Regulation Efficacy and Mechanism of the Toxicity of Microcystin-LR Targeting Protein Phosphatase 1 via the Biodegradation Pathway
Abstract
:1. Introduction
2. Results and Discussion
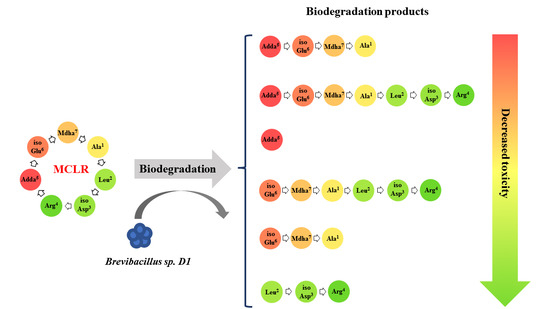
2.1. MCLR Biodegradation Products Identification
2.2. Biological Toxicity Evaluation of the MCLR Biodegradation Products Targeting PP1
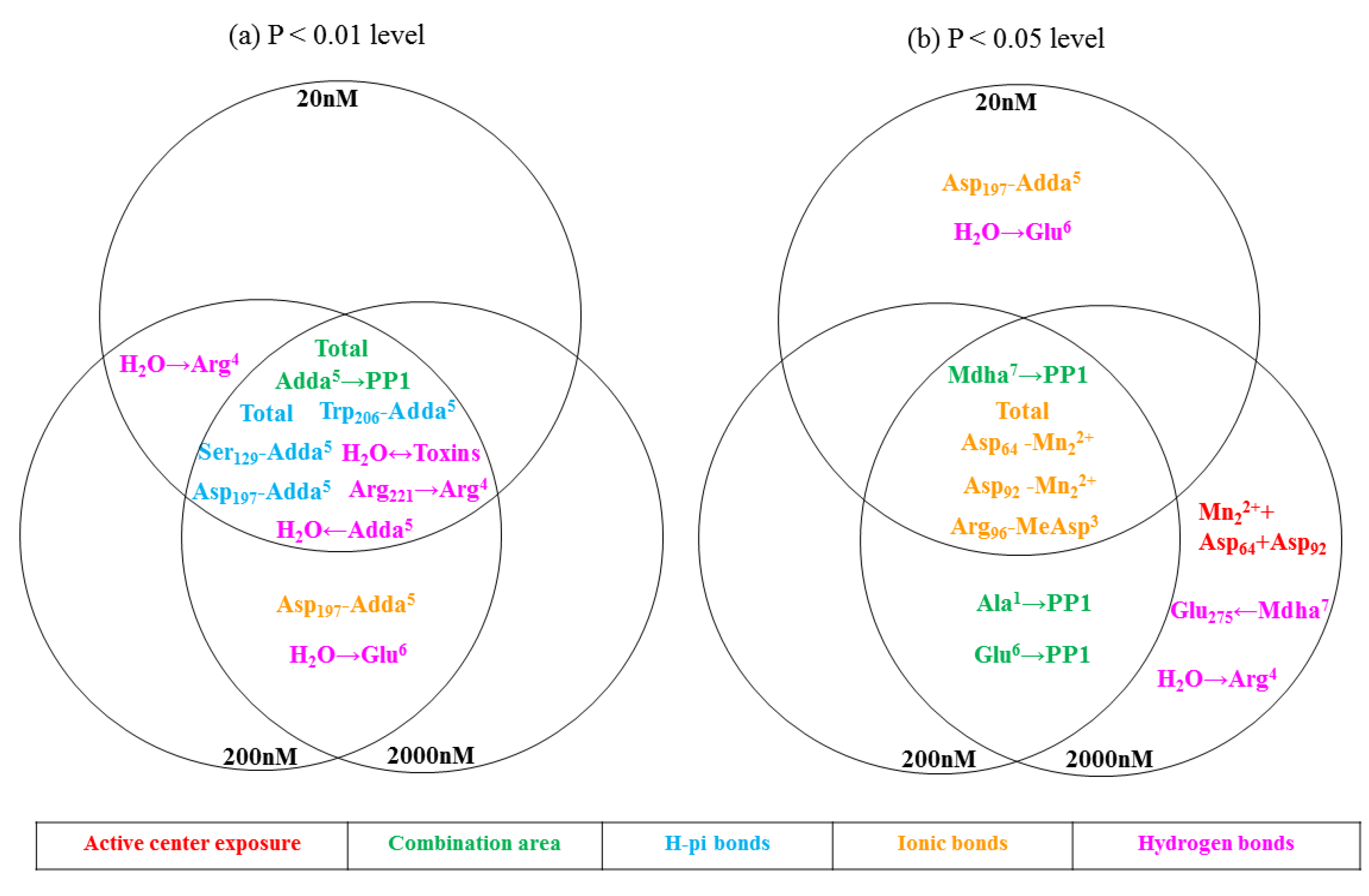
2.3. Molecular Mechanism for the Different Toxicity of MCLR and Its Biodegradation Products Targeting PP1
3. Conclusions
4. Materials and Methods
4.1. Materials
4.2. Biodegradation of MCLR
4.3. MCLR Biodegradation Products Analysis
4.4. MCLR Biodegradation Products Preparation
4.5. Protein Phosphatase 1 Inhibition Assay
4.6. Molecular Docking for the Interaction between Toxins and PP1
4.7. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Kull, T.P.J.; Backlund, P.H.; Karlsson, K.M.; Meriluoto, J.A.O. Oxidation of the cyanobacterial hepatotoxin microcystin-LR by chlorine dioxide: Reaction kinetics, characterization, and toxicity of reaction products. Environ. Sci. Technol. 2004, 38, 6025–6031. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Xie, P.; Liu, Y.; Qiu, T. Transfer, distribution and bioaccumulation of microcystins in the aquatic food web in Lake Taihu, China, with potential risks to human health. Sci. Total Environ. 2009, 407, 2191–2199. [Google Scholar] [CrossRef] [PubMed]
- de Figueiredo, D.R.; Azeiteiro, U.M.; Esteves, S.M.; Goncalves, F.J.M.; Pereira, M.J. Microcystin-producing blooms—A serious global public health issue. Ecotoxicol. Environ. Saf. 2004, 59, 151–163. [Google Scholar] [CrossRef] [PubMed]
- Antoniou, M.G.; de la Cruz, A.A.; Dionysiou, D.D. Cyanotoxins: New generation of water contaminants. J. Environ. Eng. 2005, 131, 1239–1243. [Google Scholar] [CrossRef]
- Pearson, L.; Mihali, T.; Moffitt, M.; Kellmann, R.; Neilan, B. On the chemistry, toxicology and genetics of the cyanobacterial toxins, microcystin, nodularin, saxitoxin and cylindrospermopsin. Mar. Drugs 2010, 8, 1650–1680. [Google Scholar]
- Gulledge, B.M.; Aggen, J.B.; Eng, H.; Sweimeh, K.; Chamberlin, A.R. Microcystin analogues comprised only of Adda and a single additional amino acid retain moderate activity as PP1/PP2A inhibitors. Bioorg. Med. Chem. Lett. 2003, 13, 2907–2911. [Google Scholar] [CrossRef]
- Campos, A.; Vasconcelos, V. Molecular mechanisms of microcystin toxicity in animal cells. Int. J. Mol. Sci. 2010, 11, 268–287. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Craig, M.; Luu, H.A.; McCready, T.L.; Holmes, C.F.B.; Williams, D.; Andersen, R.J. Molecular mechanisms underlying the interaction of motuporin and microcystins with type-1 and type-2A protein phosphatases. Biochem. Cell Biol. 1996, 74, 569–578. [Google Scholar] [CrossRef] [PubMed]
- Maynes, T.; Bateman, K.S.; Cherney, M.M.; Das, A.K.; Luu, H.A.; Holmes, C.F.; James, M.N. Crystal structure of the tumor-promoter okadaic acid bound to protein phosphatase-1. J. Biol. Chem. 2001, 276, 44078–44082. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zegura, B.; Sedmak, B.; Filipič, M. Microcystin-LR induces oxidative DNA damage in human hepatoma cell line HepG2. Toxicon 2003, 41, 41–48. [Google Scholar] [CrossRef]
- Li, J.; Li, R.; Li, J. Current research scenario for microcystins biodegradation—A review on fundamental knowledge, application prospects and challenges. Sci. Total Environ. 2017, 595, 615–632. [Google Scholar] [CrossRef] [PubMed]
- Duy, T.N.; Lam, P.K.S.; Shaw, G.R.; Connell, D.W. Toxicology and risk assessment of freshwater cyanobacterial (blue-green algal) toxins in water. Rev. Environ. Contam. Toxicol. 2000, 163. [Google Scholar]
- Dziga, D.; Wasylewski, M.; Wladyka, B.; Nybom, S.; Meriluoto, J. Microbial degradation of microcystins. Chem. Res. Toxicol. 2013, 26, 841–852. [Google Scholar] [CrossRef] [PubMed]
- Imanishi, S.; Kato, H.; Mizuno, M.; Tsuji, K.; Harada, K.I. Bacterial degradation of microcystins and nodularin. Chem. Res. Toxicol. 2005, 18, 591–598. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.O.D.; Blom, J.F.; Yankova, Y.; Villiger, J.; Pernthaler, J. Priming of microbial microcystin degradation in biomass-fed gravity driven membrane filtration biofilms. Syst. Appl. Microbiol. 2018, 41, 221–231. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zong, W.; Sun, F.; Sun, X. Oxidation by-products formation of microcystin-LR exposed to UV/H2O2: Toward the generative mechanism and biological toxicity. Water Res. 2013, 47, 3211–3219. [Google Scholar] [CrossRef] [PubMed]
- Zong, W.; Sun, F.; Pei, H.; Hu, W.; Pei, R. Microcystin-associated disinfection by-products: The real and non-negligible risk to drinking water subject to chlorination. Chem. Eng. J. 2015, 279, 498–506. [Google Scholar] [CrossRef]
- Zong, W.; Sun, F.; Sun, X. Evaluation on the generative mechanism and biological toxicity of microcystin-LR disinfection by-products formed by chlorination. J. Hazard. Mater. 2013, 252, 293–299. [Google Scholar] [CrossRef] [PubMed]
- Zong, W.; Wang, X.; Du, Y.; Zang, S.; Zang, Y.; Teng, Y. Molecular Mechanism for the Regulation of Microcystin Toxicity to Protein Phosphatase 1 by Glutathione Conjugation Pathway. BioMed Res. Int. 2017, 2017, 9676504. [Google Scholar] [CrossRef] [PubMed]
- Fontanillo, M.; Köhn, M. Microcystins: Synthesis and structure–activity relationship studies toward PP1 and PP2A. Bioorg. Med. Chem. 2018, 26, 1118–1126. [Google Scholar] [CrossRef] [PubMed]





| Detected Ions Identified Products | Ion Types | Linearized MCLR | Glu6-Mdha7-Ala1-Leu2-MeAsp3-Arg4 | Adda5-Glu6-Mdha7-Ala1 | Leu2-MeAsp3-Arg4 | Glu6-Mdha7-Ala1 | Adda5 |
|---|---|---|---|---|---|---|---|
| [M+H]+ | parent ions | 1013.5703 | 700.3631 | 615.3387 | 417.2452 | 302.1343 | 332.2218 |
| [PhCH2CH(OCH3)]+ | side-chain | 135.0809 | --- | √ | --- | --- | √ |
| [(-Arg4)+H]+ | y-type | 174.1110 | √ | --- | √ | --- | --- |
| [(-MeAsp3-Arg4)+H]+ | y-type | 303.1536 | √ | --- | √ | --- | --- |
| [(Adda5-)+H]+ | b-type | 315.2192 | --- | √ | --- | --- | ↑+OH |
| [(-Leu2-MeAsp3-Arg4)+H]+ | y-type | 416.2377 | √ | --- | ↑+H | --- | --- |
| [(Adda5-Glu6-)+H]+ | b-type | 444.2618 | --- | √ | --- | --- | --- |
| [(-Ala1-Leu2-MeAsp3-Arg)4+H]+ | y-type | 487.2048 | √ | --- | --- | --- | --- |
| [(Adda5-Glu6-Mdha7-)+H]+ | b-type | 527.2988 | --- | √ | --- | --- | --- |
| [(-Mdha7-Ala1-Leu2-MeAsp3-Arg4)+H]+ | y-type | 570.3119 | √ | --- | --- | --- | --- |
| [(Adda5-Glu6-Mdha7-Ala1-)+H]+ | b-type | 598.3360 | --- | ↑+OH | --- | --- | --- |
| [(-Glu6-Mdha7-Ala1-Leu2-MeAsp3-Arg4)+H]+ | y-type | 699.3545 | ↑+H | --- | --- | --- | --- |
| [(Adda5-Glu6-Mdha7-Ala1-Leu2-)+H]+ | b-type | 711.4201 | --- | --- | --- | --- | --- |
| [(Adda5-Glu6-Mdha7-Ala1-Leu2-MeAsp3-)+H]+ | b-type | 840.4627 | --- | --- | --- | --- | --- |
| [(Glu6-)+H]+ | b-type | --- | √ | --- | --- | 131.0576 | --- |
| [(-Mdha7-Ala1)+H]+ | y-type | --- | --- | √ | --- | 172.0842 | --- |
| [(Glu6-Mdha7-)+H]+ | b-type | --- | √ | --- | --- | 214.0947 | --- |
| [(-Ala1)+H]+ | y-type | --- | --- | √ | --- | 89.0470 | --- |
| Biodegradation Products | Eluted Times | Biodegradation Times | Total Collection Volumes | Final Concentrations a | Purity b |
|---|---|---|---|---|---|
| MCLR (995.5557) | 25.26 ± 0.20 min | --- | --- | --- | --- |
| Linearised MCLR (1013.5703) | 23.79 ± 0.20 min | 4, 6, 8 days | 800 × 3 = 2400 µL | 41.06 µM/L | 94.2% |
| Glu6-Mdha7-Ala1-Leu2-MeAsp3-Arg4 (700.3631) | 20.44 ± 0.20 min | 8, 10, 12 days | 800 × 3 = 2400 µL | 19.71 µM/L | 94.8% |
| Adda5-Glu6-Mdha7-Ala1 (615.3387) | 19.27 ± 0.20 min | 6, 8, 10, 12 days | 800 × 4 = 3200 µL | 15.93 µM/L | 96.0% |
| Leu2-MeAsp3-Arg4 (417.2452) | 12.65 ± 0.20 min | 8, 10, 12, 14 days | 800 × 4 = 3200 µL | 21.10 µM/L | 96.7% |
| Glu6-Mdha7-Ala1 (302.1343) | 15.22 ± 0.20 min | 12, 14, 16, 18, 20 days | 800 × 5 = 4000 µL | 7.95 µM/L | 94.7% |
| Adda5 (332.2218) | 23.18 ± 0.20 min | 8, 10, 12, 14, 16, 18, 20 days | 800 × 7 = 5600 µL | 19.88 µM/L | 97.4% |
| Pearson Correlation Analysis Data a | Binding Energy (KJ/Mol) | Binding Area (Å2) | Catalytic Center Exposure (Å2) | |||||||||
| Total | Ala1→PP1 | Leu2→PP1 | MeAsp3→PP1 | Arg4→PP1 | Adda5→PP1 | Glu6→PP1 | Mdha7→PP1 | Mn12+/Mn22+ | Mn22++ Asp64+Asp92 | |||
| Toxicity (20 nM) | R | −0.177 | 0.746 ** | 0.364 | 0.107 | −0.158 | 0.061 | 0.600 ** | 0.421 | 0.455 * | 0.001 | −0.159 |
| p | 0.444 | 0.000 | 0.104 | 0.644 | 0.494 | 0.793 | 0.004 | 0.057 | 0.038 | 0.999 | 0.492 | |
| Toxicity (200 nM) | R | −0.297 | 0.866 ** | 0.447 * | 0.101 | −0.186 | 0.045 | 0.712 ** | 0.485 * | 0.519 * | 0.001 | −0.381 |
| p | 0.190 | 0.000 | 0.042 | 0.664 | 0.420 | 0.848 | 0.000 | 0.026 | 0.016 | 0.999 | 0.088 | |
| Toxicity (2000 nM) | R | −0.401 | 0.919 ** | 0.437 * | 0.125 | −0.188 | 0.060 | 0.786 ** | 0.490 * | 0.511 * | 0.001 | −0.531 * |
| p | 0.072 | 0.000 | 0.048 | 0.589 | 0.414 | 0.795 | 0.000 | 0.024 | 0.018 | 0.999 | 0.109 | |
| Pearson Correlation Analysis Data a | H-pi bonds (KJ/Mol) | Ionic bonds (KJ/Mol) | ||||||||||
| Total | Trp206-Adda5 | Ser129-Adda5 | Asp197-Adda5 | Total | Asp64-Mn22+ | Asp92-Mn22+ | Arg96-MeAsp3 | Asp220-Arg4 | Asp197-Adda5 | Glu275-Mdha7 | ||
| Toxicity (20 nM) | R | −0.939 ** | −0.692 ** | −0.884 ** | −0.745 ** | −0.463 * | −0.455 * | −0.453 * | −0.516 * | −0.161 | −0.544 * | −0.100 |
| p | 0.000 | 0.001 | 0.000 | 0.000 | 0.034 | 0.038 | 0.039 | 0.017 | 0.484 | 0.011 | 0.667 | |
| Toxicity (200 nM) | R | −0.848 ** | −0.786 ** | −0.732 ** | −0.768 ** | −0.498 * | −0.524 * | −0.527 * | −0.534 * | −0.147 | −0.695 ** | −0.276 |
| p | 0.000 | 0.000 | 0.000 | 0.000 | 0.022 | 0.015 | 0.014 | 0.013 | 0.526 | 0.000 | 0.226 | |
| Toxicity (2000 nM) | R | −0.762 ** | −0.735 ** | −0.615 ** | −0.795 ** | −0.488 * | −0.510 * | −0.517 * | −0.491 * | −0.166 | −0.797 ** | −0.374 |
| p | 0.000 | 0.000 | 0.003 | 0.000 | 0.025 | 0.018 | 0.016 | 0.024 | 0.471 | 0.000 | 0.095 | |
| Pearson Correlation Analysis Data a | Hydrogen bonds (KJ/Mol) | |||||||||||
| Total | H2O↔Toxins | H2O←Adda5 | H2O→Arg4 | H2O→Glu6 | Asp220←Arg4 | Glu275←Arg4 | Glu275←Mdha7 | Arg96→MeAsp3 | Arg221→Arg4 | |||
| Toxicity (20 nM) | R | −0.358 | −0.843 ** | −0.748 ** | −0.807 ** | −0.487 * | −0.094 | −0.085 | 0.044 | −0.029 | −0.799 ** | |
| p | 0.111 | 0.000 | 0.000 | 0.000 | 0.025 | 0.686 | 0.737 | 0.851 | 0.901 | 0.000 | ||
| Toxicity (200 nM) | R | −0.355 | −0.761 ** | −0.668 ** | −0.624 ** | −0.606 ** | −0.112 | 0.023 | −0.220 | −0.022 | −0.669 ** | |
| p | 0.114 | 0.000 | 0.001 | 0.003 | 0.004 | 0.629 | 0.929 | 0.338 | 0.924 | 0.002 | ||
| Toxicity (2000 nM) | R | −0.306 | −0.697 ** | −0.597 ** | −0.501 * | −0.638 ** | −0.156 | 0.089 | −0.440 * | −0.103 | −0.627 ** | |
| p | 0.109 | 0.001 | 0.004 | 0.021 | 0.002 | 0.499 | 0.725 | 0.046 | 0.657 | 0.005 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ren, L.; Hu, Z.; Wang, Q.; Du, Y.; Zong, W. Regulation Efficacy and Mechanism of the Toxicity of Microcystin-LR Targeting Protein Phosphatase 1 via the Biodegradation Pathway. Toxins 2020, 12, 790. https://doi.org/10.3390/toxins12120790
Ren L, Hu Z, Wang Q, Du Y, Zong W. Regulation Efficacy and Mechanism of the Toxicity of Microcystin-LR Targeting Protein Phosphatase 1 via the Biodegradation Pathway. Toxins. 2020; 12(12):790. https://doi.org/10.3390/toxins12120790
Chicago/Turabian StyleRen, Luyao, Zhengxin Hu, Qian Wang, Yonggang Du, and Wansong Zong. 2020. "Regulation Efficacy and Mechanism of the Toxicity of Microcystin-LR Targeting Protein Phosphatase 1 via the Biodegradation Pathway" Toxins 12, no. 12: 790. https://doi.org/10.3390/toxins12120790
APA StyleRen, L., Hu, Z., Wang, Q., Du, Y., & Zong, W. (2020). Regulation Efficacy and Mechanism of the Toxicity of Microcystin-LR Targeting Protein Phosphatase 1 via the Biodegradation Pathway. Toxins, 12(12), 790. https://doi.org/10.3390/toxins12120790