Features of the First Case of Foodborne Botulism Caused by Dual-Toxin Clostridium parabotulinum Subtype A1(B5) in Spain
Abstract
1. Introduction
2. Results
2.1. Case Presentation and Management
2.2. Laboratory Diagnosis and Confirmation of a Dual-Toxin C. parabotulinum Strain
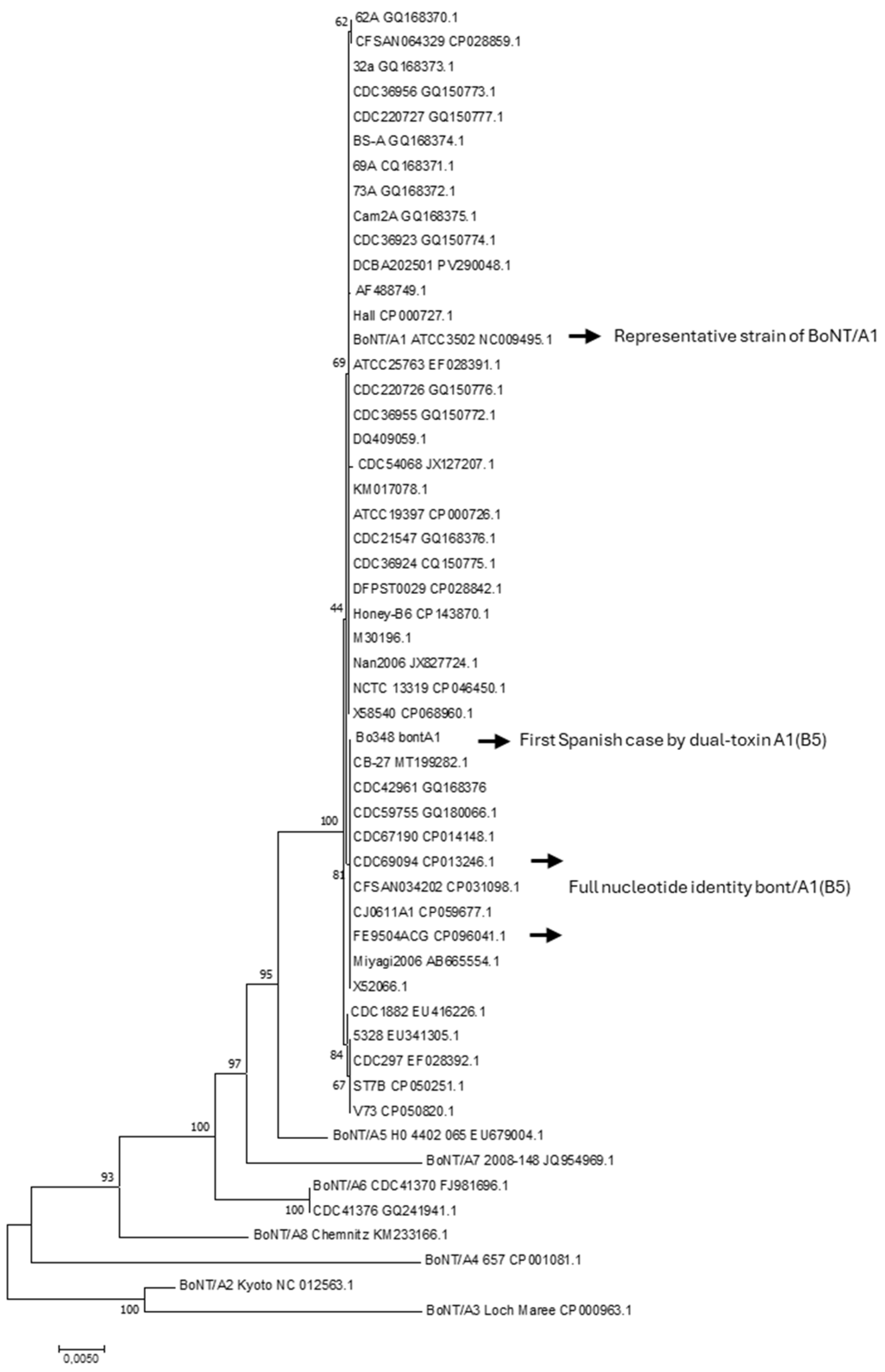
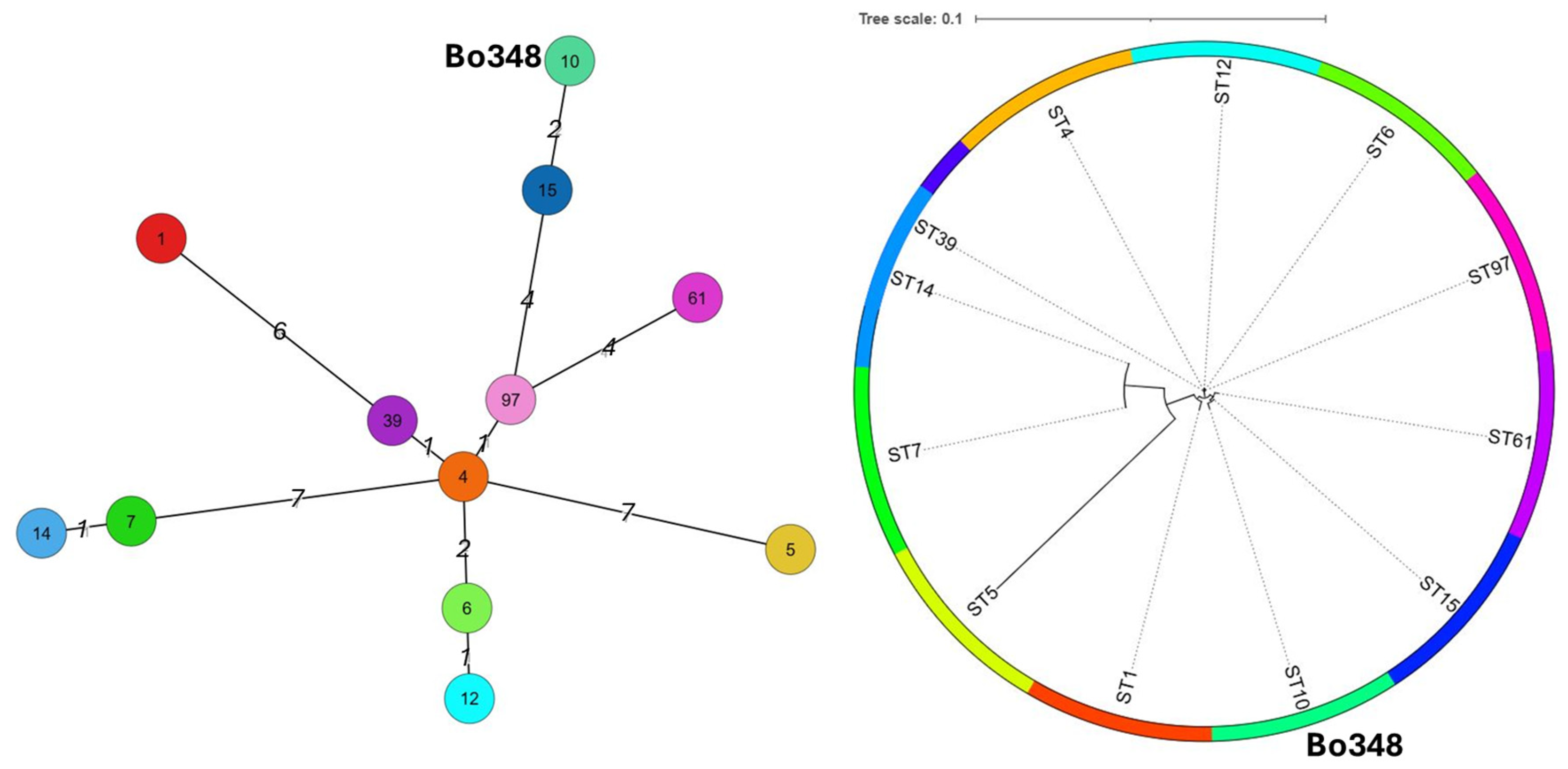
2.3. Genetic Relatedness by Flagellin A and Multi-Locus Sequence Type
3. Discussion
4. Conclusions
5. Material and Methods
5.1. Botulism Diagnosis
5.2. Botulism Confirmation by Bont, flaVR and MLST Genes
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BoNT | Botulinum neurotoxin |
| ST | Sequence type |
References
- Rawson, A.M.; Dempster, A.W.; Humphreys, C.M.; Minton, N.P. Pathogenicity and virulence of Clostridium botulinum. Virulence 2023, 14, 2205251. [Google Scholar] [CrossRef]
- Rossetto, O.; Pirazzini, M.; Montecucco, C. Botulinum neurotoxins: Genetic, structural and mechanistic insights. Nat. Rev. Microbiol. 2014, 12, 535–549. [Google Scholar] [CrossRef]
- Sobel, J. Botulism. Clin. Infect. Dis. 2005, 41, 1167–1173. [Google Scholar] [CrossRef]
- Rao, A.K.; Sobel, J.; Chatham-Stephens, K.; Luquez, C. Clinical Guidelines for Diagnosis and Treatment of Botulism, 2021. MMWR Recomm. Rep. 2021, 70, 1–30. [Google Scholar] [CrossRef]
- Hill, K.K.; Smith, T.J.; Helma, C.H.; Ticknor, L.O.; Foley, B.T.; Svensson, R.T.; Brown, J.L.; Johnson, E.A.; Smith, L.A.; Okinaka, R.T.; et al. Genetic diversity among Botulinum Neurotoxin-producing clostridial strains. J. Bacteriol. 2007, 189, 818–832. [Google Scholar] [CrossRef]
- Peck, M.W.; Smith, T.J.; Anniballi, F.; Austin, J.W.; Bano, L.; Bradshaw, M.; Cuervo, P.; Cheng, L.W.; Derman, Y.; Dorner, B.G.; et al. Historical Perspectives and Guidelines for Botulinum Neurotoxin Subtype Nomenclature. Toxins 2017, 9, 38. [Google Scholar] [CrossRef]
- Smith, T.; Williamson, C.H.D.; Hill, K.; Sahl, J.; Keim, P. Botulinum Neurotoxin-Producing Bacteria. Isn’t It Time that We Called a Species a Species? mBio 2018, 9, 10–1128. [Google Scholar] [CrossRef] [PubMed]
- Brunt, J.; van Vliet, A.H.M.; Carter, A.T.; Stringer, S.C.; Amar, C.; Grant, K.A.; Godbole, G.; Peck, M.W. Diversity of the Genomes and Neurotoxins of Strains of Clostridium botulinum Group I and Clostridium sporogenes Associated with Foodborne, Infant and Wound Botulism. Toxins 2020, 12, 586. [Google Scholar] [CrossRef] [PubMed]
- Grenda, T.; Grenda, A.; Jakubczyk, A.; Rybczyńska-Tkaczyk, K. Opportunistic Features of Non-Clostridium botulinum Strains Containing bont Gene Cluster. Pathogens 2024, 13, 780. [Google Scholar] [CrossRef] [PubMed]
- Dover, N.; Barash, J.R.; Hill, K.K.; Davenport, K.W.; Teshima, H.; Xie, G.; Arnon, S.S.; Popoff, M.R. Clostridium botulinum strain Af84 contains three neurotoxin gene clusters: Bont/A2, bont/F4 and bont/F5. PLoS ONE 2013, 8, e61205. [Google Scholar] [CrossRef]
- Franciosa, G.; Ferreira, J.L.; Hatheway, C.L. Detection of type A, B, and E botulism neurotoxin genes in Clostridium botulinum and other Clostridium species by PCR: Evidence of unexpressed type B toxin genes in type A toxigenic organisms. J. Clin. Microbiol. 1994, 32, 1911–1917. [Google Scholar] [CrossRef]
- Smith, T.J.; Tian, R.; Imanian, B.; Williamson, C.H.D.; Johnson, S.L.; Daligault, H.E.; Schill, K.M. Integration of Complete Plasmids Containing Bont Genes into Chromosomes of Clostridium parabotulinum, Clostridium sporogenes, and Clostridium argentinense. Toxins 2021, 13, 473. [Google Scholar] [CrossRef]
- Popoff, M.R.; Brüggemann, H. Regulatory Networks Controlling Neurotoxin Synthesis in Clostridium botulinum and Clostridium tetani. Toxins 2022, 14, 364. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control. Botulism—Annual Epidemiological Report for 2021; ECDC: Stockholm, Sweden, 2023. Available online: https://www.ecdc.europa.eu/en/publications-data/botulism-annual-epidemiological-report-2021 (accessed on 19 August 2025).
- Valdezate, S.; Carrasco, G.; Medina, M.J.; Garrido, N.; Del Pino, S.; Valiente, M.; Pallarés, M.P.; Villalon, P. Exploring the genetic background of the botulism neurotoxin BoNT/B2 in Spain. Microbiol. Spectr. 2023, 11, e0238023. [Google Scholar] [CrossRef]
- Lindström, M.; Korkeala, H. Laboratory diagnostics of botulism. Clin. Microbiol. Rev. 2006, 19, 298–314. [Google Scholar] [CrossRef] [PubMed]
- Lindström, M.; Keto, R.; Markkula, A.; Nevas, M.; Hielm, S.; Korkeala, H. Multiplex PCR assay for detection and identification of Clostridium botulinum types A, B, E, and F in food and fecal material. Appl. Environ. Microbiol. 2001, 67, 5694–5699. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Lúquez, C.; Raphael, B.H.; Maslanka, S.E. Neurotoxin gene clusters in Clostridium botulinum type Ab strains. Appl. Environ. Microbiol. 2009, 75, 6094–6101. [Google Scholar] [CrossRef] [PubMed]
- Halpin, J.L.; Hill, K.; Johnson, S.L.; Bruce, D.C.; Shirey, T.B.; Dykes, J.K.; Lúquez, C. Finished Whole-Genome Sequences of Two Clostridium botulinum Type A(B) Isolates. Genome Announc. 2017, 5, e00381-17. [Google Scholar] [CrossRef]
- Carter, A.T.; Paul, C.J.; Mason, D.R.; Twine, S.M.; Alston, M.J.; Logan, S.M.; Austin, J.W.; Peck, M.W. Independent evolution of neurotoxin and flagellar genetic loci in proteolytic Clostridium botulinum. BMC Genom. 2009, 10, 115. [Google Scholar] [CrossRef]
- Williamson, C.H.D.; Vazquez, A.J.; Hill, K.; Smith, T.J.; Nottingham, R.; Stone, N.E.; Sobek, C.J.; Cocking, J.H.; Fernández, R.A.; Caballero, P.A.; et al. Differentiating Botulinum Neurotoxin-Producing Clostridia with a Simple, Multiplex PCR Assay. Appl. Environ. Microbiol. 2017, 83, e00806-17. [Google Scholar] [CrossRef]
- Paul, C.J.; Twine, S.M.; Tam, K.J.; Mullen, J.A.; Kelly, J.F.; Austin, J.W.; Logan, S.M. Flagellin diversity in Clostridium botulinum groups I and II: A new strategy for strain identification. Appl. Environ. Microbiol. 2007, 73, 2963–2975. [Google Scholar] [CrossRef]
- Jacobson, M.J.; Lin, G.; Whittam, T.S.; Johnson, E.A. Phylogenetic analysis of Clostridium botulinum type A by multi-locus sequence typing. Microbiology 2008, 154 Pt 8, 2408–2415. [Google Scholar] [CrossRef]
- Jolley, K.A.; Bray, J.E.; Maiden, M.C.J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018, 3, 124. [Google Scholar] [CrossRef]
- Zhou, Z.; Alikhan, N.-F.; Sergeant, M.J.; Luhmann, N.; Vaz, C.; Francisco, A.P.; Carriço, J.A.; Achtman, M. GrapeTree: Visualization of core genomic relationships among 100,000 bacterial pathogens. Genome Res. 2018, 28, 1395–1404. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive tree of life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Vosloo, M.N.; Opperman, C.J.; Geyer, H.D.W.; Setshedi, G.M.; Allam, M.; Kwenda, S.; Ismail, A.; Khumalo, Z.T.H.; Brink, A.J.; Frean, J.A.; et al. First confirmed case of infant botulism in Africa, caused by a dual-toxin-producing Clostridium botulinum strain. Int. J. Infect. Dis. 2021, 103, 164–166. [Google Scholar] [CrossRef] [PubMed]
- Yu, P.A.; Lin, N.H.; Mahon, B.E.; Sobel, J.; Yu, Y.; Mody, R.K.; Gu, W.; Clements, J.; Kim, H.-J.; Rao, A.K. Safety and Improved Clinical Outcomes in Patients Treated with New Equine-Derived Heptavalent Botulinum Antitoxin. Clin. Infect. Dis. 2018, 66 (Suppl. S1), S57–S64. [Google Scholar] [CrossRef]
- Lonati, D.; Schicchi, A.; Crevani, M.; Buscaglia, E.; Scaravaggi, G.; Maida, F.; Cirronis, M.; Petrolini, V.M.; Locatelli, C.A. Foodborne Botulism: Clinical Diagnosis and Medical Treatment. Toxins 2020, 12, 509. [Google Scholar] [CrossRef]
- Atassi, M.Z. Immune Recognition of BoNTs A and B: How Anti-Toxin Antibodies That Bind to the Heavy Chain Obstruct Toxin Action. Toxicon 2009, 54, 600–613. [Google Scholar] [CrossRef] [PubMed]
- Poulain, B.; Lemichez, E.; Popoff, M.R. Neuronal selectivity of botulinum neurotoxins. Toxicon 2020, 178, 20–32. [Google Scholar] [CrossRef]
- Tsai, Y.C.; Maditz, R.; Kuo, C.-L.; Fishman, P.S.; Shoemaker, C.B.; Oyler, G.A.; Weissman, A.M. Targeting Botulinum Neurotoxin Persistence by the Ubiquitin-Proteasome System. Proc. Natl. Acad. Sci. USA 2010, 107, 16554–16559. [Google Scholar] [CrossRef]
- Sen, E.; Kota, K.P.; Panchal, R.G.; Bavari, S.; Kiris, E. Screening of a Focused Ubiquitin-Proteasome Pathway Inhibitor Library Identifies Small Molecules as Novel Modulators of Botulinum Neurotoxin Type A Toxicity. Front. Pharmacol. 2021, 12, 763950. [Google Scholar] [CrossRef]
- Peck, M.W. Biology and genomic analysis of Clostridium botulinum. Adv. Microb. Physiol. 2009, 55, 183–320. [Google Scholar] [CrossRef]
- Rasetti-Escargueil, C.; Lemichez, E.; Popoff, M.R. Public Health Risk Associated with Botulism as Foodborne Zoonoses. Toxins 2019, 12, 17. [Google Scholar] [CrossRef]
- Kenri, T.; Sekizuka, T.; Yamamoto, A.; Iwaki, M.; Komiya, T.; Hatakeyama, T.; Nakajima, H.; Takahashi, M.; Kuroda, M.; Shibayama, K. Genetic characterization and comparison of Clostridium botulinum isolates from botulism cases in Japan between 2006 and 2011. Appl. Environ. Microbiol. 2014, 80, 6954–6964. [Google Scholar] [CrossRef]
- Ohishi, I. Oral toxicities of Clostridium botulinum type A and B toxins from different strains. Infect. Immun. 1984, 43, 487–490. [Google Scholar] [CrossRef] [PubMed]
- Le Maréchal, C.; Huneau, A.; Solanas, S.; Avouac, A.; Jambou, L.; Roux, A.; Boulonne, L.; Rouxel, S.; Jolivet, C.; Poezevara, T.; et al. Overview of Botulinum Neurotoxin-Producing Clostridia in Soils in France. Microbiol. Spectr. 2025, 13, e0011425. [Google Scholar] [CrossRef] [PubMed]
- Le Bouquin, S.; Lucas, C.; Souillard, R.; Le Maréchal, C.; Petit, K.; Kooh, P.; Jourdan-Da Silva, N.; Meurens, F.; Guillier, L.; Mazuet, C. Human and animal botulism surveillance in France from 2008 to 2019. Front. Public. Health 2022, 24, 1003917. [Google Scholar] [CrossRef]
- Harris, R.A.; Tchao, C.; Prystajecky, N.; Weedmark, K.; Tcholakov, Y.; Lefebvre, M.; Austin, J.W. Foodborne Botulism, Canada, 2006–2021. Emerg. Infect. Dis. 2023, 29, 1730–1737. [Google Scholar] [CrossRef] [PubMed]
- Vohra, R.; Barash, J.R.; Karmarkar, E.N.; Koch-Kumar, S.; Sanchez, N.; Gore, M.; Michel, K.; Rangel, M.; Armstrong, E.; Pimentel, L.; et al. Foodborne Botulism Outbreak After Consumption of Home-Canned Cactus (Nopales)—Fresno County, California, 2024. MMWR Morb. Mortal. 2025, 74, 408–413. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Li, K.; Li, F.; Su, J.; Meng, W.; Sun, Y.; Sun, H.; Sun, J.; Yuan, Y.; Lin, Y.; et al. Tracing Foodborne Botulism Events Caused by Clostridium botulinum in Xinjiang Province, China, Using a Core Genome Sequence Typing Scheme. Microbiol. Spectr. 2022, 10, e0116422. [Google Scholar] [CrossRef] [PubMed]
- Raphael, B.H.; Shirey, T.B.; Lúquez, C.; Maslanka, S.E. Distinguishing highly-related outbreak-associated Clostridium botulinum type A(B) strains. BMC Microbiol. 2014, 14, 192. [Google Scholar] [CrossRef] [PubMed]
- Mazuet, C.; Legeay, C.; Sautereau, J.; Ma, L.; Bouchier, C.; Bouvet, P.; Popoff, M.R. Diversity of Group I and II Clostridium botulinum Strains from France Including Recently Identified Subtypes. Genome Biol. Evol. 2016, 8, 1643–1660. [Google Scholar] [CrossRef]
- Smith, T.J.; Williamson, C.H.D.; Hill, K.K.; Johnson, S.L.; Xie, G.; Anniballi, F.; Auricchio, B.; Fernández, R.A.; Caballero, P.A.; Keim, P.; et al. The Distinctive Evolution of orfX Clostridium parabotulinum Strains and Their Botulinum Neurotoxin Type A and F Gene Clusters Is Influenced by Environmental Factors and Gene Interactions via Mobile Genetic Elements. Front. Microbiol. 2021, 12, 566908. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. Botulism in the United States, 1899–1996. In Handbook for Epidemiologists, Clinicians, and Laboratory Workers; CDC: Atlanta, Georgia, 1998. Available online: https://stacks.cdc.gov/view/cdc/6673 (accessed on 23 July 2025).


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Valdezate, S.; Valiente, M.; Carrasco, G.; Medina-Pascual, M.J.; Hurtado, M.I.; de Pipaón, M.R.; Garrido, N.; Paradas, C.; Hernández-Bello, J.R.; Villalón, P. Features of the First Case of Foodborne Botulism Caused by Dual-Toxin Clostridium parabotulinum Subtype A1(B5) in Spain. Toxins 2025, 17, 429. https://doi.org/10.3390/toxins17090429
Valdezate S, Valiente M, Carrasco G, Medina-Pascual MJ, Hurtado MI, de Pipaón MR, Garrido N, Paradas C, Hernández-Bello JR, Villalón P. Features of the First Case of Foodborne Botulism Caused by Dual-Toxin Clostridium parabotulinum Subtype A1(B5) in Spain. Toxins. 2025; 17(9):429. https://doi.org/10.3390/toxins17090429
Chicago/Turabian StyleValdezate, Sylvia, Mónica Valiente, Gema Carrasco, María J. Medina-Pascual, María Isabel Hurtado, Maite Ruiz de Pipaón, Noelia Garrido, Carmen Paradas, José Ramón Hernández-Bello, and Pilar Villalón. 2025. "Features of the First Case of Foodborne Botulism Caused by Dual-Toxin Clostridium parabotulinum Subtype A1(B5) in Spain" Toxins 17, no. 9: 429. https://doi.org/10.3390/toxins17090429
APA StyleValdezate, S., Valiente, M., Carrasco, G., Medina-Pascual, M. J., Hurtado, M. I., de Pipaón, M. R., Garrido, N., Paradas, C., Hernández-Bello, J. R., & Villalón, P. (2025). Features of the First Case of Foodborne Botulism Caused by Dual-Toxin Clostridium parabotulinum Subtype A1(B5) in Spain. Toxins, 17(9), 429. https://doi.org/10.3390/toxins17090429





