A Review of Rattlesnake Venoms
Abstract
1. Introduction
2. Rattlesnake Venom Comparison
3. Venom Protein Composition Patterns
3.1. Genus Crotalus
- C. adamanteus
- C. aquilus
- C. atrox
- C. basiliscus
- C. catalinensis
- C. cerastes
- C. durissus
- C. enyo
- C. ericsmithi
- C. horridus
- C. lepidus
- C. mitchelli
- C. molossus
- C. oreganus
- C. polystictus
- C. pricei
- C. ruber
- C. scutulatus
- C. simus
- C. tigris
- C. vegrandis
- C. viridis
- C. willardi
- C. tortugenesis
- C. stejnegeri
- C. tancitarensis
- C. lannomi
- C. pusillus
- C. transversus
- C. triseriatus
- C. unicolor
- C. intermedius
3.2. Genus Sistrurus
- S. catenatus
- S. miliarius miliarius
- S. miliarius streckeri
- S. miliarius barbouri
3.3. Patterns in Venom Composition
4. Venom Constituents and Biomedical Applications
4.1. Venom Component Activities and Targets
4.1.1. Disintegrins
4.1.2. Cysteine-Rich Secretory Proteins
4.1.3. C-Type Lectins
4.1.4. Bradykinin-Potentiate Peptides
4.1.5. C-Type Natriuretic Peptides
4.1.6. Nerve Growth Factors
4.1.7. Kunitz-Type Serine Protease Inhibitors
4.1.8. Waprin
4.1.9. Snake Venom Metalloproteases
4.1.10. Snake Venom Serine Proteases
4.1.11. Phospholipases A2
4.1.12. Myotoxins
4.2. Biomedical Applications
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Fry, B.G.; Wüster, W.; Kini, R.M.; Brusic, V.; Khan, A.; Venkataraman, D.; Rooney, A.P. Molecular evolution and phylogeny of elapid snake venom three-finger toxins. J. Mol. Evol. 2003, 57, 110–129. [Google Scholar] [CrossRef] [PubMed]
- Tasoulis, T.; Isbister, G.K. A Review and Database of Snake Venom Proteomes. Toxins 2017, 9, 290. [Google Scholar] [CrossRef] [PubMed]
- Assakura, M.T.; Salomao, M.G.; Puorto, G.; Mandelbaum, F.R. Hemorrhagic, fibrinogenolytic and edema-forming activities of the venom of the colubrid snake Philodryas olfersii (green snake). Toxicon 1992, 30, 427–438. [Google Scholar] [CrossRef] [PubMed]
- Daltry, J.C.; Wuster, W.; Thorpe, R.S. Diet and snake venom evolution. Nature 1996, 379, 537–540. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G.; Wickramaratna, J.C.; Hodgson, W.C.; Alewood, P.F.; Kini, R.; Ho, H.; Wüster, W. Electrospray liquid chromatography/mass spectrometry fingerprinting of Acanthophis (death adder) venoms: Taxonomic and toxinological implications. Rapid Commun. Mass. Spectrom. 2002, 16, 600–608. [Google Scholar] [CrossRef] [PubMed]
- Glenn, J.L.; Straight, R.C.; Wolfe, M.C.; Hardy, D.L. Geographical variation in Crotalus scutulatus scutulatus (Mojave rattlesnake) venom properties. Toxicon 1983, 21, 119–130. [Google Scholar] [CrossRef]
- Jimenez Porras, J.M. Intraspecific Variations in Composition of Venom of the Jumping Viper, Bothrops nummifera. Toxicon 1964, 2, 187–195. [Google Scholar] [CrossRef]
- Creer, S.; Malhotra, A.; Thorpe, R.S.; Stocklin, R.S.; Favreau, P.S.; Hao Chou, W.S. Genetic and ecological correlates of intraspecific variation in pitviper venom composition detected using matrix-assisted laser desorption time-of-flight mass spectrometry (MALDI-TOF-MS) and isoelectric focusing. J. Mol. Evol. 2003, 56, 317–329. [Google Scholar] [CrossRef]
- Saravia, N.G.; Weigle, K.; Navas, C.; Segura, I.; Valderrama, L.; Valencia, A.Z.; Escorcia, B.; McMahon-Pratt, D. Heterogeneity, geographic distribution, and pathogenicity of serodemes of Leishmania viannia in Colombia. Am. J. Trop. Med. Hyg. 2002, 66, 738–744. [Google Scholar] [CrossRef]
- Fry, B. Venomous Reptiles and Their Toxins: Evolution, Pathophysiology and Biodiscovery; Oxford University Press: Oxford, UK, 2015. [Google Scholar]
- Tonello, F.; Simonato, M.; Aita, A.; Pizzo, P.; Fernandez, J.; Lomonte, B.; Gutierrez, J.M.; Montecucco, C. A Lys49-PLA2 myotoxin of Bothrops asper triggers a rapid death of macrophages that involves autocrine purinergic receptor signaling. Cell Death Dis. 2012, 3, e343. [Google Scholar] [CrossRef]
- Barlow, A.; Pook, C.E.; Harrison, R.A.; Wuster, W. Coevolution of diet and prey-specific venom activity supports the role of selection in snake venom evolution. Proc. Biol. Sci. 2009, 276, 2443–2449. [Google Scholar] [CrossRef] [PubMed]
- Colis-Torres, A.; Neri-Castro, E.; Strickland, J.L.; Olvera-Rodríguez, A.; Borja, M.; Calvete, J.; Jones, J.; Parkinson, C.L.; Bañuelos, J.; de León, J.L.; et al. Intraspecific venom variation of Mexican West Coast Rattlesnakes (Crotalus basiliscus) and its implications for antivenom production. Biochimie 2022, 192, 111–124. [Google Scholar] [CrossRef] [PubMed]
- Deshwal, A.; Phan, P.; Datta, J.; Kannan, R.; Kumar, T.K.S. A Meta-Analysis of the Protein Components in Rattlesnake Venom. Toxins 2021, 13, 372. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Juarez, P.; Sanz, L. Snake venomics. Strategy and applications. J. Mass. Spectrom. 2007, 42, 1405–1414. [Google Scholar] [CrossRef] [PubMed]
- Cintra, A.C.O.; De Toni, L.G.B.; Sartim, M.A.; Franco, J.J.; Caetano, R.C.; Murakami, M.T.; Sampaio, S.V. Batroxase, a new metalloproteinase from B. atrox snake venom with strong fibrinolytic activity. Toxicon 2012, 60, 70–82. [Google Scholar] [CrossRef] [PubMed]
- Fox, J.W. A brief review of the scientific history of several lesser-known snake venom proteins: L-amino acid oxidases, hyaluronidases and phosphodiesterases. Toxicon 2013, 62, 75–82. [Google Scholar] [CrossRef] [PubMed]
- Mackessy, S.P.; Sixberry, N.M.; Heyborne, W.H.; Fritts, T. Venom of the Brown Treesnake, Boiga irregularis: Ontogenetic shifts and taxa-specific toxicity. Toxicon Off. J. Int. Soc. Toxinology 2006, 47, 537–548. [Google Scholar] [CrossRef]
- Rokyta, D.R.; Margres, M.J.; Calvin, K. Post-transcriptional Mechanisms Contribute Little to Phenotypic Variation in Snake Venoms. G3 Genes. Genomes Genet. 2015, 5, 2375–2382. [Google Scholar] [CrossRef]
- Afifiyan, F.; Armugam, A.; Tan, C.H.; Gopalakrishnakone, P.; Jeyaseelan, K. Postsynaptic α-Neurotoxin Gene of the Spitting Cobra, Naja naja sputatrix: Structure, Organization, and Phylogenetic Analysis. Genome Res. 1999, 9, 259–266. [Google Scholar] [CrossRef]
- Chang, L.; Lin, S.; Huang, H.; Hsiao, M. Genetic organization of alpha-bungarotoxins from Bungarus multicinctus (Taiwan banded krait): Evidence showing that the production of alpha-bungarotoxin isotoxins is not derived from edited mRNAs. Nucleic Acids Res. 1999, 27, 3970–3975. [Google Scholar] [CrossRef]
- Kordis, D.; Gubensek, F. Adaptive evolution of animal toxin multigene families. Gene 2000, 261, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Moura-da-Silva, A.M.; Paine, M.J.I.; Diniz, M.R.V.; Theakston, R.D.G.; Crampton, J.M. The molecular cloning of a phospholipase A 2 from Bothrops jararacussu snake venom: Evolution of venom group II phospholipase A 2’s may imply gene duplications. J. Mol. Evol. 1995, 41, 174–179. [Google Scholar] [CrossRef] [PubMed]
- Slowinski, J.B.; Knight, A.; Rooney, A.P. Inferring species trees from gene trees: A phylogenetic analysis of the Elapidae (Serpentes) based on the amino acid sequences of venom proteins. Mol. Phylogenet Evol. 1997, 8, 349–362. [Google Scholar] [CrossRef] [PubMed]
- Heatwole, H.; Poran, N.S. Resistances of sympatric and allopatric eels to sea snake venoms. Copeia 1995, 1995, 136–147. [Google Scholar] [CrossRef]
- Poran, N.S.; Coss, R.G.; Benjamini, E. Resistance of California ground squirrels (Spermophilus beecheyi) to the venom of the northern Pacific rattlesnake (Crotalus viridis oreganus): A study of adaptive variation. Toxicon 1987, 25, 767–777. [Google Scholar] [CrossRef] [PubMed]
- WuÈster, W.; Daltry, J.C.; Thorpe, R.S. Can diet explain intraspecific venom variation? Reply to Sasa. Toxicon 1999, 37, 253–258. [Google Scholar]
- Seneci, L.; Zdenek, C.N.; Chowdhury, A.; Rodrigues, C.F.B.; Neri-Castro, E.; Benard-Valle, M.; Alagon, A.; Fry, B.G. A Clot Twist: Extreme Variation in Coagulotoxicity Mechanisms in Mexican Neotropical Rattlesnake Venoms. Front. Immunol. 2021, 12, 24. [Google Scholar] [CrossRef]
- Mackessy, S.P. Venom composition in rattlesnakes: Trends and biological significance. In The Biology of Rattlesnakes; Loma Linda University Press: Loma Linda, CA, USA, 2008; pp. 495–510. [Google Scholar]
- Zancolli, G.; Calvete, J.J.; Cardwell, M.D.; Greene, H.W.; Hayes, W.K.; Hegarty, M.J.; Herrmann, H.W.; Holycross, A.T.; Lannutti, D.I.; Mulley, J.F.; et al. When one phenotype is not enough: Divergent evolutionary trajectories govern venom variation in a widespread rattlesnake species. Proc. Biol. Sci. 2019, 286, 20182735. [Google Scholar] [CrossRef]
- Kocholaty, W.F.; Ledford, E.B.; Daly, J.G.; Billings, T.A. Toxicity and some enzymatic properties and activities in the venoms of Crotalidae, Elapidae and Viperidae. Toxicon 1971, 9, 131–138. [Google Scholar] [CrossRef]
- Campbell, J.A.; Lamar, W.W.; Brodie, E.D. The Venomous Reptiles of the Western Hemisphere; Comstock Pub. Associates: Ithaca, NY, USA, 2004; Volume 2. [Google Scholar]
- Boldrini-Franca, J.; Correa-Netto, C.; Silva, M.M.; Rodrigues, R.S.; De La Torre, P.; Perez, A.; Soares, A.M.; Zingali, R.B.; Nogueira, R.A.; Rodrigues, V.M.; et al. Snake venomics and antivenomics of Crotalus durissus subspecies from Brazil: Assessment of geographic variation and its implication on snakebite management. J. Proteom. 2010, 73, 1758–1776. [Google Scholar] [CrossRef]
- Calvete, J.J.; Fasoli, E.; Sanz, L.; Boschetti, E.; Righetti, P.G. Exploring the venom proteome of the western diamondback rattlesnake, Crotalus atrox, via snake venomics and combinatorial peptide ligand library approaches. J. Proteome Res. 2009, 8, 3055–3067. [Google Scholar] [CrossRef] [PubMed]
- Costa, T.R.; Burin, S.M.; Menaldo, D.L.; de Castro, F.A.; Sampaio, S.V. Snake venom L-amino acid oxidases: An overview on their antitumor effects. J. Venom. Anim. Toxins Incl. Trop. Dis. 2014, 20, 23. [Google Scholar] [CrossRef] [PubMed]
- Findrik, Z.; Vasić-Rački, Ð.; Primožič, M.; Habulin, M.; Knez, Ž. Enzymatic activity of L-amino acid oxidase from snake venom Crotalus adamanteus in supercritical CO2. Biocatal. Biotransform. 2005, 23, 315–321. [Google Scholar] [CrossRef]
- Mackessy, S.P. Fractionation of red diamond rattlesnake (Crotalus ruber ruber) venom: Protease, phosphodiesterase, L-amino acid oxidase activities and effects of metal ions and inhibitors on protease activity. Toxicon 1985, 23, 337–340. [Google Scholar] [CrossRef] [PubMed]
- Mackessy, S.P.; Leroy, J.; Mociño-Deloya, E.; Setser, K.; Bryson, R.W.; Saviola, A.J. Venom Ontogeny in the Mexican Lance-Headed Rattlesnake (Crotalus polystictus). Toxins 2018, 10, 271. [Google Scholar] [CrossRef] [PubMed]
- Saviola, A.J.; Gandara, A.J.; Bryson, R.W., Jr.; Mackessy, S.P. Venom phenotypes of the Rock Rattlesnake (Crotalus lepidus) and the Ridge-nosed Rattlesnake (Crotalus willardi) from Mexico and the United States. Toxicon 2017, 138, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Komori, Y.; Sakai, K.; Masuda, K.; Nikai, T. Isolation and Biochemical Characterization of Rubelase, a Non-Hemorrhagic Elastase from Crotalus ruber ruber (Red Rattlesnake) Venom. Toxins 2011, 3, 900–910. [Google Scholar] [CrossRef] [PubMed]
- Mackessy, S.P. Characterization of the major metalloprotease isolated from the venom of the northern pacific rattlesnake, Crotalus viridis oreganus. Toxicon 1996, 34, 1277–1285. [Google Scholar] [CrossRef]
- Smith, C.F.; Mackessy, S.P. The effects of hybridization on divergent venom phenotypes: Characterization of venom from Crotalus scutulatus scutulatus × Crotalus oreganus helleri hybrids. Toxicon 2016, 120, 110–123. [Google Scholar] [CrossRef]
- Boldrini-Franca, J.; Pinheiro-Junior, E.L.; Arantes, E.C. Functional and biological insights of rCollinein-1, a recombinant serine protease from Crotalus durissus collilineatus. J. Venom. Anim. Toxins Incl. Trop. Dis. 2019, 25, e147118. [Google Scholar] [CrossRef]
- Calvete, J.J.; Perez, A.; Lomonte, B.; Sanchez, E.E.; Sanz, L. Snake venomics of Crotalus tigris: The minimalist toxin arsenal of the deadliest Nearctic rattlesnake venom. Evolutionary Clues for generating a pan-specific antivenom against crotalid type II venoms [corrected]. J. Proteome Res. 2012, 11, 1382–1390. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Sanz, L.; Cid, P.; de la Torre, P.; Flores-Diaz, M.; Dos Santos, M.C.; Borges, A.; Bremo, A.; Angulo, Y.; Lomonte, B.; et al. Snake venomics of the Central American rattlesnake Crotalus simus and the South American Crotalus durissus complex points to neurotoxicity as an adaptive paedomorphic trend along Crotalus dispersal in South America. J. Proteome Res. 2010, 9, 528–544. [Google Scholar] [CrossRef] [PubMed]
- Almeida, J.R.; Resende, L.M.; Silva, A.G.; Ribeiro, R.I.; Stabeli, R.G.; Soares, A.M.; Calderon, L.A.; Marangoni, S.; Da Silva, S.L. Biochemical and functional studies of ColTx-I, a new myotoxic phospholipase A2 isolated from Crotalus oreganus lutosus (Great Basin rattlesnake) snake venom. Toxicon 2016, 117, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Almeida, J.R.; Lancellotti, M.; Soares, A.M.; Calderon, L.A.; Ramirez, D.; Gonzalez, W.; Marangoni, S.; Da Silva, S.L. CoaTx-II, a new dimeric Lys49 phospholipase A2 from Crotalus oreganus abyssus snake venom with bactericidal potential: Insights into its structure and biological roles. Toxicon 2016, 120, 147–158. [Google Scholar] [CrossRef] [PubMed]
- Macías-Rodríguez, E.F.; Díaz-Cárdenas, C.O.; Gatica-Colima, A.B.; Plenge-Tellechea, L.F. Seasonal variation in protein content and PLA2 activity of Crotalus molossus molossus venom from captive and wild specimens. Acta Univ. 2014, 24, 38–47. [Google Scholar] [CrossRef]
- Rivas, E.; Neri, E.; Benard, M.; Hernánez-Dávila, A.I.; Zamudio, F.; Alagón, A. General characterization of the venoms from two species of rattlesnakes and an intergrade population (C. lepidus × aquilus) from Aguascalientes and Zacatecas, Mexico. Toxicon 2017, 138, 191–195. [Google Scholar] [CrossRef] [PubMed]
- Griffin, P.R.; Aird, S.D. A new small myotoxin from the venom of the prairie rattlesnake (Crotalus viridis viridis). FEBS Lett. 1990, 274, 43–47. [Google Scholar] [CrossRef]
- Ownby, C.L.; Colberg, T.R.; White, S.P. Isolation, characterization and crystallization of a phospholipase A2 myotoxin from the venom of the prairie rattlesnake (Crotalus viridis viridis). Toxicon 1997, 35, 111–124. [Google Scholar] [CrossRef]
- Saviola, A.J.; Pla, D.; Sanz, L.; Castoe, T.A.; Calvete, J.J.; Mackessy, S.P. Comparative venomics of the Prairie Rattlesnake (Crotalus viridis viridis) from Colorado: Identification of a novel pattern of ontogenetic changes in venom composition and assessment of the immunoreactivity of the commercial antivenom CroFab(R). J. Proteom. 2015, 121, 28–43. [Google Scholar] [CrossRef]
- Strickland, J.L.; Mason, A.J.; Rokyta, D.R.; Parkinson, C.L. Phenotypic Variation in Mojave Rattlesnake (Crotalus scutulatus) Venom Is Driven by Four Toxin Families. Toxins 2018, 10, 135. [Google Scholar] [CrossRef]
- Evangelista, J.S.A.M.; Martins, A.M.C.; Nascimento, N.R.F.; Sousa, C.M.; Alves, R.S.; Toyama, D.O.; Toyama, M.H.; Evangelista, J.J.F.; Menezes, D.B.d.; Fonteles, M.C.; et al. Renal and vascular effects of the natriuretic peptide isolated from Crotalus durissus cascavella venom. Toxicon Off. J. Int. Soc. Toxinology 2008, 52, 737–744. [Google Scholar] [CrossRef] [PubMed]
- Gomes, C.L.; Konno, K.; Conceição, I.M.; Ianzer, D.; Yamanouye, N.; Prezoto, B.C.; Assakura, M.T.; Rádis-Baptista, G.; Yamane, T.; Santos, R.A.; et al. Identification of novel bradykinin-potentiating peptides (BPPs) in the venom gland of a rattlesnake allowed the evaluation of the structure-function relationship of BPPs. Biochem. Pharmacol. 2007, 74, 1350–1360. [Google Scholar] [CrossRef] [PubMed]
- Carey, C.M.; Bueno, R.; Gutierrez, D.A.; Petro, C.; Lucena, S.E.; Sanchez, E.E.; Soto, J.G. Recombinant rubistatin (r-Rub), an MVD disintegrin, inhibits cell migration and proliferation, and is a strong apoptotic inducer of the human melanoma cell line SK-Mel-28. Toxicon 2012, 59, 241–248. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, E.E.; Galan, J.A.; Russell, W.K.; Soto, J.G.; Russell, D.H.; Perez, J.C. Isolation and characterization of two disintegrins inhibiting ADP-induced human platelet aggregation from the venom of Crotalus scutulatus scutulatus (Mohave Rattlesnake). Toxicol. Appl. Pharmacol. 2006, 212, 59–68. [Google Scholar] [CrossRef] [PubMed]
- Saviola, A.J.; Burns, P.D.; Mukherjee, A.K.; Mackessy, S.P. The disintegrin tzabcanin inhibits adhesion and migration in melanoma and lung cancer cells. Int. J. Biol. Macromol. 2016, 88, 457–464. [Google Scholar] [CrossRef] [PubMed]
- Saviola, A.J.; Modahl, C.M.; Mackessy, S.P. Disintegrins of Crotalus simus tzabcan venom: Isolation, characterization and evaluation of the cytotoxic and anti-adhesion activities of tzabcanin, a new RGD disintegrin. Biochimie 2015, 116, 92–102. [Google Scholar] [CrossRef] [PubMed]
- Adade, C.M.; Carvalho, A.L.O.; Tomaz, M.A.; Costa, T.F.R.; Godinho, J.L.; Melo, P.A.; Lima, A.P.C.A.; Rodrigues, J.C.F.; Zingali, R.B.; Souto-Padrón, T. Crovirin, a snake venom cysteine-rich secretory protein (CRISP) with promising activity against Trypanosomes and Leishmania. PLoS Neglected Trop. Dis. 2014, 8, e3252. [Google Scholar] [CrossRef] [PubMed]
- Borja, M.; Neri-Castro, E.; Castaneda-Gaytan, G.; Strickland, J.L.; Parkinson, C.L.; Castaneda-Gaytan, J.; Ponce-Lopez, R.; Lomonte, B.; Olvera-Rodriguez, A.; Alagon, A.; et al. Biological and Proteolytic Variation in the Venom of Crotalus scutulatus scutulatus from Mexico. Toxins 2018, 10, 35. [Google Scholar] [CrossRef]
- Roldán-Padrón, O.; Castro-Guillén, J.L.; García-Arredondo, J.A.; Cruz-Pérez, M.S.; Díaz-Peña, L.F.; Saldaña, C.; Blanco-Labra, A.; García-Gasca, T. Snake Venom Hemotoxic Enzymes: Biochemical Comparison between Crotalus Species from Central Mexico. Molecules 2019, 24, 1489. [Google Scholar] [CrossRef]
- Casewell, N.R.; Wagstaff, S.C.; Wüster, W.; Cook, D.A.; Bolton, F.M.; King, S.I.; Pla, D.; Sanz, L.; Calvete, J.J.; Harrison, R.A. Medically important differences in snake venom composition are dictated by distinct postgenomic mechanisms. Proc. Natl. Acad. Sci. USA 2014, 111, 9205–9210. [Google Scholar] [CrossRef]
- Smith, C.F.; Nikolakis, Z.L.; Ivey, K.; Perry, B.W.; Schield, D.R.; Balchan, N.R.; Parker, J.; Hansen, K.C.; Saviola, A.J.; Castoe, T.A.; et al. Snakes on a plain: Biotic and abiotic factors determine venom compositional variation in a wide-ranging generalist rattlesnake. BMC Biol. 2023, 21, 136. [Google Scholar] [CrossRef] [PubMed]
- Rokyta, D.R.; Wray, K.P.; Margres, M.J. The genesis of an exceptionally lethal venom in the timber rattlesnake (Crotalus horridus) revealed through comparative venom-gland transcriptomics. BMC Genom. 2013, 14, 394. [Google Scholar] [CrossRef] [PubMed]
- Meik, J.M.; Deloya, E.M.; Setser, K. New distribution records for the Queretero dusky rattlesnake Crotalus aquilus (Viperidae), with comments on morphology and habitat use. West. N. Am. Nat. 2007, 67, 601–604. [Google Scholar] [CrossRef]
- Mociño-Deloya, E.; Setser, K.; Peurach, S.C.; Meik, J.M. Crotalus aquilus in the Mexican state of México consumes a diverse summer diet. Herpetol. Bull. 2008, 105, 10–12. [Google Scholar]
- Roldán-Padrón, O.; Cruz-Pérez, M.S.; Castro-Guillén, J.L.; García-Arredondo, J.A.; Mendiola-Olaya, E.; Saldaña-Gutiérrez, C.; Herrera-Paniagua, P.; Blanco-Labra, A.; García-Gasca, T. Hybridization between Crotalus aquilus and Crotalus polystictus Species: A Comparison of Their Venom Toxicity and Enzymatic Activities. Biology 2022, 11, 661. [Google Scholar] [CrossRef] [PubMed]
- Schuett, G.W.; Clark, R.W.; Repp, R.A.; Amarello, M.; Smith, C.F.; Greene, H.W. Social behavior of rattlesnakes: A shifting paradigm. Ratt. Ariz. 2016, 2, 161–244. [Google Scholar]
- Ernst, C.H.; Ernst, E.M. Venomous Reptiles of the United States, Canada, and Northern Mexico: Crotalus; JHU Press: Baltimore, MD, USA, 2011; Volume 2. [Google Scholar]
- Segura, A.; Herrera, M.; Mares, F.R.; Jaime, C.; Sanchez, A.; Vargas, M.; Villalta, M.; Gomez, A.; Gutierrez, J.M.; Leon, G. Proteomic, toxicological and immunogenic characterization of Mexican west -coast rattlesnake (Crotalus basiliscus) venom and its immunological relatedness with the venom of Central American rattlesnake (Crotalus simus). J. Proteom. 2017, 158, 62–72. [Google Scholar] [CrossRef]
- Webber, M.M.; Jezkova, T.; Rodríguez-Robles, J.A. Feeding ecology of sidewinder rattlesnakes, Crotalus cerastes (Viperidae). Herpetologica 2016, 72, 324–330. [Google Scholar] [CrossRef]
- Bosak, A.R.; Ruha, A.M.; Graeme, K.A. A case of neurotoxicity following envenomation by the Sidewinder rattlesnake, Crotalus cerastes. J. Med. Toxicol. 2014, 10, 229–231. [Google Scholar] [CrossRef]
- Frare, B.T.; Silva Resende, Y.K.; Dornelas, B.d.C.; Jorge, M.T.; Souza Ricarte, V.A.; Alves, L.M.; Izidoro, L.F.M. Clinical, laboratory, and therapeutic aspects of Crotalus durissus (South American rattlesnake) victims: A literature review. BioMed Res. Int. 2019, 2019, 1345923. [Google Scholar] [CrossRef]
- Hoyos, M.A.; Almeida-Santos, S.M. The South-American rattlesnake Crotalus durissus: Feeding ecology in the central region of Brazil. Biota Neotrop. 2016, 16, e20140027. [Google Scholar] [CrossRef]
- Taylor, E.N.; Price, A.H. Diet of the Baja California Rattlesnake, Crotalus enyo (Viperidae). Copeia 2001, 2001, 553–555. [Google Scholar] [CrossRef]
- Neri-Castro, E.; Zarzosa, V.; Colis-Torres, A.; Fry, B.G.; Olvera-Rodríguez, A.; Jones, J.; Reyes-Velasco, J.; Zamudio, F.; Borja, M.; Alagón, A.; et al. Proteomic and toxicological characterization of the venoms of the most enigmatic group of rattlesnakes: The long-tailed rattlesnakes. Biochimie 2022, 202, 226–236. [Google Scholar] [CrossRef] [PubMed]
- Campbell, J.A.; Flores-Villela, O. A new long-tailed rattlesnake (Viperidae) from Guerrero, Mexico. Herpetologica 2008, 64, 246–257. [Google Scholar] [CrossRef]
- Holycross, A.T.; Mackessy, S.P. Variation in the Diet of Sistrurus catenatus (Massasauga), with Emphasis on Sistrurus catenatus edwardsii (Desert Massasauga). J. Herpetol. 2002, 36, 454–464. [Google Scholar] [CrossRef]
- McCrystal, H.K.; McCoid, M.J. Crotalus mitchellii. Cat. Am. Amphib. Reptiles 1986, 1986, 388.1–388.4. [Google Scholar]
- Moore, R.G. Seasonal and daily activity patterns and thermoregulation in the southwestern speckled rattlesnake (Crotalus mitchelli pyrrhus) and the Colorado desert sidewinder (Crotalus cerastes laterorepens). Copeia 1978, 1978, 439–442. [Google Scholar] [CrossRef]
- Glaudas, X.; Rodriguez-Robles, J.A. Vagabond males and sedentary females: Spatial ecology and mating system of the speckled rattlesnake (Crotalus mitchellii). Biol. J. Linn. Soc. 2011, 103, 681–695. [Google Scholar] [CrossRef]
- Arnaud-Franco, G.; Cordero-Tapia, A.; Ortiz-Avila, V.; Moctezuma-Gonzalez, C.L.; Tejocote-Perez, M.; Carbajal-Saucedo, A. Comparison of biological and biochemical characteristics of venom from rattlesnakes in the southern Baja California Peninsula. Toxicon 2018, 148, 197–201. [Google Scholar] [CrossRef]
- Holzer, M.; Mackessy, S.P. An aqueous endpoint assay of snake venom phospholipase A2. Toxicon 1996, 34, 1149–1155. [Google Scholar] [CrossRef]
- Tan, K.K.; Ler, S.G.; Gunaratne, J.; Bay, B.H.; Ponnampalam, G. In Vitro Cytotoxicity of L-amino Acid Oxidase from the Venom of Crotalus mitchellii pyrrhus. Toxicon Off. J. Int. Soc. Toxinol. 2017, 139, 20–30. [Google Scholar] [CrossRef] [PubMed]
- Carbajal-Márquez, R.A.; Sigala-Rodríguez, J.J.; Reyes-Velasco, J.; Jones, J.M.; Montaño-Ruvalcaba, C.; Fernandez-Badillo, L.; Borja-Jiménez, J.M. New dietary records for three species in the Crotalus molossus species complex (Serpentes: Viperidae). Phyllomedusa 2023, 22, 81–86. [Google Scholar] [CrossRef]
- Anderson, C.G.; Greenbaum, E. Phylogeography of northern populations of the black-tailed rattlesnake (Crotalus molossus Baird and Girard, 1853), with the revalidation of C. ornatus Hallowell, 1854. Herpetol. Monogr. 2012, 26, 19–57. [Google Scholar] [CrossRef]
- Sparks, A.M.; Lind, C.; Taylor, E.N. Diet of the Northern Pacific rattlesnake (Crotalus o. oreganus) in California. Herpetol. Rev. 2015, 46, 161–165. [Google Scholar]
- Hecker, L.J.; Bean, W.T.; Marks, S.B. Compensatory Microhabitat Selection by Northern Pacific Rattlesnakes (Crotalus oreganus oreganus) in a Cool and Wet Macroclimate. J. Herpetol. 2020, 54, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Grabowsky, E.R.; Mackessy, S.P. Predator-prey interactions and venom composition in a high elevation lizard specialist, Crotalus pricei (Twin-spotted Rattlesnake). Toxicon Off. J. Int. Soc. Toxinol. 2019, 170, 29–40. [Google Scholar] [CrossRef] [PubMed]
- Brown, T.K.; Lemm, J.M.; Montagne, J.P.; Tracey, J.A.; Alberts, A.C. Spatial ecology, habitat use, and survivorship of resident and translocated red diamond rattlesnakes (Crotalus ruber). In The Biology of Rattlesnakes; Loma Linda University Press: Loma Linda, CA, USA, 2008. [Google Scholar]
- Dobson, J.; Yang, D.C.; Op den Brouw, B.; Cochran, C.; Huynh, T.; Kurrupu, S.; Sanchez, E.E.; Massey, D.J.; Baumann, K.; Jackson, T.N.W.; et al. Rattling the border wall: Pathophysiological implications of functional and proteomic venom variation between Mexican and US subspecies of the desert rattlesnake Crotalus scutulatus. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2018, 205, 62–69. [Google Scholar] [CrossRef]
- Neri-Castro, E.; Bénard-Valle, M.; Paniagua, D.; Boyer, L.V.; Possani, L.D.; López-Casillas, F.; Olvera, A.; Romero, C.; Zamudio, F.; Alagón, A. Neotropical Rattlesnake Crotalus simus Venom Pharmacokinetics in Lymph and Blood Using an Ovine Model. Toxins 2020, 12, 455. [Google Scholar] [CrossRef]
- Goode, M. Conservation Genetics of the Tiger Rattlesnake (Crotalus tigris) in the Context of Long-Term Ecological Data. Doctoral Dissertation, The University of Arizona, Tucson, AZ, USA, 2015. [Google Scholar]
- Viala, V.L.; Hildebrand, D.; Fucase, T.M.; Sciani, J.M.; Prezotto-Neto, J.P.; Riedner, M.; Sanches, L.; Nishimura, P.J.; Oguiura, N.; Pimenta, D.C.; et al. Proteomic analysis of the rare Uracoan rattlesnake Crotalus vegrandis venom: Evidence of a broad arsenal of toxins. Toxicon 2015, 107, 234–251. [Google Scholar] [CrossRef]
- Gardiner, L.E.; Somers, C.M.; Parker, D.L.; Poulin, R.G. Microhabitat Selection by Prairie Rattlesnakes (Crotalus viridis) at the Northern Extreme of their Geographic Range. J. Herpetol. 2015, 49, 131–137. [Google Scholar] [CrossRef]
- Holycross, A.T.; Goldberg, S.R. Reproduction in northern populations of the ridgenose rattlesnake, Crotalus willardi (Serpentes: Viperidae). Copeia 2001, 2001, 473–481. [Google Scholar] [CrossRef]
- Mociño-Deloya, E.; Setser, K.; Heacker, M.; Peurach, S. Diet of New Mexico Ridge-nosed Rattlesnake (Crotalus willardi obscurus) in the Sierra San Luis and Sierra Pan Duro, México. J. Herpetol. 2015, 49, 104–107. [Google Scholar] [CrossRef] [PubMed]
- van der Heiden, A.M.; Flores-Villela, O. New records of the rare Sinaloan Long-tailed Rattlesnake, Crotalus stejnegeri, from southern Sinaloa, Mexico. Rev. Mex. Biodivers. 2013, 84, 1343–1348. [Google Scholar] [CrossRef]
- Grabowsky, E.R.; Saviola, A.J.; Alvarado-Díaz, J.; Mascareñas, A.Q.; Hansen, K.C.; Yates, J.; Mackessy, S.P. Montane Rattlesnakes in Mexico: Venoms of Crotalus tancitarensis and Related Species within the Crotalus intermedius Group. Toxins 2023, 15, 72. [Google Scholar] [CrossRef] [PubMed]
- Reyes-Velasco, J.; Grünwald, C.I.; Jones, J.M.; Weatherman, G.N. Rediscovery of the rare Autlán long-tailed rattlesnake, Crotalus lannomi. Herpetol. Rev. 2010, 41, 19. [Google Scholar]
- McCranie, J.R. Crotalus pusillus. Cat. Am. Amphib. Reptiles 1983, 1983, 313.1–313.2. [Google Scholar]
- Campbell, J.A. Crotalus pusillus. Cat. Am. Amphib. Reptiles 1988, 1988, 450.1–450.3. [Google Scholar]
- Jaramillo-Alba, J.; de la Vega-Pérez, A.H.D.; Bucio-Jiménez, L.E.; Méndez-De la Cruz, F.R.; Pérez-Mendoza, H.A. Comparative thermal ecology parameters of the mexican dusky rattlesnake (Crotalus triseriatus). J. Therm. Biol. 2020, 92, 102695. [Google Scholar] [CrossRef]
- Mociño-Deloya, E.; Setser, K.; Pérez-Ramos, E. Observations on the diet of Crotalus triseriatus (Mexican dusky rattlesnake). Rev. Mex. Biodivers. 2014, 85, 1289–1291. [Google Scholar] [CrossRef]
- Reinert, H.K.; Bushar, L.M.; Rocco, G.L.; Goode, M.; Odum, R.A. Distribution of the Aruba Island rattlesnake, Crotalus unicolor, on Aruba, Dutch West Indies. Caribb. J. Sci. 2002, 38, 126–128. [Google Scholar]
- Reinert, H.K.; Kodrich, W.R. Movements and Habitat Utilization by the Massasauga, Sistrurus-Catenatus Catenatus. J. Herpetol. 1982, 16, 162–171. [Google Scholar] [CrossRef]
- Keenlyne, K.; Beer, J. Food habits of Sistrurus catenatus catenatus. J. Herpetol. 1973, 7, 382–384. [Google Scholar] [CrossRef]
- Gibbs, H.L.; Sanz, L.; Sovic, M.G.; Calvete, J.J. Phylogeny-based comparative analysis of venom proteome variation in a clade of rattlesnakes (Sistrurus sp.). PLoS ONE 2013, 8, e67220. [Google Scholar] [CrossRef] [PubMed]
- Palmer, W.M. Sistrurus miliarius. Cat. Am. Amphib. Reptiles 1978, 1978, 220.1–220.2. [Google Scholar]
- Gibbs, H.L.; Sanz, L.; Chiucchi, J.E.; Farrell, T.M.; Calvete, J.J. Proteomic analysis of ontogenetic and diet-related changes in venom composition of juvenile and adult Dusky Pigmy rattlesnakes (Sistrurus miliarius barbouri). J. Proteom. 2011, 74, 2169–2179. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Moreno-Murciano, M.P.; Theakston, R.D.; Kisiel, D.G.; Marcinkiewicz, C. Snake venom disintegrins: Novel dimeric disintegrins and structural diversification by disulphide bond engineering. Biochem. J. 2003, 372, 725–734. [Google Scholar] [CrossRef]
- Soto, J.G.; White, S.A.; Reyes, S.R.; Regalado, R.; Sanchez, E.E.; Perez, J.C. Molecular evolution of PIII-SVMP and RGD disintegrin genes from the genus Crotalus. Gene 2007, 389, 66–72. [Google Scholar] [CrossRef]
- Calvete, J.J.; Marcinkiewicz, C.; Monleon, D.; Esteve, V.; Celda, B.; Juarez, P.; Sanz, L. Snake venom disintegrins: Evolution of structure and function. Toxicon 2005, 45, 1063–1074. [Google Scholar] [CrossRef]
- Fry, B.G. From genome to “venome”: Molecular origin and evolution of the snake venom proteome inferred from phylogenetic analysis of toxin sequences and related body proteins. Genome Res. 2005, 15, 403–420. [Google Scholar] [CrossRef]
- Galán, J.A.; Sánchez, E.E.; Bashir, S.a.; Pérez, J.C. Characterization and identification of disintegrins in Crotalus horridus venom by liquid chromatography and tandem matrix-assisted laser desorption ionization-quadrupole ion trap time-of-flight (MALDI-QIT-TOF) mass spectrometry. Can. J. Chem. 2005, 83, 1124–1131. [Google Scholar] [CrossRef]
- Galán, J.A.; Sánchez, E.E.; Rodríguez-Acosta, A.; Soto, J.G.; Bashir, S.; McLane, M.A.; Paquette-Straub, C.; Pérez, J.C. Inhibition of lung tumor colonization and cell migration with the disintegrin crotatroxin 2 isolated from the venom of Crotalus atrox. Toxicon Off. J. Int. Soc. Toxinol. 2008, 51, 1186–1196. [Google Scholar] [CrossRef] [PubMed]
- Borja, M.; Galan, J.A.; Cantu, E.; Zugasti-Cruz, A.; Rodriguez-Acosta, A.; Lazcano, D.; Lucena, S.; Suntravat, M.; Sanchez, Y.E.E. Morulustatin, A Disintegrin that Inhibits ADP-Induced Platelet Aggregation, Isolated from the Mexican Tamaulipan Rock Rattlesnake (Crotalus lepidus morulus). Rev. Cient. 2016, 26, 86–94. [Google Scholar]
- Sanchez, E.E.; Galan, J.A.; Powell, R.L.; Reyes, S.R.; Soto, J.G.; Russell, W.K.; Russell, D.H.; Perez, J.C. Disintegrin, hemorrhagic, and proteolytic activities of Mohave rattlesnake, Crotalus scutulatus scutulatus venoms lacking Mojave toxin. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2005, 141, 124–132. [Google Scholar] [CrossRef] [PubMed]
- Markland, F.S.; Swenson, S. Snake venom metalloproteinases. Toxicon 2013, 62, 3–18. [Google Scholar] [CrossRef] [PubMed]
- Suntravat, M.; Jia, Y.; Lucena, S.E.; Sánchez, E.E.; Pérez, J.C. cDNA cloning of a snake venom metalloproteinase from the eastern diamondback rattlesnake (Crotalus adamanteus), and the expression of its disintegrin domain with anti-platelet effects. Toxicon Off. J. Int. Soc. Toxinol. 2013, 64, 43–54. [Google Scholar] [CrossRef] [PubMed]
- Urra, F.A.; Araya-Maturana, R. Targeting Metastasis with Snake Toxins: Molecular Mechanisms. Toxins 2017, 9, 390. [Google Scholar] [CrossRef] [PubMed]
- Rokyta, D.R.; Lemmon, A.R.; Margres, M.J.; Aronow, K. The venom-gland transcriptome of the eastern diamondback rattlesnake (Crotalus adamanteus). BMC Genom. 2012, 13, 312. [Google Scholar] [CrossRef] [PubMed]
- Lecht, S.; Chiaverelli, R.A.; Gerstenhaber, J.; Calvete, J.J.; Lazarovici, P.; Casewell, N.R.; Harrison, R.; Lelkes, P.I.; Marcinkiewicz, C. Anti-angiogenic activities of snake venom CRISP isolated from Echis carinatus sochureki. Biochim. Biophys. Acta (BBA)—Gen. Subj. 2015, 1850, 1169–1179. [Google Scholar] [CrossRef]
- Arruda Macedo, J.K.; Fox, J.W.; de Souza Castro, M. Disintegrins from snake venoms and their applications in cancer research and therapy. Curr. Protein Pept. Sci. 2015, 16, 532–548. [Google Scholar] [CrossRef]
- Da Silva, M.; Lucena, S.; Aguilar, I.; Rodriguez-Acosta, A.; Salazar, A.M.; Sanchez, E.E.; Giron, M.E.; Carvajal, Z.; Arocha-Pinango, C.L.; Guerrero, B. Anti-platelet effect of cumanastatin 1, a disintegrin isolated from venom of South American Crotalus rattlesnake. Thromb. Res. 2009, 123, 731–739. [Google Scholar] [CrossRef]
- Fox, J.W.; Serrano, S.M. Approaching the golden age of natural product pharmaceuticals from venom libraries: An overview of toxins and toxin-derivatives currently involved in therapeutic or diagnostic applications. Curr. Pharm. Des. 2007, 13, 2927–2934. [Google Scholar] [CrossRef] [PubMed]
- Lucena, S.E.; Romo, K.; Suntravat, M.; Sánchez, E.E. Anti-angiogenic activities of two recombinant disintegrins derived from the Mohave and Prairie rattlesnakes. Toxicon 2014, 78, 10–17. [Google Scholar] [CrossRef] [PubMed]
- Mackessy, S.; Saviola, A.; Mukherjee, A. Venom toxins to drugs: Anti-thrombotic and anti-metastasis compounds from snake venoms. Toxicon 2018, 150, 320. [Google Scholar] [CrossRef]
- Suntravat, M.; Barret, H.S.; Jurica, C.A.; Lucena, S.E.; Perez, J.C.; Sánchez, E.E. Recombinant disintegrin (r-Cam-dis) from Crotalus adamanteus inhibits adhesion of human pancreatic cancer cell lines to laminin-1 and vitronectin. J. Venom. Res. 2015, 6, 1–10. [Google Scholar] [PubMed]
- Marquez, J.; Parra, V.; Suntravat, M.; Sanchez, E.E. Purification and characterization of cysteine rich-secretory proteins (CRiSPs) from the venom of the Southern Pacific rattlesnake (Crotalus oreganus helleri): Their role on blood and lymphatic endothelial cell permeability. Toxicon 2018, 150, 315–334. [Google Scholar] [CrossRef]
- Sanz, L.; Gibbs, H.L.; Mackessy, S.P.; Calvete, J.J. Venom Proteomes of Closely Related Sistrurus Rattlesnakes with Divergent Diets. J. Proteome Res. 2006, 5, 2098–2112. [Google Scholar] [CrossRef] [PubMed]
- Tadokoro, T.M.; Modahl, C.; Maenaka, K.; Aoki-Shioi, N. Cysteine-Rich Secretory Proteins (CRISPs) From Venomous Snakes: An Overview of the Functional Diversity in A Large and Underappreciated Superfamily. Toxins 2020, 12, 175. [Google Scholar] [CrossRef]
- Wang, Y.-L.; Goh, K.-X.; Wu, W.-G.; Chen, C.-J. Purification, crystallization and preliminary X-ray crystallographic analysis of a cysteine-rich secretory protein (CRISP) from Naja atra venom. Acta Crystallographica. Sect. D Biol. Crystallogr. 2004, 60, 1912–1915. [Google Scholar] [CrossRef]
- Peichoto, M.E.; Mackessy, S.P.; Teibler, P.; Tavares, F.L.; Burckhardt, P.L.; Breno, M.C.; Acosta, O.; Santoro, M.L. Purification and characterization of a cysteine-rich secretory protein from Philodryas patagoniensis snake venom. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2009, 150, 79–84. [Google Scholar] [CrossRef]
- Yamazaki, Y.; Morita, T. Structure and function of snake venom cysteine-rich secretory proteins. Toxicon 2004, 44, 227–231. [Google Scholar] [CrossRef]
- Shikamoto, Y.; Suto, K.; Yamazaki, Y.; Morita, T.; Mizuno, H. Crystal structure of a CRISP family Ca2+-channel blocker derived from snake venom. J. Mol. Biol. 2005, 350, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Lodovicho, M.E.; Costa, T.; Bernardes, C.P.; Menaldo, D.L.; Zoccal, K.F.; Carone, S.E.; Rosa, J.C.; Pucca, M.B.; Cerni, F.A.; Arantes, E.C.; et al. Investigating possible biological targets of Bj-CRP, the first cysteine-rich secretory protein (CRISP) isolated from Bothrops jararaca snake venom. Toxicol. Lett. 2017, 265, 156–169. [Google Scholar] [CrossRef] [PubMed]
- Brown, R.L.; Lynch, L.L.; Haley, T.L.; Arsanjani, R. Pseudechetoxin Binds to the Pore Turret of Cyclic Nucleotide–gated Ion Channels. J. Gen. Physiol. 2003, 122, 749–760. [Google Scholar] [CrossRef] [PubMed]
- Yamazaki, Y.; Brown, L.R.; Morita, T. Purification and Cloning of Toxins from Elapid Venoms that Target Cyclic Nucleotide-Gated Ion Channels. Biochemistry 2002, 41, 11331–11337. [Google Scholar] [CrossRef] [PubMed]
- Yamazaki, Y.; Koike, H.; Sugiyama, Y.; Motoyoshi, K.; Wada, T.; Hishinuma, S.; Mita, M.; Morita, T. Cloning and characterization of novel snake venom proteins that block smooth muscle contraction. Eur. J. Biochem. 2002, 269, 2708–2715. [Google Scholar] [CrossRef]
- Clemetson, K.J. Snaclecs (snake C-type lectins) that inhibit or activate platelets by binding to receptors. Toxicon 2010, 56, 1236–1246. [Google Scholar] [CrossRef] [PubMed]
- Morita, T. Structures and functions of snake venom CLPs (C-type lectin-like proteins) with anticoagulant-, procoagulant-, and platelet-modulating activities. Toxicon 2005, 45, 1099–1114. [Google Scholar] [CrossRef] [PubMed]
- Rini, J.M. Lectin structure. Annu. Rev. Biophys. Biomol. Struct. 1995, 24, 551–577. [Google Scholar] [CrossRef]
- Sharon, N.; Lis, H. History of lectins: From hemagglutinins to biological recognition molecules. Glycobiology 2004, 14, 53r–62r. [Google Scholar] [CrossRef]
- Drickamer, K. Engineering galactose-binding activity into a C-type mannose-binding protein. Nature 1992, 360, 183–186. [Google Scholar] [CrossRef]
- Abreu, P.A.; Albuquerque, M.G.; Rodrigues, C.R.; Castro, H.C. Structure-function inferences based on molecular modeling, sequence-based methods and biological data analysis of snake venom lectins. Toxicon 2006, 48, 690–701. [Google Scholar] [CrossRef] [PubMed]
- Hirabayashi, J.; Kusunoki, T.; Kasai, K. Complete primary structure of a galactose-specific lectin from the venom of the rattlesnake Crotalus atrox. Homologies with Ca2(+)-dependent-type lectins. J. Biol. Chem. 1991, 266, 2320–2326. [Google Scholar] [CrossRef] [PubMed]
- Sartim, M.A.; Sampaio, S.V. Snake venom galactoside-binding lectins: A structural and functional overview. J. Venom. Anim. Toxins Incl. Trop. Dis. 2015, 21, 35. [Google Scholar] [CrossRef] [PubMed]
- Walker, J.R.; Nagar, B.; Young, N.M.; Hirama, T.; Rini, J.M. X-ray crystal structure of a galactose-specific C-type lectin possessing a novel decameric quaternary structure. Biochemistry 2004, 43, 3783–3792. [Google Scholar] [CrossRef]
- Durban, J.; Sanz, L.; Trevisan-Silva, D.; Neri-Castro, E.; Alagon, A.; Calvete, J.J. Integrated Venomics and Venom Gland Transcriptome Analysis of Juvenile and Adult Mexican Rattlesnakes Crotalus simus, C. tzabcan, and C. culminatus Revealed miRNA-modulated Ontogenetic Shifts. J. Proteome Res. 2017, 16, 3370–3390. [Google Scholar] [CrossRef]
- Rokyta, D.R.; Wray, K.P.; Lemmon, A.R.; Lemmon, E.M.; Caudle, S.B. A high-throughput venom-gland transcriptome for the Eastern Diamondback Rattlesnake (Crotalus adamanteus) and evidence for pervasive positive selection across toxin classes. Toxicon Off. J. Int. Soc. Toxinol. 2011, 57, 657–671. [Google Scholar] [CrossRef]
- Margres, M.J.; McGivern, J.J.; Wray, K.P.; Seavy, M.; Calvin, K.; Rokyta, D.R. Linking the transcriptome and proteome to characterize the venom of the eastern diamondback rattlesnake (Crotalus adamanteus). J. Proteom. 2014, 96, 145–158. [Google Scholar] [CrossRef]
- Mastro, A.M.; Hurley, D.J.; Winning, R.K.; Filipowski, R.; Ogilvie, M.L.; Gartner, T.K. Mitogenic activity of snake venom lectins. Cell Tissue Kinet. 1986, 19, 557–566. [Google Scholar] [CrossRef]
- Nikai, T.; Kato, S.; Komori, Y.; Sugihara, H. Amino acid sequence and biological properties of the lectin from the venom of Trimeresurus okinavensis (Himehabu). Toxicon 2000, 38, 707–711. [Google Scholar] [CrossRef]
- Lomonte, B.; Rojas, G.; Gutierrez, J.M.; Ramirez, G. Isolation of a galactose-binding lectin from the venom of the snake Bothrops godmani (Godmann’s pit viper). Toxicon 1990, 28, 75–81. [Google Scholar] [CrossRef]
- Ogilvie, M.L.; Byl, J.W.; Gartner, T.K. Platelet-aggregation is stimulated by lactose-inhibitable snake venom lectins. Thromb. Haemost. 1989, 62, 704–707. [Google Scholar] [PubMed]
- Castanheira, L.E.; Nunes, D.C.; Cardoso, T.M.; Santos Pde, S.; Goulart, L.R.; Rodrigues, R.S.; Richardson, M.; Borges, M.H.; Yoneyama, K.A.; Rodrigues, V.M. Biochemical and functional characterization of a C-type lectin (BpLec) from Bothrops pauloensis snake venom. Int. J. Biol. Macromol. 2013, 54, 57–64. [Google Scholar] [CrossRef] [PubMed]
- Nunes Edos, S.; de Souza, M.A.; Vaz, A.F.; Santana, G.M.; Gomes, F.S.; Coelho, L.C.; Paiva, P.M.; da Silva, R.M.; Silva-Lucca, R.A.; Oliva, M.L.; et al. Purification of a lectin with antibacterial activity from Bothrops leucurus snake venom. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2011, 159, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Damasio Dde, C.; Nolte, S.; Polak, L.P.; Brandt, A.P.; Bonan, N.B.; Zischler, L.; Stuelp-Campelo, P.M.; Cadena, S.M.; Noronha, L.; Elifio-Esposito, S.L.; et al. The lectin BJcuL induces apoptosis through TRAIL expression, caspase cascade activation and mitochondrial membrane permeability in a human colon adenocarcinoma cell line. Toxicon 2014, 90, 299–307. [Google Scholar] [CrossRef] [PubMed]
- Nolte, S.; de Castro Damasio, D.; Barea, A.C.; Gomes, J.; Magalhaes, A.; Mello Zischler, L.F.; Stuelp-Campelo, P.M.; Elifio-Esposito, S.L.; Roque-Barreira, M.C.; Reis, C.A.; et al. BJcuL, a lectin purified from Bothrops jararacussu venom, induces apoptosis in human gastric carcinoma cells accompanied by inhibition of cell adhesion and actin cytoskeleton disassembly. Toxicon 2012, 59, 81–85. [Google Scholar] [CrossRef] [PubMed]
- Nunes, E.S.; Souza, M.A.A.; Vaz, A.F.M.; Silva, T.G.; Aguiar, J.S.; Batista, A.M.; Guerra, M.M.P.; Guarnieri, M.C.; Coelho, L.C.B.B.; Correia, M.T.S. Cytotoxic effect and apoptosis induction by Bothrops leucurus venom lectin on tumor cell lines. Toxicon 1962, 59, 667–671. [Google Scholar] [CrossRef] [PubMed]
- Rebello Horta, C.C.; Chatzaki, M.; Rezende, B.A.; Magalhaes Bde, F.; Duarte, C.G.; Felicori, L.F.; Ribeiro Oliveira-Mendes, B.B.; do Carmo, A.O.; Chavez-Olortegui, C.; Kalapothakis, E. Cardiovascular-Active Venom Toxins: An Overview. Curr. Med. Chem. 2016, 23, 603–622. [Google Scholar] [CrossRef] [PubMed]
- Higuchi, S.; Murayama, N.; Saguchi, K.; Ohi, H.; Fujita, Y.; Camargo, A.C.; Ogawa, T.; Deshimaru, M.; Ohno, M. Bradykinin-potentiating peptides and C-type natriuretic peptides from snake venom. Immunopharmacology 1999, 44, 129–135. [Google Scholar] [CrossRef]
- Lopes, D.M.; Junior, N.E.; Costa, P.P.; Martins, P.L.; Santos, C.F.; Carvalho, E.D.; Carvalho, M.D.; Pimenta, D.C.; Cardi, B.A.; Fonteles, M.C.; et al. A new structurally atypical bradykinin-potentiating peptide isolated from Crotalus durissus cascavella venom (South American rattlesnake). Toxicon 2014, 90, 36–44. [Google Scholar] [CrossRef]
- Cotton, J.; Hayashi, M.A.; Cuniasse, P.; Vazeux, G.; Ianzer, D.; De Camargo, A.C.; Dive, V. Selective Inhibition of the C-Domain of Angiotensin I Converting Enzyme by Bradykinin Potentiating Peptides. Biochemistry 2002, 41, 6065–6071. [Google Scholar] [CrossRef]
- Ferreira, S.H.; Bartelt, D.C.; Greene, L.J. Isolation of bradykinin-potentiating peptides from Bothrops jararaca venom. Biochemistry 1970, 9, 2583–2593. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, L.A.; Henriques, O.B.; Lebrun, I.; Batista, M.B.; Prezoto, B.C.; Andreoni, A.S.; Zelnik, R.; Habermehl, G. A new bradykinin-potentiating peptide (peptide P) isolated from the venom of Bothrops jararacussu (jararacucu tapete, urutu dourado). Toxicon 1992, 30, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Camargo, A.; Ferreira, S.H. Action of bradykinin potentiating factor (BPF) and dimercaprol (BAL) on the responses to bradykinin of isolated preparations of rat intestines. Br. J. Pharmacol. 1971, 42, 305–307. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, S.H.; Rocha e Silva, M. Potentiation of bradykinin and eledoisin by BPF (bradykinin potentiating factor) from Bothrops jararaca venom. Experientia 1965, 21, 347–349. [Google Scholar] [CrossRef] [PubMed]
- Cushman, D.W.; Ondetti, M.A. History of the design of captopril and related inhibitors of angiotensin converting enzyme. Hypertension 1991, 17, 589–592. [Google Scholar] [CrossRef] [PubMed]
- Ondetti, M.A.; Rubin, B.; Cushman, D.W. Design of specific inhibitors of angiotensin-converting enzyme: New class of orally active antihypertensive agents. Science 1977, 196, 441–444. [Google Scholar] [CrossRef] [PubMed]
- Ianzer, D.; Santos, R.A.; Etelvino, G.M.; Xavier, C.H.; de Almeida Santos, J.; Mendes, E.P.; Machado, L.T.; Prezoto, B.C.; Dive, V.; de Camargo, A.C. Do the cardiovascular effects of angiotensin-converting enzyme (ACE) I involve ACE-independent mechanisms? new insights from proline-rich peptides of Bothrops jararaca. J. Pharmacol. Exp. Ther. 2007, 322, 795–805. [Google Scholar] [CrossRef]
- Wei, L.; Alhenc-Gelas, F.; Corvol, P.; Clauser, E. The two homologous domains of human angiotensin I-converting enzyme are both catalytically active. J. Biol. Chem. 1991, 266, 9002–9008. [Google Scholar] [CrossRef]
- Murayama, N.; Hayashi, M.A.; Ohi, H.; Ferreira, L.A.; Hermann, V.V.; Saito, H.; Fujita, Y.; Higuchi, S.; Fernandes, B.L.; Yamane, T.; et al. Cloning and sequence analysis of a Bothrops jararaca cDNA encoding a precursor of seven bradykinin-potentiating peptides and a C-type natriuretic peptide. Proc. Natl. Acad. Sci. USA 1997, 94, 1189–1193. [Google Scholar] [CrossRef]
- Fry, B.G.; Wüster, W. Assembling an arsenal: Origin and evolution of the snake venom proteome inferred from phylogenetic analysis of toxin sequences. Mol. Biol. Evol. 2004, 21, 870–883. [Google Scholar] [CrossRef]
- Alves, R.S.; Ximenes, R.M.; Jorge, A.R.C.; Nascimento, N.R.F.; Martins, R.D.; Rabello, M.M.; Hernandes, M.Z.; Toyama, D.O.; Toyama, M.H.; Martins, A.M.C.; et al. Isolation, homology modeling and renal effects of a C-type natriuretic peptide from the venom of the Brazilian yellow scorpion (Tityus serrulatus). Toxicon 2013, 74, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Dickey, D.M.; Potter, L.R. Dendroaspis natriuretic peptide and the designer natriuretic peptide, CD-NP, are resistant to proteolytic inactivation. J. Mol. Cell. Cardiol. 2011, 51, 67–71. [Google Scholar] [CrossRef] [PubMed]
- Ho, P.L.; Soares, M.B.; Maack, T.; Gimenez, I.; Puorto, G.; Furtado, M.F.; Raw, I. Cloning of an unusual natriuretic peptide from the South American coral snake Micrurus corallinus. Eur. J. Biochem. 1997, 250, 144–149. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.-C. Dendroaspis natriuretic peptide is the most potent natriuretic peptide to cause relaxation of lower esophageal sphincter. Regul. Pept. 2011, 167, 246–249. [Google Scholar] [CrossRef] [PubMed]
- Lisy, O.; Huntley, B.K.; McCormick, D.J.; Kurlansky, P.A.; Burnett, J.C., Jr. Design, synthesis, and actions of a novel chimeric natriuretic peptide: CD-NP. J. Am. Coll. Cardiol. 2008, 52, 60–68. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, S.L.; Dias-Junior, C.A.; Baldasso, P.A.; Damico, D.C.S.; Carvalho, B.M.A.; Garanto, A.; Acosta, G.; Oliveira, E.; Albericio, F.; Soares, A.M.; et al. Vascular effects and electrolyte homeostasis of the natriuretic peptide isolated from Crotalus oreganus abyssus (North American Grand Canyon rattlesnake) venom. Peptides 2012, 36, 206–212. [Google Scholar] [CrossRef] [PubMed]
- Reeks, T.; Jones, A.; Brust, A.; Sridharan, S.; Corcilius, L.; Wilkinson, B.L.; Thaysen-Andersen, M.; Payne, R.J.; Kini, R.M.; Daly, N.L.; et al. A defined alpha-helix in the bifunctional O-glycosylated natriuretic peptide TcNPa from the venom of Tropidechis carinatus. Angew. Chem. Int. Ed. Engl. 2015, 54, 4828–4831. [Google Scholar] [CrossRef] [PubMed]
- Ichiki, T.; Dzhoyashvili, N.; Burnett, J.C. Natriuretic peptide based therapeutics for heart failure: Cenderitide: A novel first-in-class designer natriuretic peptide. Int. J. Cardiol. 2019, 281, 166–171. [Google Scholar] [CrossRef]
- Tourki, B.; Matéo, P.; Morand, J.; Elayeb, M.; Godin-Ribuot, D.; Marrakchi, N.; Belaidi, E.; Messadi, E. Lebetin 2, a Snake Venom-Derived Natriuretic Peptide, Attenuates Acute Myocardial Ischemic Injury through the Modulation of Mitochondrial Permeability Transition Pore at the Time of Reperfusion. PLoS ONE 2016, 11, e0162632. [Google Scholar] [CrossRef]
- Vink, S.; Jin, A.H.; Poth, K.J.; Head, G.A.; Alewood, P.F. Natriuretic peptide drug leads from snake venom. Toxicon Off. J. Int. Soc. Toxinol. 2012, 59, 434–445. [Google Scholar] [CrossRef]
- Lee, C.Y.; Chen, H.H.; Lisy, O.; Swan, S.; Cannon, C.; Lieu, H.D.; Burnett, J.C., Jr. Pharmacodynamics of a novel designer natriuretic peptide, CD-NP, in a first-in-human clinical trial in healthy subjects. J. Clin. Pharmacol. 2009, 49, 668–673. [Google Scholar] [CrossRef] [PubMed]
- Schweitz, H.; Vigne, P.; Moinier, D.; Frelin, C.; Lazdunski, M. A new member of the natriuretic peptide family is present in the venom of the green mamba (Dendroaspis angusticeps). J. Biol. Chem. 1992, 267, 13928–13932. [Google Scholar] [CrossRef] [PubMed]
- Silva, S.L.d.; Almeida, J.R.; Resende, L.M.H.; Martins, W.K.; Henriques, F.F.; Baldasso, P.A.; Soares, A.M.; Taranto, A.G.; Resende, R.R.; Marangoni, S.; et al. Isolation and Characterization of a Natriuretic Peptide from Crotalus oreganus abyssus (Grand Canyon Rattlesnake) and its Effects on Systemic Blood Pressure and Nitrite Levels. Int. J. Pept. Res. Ther. 2011, 17, 165–173. [Google Scholar] [CrossRef]
- Gibbs, H.L.; Mackessy, S.P. Functional basis of a molecular adaptation: Prey-specific toxic effects of venom from Sistrurus rattlesnakes. Toxicon 2009, 53, 672–679. [Google Scholar] [CrossRef] [PubMed]
- Bian, L.-j.; Wu, P.; Yang, X.-y. Two-step chromatographic method for separation and purification of nerve growth factor from venom of Chinese cobra. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2004, 805, 119–125. [Google Scholar] [CrossRef]
- Gu, L.; Shen, Y.; Xu, S.; Shu, Y.; Jiang, T.; Lin, Z. Crystallization and preliminary X-ray diffraction studies of cobra venom beta-nerve growth factor. Biochem. Biophys. Res. Commun. 2002, 297, 1008–1010. [Google Scholar] [CrossRef]
- Mariam, K.; Tu, A.T. Extremely low nerve growth facior (NGF) activity of sea snake (Hydrophiidae) venoms. J. Nat. Toxins 2002, 11, 393–398. [Google Scholar]
- Osipov, A.V.; Terpinskaya, T.I.; Kuznetsova, T.E.; Ryzhkovskaya, E.L.; Lukashevich, V.S.; Rudnichenko, J.A.; Ulashchyk, V.S.; Starkov, V.G.; Utkin, Y.N. Cobra Venom Factor and Ketoprofen Abolish the Antitumor Effect of Nerve Growth Factor from Cobra Venom. Toxins 2017, 9, 274. [Google Scholar] [CrossRef]
- Perez-Polo, J.R.; Bomar, H.; Beck, C.; Hall, K. Nerve growth factor from Crotalus adamenteus snake venom. J. Biol. Chem. 1978, 253, 6140–6148. [Google Scholar] [CrossRef]
- Sunagar, K.; Fry, B.G.; Jackson, T.N.W.; Casewell, N.R.; Undheim, E.A.B.; Vidal, N.; Ali, S.A.; King, G.F.; Vasudevan, K.; Vasconcelos, V.; et al. Molecular Evolution of Vertebrate Neurotrophins: Co-Option of the Highly Conserved Nerve Growth Factor Gene into the Advanced Snake Venom Arsenalf. PLoS ONE 2013, 8, e81827. [Google Scholar] [CrossRef]
- Lu, Z.; Lei, D.; Jiang, T.; Yang, L.; Zheng, L.; Zhao, J. Nerve growth factor from Chinese cobra venom stimulates chondrogenic differentiation of mesenchymal stem cells. Cell Death Dis. 2017, 8, e2801. [Google Scholar] [CrossRef]
- Osipov, A.V.; Terpinskaya, T.I.; Ulaschik, V.S.; Tsetlin, V.I.; Utkin, Y.N. Nerve growth factor suppresses Ehrlich carcinoma growth. Dokl. Biochem. Biophys. 2013, 451, 207–208. [Google Scholar] [CrossRef] [PubMed]
- Komatsu, K.; Hasegawa, H.; Honda, T.; Yabashi, A.; Kawasaki, T. Nerve Growth Factor in Saliva Stimulated by Mastication. Oral. Sci. Int. 2008, 5, 78–84. [Google Scholar] [CrossRef]
- Wijeyewickrema, L.C.; Gardiner, E.E.; Gladigau, E.L.; Berndt, M.C.; Andrews, R.K. Nerve growth factor inhibits metalloproteinase-disintegrins and blocks ectodomain shedding of platelet glycoprotein VI. J. Biol. Chem. 2010, 285, 11793–11799. [Google Scholar] [CrossRef] [PubMed]
- Kerkkamp, H.M.; Kini, R.M.; Pospelov, A.S.; Vonk, F.J.; Henkel, C.V.; Richardson, M.K. Snake Genome Sequencing: Results and Future Prospects. Toxins 2016, 8, 360. [Google Scholar] [CrossRef]
- Kostiza, T.; Meier, J. Nerve growth factors from snake venoms: Chemical properties, mode of action and biological significance. Toxicon Off. J. Int. Soc. Toxinology 1996, 34, 787–806. [Google Scholar] [CrossRef]
- Boldrini-França, J.; Cologna, C.T.; Pucca, M.B.; Bordon, K.d.C.F.; Amorim, F.G.; Anjolette, F.A.P.; Cordeiro, F.A.; Wiezel, G.A.; Cerni, F.A.; Pinheiro-Junior, E.L.; et al. Minor snake venom proteins: Structure, function and potential applications. Biochim. Biophys. Acta (BBA)—Gen. Subj. 2017, 1861, 824–838. [Google Scholar] [CrossRef]
- Kawamoto, K.; Matsuda, H. Nerve growth factor and wound healing. Prog. Brain Res. 2004, 146, 369–384. [Google Scholar] [CrossRef]
- Villoslada, P.; Genain, C.P. Role of nerve growth factor and other trophic factors in brain inflammation. Prog. Brain Res. 2004, 146, 403–414. [Google Scholar] [CrossRef]
- Reyes-Velasco, J.; Meik, J.M.; Smith, E.N.; Castoe, T.A. Phylogenetic relationships of the enigmatic longtailed rattlesnakes (Crotalus ericsmithi, C. lannomi, and C. stejnegeri). Mol. Phylogenet Evol. 2013, 69, 524–534. [Google Scholar] [CrossRef]
- Katzir, I.; Shani, J.; Goshen, G.; Sela, J.; Ninary, E.; Dogonovski, A.M.; Shabashov, D.; Inoue, S.; Ikeda, K.; Hayashi, K.; et al. Characterization of nerve growth factors (NGFs) from snake venoms by use of a novel, quantitative bioassay utilizing pheochromocytoma (PC12) cells overexpressing human trkA receptors. Toxicon 2003, 42, 481–490. [Google Scholar] [CrossRef] [PubMed]
- Akiyama, N.; Hatori, Y.; Takashiro, Y.; Hirabayashi, T.; Saito, T.; Murayama, T. Nerve growth factor-induced up-regulation of cytosolic phospholipase A2alpha level in rat PC12 cells. Neurosci. Lett. 2004, 365, 218–222. [Google Scholar] [CrossRef]
- Tong, Q.; Wang, F.; Zhou, H.-Z.; Sun, H.-L.; Song, H.; Shu, Y.-Y.; Gong, Y.; Zhang, W.-T.; Cai, T.-X.; Yang, F.-Q.; et al. Structural and functional insights into lipid-bound nerve growth factors. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2012, 26, 3811–3821. [Google Scholar] [CrossRef] [PubMed]
- Osipov, A.V.; Terpinskaya, T.I.; Kryukova, E.V.; Ulaschik, V.S.; Paulovets, L.V.; Petrova, E.A.; Blagun, E.V.; Starkov, V.G.; Utkin, Y.N. Nerve Growth Factor from Cobra Venom Inhibits the Growth of Ehrlich Tumor in Mice. Toxins 2014, 6, 784–795. [Google Scholar] [CrossRef] [PubMed]
- Chiarenza, A.; Lazarovici, P.; Lempereur, L.; Cantarella, G.; Bianchi, A.; Bernardini, R. Tamoxifen Inhibits Nerve Growth Factor-induced Proliferation of the Human Breast Cancerous Cell Line MCF-7. Cancer Res. 2001, 61, 3002–3008. [Google Scholar] [PubMed]
- Hampel, H.; Mesulam, M.M.; Cuello, A.C.; Farlow, M.R.; Giacobini, E.; Grossberg, G.T.; Khachaturian, A.S.; Vergallo, A.; Cavedo, E.; Snyder, P.J.; et al. The cholinergic system in the pathophysiology and treatment of Alzheimer’s disease. Brain 2018, 141, 1917–1933. [Google Scholar] [CrossRef] [PubMed]
- Trummal, K.; Tõnismägi, K.; Paalme, V.; Järvekülg, L.; Siigur, J.; Siigur, E. Molecular diversity of snake venom nerve growth factors. Toxicon 2011, 58, 363–368. [Google Scholar] [CrossRef]
- McCleary, R.J.R.; Kini, R.M. Non-enzymatic proteins from snake venoms: A gold mine of pharmacological tools and drug leads. Toxicon 2013, 62, 56–74. [Google Scholar] [CrossRef]
- Munawar, A.; Ali, S.A.; Akrem, A.; Betzel, C. Snake Venom Peptides: Tools of Biodiscovery. Toxins 2018, 10, 474. [Google Scholar] [CrossRef]
- Millers, E.K.; Johnson, L.A.; Birrell, G.W.; Masci, P.P.; Lavin, M.F.; de Jersey, J.; Guddat, L.W. The structure of human microplasmin in complex with textilinin-1, an aprotinin-like inhibitor from the Australian brown snake. PLoS ONE 2013, 8, e54104. [Google Scholar] [CrossRef]
- Zupunski, V.; Kordis, D.; Gubensek, F. Adaptive evolution in the snake venom Kunitz/BPTI protein family. FEBS Lett. 2003, 547, 131–136. [Google Scholar] [CrossRef] [PubMed]
- Millers, E.K.; Trabi, M.; Masci, P.P.; Lavin, M.F.; de Jersey, J.; Guddat, L.W. Crystal structure of textilinin-1, a Kunitz-type serine protease inhibitor from the venom of the Australian common brown snake (Pseudonaja textilis). FEBS J. 2009, 276, 3163–3175. [Google Scholar] [CrossRef] [PubMed]
- Earl, S.T.H.; Richards, R.; Johnson, L.A.; Flight, S.; Anderson, S.; Liao, A.; de Jersey, J.; Masci, P.P.; Lavin, M.F. Identification and characterisation of Kunitz-type plasma kallikrein inhibitors unique to Oxyuranus sp. snake venoms. Biochimie 2012, 94, 365–373. [Google Scholar] [CrossRef] [PubMed]
- Vivas, J.; Ibarra, C.; Salazar, A.M.; Neves-Ferreira, A.G.; Sánchez, E.E.; Perales, J.; Rodríguez-Acosta, A.; Guerrero, B. Purification and characterization of tenerplasminin-1, a serine peptidase inhibitor with antiplasmin activity from the coral snake (Micrurus tener tener) venom. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2016, 179, 107–115. [Google Scholar] [CrossRef] [PubMed]
- St Pierre, L.; Earl, S.T.; Filippovich, I.; Sorokina, N.; Masci, P.P.; De Jersey, J.; Lavin, M.F. Common evolution of waprin and kunitz-like toxin families in Australian venomous snakes. Cell. Mol. Life Sci. 2008, 65, 4039–4054. [Google Scholar] [CrossRef] [PubMed]
- Pahari, S.; Mackessy, S.P.; Kini, R.M. The venom gland transcriptome of the Desert Massasauga rattlesnake (Sistrurus catenatus edwardsii): Towards an understanding of venom composition among advanced snakes (Superfamily Colubroidea). BMC Mol. Biol. 2007, 8, 115. [Google Scholar] [CrossRef] [PubMed]
- Torres, A.M.; Wong, H.Y.; Desai, M.; Moochhala, S.; Kuchel, P.W.; Kini, R.M. Identification of a novel family of proteins in snake venoms. Purification and structural characterization of nawaprin from Naja nigricollis snake venom. J. Biol. Chem. 2003, 278, 40097–40104. [Google Scholar] [CrossRef] [PubMed]
- Nair, D.G.; Fry, B.G.; Alewood, P.; Kumar, P.P.; Kini, R.M. Antimicrobial activity of omwaprin, a new member of the waprin family of snake venom proteins. Biochem. J. 2007, 402, 93–104. [Google Scholar] [CrossRef]
- Oguiura, N.; Sanches, L.; Duarte, P.V.; Sulca-López, M.A.; Machini, M.T. Past, Present, and Future of Naturally Occurring Antimicrobials Related to Snake Venoms. Animals 2023, 13, 744. [Google Scholar] [CrossRef]
- Takeya, H.; Onikura, A.; Nikai, T.; Sugihara, H.; Iwanaga, S. Primary structure of a hemorrhagic metalloproteinase, HT-2, isolated from the venom of Crotalus ruber ruber. J. Biochem. 1990, 108, 711–719. [Google Scholar] [CrossRef]
- Takeda, S. ADAM and ADAMTS Family Proteins and Snake Venom Metalloproteinases: A Structural Overview. Toxins 2016, 8, 155. [Google Scholar] [CrossRef] [PubMed]
- Escalante, T.; Rucavado, A.; Fox, J.W.; Gutiérrez, J.M. Key events in microvascular damage induced by snake venom hemorrhagic metalloproteinases. J. Proteom. 2011, 74, 1781–1794. [Google Scholar] [CrossRef]
- Kikushima, E.; Nakamura, S.; Oshima, Y.; Shibuya, T.; Miao, J.Y.; Hayashi, H.; Nikai, T.; Araki, S. Hemorrhagic activity of the vascular apoptosis-inducing proteins VAP1 and VAP2 from Crotalus atrox. Toxicon Off. J. Int. Soc. Toxinol. 2008, 52, 589–593. [Google Scholar] [CrossRef]
- Bjarnason, J.B.; Fox, J.W. Proteolytic specificity and cobalt exchange of hemorrhagic toxin e, a zinc protease isolated from the venom of the western diamondback rattlesnake (Crotalus atrox). Biochemistry 1983, 22, 3770–3778. [Google Scholar] [CrossRef] [PubMed]
- Bjarnason, J.B.; Fox, J.W. Hemorrhagic metalloproteinases from snake venoms. Pharmacol. Ther. 1994, 62, 325–372. [Google Scholar] [CrossRef] [PubMed]
- Ferraz, C.R.; Arrahman, A.; Xie, C.F.; Casewell, N.R.; Lewis, R.J.; Kool, J.; Cardoso, F.C. Multifunctional Toxins in Snake Venoms and Therapeutic Implications: From Pain to Hemorrhage and Necrosis. Front. Ecol. Evol. 2019, 7, 218. [Google Scholar] [CrossRef]
- Takeda, S.; Takeya, H.; Iwanaga, S. Snake venom metalloproteinases: Structure, function and relevance to the mammalian ADAM/ADAMTS family proteins. Biochim. Biophys. Acta (BBA)—Proteins Proteom. 2012, 1824, 164–176. [Google Scholar] [CrossRef] [PubMed]
- da Silva, I.R.F.; Lorenzetti, R.; Rennó, A.L.; Baldissera, L.; Zelanis, A.; Serrano, S.M.d.T.; Hyslop, S. BJ-PI2, A non-hemorrhagic metalloproteinase from Bothrops jararaca snake venom. Biochim. Biophys. Acta (BBA)—Gen. Subj. 2012, 1820, 1809–1821. [Google Scholar] [CrossRef]
- Gomis-Rüth, F.X.; Kress, L.F.; Kellermann, J.; Mayr, I.; Lee, X.; Huber, R.; Bode, W. Refined 2·0 Å X-ray Crystal Structure of the Snake Venom Zinc-endopeptidase Adamalysin II: Primary and Tertiary Structure Determination, Refinement, Molecular Structure and Comparison with Astacin, Collagenase and Thermolysin. J. Mol. Biol. 1994, 239, 513–544. [Google Scholar] [CrossRef]
- Igarashi, T.; Araki, S.; Mori, H.; Takeda, S. Crystal structures of catrocollastatin/VAP2B reveal a dynamic, modular architecture of ADAM/adamalysin/reprolysin family proteins. FEBS Lett. 2007, 581, 2416–2422. [Google Scholar] [CrossRef]
- Wallnoefer, H.G.; Lingott, T.; Gutierrez, J.M.; Merfort, I.; Liedl, K.R. Backbone flexibility controls the activity and specificity of a protein-protein interface: Specificity in snake venom metalloproteases. J. Am. Chem. Soc. 2010, 132, 10330–10337. [Google Scholar] [CrossRef] [PubMed]
- Chapeaurouge, A.; Reza, M.A.; Mackessy, S.P.; Carvalho, P.C.; Valente, R.H.; Teixeira-Ferreira, A.; Perales, J.; Lin, Q.; Kini, R.M. Interrogating the Venom of the Viperid Snake Sistrurus catenatus edwardsii by a Combined Approach of Electrospray and MALDI Mass Spectrometry. PLoS ONE 2015, 10, e0092091. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G.; Scheib, H.; van der Weerd, L.; Young, B.; McNaughtan, J.; Ramjan, S.F.R.; Vidal, N.; Poelmann, R.E.; Norman, J.A. Evolution of an arsenal: Structural and functional diversification of the venom system in the advanced snakes (Caenophidia). Mol. Cell. Proteom. 2008, 7, 215–246. [Google Scholar] [CrossRef] [PubMed]
- Urra, F.A.; Pulgar, R.; Gutierrez, R.; Hodar, C.; Cambiazo, V.; Labra, A. Identification and molecular characterization of five putative toxins from the venom gland of the snake Philodryas chamissonis (Serpentes: Dipsadidae). Toxicon 2015, 108, 19–31. [Google Scholar] [CrossRef] [PubMed]
- Polgár, L. On the mechanism of proton transfer in the catalysis by serine proteases. J. Theor. Biol. 1971, 31, 165–169. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, S.A.; Chen, Y.; Müller, J.D.; Gratton, E.; Hazlett, T.L. Solution and interface aggregation states of Crotalus atrox venom phospholipase A2 by two-photon excitation fluorescence correlation spectroscopy. Biochemistry 2001, 40, 6903–6911. [Google Scholar] [CrossRef]
- Rodrigues, C.R.; Molina, D.A.M.; de Souza, D.L.N.; Cardenas, J.; Costal-Oliveira, F.; Guerra-Duarte, C.; Chávez-Olórtegui, C. Biological and proteomic characterization of the venom from Peruvian Andes rattlesnake Crotalus durissus. Toxicon 2022, 207, 31–42. [Google Scholar] [CrossRef]
- Quach, N.D.; Arnold, R.D.; Cummings, B.S. Secretory phospholipase A2 enzymes as pharmacological targets for treatment of disease. Biochem. Pharmacol. 2014, 90, 338–348. [Google Scholar] [CrossRef]
- Almeida, J.R.; Mendes, B.; Lancellotti, M.; Marangoni, S.; Vale, N.; Passos, O.; Ramos, M.J.; Fernandes, P.A.; Gomes, P.; Da Silva, S.L. A novel synthetic peptide inspired on Lys49 phospholipase A2 from Crotalus oreganus abyssus snake venom active against multidrug-resistant clinical isolates. Eur. J. Med. Chem. 2018, 149, 248–256. [Google Scholar] [CrossRef]
- Hendon, R.A.; Fraenkel-Conrat, H. Biological roles of the two components of crotoxin. Proc. Natl. Acad. Sci. USA 1971, 68, 1560–1563. [Google Scholar] [CrossRef]
- Cavalcante, W.L.; Ponce-Soto, L.A.; Marangoni, S.; Gallacci, M. Neuromuscular effects of venoms and crotoxin-like proteins from Crotalus durissus ruruima and Crotalus durissus cumanensis. Toxicon 2015, 96, 46–49. [Google Scholar] [CrossRef] [PubMed]
- Cavalcante, W.L.G.; Noronha-Matos, J.B.; Timoteo, M.A.; Fontes, M.R.M.; Gallacci, M.; Correia-de-Sa, P. Neuromuscular paralysis by the basic phospholipase A2 subunit of crotoxin from Crotalus durissus terrificus snake venom needs its acid chaperone to concurrently inhibit acetylcholine release and produce muscle blockage. Toxicol. Appl. Pharmacol. 2017, 334, 8–17. [Google Scholar] [CrossRef] [PubMed]
- Russo, R.R.; Dos Santos Junior, N.N.; Cintra, A.C.O.; Figueiredo, L.T.M.; Sampaio, S.V.; Aquino, V.H. Expression, purification and virucidal activity of two recombinant isoforms of phospholipase A2 from Crotalus durissus terrificus venom. Arch. Virol. 2019, 164, 1159–1171. [Google Scholar] [CrossRef] [PubMed]
- Terra, A.L.C.; Moreira-Dill, L.; Simões-Silva, R.; Monteiro, J.R.N.; Cavalcante, W.L.G.; Gallacci, M.; Barros, N.B.; Nicolete, R.; Teles, C.B.G.; Medeiros, P.S.M.; et al. Biological characterization of the Amazon coral Micrurus spixii snake venom: Isolation of a new neurotoxic phospholipase A2. Toxicon 2015, 103, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G. Carbon monoxide inhibits the anticoagulant activity of phospholipase A2 purified from Crotalus adamanteus venom. J. Thromb. Thrombolysis 2019, 47, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Melendez-Martinez, D.; Macias-Rodriguez, E.; Vazquez-Briones, R.; Lopez-Vera, E.; Sandra Cruz-Perez, M.; Vargas-Caraveo, A.; Gatica-Colima, A.; Fernando Plenge-Tellechea, L. In vitro hemotoxic, alpha-neurotoxic and vasculotoxic effects of the Mexican black-tailed rattlesnake (Crotalus molossus nigrescens) venom. J. Venom. Res. 2017, 8, 1–8. [Google Scholar] [PubMed]
- Moore, S.W.; Bhat, V.K.; Flatt, P.R.; Gault, V.A.; McClean, S. Isolation and characterisation of insulin-releasing compounds from Crotalus adamanteus, Crotalus vegrandis and Bitis nasicornis venom. Toxicon 2015, 101, 48–54. [Google Scholar] [CrossRef] [PubMed]
- Peigneur, S.; Orts, D.J.; Prieto da Silva, A.R.; Oguiura, N.; Boni-Mitake, M.; de Oliveira, E.B.; Zaharenko, A.J.; de Freitas, J.C.; Tytgat, J. Crotamine pharmacology revisited: Novel insights based on the inhibition of KV channels. Mol. Pharmacol. 2012, 82, 90–96. [Google Scholar] [CrossRef]
- Yamane, E.S.; Bizerra, F.C.; Oliveira, E.B.; Moreira, J.; Rajabi, M.; Nunes, G.L.C.; de Souza, A.O.; da Silva, I.D.C.G.; Yamane, T.; Karpel, R.L.; et al. Unraveling the antifungal activity of a South American rattlesnake toxin crotamine. Biochimie 2013, 95, 231–240. [Google Scholar] [CrossRef]
- Whittington, C.M.; Papenfuss, A.T.; Bansal, P.; Torres, A.M.; Wong, E.S.W.; Deakin, J.E.; Graves, T.; Alsop, A.; Schatzkamer, K.; Kremitzki, C.; et al. Defensins and the convergent evolution of platypus and reptile venom genes. Genome Res. 2008, 18, 986–994. [Google Scholar] [CrossRef]
- Bober, M.A.; Glenn, J.L.; Straight, R.C.; Ownby, C.L. Detection of myotoxin alpha-like proteins in various snake venoms. Toxicon 1988, 26, 665–673. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, E.E.; Gonzalez, R.; Lucena, S.; Garcia, S.; Finol, H.J.; Suntravat, M.; Giron, M.E.; Fernandez, I.; Rodriguez-Acosta, A. Crotamine-like from Southern Pacific rattlesnake (Crotalus oreganus helleri) Venom acts on human leukemia (K-562) cell lines and produces ultrastructural changes on mice adrenal gland. Ultrastruct. Pathol. 2018, 42, 116–123. [Google Scholar] [CrossRef] [PubMed]
- Laure, C.J. Die primärstruktur des crotamins. Hoppe-Seyler’s Z. Physiol. Chem. 1975, 356, 213–215. [Google Scholar]
- de Oliveira, S.A.M.; Magalhães, M.R.; Salazar, V.C.R.; Valadares, M.C.; da Cunha, L.C. Identification of crotamine in the venom of Crotalus durissus collilineatus by three different methods. Toxicon 2015, 95, 46–51. [Google Scholar] [CrossRef]
- Correa, P.G.; Oguiura, N. Phylogenetic analysis of β-defensin-like genes of Bothrops, Crotalus and Lachesis snakes. Toxicon 2013, 69, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Norris, J.W.; Fry, R.M.; Tu, A.T. The nucleotide sequence of the translated and untranslated regions of a cDNA for myotoxin a from the venom of prairie rattlesnake (Crotalus viridis viridis). Biochem. Biophys. Res. Commun. 1997, 230, 607–610. [Google Scholar] [CrossRef] [PubMed]
- Suntravat, M.; Langlais, P.R.; Sánchez, E.E.; Nielsen, V.G. CatroxMP-II: A heme-modulated fibrinogenolytic metalloproteinase isolated from Crotalus atrox venom. Biomet. Int. J. Role Met. Ions Biol. Biochem. Med. 2018, 31, 585–593. [Google Scholar] [CrossRef] [PubMed]
- Santos, W.S.; Montoni, F.; Eichler, R.A.S.; Arcos, S.S.S.; Andreotti, D.Z.; Kisaki, C.Y.; Evangelista, K.B.; Calacina, H.M.; Lima, I.F.; Soares, M.A.M.; et al. Proteomic analysis reveals rattlesnake venom modulation of proteins associated with cardiac tissue damage in mouse hearts. J. Proteom. 2022, 258, 104530. [Google Scholar] [CrossRef]
- Rodrigues, M.; Santos, A.; Santos, N.C.; de la Torre, B.G.; Rádis-Baptista, G.; Andreu, D. Molecular characterization of the interaction of crotamine-derived nucleolar targeting peptides with lipid membranes. Biochim. Biophys. Acta 2012, 1818, 2707–2717. [Google Scholar] [CrossRef][Green Version]
- Oguiura, N.; Boni-Mitake, M.; Radis-Baptista, G. New view on crotamine, a small basic polypeptide myotoxin from South American rattlesnake venom. Toxicon 2005, 46, 363–370. [Google Scholar] [CrossRef]
- Utaisincharoen, P.; Baker, B.; Tu, A.T. Binding of myotoxin a to sarcoplasmic reticulum calcium-ATPase: A structural study. Biochemistry 1991, 30, 8211–8216. [Google Scholar] [CrossRef] [PubMed]
- Fadel, V.; Bettendorff, P.; Herrmann, T.; de Azevedo, W.F., Jr.; Oliveira, E.B.; Yamane, T.; Wuthrich, K. Automated NMR structure determination and disulfide bond identification of the myotoxin crotamine from Crotalus durissus terrificus. Toxicon 2005, 46, 759–767. [Google Scholar] [CrossRef] [PubMed]
- Pereira, A.; Kerkis, A.; Hayashi, M.A.; Pereira, A.S.; Silva, F.S.; Oliveira, E.B.; Prieto da Silva, A.R.; Yamane, T.; Rádis-Baptista, G.; Kerkis, I. Crotamine toxicity and efficacy in mouse models of melanoma. Expert. Opin. Investig. Drugs 2011, 20, 1189–1200. [Google Scholar] [CrossRef] [PubMed]
- Mambelli-Lisboa, N.C.; Sciani, J.M.; Brandão Prieto da Silva, A.R.; Kerkis, I. Co-Localization of Crotamine with Internal Membranes and Accentuated Accumulation in Tumor Cells. Molecules 2018, 23, 968. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira, S.A.; Magalhaes, M.R.; de Oliveira, L.P.; da Cunha, L.C. Identification of antinociceptive fraction of snake venom from Crotalus durissus collilineatus crotamine-negative and its acute toxicity evaluation. Toxicon 2016, 122, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Teixeira-Araújo, R.; Castanheira, P.; Brazil-Más, L.; Pontes, F.; Leitão de Araújo, M.; Machado Alves, M.L.; Zingali, R.B.; Correa-Netto, C. Antivenomics as a tool to improve the neutralizing capacity of the crotalic antivenom: A study with crotamine. J. Venom. Anim. Toxins Incl. Trop. Dis. 2017, 23, 28. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kerkis, I.; Silva, F.D.; Pereira, A.; Kerkis, A.; Rádis-Baptista, G. Biological versatility of crotamine—A cationic peptide from the venom of a South American rattlesnake. Expert. Opin. Investig. Drugs 2010, 19, 1515–1525. [Google Scholar] [CrossRef]
- Gibbs, H.L.; Sanz, L.; Calvete, J.J. Snake population venomics: Proteomics-based analyses of individual variation reveals significant gene regulation effects on venom protein expression in Sistrurus rattlesnakes. J. Mol. Evol. 2009, 68, 113–125. [Google Scholar] [CrossRef]
- Gómez, A.; Solano, G.; Chang-Castillo, A.; Chacón, D.; Corrales, G.; Segura, A.; Estrada, R.; León, G. Intraspecific variability of the Central American rattlesnake (Crotalus simus) venom and its usefulness to obtain a representative standard venom. Toxicon 2021, 202, 20–26. [Google Scholar] [CrossRef]
- Munawar, A.; Trusch, M.; Georgieva, D.; Hildebrand, D.; Kwiatkowski, M.; Behnken, H.; Harder, S.; Arni, R.; Spencer, P.; Schluter, H.; et al. Elapid snake venom analyses show the specificity of the peptide composition at the level of genera Naja and Notechis. Toxins 2014, 6, 850–868. [Google Scholar] [CrossRef]
- Aloe, L.; Rocco, M.L.; Balzamino, B.O.; Micera, A. Nerve Growth Factor: A Focus on Neuroscience and Therapy. Curr. Neuropharmacol. 2015, 13, 294–303. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, M.P.; Mantelli, F.; Sacchetti, M.; Antonangeli, M.I.; Cattani, F.; D’Anniballe, G.; Sinigaglia, F.; Ruffini, P.A.; Lambiase, A. Safety and Pharmacokinetics of Escalating Doses of Human Recombinant Nerve Growth Factor Eye Drops in a Double-Masked, Randomized Clinical Trial. Biodrugs 2014, 28, 275–283. [Google Scholar] [CrossRef]
- Sacchetti, M.; Lambiase, A.; Schmidl, D.; Schmetterer, L.; Ferrari, M.; Mantelli, F.; Allegretti, M.; Garhoefer, G. Effect of recombinant human nerve growth factor eye drops in patients with dry eye: A phase IIa, open label, multiple-dose study. Br. J. Ophthalmol. 2020, 104, 127–135. [Google Scholar] [CrossRef] [PubMed]
- Beykin, G.; Stell, L.; Halim, M.S.; Nuñez, M.; Popova, L.; Nguyen, B.T.; Groth, S.L.; Dennis, A.; Li, Z.Q.; Atkins, M.; et al. Phase 1b Randomized Controlled Study of Short Course Topical Recombinant Human Nerve Growth Factor (Rhngf) for Neuroenhancement in Glaucoma: Safety, Tolerability, and Efficacy Measure Outcomes. Am. J. Ophthalmol. 2022, 234, 223–234. [Google Scholar] [CrossRef] [PubMed]
- Tuszynski, M.H.; Thal, L.; Pay, M.; Salmon, D.P.; Sang, U.H.; Bakay, R.; Patel, P.; Blesch, A.; Vahlsing, H.L.; Ho, G.; et al. A phase 1 clinical trial of nerve growth factor gene therapy for Alzheimer disease. Nat. Med. 2005, 11, 551–555. [Google Scholar] [CrossRef] [PubMed]
- Tuszynski, M.H.; Yang, J.H.; Barba, D.; Hoi-Sang, U.; Bakay, R.A.E.; Pay, M.M.; Masliah, E.; Conner, J.M.; Kobalka, P.; Roy, S.; et al. Nerve Growth Factor Gene Therapy Activation of Neuronal Responses in Alzheimer Disease. JAMA Neurol. 2015, 72, 1139–1147. [Google Scholar] [CrossRef] [PubMed]
- Castle, M.J.; Baltanas, F.C.; Kovacs, I.; Nagahara, A.H.; Barba, D.; Tuszynski, M.H. Postmortem Analysis in a Clinical Trial of AAV2-NGF Gene Therapy for Alzheimer’s Disease Identifies a Need for Improved Vector Delivery. Hum. Gene Ther. 2020, 31, 415–422. [Google Scholar] [CrossRef]
- Rafii, M.S.; Tuszynski, M.H.; Thomas, R.G.; Barba, D.; Brewer, J.B.; Rissman, R.A.; Siffert, J.; Aisen, P.S.; Team, A.N.S. Adeno-Associated Viral Vector (Serotype 2)-Nerve Growth Factor for Patients With Alzheimer Disease A Randomized Clinical Trial. JAMA Neurol. 2018, 75, 834–841. [Google Scholar] [CrossRef]
- Muller, V.D.; Soares, R.O.; dos Santos, N.N., Jr.; Trabuco, A.C.; Cintra, A.C.; Figueiredo, L.T.; Caliri, A.; Sampaio, S.V.; Aquino, V.H. Phospholipase A2 isolated from the venom of Crotalus durissus terrificus inactivates dengue virus and other enveloped viruses by disrupting the viral envelope. PLoS ONE 2014, 9, e112351. [Google Scholar] [CrossRef]
- Oliveira, A.L.; Viegas, M.F.; da Silva, S.L.; Soares, A.M.; Ramos, M.J.; Fernandes, P.A. The chemistry of snake venom and its medicinal potential. Nat. Rev. Chem. 2022, 6, 451–469. [Google Scholar] [CrossRef]
- Querobino, S.M.; Ribeiro, C.A.J.; Alberto-Silva, C. Bradykinin-potentiating PEPTIDE-10C, an argininosuccinate synthetase activator, protects against H2O2-induced oxidative stress in SH-SY5Y neuroblastoma cells. Peptides 2018, 103, 90–97. [Google Scholar] [CrossRef] [PubMed]
- Averin, A.S.; Utkin, Y.N. Cardiovascular Effects of Snake Toxins: Cardiotoxicity and Cardioprotection. Acta Naturae 2021, 13, 4–14. [Google Scholar] [CrossRef] [PubMed]
- Estevão-Costa, M.I.; Sanz-Soler, R.; Johanningmeier, B.; Eble, J.A. Snake venom components in medicine: From the symbolic rod of Asclepius to tangible medical research and application. Int. J. Biochem. Cell Biol. 2018, 104, 94–113. [Google Scholar] [CrossRef] [PubMed]
- Gan, Z.R.; Gould, R.J.; Jacobs, J.W.; Friedman, P.A.; Polokoff, M.A. Echistatin. A potent platelet aggregation inhibitor from the venom of the viper, Echis carinatus. J. Biol. Chem. 1988, 263, 19827–19832. [Google Scholar] [CrossRef] [PubMed]
- Garsky, V.M.; Lumma, P.K.; Freidinger, R.F.; Pitzenberger, S.M.; Randall, W.C.; Veber, D.F.; Gould, R.J.; Friedman, P.A. Chemical synthesis of echistatin, a potent inhibitor of platelet aggregation from Echis carinatus: Synthesis and biological activity of selected analogs. Proc. Natl. Acad. Sci. USA 1989, 86, 4022–4026. [Google Scholar] [CrossRef] [PubMed]
- Lazarovici, P.; Marcinkiewicz, C.; Lelkes, P.I. From Snake Venom’s Disintegrins and C-Type Lectins to Anti-Platelet Drugs. Toxins 2019, 11, 303. [Google Scholar] [CrossRef] [PubMed]
- Hartman, G.D.; Egbertson, M.S.; Halczenko, W.; Laswell, W.L.; Duggan, M.E.; Smith, R.L.; Naylor, A.M.; Manno, P.D.; Lynch, R.J.; Zhang, G. Non-peptide fibrinogen receptor antagonists. 1. Discovery and design of exosite inhibitors. J. Med. Chem. 1992, 35, 4640–4642. [Google Scholar] [CrossRef] [PubMed]
- Abd El-Aziz, T.M.; Soares, A.G.; Stockand, J.D. Snake Venoms in Drug Discovery: Valuable Therapeutic Tools for Life Saving. Toxins 2019, 11, 564. [Google Scholar] [CrossRef]
- Scarborough, R.M.; Rose, J.W.; Naughton, M.A.; Phillips, D.R.; Nannizzi, L.; Arfsten, A.; Campbell, A.M.; Charo, I.F. Characterization of the Integrin Specificities of Disintegrins Isolated From American Pit Viper Venoms. J. Biol. Chem. 1993, 268, 1058–1065. [Google Scholar] [CrossRef]
- Scarborough, R.M.; Rose, J.W.; Hsu, M.A.; Phillips, D.R.; Fried, V.A.; Campbell, A.M.; Nannizzi, L.; Charo, I.F. Barbourin. A GPIIb-IIIa-specific integrin antagonist from the venom of Sistrurus m. barbouri. J. Biol. Chem. 1991, 266, 9359–9362. [Google Scholar] [CrossRef]
- Scarborough, R.M. Development of eptifibatide. Am. Heart J. 1999, 138, 1093–1104. [Google Scholar] [CrossRef]
- Ferreira, B.A.; De Moura, F.B.R.; Tomiosso, T.C.; Corrêa, N.C.R.; Goulart, L.R.; Barcelos, L.S.; Clissa, P.B.; Araújo, F.A. Jararhagin-C, a disintegrin-like protein, improves wound healing in mice through stimulation of M2-like macrophage, angiogenesis and collagen deposition. Int. Immunopharmacol. 2021, 101, 108224. [Google Scholar] [CrossRef] [PubMed]
- Morjen, M.; Kallech-Ziri, O.; Bazaa, A.; Othman, H.; Mabrouk, K.; Zouari-Kessentini, R.; Sanz, L.; Calvete, J.J.; Srairi-Abid, N.; El Ayeb, M.; et al. PIVL, a new serine protease inhibitor from Macrovipera lebetina transmediterranea venom, impairs motility of human glioblastoma cells. Matrix Biol. J. Int. Soc. Matrix Biol. 2013, 32, 52–62. [Google Scholar] [CrossRef] [PubMed]
- Ageitos, L.; Torres, M.D.T.; de la Fuente-nunez, C. Biologically Active Peptides from Venoms: Applications in Antibiotic Resistance, Cancer, and Beyond. Int. J. Mol. Sci. 2022, 23, 15437. [Google Scholar] [CrossRef] [PubMed]
- Retzios, A.D.; Markland, F.S., Jr. A direct-acting fibrinolytic enzyme from the venom of Agkistrodon contortrix contortrix: Effects on various components of the human blood coagulation and fibrinolysis systems. Thromb. Res. 1988, 52, 541–552. [Google Scholar] [CrossRef]
- Swenson, S.; Markland, F.S., Jr. Snake venom fibrin(ogen)olytic enzymes. Toxicon 2005, 45, 1021–1039. [Google Scholar] [CrossRef]
- Rau, J.C.; Beaulieu, L.M.; Huntington, J.A.; Church, F.C. Serpins in thrombosis, hemostasis and fibrinolysis. J. Thromb. Haemost. 2007, 5, 102–115. [Google Scholar] [CrossRef]
- Deitcher, S.R.; Funk, W.D.; Buchanan, J.; Liu, S.; Levy, M.D.; Toombs, C.F. Alfimeprase: A novel recombinant direct-acting fibrinolytic. Expert. Opin. Biol. Ther. 2006, 6, 1361–1369. [Google Scholar] [CrossRef]
- Markland, F.S.; Swenson, S. Fibrolase: Trials and Tribulations. Toxins 2010, 2, 793–808. [Google Scholar] [CrossRef]
- Ramesh, K.V.; Rao, C.M.; Bairy, K.L.; Ramanarayan, K.; Kulkarni, D.R. EFFECTS OF PROCOAGULANTS ON WOUND-HEALING. Indian. J. Exp. Biol. 1990, 28, 43–45. [Google Scholar]
- Gupta, G.; Muthusekhar, M.R.; Kumar, S.P. Efficacy of Hemocoagulase as a Topical Hemostatic Agent After Dental Extractions: A Systematic Review. Cureus J. Med. Sci. 2018, 10, e2398. [Google Scholar] [CrossRef] [PubMed]
- Aronson, D.L. Comparison of Actions of Thrombin and Thrombin-Like Venom Enzymes Ancrod and Batroxobin. Thromb. Haemost. 1976, 36, 9–13. [Google Scholar] [CrossRef] [PubMed]
- Vu, T.T.; Stafford, A.R.; Leslie, B.A.; Kim, P.Y.; Fredenburgh, J.C.; Weitz, J.I. Batroxobin binds fibrin with higher affinity and promotes clot expansion to a greater extent than thrombin. J. Biol. Chem. 2013, 288, 16862–16871. [Google Scholar] [CrossRef] [PubMed]
- Levy, D.E.; Del Zoppo, G.J. Ancrod: A potential treatment for acute, ischemic stroke from snake venom. Toxin Rev. 2006, 25, 323–333. [Google Scholar] [CrossRef]
- Liu, S.; Marder, V.J.; Levy, D.E.; Wang, S.J.; Yang, F.; Paganini-Hill, A.; Fisher, M.J. Ancrod and Fibrin Formation Perspectives on Mechanisms of Action. Stroke 2011, 42, 3277–3280. [Google Scholar] [CrossRef]
- Lollar, P.; Parker, C.G.; Kajenski, P.J.; Litwiller, R.D.; Fass, D.N. Degradation of Coagulation Proteins by an Enzyme from Malayan Pit Viper (Akistrodon-Rhodostoma) Venom. Biochemistry 1987, 26, 7627–7636. [Google Scholar] [CrossRef]
- Siniavin, A.E.; Streltsova, M.A.; Nikiforova, M.A.; Kudryavtsev, D.S.; Grinkina, S.D.; Gushchin, V.A.; Mozhaeva, V.A.; Starkov, V.G.; Osipov, A.V.; Lummis, S.C.R.; et al. Snake venom phospholipase A2s exhibit strong virucidal activity against SARS-CoV-2 and inhibit the viral spike glycoprotein interaction with ACE2. Cell. Mol. Life Sci. 2021, 78, 7777–7794. [Google Scholar] [CrossRef]
- Brglez, V.; Pucer, A.; Pungerčar, J.; Lambeau, G.; Petan, T. Secreted phospholipases A2 are differentially expressed and epigenetically silenced in human breast cancer cells. Biochem. Biophys. Res. Commun. 2014, 445, 230–235. [Google Scholar] [CrossRef]
- Yu, J.A.; Mauchley, D.; Li, H.; Meng, X.; Nemenoff, R.A.; Fullerton, D.A.; Weyant, M.J. Knockdown of secretory phospholipase A2 IIa reduces lung cancer growth in vitro and in vivo. J. Thorac. Cardiovasc. Surg. 2012, 144, 1185–1191. [Google Scholar] [CrossRef][Green Version]
- Eberle, R.J.; Gering, I.; Tusche, M.; Ostermann, P.N.; Müller, L.; Adams, O.; Schaal, H.; Olivier, D.S.; Amaral, M.S.; Arni, R.K.; et al. Design of D-Amino Acids SARS-CoV-2 Main Protease Inhibitors Using the Cationic Peptide from Rattlesnake Venom as a Scaffold. Pharmaceuticals 2022, 15, 540. [Google Scholar] [CrossRef]
- Kerkis, A.; Hayashi, M.A.F.; Yamane, T.; Kerkis, I. Properties of cell penetrating peptides (CPPs). Iubmb Life 2006, 58, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, M.A.F.; Nascimento, F.D.; Kerkis, A.; Oliveira, V.; Oliveira, E.B.; Pereira, A.; Rádis-Baptista, G.; Nader, H.B.; Yamane, T.; Kerkis, I.; et al. Cytotoxic effects of crotamine are mediated through lysosomal membrane permeabilization. Toxicon 2008, 52, 508–517. [Google Scholar] [CrossRef] [PubMed]
- Moreira, L.A.; Oliveira, L.P.; Magalhães, M.R.; Oliveira, S.A.M.; Oliveira-Neto, J.R.; Carvalho, P.M.G.; Carvalho, A.A.V.; Fajemiroye, J.O.; Cruz, A.C.; Cunha, L.C. Acute toxicity, antinociceptive, and anti-inflammatory activities of the orally administered crotamine in mice. Naunyn-Schmiedeberg’s Arch. Pharmacol. 2021, 394, 1703–1711. [Google Scholar] [CrossRef] [PubMed]
- Radis-Baptista, G.; Kerkis, I. Crotamine, a Small Basic Polypeptide Myotoxin from Rattlesnake Venom with Cell-Penetrating Properties. Curr. Pharm. Des. 2011, 17, 4351–4361. [Google Scholar] [CrossRef] [PubMed]
- Read, M.S.; Shermer, R.W.; Brinkhous, K.M. Venom Coagglutinin—Activator of Platelet-Aggregation Dependent on Vonwillebrand Factor. Proc. Natl. Acad. Sci. USA 1978, 75, 4514–4518. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, K.; Doggett, T.; Laurenzi, I.J.; Liddington, R.C.; Diacovo, T.G. The snake venom protein botrocetin acts as a biological brace to promote dysfunctional platelet aggregation. Nat. Struct. Mol. Biol. 2005, 12, 152–159. [Google Scholar] [CrossRef] [PubMed]
- Stocker, K.; Fischer, H.; Meier, J.; Brogli, M.; Svendsen, L. Characterization of the Protein-C Activator Protac from the Venom of the Southern Copperhead (Agkistrodon-Contortrix) Snake. Toxicon 1987, 25, 239–252. [Google Scholar] [CrossRef] [PubMed]
- Johnstone, I.B.; Martin, C.A. Comparative effects of the human protein C activator, Protac, on the activated partial thromboplastin clotting times of plasmas, with special reference to the dog. Can. J. Vet. Res. 2000, 64, 117–122. [Google Scholar]
- Takeya, H.; Nishida, S.; Miyata, T.; Kawada, S.I.; Saisaka, Y.; Morita, T.; Iwanaga, S. Coagulation Factor-X Activating Enzyme From Russells Viper Venom (Rvv-X)—A Novel Metalloproteinase with Disintegrin (Platelet-Aggregation Inhibitor)-Like and C-Type Lectin-Like Domains. J. Biol. Chem. 1992, 267, 14109–14117. [Google Scholar] [CrossRef]
- Marsh, N.A. Diagnostic uses of snake venom. Haemostasis 2001, 31, 211–217. [Google Scholar] [CrossRef]
- Nakayama, D.; Ben Ammar, Y.; Miyata, T.; Takeda, S. Structural basis of coagulation factor V recognition for cleavage by RVV-V. Febs Lett. 2011, 585, 3020–3025. [Google Scholar] [CrossRef] [PubMed]
- Merlin, M.; Gecchele, E.; Capaldi, S.; Pezzotti, M.; Avesani, L. Comparative Evaluation of Recombinant Protein Production in Different Biofactories: The Green Perspective. Biomed. Res. Int. 2014, 2014, 136419. [Google Scholar] [CrossRef] [PubMed]
- Bordon, K.D.F.; Cologna, C.T.; Fornari-Baldo, E.C.; Pinheiro, E.L.; Cerni, F.A.; Amorim, F.G.; Anjolette, F.A.P.; Cordeiro, F.A.; Wiezel, G.A.; Cardoso, I.A.; et al. From Animal Poisons and Venoms to Medicines: Achievements, Challenges and Perspectives in Drug Discovery. Front. Pharmacol. 2020, 11, 1132. [Google Scholar] [CrossRef] [PubMed]
- Kerkis, I.; Hayashi, M.A.F.; da Silva, A.; Pereira, A.; De Sa, P.L.; Zaharenko, A.J.; Rádis-Baptista, G.; Kerkis, A.; Yamane, T. State of the Art in the Studies on Crotamine, a Cell Penetrating Peptide from South American Rattlesnake. Biomed. Res. Int. 2014, 2014, 675985. [Google Scholar] [CrossRef]
- Xhima, K.; Aubert, I. The therapeutic potential of nerve growth factor combined with blood-brain barrier modulation by focused ultrasound for neurodegenerative disorders. Neural Regen. Res. 2021, 16, 1783–1785. [Google Scholar] [CrossRef]
- Rokyta, D.R.; Margres, M.J.; Ward, M.J.; Sanchez, E.E. The genetics of venom ontogeny in the eastern diamondback rattlesnake (Crotalus adamanteus). PeerJ 2017, 5, e3249. [Google Scholar] [CrossRef]
- Buschek, S.; Ignjatovic, V.; Summerhayes, R.; Lowe, R. The effect of different snake venoms and anti-venoms on thrombin clotting time in human plasma. Thromb. Res. 2010, 125, e149-152. [Google Scholar] [CrossRef]
- Castro, E.N.; Lomonte, B.; del Carmen Gutierrez, M.; Alagon, A.; Gutierrez, J.M. Intraspecies variation in the venom of the rattlesnake Crotalus simus from Mexico: Different expression of crotoxin results in highly variable toxicity in the venoms of three subspecies. J. Proteom. 2013, 87, 103–121. [Google Scholar] [CrossRef]
- Heinrikson, R.L.; Krueger, E.T.; Keim, P.S. Amino acid sequence of phospholipase A2-alpha from the venom of Crotalus adamanteus. A new classification of phospholipases A2 based upon structural determinants. J. Biol. Chem. 1977, 252, 4913–4921. [Google Scholar] [CrossRef]
- Massey, V.; Curti, B. On the Reaction Mechanism of Crotalus adamanteus l-Amino Acid Oxidase. J. Biol. Chem. 1967, 242, 1259–1264. [Google Scholar] [CrossRef]
- Samy, R.P.; Kandasamy, M.; Gopalakrishnakone, P.; Stiles, B.G.; Rowan, E.G.; Becker, D.; Shanmugam, M.K.; Sethi, G.; Chow, V.T.K. Wound healing activity and mechanisms of action of an antibacterial protein from the venom of the eastern diamondback rattlesnake (Crotalus adamanteus). PLoS ONE 2014, 9, e80199. [Google Scholar] [CrossRef] [PubMed]
- Wellner, D.; Meister, A. Crystalline L-amino acid oxidase of Crotalus adamanteus. J. Biol. Chem. 1960, 235, 2013–2018. [Google Scholar] [CrossRef] [PubMed]
- Schonour, R.B.; Huff, E.M.; Holding, M.L.; Claunch, N.M.; Ellsworth, S.A.; Hogan, M.P.; Wray, K.; McGivern, J.; Margres, M.J.; Colston, T.J.; et al. Gradual and Discrete Ontogenetic Shifts in Rattlesnake Venom Composition and Assessment of Hormonal and Ecological Correlates. Toxins 2020, 12, 659. [Google Scholar] [CrossRef] [PubMed]
- Jia, Y.; Olvera, P.; Rangel, F.; Mendez, B.; Reddy, S. Rapid Identification of Phospholipase A2 Transcripts from Snake Venoms. Toxins 2019, 11, 69. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; He, Q.Y.; Yu, X.D.; Li, B.; Song, Z.W. The anti-ovarian carcinoma activity of l-amino acid oxidase from Crotalus adamanteusvenom in vivo and in vitro. Med. Oncol. 2022, 39, 112. [Google Scholar] [CrossRef] [PubMed]
- Fox, J.W.; Serrano, S.M.T. Timeline of key events in snake venom metalloproteinase research. J. Proteom. 2009, 72, 200–209. [Google Scholar] [CrossRef]
- Bjarnason, J.B.; Barish, A.; Direnzo, G.S.; Campbell, R.; Fox, J.W. Kallikrein-like Enzymes From Crotalus Atrox Venom. J. Biol. Chem. 1983, 258, 12566–12573. [Google Scholar] [CrossRef]
- Sultan, M.T.; Li, H.-M.; Lee, Y.Z.; Lim, S.S.; Song, D.-K. Identification of Lys49-PLA2 from crude venom of Crotalus atrox as a human neutrophil-calcium modulating protein. Korean J. Physiol. Pharmacol. 2016, 20, 177–183. [Google Scholar] [CrossRef]
- Torii, S.; Naito, M.; Tsuruo, T. Apoxin I, a novel apoptosis-inducing factor with L-amino acid oxidase activity purified from Western diamondback rattlesnake venom. J. Biol. Chem. 1997, 272, 9539–9542. [Google Scholar] [CrossRef]
- Slagboom, J.; Derks, R.J.E.; Sadighi, R.; Somsen, G.W.; Ulens, C.; Casewell, N.R.; Kool, J. High-Throughput Venomics. J. Proteome Res. 2023, 22, 1734–1746. [Google Scholar] [CrossRef]
- Retzios, A.D.; Markland, J.F.S. Purification, characterization, and fibrinogen cleavage sites of three fibrinolytic enzymes from the venom of Crotalus basiliscus basiliscus. Biochemistry 1992, 31, 4547–4557. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.H.; Wang, Y.M.; Hseu, M.K.; Tsai, I.H. Molecular evolution and structure-function relationships of crotoxin-like and asparagine-6-containing phospholipases A2 in pit viper venoms. Biochem. J. 2004, 381, 25–34. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, M.; Solano, G.; Vargas, M.; Reta-Mares, F.; Neri-Castro, E.; Alagón, A.; Sánchez, A.; Villalta, M.; León, G.; Segura, A. Toxicological profile of medically relevant Crotalus species from Mexico and their neutralization by a Crotalus basiliscus Bothrops asper antivenom. Toxicon 2020, 179, 92–100. [Google Scholar] [CrossRef] [PubMed]
- Datta, G.; Dong, A.; Witt, J.; Tu, A.T. Biochemical characterization of basilase, a fibrinolytic enzyme from Crotalus basiliscus basiliscus. Arch. Biochem. Biophys. 1995, 317, 365–373. [Google Scholar] [CrossRef] [PubMed]
- Arnaud, G.; García-de León, F.J.; Beltrán, L.F.; Carbajal-Saucedo, A. Proteomic comparison of adult and juvenile Santa Catalina rattlesnake (Crotalus catalinensis) venom. Toxicon Off. J. Int. Soc. Toxinol. 2021, 193, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Glenn, J.L.; Straight, R.C. Venom properties of the rattlesnakes (Crotalus) inhabiting the Baja California region of Mexico. Toxicon 1985, 23, 769–775. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, E.P.; Rautsaw, R.M.; Strickland, J.L.; Holding, M.L.; Hogan, M.P.; Mason, A.J.; Rokyta, D.R.; Parkinson, C.L. Comparative venom-gland transcriptomics and venom proteomics of four Sidewinder Rattlesnake (Crotalus cerastes) lineages reveal little differential expression despite individual variation. Sci. Rep. 2018, 8, 15534. [Google Scholar] [CrossRef] [PubMed]
- Rautsaw, R.M.; Hofmann, E.P.; Margres, M.J.; Holding, M.L.; Strickland, J.L.; Mason, A.J.; Rokyta, D.R.; Parkinson, C.L. Intraspecific sequence and gene expression variation contribute little to venom diversity in sidewinder rattlesnakes (Crotalus cerastes). Proc. Biol. Sci. 2019, 286, 20190810. [Google Scholar] [CrossRef]
- de Oliveira, L.A.; Ferreira, R.S., Jr.; Barraviera, B.; de Carvalho, F.C.T.; de Barros, L.C.; Dos Santos, L.D.; Pimenta, D.C. Crotalus durissus terrificus crotapotin naturally displays preferred positions for amino acid substitutions. J. Venom. Anim. Toxins Incl. Trop. Dis. 2017, 23, 46. [Google Scholar] [CrossRef]
- Bandeira, I.C.J.; Bandeira-Lima, D.; Mello, C.P.; Pereira, T.P.; De Menezes, R.; Sampaio, T.L.; Falcao, C.B.; Radis-Baptista, G.; Martins, A.M.C. Antichagasic effect of crotalicidin, a cathelicidin-like vipericidin, found in Crotalus durissus terrificus rattlesnake’s venom gland. Parasitology 2018, 145, 1059–1064. [Google Scholar] [CrossRef]
- Bordon, K.C.; Perino, M.G.; Giglio, J.R.; Arantes, E.C. Isolation, enzymatic characterization and antiedematogenic activity of the first reported rattlesnake hyaluronidase from Crotalus durissus terrificus venom. Biochimie 2012, 94, 2740–2748. [Google Scholar] [CrossRef]
- Costa, C.R.C.; Belchor, M.N.; Rodrigues, C.F.B.; Toyama, D.O.; de Oliveira, M.A.; Novaes, D.P.; Toyama, M.H. Edema Induced by a Crotalus durissus terrificus Venom Serine Protease (Cdtsp 2) Involves the PAR Pathway and PKC and PLC Activation. Int. J. Mol. Sci. 2018, 19, 2405. [Google Scholar] [CrossRef] [PubMed]
- Freitas, A.P.; Favoretto, B.C.; Clissa, P.B.; Sampaio, S.C.; Faquim-Mauro, E.L. Crotoxin Isolated from Crotalus durissus terrificus Venom Modulates the Functional Activity of Dendritic Cells via Formyl Peptide Receptors. J. Immunol. Res. 2018, 2018, 7873257. [Google Scholar] [CrossRef] [PubMed]
- Marcussi, S.; Santos, P.R.S.; Menaldo, D.L.; Silveira, L.B.; Santos-Filho, N.; Mazzi, M.V.; da Silva, S.L.; Stábeli, R.G.; Antunes, L.M.G.; Soares, A.M. Evaluation of the genotoxicity of Crotalus durissus terrificus snake venom and its isolated toxins on human lymphocytes. Mutat. Res. 2011, 724, 59–63. [Google Scholar] [CrossRef]
- Pereañez, J.A.; Núñez, V.; Huancahuire-Vega, S.; Marangoni, S.; Ponce-Soto, L.A. Biochemical and Biological Characterization of a PLA2 From Crotoxin Complex of Crotalus Durissus Cumanensis. Toxicon Off. J. Int. Soc. Toxinol. 2009, 53, 534–542. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, J.F.; Pereira, C.M.; Bittar, C.; Batista, M.N.; Campos, G.R.F.; da Silva, S.; Cintra, A.C.O.; Zothner, C.; Harris, M.; Sampaio, S.V.; et al. Multiple effects of toxins isolated from Crotalus durissus terrificus on the hepatitis C virus life cycle. PLoS ONE 2017, 12, e0187857. [Google Scholar] [CrossRef] [PubMed]
- Vargas, L.J.; Quintana, J.C.; Pereanez, J.A.; Nunez, V.; Sanz, L.; Calvete, J. Cloning and characterization of an antibacterial L-amino acid oxidase from Crotalus durissus cumanensis venom. Toxicon 2013, 64, 1–11. [Google Scholar] [CrossRef]
- Wiezel, G.A.; Shibao, P.Y.T.; Cologna, C.T.; Morandi Filho, R.; Ueira-Vieira, C.; De Pauw, E.; Quinton, L.; Arantes, E.C. In-Depth Venome of the Brazilian Rattlesnake Crotalus durissus terrificus: An Integrative Approach Combining Its Venom Gland Transcriptome and Venom Proteome. J. Proteome Res. 2018, 17, 3941–3958. [Google Scholar] [CrossRef]
- Rokyta, D.R.; Wray, K.P.; McGivern, J.J.; Margres, M.J. The transcriptomic and proteomic basis for the evolution of a novel venom phenotype within the Timber Rattlesnake (Crotalus horridus). Toxicon 2015, 98, 34–48. [Google Scholar] [CrossRef]
- Martinez-Romero, G.; Rucavado, A.; Lazcano, D.; Gutierrez, J.M.; Borja, M.; Lomonte, B.; Garza-Garcia, Y.; Zugasti-Cruz, A. Comparison of venom composition and biological activities of the subspecies Crotalus lepidus lepidus, Crotalus lepidus klauberi and Crotalus lepidus morulus from Mexico. Toxicon 2013, 71, 84–95. [Google Scholar] [CrossRef]
- Borja, M.; Lazcano, D.; Martínez-Romero, G.; Morlett, J.; Sánchez, E.; Cepeda-Nieto, A.C.; Garza-García, Y.; Zugasti-Cruz, A. Intra-specific Variation in the Protein Composition and Proteolytic Activity of Venom of Crotalus lepidus morulus from the Northeast of Mexico. Copeia 2013, 2013, 707–716. [Google Scholar] [CrossRef]
- Meléndez-Martínez, D.; Muñoz, J.M.; Barraza-Garza, G.; Cruz-Peréz, M.S.; Gatica-Colima, A.; Alvarez-Parrilla, E.; Plenge-Tellechea, L.F. Rattlesnake Crotalus molossus nigrescens venom induces oxidative stress on human erythrocytes. J. Venom. Anim. Toxins Incl. Trop. Dis. 2017, 23. [Google Scholar] [CrossRef] [PubMed]
- Ponce-López, R.; Neri-Castro, E.; Olvera-Rodríguez, F.; Sánchez, E.E.; Alagón, A.; Olvera-Rodríguez, A. Neutralization of crotamine by polyclonal antibodies generated against two whole rattlesnake venoms and a novel recombinant fusion protein. Toxicon Off. J. Int. Soc. Toxinol. 2021, 197, 70–78. [Google Scholar] [CrossRef] [PubMed]
- Borja, M.; Neri-Castro, E.; Gutiérrez-Martínez, A.; Bledsoe, R.; Zarzosa, V.; Rodriguez-López, B.; Strickland, J.L.; Becerra-López, J.; Valenzuela-Ceballos, S.; Parkinson, C.L.; et al. Ontogenetic change in the venom composition of one Mexican black-tailed rattlesnake (Crotalus molossus nigrescens) from Durango, Mexico. Toxicon 2023, 234, 107280. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Rael, E.D. Purification of M5, a fibrinolytic proteinase from Crotalus molossus molossus venom that attacks complement. Int. J. Biochem. Cell Biol. 1997, 29, 789–799. [Google Scholar] [CrossRef] [PubMed]
- Rael, E.D.; Martinez, M.; Molina, O. Isolation of a fibrinolytic protease, M4, from venom of Crotalus molossus molossus (northern blacktail rattlesnake). Haemostasis 1992, 22, 41–49. [Google Scholar] [CrossRef] [PubMed]
- Borja, M.; Neri-Castro, E.; Pérez-Morales, R.; Strickland, J.L.; Ponce-López, R.; Parkinson, C.L.; Espinosa-Fematt, J.; Sáenz-Mata, J.; Flores-Martínez, E.; Alagón, A.; et al. Ontogenetic Change in the Venom of Mexican Black-Tailed Rattlesnakes (Crotalus molossus nigrescens). Toxins 2018, 10, 501. [Google Scholar] [CrossRef] [PubMed]
- Martins, W.; Baldasso, P.A.; Honorio, K.M.; Maltarollo, V.G.; Ribeiro, R.I.; Carvalho, B.M.; Soares, A.M.; Calderon, L.A.; Stabeli, R.G.; Caballol, M.A.; et al. A novel phospholipase A2 (D49) from the venom of the Crotalus oreganus abyssus (North American Grand canyon rattlesnake). Biomed. Res. Int. 2014, 2014, 654170. [Google Scholar] [CrossRef]
- Salazar, A.M.; Guerrero, B.; Cantu, B.; Cantu, E.; Rodríguez-Acosta, A.; Pérez, J.C.; Galán, J.A.; Tao, A.; Sánchez, E.E. Venom Variation in Hemostasis of the Southern Pacific Rattlesnake (Crotalus Oreganus Helleri): Isolation of Hellerase. Comp. Biochem. Physiology. Toxicol. Pharmacol. 2009, 149, 307–316. [Google Scholar] [CrossRef]
- Sunagar, K.; Undheim, E.A.; Scheib, H.; Gren, E.C.; Cochran, C.; Person, C.E.; Koludarov, I.; Kelln, W.; Hayes, W.K.; King, G.F.; et al. Intraspecific Venom Variation in the Medically Significant Southern Pacific Rattlesnake (Crotalus Oreganus Helleri): Biodiscovery, Clinical and Evolutionary Implications. J. Proteom. 2014, 99, 68–83. [Google Scholar] [CrossRef]
- Minton, S.A.; Weinstein, S.A. Protease activity and lethal toxicity of venoms from some little known rattlesnakes. Toxicon 1984, 22, 828–830. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, E.E.; García, C.; Pérez, J.C.; de la Zerda, S.J. The detection of hemorrhagic proteins in snake venoms using monoclonal antibodies against Virginia opossum (Didelphis virginiana) serum. Toxicon 1998, 36, 1451–1459. [Google Scholar] [CrossRef] [PubMed]
- Straight, R.C.; Glenn, J.L.; Wolt, T.B.; Wolfe, M.C. North-south regional variation in phospholipase A activity in the venom of Crotalus ruber. Comp. Biochem. Physiol. B 1992, 103, 635–639. [Google Scholar] [CrossRef] [PubMed]
- Mori, N.; Nikai, T.; Sugihara, H.; Tu, A.T. Biochemical characterization of hemorrhagic toxins with fibrinogenase activity isolated from Crotalus ruber ruber venom. Arch. Biochem. Biophys. 1987, 253, 108–121. [Google Scholar] [CrossRef] [PubMed]
- Mori, N.; Nikai, T.; Sugihara, H. Phosphodiesterase from the venom of Crotalus ruber ruber. Int. J. Biochem. 1987, 19, 115–119. [Google Scholar] [CrossRef] [PubMed]
- Mori, N.; Sugihara, H. Comparative study of two arginine ester hydrolases, E-I and E-II from the venom of Crotalus ruber ruber (red rattlesnake). Comp. Biochem. Physiol. B 1989, 92, 537–547. [Google Scholar] [CrossRef] [PubMed]
- Mori, N.; Sugihara, H. Characterization of kallikrein-like enzyme from Crotalus ruber ruber (red rattlesnake) venom. Int. J. Biochem. 1989, 21, 83–90. [Google Scholar] [CrossRef] [PubMed]
- Hamako, J.; Suzuki, Y.; Hayashi, N.; Kimura, M.; Ozeki, Y.; Hashimoto, K.; Matsui, T. Amino acid sequence and characterization of C-type lectin purified from the snake venom of Crotalus ruber. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2007, 146, 299–306. [Google Scholar] [CrossRef]
- Szteiter, S.S.; Diego, I.N.; Ortegon, J.; Salinas, E.M.; Cirilo, A.; Reyes, A.; Sanchez, O.; Suntravat, M.; Salazar, E.; Sanchez, E.E.; et al. Examination of the Efficacy and Cross-Reactivity of a Novel Polyclonal Antibody Targeting the Disintegrin Domain in SVMPs to Neutralize Snake Venom. Toxins 2021, 13, 19. [Google Scholar] [CrossRef]
- Borja, M.; Castañeda, G.; Espinosa, J.; Neri, E.; Carbajal, A.; Clement, H.; García, O.; Alagon, A. Mojave Rattlesnake (Crotalus scutulatus scutulatus) with Type B Venom from Mexico. Copeia 2014, 2014, 7–13. [Google Scholar] [CrossRef]
- Ho, C.L.; Lee, C.Y. Presynaptic actions of Mojave toxin isolated from Mojave rattlesnake (crotalus scutulatus) venom. Toxicon 1981, 19, 889–892. [Google Scholar] [CrossRef] [PubMed]
- Hinson, M.W.; Childs, C.; Johnson, B.D.; Sifford, D.H. Concanavalin A-Binding Enzymes of Crotalus scutulatus scutulatus Venom. J. Ark. Acad. Sci. 1985, 39, 50–54. [Google Scholar]
- Massey, D.J.; Calvete, J.J.; Sanchez, E.E.; Sanz, L.; Richards, K.; Curtis, R.; Boesen, K. Venom variability and envenoming severity outcomes of the Crotalus scutulatus scutulatus (Mojave rattlesnake) from Southern Arizona. J. Proteom. 2012, 75, 2576–2587. [Google Scholar] [CrossRef] [PubMed]
- Modahl, C.M.; Mackessy, S.P. Full-Length Venom Protein cDNA Sequences from Venom-Derived mRNA: Exploring Compositional Variation and Adaptive Multigene Evolution. PLoS Negl. Trop. Dis. 2016, 10, e0004587. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G.; Boyer, L.V.; Redford, D.T.; Ford, P. Thrombelastographic characterization of the thrombin-like activity of Crotalus simus and Bothrops asper venoms. Blood Coagul. Fibrinolysis. 2017, 28, 211–217. [Google Scholar] [CrossRef] [PubMed]
- Weinstein, S.A.; Smith, L.A. Preliminary fractionation of tiger rattlesnake (Crotalus tigris) venom. Toxicon 1990, 28, 1447–1455. [Google Scholar] [CrossRef]
- Powell, R.L.; Lieb, C.S.; Rael, E.D. Identification of a neurotoxic venom component in the Tiger rattlesnake, Crotalus tigris. J. Herpetol. 2004, 38, 149–152. [Google Scholar] [CrossRef]
- Aguilar, I.; Giron, M.E.; Rodriguez-Acosta, A. Purification and characterisation of a haemorrhagic fraction from the venom of the Uracoan rattlesnake Crotalus vegrandis. Biochim. Biophys. Acta 2001, 1548, 57–65. [Google Scholar] [CrossRef]
- Giron, M.; Pinto, A.; Finol, H.J.; Aguilar, I.; Rodriguez-Acosta, A. Kidney structural and ultrastructural pathological changes induced by uracoan rattlesnake (Crotalus vegrandis Klauber 1941) venom. J. Submicrosc. Cytol. Pathol. 2002, 34, 447–459. [Google Scholar]
- Rodriguez-Acosta, A.; Aguilar, I.; Giron, M.; Rodriguez-Pulido, V. Haemorrhagic activity of neotropical rattlesnake (Crotalus vegrandis Klauber, 1941) venom. Nat. Toxins 1998, 6, 15–18. [Google Scholar] [CrossRef]
- Scannone, H.R.; Rodriguez, O.G.; Lancini, A.R. Enzymatic activities and other characteristics of Crotalus vegrandis snake venom. Toxins 1978, 223–229. [Google Scholar] [CrossRef]
- Kaiser, I.I.; Aird, S.D. A crotoxin homolog from the venom of the Uracoan rattlesnake (Crotalus vegrandis). Toxicon 1987, 25, 1113–1120. [Google Scholar] [CrossRef] [PubMed]
- Mackessy, S.P. Evolutionary trends in venom composition in the western rattlesnakes (Crotalus viridis sensu lato): Toxicity vs. tenderizers. Toxicon 2010, 55, 1463–1474. [Google Scholar] [CrossRef] [PubMed]
- Adade, C.; Fernandes Anne Cristine, S.; Carvalho Ana Lúcia, O.; Zingali, R.; Souto-Padrón, T. Leishmanicidal Effects of a Phospholipase A2 Isolated from Crotalus viridis viridis Snake Venom. Toxicon 2012, 60, 117. [Google Scholar] [CrossRef]
- Adade, C.M.; Cons, B.L.; Melo, P.A.; Souto-Padron, T. Effect of Crotalus viridis viridis snake venom on the ultrastructure and intracellular survival of Trypanosoma cruzi. Parasitology 2011, 138, 46–58. [Google Scholar] [CrossRef] [PubMed]
- Zancolli, G.; Baker, T.G.; Barlow, A.; Bradley, R.K.; Calvete, J.J.; Carter, K.C.; de Jager, K.; Owens, J.B.; Price, J.F.; Sanz, L.; et al. Is Hybridization a Source of Adaptive Venom Variation in Rattlesnakes? A Test, Using a Crotalus scutulatus × viridis Hybrid Zone in Southwestern New Mexico. Toxins 2016, 8, 188. [Google Scholar] [CrossRef]
- Tan, N.H.; Ponnudurai, G. A Comparative Study of the Biological Activities of Rattlesnake (Genera Crotalus and Sistrurus) Venoms. Comp. Biochem. Physiology. C Comp. Pharmacol. Toxicol. 1991, 98, 455–461. [Google Scholar] [CrossRef]
- Doley, R.; Pahari, S.; Mackessy, S.P.; Kini, R.M. Accelerated exchange of exon segments in Viperid three-finger toxin genes (Sistrurus catenatus edwardsii; Desert Massasauga). BMC Evol. Biol. 2008, 8, 196. [Google Scholar] [CrossRef]
- Sunagar, K.; Jackson, T.N.; Undheim, E.A.; Ali, S.A.; Antunes, A.; Fry, B.G. Three-fingered RAVERs: Rapid Accumulation of Variations in Exposed Residues of snake venom toxins. Toxins 2013, 5, 2172–2208. [Google Scholar] [CrossRef]
- Juarez, P.; Sanz, L.; Calvete, J.J. Snake venomics: Characterization of protein families in Sistrurus barbouri venom by cysteine mapping, N-terminal sequencing, and tandem mass spectrometry analysis. Proteomics 2004, 4, 327–338. [Google Scholar] [CrossRef]
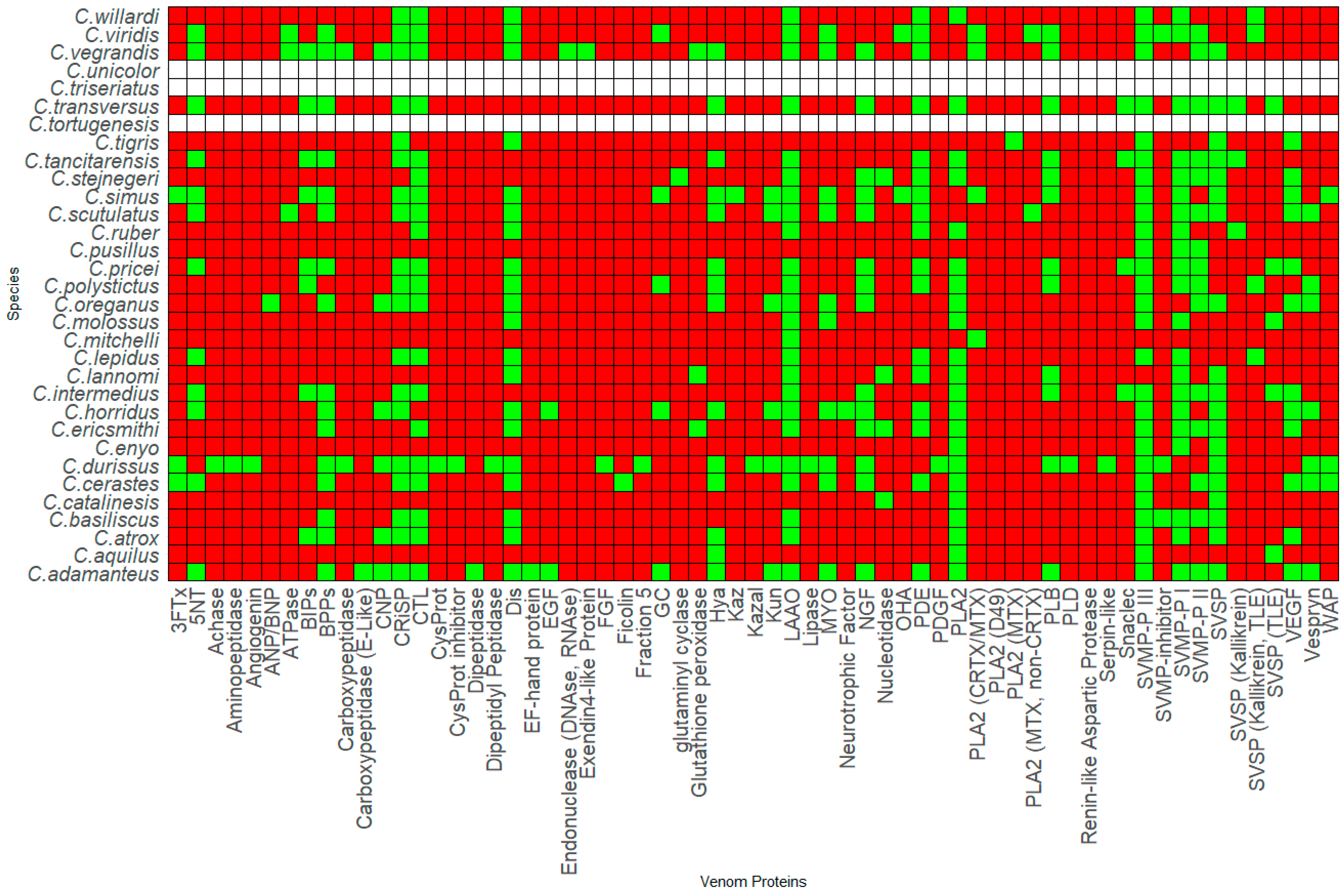
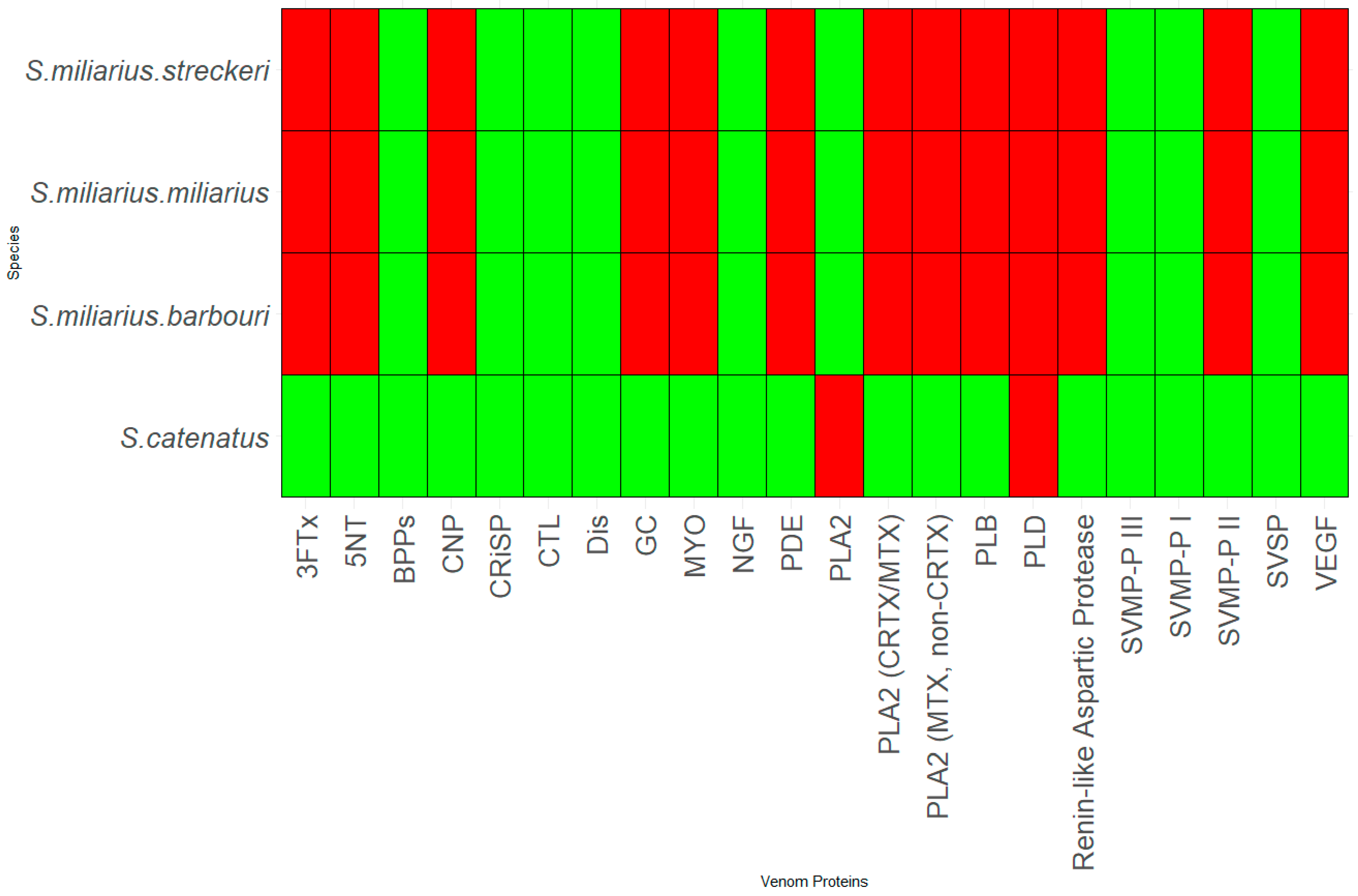
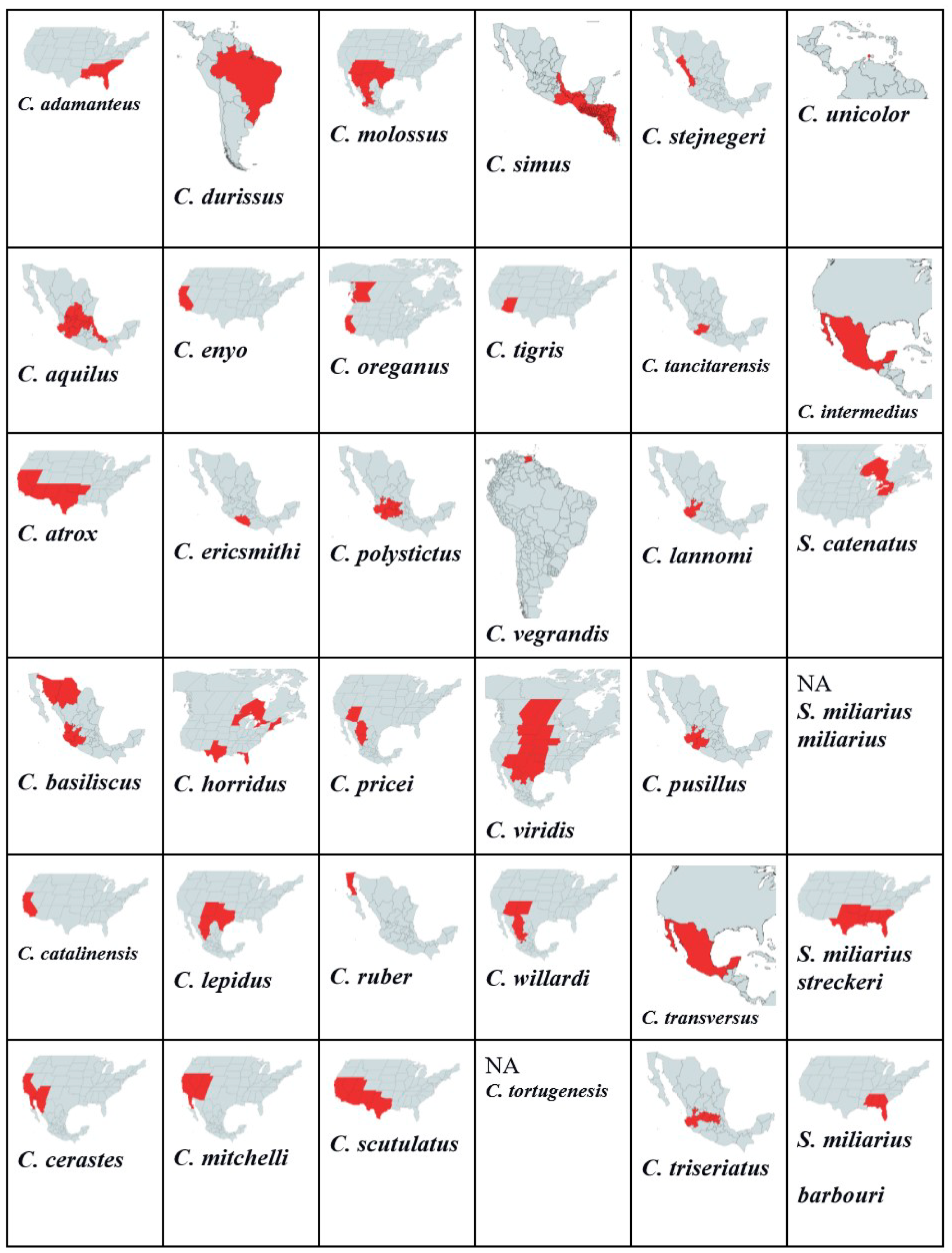
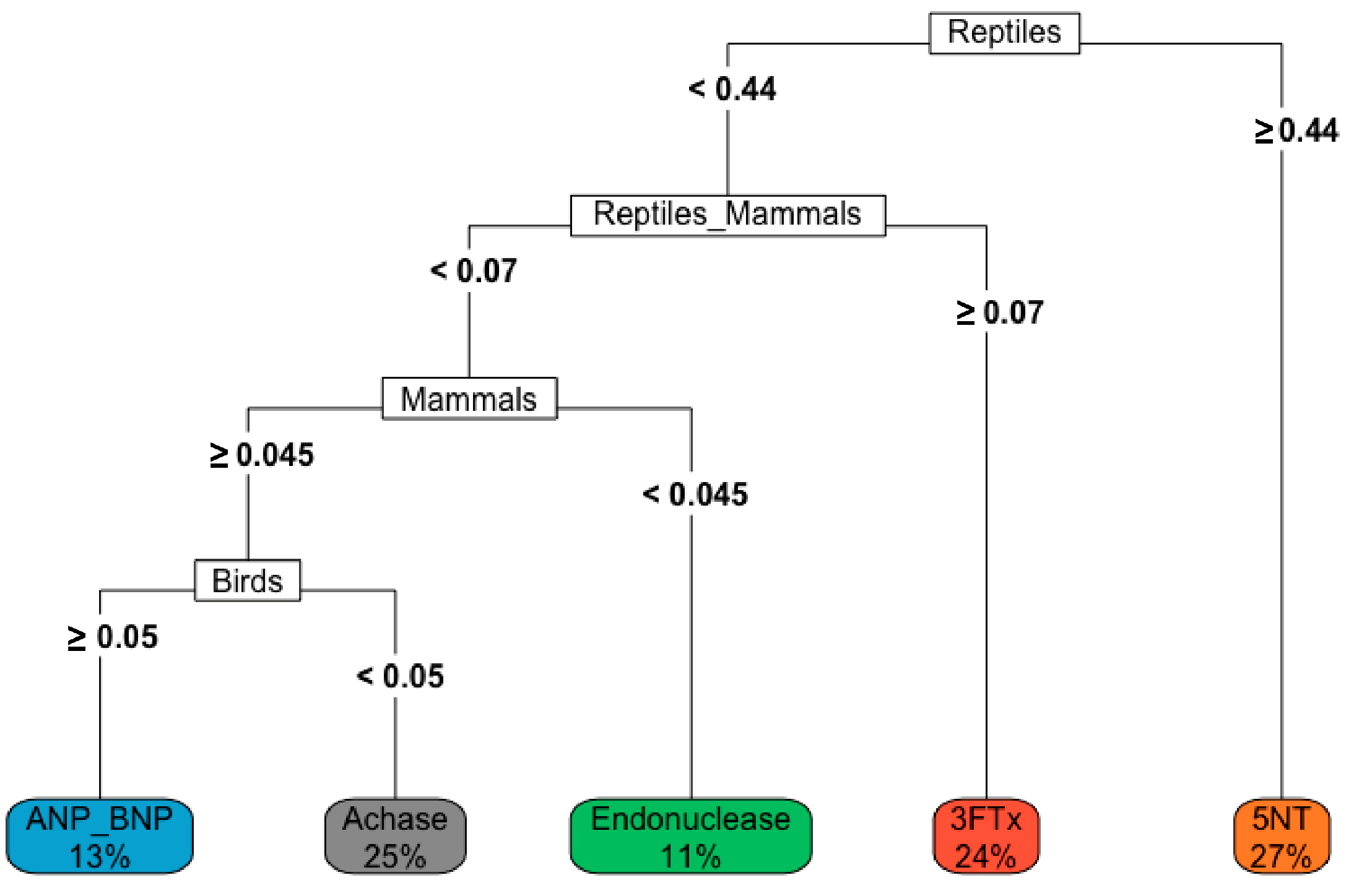
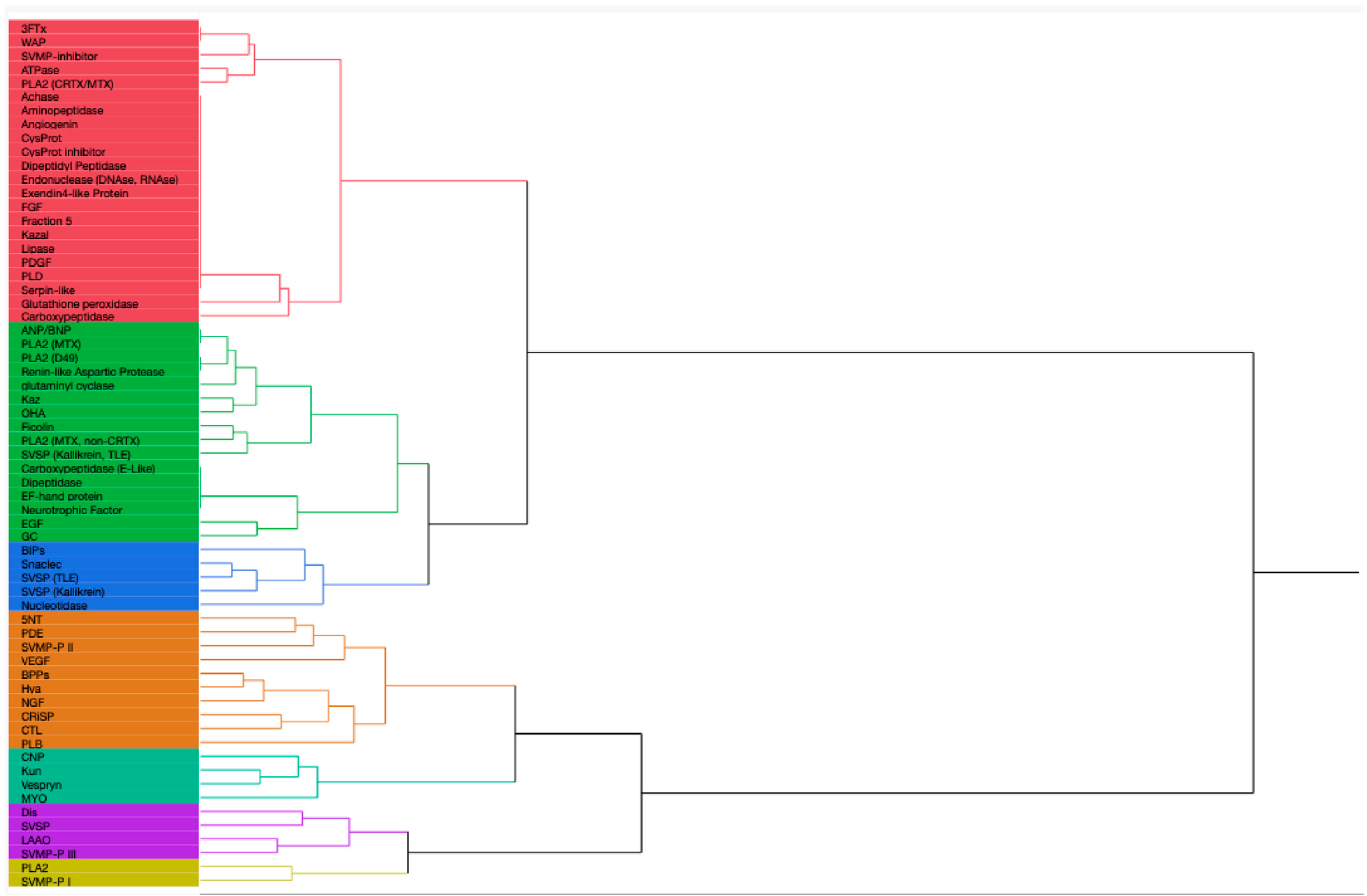
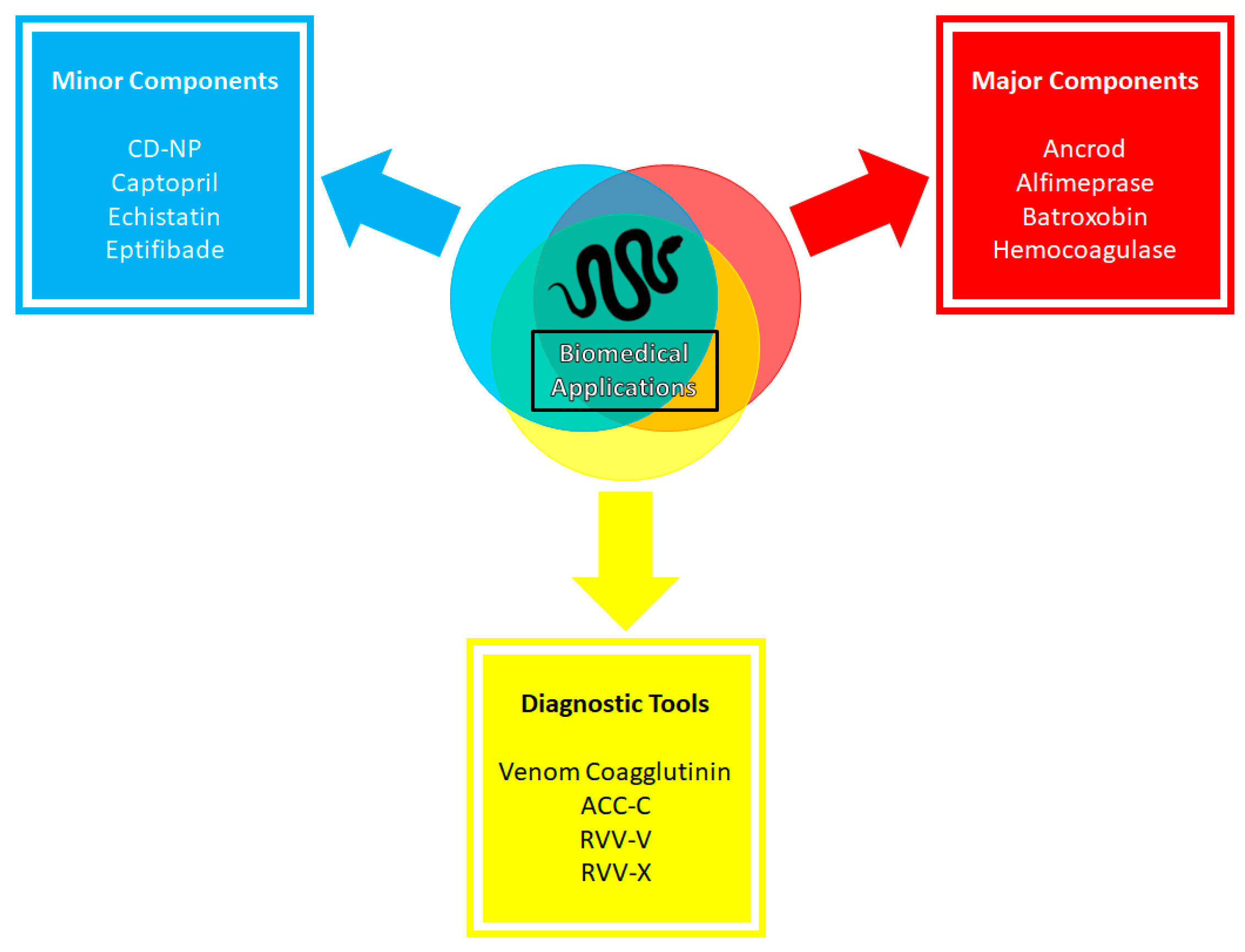
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Phan, P.; Deshwal, A.; McMahon, T.A.; Slikas, M.; Andrews, E.; Becker, B.; Kumar, T.K.S. A Review of Rattlesnake Venoms. Toxins 2024, 16, 2. https://doi.org/10.3390/toxins16010002
Phan P, Deshwal A, McMahon TA, Slikas M, Andrews E, Becker B, Kumar TKS. A Review of Rattlesnake Venoms. Toxins. 2024; 16(1):2. https://doi.org/10.3390/toxins16010002
Chicago/Turabian StylePhan, Phuc, Anant Deshwal, Tyler Anthony McMahon, Matthew Slikas, Elodie Andrews, Brian Becker, and Thallapuranam Krishnaswamy Suresh Kumar. 2024. "A Review of Rattlesnake Venoms" Toxins 16, no. 1: 2. https://doi.org/10.3390/toxins16010002
APA StylePhan, P., Deshwal, A., McMahon, T. A., Slikas, M., Andrews, E., Becker, B., & Kumar, T. K. S. (2024). A Review of Rattlesnake Venoms. Toxins, 16(1), 2. https://doi.org/10.3390/toxins16010002





