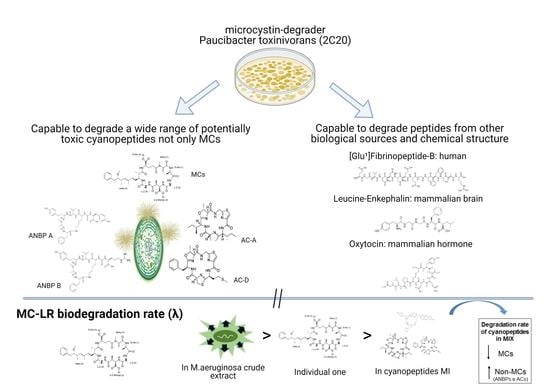
Degradation of Multiple Peptides by Microcystin-Degrader Paucibacter toxinivorans (2C20)
Abstract
1. Introduction
2. Results
2.1. Biodegradation of Individual Cyanopeptides
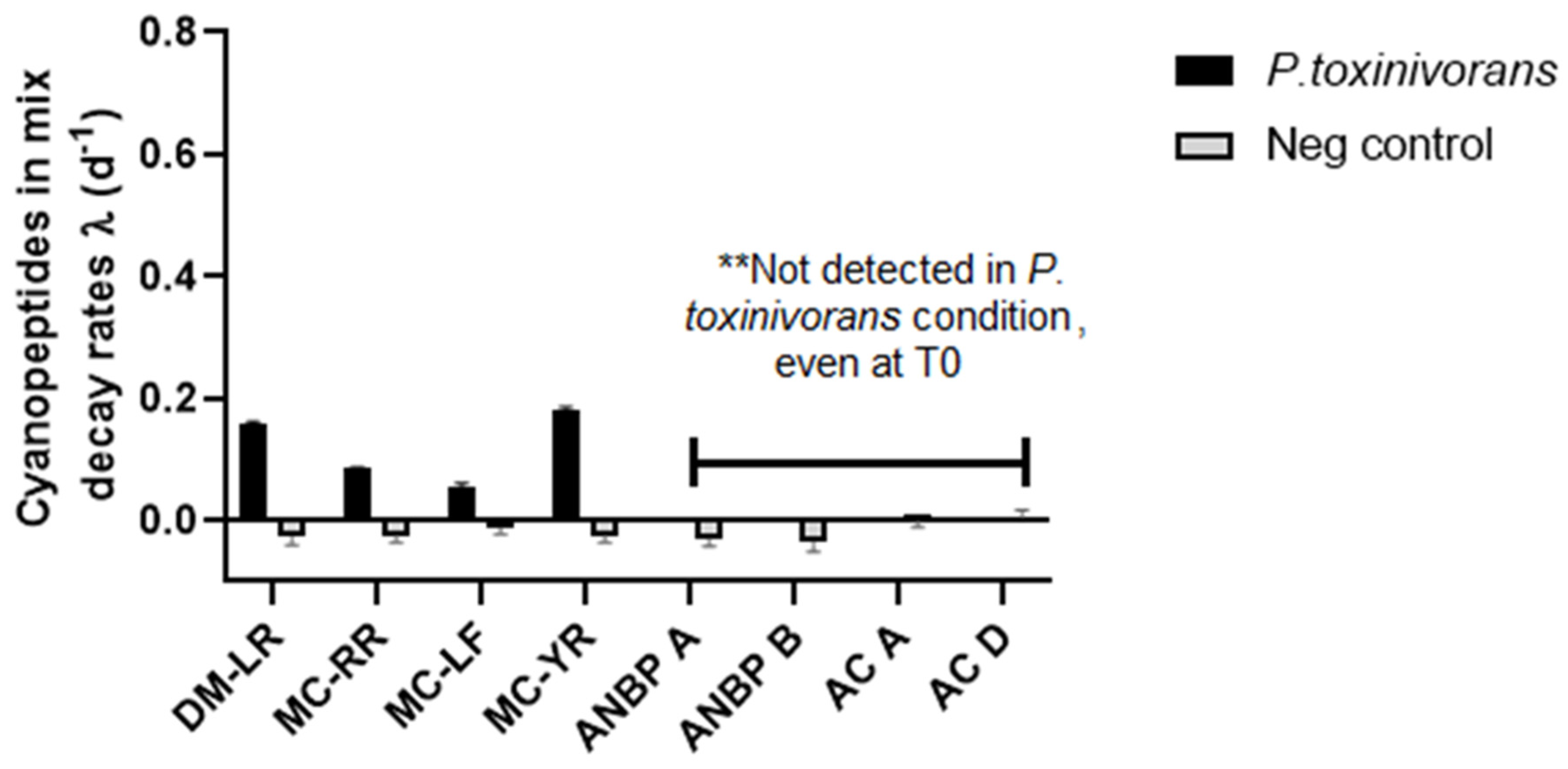
2.2. MC-LR Biodegradation in Different Complex Mixtures
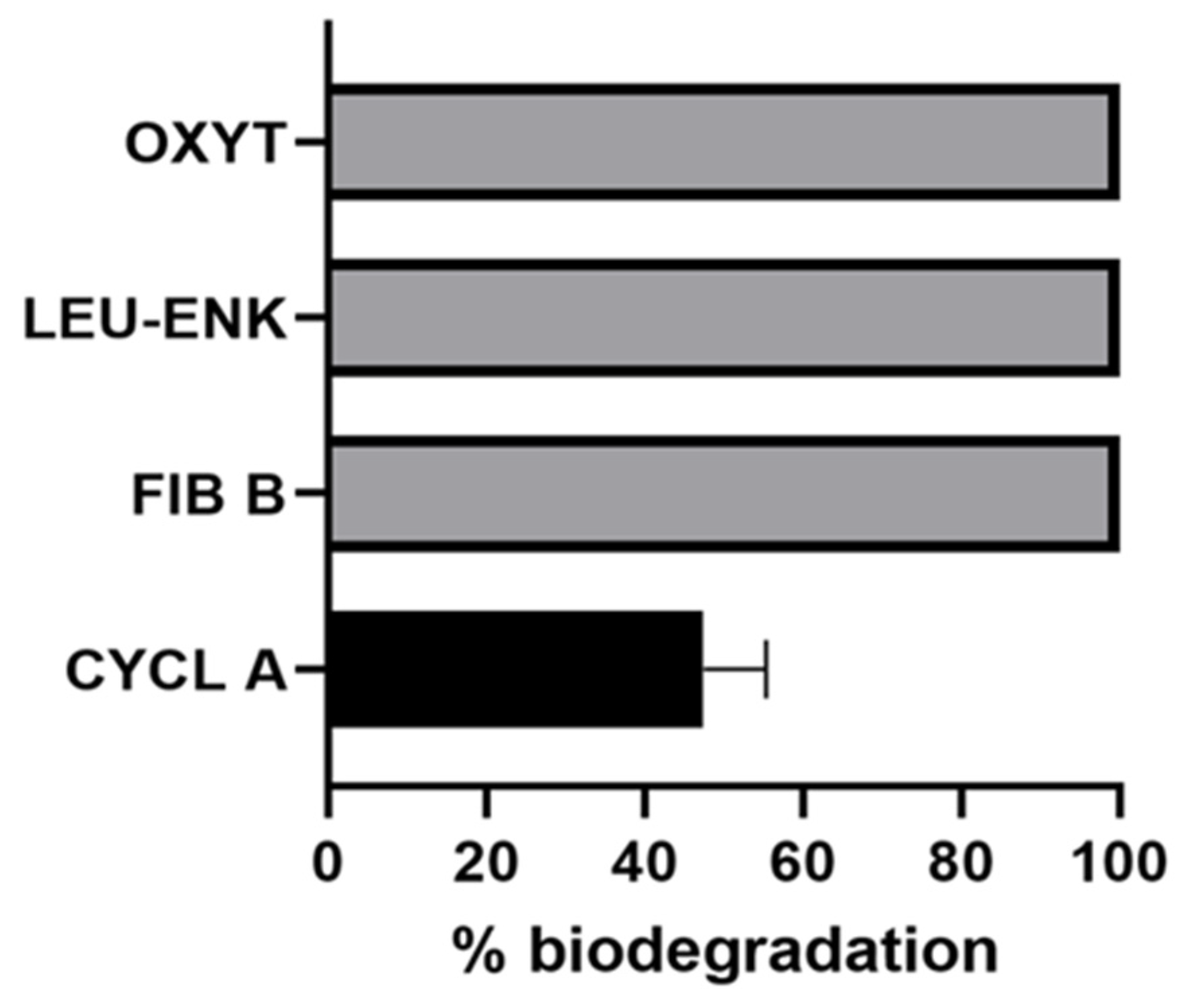
2.3. Biodegradation of Non-Cyanobacterial Peptides
2.4. Identification of MC Biodegradation Products by UPLC–QTOF–MS/MS
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Chemicals and Reagents
5.2. Bacterial Cultivation
5.3. High-Throughput 96-Well Microplate Biodegradation Experiments
5.3.1. Biodegradation of Individual Cyanopeptides
5.3.2. Biodegradation Rate of MC-LR in Complex Mixtures
5.3.3. Biodegradation of Non-Cyanobacterial Peptides
5.4. Analysis of Peptides by Ultrahigh Performance Liquid Chromatography Coupled to Photodiode Array Detection and Tandem Mass Spectrometry (UPLC–PDA–MS/MS)
5.5. Analysis of Peptide Biodegradation Products by UPLC–PDA Coupled to Quadrupole Time of Flight Mass Spectrometry (QTOF-MSE, QTOF-MS/MS)
5.6. Biodegradation Rate, Half-Life, and Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Paerl, H.W.; Otten, T.G. Harmful Cyanobacterial Blooms: Causes, Consequences, and Controls. Microb. Ecol. 2013, 65, 995–1010. [Google Scholar] [CrossRef]
- Szlag, D.C.; Sinclair, J.L.; Southwell, B.J.; Westrick, J.A. Cyanobacteria and Cyanotoxins Occurrence and Removal from Five High-Risk Conventional Treatment Drinking Water Plants. Toxins 2015, 7, 2198–2220. [Google Scholar] [CrossRef] [PubMed]
- Meriluoto, J.; Spoof, L.; Codd, G.A. Handbook of Cyanobacterial Monitoring and Cyanotoxin Analysis, 1st ed; John Wiley & Sons: Hoboken, NJ, USA, 2017; 576p, ISBN 978-1-119-06868-6. [Google Scholar]
- Bouaïcha, N.; Miles, C.O.; Beach, D.G.; Labidi, Z.; Djabri, A.; Benayache, N.Y.; Nguyen-Quang, T. Structural Diversity, Characterization and Toxicology of Microcystins. Toxins 2019, 11, 714. [Google Scholar] [CrossRef]
- Chernoff, N.; Hill, D.; Lang, J.; Schmid, J.; Le, T.; Farthing, A.; Huang, H. The Comparative Toxicity of 10 Microcystin Congeners Administered Orally to Mice: Clinical Effects and Organ Toxicity. Toxins 2020, 12, 403. [Google Scholar] [CrossRef] [PubMed]
- Janssen, E.M.-L. Cyanobacterial peptides beyond microcystins—A review on co-occurrence, toxicity, and challenges for risk assessment. Water Res. 2019, 151, 488–499. [Google Scholar] [CrossRef] [PubMed]
- Smutná, M.; Babica, P.; Jarque, S.; Hilscherová, K.; Maršálek, B.; Haeba, M.; Bláha, L. Acute, chronic and reproductive toxicity of complex cyanobacterial blooms in Daphnia magna and the role of microcystins. Toxicon 2014, 79, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Le Manach, S.; Khenfech, N.; Huet, H.; Qiao, Q.; Duval, C.; Marie, A.; Bolbach, G.; Clodic, G.; Djediat, C.; Bernard, C.; et al. Gender-Specific Toxicological Effects of Chronic Exposure to Pure Microcystin-LR or Complex Microcystis aeruginosa Extracts on Adult Medaka Fish. Environ. Sci. Technol. 2016, 50, 8324–8334. [Google Scholar] [CrossRef]
- Beversdorf, L.J.; Weirich, C.A.; Bartlett, S.L.; Miller, T.R. Variable Cyanobacterial Toxin and Metabolite Profiles across Six Eutrophic Lakes of Differing Physiochemical Characteristics. Toxins 2017, 9, 62. [Google Scholar] [CrossRef] [PubMed]
- Gkelis, S.; Lanaras, T.; Sivonen, K. The presence of microcystins and other cyanobacterial bioactive peptides in aquatic fauna collected from Greek freshwaters. Aquat. Toxicol. 2006, 78, 32–41. [Google Scholar] [CrossRef]
- Dao, T.-S.; Ortiz-Rodríguez, R.; Do-Hong, L.-C.; Wiegand, C. Non-microcystin and non-cylindrospermopsin producing cyanobacteria affect the biochemical responses and behavior of Daphnia magna. Int. Rev. Hydrobiol. 2013, 98, 235–244. [Google Scholar] [CrossRef]
- Kohler, E.; Grundler, V.; Häussinger, D.; Kurmayer, R.; Gademann, K.; Pernthaler, J.; Blom, J.F. The toxicity and enzyme activity of a chlorine and sulfate containing aeruginosin isolated from a non-microcystin-producing Planktothrix strain. Harmful Algae 2014, 39, 154–160. [Google Scholar] [CrossRef]
- Schreuder, H.; Liesum, A.; Lönze, P.; Stump, H.; Hoffmann, H.; Schiell, M.; Kurz, M.; Toti, L.; Bauer, A.; Kallus, C.; et al. Isolation, Co-Crystallization and Structure-Based Characterization of Anabaenopeptins as Highly Potent Inhibitors of Activated Thrombin Activatable Fibrinolysis Inhibitor (TAFIa). Sci. Rep. 2016, 6, 32958. [Google Scholar] [CrossRef]
- Kurmayer, R.; Deng, L.; Entfellner, E. Role of toxic and bioactive secondary metabolites in colonization and bloom formation by filamentous cyanobacteria Planktothrix. Harmful Algae 2016, 54, 69–86. [Google Scholar] [CrossRef]
- Otten, T.G.; Paerl, H.W.; Dreher, T.W.; Kimmerer, W.J.; Parker, A.E. The molecular ecology ofMicrocystissp. blooms in the San Francisco Estuary. Environ. Microbiol. 2017, 19, 3619–3637. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Hegde, K.; Brar, S.K.; Cledon, M.; Kermanshahi-Pour, A. Potential of biological approaches for cyanotoxin removal from drinking water: A review. Ecotoxicol. Environ. Saf. 2019, 172, 488–503. [Google Scholar] [CrossRef] [PubMed]
- Valeria, A.M.; Ricardo, E.J.; Stephan, P.; Alberto, W.D. Degradation of Microcystin-RR by Sphingomonas sp. CBA4 isolated from San Roque reservoir (Córdoba–Argentina). Biodegradation 2006, 17, 447–455. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Yang, X.; Yang, L.; Xiao, B.; Wu, X.; Wang, J.; Wan, H. An effective pathway for the removal of microcystin LR via anoxic biodegradation in lake sediments. Water Res. 2010, 44, 1884–1892. [Google Scholar] [CrossRef] [PubMed]
- Edwards, C.; Graham, D.; Fowler, N.; Lawton, L.A. Biodegradation of microcystins and nodularin in freshwaters. Chemosphere 2008, 73, 1315–1321. [Google Scholar] [CrossRef]
- Harada, K.; Fujii, K.; Shimada, T.; Suzuki, M.; Sano, H.; Adachi, K.; Carmichael, W.W. 2 cyclic-peptides, anabaenopeptins, a 3rd group of bioactive compounds from the cyanobacterium Anabaena-flos-aquae NRC-525-17. Tetrahedron Lett. 1995, 36, 1511–1514. [Google Scholar] [CrossRef]
- Dexter, J.; McCormick, A.J.; Fu, P.; Dziga, D. Microcystinase—A review of the natural occurrence, heterologous expression, and biotechnological application of MlrA. Water Res. 2021, 189, 116646. [Google Scholar] [CrossRef]
- Jones, G.J.; Bourne, D.G.; Blakeley, R.L.; Doelle, H. Degradation of the cyanobacterial hepatotoxin microcystin by aquatic bacteria. Nat. Toxins 1994, 2, 228–235. [Google Scholar] [CrossRef] [PubMed]
- Bourne, D.G.; Jones, G.J.; Blakeley, R.L.; Jones, G.J.; Negri, A.P.; Riddles, P. Enzymatic pathway for the bacterial deg-radation of the cyanobacterial cyclic peptide toxin microcystin LR. Appl. Environ. Microbiol. 1996, 62, 4086–4094. [Google Scholar] [CrossRef] [PubMed]
- Bourne, D.G.; Riddles, P.; Jones, G.J.; Smith, W.; Blakeley, R.L. Characterization of a gene cluster involved in bacterial degradation of the cyanobacterial toxin microcystin LR. Environ. Toxicol. 2001, 16, 523–534. [Google Scholar] [CrossRef] [PubMed]
- Dziga, D.; Wasylewski, M.; Wladyka, B.; Nybom, S.; Meriluoto, J. Microbial Degradation of Microcystins. Chem. Res. Toxicol. 2013, 26, 841–852. [Google Scholar] [CrossRef] [PubMed]
- Dziga, D.; Zielinska, G.; Wladyka, B.; Bochenska, O.; Maksylewicz, A.; Strzalka, W.; Meriluoto, J. Characterization of Enzymatic Activity of MlrB and MlrC Proteins Involved in Bacterial Degradation of Cyanotoxins Microcystins. Toxins 2016, 8, 76. [Google Scholar] [CrossRef]
- Dziga, D.; Nybom, S.; Gagala, I.; Wasylewski, M. Biological Treatment for the Destruction of Cyanotoxins. In Water Treatment for Purification from Cyanobacteria and Cyanotoxins; John Wiley & Sons: Hoboken, NJ, USA, 2020; Chapter 5; pp. 117–153. [Google Scholar] [CrossRef]
- Lezcano, M.Á.; Morón-López, J.; Agha, R.; López-Heras, I.; Nozal, L.; Quesada, A.; El-Shehawy, R. Presence or Absence of mlr Genes and Nutrient Concentrations Co-Determine the Microcystin Biodegradation Efficiency of a Natural Bacterial Community. Toxins 2016, 8, 318. [Google Scholar] [CrossRef] [PubMed]
- Morón-López, J.; Nieto-Reyes, L.; El-Shehawy, R. Assessment of the influence of key abiotic factors on the alternative microcystin degradation pathway(s) (mlr−): A detailed comparison with the mlr route (mlr+). Sci. Total. Environ. 2017, 599–600, 1945–1953. [Google Scholar] [CrossRef]
- Mou, X.; Lu, X.; Jacob, J.; Sun, S.; Heath, R. Metagenomic Identification of Bacterioplankton Taxa and Pathways Involved in Microcystin Degradation in Lake Erie. PLoS ONE 2013, 8, e61890. [Google Scholar] [CrossRef]
- Zhu, X.; Shen, Y.; Chen, X.; Hu, Y.O.; Xiang, H.; Tao, J.; Ling, Y. Biodegradation mechanism of microcystin-LR by a novel isolate of Rhizobium sp. TH and the evolutionary origin of the mlrA gene. Int. Biodeterior. Biodegrad. 2016, 115, 17–25. [Google Scholar] [CrossRef]
- Qin, L.; Zhang, X.; Chen, X.; Wang, K.; Shen, Y.; Li, D. Isolation of a Novel Microcystin-Degrading Bacterium and the Evolutionary Origin of mlr Gene Cluster. Toxins 2019, 11, 269. [Google Scholar] [CrossRef]
- Yang, F.; Huang, F.; Feng, H.; Wei, J.; Massey, I.Y.; Liang, G.; Zhang, F.; Yin, L.; Kacew, S.; Zhang, X.; et al. A complete route for biodegradation of potentially carcinogenic cyanotoxin microcystin-LR in a novel indigenous bacterium. Water Res. 2020, 174, 115638. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Yang, F.; Chen, L.; Feng, H.; Yin, S.; Chen, M. Insights into ecological roles and potential evolution of Mlr-dependent microcystin-degrading bacteria. Sci. Total. Environ. 2020, 710, 136401. [Google Scholar] [CrossRef] [PubMed]
- Kreitz, J. Evidence for Functional Drift of Bacterial Isolates in Response to Cyanobacterial Microcystin-lr and Multiple Peptide Degradation in Paucibacter Toxinivorans. MPhil Thesis, Robert Gordon University, Aberdeen, UK, January 2016. [Google Scholar]
- Park, H.-D.; Sasaki, Y.; Maruyama, T.; Yanagisawa, E.; Hiraishi, A.; Kato, K. Degradation of the cyanobacterial hepatotoxin microcystin by a new bacterium isolated from a hypertrophic lake. Environ. Toxicol. 2001, 16, 337–343. [Google Scholar] [CrossRef] [PubMed]
- Rapala, J.; Berg, K.A.; Lyra, C.; Niemi, R.M.; Manz, W.; Suomalainen, S.; Paulin, L.; Lahti, K. Paucibacter toxinivorans gen. nov., sp. nov., a bacterium that degrades cyclic cyanobacterial hepatotoxins microcystins and nodularin. Int. J. Syst. Evol. Microbiol. 2005, 55, 1563–1568. [Google Scholar] [CrossRef]
- Kato, H.; Imanishi, S.Y.; Tsuji, K.; Harada, K.-I. Microbial degradation of cyanobacterial cyclic peptides. Water Res. 2007, 41, 1754–1762. [Google Scholar] [CrossRef]
- Briand, E.; Humbert, J.; Tambosco, K.; Bormans, M.; Gerwick, W.H. Role of bacteria in the production and degradation of Microcystis cyanopeptides. Microbiologyopen 2016, 5, 469–478. [Google Scholar] [CrossRef]
- Toruńska-Sitarz, A.; Kotlarska, E.; Mazur-Marzec, H. Biodegradation of nodularin and other nonribosomal peptides by the Baltic bacteria. Int. Biodeterior. Biodegrad. 2018, 134, 48–57. [Google Scholar] [CrossRef]
- Lawton, L.A.; Welgamage, A.; Manage, P.M.; Edwards, C. Novel bacterial strains for the removal of microcystins from drinking water. Water Sci. Technol. 2011, 63, 1137–1142. [Google Scholar] [CrossRef]
- Seo, J.-S.; Keum, Y.-S.; Li, Q.X. Bacterial Degradation of Aromatic Compounds. Int. J. Environ. Res. Public Health 2009, 6, 278–309. [Google Scholar] [CrossRef]
- Wackett, L.P. The Metabolic Pathways of Biodegradation. Prokaryotes 2013, 383–393. [Google Scholar] [CrossRef]
- Yang, F.; Zhou, Y.; Sun, R.; Wei, H.; Li, Y.; Yin, L.; Pu, Y. Biodegradation of microcystin-LR and-RR by a novel microcystin-degrading bacterium isolated from Lake Taihu. Biodegradation 2014, 25, 447–457. [Google Scholar] [CrossRef]
- Rapala, J.; Lahti, K.; Sivonen, K.; Niemelä, S.I. Biodegradability and adsorption on lake sediments of cyanobacterial hepatotoxins and anatoxin-a. Lett. Appl. Microbiol. 1994, 19, 423–428. [Google Scholar] [CrossRef]
- Christoffersen, K.; Lyck, S.; Winding, A. Microbial activity and bacterial community structure during degradation of microcystins. Aquat. Microb. Ecol. 2002, 27, 125–136. [Google Scholar] [CrossRef]
- Le Manach, S.; Duval, C.; Marie, A.; Djediat, C.; Catherine, A.; Edery, M.; Bernard, C.; Marie, B. Global Metabolomic Characterizations of Microcystis spp. Highlights Clonal Diversity in Natural Bloom-Forming Populations and Expands Metabolite Structural Diversity. Front. Microbiol. 2019, 10, 791. [Google Scholar] [CrossRef] [PubMed]
- Sarma, H.; Prasad, M. Phytomanagement of Polycyclic Aromatic Hydrocarbons and Heavy Metals-Contaminated Sites in Assam, North Eastern State of India, for Boosting Bioeconomy. Bioremediat. Bioecon. 2016, 609–626. [Google Scholar] [CrossRef]
- Portmann, C.; Blom, J.F.; Gademann, K.; Jüttner, F. Aerucyclamides A and B: Isolation and Synthesis of Toxic Ribosomal Heterocyclic Peptides from the CyanobacteriumMicrocystis aeruginosaPCC 7806. J. Nat. Prod. 2008, 71, 1193–1196. [Google Scholar] [CrossRef]
- Welker, M.; Maršálek, B.; Šejnohová, L.; Von Döhren, H. Detection and identification of oligopeptides in Microcystis (cyanobacteria) colonies: Toward an understanding of metabolic diversity. Peptides 2006, 27, 2090–2103. [Google Scholar] [CrossRef]
- Fotiou, T.; Triantis, T.M.; Hiskia, A.E.; Dziga, D.; Merel, S.; Edwards, C.; Antoniou, M.G. Transformation Products (TPs) of Cyanobacterial Metabolites During Treatment. In Water Treatment for Purification from Cyanobacteria and Cyanotoxins; John Wiley & Sons: Hoboken, NJ, USA, 2020; pp. 231–305. [Google Scholar] [CrossRef]
- Xu, H.; Wang, H.; Xu, Q.; Lv, L.; Yin, C.; Liu, X.; Du, H.; Yan, H. Pathway for Biodegrading Microcystin-YR by Sphingopyxis sp. USTB-05. PLoS ONE 2015, 10, e0124425. [Google Scholar] [CrossRef]
- Imanishi, S.; Kato, H.; Mizuno, M.; Tsuji, K.; Harada, K.-I. Bacterial Degradation of Microcystins and Nodularin. Chem. Res. Toxicol. 2005, 18, 591–598. [Google Scholar] [CrossRef]
- du Vigneaud, V.; Ressler, C.; Trippett, S. The sequence of amino acids in oxytocin, with a proposal for the structure of oxytocin. J. Biol. Chem. 1953, 205, 949–957. [Google Scholar] [CrossRef]
- Wiśniewski, K.; Finnman, J.; Flipo, M.; Galyean, R.; Schteingart, C.D. On the mechanism of degradation of oxytocin and its analogues in aqueous solution. Biopolymers 2013, 100, 408–421. [Google Scholar] [CrossRef]
- Reasoner, D.J.; E Geldreich, E. A new medium for the enumeration and subculture of bacteria from potable water. Appl. Environ. Microbiol. 1985, 49, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Turner, A.D.; Waack, J.; Lewis, A.; Edwards, C.; Lawton, L. Development and single-laboratory validation of a UHPLC-MS/MS method for quantitation of microcystins and nodularin in natural water, cyanobacteria, shellfish and algal supplement tablet powders. J. Chromatogr. B 2018, 1074–1075, 111–123. [Google Scholar] [CrossRef]
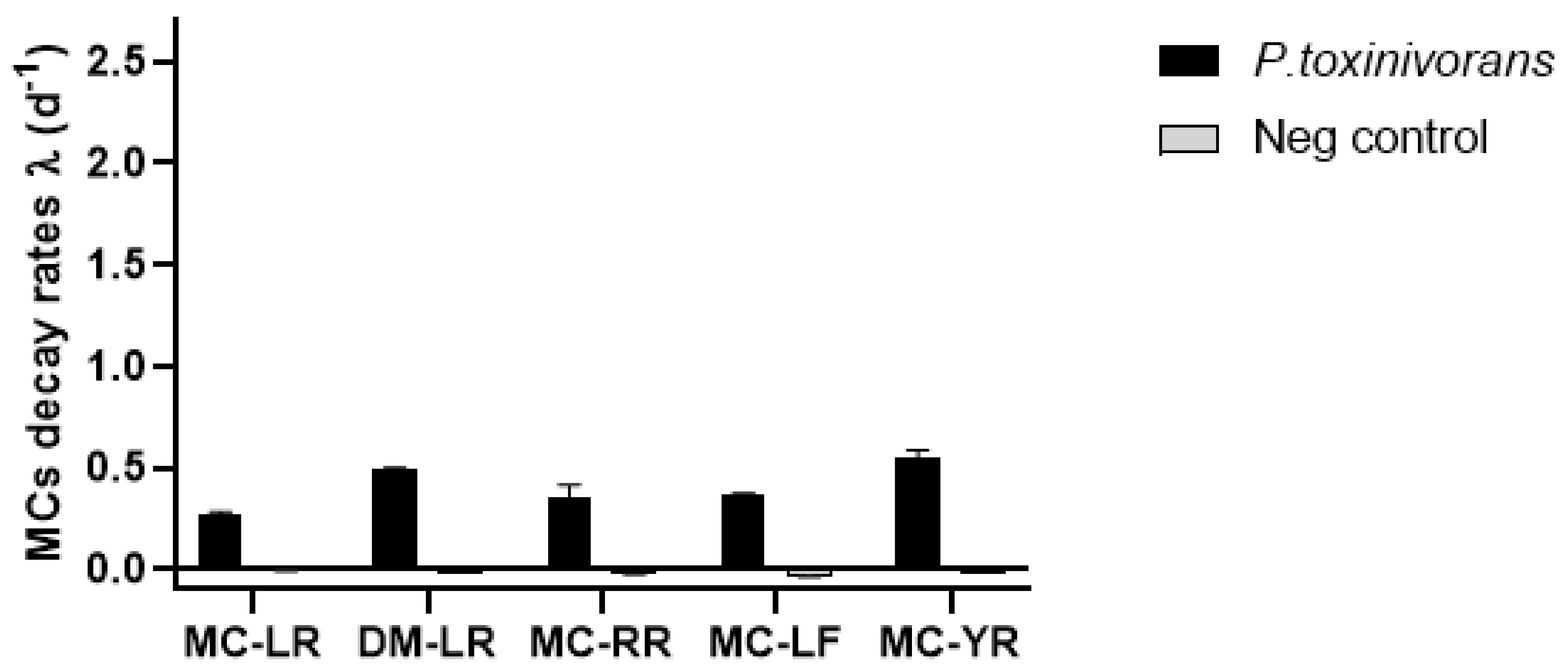
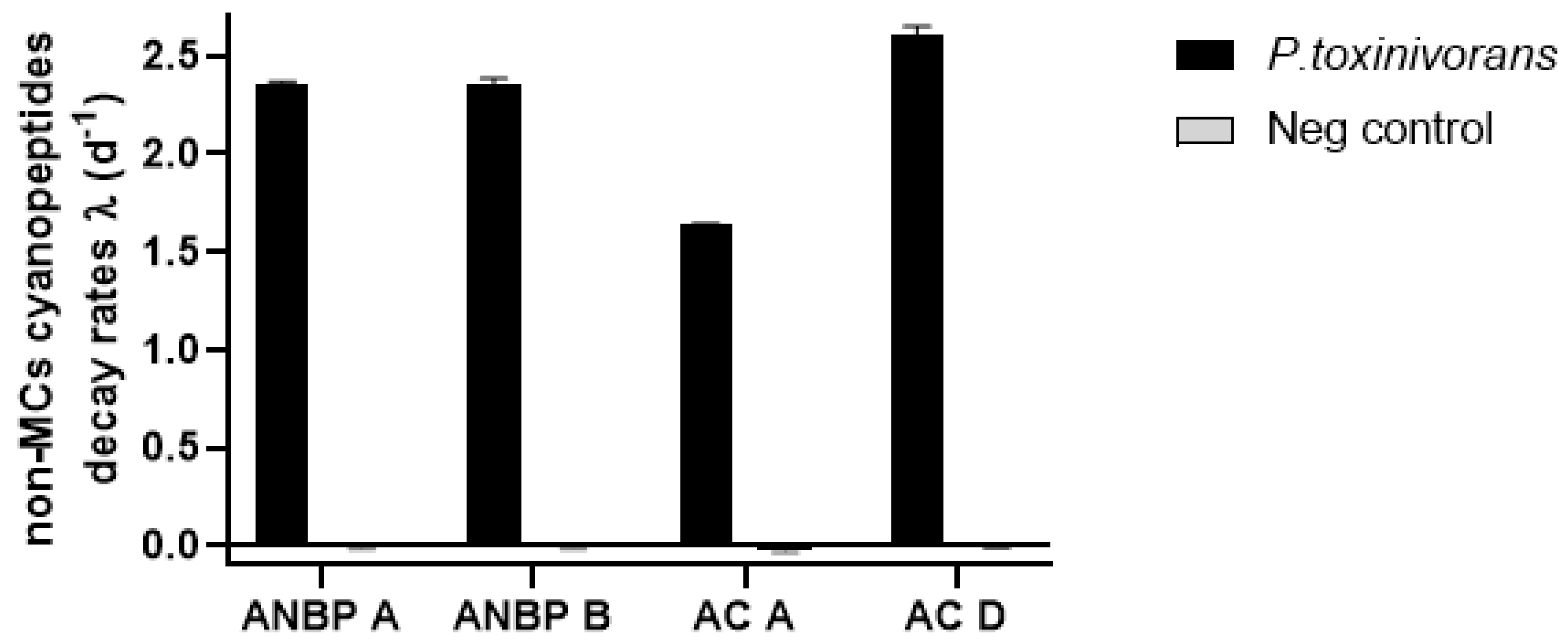
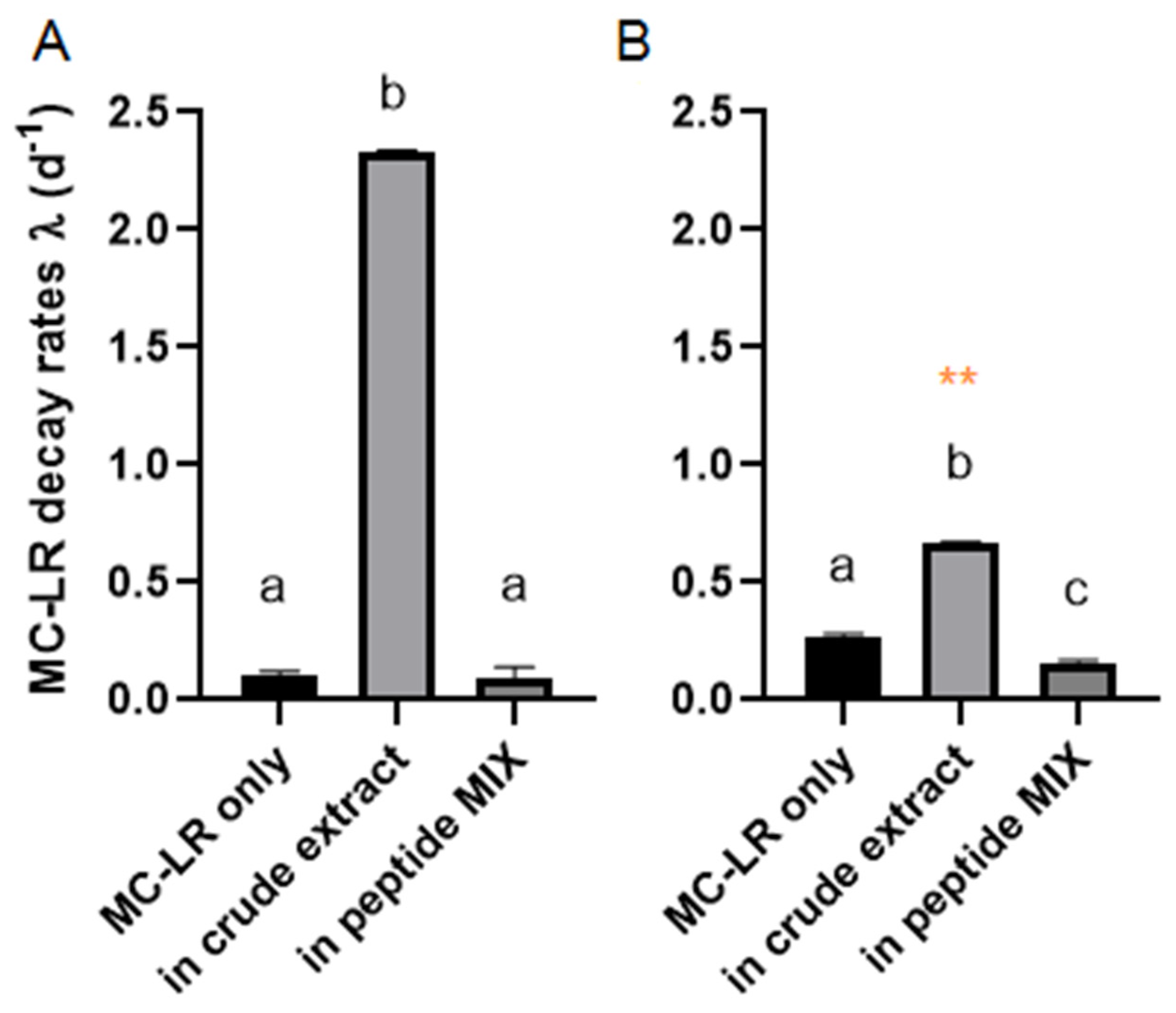
| Peptide | Peptide Remaining Concentration (%) | |||
|---|---|---|---|---|
| Paucibacter toxinivorans | Negative Control | |||
| 3rd Day | 7th Day | 3rd Day | 7th Day | |
| Individual | ||||
| MC-LR | 72 ± 2.8 | 15 ± 1.1 | 104 ± 2.8 | 106 ± 3.4 |
| DM-LR | 54 ± 0.8 | 0.7 ± 1.3 | 106 ± 2.2 | 119 ± 0.3 |
| MC-RR | 49 ± 6.1 | 9 ± 4.1 | 101 ± 8.2 | 119 ± 4.4 |
| MC-LF | 49 ± 2.5 | nd | 143 ± 4.5 | 126 ± 5.5 |
| MC-YR | 59 ± 4.0 | 1 ± 1.7 | 115 ± 7.5 | 119 ± 0.4 |
| ANBP-A | nd | nd | 112 ± 8.1 | 112 ± 10.1 |
| ANBP-B | nd | nd | 100 ± 17.1 | 99 ± 18.1 |
| AC-A | nd | nd | 125 ± 14.1 | 126 ± 14.3 |
| AC-D | nd | nd | 113 ± 1.6 | 102 ± 4.6 |
| In MIX | ||||
| MC-LR | 77 ± 11.6 | 32 ± 1.9 | 112 ± 6.8 | 121 ± 11.9 |
| DM-LR | 77 ± 5.4 | 33 ± 1.1 | 108 ± 5.2 | 121 ± 11.3 |
| MC-RR | 81 ± 5.9 | 54 ± 0.7 | 109 ± 6.2 | 119 ± 9.9 |
| MC-LF | 94 ± 7.2 | 68 ± 3.2 | 107 ± 3.4 | 109 ± 8.2 |
| MC-YR | 72 ± 4.3 | 28 ± 0.7 | 107 ± 5.1 | 118 ± 10.3 |
| ANBP-A | nd * | nd * | 111 ± 7.7 | 123 ± 11.1 |
| ANBP-B | nd * | nd * | 116 ± 9.5 | 129 ± 4.5 |
| AC-A | nd * | nd * | 109 ± 5.6 | 101 ± 7.6 |
| AC-D | nd * | nd * | 103 ± 6.9 | 97 ± 8.2 |
| In crude extract of M. aeruginosa | ||||
| MC-LR | 0.9 ± 0.02 | 0.9 ± 0.02 | 100 ± 3.9 | 104 ± 7.6 |
| (A) | |||
| MC-LR Linear By-Product A | MC-LR Linear By-Product B | ||
| m/z | Identified Structure | m/z | Identified Structure |
| 1013.5646 | M+H, Adda-Glu-Mdha-Ala-Leu-Masp-Arg-OH+H | 1013.5649 | M+H, Adda-Glu-Mdha-Ala-Leu-Masp-Arg-OH+H |
| 862.4684 | M+H-NH2-PhCH2CHOMe | 879.5062 | M+H-PhCH2CHOMe |
| 756.3956 | CH3-CHCO-Glu-Mdha-Ala-Leu-Masp-Arg-OH+2H | 821.4431 | Not detected |
| 726.3453 | CO-Glu-Mdha-Ala-Leu-Masp-Arg-OH+2H | 743.3694 | Not detected |
| - | - | 663.3569 | Not detected |
| 571.3195 | Mdha-Ala-Leu-Masp-Arg-OH+2H | 588.3467 | Not detected |
| 488.2766 | Ala-Leu-Masp-Arg-OH+2H | 488.2812 | Ala-Leu-Masp-Arg-OH+2H |
| 375.1940 | C11H14O-Glu-Mdha+H | 363.1936 | Not detected |
| 304.1622 | Masp-Arg-OH+2H | 304.1642 | Masp-Arg-OH+2H |
| 286.1524 | Masp-Arg+H | 286.1516 | Masp-Arg+H |
| 175.1189 | Arg-OH+2H | 175.1196 | Arg-OH+2H |
| 135.0814 | PhCH2CHOMe | 135.0719 | PhCH2CHOMe |
| (B) | |||
| MC-YR Linear By-Product C | MC-YR Linear By-Product D | ||
| m/z | Identified Structure | m/z | Identified Structure |
| 1063.6191 | M+H, Adda-Glu-Mdha-Ala-Tyr-Masp-Arg-OH+H | 1063.6200 | M+H, Adda-Glu-Mdha-Ala-Tyr-Masp-Arg-OH+H |
| 912.4456 | M+H-NH2-PhCH2CHOMe | 929.4713 | M+H-PhCH2CHOMe |
| - | - | 893.4518 | Not detected |
| 621.3013 | Mdha-Ala-Tyr-Masp-Arg-OH+2H | 731.3482 | Not detected |
| - | - | 638.3301 | Not detected |
| 538.2344 | Ala-Tyr-Masp-Arg-OH+2H | 538.2687 | Ala-Tyr-Masp-Arg-OH+2H |
| 450.2138 | Not detected | - | - |
| 375.1960 | C11H14O-Glu-Mdha+H | 363.1893 | Not detected |
| 304.1624 | Masp-Arg-OH+2H | 304.1621 | Masp-Arg-OH+2H |
| 286.1526 | Masp-Arg+H | 286.1523 | Masp-Arg+H |
| 213.1116 | Glu-Mdha+H | - | - |
| 175.1196 | Arg-OH+2H | 175.1199 | Arg-OH+2H |
| (C) | |||
| MC-LF Linear By-Product E | |||
| m/z | Identified Structure | ||
| 1004.5352 | M+H, Adda-Glu-Mdha-Ala-Leu-Masp-Phe-OH+H | ||
| 870.4624 | M+H-PhCH2CHOMe | ||
| 853.4357 | M+H-NH2-PhCH2CHOMe, (M+H-151) | ||
| 705.3755 | (Masp-Leu-Ala-Mhda-Glu-Adda-OH-H)-151 | ||
| 576.3404 | (Leu-Ala-Mhda-Glu-Adda-OH-H)-151 | ||
| 463.2590 | (Ala-Mhda-Glu-Adda-OH-H)-151 | ||
| 435.2520 | Not detected | ||
| 418.2407 | Not detected | ||
| 375.1956 | C11H14O-Glu-Mdha+H | ||
| 363.1892 | Not detected | ||
| 335.1969 | Not detected | ||
| 295.1309 | Masp-Phe-OH+2H | ||
| 206.1505 | Not detected | ||
| 163.1154 | PhCH2-CHOCH3CHCH3 | ||
| 135.1164 | PhCH2CHOMe | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Santos, A.A.; Soldatou, S.; de Magalhães, V.F.; Azevedo, S.M.F.O.; Camacho-Muñoz, D.; Lawton, L.A.; Edwards, C. Degradation of Multiple Peptides by Microcystin-Degrader Paucibacter toxinivorans (2C20). Toxins 2021, 13, 265. https://doi.org/10.3390/toxins13040265
Santos AA, Soldatou S, de Magalhães VF, Azevedo SMFO, Camacho-Muñoz D, Lawton LA, Edwards C. Degradation of Multiple Peptides by Microcystin-Degrader Paucibacter toxinivorans (2C20). Toxins. 2021; 13(4):265. https://doi.org/10.3390/toxins13040265
Chicago/Turabian StyleSantos, Allan A., Sylvia Soldatou, Valeria Freitas de Magalhães, Sandra M. F. O. Azevedo, Dolores Camacho-Muñoz, Linda A. Lawton, and Christine Edwards. 2021. "Degradation of Multiple Peptides by Microcystin-Degrader Paucibacter toxinivorans (2C20)" Toxins 13, no. 4: 265. https://doi.org/10.3390/toxins13040265
APA StyleSantos, A. A., Soldatou, S., de Magalhães, V. F., Azevedo, S. M. F. O., Camacho-Muñoz, D., Lawton, L. A., & Edwards, C. (2021). Degradation of Multiple Peptides by Microcystin-Degrader Paucibacter toxinivorans (2C20). Toxins, 13(4), 265. https://doi.org/10.3390/toxins13040265