Cell. Proteomics 2008, 7, 215–246. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G.; Wuster, W.; Kini, R.M.; Brusic, V.; Khan, A.; Venkataraman, D.; Rooney, A.P. Molecular evolution and phylogeny of elapid snake venom three-finger toxins. J. Mol. Evol. 2003, 57, 110–129. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, H.L.; Mackessy, S.P. Functional basis of a molecular adaptation: Prey-specific toxic effects of venom from Sistrurus rattlesnakes. Toxicon 2009, 53, 672–679. [Google Scholar] [CrossRef] [PubMed]
- Pawlak, J.; Mackessy, S.P.; Sixberry, N.M.; Stura, E.A.; le Du, M.H.; Menez, R.; Foo, C.S.; Menez, A.; Nirthanan, S.; Kini, R.M. Irditoxin, a novel covalently linked heterodimeric three-finger toxin with high taxon-specific neurotoxicity. FASEB J. 2009, 23, 534–545. [Google Scholar] [CrossRef]
- Sunagar, K.; Johnson, W.E.; O’Brien, S.J.; Vasconcelos, V.; Antunes, A. Evolution of CRISPs associated with toxicoferan-reptilian venom and mammalian reproduction. Mol. Biol. Evol. 2012, 29, 1807–1822. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.A.; Jackson, T.N.; Casewell, N.R.; Low, D.H.; Rossi, S.; Baumann, K.; Fathinia, B.; Visser, J.; Nouwens, A.; Hendrikx, I.; Jones, A.; Fry, B.G. Extreme venom variation in Middle Eastern vipers: A proteomics comparison of Eristicophis macmahonii, Pseudocerastes fieldi and Pseudocerastes persicus. J. Proteomics 2014. [Google Scholar] [CrossRef]
- Fry, B.G.; Roelants, K.; Winter, K.; Hodgson, W.C.; Griesman, L.; Kwok, H.F.; Scanlon, D.; Karas, J.; Shaw, C.; Wong, L.; et al. Novel venom proteins produced by differential domain-expression strategies in beaded lizards and gila monsters (genus Heloderma). Mol. Biol. Evol. 2010, 27, 395–407. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G.; Winter, K.; Norman, J.A.; Roelants, K.; Nabuurs, R.J.; van Osch, M.J.; Teeuwisse, W.M.; van der Weerd, L.; McNaughtan, J.E.; Kwok, H.F.; et al. Functional and structural diversification of the Anguimorpha lizard venom system. Mol. Cell. Proteomics 2010, 9, 2369–2390. [Google Scholar] [CrossRef] [PubMed]
- Beck, D.D. Ecology and behavior of the gila monster in southwestern Utah. J. Herpetol. 1990, 24, 54–68. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Koludarov, I.; Jackson, T.N.W.; Sunagar, K.; Nouwens, A.; Hendrikx, I.; Fry, B.G. Fossilized Venom: The Unusually Conserved Venom Profiles of Heloderma Species (Beaded Lizards and Gila Monsters). Toxins 2014, 6, 3582-3595. https://doi.org/10.3390/toxins6123582
Koludarov I, Jackson TNW, Sunagar K, Nouwens A, Hendrikx I, Fry BG. Fossilized Venom: The Unusually Conserved Venom Profiles of Heloderma Species (Beaded Lizards and Gila Monsters). Toxins. 2014; 6(12):3582-3595. https://doi.org/10.3390/toxins6123582
Chicago/Turabian StyleKoludarov, Ivan, Timothy N. W. Jackson, Kartik Sunagar, Amanda Nouwens, Iwan Hendrikx, and Bryan G. Fry. 2014. "Fossilized Venom: The Unusually Conserved Venom Profiles of Heloderma Species (Beaded Lizards and Gila Monsters)" Toxins 6, no. 12: 3582-3595. https://doi.org/10.3390/toxins6123582
APA StyleKoludarov, I., Jackson, T. N. W., Sunagar, K., Nouwens, A., Hendrikx, I., & Fry, B. G. (2014). Fossilized Venom: The Unusually Conserved Venom Profiles of Heloderma Species (Beaded Lizards and Gila Monsters). Toxins, 6(12), 3582-3595. https://doi.org/10.3390/toxins6123582







