Characterization of Ugandan Endemic Aspergillus Species and Identification of Non-Aflatoxigenic Isolates for Potential Biocontrol of Aflatoxins
Abstract
:1. Introduction
2. Results
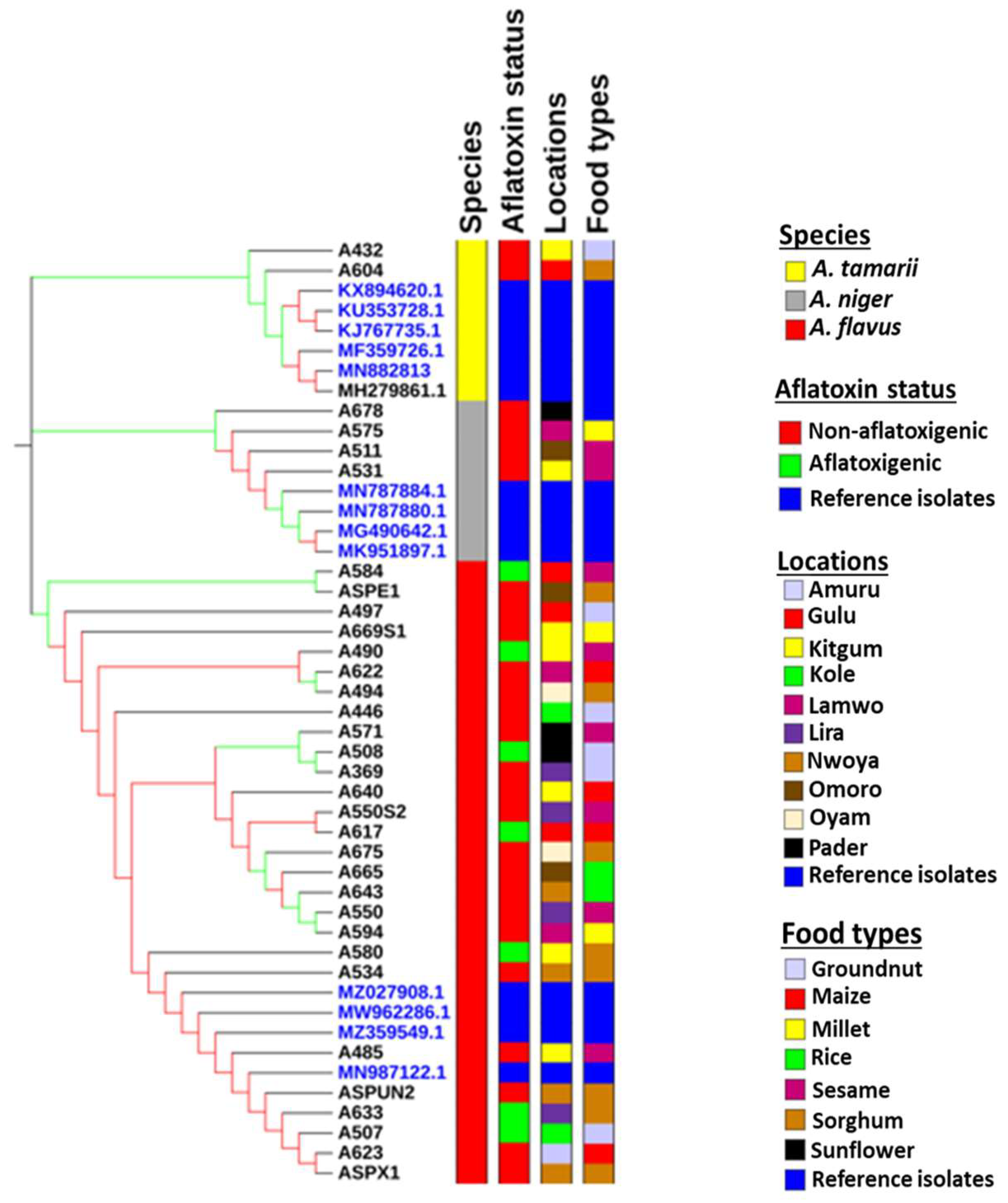
2.1. Characterization of Aspergillus Species
2.2. Distribution of Aspergillus Species by Food Types
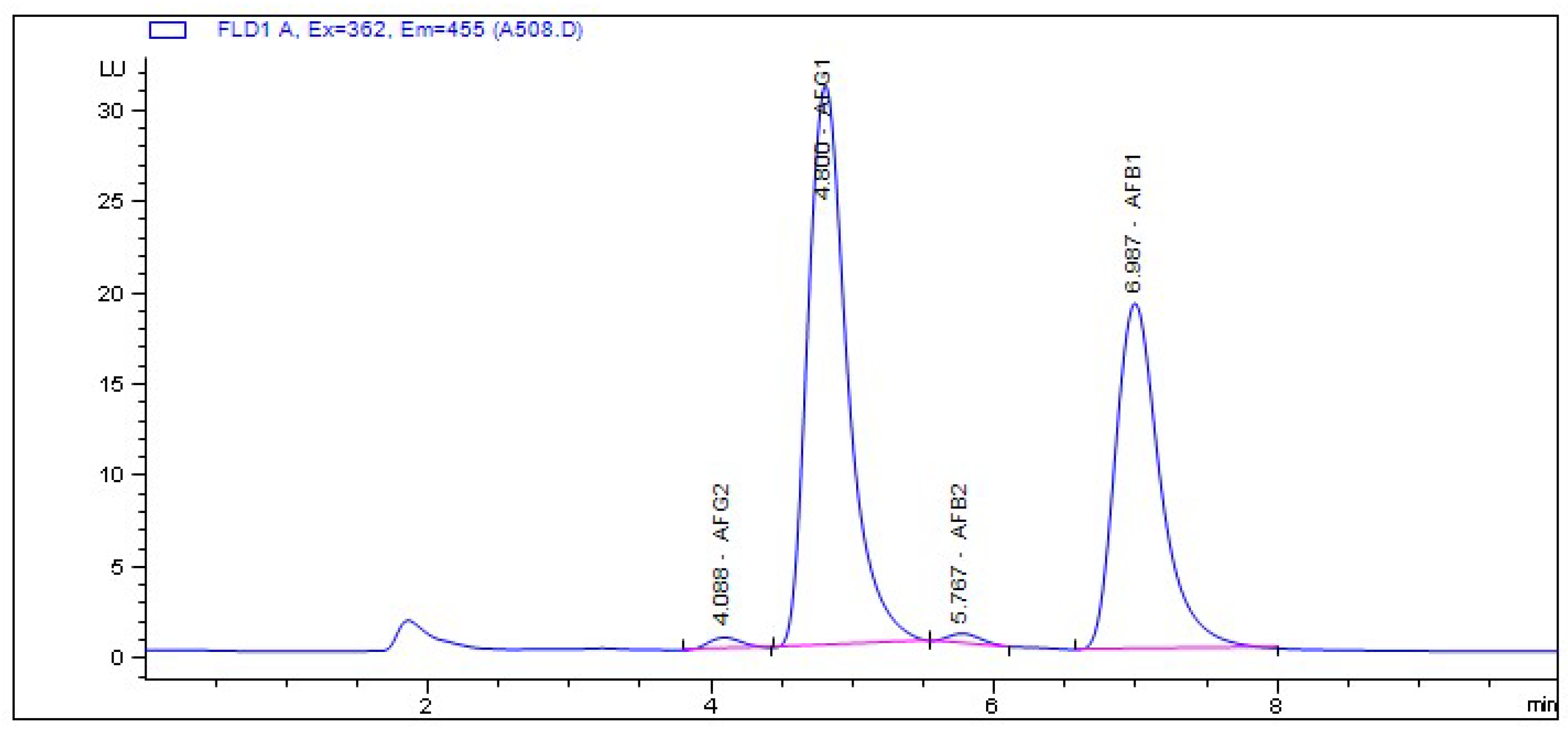
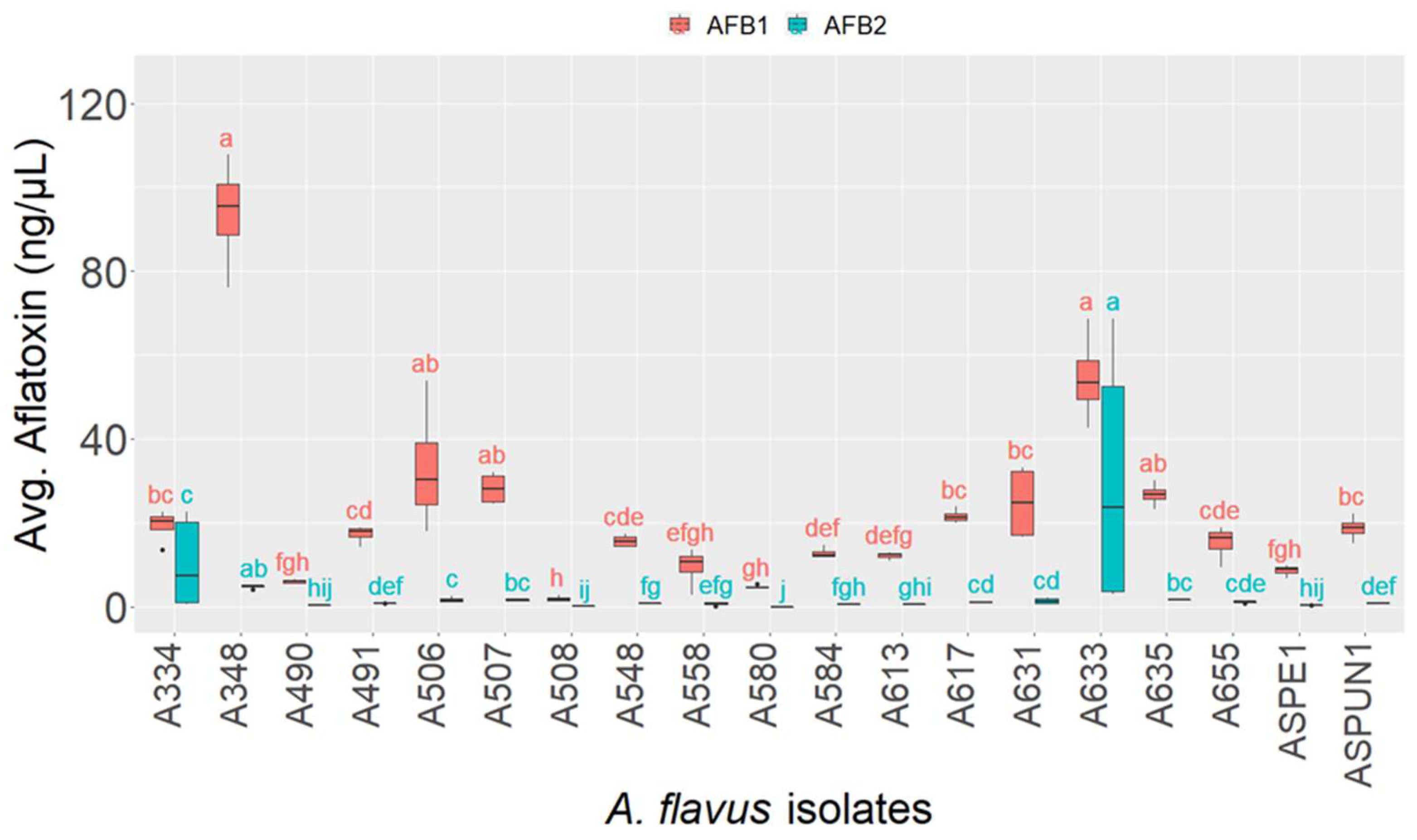
2.3. Chemotyping of Aspergillus Isolates
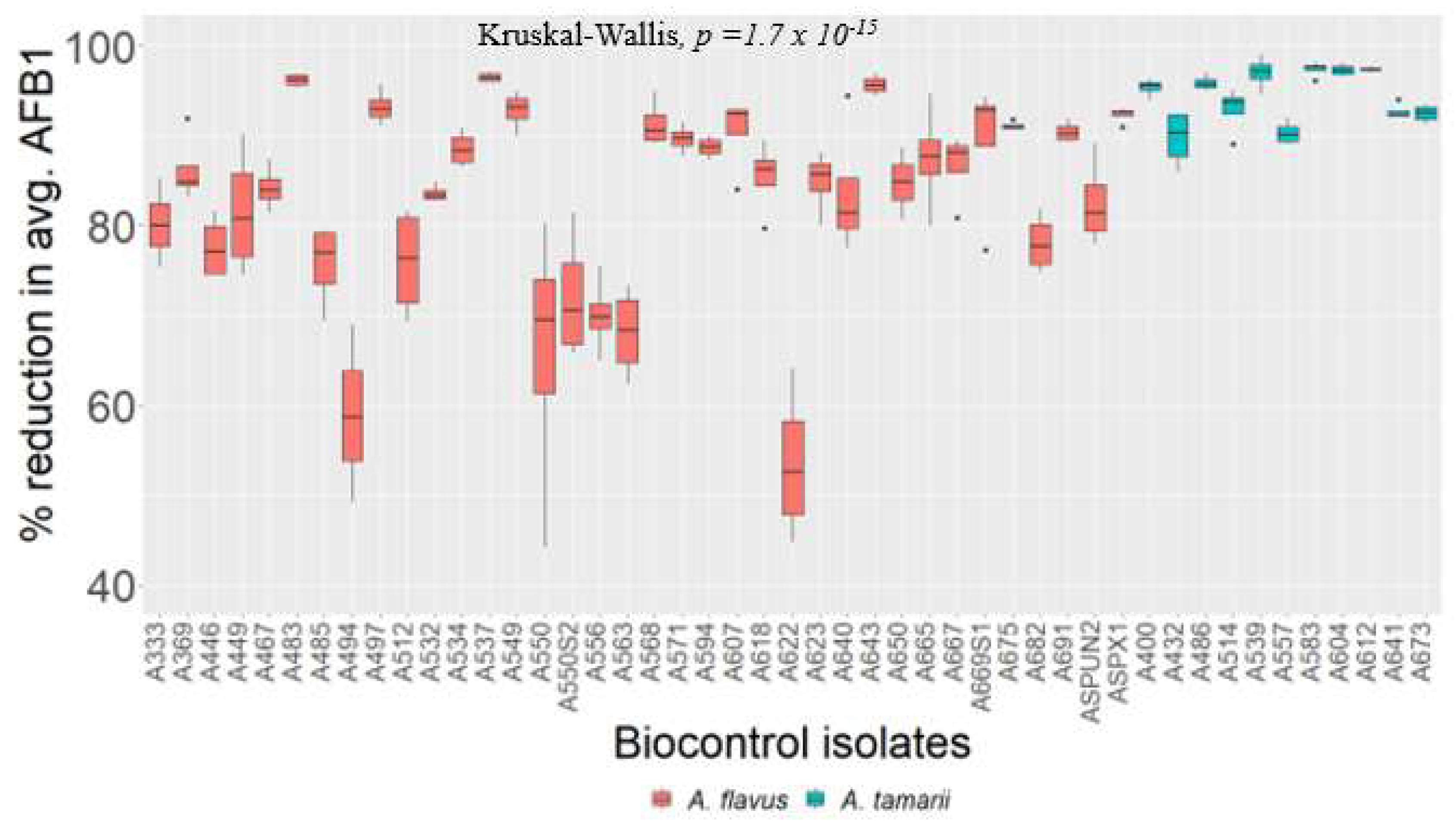
2.4. Testing the Biocontrol Potential of Non-Aflatoxigenic Aspergilli on Maize Grains
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Study Area
5.2. Sample Types
5.3. Isolation and Characterization of Aspergillus Isolates from Household Grains
5.4. Molecular Confirmation of Species Identities
5.5. Phylogenetic Tree Construction
5.6. Chemical Profiling of Isolates for Aflatoxin Production
5.7. Testing the Biocontrol Potential of Non-Aflatoxigenic Aspergilli on Maize Grains
5.8. Extraction of Aflatoxins from the Experimental Setup
5.9. Detection and Quantification of Aflatoxins
5.10. Data Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Murphy, P.A.; Hendrich, S.; Landgren, C.; Bryant, C.M. Food mycotoxins: An update. J. Food Sci. 2006, 71, 51–65. [Google Scholar] [CrossRef]
- Bandyopadhyay, R.; Kumar, M.; Leslie, J. Relative severity of aflatoxin contamination of cereal crops in West Africa. Food Addit. Contam. 2007, 24, 1109–1114. [Google Scholar] [CrossRef] [PubMed]
- Kaaya, A.N.; Harris, C.; Eigel, W. Peanut aflatoxin levels on farms and in markets of Uganda. Peanut Sci. 2006, 33, 68–75. [Google Scholar] [CrossRef] [Green Version]
- Joint FAO/WHO Expert Committee on Food Additives. Meeting 57th : 2001 : Geneva S, International Program on Chemical Safety, World Health Organization. Safety Evaluation of Certain Food Additives and Contaminants; World Health Organization: Geneva, Switzerland, 2002; 692p. [Google Scholar]
- EC (European Commission). Regulation (EC) No 1881/2006 of 19 December 2006 setting maximum levels for certain contaminants in foodstuffs. Off. J. Eur. Union 2006, 364, 5. [Google Scholar]
- Serck-Hanssen, A. Aflatoxin-Induced Fatal Hepatitis? Arch. Environ. Health Int. J. 1970, 20, 729–731. [Google Scholar] [CrossRef]
- Lauer, J.M.; Duggan, C.P.; Ausman, L.M.; Griffiths, J.K.; Webb, P.; Wang, J.S.; Xue, K.S.; Agaba, E.; Nshakira, N.; Ghosh, S. Maternal aflatoxin exposure during pregnancy and adverse birth outcomes in Uganda. Matern. Child Nutr. 2019, 15, e12701. [Google Scholar] [CrossRef] [Green Version]
- Asiki, G.; Seeley, J.; Srey, C.; Baisley, K.; Lightfoot, T.; Archileo, K.; Agol, D.; Abaasa, A.; Wakeham, K.; Routledge, M.N.; et al. A pilot study to evaluate aflatoxin exposure in a rural Ugandan population. Trop. Med. Int. Health 2014, 19, 592–599. [Google Scholar] [CrossRef]
- Lauer, J.M.; Natamba, B.K.; Ghosh, S.; Webb, P.; Wang, J.S.; Griffiths, J.K. Aflatoxin exposure in pregnant women of mixed status of human immunodeficiency virus infection and rate of gestational weight gain: A Ugandan cohort study. Trop. Med. Int. Health 2020, 25, 1145–1154. [Google Scholar] [CrossRef]
- Kang, M.S.; Nkurunziza, P.; Muwanika, R.; Qian, G.; Tang, L.; Song, X.; Xue, K.; Nkwata, A.; Ssempebwa, J.; Lutalo, T.; et al. Longitudinal evaluation of aflatoxin exposure in two cohorts in south-western Uganda. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2015, 32, 1322–1330. [Google Scholar] [CrossRef] [Green Version]
- Omara, T.; Nassazi, W.; Omute, T.; Awath, A.; Laker, F.; Kalukusu, R.; Musau, B.; Victoria Nakabuye, B.; Kagoya, S.; Otim, G.; et al. Aflatoxins in Uganda: An Encyclopedic Review of the Etiology, Epidemiology, Detection, Quantification, Exposure Assessment, Reduction, and Control. Int. J. Microbiol. 2020, 2020, 4723612. [Google Scholar] [CrossRef] [Green Version]
- Lukwago, F.B.; Mukisa, I.M.; Atukwase, A.; Kaaya, A.N.; Tumwebaze, S. Mycotoxins contamination in foods consumed in Uganda: A 12-year review (2006–18). Sci. Afr. 2019, 3, e00054. [Google Scholar] [CrossRef]
- Kaaya, A.N.; Warren, H.L. A review of past and present research on aflatoxin in Uganda. Afr. J. Food Agric. Nutr. Dev. 2005, 5, 1–18. [Google Scholar] [CrossRef]
- Nasir, U.; Naeem, I.; Asif, M.; Ismail, A.; Gong, Y.Y.; Routledgede, M.N.; Amjada, A.; Fazala, A.; Ismaila, Z. Assessment of aflatoxins exposure through urinary biomarker approach and the evaluation of the impacts of aflatoxins exposure on the selected health parameters of the children of Multan city of Pakistan. Food Control 2021, 123, 107863. [Google Scholar] [CrossRef]
- Frisvad, J.C.; Hubka, V.; Ezekiel, C.N.; Hong, S.B.; Nováková, A.; Chen, A.J.; Arzanlou, M.; Larsen, T.O.; Sklenář, F.; Mahakarnchanakul, W.; et al. Taxonomy of Aspergillus section Flavi and their production of aflatoxins, ochratoxins and other mycotoxins. Stud. Mycol. 2019, 93, 1–63. [Google Scholar] [CrossRef]
- Cotty, P.J. Aflatoxin-producing potential of communities of Aspergillus section Flavi from cotton producing areas in the United States. Mycol. Res. 1997, 101, 698–704. [Google Scholar] [CrossRef] [Green Version]
- Ehrlich, K.C.; Cotty, P.J. An isolate of Aspergillus flavus used to reduce aflatoxin contamination in cottonseed has a defective polyketide synthase gene. Appl. Microbiol. Biotechnol. 2004, 65, 473–478. [Google Scholar] [CrossRef]
- Atehnkeng, J.; Ojiambo, P.S.; Donner, M.; Ikotun, T.; Sikora, R.A.; Cotty, P.J.; Bandyopadhyay, R. Distribution and toxigenicity of Aspergillus species isolated from maize kernels from three agro-ecological zones in Nigeria. Int. J. Food Microbiol. 2008, 122, 74–84. [Google Scholar] [CrossRef]
- Cardwell, K.F.; Cotty, P.J. Distribution of Aspergillus section Flavi among field soils from the four agroecological zones of the Republic of Bénin, West Africa. Plant Dis. 2002, 86, 434–439. [Google Scholar] [CrossRef] [Green Version]
- Probst, C.; Schulthess, F.; Cotty, P.J. Impact of Aspergillus section Flavi community structure on the development of lethal levels of aflatoxins in Kenyan maize (Zea mays). J. Appl. Microbiol. 2010, 108, 600–610. [Google Scholar] [CrossRef]
- Atehnkeng, J.; Ojiambo, P.S.; Ikotun, T.; Sikora, R.A.; Cotty, P.J.; Bandyopadhyay, R. Evaluation of atoxigenic isolates of Aspergillus flavus as potential biocontrol agents for aflatoxin in maize. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2008, 25, 1264–1271. [Google Scholar] [CrossRef]
- Atehnkeng, J.; Ojiambo, P.S.; Cotty, P.J.; Bandyopadhyay, R. Field efficacy of a mixture of atoxigenic Aspergillus flavus Link: FR vegetative compatibility groups in preventing aflatoxin contamination in maize (Zea mays L.). Biol. Control 2014, 72, 62–70. [Google Scholar] [CrossRef]
- Pankaj, S.K.; Shi, H.; Keener, K.M. A review of novel physical and chemical decontamination technologies for aflatoxin in food. Trends Food Sci. Technol. 2018, 71, 73–83. [Google Scholar] [CrossRef]
- Wang, B.; Mahoney, N.E.; Pan, Z.; Khir, R.; Wu, B.; Ma, H.; Zhao, L. Effectiveness of pulsed light treatment for degradation and detoxification of aflatoxin B1 and B2 in rough rice and rice bran. Food Control. 2016, 59, 461–467. [Google Scholar] [CrossRef]
- Luo, X.; Wang, R.; Wang, L.; Li, Y.; Bian, Y.; Chen, Z. Effect of ozone treatment on aflatoxin B1 and safety evaluation of ozonized corn. Food Control. 2014, 37, 171–176. [Google Scholar] [CrossRef]
- Kamala, A.; Kimanya, M.; Lachat, C.; Jacxsens, L.; Haesaert, G.; Kolsteren, P.; Ortiz, J.; Tiisekwa, B.; Meulenaer, B.D. Risk of Exposure to Multiple Mycotoxins from Maize-Based Complementary Foods in Tanzania. J. Agric. Food Chem. 2017, 65, 7106–7114. [Google Scholar] [CrossRef]
- Chulze, S.N. Strategies to reduce mycotoxin levels in maize during storage: A review. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2010, 27, 651–657. [Google Scholar] [CrossRef]
- Degola, F.; Berni, E.; Restivo, F.M. Laboratory tests for assessing efficacy of atoxigenic Aspergillus flavus strains as biocontrol agents. Int. J. Food Microbiol. 2011, 146, 235–243. [Google Scholar] [CrossRef] [PubMed]
- Bandyopadhyay, R.; Ortega-Beltran, A.; Akande, A.; Mutegi, C.; Atehnkeng, J.; Kaptoge, L.; Senghor, A.L.; Adhikari, B.N.; Cotty, P.J. Biological control of aflatoxins in Africa: Current status and potential challenges in the face of climate change. World Mycotoxin J. 2016, 9, 771–789. [Google Scholar] [CrossRef] [Green Version]
- Ezekiel, C.N.; Atehnkeng, J.; Odebode, A.C.; Bandyopadhyay, R. Distribution of aflatoxigenic Aspergillus section Flavi in commercial poultry feed in Nigeria. Int. J. Food Microbiol. 2014, 189, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Moore, G.G.; Olarte, R.A.; Horn, B.W.; Elliott, J.L.; Singh, R.; O’neal, C.J.; Carbone, I. Global population structure and adaptive evolution of aflatoxin-producing fungi. Ecol. Evol. 2017, 7, 9179–9191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- O’Donnell, K.; Corby Kistler, H.; Cigelnik, E.; Ploetz, R.C. Multiple Evolutionary Origins of the Fungus Causing Panama Disease of Banana: Concordant Evidence from Nuclear and Mitochondrial Gene Genealogies. Volume 95, Applied Biological Sciences Communicated by R. 1998. Available online: www.pnas.org (accessed on 2 June 2021).
- Glass, N.L.; Donaldson, G.C. Development of Primer Sets Designed for Use with the PCR to Amplify Conserved Genes from Filamentous Ascomycetes. Appl. Environ. Microbiol. 1995, 61, 1323–1330. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Probst, C.; Bandyopadhyay, R.; Cotty, P.J. Diversity of aflatoxin-producing fungi and their impact on food safety in sub-Saharan Africa. Int. J. Food Microbiol. 2014, 174, 113–122. [Google Scholar] [CrossRef]
- Valencia-Quintana, R.; Milić, M.; Jakšić, D.; Klarić, M.Š.; Tenorio-Arvide, M.G.; Pérez-Flores, G.A.; Bonassi, S.; Sánchez-Alarcón, J. Environment changes, aflatoxins, and health issues, a review. Int. J. Environ. Res. Public Health 2020, 17, 7850. [Google Scholar] [CrossRef]
- Lillehoj, E.B.; Wall, J.H.; Bowers, E.J. Preharvest Aflatoxin Contamination: Effect of Moisture and Substrate Variation in Developing Cottonseed and Corn Kernels. Appl. Environ. Microbiol. 1987, 53, 584–586. [Google Scholar] [CrossRef] [Green Version]
- Kim, N.Y.; Lee, J.H.; Lee, I.; Ji, G.E. An evaluation of aflatoxin and cyclopiazonic acid production in Aspergillus oryzae. J. Food Prot. 2014, 77, 1010–1016. [Google Scholar] [CrossRef]
- de Almeida, Â.B.; Corrêa, I.P.; Furuie, J.L.; de Farias Pires, T.; do Rocio Dalzoto, P.; Pimentel, I.C. Inhibition of growth and ochratoxin A production in Aspergillus species by fungi isolated from coffee beans. Braz. J. Microbiol. 2019, 50, 1091–1098. [Google Scholar] [CrossRef]
- Use of Biocontrol for Aflatoxin Prevention and Control in the EAC. In EAC Policy Brief on Aflatoxin Prevention and Control Policy Brief No. 5; EAC: Arusha, United Republic of Tanzania, 2018.
- Okayo, R.O.; Andika, D.O.; Dida, M.M.; K’otuto, G.O.; Gichimu, B.M. Morphological and Molecular Characterization of Toxigenic Aspergillus flavus from Groundnut Kernels in Kenya. Int. J. Microbiol. 2020, 2020, 8854718. [Google Scholar] [CrossRef]
- Tam, E.W.T.; Chen, J.H.K.; Lau, E.C.L.; Ngan, A.H.Y.; Fung, K.S.C.; Lee, K.C.; Lam, C.-W.; Yuen, K.-Y.; Lau, S.K.P.; Woo, P.C.Y. Misidentification of Aspergillus nomius and Aspergillus tamarii as Aspergillus flavus: Characterization by internal transcribed spacer, β-tubulin, and calmodulin gene sequencing, metabolic fingerprinting, and matrix-assisted laser desorption ionization-ti. J. Clin. Microbiol. 2014, 52, 1153–1160. [Google Scholar] [CrossRef] [Green Version]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef] [PubMed]
- Khalil, A.M.A.; Hashem, A.H. Morphological changes of conidiogenesis in two Aspergillus species. J. Pure Appl. Microbiol. 2018, 12, 2041–2048. [Google Scholar] [CrossRef] [Green Version]
- Yang, X.; An, R. Determination of Aflatoxins (B1, B2, G1 and G2) in Corn and Peanut Butter by Immunoaffinity Cleanup and Post-Column Electrochemical Derivatization. J AOAC Int. 2007, 91, 511–523. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]

| Percentage (%) | |||
|---|---|---|---|
| Aspergillus flavus | Aspergillus niger | Aspergillus tamarii | |
| Cassava | 0.00 | 1.23 | 0.00 |
| Groundnuts | 7.41 | 3.70 | 1.23 |
| Maize | 11.11 | 3.70 | 4.94 |
| Millet | 3.70 | 2.47 | 0.00 |
| Rice | 8.64 | 1.23 | 2.47 |
| Sesame | 13.58 | 4.94 | 1.23 |
| Sorghum | 18.52 | 4.94 | 3.70 |
| Sunflower | 0.00 | 1.23 | 0.00 |
| Isolates | Locations | Food Types | Aflatoxin Chemotypes | |||
|---|---|---|---|---|---|---|
| AFG2 | AFG1 | AFB2 | AFB1 | |||
| A635 | Lamwo | Sorghum | N | N | P | P |
| A348 | Kitgum | Maize | N | N | P | P |
| A584 | Gulu | Sesame | N | N | P | P |
| A490 | Kitgum | Sesame | N | N | P | P |
| A655 | Lira | Maize | N | N | P | P |
| A558 | Pader | Maize | N | N | P | P |
| ASPUN1 | Nwoya | Sorghum | N | N | P | P |
| A508 | Pader | Groundnuts | P | P | P | P |
| A617 | Gulu | Maize | N | N | P | P |
| A613 | Pader | Rice | N | N | P | P |
| ASPE1 | Omoro | Sorghum | N | N | P | P |
| A548 | Amuru | Maize | N | N | P | P |
| A506 | Oyam | Maize | N | N | P | P |
| A491 | Kole | Sesame | N | N | P | P |
| A580 | Kitgum | Sorghum | N | N | P | P |
| A507 | Kole | Groundnuts | N | N | P | P |
| A631 | Kitgum | Millet | N | N | P | P |
| A334 | Gulu | Maize | N | N | P | P |
| A633 | Lira | Sorghum | N | N | P | P |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wokorach, G.; Landschoot, S.; Lakot, A.; Karyeija, S.A.; Audenaert, K.; Echodu, R.; Haesaert, G. Characterization of Ugandan Endemic Aspergillus Species and Identification of Non-Aflatoxigenic Isolates for Potential Biocontrol of Aflatoxins. Toxins 2022, 14, 304. https://doi.org/10.3390/toxins14050304
Wokorach G, Landschoot S, Lakot A, Karyeija SA, Audenaert K, Echodu R, Haesaert G. Characterization of Ugandan Endemic Aspergillus Species and Identification of Non-Aflatoxigenic Isolates for Potential Biocontrol of Aflatoxins. Toxins. 2022; 14(5):304. https://doi.org/10.3390/toxins14050304
Chicago/Turabian StyleWokorach, Godfrey, Sofie Landschoot, Amerida Lakot, Sidney Arihona Karyeija, Kris Audenaert, Richard Echodu, and Geert Haesaert. 2022. "Characterization of Ugandan Endemic Aspergillus Species and Identification of Non-Aflatoxigenic Isolates for Potential Biocontrol of Aflatoxins" Toxins 14, no. 5: 304. https://doi.org/10.3390/toxins14050304
APA StyleWokorach, G., Landschoot, S., Lakot, A., Karyeija, S. A., Audenaert, K., Echodu, R., & Haesaert, G. (2022). Characterization of Ugandan Endemic Aspergillus Species and Identification of Non-Aflatoxigenic Isolates for Potential Biocontrol of Aflatoxins. Toxins, 14(5), 304. https://doi.org/10.3390/toxins14050304






