Abstract
The occurrence of mycotoxins on grapes poses a high risk for food safety; thus, it is necessary to implement effective prevention methods. In this work, a metagenomic approach revealed the presence of important mycotoxigenic fungi in grape berries, including Aspergillus flavus, Aspergillus niger aggregate species, or Aspergillus section Circumdati. However, A. carbonarius was not detected in any sample. One of the samples was not contaminated by any mycotoxigenic species, and, therefore, it was selected for the isolation of potential biocontrol agents. In this context, Hanseniaspora uvarum U1 was selected for biocontrol in vitro assays. The results showed that this yeast is able to reduce the growth rate of the main ochratoxigenic and aflatoxigenic Aspergillus spp. occurring on grapes. Moreover, H. uvarum U1 seems to be an effective detoxifying agent for aflatoxin B1 and ochratoxin A, probably mediated by the mechanisms of adsorption to the cell wall and other active mechanisms. Therefore, H. uvarum U1 should be considered in an integrated approach to preventing AFB1 and OTA in grapes due to its potential as a biocontrol and detoxifying agent.
Keywords:
metagenomics; biological control; Hanseniaspora uvarum; detoxification; aflatoxin B1; ochratoxin A Key Contribution:
The study of the mycobiota revealed a change in the distribution of toxigenic species with an increased relevance of aflatoxigenic species and an absence of Aspergillus carbonarius in Spanish grapes. Hanseniaspora uvarum U1 has been proposed as an effective agent both to control fungal growth and to detoxify aflatoxin B1 and OTA.
1. Introduction
Mycotoxins are secondary metabolites produced by a variety of filamentous fungi such as Aspergillus spp., Penicillium spp., Alternaria spp., and Fusarium spp. There are more than 500 types of mycotoxins, which are toxic to vertebrates and other animals after exposure [1]. Their ingestion may cause acute or chronic diseases depending on the dose and exposure time, and they have mutagenic, teratogenic, and/or carcinogenic properties [2]. Recent studies estimated that about 60–80% of global crops are contaminated by mycotoxins [3].
Spain is the third-largest wine producer worldwide and represents 14% of the global vineyard areas [4]. Several mycotoxins have been reported on grapes such as ochratoxin A (OTA), aflatoxin B1 (AFB1), patulin, fumonisin B2, alternariol, alternariol monomethyl ether, tenuazonic acid, and citrinin [5]. These mycotoxins are produced by different fungi reported on grapes such as Aspergillus spp., Penicillium spp., or Alternaria spp. [5]. Nevertheless, OTA remains to be the most commonly reported mycotoxin on grapes and their derivatives [6]. Due to its frequent occurrence and important toxic effects, OTA levels in grape products are strongly controlled by the current European legislation, which limits its concentration to 2 μg/kg in wine and grape juice [7]. Even though AFB1 on grapes is not as relevant as OTA and its presence on wine is not regulated yet, it cannot be ignored. Several authors have described its presence on grapes and wine from various regions, probably due to the wider distribution of its producing fungi due to climate change [8,9,10,11].
In this context, Aspergillus ochratoxigenic species are the predominant mycotoxin producers in grapes and their distribution is related to specific vineyard geographic locations and climatic conditions [5]. Aspergillus carbonarius and Aspergillus niger aggregate species are considered the most prevalent OTA producers in southern European regions and, specifically, in Spain [6,12,13]. Species belonging to the Aspergillus section Circumdati such as Aspergillus westerdijkiae or Aspergillus steynii have also been isolated from grapes, although their contribution to OTA contamination on grapes and derivatives is low [6]. The emerging occurrence of AFB1 is predominantly due to Aspergillus parasiticus and Aspergillus flavus, which are both occasionally isolated from grapes [11].
Control of mycotoxin contamination is focused on two strategies based on prevention of the development of mycotoxigenic fungi and/or detoxification [14]. The application of agricultural practices such as proper use of antifungals or controlling harvesting time is known to prevent the development of mycotoxigenic fungi in vineyards [5,15,16]. Currently, consumers demand harmless foodstuffs with an environmentally friendly production, which has led research to focus on the development of novel methods to prevent mycotoxin contamination [17]. Among these new approaches, biological control using antagonistic microorganisms such as yeasts or lactic acid bacteria is an emerging and very promising alternative [18,19].
Grape microbiota is extremely complex and includes a variety of bacteria, yeasts, and filamentous fungi. Among them, different mycotoxigenic species might occur but, in fact, these plant communities are also the source of most biocontrol agents described up to now [20]. Next-generation sequencing provides a useful tool to understand the complexity of grape microbiota and might be of interest in the search for new biological control agents (BCA) [20,21,22]. Yeasts represent the predominant group of microorganisms on the grape surface and are mainly represented by the yeast-like fungus Aureobasidium pullulans, oxidative or weakly fermentative ascomycetous (i.e., Hanseniaspora spp., Candida spp., Metschnikowia spp., or Pichia spp.), basidiomycetous populations (i.e., Cryptococcus, Rhodotorula spp., or Sporobolomyces spp.), and, in some cases, highly fermentative wine spoilage yeasts such a Zygosaccharomyces spp. or Torulaspora spp. [23].
Different biological methods have been proposed for controlling OTA in food products based on the application of yeasts, bacteria, or nontoxic fungi, which reduce fungal growth or decrease OTA levels [24]. In this context, epiphytic yeasts represent the most effective group of BCA due to their ability to colonize grapes and compete for space and nutrients with other microorganisms, such as mycotoxigenic fungi [15,25]. As mentioned before, epiphytic yeasts represent the largest community on grape berries and are highly adapted to this ecological niche; therefore, they might be considered effective biocontrol agents for their application in vineyards [22]. Accordingly, Bleve et al. [26] selected several yeasts that naturally occur on grapes (Issatchenkia orientalis, Metschnikowia pulcherrima, and Candida incommunis), which significantly controlled the fungal growth of ochratoxigenic isolates of A. niger and A. carbonarius. In addition, the ability of A. pullulans and Lachacea thermotolerans to reduce fungal growth and OTA biosynthesis by Aspergillus section Nigri species has also been reported [27,28].
Despite the application of control methods to avoid fungal growth, the presence of mycotoxins in foods cannot always be prevented. Moreover, the coexistence of several mycotoxins in the same product is a matter of concern because it is well-known that they have a cumulative and synergistic effect on toxicity and carcinogenicity, even at low levels of exposure [29,30]. Thus, it is necessary to develop new strategies of detoxification to eliminate mycotoxins from food.
Detoxification includes diverse methods (physical, chemical, or biological) that eliminate mycotoxins without generating toxic products or altering the nutritional value and organoleptic properties of foods [31,32]. In general, chemical detoxification is not allowed in the European Union due to biosafety concerns [14]. In the case of wine, OTA detoxification using physical adsorbents such as activated carbon during the fining process has been tested with positive results, although this treatment usually affects wine quality [33]. Considering the limitations of chemical and physical processes, biological detoxification is the most promising approach because it is safe from environmental and health perspectives, and it also maintains the organoleptic properties of foods [34]. These biological methods use microorganisms or their enzymes to degrade, transform, or adsorb mycotoxins and, therefore, avoid the toxic effects produced by the ingestion of contaminated food [35].
In this type of detoxification, it is indispensable that the microorganisms used as well as their degradation products are harmless to both humans and animals [35]. For this reason, current studies are focused on the selection of candidates for mycotoxin detoxification among probiotic microorganisms and other QPS (qualified presumption of safety) species [36]. Several microorganisms, including yeasts such as Saccharomyces cerevisiae, have shown mycotoxin detoxification properties in wines [19,37,38,39].
The aim of this work is to determine, through a metagenomic approach, the diversity of grape berry microbiota to study the occurrence of mycotoxigenic species and the presence of potential BCA candidates. The most relevant potential BCA, Hanseniaspora uvarum U1, was selected for subsequent studies to characterize its ability to control fungal growth of the main OTA- and AFB1-producing fungi found on grapes (A. carbonarius, A. welwitschiae, A. steynii, A. westerdijkiae, A. flavus, and A. parasiticus), as well as its detoxification potential relevant to these mycotoxins.
2. Results
2.1. Analysis of the Mycobiota of Grape Berries
A metagenomic approach was used to study the mycobiota present in grapes using the sequence of the ITS2 region of the rDNA. Rarefaction curves were constructed (Supplementary Figure S1) as a function of sampling depth. All curves reached the plateau and were close to saturation; therefore, the number of clones sequenced was sufficient to fully observe the richness of the samples. The complete results of relative abundance at genus level are included in Figure S2.
Alpha-diversity was studied using Shannon, Chao1, and inverse Simpson indexes (Table 1). No statistically significant differences were found in any diversity index when the effect of the location or the vineyard management was evaluated; all samples analyzed showed similar richness and evenness. The high levels of Chao1 and Shannon indexes are related to rich and uniform fungal communities, and inverse Simpson values indicate no clear dominance of one or more species. Although no differences were reported, M2 showed lower values of the three indexes evaluated, possibly due to the high predominance of Penicillium spp., representing 98% of the total diversity.

Table 1.
Metagenomic analysis of the mycobiota in grape berries. The percentage of abundance related to the total mycobiota of toxigenic fungi and yeasts that could be used as potential biological control agents are shown. Diversity indexes for each sample are also included.
Different mycotoxigenic species were detected in grape berries and their distribution widely varies among samples. Aspergillus section Nigri species were frequently found but represented very different percentages of the total fungal diversity, and A. carbonarius was not detected in any sample. It is important to highlight the presence of A. flavus in all but one grape sample, indicating a potential risk of aflatoxin contamination. Moreover, potential ochratoxigenic Aspergillus section Circumdati species (A. westerdijkiae and A. steynii) were also detected in two of the eight samples. Sample M2 was the only one not contaminated by any potential toxigenic species.
The presence of species that have been reported as potential biocontrol agents against mycotoxigenic fungi was also evaluated. Most of the diversity in all samples was represented by the yeast-like fungus Aureobasidium pullulans, with up to 54% of the total diversity. The yeast Hanseniaspora uvarum was also detected in 75% of the grape samples and, in some cases, represented high percentages of the diversity of the total mycobiota. Other significant yeasts detected that had been described previously as potential biocontrol agents were Debaryomyces hansenii, Lanchacea sp., and Candida sp.
Considering that sample M2 was not contaminated by any mycotoxigenic species, it was selected for the isolation of potential biocontrol agents since the yeasts that occur there might influence the occurrence of these fungi. The most frequently isolated yeast from rose bengal chloramphenicol agar plates was Hanseniaspora uvarum and the isolate U1 was selected for subsequent studies. The identification was confirmed using sequencing of the D1–D2 region. The sequence was deposited in the NCBI database (accession number OK039223).
2.2. Biocontrol
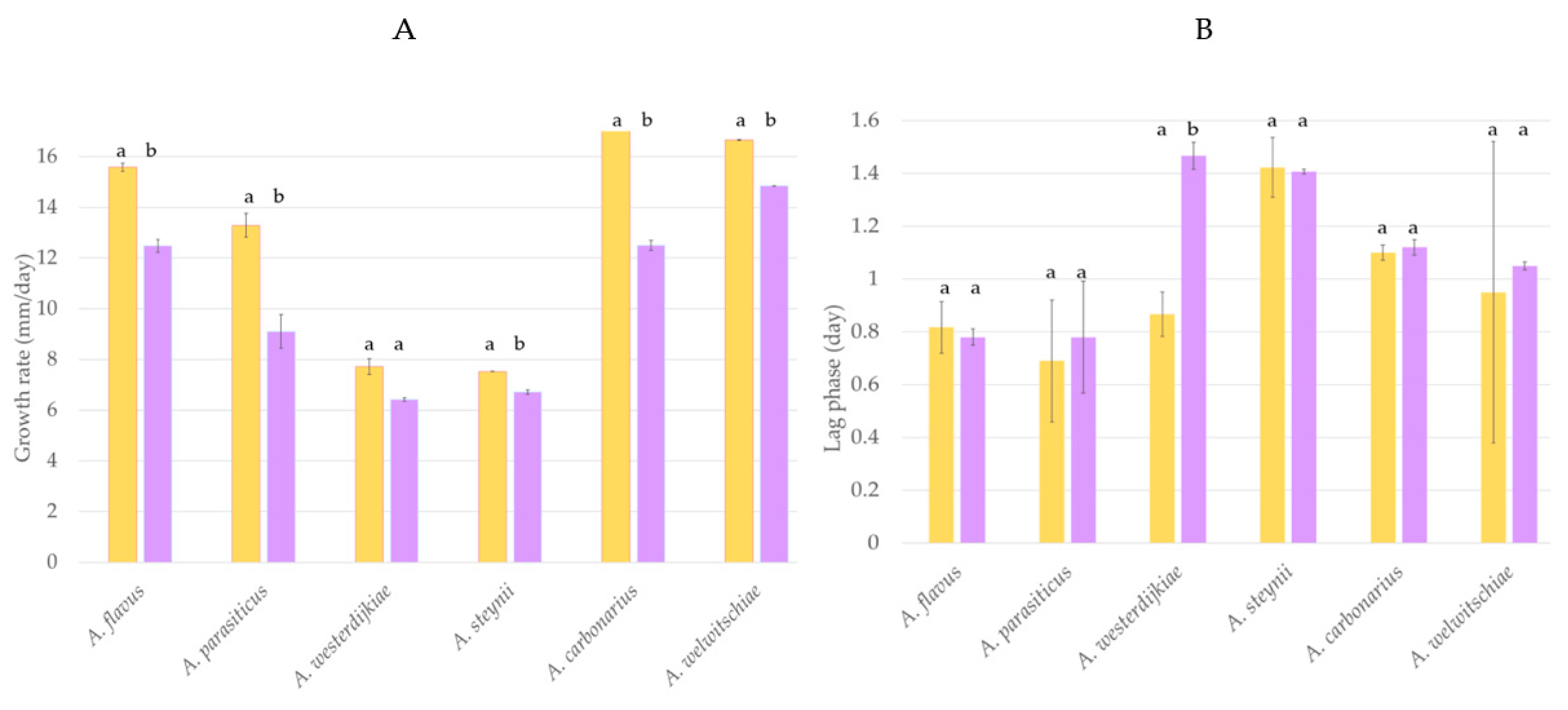
The potential of H. uvarum U1 to prevent fungal growth was evaluated using A. carbonarius, A. welwitschiae, A. westerdijkiae, A. steynii, A. flavus, and A. parasiticus. Results regarding the influence of H. uvarum U1 on the lag phase and growth rate of each fungus are shown in Figure 1. The growth rate of most fungi tested was significantly reduced when H. uvarum U1 was added into the CYA plates. The reduction percentages were 34, 31, and 19% in the case of A. carbonarius, A. parasiticus, and A. flavus, respectively, whereas the growth rates of A. steynii and A. welwitschiae were reduced by 11%. The growth rate of A. westerdijkiae was not affected by the presence of H. uvarum U1, but its lag phase was extended by 75%, suggesting that the potential BCA can control fungal growth in the first steps of the cycle (Figure 1 and Figure S3).
Figure 1.
Growth rate (A) and lag phase (B) of A. flavus, A. parasiticus, A. westerdijkiae, A. steynii, A. carbonarius, and A. welwitschiae cultured in control CYA plates (yellow) and supplemented with H. uvarum U1 (purple). Each value is the mean of two replicates. Thin vertical bars represent the standard error of the corresponding data. Groups with the same letter are not significantly different (p > 0.05).
2.3. Detoxification Ability of Aflatoxin B1 and Ochratoxin A by Hanseniaspora uvarum U1
First, the effect of mycotoxins on yeast development and viability was evaluated. For this purpose, the growth of H. uvarum U1 was measured at a different pH (3.0, 5.5, and 7.0) at 630 nm in the presence and absence of the analyzed concentration of aflatoxin B1 (AFB1) and ochratoxin A (OTA). In addition, the effect of the mycotoxin solvent (methanol) on H. uvarum U1 growth was evaluated. Table 2 shows viable cell (VC) absorbance values, which correspond to the growth of H. uvarum U1. Yeast growth at pH 3.0, 5.5, and 7.0 was not affected by the amount of AFB1, OTA, or methanol used (p-value for pH 3.0 = 0.377; p-value for pH 5.5 = 0.539; p-value for pH 7 = 0.967). Therefore, neither mycotoxins nor methanol reduced the growth of H. uvarum U1.

Table 2.
Absorbance values at 630 nm after culturing Hanseniaspora uvarum U1 on PDB at different pH (3.0, 5.5, and 7.0) and treatments (methanol, AFB1, OTA) after 48 h at 30 °C. Results are mean values ± standard deviations of two samples. For each pH, comparisons were performed between different treatments. There were no significant differences between control and treatments (methanol and mycotoxins) in any case (p < 0.05).
The ability of viable cells (VC) and heat-inactivated cells (HIC) of H. uvarum U1 to remove AFB1 and OTA at different pH (3.0, 5.5, and 7.0) was evaluated by ELISA. As shown in Table 3, H. uvarum U1 cells were able to significantly reduce AFB1 and OTA at all pH and treatments tested. In general, the highest reduction values were found for AFB1. The greatest AFB1 removal was observed at pH 5.5 (VC = 99.08%; HIC = 97.68%). There were significant differences in the removal percentage of AFB1 between VC and HIC H. uvarum U1 when incubated at pH 3 (VC = 93.68%; HIC = 83.84%) and pH 7.0 (VC = 98.29%; HIC = 95.78%), and VC was more efficient at detoxifying AFB1. These differences between VC and HIC were not observed at pH 5.5.

Table 3.
Percentages of mycotoxin reduction by viable cells (VC) and heat-inactivated cells (HIC) of H. uvarum U1 obtained at different pH (3.0, 5.5, and 7.0) after 48 h of incubation at 30 °C and 30 min. Results are mean values ± standard deviations of two samples. Comparisons were performed for each mycotoxin independently. Different letters (a–c) in the same column (pH) indicate a significant effect among control, VC, and HIC. Different letters (y–z) in the same row (treatment) indicate a significant effect of pH (p < 0.05).
In the case of OTA, the maximum removal ability of VC was observed at pH 7.0 (82.96%), whereas for HIC, the highest percentage of reduction occurred at pH 5.5 (79.87%). Significant differences were found at pH 3.0, and the highest reduction percentage was found in the case of VC.
In general, the percentage reduction of AFB1 and OTA was higher in viable cells than in HIC. The lowest percentages of AFB1 and OTA detoxification occurred at pH 3.0 and with inactivated cells.
3. Discussion
Ochratoxin A (OTA) is considered the most important mycotoxin on grapes and wines [6]. However, new weather conditions as a result of climate change favor the growth of aflatoxin B1-producing fungi and, therefore, AFB1 is an emerging mycotoxin in grapes and grape products [11]. The impact of these mycotoxins on health and the economy has led to an increase in the search for effective measures to reduce mycotoxigenic fungi. Although the effectiveness of chemical methods to control mycotoxigenic fungi has been established, there is a growing public demand and deep concern within the European Union to find safer and more environmentally friendly alternatives to avoid mycotoxins in foodstuffs [40]. Biological control is now considered a suitable approach to prevent fungal pathogens on grapes in both preharvest and postharvest [41,42] and has even been proposed as an effective measure to reduce the development of mycotoxigenic fungi [16,24]. Among potential biological control agents (BCA), yeasts present particular characteristics that make them considered the most adaptable organisms for use in grapes [22].
The metagenomic approach performed in this work was useful for studying the fungal mycobiota on grape berries and, specifically, the occurrence of potential mycotoxigenic fungi. Although the presence of aflatoxigenic species has not been studied systematically, there are several reports available that describe the presence of A. flavus and A. parasiticus on grapes [11]. In this work, A. flavus was consistently detected in almost all samples analyzed, which might be related to an important risk of contamination by aflatoxins. Several articles have pointed out that A. flavus is an emerging problem mainly in temperate, southern regions of Europe, since it is able to grow and produce aflatoxins under these conditions [43]. Therefore, this work might be a reflection of the expansion of A. flavus distribution across Europe and its increased occurrence in unconventional matrices.
Surprisingly, A. carbonarius was not detected in any of the samples despite being reported for a long time as the predominant ochratoxigenic fungus in grapes, mainly in Mediterranean vineyards [5,6,44]. The findings of this work might agree with those reported by Cervini et al. [45], who demonstrated that the climate change scenario predicted for this region might affect A. carbonarius development and its ability to produce OTA in grapes. However, the risk posed by OTA contamination should not be disregarded since other important producers included in the section Circumdati such as A. steynii and A. westerdijkiae have been detected in several samples. Aspergillus niger aggregate species were also present in some grape samples and, although their importance as OTA producers is discussed, their presence might lead to FB2 contamination promoted by the new forecast conditions of the climate change scenario [10].
The metagenomic analysis of fungal microbiota also allowed the identification of potential biocontrol agents among the epiphytic yeasts. The results regarding yeast occurrence on grape berries are similar to those previously reported by other authors [23]. Aureobasidium pullulans yeast-like fungus was by far the most representative group of fungal microbiota. This is an indicator of its ability to establish and dominate in this ecosystem, which is one of the required characteristics of a potential biocontrol agent. However, A. pullulans has been related to human infections in immunocompromised patients and it is not included in the GRAS and QPS lists. Therefore, its potential use in the environment is not advisable without performing prior in-depth studies [46,47]. Moreover, this yeast-like fungus is not included in the GRAS and QPS lists, and its inclusion is unlikely due to its possibility of being a human pathogen. The second most frequent yeast in grape berries was Hanseniaspora uvarum. Several works have reported that this yeast can occupy 50% of the total population on grapes [48,49]. Non-Saccharomyces yeasts such as H. uvarum play an important role in the early stages of fermentation as they can release hydrolytic enzymes that improve the aromatic and sensory properties of wines [49,50]. Moreover, H. uvarum has been previously proposed as an effective BCA against different fungal pathogens in grapes included in the genera Aspergillus, Penicillium, Alternaria, and Botrytis [51,52,53,54,55]. Therefore, this species was considered for isolation from grape samples since it could be a suitable candidate as a biocontrol agent against mycotoxigenic fungi on grapes.
Hanseniaspora uvarum U1 was isolated from grapes sampled in a conventionally managed vineyard from Spain. Although H. uvarum was detected in all samples, the grapes sampled in this vineyard (M2) were not contaminated by any mycotoxigenic species and, therefore, it could be possible that some effective BCA was present and controlling fungal development. The presence of H. uvarum U1 was demonstrated to affect the fungal growth of important species frequently occurring in grapes that are able to produce both aflatoxins and OTA. Aspergillus westerdijkiae was the only fungi not significantly affected by the presence of H. uvarum. However, a potential BCA for controlling mycotoxins in foodstuffs might not only affect fungal growth and its ability to colonize the product but also inhibit or reduce mycotoxin biosynthesis [56]. Prendes et al. [55] recently reported that though H. uvarum can stimulate Alternaria alternata growth, it also produces high inhibition rates of mycotoxin production. Therefore, further studies are necessary to unravel the effect of H. uvarum U1 on the ability to produce mycotoxins by these Aspergillus species.
Although controlling fungal growth is considered the most effective method for avoiding mycotoxins, in some cases and conditions, it cannot be implemented. In this context, some yeasts present desirable characteristics for application as biological detoxifying agents by adsorption or degradation of mycotoxins [19]. Therefore, in this work, the potential of H. uvarum U1 to detoxify AFB1 and OTA was also tested. Some mycotoxins present cytotoxic effects, and some yeasts can be highly sensitive; therefore, yeast growth and survival should be tested before the optimization of a detoxifying agent [19]. At the doses tested, neither AFB1 nor OTA affected H. uvarum development, which is an advantage for the future application of viable cells for mycotoxin decontamination.
To our knowledge, there is only one previous report that describes the ability of this yeast to detoxify mycotoxins [57]. In a preliminary study, H. uvarum U1 showed a high mycotoxin detoxification ability since it was able to remove 99% and 83% of AFB1 and OTA, respectively.
Mycotoxin removal by microorganisms can occur by biodegradation and/or by adsorption to the cell walls [19,58,59]. To determine its application in food products, it is essential to evaluate whether one or both mechanisms are involved in the elimination of AFB1 and OTA by H. uvarum U1; therefore, experiments were performed using viable and heat-inactivated cells. AFB1 and OTA were efficiently removed by both cells, which indicates that adsorption may be one of the mechanisms used by H. uvarum U1 to detoxify these mycotoxins. Adsorption is a physical process that does not depend on the viability or physiological state of the cell and can also occur in inactivated cells. However, it has been claimed that heat treatment might result in an increased exposure of binding sites on nonviable cells [60], although this effect has not been observed in the case of H. uvarum U1. In general, viable cells were more efficient in mycotoxin removal than heat-inactivated yeasts and, therefore, other mechanisms might be involved in H. uvarum U1′s detoxification capacity.
Subsequently, in this work, the ability of H. uvarum U1 to detoxify AFB1 and OTA at different pH was analyzed. Several authors described pH as an important factor involved in mycotoxin adsorption, especially when electrostatic interactions are involved [61]. Under these circumstances, pH can affect the degree of ionization of mycotoxins and the charge distribution of microorganism wall components. AFB1 is a nonionizable molecule and its adsorption should not be drastically affected by pH [61]. The results of this work indicate a lower reduction of AFB1 at pH 3.0, although the differences in adsorption with respect to other pH are not as pronounced as those observed in OTA. In the case of OTA adsorption by yeasts, several studies reported that acidic conditions increase detoxification values when compared to alkaline pH [62,63]. The acidic character of OTA (pKa 4.4) causes the protonated and uncharged forms to be present in acidic solutions (pH < 4.0), which is supposed to enable its binding to yeast wall components such as glucans [64]. These components provide numerous binding sites for mycotoxin adhesion. Glucans represent up to 60% of the cell wall of Saccharomyces cerevisiae, and, therefore, most research has been focused on them [60,65,66]. However, the ability of H. uvarum U1 to detoxify OTA was strongly reduced in acidic conditions, which might be due to its unusual wall composition, being mostly enriched with proteins [67]. Moreover interactions involving basic amino acids are highly pH-dependent, which may affect in some way the OTA binding to the cell wall of H. uvarum when subjected to such an acidic pH
The adsorption process also depends on the three-dimensional conformation of the cell wall components. Yiannikouris et al. [68] described a higher affinity of AFB1 for the conformation of binding sites offered by β-d-glucans compared to OTA. Although the percentage of glucans in H. uvarum cell wall is low, this fact could explain why, under the same pH conditions, elimination is higher in the case of AFB1.
In this study, viable H. uvarum U1 cells showed a greater ability to reduce AFB1 and OTA than heat-inactivated cells. These results indicate that the detoxification process also involved an active mechanism, such as degradation. Angioni et al. [57] described the ability of H. uvarum to degrade OTA but did not unravel the pathway involved. Other articles reported that Trichosporon, Rhodotorula, Cryptococcus, and Aureobasidium pullulans are able to degrade OTA by cleavage of the amide bond [69,70]. Hydrolytic enzymes such as carboxypeptidases, ochratoxinases, proteases, and lipases have been identified in most of the described microorganisms as agents capable of degrading OTA into less-toxic by-products [35,71]. The highest OTA removal by H. uvarum U1 was found at conditions in which these enzymes have maximum hydrolytic activity (pH between 6.0 and 7.5) [71,72] and, therefore, a degradation mediated by these hydrolytic enzymes might be involved in the detoxification ability of this yeast. Recent work reported that strains of Pichia occidentalis, Candida sorboxylosa, and Hanseniaspora opuntiae isolated from kombucha are able to degrade AFB1 into less-toxic products [73]. Furthermore, Li et al. [74] postulated the mechanism by which Candida versatilis CGMCC 3790 degrades AFB1 and found that the optimal degradation condition was at pH 5.0. These results agree with those observed in H. uvarum U1, although more research is needed to characterize the degradation process.
Vanhoutte et al. [75] established the ideal features of a detoxifying agent that include the following: (I) the process should be fast and efficient, (II) the process should occur on more than one mycotoxin, (III) the agent must not be pathogenic, (IV) the resulting degradation products must not be toxic, and (V) the degradation must occur under conditions that are relevant to the contaminated matrix. Hanseniaspora uvarum U1 accomplishes most of these tasks, since it is able to remove both AFB1 and OTA very effectively in a short time and, additionally, it is qualified as a QPS agent [76].
4. Conclusions
This work presents an integrated approach to prevent AFB1 and OTA in grapes. The study of the mycobiota by a metagenomic approach allowed the characterization of both mycotoxigenic risk and potential biocontrol agents. Moreover, Hanseniaspora uvarum U1 was demonstrated to be a promising biocontrol agent since it is capable of controlling fungal growth in different aflatoxigenic and ochratoxigenic species. Moreover, this yeast is able to detoxify AFB1 and OTA extracts in a wide range of conditions, and the most probable detoxification pathway includes both adsorption to the cell wall and an active mechanism.
5. Materials and Methods
5.1. Sample Collection
Samples were collected in four locations in Spain (Toledo, M1 and M2; Valencia, M3 and M4; Burgos, M5 and M6; Madrid, M7 and M8), and two vineyards were selected from each site to include both ecological and conventional practices (even and odd sample number, respectively). For each vineyard, the samples were collected at random, taking one grape cluster every 5 m along the plot, reaching a total of approximately 3 kg of grapes. At the laboratory, all samples were manually destemmed, mixed, and divided into lots of 100 g. Subsequently, the subsamples were processed using a lyophilizer (Cryodos, Telstar, Madrid, Spain) and stored at 4 °C until analysis.
5.2. DNA Extraction and Metagenomic Analysis
DNA isolation from lyophilized grapes was performed using the DNeasy Plant Mini Kit (QIAgen, Hilden, Germany) following the manufacturer’s instructions. Three DNA extractions were carried out for each sample and were subsequently mixed and concentrated using a vacuum concentrator (Concentrator plus, Eppendorf, Hamburg, Germany).
For metagenomic analysis of fungal microbiota, the ITS2 region of ribosomal DNA was selected and amplified using ITS3 and ITS4 universal primers [77]. Amplicon libraries were prepared using Herculase II Fusion DNA Polymerase Nextera XT Index Kit V2 (Illumina, Foster City, USA) and sequenced on a MiSeq Illumina Platform in Macrogen facilities (Macrogen Inc., Seoul, Korea).
Data processing and analysis were performed using different bioinformatic software, which processed and analyzed the obtained sequence data. Preliminary processing of the data was performed using FLASH 1.2.11 [78] to merge paired-end reads. Subsequently, CD-HIT-OTU 4.5.5 [79] was used to filter and trim raw sequences, which were then clustered into operational taxonomical units (OTU). Taxonomy assignments were performed using the UNITE database [80]. Further analyses were carried out using QIIME v. 1.8 [81], and diversity indexes such as alpha diversity, Shannon, and Chao1 were estimated.
5.3. Isolation and Identification of Hanseniaspora uvarum Isolated from Grapes
The source of yeast isolation was sample M2 from Toledo taken in a conventionally managed vineyard. This sample was selected due to the absence of mycotoxigenic fungi, which might be related to the presence of established microorganisms that can effectively compete in this substrate. For this purpose, surface-disinfected grapes were cultured on rose bengal chloramphenicol agar plates (Pronadisa, Madrid, Spain) for 48 h at 28 °C to recover the maximum number of yeast species present in the grapes. Isolated colonies were then selected and reisolated on potato dextrose agar plates (PDA, Pronadisa, Madrid, Spain). All yeasts were stored in 15% glycerol (Fisher Chemical, Loughborough, UK) at −80 °C until required.
Before DNA extraction, the isolated yeasts were cultured in 20 mL of potato dextrose broth (PDB) (Pronadisa, Madrid, Spain) at 28 ± 1 °C in an orbital shaker SK-0330-PRO (140 rpm) (Labolan, Esparza, Spain) in darkness. Subsequently, 1 mL of the culture was centrifuged, the supernatant removed, and DNA extraction was performed. Genomic DNA extraction from yeast isolates was carried out using a protocol described elsewhere [34].
The identification was performed by sequencing the variable D1–D2 domain of the 26S subunit region using the protocols described by Kurtzman and Robnett [82]. PCR assays were performed in an Eppendorf Mastercycler Gradient (Eppendorf, Hamburg, Germany). All amplification reactions were carried out in a final volume of 25 μL, containing 100 ng of sample DNA, 1 μL of each primer (20 μM) (Metabion, Planneg, Germany), and 12.5 μL NZYTaq II 2× Green Master Mix (NZYTech, Lisbon, Portugal). PCR-amplified products were visualized in a 1.5% agarose gel electrophoresis (Pronadisa, Madrid, Spain) using 1× TAE buffer (Tris-acetate 40 mM and EDTA 1.0 mM) and 3 μL of Green Safe Premium (1 μg/mL) (NZYTech, Lisbon, Portugal). The NZYDNA Ladder V (NZYTech, Lisbon, Portugal) was used as a molecular size marker. The electrophoresis was performed at 80 V for 25 min and then visualized under UV light (ETX-20-M, Vilber Lourmat, France).
The amplification products were purified with a NZYGelpure Kit (NZYTech, Lisbon, Portugal). Sequencing was carried out on an ABI PRISM 3730XL DNA sequencer (Applied Biosystems, Foster City, USA) according to the manufacturer’s instructions in Macrogen facilities (Madrid, Spain). All amplification products were sequenced in both directions.
The sequences were assembled using the UGENE v33.0 package. Sequences were compared with those deposited on the NCBI nucleotide database using Blast to extend their identification to species level.
5.4. Biological Control Using Hanseniaspora uvarum U1
Aflatoxin-producing strains of A. flavus (A23) and A. parasiticus (A19) as well as OTA-producing strains of A. carbonarius (350), A. westerdijkiae (AODP 16-1), A. welwitschiae (N5-M16), and A. steynii (3.53) were selected to test the potential of H. uvarum U1 to reduce fungal growth in vitro. Fungal strains were cultured on potato dextrose agar (PDA) and incubated at 28 °C for five days. Subsequently, spore suspensions were prepared in sterile saline solution (9 g/L sodium chloride), and the concentration was determined using a Thoma counting chamber (Marienfeld, Lauda-Königshofen, Germany) and adjusted to a final concentration of 106 spores/mL. Hanseniaspora uvarum was cultured in PDA plates for 48 h at 28 °C and a cell suspension was prepared following the same method used for spore suspensions.
The effect of the presence of yeast on fungal growth was evaluated in CYA medium (45.5 g/L of modified Czapek–Dox agar (Pronadisa, Madrid, Spain) and 5 g/L of yeast extract (Pronadisa, Madrid, Spain)). Hanseniaspora uvarum U1 was added to the medium to obtain a final concentration of 103 cells/mL. The same amount of saline solution was included in control plates instead of H. uvarum U1. Both plates were inoculated with 2 µL (4 mm diameter) of each fungal spore suspension in the center of the plate. Fungal colony diameter was measured daily in two directions until the surface of the control plates was fully colonized, which took 5 days for A. carbonarius; 6 days for A. flavus, A. parasiticus, and A. welwitschiae; and 9 days for A. westerdijkiae and A. steynii. Growth parameters were calculated from a linear model by plotting diameter (mm) against time (day). The lag phase (days prior to mycelial growth) and growth rate (mm/day) were calculated both for control plates and H. uvarum-supplemented ones.
5.5. Study of the Detoxification Capacity of Ochratoxin A and Aflatoxin B1 by Hanseniaspora uvarum U1
5.5.1. Preparation of Yeast and Toxin Extracts
Stock solutions of OTA and AFB1 (Sigma-Aldrich, Darmstadt, Germany) were prepared in 5 mL of methanol, to a final concentration of 200,000 μg/L. These initial stock solutions were diluted up to 0.5 μg/L and 1 μg/L, respectively. Concentrations were standardized to add 2.5 μL of AFB1 and OTA to each well.
Hanseniaspora uvarum U1 suspensions were prepared as mentioned above to a final concentration of 1.5 × 108 CFU/mL. These resulting suspensions constituted the viable cells (VC).
5.5.2. Detoxification Assays
The analysis of the detoxification capacity of H. uvarum U1 for AFB1 and OTA was performed in 96-well polystyrene microplates (CorningTM, USA). The viable cells experiment included the following conditions: (I) A growth control—25 μL of the H. uvarum suspension; 200 μL of potato dextrose broth (PDB) (Pronadisa, Madrid, Spain) at pH 3.0, 5.5, or 7.0, and 25 μL of 0.9% w/v NaCl solution to complete the volume. (II) A test sample containing 25 μL of the yeast suspension; 200 μL of PDB at pH 3.0, 5.5, or 7.0; 2.5 μL of AFB1 or OTA; and 22.5 μL of saline solution. (III) Methanol control for toxicity, which included the same components as (i), replacing the 2.5 μL of mycotoxin with methanol. (IV) Mycotoxin control with 200 μL of PDB medium at pH 3.0, 5.5, or 7.0; 2.5 μL of AFB1 or OTA; and 47.5 μL of saline solution. All samples and controls described before were analyzed by duplicate. The microplates were incubated for 48 h at 30 °C without shaking and yeast growth was then evaluated by measuring the turbidity in each well at 630 nm with a plate reader (Dutscher, Bernolsheim, France). Subsequently, samples and controls containing mycotoxins were filtered using 0.22 μm filters (Fisherbrand, Thermo Fisher Scientific, Madrid, Spain), and the filtered products were stored in 1.5 mL Eppendorf tubes at -20 °C until mycotoxin quantification by ELISA.
A similar approach was performed to evaluate the ability of heat-inactivated cells (HIC) of H. uvarum U1 to detoxify OTA and AFB1. As the first step, 25 μL of the H. uvarum U1 suspension was incubated in multi-well plates with 200 μL of PDB and 25 μL of saline solution for 48 h at 30 °C. In this way, there was the same number of cells in both experiments. After incubation, mixtures were inactivated by heat treatment at 121 °C for 20 min in an autoclave and, subsequently, 2.5 μL of AFB1 and OTA extracts were added to each one. These samples were incubated for 30 min at room temperature and then filtered. The filtered products were finally stored at −20 °C until quantification by ELISA.
5.5.3. Quantification of Mycotoxins by ELISA
Quantification of AFB1 and OTA was performed using the RIDASCREEN® Aflatoxin B1 30/15 Art. No. R1211 and RIDASCREEN® Ochratoxin A 30/15 Art. No. R1311 (R-Biopharm, Darmstadt, Germany), respectively, following the manufacturer’s instructions. These kits are based on a competitive ELISA assay, in which the absorbance measured is inversely proportional to the mycotoxin concentration of the sample. The resulting color reaction was measured at 450 nm with a plate reader (Dutscher, Bernolsheim, France).
A six-point standard curve was prepared using samples with different concentrations of AFB1 (0; 1; 5; 10; 20; 50 μg/L) (%(B/B0) = −35,291 log(concentration) + 76,952; R2 = 0.9884) and OTA (0; 0.05; 0.1; 0.3; 0.9; 1.800 μg/L) (%(B/B0) = −50,442 log(concentration) + 167.98; R2 = 0.9607) included in each RIDASCREEN® ELISA kit. First, the percentage of absorbance (% (B/B0)) was calculated by the following formula:
% (B/B0) = (Standard absorbance or sample (B)/Absorbance of blank (B0)) × 100
The absorbance data obtained in the detoxification test were interpolated from the line to obtain the concentration value of AFB1 and OTA present in the samples after incubation. The standard line was made using Microsoft Excel® software (Microsoft Corporation, Washington, USA).
5.6. Statistical Analysis
Statistical analyses were performed with StatGraphics Centurion XVII V.17.2.04 software (Statpoint Technologies Inc., Warrenton, VA, USA). Normality and homoscedasticity of the data were tested by the Shapiro–Wilk and Bartlett tests. In the case of diversity indexes, percentage of mycotoxin reduction in detoxification assays, and growth absorbance values, an analysis of variance (ANOVA) was performed followed by Fisher’s LSD post hoc test for checking differences among group means. For lag phase and growth rate in biological control experiments, a statistical analysis was performed independently for each mycotoxigenic strain using Student’s t-test. In all cases, the significance level was set at p < 0.05.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/toxins13090649/s1, Figure S1, rarefaction curves. Figure S2, relative abundance of the main fungal genera in grape samples, based on the metagenomic analysis of ITS2 sequences. Figure S3, colony growth of A. parasiticus and A. carbonarius on control plates and when incubated in the presence of H. uvarum U1.
Author Contributions
Laboratory work, C.G.-A. and C.M.; data analysis, J.G.-S. and C.G.-A.; writing—original draft preparation, J.G.-S., C.G.-A. and C.M.; writing—review and editing, J.G.-S., B.P. and C.V.; project administration, B.P.; funding acquisition, B.P. and C.V. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Spanish Ministry of Science and Innovation, grant number RTI 2018-097593-B-C21R and the authors thank the AgroBank-UdL Chair “Quality and Innovation in the Agri-Food Sector” for an economic aid for the realization of this work. C.G.A. was funded by a FPI fellowship by the Spanish Ministry of Science and Innovation (PRE-2019-087768). This work conforms the Nagoya Protocol ESNC60 (ABSCH-CNA-ES-240783-2).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Raw data are available upon request, please contact the corresponding author.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Bennett, J.W.; Klich, M. Mycotoxins. Clin. Microbiol. Rev. 2003, 16, 497–516. [Google Scholar] [CrossRef] [Green Version]
- Cimbalo, A.; Alonso-Garrido, M.; Font, G.; Manyes, L. Toxicity of mycotoxins in vivo on vertebrate organisms: A review. Food Chem. Toxicol. 2020, 137, 111161. [Google Scholar] [CrossRef]
- Eskola, M.; Kos, G.; Elliott, C.T.; Hajslova, J.; Mayar, S.; Krska, R. Worldwide contamination of food-crops with mycotoxins: Validity of the widely cited ‘FAO estimate’ of 25%. Crit. Rev. Food Sci. Nutr. 2020, 60, 2773–2789. [Google Scholar] [CrossRef] [PubMed]
- Note de Conjoncture Vitivinicole Mondiale. 2020. Available online: https://www.oiv.int/js/lib/pdfjs/web/viewer.html?file=/public/medias/7899/oiv-note-de-conjoncture-vitivinicole-mondiale-2020.pdf (accessed on 5 July 2021).
- Welke, J.E. Fungal and mycotoxin problems in grape juice and wine industries. Curr. Opin. Food Sci. 2019, 29, 7–13. [Google Scholar] [CrossRef]
- Gil-Serna, J.; Vázquez, C.; González-Jaén, M.T.; Patiño, B. Wine Contamination with Ochratoxins: A Review. Beverages 2018, 4, 6. [Google Scholar] [CrossRef] [Green Version]
- European Commission. Regulation Nº 1881/2006 setting maximum levels for certain contaminants in foodstuffs. Off. J. Eur. Union 2006, 50, 8–12. [Google Scholar]
- Di Stefano, V.; Pitonzo, R.; Avellone, G.; Di Fiore, A.; Monte, L.; Ogorka, A.Z.T. Determination of Aflatoxins and Ochratoxins in Sicilian Sweet Wines by High-Performance Liquid Chromatography with Fluorometric Detection and Immunoaffinity Cleanup. Food Anal. Methods 2015, 8, 569–577. [Google Scholar] [CrossRef]
- Khoury, A.E.; Rizk, T.; Lteif, R.; Azouri, H.; Delia, M.-L.; Lebrihi, A. Fungal contamination and Aflatoxin B1 and Ochratoxin A in Lebanese wine–grapes and musts. Food Chem. Toxicol. 2008, 46, 2244–2250. [Google Scholar] [CrossRef]
- García-Cela, E.; Crespo-Sempere, A.; Gil-Serna, J.; Porqueres, A.; Marin, S. Fungal diversity, incidence and mycotoxin contamination in grapes from two agro-climatic Spanish regions with emphasis on Aspergillus species. J. Sci. Food Agric. 2015, 95, 1716–1729. [Google Scholar] [CrossRef]
- Paterson, R.R.M.; Venâncio, A.; Lima, N.; Guilloux-Bénatier, M.; Rousseaux, S. Predominant mycotoxins, mycotoxigenic fungi and climate change related to wine. Food Res. Int. 2018, 103, 478–491. [Google Scholar] [CrossRef] [Green Version]
- García-Cela, E.; Crespo-Sempere, A.; Ramos, A.J.; Sanchis, V.; Marín, S. Ecophysiological characterization of Aspergillus carbonarius, Aspergillus tubingensis and Aspergillus niger isolated from grapes in Spanish vineyards. Int. J. Food Microbiol. 2014, 173, 89–98. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hocking, A.D.; Leong, S.-L.L.; Kazi, B.A.; Emmett, R.W.; Scott, E.S. Fungi and mycotoxins in vineyards and grape products. Int. J. Food Microbiol. 2007, 119, 84–88. [Google Scholar] [CrossRef]
- Kabak, B.; Dobson, A.; Var, I. Strategies to Prevent Mycotoxin Contamination of Food and Animal Feed: A Review. Crit. Rev. Food Sci. Nutr. 2006, 46, 593–619. [Google Scholar] [CrossRef]
- Ponsone, M.L.; Chiotta, M.L.; Palazzini, J.M.; Combina, M.; Chulze, S. Control of Ochratoxin a Production in Grapes. Toxins 2012, 4, 364–372. [Google Scholar] [CrossRef] [Green Version]
- Gonçalves, A.; Palumbo, R.; Guimarães, A.; Gkrillas, A.; Dall’Asta, C.; Dorne, J.-L.; Battilani, P.; Venâncio, A. The Route of Mycotoxins in the Grape Food Chain. Am. J. Enol. Vitic. 2019, 71, 89–104. [Google Scholar] [CrossRef]
- Dukare, A.S.; Paul, S.; Nambi, V.E.; Gupta, R.K.; Singh, R.; Sharma, K.; Vishwakarma, R.K. Exploitation of microbial antagonists for the control of postharvest diseases of fruits: A review. Crit. Rev. Food Sci. Nutr. 2019, 59, 1498–1513. [Google Scholar] [CrossRef] [PubMed]
- Taroub, B.; Salma, L.; Manel, Z.; Ouzari, H.-I.; Hamdi, Z.; Moktar, H. Isolation of lactic acid bacteria from grape fruit: Antifungal activities, probiotic properties, and in vitro detoxification of ochratoxin A. Ann. Microbiol. 2019, 69, 17–27. [Google Scholar] [CrossRef]
- Pfliegler, W.P.; Pusztahelyi, T.; Pócsi, I. Mycotoxins—Prevention and decontamination by yeasts. J. Basic Microbiol. 2015, 55, 805–818. [Google Scholar] [CrossRef]
- Massart, S.; Martinez-Medina, M.; Jijakli, M.H. Biological control in the microbiome era: Challenges and opportunities. Biol. Control 2015, 89, 98–108. [Google Scholar] [CrossRef]
- Belda, I.; Zarraonaindia, I.; Perisin, M.; Palacios, A.; Acedo, A. From Vineyard Soil to Wine Fermentation: Microbiome Approximations to Explain the “terroir” Concept. Front. Microbiol. 2017, 8, 821. [Google Scholar] [CrossRef] [Green Version]
- Sarrocco, S.; Vannacci, G. Preharvest application of beneficial fungi as a strategy to prevent postharvest mycotoxin contamination: A review. Crop. Prot. 2018, 110, 160–170. [Google Scholar] [CrossRef]
- Barata, A.; Malfeito-Ferreira, M.; Loureiro, V. The microbial ecology of wine grape berries. Int. J. Food Microbiol. 2012, 153, 243–259. [Google Scholar] [CrossRef]
- Zhang, H.; Apaliya, M.T.; Mahunu, G.K.; Chen, L.; Li, W. Control of ochratoxin A-producing fungi in grape berry by microbial antagonists: A review. Trends Food Sci. Technol. 2016, 51, 88–97. [Google Scholar] [CrossRef]
- Liu, J.; Sui, Y.; Wisniewski, M.; Droby, S.; Liu, Y. Review: Utilization of antagonistic yeasts to manage postharvest fungal diseases of fruit. Int. J. Food Microbiol. 2013, 167, 153–160. [Google Scholar] [CrossRef] [PubMed]
- Bleve, G.; Grieco, F.; Cozzi, G.; Logrieco, A.F.; Visconti, A. Isolation of epiphytic yeasts with potential for biocontrol of Aspergillus carbonarius and A. niger on grape. Int. J. Food Microbiol. 2006, 108, 204–209. [Google Scholar] [CrossRef]
- Dimakopoulou, M.; Tjamos, S.E.; Antoniou, P.P.; Pietri, A.; Battilani, P.; Avramidis, N.; Markakis, E.; Tjamos, E.C. Phyllosphere grapevine yeast Aureobasidium pullulans reduces Aspergillus carbonarius (sour rot) incidence in wine-producing vineyards in Greece. Biol. Control 2008, 46, 158–165. [Google Scholar] [CrossRef]
- Ponsone, M.L.; Chiotta, M.L.; Combina, M.; Dalcero, A.; Chulze, S. Biocontrol as a strategy to reduce the impact of ochratoxin A and Aspergillus section Nigri in grapes. Int. J. Food Microbiol. 2011, 151, 70–77. [Google Scholar] [CrossRef]
- Grenier, B.; Oswald, I. Mycotoxin co-contamination of food and feed: Meta-analysis of publications describing toxicological interactions. World Mycotoxin J. 2011, 4, 285–313. [Google Scholar] [CrossRef]
- Stoev, S.D. Foodborne mycotoxicoses, risk assessment and underestimated hazard of masked mycotoxins and joint mycotoxin effects or interaction. Environ. Toxicol. Pharmacol. 2015, 39, 794–809. [Google Scholar] [CrossRef] [PubMed]
- Jard, G.; Liboz, T.; Mathieu, F.; Guyonvarch, A.; Lebrihi, A. Review of mycotoxin reduction in food and feed: From prevention in the field to detoxification by adsorption or transformation. Food Addit. Contam. Part. A 2011, 28, 1590–1609. [Google Scholar] [CrossRef] [PubMed]
- Agriopoulou, S.; Stamatelopoulou, E.; Varzakas, T. Advances in Occurrence, Importance, and Mycotoxin Control Strategies: Prevention and Detoxification in Foods. Foods 2020, 9, 137. [Google Scholar] [CrossRef]
- Castellari, M.; Versari, A.; Fabiani, A.; Parpinello, A.G.P.; Galassi, S. Removal of Ochratoxin A in Red Wines by Means of Adsorption Treatments with Commercial Fining Agents. J. Agric. Food Chem. 2001, 49, 3917–3921. [Google Scholar] [CrossRef] [PubMed]
- Hathout, A.S.; Aly, S. Biological detoxification of mycotoxins: A review. Ann. Microbiol. 2014, 64, 905–919. [Google Scholar] [CrossRef]
- Abrunhosa, L.; Paterson, R.R.M.; Venâncio, A. Biodegradation of Ochratoxin A for Food and Feed Decontamination. Toxins 2010, 2, 1078–1099. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pawlowska, A.M.; Zannini, E.; Coffey, A.; Arendt, E.K. “Green preservatives”: Combating fungi in the food and feed industry by applying antifungal lactic acid bacteria. In Advances Food and Nutrition Research; Jeyakuman, H., Ed.; Academic Press: Cambridge, MA, USA, 2012; Volume 66, pp. 217–238. [Google Scholar]
- Cecchini, F.; Morassut, M.; Moruno, E.G.; Di Stefano, R. Influence of yeast strain on ochratoxin A content during fermentation of white and red must. Food Microbiol. 2006, 23, 411–417. [Google Scholar] [CrossRef] [PubMed]
- Meca, G.; Blaiotta, G.; Ritieni, A. Reduction of ochratoxin A during the fermentation of Italian red wine Moscato. Food Control 2010, 21, 579–583. [Google Scholar] [CrossRef]
- Quintela, S.; Villarán, M.C.; de Armentia, I.L.; Elejalde, E. Ochratoxin A removal in wine: A review. Food Control 2013, 30, 439–445. [Google Scholar] [CrossRef]
- European Commission. A Farm to Fork Strategy for a Fair, Healthy and Environmentally-Friendly Food System. Available online: https://ec.europa.eu/food/system/files/2020-05/f2f_action-plan_2020_strategy-info_en.pdf (accessed on 5 July 2021).
- Zhang, H.; Godana, E.A.; Sui, Y.; Yang, Q.; Zhang, X.; Zhao, L. Biological control as an alternative to synthetic fungicides for the management of grey and blue mould diseases of table grapes: A review. Crit. Rev. Microbiol. 2020, 46, 450–462. [Google Scholar] [CrossRef]
- De Simone, N.; Pace, B.; Grieco, F.; Chimienti, M.; Tyibilika, V.; Santoro, V.; Capozzi, V.; Colelli, G.; Spano, G.; Russo, P. Botrytis cinerea and Table Grapes: A Review of the Main Physical, Chemical, and Bio-Based Control Treatments in Post-Harvest. Foods 2020, 9, 1138. [Google Scholar] [CrossRef]
- Medina-Vaya, A.; Rodriguez, A.; Magan, N. Climate change and mycotoxigenic fungi: Impacts on mycotoxin production. Curr. Opin. Food Sci. 2015, 5, 99–104. [Google Scholar] [CrossRef]
- Varga, J.; Kozakiewicz, Z. Ochratoxin A in grapes and grape-derived products. Trends Food Sci. Technol. 2006, 17, 72–81. [Google Scholar] [CrossRef]
- Cervini, C.; Gallo, A.; Piemontese, L.; Magistà, D.; Logrieco, A.F.; Ferrara, M.; Solfrizzo, M.; Perrone, G. Effects of temperature and water activity change on ecophysiology of ochratoxigenic Aspergillus carbonarius in field-simulating conditions. Int. J. Food Microbiol. 2020, 315, 108420. [Google Scholar] [CrossRef] [PubMed]
- Gostinčar, C.; Grube, M.; Gunde-Cimerman, N. Evolution of Fungal Pathogens in Domestic Environments? Fungal Biol. 2011, 115, 1008–1018. [Google Scholar] [CrossRef] [PubMed]
- Zajc, J.; Gostinčar, C.; Černoša, A.; Gunde-Cimerman, N. Stress-Tolerant Yeasts: Opportunistic Pathogenicity Versus Biocontrol Potential. Genes 2019, 10, 42. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zott, K.; Miot-Sertier, C.; Claisse, O.; Lonvaud, A.; Masneuf-Pomarede, I. Dynamics and diversity of non-Saccharomyces yeasts during the early stages in winemaking. Int. J. Food Microbiol. 2008, 125, 197–203. [Google Scholar] [CrossRef]
- Belda, I.; Ruiz, J.; Alastruey-Izquierdo, A.; Navascués, E.; Marquina, D.; Santos, A. Unraveling the Enzymatic Basis of Wine “Flavorome”: A Phylo-Functional Study of Wine Related Yeast Species. Front. Microbiol. 2016, 7, 12. [Google Scholar] [CrossRef] [Green Version]
- Jolly, N.P.; Varela, C.; Pretorius, I.S. Not your ordinary yeast: Non-Saccharomyces yeasts in wine production uncovered. FEMS Yeast Res. 2014, 14, 215–237. [Google Scholar] [CrossRef] [Green Version]
- Qin, X.; Xiao, H.; Xue, C.; Yu, Z.; Yang, R.; Cai, Z.; Si, L. Biocontrol of gray mold in grapes with the yeast Hanseniaspora uvarum alone and in combination with salicylic acid or sodium bicarbonate. Postharvest Biol. Technol. 2015, 100, 160–167. [Google Scholar] [CrossRef]
- Liu, H.M.; Guo, J.H.; Cheng, Y.J.; Luo, L.; Liu, P.; Wang, B.Q.; Deng, B.X.; Long, C.A. Control of gray mold of grape by Hanseniaspora uvarum and its effects on postharvest quality parameters. Ann. Microbiol. 2010, 60, 31–35. [Google Scholar] [CrossRef]
- Cordero-Bueso, G.; Mangieri, N.; Maghradze, D.; Foschino, R.; Valdetara, F.; Cantoral, J.M.; Vigentini, I. Wild Grape-Associated Yeasts as Promising Biocontrol Agents against Vitis vinifera Fungal Pathogens. Front. Microbiol. 2017, 8, 2025. [Google Scholar] [CrossRef] [Green Version]
- Apaliya, M.T.; Zhang, H.; Zheng, X.; Yang, Q.; Mahunu, G.K.; Kwaw, E. Exogenous trehalose enhanced the biocontrol efficacy of Hanseniaspora uvarum against grape berry rots caused by Aspergillus tubingensis and Penicillium commune. J. Sci. Food Agric. 2018, 98, 4665–4672. [Google Scholar] [CrossRef]
- Prendes, L.P.; Merín, M.G.; Zachetti, V.G.; Pereyra, A.; Ramirez, M.L.; de Ambrosini, V.I.M. Impact of antagonistic yeasts from wine grapes on growth and mycotoxin production by Alternaria alternata. J. Appl. Microbiol. 2021. [Google Scholar] [CrossRef] [PubMed]
- Medina, A.; Mohale, S.; Samsudin, N.I.P.; Rodriguez-Sixtos, A.; Rodriguez, A.; Magan, N. Biocontrol of mycotoxins: Dynamics and mechanisms of action. Curr. Opin. Food Sci. 2017, 17, 41–48. [Google Scholar] [CrossRef]
- Angioni, A.; Caboni, P.; Garau, A.; Farris, A.; Orro, D.; Budroni, M.; Cabras, P. In Vitro Interaction between Ochratoxin A and Different Strains of Saccharomyces cerevisiae and Kloeckera apiculata. J. Agric. Food Chem. 2007, 55, 2043–2048. [Google Scholar] [CrossRef] [PubMed]
- Piotrowska, M.; Żakowska, Z. The biodegradation of ochratoxin A in food products by lactic acid bacteria and baker’s yeast. Prog. Biotechnol. 2000, 17, 307–310. [Google Scholar] [CrossRef]
- Chlebicz, A.; Śliżewska, K. In Vitro Detoxification of Aflatoxin B1, Deoxynivalenol, Fumonisins, T-2 Toxin and Zearalenone by Probiotic Bacteria from Genus Lactobacillus and Saccharomyces cerevisiae Yeast. Probiotics Antimicrob. Proteins 2020, 12, 289–301. [Google Scholar] [CrossRef] [Green Version]
- Bejaoui, H.; Mathieu, F.; Taillandier, P.; Lebrihi, A. Ochratoxin A removal in synthetic and natural grape juices by selected oenological Saccharomyces strains. J. Appl. Microbiol. 2004, 97, 1038–1044. [Google Scholar] [CrossRef] [Green Version]
- Avantaggiato, G.; Greco, D.; Damascelli, A.; Solfrizzo, M.; Visconti, A. Assessment of Multi-mycotoxin Adsorption Efficacy of Grape Pomace. J. Agric. Food Chem. 2014, 62, 497–507. [Google Scholar] [CrossRef]
- Gil-Serna, J.; Patiño, B.; Cortes, L.; González-Jaén, M.T.; Vázquez, C. Mechanisms involved in reduction of ochratoxin A produced by Aspergillus westerdijkiae using Debaryomyces hansenii CYC 1244. Int. J. Food Microbiol. 2011, 151, 113–118. [Google Scholar] [CrossRef]
- Piotrowska, M.; Masek, A. Saccharomyces Cerevisiae Cell Wall Components as Tools for Ochratoxin a Decontamination. Toxins 2015, 7, 1151–1162. [Google Scholar] [CrossRef] [Green Version]
- Faucet-Marquis, V.; Joannis-Cassan, C.; Hadjeba-Medjdoub, K.; Ballet, N.; Pfohl-Leszkowicz, A. Development of an in vitro method for the prediction of mycotoxin binding on yeast-based products: Case of aflatoxin B1, zearalenone and ochratoxin A. Appl. Microbiol. Biotechnol. 2014, 98, 7583–7596. [Google Scholar] [CrossRef] [Green Version]
- Shetty, P.H.; Hald, B.; Jespersen, L. Surface binding of aflatoxin B1 by Saccharomyces cerevisiae strains with potential decontaminating abilities in indigenous fermented foods. Int. J. Food Microbiol. 2007, 113, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Jouany, J.; Yiannikouris, A.; Bertin, G. The chemical bonds between mycotoxins and cell wall components of Saccharomyces cerevisiae have been identified. Arch. Zootech. 2005, 8, 26–50. [Google Scholar]
- Moore, J.P.; Zhang, S.-L.; Nieuwoudt, H.; Divol, B.; Trygg, J.; Bauer, F.F. A Multivariate Approach Using Attenuated Total Reflectance Mid-infrared Spectroscopy To Measure the Surface Mannoproteins and β-Glucans of Yeast Cell Walls during Wine Fermentations. J. Agric. Food Chem. 2015, 63, 10054–10063. [Google Scholar] [CrossRef] [PubMed]
- Yiannikouris, A.; André, G.; Poughon, L.; François, J.M.; Dussap, C.-G.; Jeminet, G.; Bertin, G.; Jouany, J.-P. Chemical and Conformational Study of the Interactions Involved in Mycotoxin Complexation with β-d-Glucans. Biomacromolecules 2006, 7, 1147–1155. [Google Scholar] [CrossRef] [PubMed]
- Schatzmayr, G.; Heidler, D.; Fuchs, E.; Nitsch, S.; Mohnl, M.; Täubel, M.; Loibner, A.P.; Braun, R.; Binder, E.M. Investigation of different yeast strains for the detoxification of ochratoxin A. Mycotoxin Res. 2003, 19, 124–128. [Google Scholar] [CrossRef]
- De Felice, D.V.; Solfrizzo, M.; De Curtis, F.; Lima, G.; Visconti, A.; Castoria, R. Strains of Aureobasidium pullulans Can Lower Ochratoxin a Contamination in Wine Grapes. Phytopathology 2008, 98, 1261–1270. [Google Scholar] [CrossRef] [Green Version]
- Dobritzsch, D.; Wang, H.; Schneider, G.; Yu, S. Structural and functional characterization of ochratoxinase, a novel mycotoxin-degrading enzyme. Biochem. J. 2014, 462, 441–452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abrunhosa, L.; Santos, L.; Venâncio, A. Degradation of Ochratoxin A by Proteases and by a Crude Enzyme of Aspergillus niger. Food Biotechnol. 2006, 20, 231–242. [Google Scholar] [CrossRef] [Green Version]
- Ben Taheur, F.; Mansour, C.; Ben Jeddou, K.; Machreki, Y.; Kouidhi, B.; Abdulhakim, J.A.; Chaieb, K. Aflatoxin B1 degradation by microorganisms isolated from Kombucha culture. Toxicon 2020, 179, 76–83. [Google Scholar] [CrossRef]
- Li, J.; Huang, J.; Jin, Y.; Wu, C.; Shen, D.; Zhang, S.; Zhou, R. Mechanism and kinetics of degrading aflatoxin B1 by salt tolerant Candida versatilis CGMCC 3790. J. Hazard. Mater. 2018, 359, 382–387. [Google Scholar] [CrossRef]
- Vanhoutte, I.; Audenaert, K.; De Gelder, L. Biodegradation of Mycotoxins: Tales from Known and Unexplored Worlds. Front. Microbiol. 2016, 7, 561. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- EFSA Panel. The list of QPS status recommended biological agents for safety risk assessments carried out by EFSA. EFSA J. 2021, 19, 6377. [Google Scholar]
- White, T.J.; Burns, T.; Lee, S.; Taylor, J.W. Amplification and direct sequencing of fungal ribosomal RNA genes for phyloge-netics. In PCR Protocols: A Guide to Method and Application; Innis, M.A., Gelfald, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Magoč, T.; Salzberg, S. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011, 27, 2957–2963. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Fu, L.; Niu, B.; Wu, S.; Wooley, J. Ultrafast clustering algorithms for metagenomic sequence analysis. Brief. Bioinform. 2012, 13, 656–668. [Google Scholar] [CrossRef] [PubMed]
- Kõljalg, U.; Nilsson, H.; Abarenkov, K.; Tedersoo, L.; Taylor, A.F.S.; Bahram, M.; Bates, S.T.; Bruns, T.D.; Bengtsson-Palme, J.; Callaghan, T.M.; et al. Towards a unified paradigm for sequence-based identification of fungi. Mol. Ecol. 2013, 22, 5271–5277. [Google Scholar] [CrossRef] [Green Version]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [Green Version]
- Kurtzman, C.P.; Robnett, C.J. Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Anton. Leeuw. Int. J. G. 1998, 73, 331–371. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).