Abstract
Nowadays, proliferation of jellyfish has become a severe matter in many coastal areas around the world. Jellyfish Nemopilema nomurai is one of the most perilous organisms and leads to significant deleterious outcomes such as harm to the fishery, damage the coastal equipment, and moreover, its envenomation can be hazardous to the victims. Till now, the components of Nemopilema nomurai venom (NnV) are unknown owing to scant transcriptomics and genomic data. In the current research, we have explored a proteomic approach to identify NnV components and their interrelation with pathological effects caused by the jellyfish sting. Altogether, 150 proteins were identified, comprising toxins and other distinct proteins that are substantial in nematocyst genesis and nematocyte growth by employing two-dimensional gel electrophoresis and matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI/TOF/MS). The identified toxins are phospholipase A2, phospholipase D Li Sic Tox beta IDI, a serine protease, putative Kunitz-type serine protease inhibitor, disintegrin and metalloproteinase, hemolysin, leukotoxin, three finger toxin MALT0044C, allergens, venom prothrombin activator trocarin D, tripeptide Gsp 9.1, and along with other toxin proteins. These toxins are relatively well characterized in the venoms of other poisonous species to induce pathogenesis, hemolysis, inflammation, proteolysis, blood coagulation, cytolysis, hemorrhagic activity, and type 1 hypersensitivity, suggesting that these toxins in NnV can also cause similar deleterious consequences. Our proteomic works indicate that NnV protein profile represents valuable source which leads to better understanding the clinical features of the jellyfish stings. As one of the largest jellyfish in the world, Nemopilema nomurai sting is considered to be harmful to humans due to its potent toxicity. The identification and functional characterization of its venom components have been poorly described and are beyond our knowledge. Here is the first report demonstrating the methodical overview of NnV proteomics research, providing significant information to understand the mechanism of NnV envenomation. Our proteomics findings can provide a platform for novel protein discovery and development of practical ways to deal with jellyfish stings on human beings.
Key Contribution:
This study is the first to examine comprehensively the proteome of poisonous Nemopilema nomurai jellyfish venom, which can be valuable information in understanding the venom toxin components and their mechanism of actions as well as helpful for treating the poisoned patients.
1. Introduction
Over the recent decades, there is a huge expansion of jellyfish blooms worldwide, which cause severe damage to the fishery and disturb marine ecosystem [1]. Since 1983, more than 2000 cases of N. nomurai jellyfish accidents have been reported in the coastal areas of China, Korea, and Japan, and life-threatening cases were also observed in humans [2]. N. nomurai known as the giant jellyfish is one of the most dangerous species belonging to the Phylum Cnidaria, being their diagnostic feature presence of stinging organelles called nematocysts, located mostly on the tentacles of jellyfish [3,4]. On sudden stimulation, nematocysts explosively discharge various venom constituents into the preys or victims [4]. Many scientists performed toxicological research on NnV, which includes cardiotoxic, hepatotoxic, hemolytic and cytotoxic biological activities [5,6,7,8]. Edema, itching, burning sensation, and small vesicles with erythematous eruption appeared at the site of contact on the victim’s body after NnV envenomation [9,10]. In the previous research, we have evaluated the cardiotoxic effect of NnV in H9c2 cells using a proteomic strategy [6]. Beside the toxicological and pharmacological importance of NnV, till now its venom composition has not been well defined. Classification and isolation of toxic proteins is a hard and laborious process.
In many venomous creatures such as snakes, spiders, scorpions, and cone snails, proteomic approaches have been auspiciously exploited to purify and characterize their venom components [11,12,13,14]. A few investigators have also attempted to fractionate and identify jellyfish venom toxins [15,16,17,18]. However, it is not so astonishing that jellyfish venom transcriptomics, and genomic data are rarely available, that, if any, may aid in the detection of bioactive venom components individually. For the present study, we have used the proteomic approach by utilizing two-dimensional gel electrophoresis, followed by matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI/TOF/MS) and bioinformatics analyses to determine NnV composition that could explain life-threatening and undesirable consequences in human envenomation.
2. Results
2.1. Identification of N. nomurai Nematocyst Proteins by Proteomic Characterization
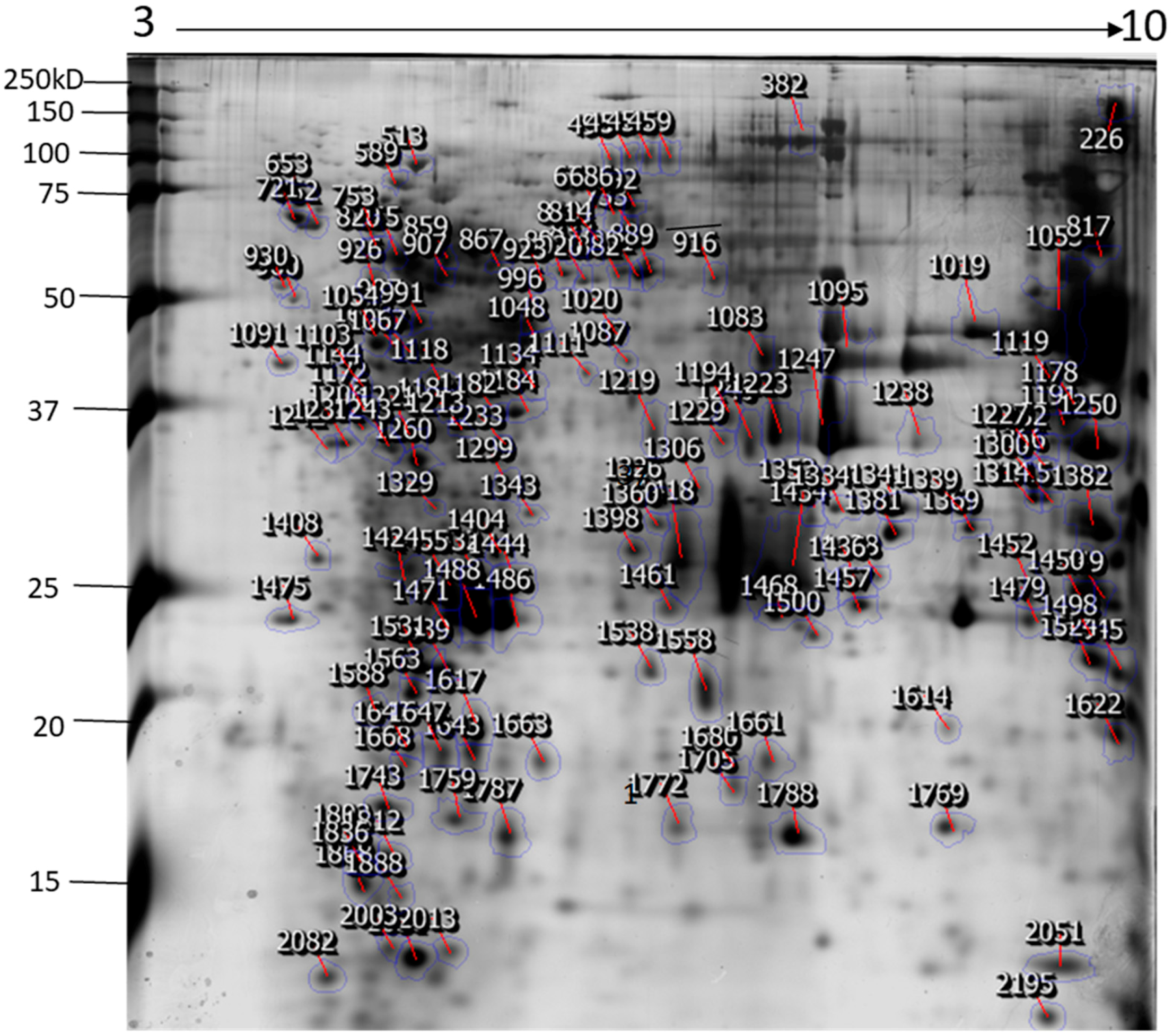
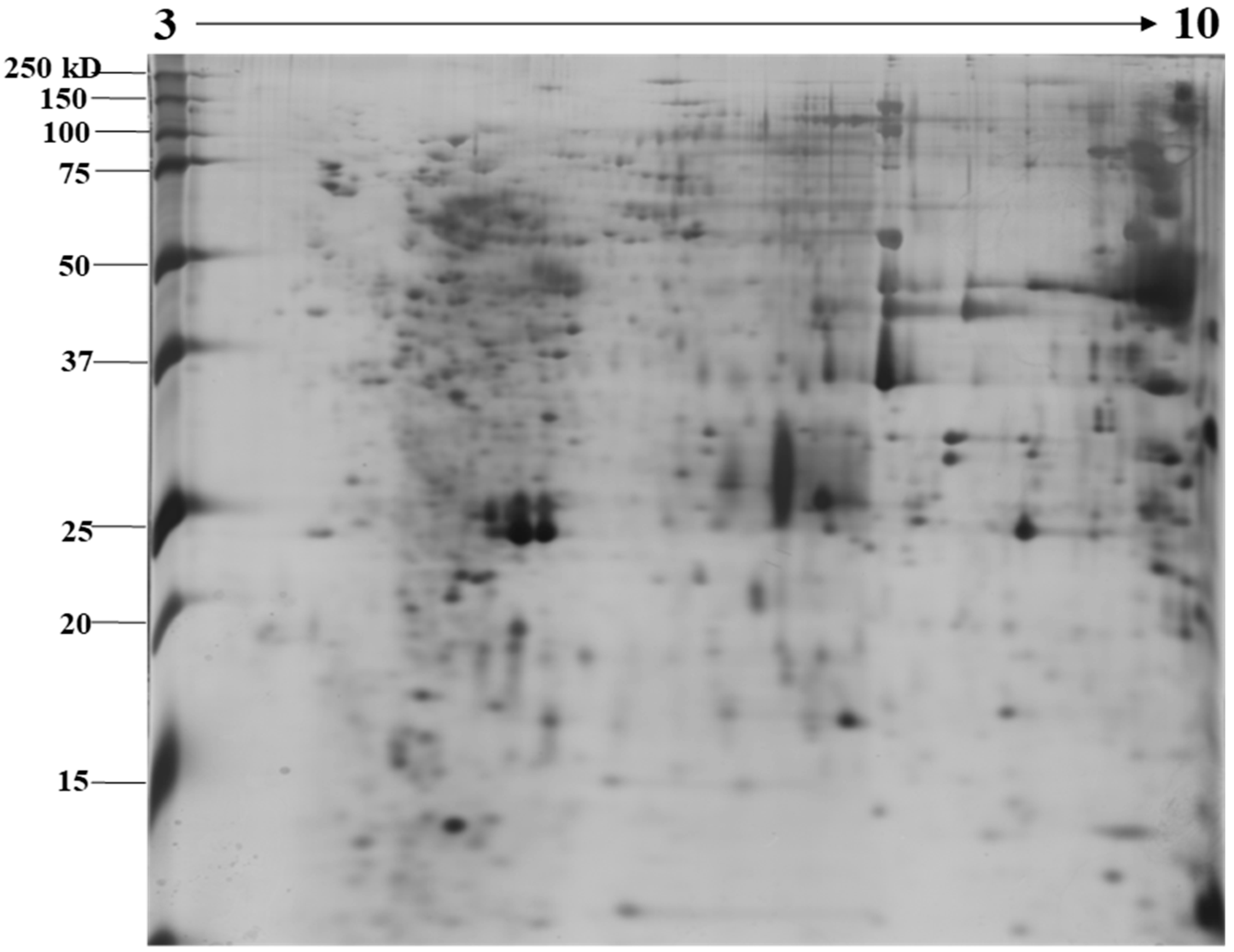
The identification and functional characterization of NnV components have been poorly characterized. Herein, we described the proteomic profile of the NnV. Our present study has successfully demonstrated the proteomic characterization of NnV by utilizing 2-DE and MALDI/TOF/MS. 2-DE gel electrophoresis, revealing the venom components ranging from PI 3–10 and molecular weights between 15–250 kDa, as shown in Figure 1. The scanned 2-DE image was marked with arrows and boundaries, generated by Progenesis Same Spots software (Nonlinear Dynamics, New Castle, UK), as shown in Figure 2. In this study, a total of 150 proteins identified from the nematocysts of NnV, including some toxins and another distinct type of proteins which are substantial in nematocyst and nematocyte generation (Table 1, Supplement Table). Interestingly, the identified toxins from N. nomurai jellyfish have shown high 71 sequence similarity with those of other venomous and poisonous animals. Mainly, it is composed of Phospholipase A2, Phospholipase D Li Sic Tox beta IDI, Serine protease, Putative Kunitz-type Serine protease inhibitor, Disintigrin and Metalloproteinase, Hemolysin, Leukotoxin, Three-finger toxin MALT0044C, allergens, Venom prothrombin activator trocarin D, Tripeptide Gsp 9.1 and Cell death abnormality protein 1 from Caenorhabditis elegans. Several toxin proteins shared homology with proteins from microorganisms such as RTX-III toxin determinant A from serotype 2 from Actinobacillus pleuropneumoniae, Shiga-like toxin 1 subunit A from Enterobacteria phage, Leukotoxin from Pasteurella haemolytica, Cell death abnormality protein 12 from Saccharomyces cerevisiae, and Pro-apoptotic serine protease nma111 from Neurospora crassa. Interestingly, it also includes non-toxic proteins during NnV profiling, such as PCNA-interacting partner, Fukutin, Cell death abnormality protein 1, UvrABC system protein A, Division control protein 7, and OTU domain-containing protein 7B.
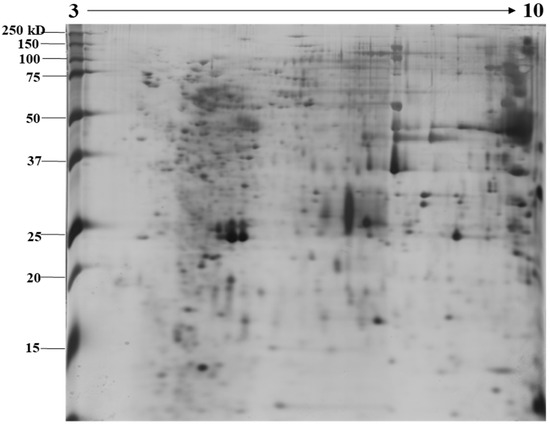
Figure 1.
Image showing proteomic analysis of 2-DE of NnV. Representative 2-DE image of N. nomurai jellyfish venom (A). For the first dimension, 500 µg of proteins were resolved on an 18 cm, IPG dry strips (pH 3–10) and 12% SDS-PAGE gels were used to run second dimension. 2-DE gels were stained using the silver staining method and the Epson perfection V 700 photo scanner was used for scanning the stained gels. For statistical analysis, three independent replicate gels were run.
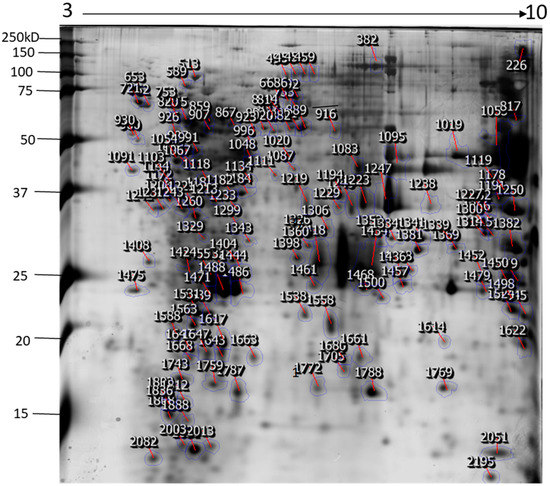

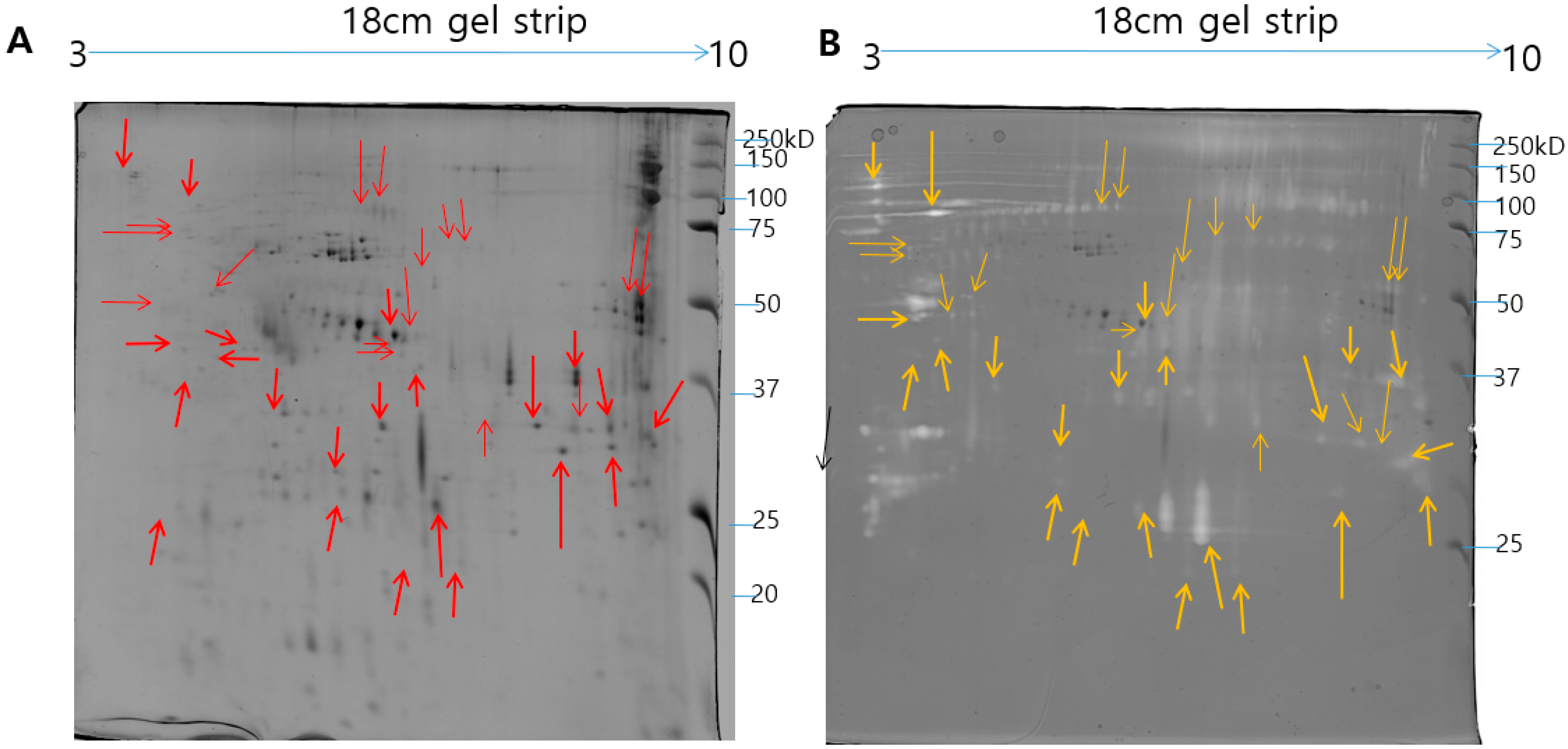
Figure 2.
The representative 2-DE image of NnV proteins generated by Progenesis Same Spots software. Boundaries and arrows signify the position of differentially expressed proteins. Putative numbers were assigned to each protein spot.

Table 1.
Protein identified in N. nomurai venom by using MALDI/TOF/MS.
2.2. Modified Zymography Identify Metalloproteinase and PLA2 in NnV
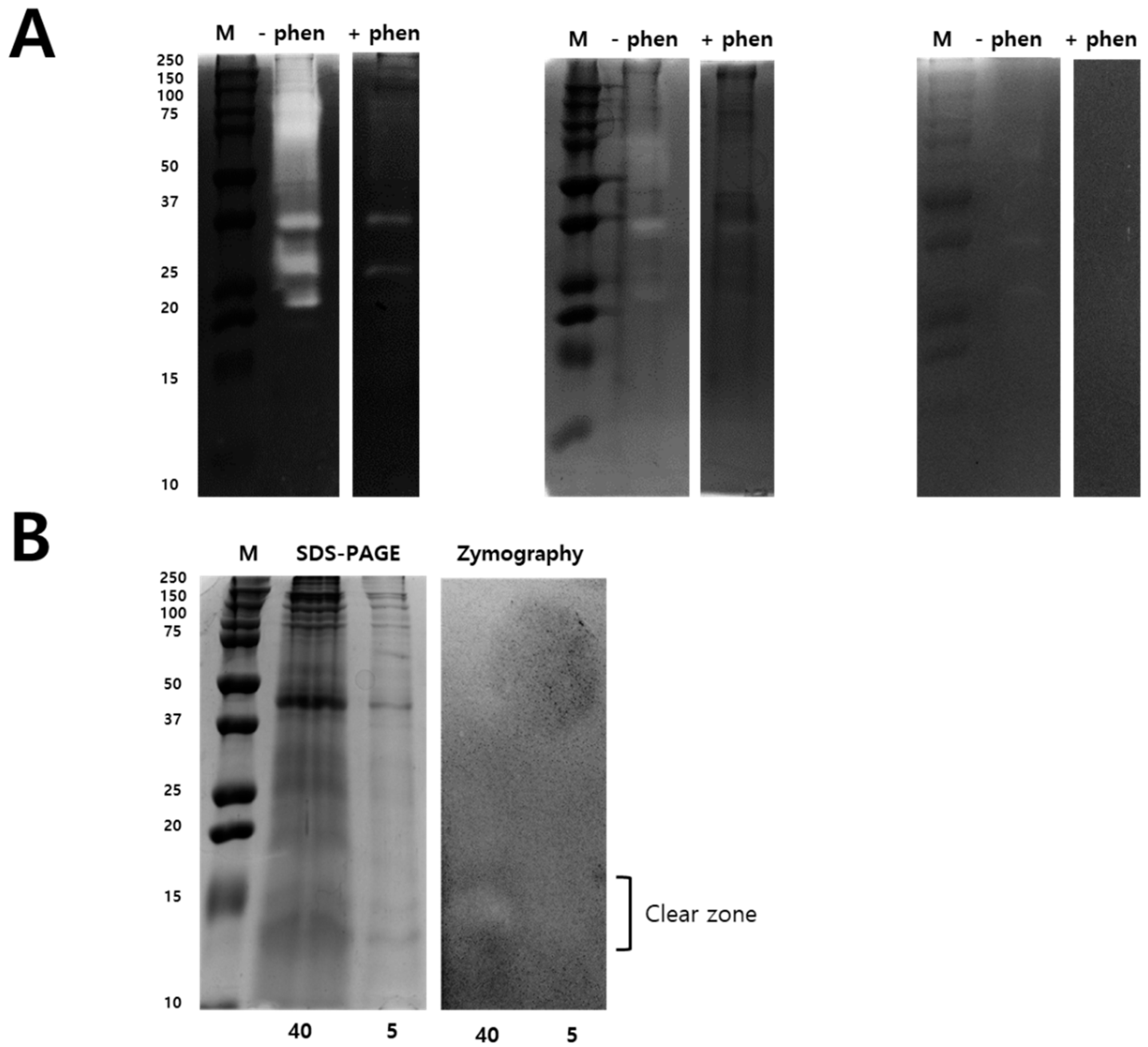
Zymography assay was performed to identify the N. nomurai jellyfish venom proteolytic activity using various types of substrates such as gelatin, casein, and fibrin, as shown in Figure 3. Interestingly, N. nomurai jellyfish venom possesses higher gelatinolytic, caseinolytic, and fibrinolytic behavior. The gelatinolytic activities of NnV showed the greatest proteolytic activity as comparision to casein and fibrin zymography and displayed different protein banding pattern between 150–25 kDa. The majority of the gelatinolytic activities were inhibited in the presence of broad-spectrum metalloproteinase inhibitor (1,10-phenanthroline), confirming that metalloproteinase-like enzymes are present in N. nomurai jellyfish venom. Furthermore, casein zymography showed strong enzymatic activity, which could be evaluated in between the range of 70–20 kDa. In fibrin zymography, weaker fibrinolytic activity was observed at 70–25 kDa. The caseinolytic and fibrinolytic activities disappeared in the presence of 1,10-phenanthroline. Despite this, we have performed 2-DE zymography under the non-reducing condition to determine the proteolytic patterns, as shown in Figure 4.
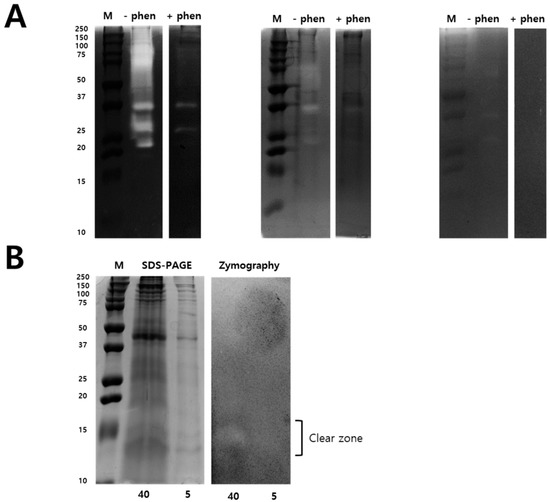
Figure 3.
Zymography identify Metalloproteinase and PLA2 activity in NnV. Different types of zymography assays were performed to determining proteolytic activity of NnV (2 mg/mL) using gelatin, casein, and fibrin as a substrate and copolymerized in non-reducing SDS-PAGE (A). Zymography assays were performed in the presence of a metalloproteinase inhibitor (1,10-phenanthroline, 10 mM) during the enzymatic reaction time. Comparison of SDS-PAGE and zymography of NnV under non-reducing conditions (B). NnV proteins (40 mg and 5 mg of total protein) were run on a 12% SDS gel and gels were Coomassie stained. M: Protein molecular size marker. Clear zones in the gel indicate regions of proteolytic activity.
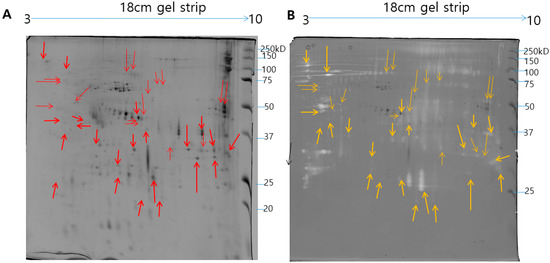
Figure 4.
Comparison of 2-DE pattern and 2-DE gelatin zymogram of N. nomurai venom. The N. nomurai venom (500 µg) separated on 18 cm IPG dry strips (pH 3–10) in the first dimension then followed by second dimension (A). For the 2-DE gelatin zymography, the gels were copolymerized with gelatin and the second dimension was performed in a 12% SDS-PAGE gels under non-reducing conditions. Proteolytic activity is understood as clear zones of lysis against the dark background (B).
2.3. Ontological Classification of Differentially Expressed Proteins
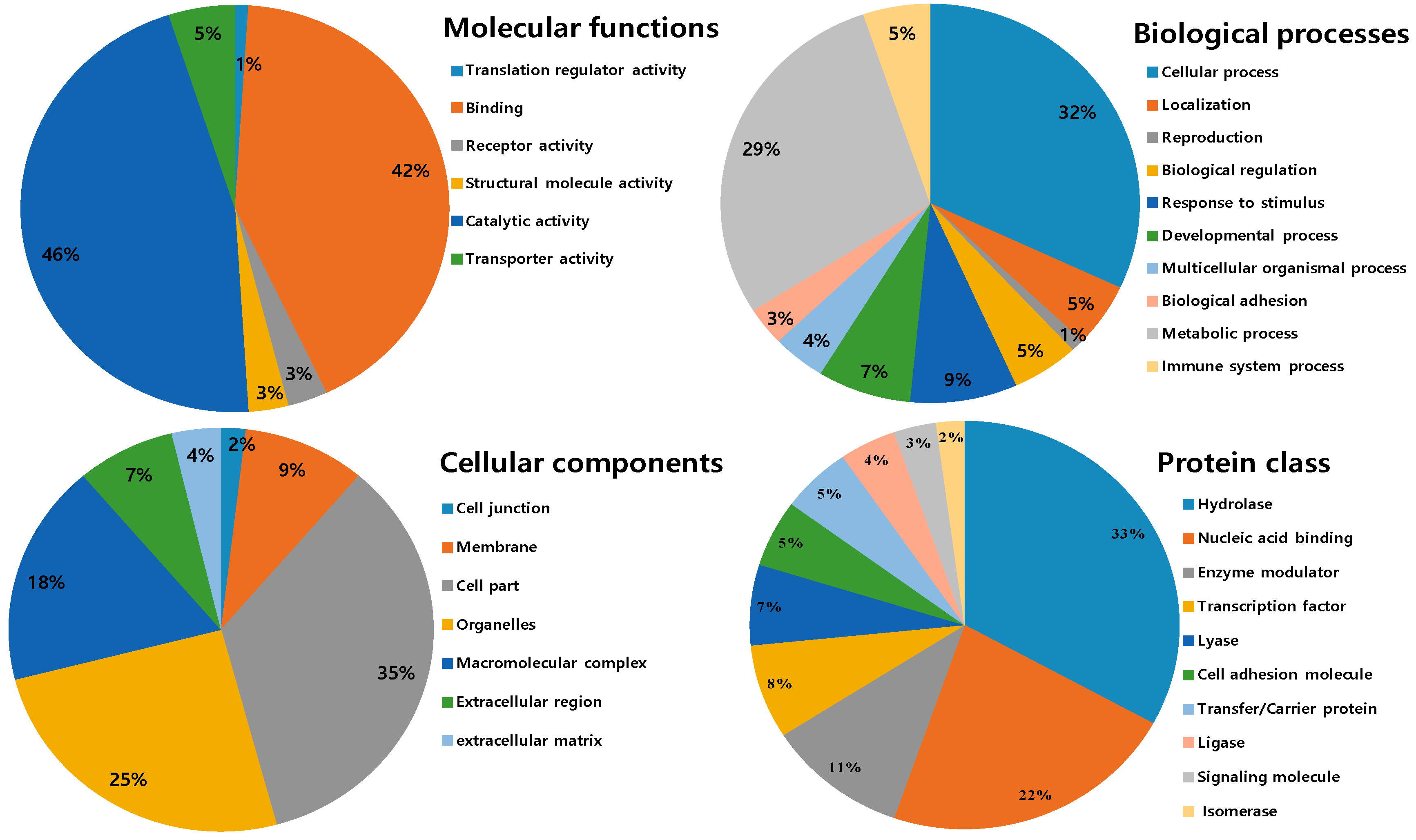
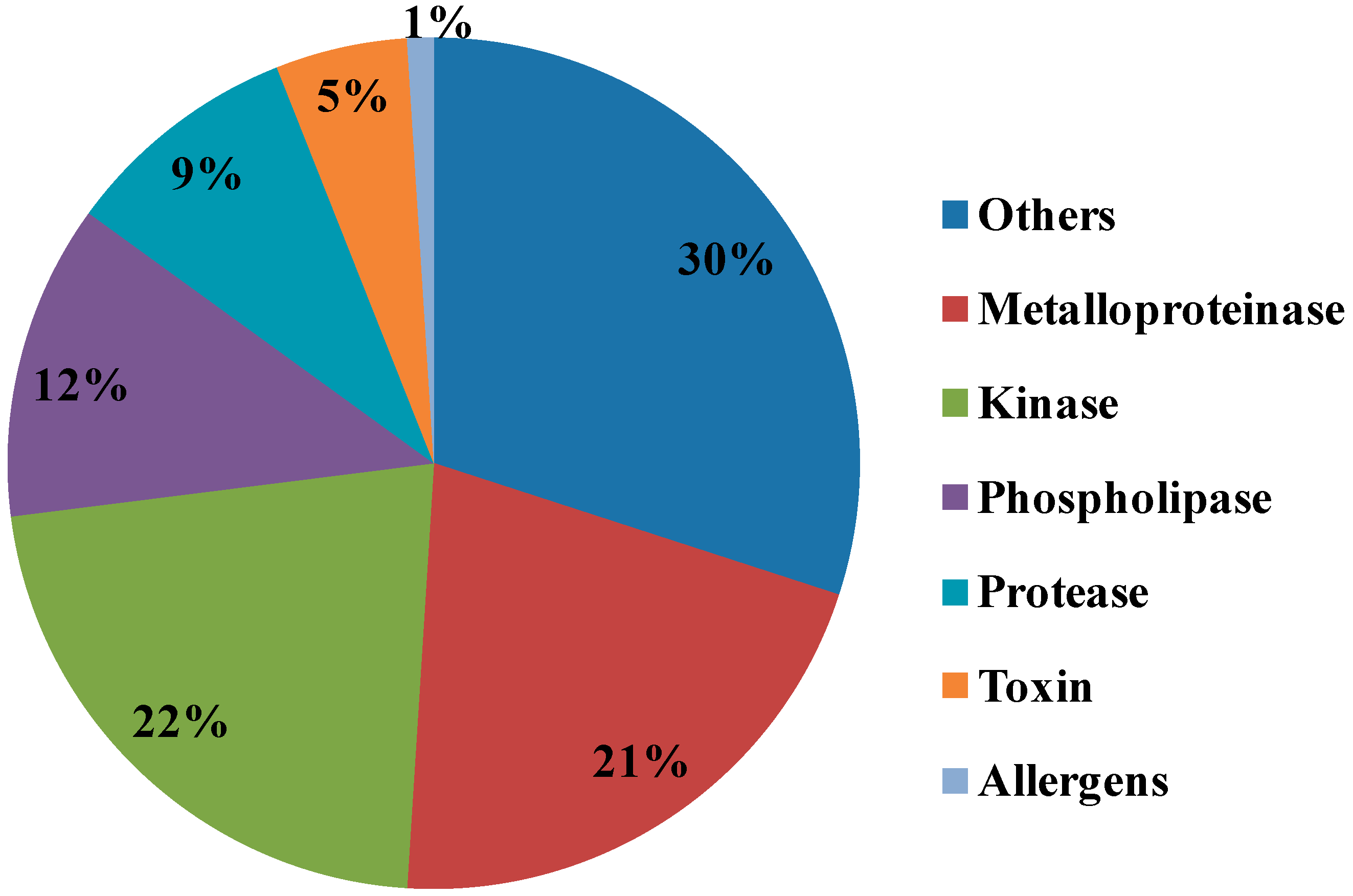
All identified proteins were subsequently classified into four ontologies according to their molecular functions, biological processes, cell components, and protein classes. In the molecular function classification, the most abundant protein category is of catalytic activity (46%) and binding (42%). Few proteins are associated with transporter activity (5%), receptor activity and structural molecular activity (3%) and translation regulator activity (1%). For the molecular function catalytic and binding activities designated the first rank, which can be related to the toxins components in NnV and hint towards strong toxicity. In the biological process category, the two significant groups include a cellular process (31%) and the metabolic process (28%). Followed by the response to stimulus (8%), developmental process (7%), biological regulation (5%), immune system process (5%), localization (5%), and multicellular organismal process (4%). The small number of proteins are assigned with biological adhesion (3%) and reproduction (1S%), as shown in Figure 5. According to cellular components, most of the proteins are assigned to the cell part (37%), followed by the organelles (20%) and macromolecular complex (19%). A minority of these proteins are localized in the membrane (10%), extracellular region (8%), extracellular matrix (4%), and (2%) cell junction. In the category of the protein class hydrolase represented the dominant class (31%). Followed by nucleic acid binding (17%), enzyme modulator (10%), transcription factor (7%), lyase (6%), cell adhesion molecule and transfer/carrier protein (5%), ligase (4%), signaling molecule (3%), and isomerase (2%), as shown in Figure 5.
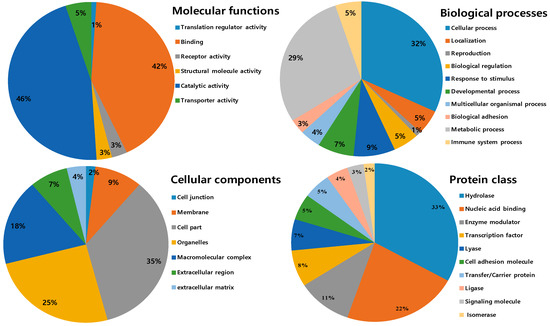
Figure 5.
Gene ontology analysis of the identified venom proteins according to their molecular functions, biological processes, cell components, and protein classes.
3. Discussion
Over the last few decades, jellyfish envenomation has turned into a universal health issue and gave rise to approximately 150 million envenomation cases yearly [19]. Jellyfish stings display a wide range of clinical symptoms of severe pain, skin inflammation, dermatitis, nausea, emesis, cardiovascular, and respiratory distress [19]. In the current study, we have explored a proteomic approach to identify NnV components by performing two-dimensional gel electrophoresis, in-gel digestion, and MALDI/TOF/MS. In this study, a total of 150 proteins were identified from the nematocysts containing NnV. These proteins include toxins that might cause severe effects after envenomation and another distinct type of proteins.
Metalloproteases are well known for hemorrhagic activities such as fibrinolysis, apoptosis, inhibit platelet aggregation, interfere with blood coagulation, cause pro-inflammatory activity and deactivate blood serine protease inhibitors [20]. Our proteomics results identified various Metalloproteinases in the NnV, matched with Zinc-metalloproteinase-disintegrin-like or snake venom metalloproteinase from Crotalus durissus snake species, A Disintegrin and Metalloproteinase with thrombospondin motifs 10, A Disintegrin and Metalloproteinase with thrombospondin motifs 23 from Mus musculus, along with ATP-dependent Zinc Metalloprotease FtsH from Oenococcus oeni. In the present study, our proteomic analysis showed that metalloproteinase are the second most abundant components of NnV, comprising 21% of venom proteome, as shown in Figure 6. Further, 1-D zymography assay (Figure 3) and 2-DE zymography assay (Figure 4) determined the proteolytic activity of NnV, which was visualized as clear white bands or spots, respectively, on a blue background.
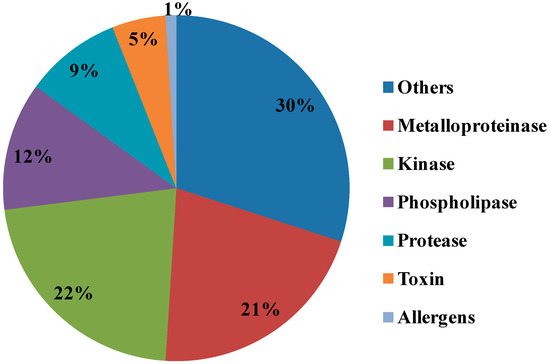
Figure 6.
Comparison of relative composition of N. nomurai venom according to protein families by a proteomics approach. The pie chart is showing relative abundance of different protein families identified by MALDI/TOF/MS in NnV.
Moreover, in our previous study, we have found that the venoms of the scyphozoan jellyfish species including Nemopilema nomurai, Cyanea nozakii, Rhopilema esculenta, and Aurelia aurita contains various Metalloproteinases [21], which shows proteolytic activity and induced cytotoxicity in NIH 3T3 cells [21]. Therefore, Metalloproteinases in the venom of jellyfish Nemopilema nomurai might be responsible for swelling, inflammation, and dermonecrosis [22]. Previously, matrix Metalloproteinases from the venom of Nemopilema nomurai were also reported to induce dermal toxicity in both the in-vivo and in-vitro animal model [22]. In future, search and application of Metalloprotease inhibitor on jellyfish venom will be beneficial in the management of jellyfish envenomation and development of novel therapeutic source against venom toxicity.
Several phospholipases were also identified in NnV in current research, such as Phospholipase D LiSicTox-betaIDI, also known as LiRecDT5 homologous to PLA2 from Loxosceles intermedia spider venom, 85/88 kDa Calcium-independent Phospholipase A2 and Phospholipase A-2-activating protein from Rattus norvegicus, Calcium-independent Phospholipase A2-gamma from Mus musculus. Phospholipase D LiSicTox-betaIDI is the novel member of loxosceles dermonecrotic toxin family, which has diverse biological activities such as dermonecrosis, increased vessel permeability, platelet aggregation, induce an inflammatory response, and cause mortality in animal models [23,24]. The presence of phospholipases in NnV can be corelated to pathogenesis, hemolysis, and other harmful consequences, for example, burning sensation and an erythematous eruption with small vesicles after proper provocation. Our proteomic analysis revealed that phospholipases is another abundant toxin protein constituting 12% in whole venom proteome (Figure 6).
Proteases constituted a vital portion of NnV counting 5% of the venom proteome (Figure 6). Serine proteases are well studied in various venom sources, typically in the snake, cone snail, scorpion, and spider venoms [25,26,27,28]. Serine proteases were also reported in the marine animals such as sea snake, stingrays, sponge and jellyfish [29,30,31]. Serine proteases mainly cause haemotoxicity by affecting the blood coagulation system and triggers fibrinolysis, platelet aggregation, and edema [32]. In this study, we have identified several serine proteases matched with snake venom serine protease Nikobin from Vipera nikolskii, serine protease HtrA-like from Staphylococcus aureus (strain NCTC 8325), Pro-apoptotic serine protease nma111 from Neurospora crassa (strain ATCC 24698/74-OR23-1A/CBS 708.71/DSM 1257/FGSC 987). Snake venom Serine Protease Nikobin can acts on the hemostasis system of the prey by affecting several physiological processes that include blood coagulation, fibrinolysis, and blood pressure [32,33]. Accordingly, serine proteases obtained from NnV might be responsible for pathogenesis during NnV envenomation.
Putative Kunitz-type Serine protease inhibitor is efficiently characterized in well-known venomous animals, for example snakes, spiders, scorpions, centipedes, cone snails, and sea anemone [34,35,36,37,38,39]. Kunitz-type Serine protease inhibitor plays a crucial role in various physiological activities namely blood coagulation, fibrinolysis, inflammation and ion channel blocking [37,38,39,40]. Kunitz-type Serine protease inhibitor had inhibitory activity against trypsin, chymotrypsin, and demonstrated its antifibrinolytic activity in snake venom [39,40,41]. Additionally, recent studies revealed that spider Kunitz-type Serine protease inhibitor exhibits inhibitory activity against trypsin, chymotrypsin, plasmin, and neutrophil elastase [42]. In the present study, our proteomic data showed that NnV was also comprised of such a Putative Kunitz-type Serine protease inhibitor, which might contribute to NnV toxicity.
Other toxin protein types present in NnV are homologous to the three-finger toxin MALT0044C, hemolysins, leukotoxins, major pollen allergen Lol p 5a, Allergen Mag, venom prothrombin activator trocarin D, RTX-III toxin determinant A from serotype 2, and Shiga-like toxin 1 subunit A. Three-finger toxin MALT0044C belongs to the snake three finger toxin family, it has a predominant role in neurotoxicity by inducing peripheral paralysis and finally leads to respiratory arrest and death [41]. The proteins in the 3FTx family have high functional diversity; they act as neurotoxins, which can target the cholinergic system and also block the L-type Ca2+ channels [42]. Therefore, three-finger toxin in NnV can also result in lethal health outcomes.
Various types of Hemolysins were reported in the venom of different kinds of jellyfish, including Ryncolin-2, Ryncolin-3, Neoverrucotoxin subunit beta, and Veficolin-1 were identified in the venom of jellyfish S. meleagris [43]. Moreover, CfTX-1 and CfTX-2 are potent Haemolysins were identified in C. fleckeri venom, which share sequence homology to toxins related to 4 cubozoan jellyfish species [44]. Other hemolytic proteins include CaTX-1 from C. alata, CrTX-1 from C. rastoni, and CcTX-1 from C. capillata were also reported in previous studies [45,46,47,48,49]. Former research indicated that all these hemolytic proteins are responsible for the hemolytic action and induce inflammation, pain, dermonecrosis, and death in animal models [45,46,47,48,49]. Presence of Hemolysins in NnV supports its previously reported evenomation symptoms like inflammation, pain, and dermonecrosis.
Hemolysin transporter protein was also identified in NnV, which is homologous to ShlB of Serratia marcescens. Hemolysin triggers the release of the inflammatory mediators, increase vascular permeability, cause edema formation, granulocyte accumulation, and finally contribute pathogenicity of Serratia species [50,51]. Therefore, it can be inferred that the Hemolysin present in NnV might impair hemostasis, and cause edema and hypotension after envenomation.
Pasteurella Leukotoxins are exotoxins that attack host leukocytes, chiefly polymorphonuclear cells, by inducing cell rupture [52]. The Leukotoxin binds to the host LFA-1 integrin and triggers a signaling cascade cause following biological effects such as tyrosine phosphorylation of the CD18 tail, increasing the intracellular Ca2+, finally results in host cell rupture [52]. We also identified Leukotoxins in the proteomics analysis of NnV homolog to Pasteurella Haemo-lytica-like sp. (strain 5943b), which may play a significant role in NnV envenomation and cause harmful consequences.
Two types of allergens were also detected in the NnV proteomics known as Major pollen allergen Lol p 5a from Lolium perenne and Allergen Mag (fragment) from Dermatophagoides farina. Lol p 5a is a major allergen of rye-grass pollen (Lolium perenne pollen), which mediate type I hypersensitivity, cause an allergic reaction in humans, hay fever, and triggers allergic asthma. House dust fly (Dermatophagoides farina) is the one of the major agent causing allergic ailment by way of bronchial, asthma, atopic dermatitis and rhinitis [53,54]. Allergen Mag is the major allergen found in Dermatophagoides farina, which binds to IgE and causes an allergic reaction in humans [55,56,57]. It releases histamine from washed blood cells of the mite-allergic patients. Hence, the allergens identified in NnV may be related to symptoms like dermatitis, utricularia, pruritus, and fever caused by jellyfish envenoming.
Venom prothrombin activator trocarin D is the snake prothrombin activator from Tropidechis carinatus, acts as a toxin component of the snake venom and disturbs the hemostatic system of prey [58]. It was reported that Venom prothrombin activator trocarin D induces cyanosis and death in mice at 1mg/kg body weight and display potent procoagulant effects [58]. Fortunately, we also identified the venom prothrombin activator trocarin D from NnV matched with that of Tropidechis carinatus. Therefore, venom prothrombin activator trocarin D is one of the important toxic protein in NnV which might cause severe harmful consequences such as shock, cardiovascular instability, and sudden death. RTX-III toxin determinant A from serotype 2 and RTX-I toxin determinant A from serotypes 1/9 are an essential virulence factor for A. pleuropneumoniae and cause swine pleuropneumonia [59].
RTX-III PROTEIN are toxic to porcine lung macrophages and erythrocytes, and it possesses strong hemolytic activity and causes cytotoxicity in alveolar macrophages and neutrophils hence play a significant role in pathogenesis [60]. Noteworthy, RTX-III toxin determinant A from serotype 2 was also found in NnV venom, and therefore, RTX-III toxin may account for hemolytic activity and cytotoxicity in jellyfish victims.
It was reported earlier that a Shiga-like toxin subunit A negatively regulates the translation process by catalytic inactivation of 60S ribosomal subunit and responsible for inhibiting protein synthesis [61]. The shiga-like toxin (SLT)-producing E. coli is associated with diarrhea, hemorrhagic colitis, and hemolytic uremic syndrome [61,62,63]. We also identified Shiga- like toxin subunit A in the proteomic analysis. So Shiga- like toxin subunit A could be responsible for cramps, abdominal pain, and other hemorrhagic activities after NnV stings.
Surprisingly, along with toxic proteins, we have also recognized many nontoxic proteins during NnV profiling, which plays an important role in signaling and other metabolic processes. Kinases itself constitute the major part of NnV, they hold 22% of known NnV proteins. Other proteins are 30% of total proteins found in NnV. OTU domain-containing protein 7B is one of the nontoxic proteins found in NnV. OTU domain-containing protein 7B is the negative regulator of the non-canonical NF-kappa-B pathway and displays the anticancer effect by suppressing the NF-kappa-B pathway in HCC cells [64]. OTU domain-containing protein 7B act as an anticancer target in liver cancer cells [64,65]. Hence NnV can exert an anticancer effect, and extensive studies need to investigate the therapeutic potential of NnV. Cell death abnormality protein 1, related to Caenorhabditis elegans, was also identified in NnV, and Cell death abnormality protein 1 necessitates the engulfment of cells undergoing programmed cell death [66,67]. It activates the expression of unfolded protein response genes and has a defense response to bacteria [68]. However, cell death abnormality protein 1 might play an essential role in nematocyst genesis and developments.
4. Conclusions
Jellyfish Nemopilema nomurai is one of the largest jellyfish and its evenomation can lead to harmful consequences, even death. Till now the identification and functional characterization of NnV components have been poorly described and remain beyond our knowledge. This is the first report that is able to identify around 150 proteins in Nemopilema nomurai jellyfish venom, which include metalloproteinases, kinases, phospholipases, proteases, toxins, and allergens. Some of these components are considered to play an important role in the cardiovascular, hepatotoxic, hematological, cytotoxic, and allergenic effects of the venom. These findings are beneficial in recognizing the mechanism of jellyfish envenomation and provide valuable means to establish new methods or treatments to handle jellyfish stings. The overall comprehensive characterization of NnV, provide a promising approach to explore the rich source of bioactive toxin components as therapeutic agents in the future.
5. Materials and Methods
5.1. Chemicals and Reagents
Immobiline™ Drystrip (pH 4–7, 18 cm), dithiothreitol (DTT), and iodoacetamide were bought from GE Healthcare life sciences (Marlborough, MA, USA). Acetonitrile (ACN), trifluoroacetic acid was purchased from Merck Chemicals, Darmstadt, Germany. Sequencing grade modified trypsin was purchased from Promega Corporation, Madison, WI, USA, formic acid from Acros Organics BVBA, Geel, Belgium. Other analytical grade chemicals and reagents were used and purchased from Sigma Aldrich Corporation (St. Louis, MO, USA).
5.2. Sample Collection and Preparation
N. nomurai jellyfish specimens were collected from the Yellow Sea near the coast of Gunsan, South Korea. After taking out of water tentacles were dissected and transferred immediately in ice to the laboratory for further processing. Isolation of nematocysts was done using the previously described method [69]. In short, dissected tentacles were rinsed with cold seawater to eliminate any debris, and then 3 volumes (v/v) of cold sea water were added and at 4 °C put on shaker for one day. After that, the tentacle-free seawater was harvested and centrifuged at 1000× g for 5 min; then the nematocyst-rich pellet was washed thrice with fresh sea water. The remaining sediment tentacles were later autolyzed in fresh seawater at 4 °C for one day, as described above, and the autolysis process was repeated for 3–4 days. At last, the nematocysts that settled down were collected and seawater was used to wash them several times. Later, the nematocysts were centifuged at 500× g for 5 min, and pellets (nematocysts) were lyophilized and stored at −20 °C until further use.
5.3. Venom Extraction
The technique described by Carrette and Seymour was used to extract venom from the freeze-dried nematocysts [69] with slight modification. In short, venom was extracted from 50 mg of nematocyst using glass beads (approximately 8000 beads; 0.5 mm in diameter), and 1 mL of ice-cold phosphate buffered saline (PBS, pH 7.4). Further, samples were kept shaking on the mini bead mill at 3000 rpm for 30 s, the above step was repeated ten times with intermittent cooling on ice. The venom extracts were then transferred to a new Eppendorf tube and centrifuged (22,000 g) at 4 °C for 30 min. The above supernatant was used as NnV for the present study. Bradford assay (Bio-Rad, Hercules, CA, USA) [70] was used to determine the protein concentration of the venom. Finally, the venom was utilized in research, based on its protein concentration.
5.4. Two-Dimensional Gel Electrophoresis under Reducing Conditions and Image Analysis
For two-dimensional gel electrophoresis, 500 µg of protein samples were resuspended in 2-DE sample buffer (7 M urea, 2 M thiourea, 4% (w/v) CHAPS, 10 mg/mL DTT, 1% pharmalytes 3–10 and few grains of bromophenol blue). Then, protein samples were applied to immobiline TM Dry strip 18 cm, pH gradient (IPG) strips (pH 3–10) and passively rehydrated overnight at room temperature. The isoelectric focusing (IEF) was performed at a constant temperature of 20 °C using Ettan IPGphor system (GE Healthcare), with the following procedure: 50 V for 1:00 h, 200 V for 1:00 h, 500 V for 0:30 h, gradient 4000 V for 0:30 h, 4000 V for 1:00 h, gradient 10,000 V for 1:00 h, 10,000 V for 13:00 h, and 50 V for 3:00 h. Before the second dimension, the focused strips were firstly reduced with equilibration buffer (50 mM Tris-HCl (pH 8.8), 6 M urea, 30% glycerol, 2% SDS and 0.01% bromophenol blue containing 1% w/v DTT) for 15 min. For the second time, the gel strips were washed with the same equilibration buffer containing 2.5% w/v iodoacetamide for 15 min. The focused and equilibrated strips were inserted on top of the 12% SDS-PAGE and sealed with 0.5% w/v agarose gel. After that, electrophoresis was performed in a PROTEAN II xi cell gel electrophoresis unit. The gels were run at 10 mA/gel for 15 min for the initial migration and increased to 20 mA/gel at 20 °C for separation until the dye front reached the bottom of the gels. Silver staining was performed to visualize the protein spots as similar to the method reported by Mortz et al. [71], and this experiment was repeated thrice. The 2-DE gel images were digitalized by using Epson perfection V 700 photo scanner (Epson, Long Beach, CA, USA), and the acquired images were analyzed using Progenesis Same Spots software (Nonlinear Dynamics, Newcastle, UK).
5.5. In-gel Digestion
In-gel digestion of the proteins was performed by the method described previously [72]. The protein spots of interest were manually excised from preparative gels to perform in-gel digestion. In brief, the gel pieces were washed with pure water for several times and then destained with 30 mM potassium ferricyanide and 100 mM Na2S2O3 (50%/50% v/v) for 10 min with shaking. Further, the gel pieces were dehydrated for 10 min by incubating in 100 µl of 100% acetonitrile and were then dried in a lypholizer equipment for 15 min. The reduction step was performed by adding 50 µL reduction solution (10 mM DTT in 100 mM ammonium bicarbonate) to the dried gel pieces and incubated for 45 min at room temperature. After that, the gel pieces were alkylated with 100 mM iodoacetamide in the dark for 45 min at room temperature. The gel pieces were dried, incubated in trypsin (Promega, Madison, WI, USA) at a final concentration of 2 ng/µL in 10 µL of 50 mM NH4HCO3 and incubated on ice for 45 min. The excessive liquid was removed, and the proteolysis of protein was performed by adding the exact amount of 50 mM NH4HCO3. After overnight digestion with trypsin at 37 °C, the tryptic peptide mixture was pooled with extraction buffer containing 100% acetonitrile and 50% trifluoroacetic acid and concentrated in a speed vacuum.
5.6. MALDI/TOF/MS Analysis and Database Searching
The peptide samples were mixed with 1 µL of HCCAs matrix solution (α-acyano-4-hydroxycinnamic acid) and 1 µL of extraction buffer. 1 µL peptide mixture was spotted onto a freshly cleaned MALDI/TOF plate and dried at room temperature. Mass spectra were measured by using Voyager-DE STR mass spectrometer (Applied Biosystems, Franklin Lakes, NJ, USA).
Reflection/delayed extraction mode acquired the spectra. Monoisotopic peptide masses were chosen over a mass range of 800–3000 Da. Peptide mass fingerprinting (PMF) was performed to identify proteins using the MS-Fit program (http://prospector.ucsf.edu), and Mascot (Matrix science http://www.matrixscience.com) in the Swiss-Prot databases. For the peptide search following parameters were considered carbamidomethylation of cysteines as a fixed modification, oxidation of methionine as a variable modification, peptide mass tolerance of ±30 ppm for the fragment ions, trypsin with one missed cleavage was allowed. Criteria used for protein identification includes a number of the matched peptide, the extent of sequence coverage and probability based Mowse score was considered before accepting the identification.
5.7. Proteolytic Activity Assay
The proteolytic zymography assay was performed by using different types of substrates such as fibrin, gelatin, and casein, as the method described by References [73,74]. Respective zymography gels were prepared by copolymerized gelatin (2 mg/mL), casein (2 mg/mL), or fibrinogen (0.5 mg/mL)–thrombin (0.01 unit/mL), dissolved in 20 mM sodium phosphate buffer (pH 7.4) with 12% polyacrylamide. The protein samples were dissolved in non reducing sample buffer and the zymography gels were run under cold condition (4 °C) at 15 mA/gel. When the electrophoresis was over, the gels were washed with 2.5% Triton X-100 for 20 min to remove the SDS. After washing the gels were incubated with zymography reaction buffer (20 mM Tris (pH 7.4), 0.5 mM calcium chloride) at 37 °C overnight and stained with 0.125% coomassie blue. To study the protease inhibitor effect 1,10-phenanthroline was prepared as a 300 mM stock solution in methanol and added fresh to each incubation buffer to give a final concentration of 10 mM.
5.8. Modified Zymography Assays to Identify PLA2 Activity in NnV
NnV was run under non-reducing conditions on 12% SDS-PAGE and stained with Coomassie Brilliant Blue R-250. Blood and egg yolk zymography method [75] with slight modifications was to determine the molecular weight of PLA2. Briefly, to eradicate the SDS from the gel, the gel was washed for 1 h with 500 mM Tris-HCl (pH 7.4) containing 2% Triton X-100, and then for 1 h with 100 mM Tris-HCl (pH 7.4) along with 1% Triton X-100. Further gels were washed with 50 mM Tris-HCl (pH 7.4), 140 mM NaCl, and 2.5 mM CaCl2. For zymography on blood and egg-yolk, the washed SDS-PAGE gel was placed directly on a 1% agarose gel (50 mM Tris-HCl (pH 7.4), 140 mM NaCl, 2.5 mM CaCl2, 2.4% human erythrocytes, and 2% egg yolk. After overnight incubation at 37 °C, the opaque zone indicated the presence of PLA2 [76].
5.9. Two-Dimensional Gel Electrophoresis under Non-Reducing Conditions and 2-DE Gelatin Zymography
The NnV samples were prepared under non reducing conditions for 2-DE and gelatin 2-DE zymography. The 500 µg of protein samples were dissolved in 7 M urea, 2 M thiourea, 4% (w/v) CHAPS, 1% pharmalytes 3–10 and few grains of bromophenol blue) without DTT and the strips were rehydrated overnight, and the isoelectric focusing was performed as the method described above. After IEF the focused strips were washed with equilibration buffer (50 mM Tris-HCl (pH 8.8), 6 M urea, 30% glycerol, 2% SDS, and 0.01% bromophenol blue) for 15 min followed by electrophoresis. For 2-DE gelatin zymography 12% SDS-polyacrylamide gels were copolymerized with gelatin (1 mg/mL), and electrophoresis was performed at 4 °C. The gels were washed twice after electrophoresis for 30 min in 2.5% Triton X-100 to remove SDS. Triton X-100 traces were removed from the gels by washing with deionized water. After that, the gels were incubated in 20 mM Tris (pH 7.4), 0.5 mM calcium chloride at 37 °C for 16 h and stained with 0.125% coomassie blue. Clear zones of hydrolysis in the gel against blue background showed gelatin proteolytic activity.
5.10. Gene Ontology Analysis of the Identified Venom Proteins
The proteomic data was analyzed using the Panther classification system (http:www.pantherdb.org/) to classify regarding molecular function, biological process, protein class, and cellular components [77].
Supplementary Materials
The following are available online at https://www.mdpi.com/2072-6651/11/3/153/s1, Table S1: Nnv proteomics.
Author Contributions
E.K. conceived and designed the study. I.C. performed the experiments. D.H.H., H.L. and C.K. analyzed and interpreted the data. S.Y., J.C. and C.H.H. and W.D.Y. advised jellyfish sampling and species identification. I.C. wrote the manuscript. All authors have read and approved the final manuscript.
Funding
This work was supported by grants from the National Research Foundation of Korea (NRF-2017R1D1A1B03036005) from the Korean government. This research was also a part of the project titled “Development of Biomedical materials based on marine proteins”, funded by the Ministry of Oceans and Fisheries, Korea and “Management of Marine Organisms causing Ecological Disturbance and Harmful Effects” funded by the Korea Institute of Marine Science and Technology Promotion (KIMST)/Ministry of Oceans and Fisheries (MOF).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Uye, S.-I. Blooms of the giant jellyfish Nemopilema nomurai: A threat to the fisheries sustainability of the East Asian Marginal Seas. Plankton Benthos Res. 2008, 3, 125–131. [Google Scholar] [CrossRef]
- Dong, Z.; Liu, D.; Keesing, J.K. Jellyfish blooms in China: Dominant species, causes and consequences. Mar. Pollut. Bull. 2010, 60, 954–963. [Google Scholar] [CrossRef] [PubMed]
- Tibballs, J. Australian venomous jellyfish, envenomation syndromes, toxins and therapy. Toxicon 2006, 48, 830–859. [Google Scholar] [CrossRef] [PubMed]
- Burnett, J.W.; Fenner, P.J.; Rifkin, J.F. Venomous and Poisonous Marine Animals: A Medical and Biological Handbook; UNSW Press: Sydney, Australia, 1996. [Google Scholar]
- Kim, E.; Lee, S.; Kim, J.-S.; Yoon, W.D.; Lim, D.; Hart, A.J.; Hodgson, W.C. Cardiovascular effects of Nemopilema nomurai (Scyphozoa: Rhizostomeae) jellyfish venom in rats. Toxicol. Lett. 2006, 167, 205–211. [Google Scholar] [CrossRef] [PubMed]
- Choudhary, I.; Lee, H.; Pyo, M.-J.; Heo, Y.; Bae, S.K.; Kwon, Y.C.; Yoon, W.D.; Kang, C.; Kim, E. Proteomics approach to examine the cardiotoxic effects of Nemopilema nomurai jellyfish venom. J. Proteom. 2015, 128, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Kang, C.; Munawir, A.; Cha, M.; Sohn, E.-T.; Lee, H.; Kim, J.-S.; Yoon, W.D.; Lim, D.; Kim, E. Cytotoxicity and hemolytic activity of jellyfish Nemopilema nomurai (Scyphozoa: Rhizostomeae) venom. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2009, 150, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Choudhary, I.; Lee, H.; Pyo, M.-J.; Heo, Y.; Bae, S.K.; Kwon, Y.C.; Yoon, W.D.; Kang, C.; Kim, E. Nemopilema nomurai jellyfish venom treatment leads to alterations in rat cardiomyocytes proteome. Data Brief 2015, 5, 884–887. [Google Scholar] [CrossRef] [PubMed]
- Kang, C.; Kim, Y.K.; Lee, H.; Cha, M.; Sohn, E.-T.; Jung, E.-S.; Song, C.; Kim, M.; Lee, H.C.; Kim, J.-S. Target organ identification of jellyfish envenomation using systemic and integrative analyses in anesthetized dogs. J. Pharmacol. Toxicol. Methods 2011, 64, 173–179. [Google Scholar] [CrossRef] [PubMed]
- Kawahara, M.; Uye, S.; Burnett, J.; Mianzan, H. Stings of edible jellyfish (Rhopilema hispidum, Rhopilema esculentum and Nemopilema nomurai) in Japanese waters. Toxicon 2006, 48, 713–716. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira, S.A.M.; Magalhães, M.R.; de Oliveira, L.P.; da Cunha, L.C. Identification of antinociceptive fraction of snake venom from Crotalus durissus collilineatus crotamine-negative and its acute toxicity evaluation. Toxicon 2016, 122, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, F.C.; Dekan, Z.; Rosengren, K.J.; Erickson, A.; Vetter, I.; Deuis, J.; Herzig, V.; Alewood, P.; King, G.F.; Lewis, R.J. Identification and characterization of ProTx-III [μ-TRTX-Tp1a], a new voltage-gated sodium channel inhibitor from venom of the tarantula Thrixopelma pruriens. Mol. Pharmacol. 2015, 88, 291–303. [Google Scholar] [CrossRef] [PubMed]
- Lima, P.C.; Bordon, K.C.; Pucca, M.B.; Cerni, F.A.; Zoccal, K.F.; Faccioli, L.H.; Arantes, E.C. Partial purification and functional characterization of Ts19 Frag-I, a novel toxin from Tityus serrulatus scorpion venom. J. Venom. Anim. Toxins Incl. Trop. Dis. 2015, 21, 49. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, B.; Le Caer, J.-P.; Aráoz, R.; Thai, R.; Lamthanh, H.; Benoit, E.; Molgó, J. Isolation, purification and functional characterization of α-BnIA from Conus bandanus venom. Toxicon 2014, 91, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Yu, H.; Yue, Y.; Liu, S.; Xing, R.; Chen, X.; Li, P. Combined proteomics and transcriptomics identifies sting-related toxins of jellyfish Cyanea nozakii. J. Proteom. 2016, 148, 57–64. [Google Scholar] [CrossRef] [PubMed]
- Brinkman, D.L.; Jia, X.; Potriquet, J.; Kumar, D.; Dash, D.; Kvaskoff, D.; Mulvenna, J. Transcriptome and venom proteome of the box jellyfish Chironex fleckeri. BMC Genom. 2015, 16, 407. [Google Scholar] [CrossRef] [PubMed]
- Weston, A.J.; Chung, R.; Dunlap, W.C.; Morandini, A.C.; Marques, A.C.; Moura-da-Silva, A.M.; Ward, M.; Padilla, G.; da Silva, L.F.; Andreakis, N. Proteomic characterisation of toxins isolated from nematocysts of the South Atlantic jellyfish Olindias sambaquiensis. Toxicon 2013, 71, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Bloom, D.A.; Burnett, J.W.; Alderslade, P. Partial purification of box jellyfish (Chironex fleckeri) nematocyst venom isolated at the beachside. Toxicon 1998, 36, 1075–1085. [Google Scholar] [CrossRef]
- Nomura, J.T.; Sato, R.L.; Ahern, R.M.; Snow, J.L.; Kuwaye, T.T.; Yamamoto, L.G. A randomized paired comparison trial of cutaneous treatments for acute jellyfish (Carybdea alata) stings. Am. J. Emerg. Med. 2002, 20, 624–626. [Google Scholar] [CrossRef] [PubMed]
- Kini, R.; Koh, C. Metalloproteases affecting blood coagulation, fibrinolysis and platelet aggregation from snake venoms: Definition and nomenclature of interaction sites. Toxins 2016, 8, 284. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Jung, E.-S.; Kang, C.; Yoon, W.D.; Kim, J.-S.; Kim, E. Scyphozoan jellyfish venom metalloproteinases and their role in the cytotoxicity. Toxicon 2011, 58, 277–284. [Google Scholar] [CrossRef] [PubMed]
- Kang, C.; Jin, Y.B.; Kwak, J.; Jung, H.; Yoon, W.D.; Yoon, T.-J.; Kim, J.-S.; Kim, E. protective effect of tetracycline against dermal toxicity induced by Jellyfish venom. PLoS ONE 2013, 8, e57658. [Google Scholar] [CrossRef] [PubMed]
- da Silveira, R.B.; Pigozzo, R.B.; Chaim, O.M.; Appel, M.H.; Silva, D.T.; Dreyfuss, J.L.; Toma, L.; Dietrich, C.P.; Nader, H.B.; Veiga, S.S. Two novel dermonecrotic toxins LiRecDT4 and LiRecDT5 from brown spider (Loxosceles intermedia) venom: From cloning to functional characterization. Biochimie 2007, 89, 289–300. [Google Scholar] [CrossRef] [PubMed]
- Chaves-Moreira, D.; Souza, F.N.; Fogaça, R.T.; Mangili, O.C.; Gremski, W.; Senff-Ribeiro, A.; Chaim, O.M.; Veiga, S.S. The relationship between calcium and the metabolism of plasma membrane phospholipids in hemolysis induced by brown spider venom phospholipase-D toxin. J. Cell. Biochem. 2011, 112, 2529–2540. [Google Scholar] [CrossRef] [PubMed]
- Ghorbanpur, M.; Zare Mirakabadi, A.; Zokaee, F.; Zolfagarrian, H.; Rabiei, H. Purification and partial characterization of a coagulant serine protease from the venom of the Iranian snake Agkistrodon halys. J. Venom. Anim. Toxins Incl. Trop. Dis. 2009, 15, 411–423. [Google Scholar] [CrossRef]
- Möller, C.; Vanderweit, N.; Bubis, J.; Marí, F. Comparative analysis of proteases in the injected and dissected venom of cone snail species. Toxicon 2013, 65, 59–67. [Google Scholar] [CrossRef] [PubMed]
- Veiga, S.S.; da Silveira, R.B.; Dreyfuss, J.L.; Haoach, J.; Pereira, A.M.; Mangili, O.C.; Gremski, W. Identification of high molecular weight serine-proteases in Loxosceles intermedia (brown spider) venom. Toxicon 2000, 38, 825–839. [Google Scholar] [CrossRef]
- Ma, Y.; Zhao, R.; He, Y.; Li, S.; Liu, J.; Wu, Y.; Cao, Z.; Li, W. Transcriptome analysis of the venom gland of the scorpion Scorpiops jendeki: Implication for the evolution of the scorpion venom arsenal. BMC Genom. 2009, 10, 290. [Google Scholar] [CrossRef] [PubMed]
- Rojas, A.; Doolittle, R.F. The occurrence of type S1A serine proteases in sponge and jellyfish. J. Mol. Evol. 2002, 55, 790–794. [Google Scholar] [CrossRef]
- He, J.; Chen, S.; Gu, J. Identification and characterization of Harobin, a novel fibrino (geno) lytic serine protease from a sea snake (Lapemis hardwickii). FEBS Lett. 2007, 581, 2965–2973. [Google Scholar] [CrossRef] [PubMed]
- Zaqueo, K.D.; Kayano, A.M.; Simões-Silva, R.; Moreira-Dill, L.S.; Fernandes, C.F.; Fuly, A.L.; Maltarollo, V.G.; Honório, K.M.; da Silva, S.L.; Acosta, G. Isolation and biochemical characterization of a new thrombin-like serine protease from Bothrops pirajai snake venom. BioMed Res. Int. 2014, 2014, 595186. [Google Scholar] [CrossRef] [PubMed]
- Markland, F.S. Snake venoms and the hemostatic system. Toxicon 1998, 36, 1749–1800. [Google Scholar] [CrossRef]
- Tsai, I.-H.; Wang, Y.-M.; Cheng, A.C.; Starkov, V.; Osipov, A.; Nikitin, I.; Makarova, Y.; Ziganshin, R.; Utkin, Y. cDNA cloning, structural, and functional analyses of venom phospholipases A2 and a Kunitz-type protease inhibitor from steppe viper Vipera ursinii renardi. Toxicon 2011, 57, 332–341. [Google Scholar] [CrossRef] [PubMed]
- Fahrenkrog, B. Nma111p, the Pro-Apoptotic HtrA-Like Nuclear Serine Protease in Saccharomyces Cerevisiae: A Short Survey; In Portland Press Limited: London, UK, 2011. [Google Scholar]
- Lu, J.; Yang, H.; Yu, H.; Gao, W.; Lai, R.; Liu, J.; Liang, X. A novel serine protease inhibitor from Bungarus fasciatus venom. Peptides 2008, 29, 369–374. [Google Scholar] [CrossRef] [PubMed]
- Yuan, C.-H.; He, Q.-Y.; Peng, K.; Diao, J.-B.; Jiang, L.-P.; Tang, X.; Liang, S.-P. Discovery of a distinct superfamily of Kunitz-type toxin (KTT) from tarantulas. PLoS ONE 2008, 3, e3414. [Google Scholar] [CrossRef]
- Zhao, R.; Dai, H.; Qiu, S.; Li, T.; He, Y.; Ma, Y.; Chen, Z.; Wu, Y.; Li, W.; Cao, Z. SdPI, the first functionally characterized Kunitz-type trypsin inhibitor from scorpion venom. PLoS ONE 2011, 6, e27548. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Liu, Z.; Xiao, Y.; Li, Y.; Rong, M.; Liang, S.; Zhang, Z.; Yu, H.; King, G.F.; Lai, R. Chemical punch packed in venoms makes centipedes excellent predators. Mol. Cell. Proteom. 2012, 11, 640–650. [Google Scholar] [CrossRef] [PubMed]
- Mourão, C.; Schwartz, E. Protease inhibitors from marine venomous animals and their counterparts in terrestrial venomous animals. Mar. Drugs 2013, 11, 2069–2112. [Google Scholar] [CrossRef] [PubMed]
- Wan, H.; Lee, K.S.; Kim, B.Y.; Zou, F.M.; Yoon, H.J.; Je, Y.H.; Li, J.; Jin, B.R. A spider-derived Kunitz-type serine protease inhibitor that acts as a plasmin inhibitor and an elastase inhibitor. PLoS ONE 2013, 8, e53343. [Google Scholar] [CrossRef] [PubMed]
- Kini, R.M.; Doley, R. Structure, function and evolution of three-finger toxins: Mini proteins with multiple targets. Toxicon 2010, 56, 855–867. [Google Scholar] [CrossRef] [PubMed]
- Corrêa-Netto, C.; de LM Junqueira-de-Azevedo, I.; Silva, D.A.; Ho, P.L.; Leitão-de-Araújo, M.; Alves, M.L.M.; Sanz, L.; Foguel, D.; Zingali, R.B.; Calvete, J.J. Snake venomics and venom gland transcriptomic analysis of Brazilian coral snakes, Micrurus altirostris and M. corallinus. J. Proteom. 2011, 74, 1795–1809. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Yu, H.; Xue, W.; Yue, Y.; Liu, S.; Xing, R.; Li, P. Jellyfish venomics and venom gland transcriptomics analysis of Stomolophus meleagris to reveal the toxins associated with sting. J. Proteom. 2014, 106, 17–29. [Google Scholar] [CrossRef] [PubMed]
- Brinkman, D.; Burnell, J. Identification, cloning and sequencing of two major venom proteins from the box jellyfish, Chironex fleckeri. Toxicon 2007, 50, 850–860. [Google Scholar] [CrossRef] [PubMed]
- Brinkman, D.; Burnell, J. Partial purification of cytolytic venom proteins from the box jellyfish, Chironex fleckeri. Toxicon 2008, 51, 853–863. [Google Scholar] [CrossRef] [PubMed]
- Lassen, S.; Helmholz, H.; Ruhnau, C.; Prange, A. A novel proteinaceous cytotoxin from the northern Scyphozoa Cyanea capillata (L.) with structural homology to cubozoan haemolysins. Toxicon 2011, 57, 721–729. [Google Scholar] [CrossRef] [PubMed]
- Nagai, H.; Takuwa, K.; Nakao, M.; Sakamoto, B.; Crow, G.L.; Nakajima, T. Isolation and characterization of a novel protein toxin from the Hawaiian box jellyfish (sea wasp) Carybdea alata. Biochem. Biophys. Res. Commun. 2000, 275, 589–594. [Google Scholar] [CrossRef] [PubMed]
- Azuma, H.; Sekizaki, S.; Satoh, A.; Nakajima, T. Platelet aggregation caused by carybdea rastonii toxins (CrTX-I, II, and III) obtained from a jellyfish, carybdea rastonii. Proc. Soc. Exp. Biol. Med. 1986, 182, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Nagai, H.; Takuwa, K.; Nakao, M.; Ito, E.; Miyake, M.; Noda, M.; Nakajima, T. Novel proteinaceous toxins from the box jellyfish (sea wasp) Carybdea rastoni. Biochem. Biophys. Res. Commun. 2000, 275, 582–588. [Google Scholar] [CrossRef] [PubMed]
- König, W.; Faltin, Y.; Scheffer, J.; Schöffler, H.; Braun, V. Role of cell-bound hemolysin as a pathogenicity factor for Serratia infections. Infect. Immunity 1987, 55, 2554–2561. [Google Scholar]
- Poole, K.; Schiebel, E.; Braun, V. Molecular characterization of the hemolysin determinant of Serratia marcescens. J. Bacteriol. 1988, 170, 3177–3188. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.; Ma, D.; Shi, J.; Chengappa, M. Molecular characterization of a leukotoxin gene from a Pasteurella haemolytica-like organism, encoding a new member of the RTX toxin family. Infect. Immunity 1993, 61, 2089–2095. [Google Scholar]
- Singh, M.B.; Hough, T.; Theerakulpisut, P.; Avjioglu, A.; Davies, S.; Smith, P.M.; Taylor, P.; Simpson, R.J.; Ward, L.D.; McCluskey, J. Isolation of cDNA encoding a newly identified major allergenic protein of rye-grass pollen: Intracellular targeting to the amyloplast. Proc. Natl. Acad. Sci. USA 1991, 88, 1384–1388. [Google Scholar] [CrossRef] [PubMed]
- Bungy, G.A.; Rodda, S.; Roitt, I.; Brostoff, J. Mapping of T cell epitopes of the major fraction of rye grass using peripheral blood mononuclear cells from atopics and non-atopics. II. Isoallergen clone 5A of Loliuum perenne group I (Lol p I). Eur. J. Immunol. 1994, 24, 2098–2103. [Google Scholar] [CrossRef] [PubMed]
- Fujikawa, A.; Ishimaru, N.; Seto, A.; Yamada, H.; Aki, T.; Shigeta, S.; Wada, T.; Jyo, T.; Murooka, Y.; Oka, S. Cloning and characterization of a new allergen, Mag 3, from the house dust mite, Dermatophagoides farinae: Cross-reactivity with high-molecular-weight allergen. Mol. Immunol. 1996, 33, 311–319. [Google Scholar] [CrossRef]
- Arlian, L.G. House-dust-mite allergens: A review. Exp. Appl. Acarol. 1991, 10, 167–186. [Google Scholar] [CrossRef] [PubMed]
- Aki, T.; Ono, K.; Paik, S.-Y.; Wada, T.; Jyo, T.; Shigeta, S.; Murooka, Y.; Oka, S. Cloning and characterization of cDNA coding for a new allergen from the house dust mite, Dermatophagoides farinae. Int. Arch. Allergy Immunol. 1994, 103, 349–356. [Google Scholar] [CrossRef] [PubMed]
- Joseph, J.S.; Chung, M.C.; Jeyaseelan, K.; Kini, R.M. Amino Acid Sequence of Trocarin, a Prothrombin Activator From Tropidechis carinatus Venom: Its Structural Similarity to Coagulation Factor Xa. Blood 1999, 94, 621–631. [Google Scholar] [PubMed]
- Jansen, R.; Briaire, J.; Kamp, E.; Gielkens, A.; Smits, M. Structural analysis of the Actinobacillus pleuropneumoniae-RTX-toxin I (ApxI) operon. Infect. Immunity 1993, 61, 3688–3695. [Google Scholar]
- Kamp, E.M.; Popma, J.K.; Anakotta, J.; Smits, M.A. Identification of hemolytic and cytotoxic proteins of Actinobacillus pleuropneumoniae by use of monoclonal antibodies. Infect. Immunity 1991, 59, 3079–3085. [Google Scholar]
- De Grandis, S.; Ginsberg, J.; Toone, M.; Climie, S.; Friesen, J.; Brunton, J. Nucleotide sequence and promoter mapping of the Escherichia coli Shiga-like toxin operon of bacteriophage H-19B. J. Bacteriol. 1987, 169, 4313–4319. [Google Scholar] [CrossRef] [PubMed]
- Karmali, M.A.; Petric, M.; Lim, C.; Fleming, P.C.; Arbus, G.S.; Lior, H. The association between idiopathic hemolytic uremic syndrome and infection by verotoxin-producing Escherichia coli. J. Infect. Dis. 1985, 151, 775–782. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, A.; Lively, T.; Chen, M.; Rothman, S.; Formal, S. Escherichia coli 0157: H7 strains associated with haemorrhagic colitis in the United States produce a Shigella dysenteriae 1 (Shiga) like cytotoxin. Lancet 1983, 321, 702. [Google Scholar] [CrossRef]
- Kanki, K.; Akechi, Y.; Ueda, C.; Tsuchiya, H.; Shimizu, H.; Ishijima, N.; Toriguchi, K.; Hatano, E.; Endo, K.; Hirooka, Y. Biological and clinical implications of retinoic acid-responsive genes in human hepatocellular carcinoma cells. J. Hepatol. 2013, 59, 1037–1044. [Google Scholar] [CrossRef] [PubMed]
- García-Santisteban, I.; Bañuelos, S.; Rodríguez, J.A. A global survey of CRM1-dependent nuclear export sequences in the human deubiquitinase family. Biochem. J. 2012, 441, 209–217. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Hartwieg, E.; Horvitz, H.R. CED-1 is a transmembrane receptor that mediates cell corpse engulfment in C. elegans. Cell 2001, 104, 43–56. [Google Scholar] [CrossRef]
- Ellis, R.E.; Jacobson, D.M.; Horvitz, H.R. Genes required for the engulfment of cell corpses during programmed cell death in Caenorhabditis elegans. Genetics 1991, 129, 79–94. [Google Scholar] [PubMed]
- Yu, X.; Odera, S.; Chuang, C.-H.; Lu, N.; Zhou, Z. C. elegans Dynamin mediates the signaling of phagocytic receptor CED-1 for the engulfment and degradation of apoptotic cells. Dev. Cell 2006, 10, 743–757. [Google Scholar] [CrossRef] [PubMed]
- Carrette, T.; Seymour, J. A rapid and repeatable method for venom extraction from Cubozoan nematocysts. Toxicon 2004, 44, 135–139. [Google Scholar] [CrossRef] [PubMed]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Mortz, E.; Krogh, T.N.; Vorum, H.; Görg, A. Improved silver staining protocols for high sensitivity protein identification using matrix-assisted laser desorption/ionization-time of flight analysis. Proteomics 2001, 1, 1359–1363. [Google Scholar] [CrossRef]
- Shevchenko, A.; Jensen, O.N.; Podtelejnikov, A.V.; Sagliocco, F.; Wilm, M.; Vorm, O.; Mortensen, P.; Shevchenko, A.; Boucherie, H.; Mann, M. Linking genome and proteome by mass spectrometry: Large-scale identification of yeast proteins from two dimensional gels. Proc. Natl. Acad. Sci. USA 1996, 93, 14440–14445. [Google Scholar] [CrossRef] [PubMed]
- Choi, N.-S.; Kim, S.-H. Application of fibrin zymography for determining the optimum culture time for protease activity. Biotechnol. Tech. 1999, 13, 899–901. [Google Scholar] [CrossRef]
- d’Avila-Levy, C.M.; Santos, A.L.; Cuervo, P.; de Jesus, J.B.; Branquinha, M.H. Applications of zymography (substrate-SDS-PAGE) for peptidase screening in a post-genomic era. In Gel Electrophoresis-Advanced Techniques; InTech: Rijeka, Croatia, 2012. [Google Scholar]
- Díaz-García, A.; Ruiz-Fuentes, J.L.; Yglesias-Rivera, A.; Rodríguez-Sánchez, H.; Garlobo, Y.R.; Martinez, O.F.; Castro, J.A.F. Enzymatic analysis of venom from Cuban scorpion Rhopalurus junceus. J. Venom Res. 2015, 6, 11. [Google Scholar] [PubMed]
- Muller, V.D.; Soares, R.O.; dos Santos-Junior, N.N.; Trabuco, A.C.; Cintra, A.C.; Figueiredo, L.T.; Caliri, A.; Sampaio, S.V.; Aquino, V.H. Phospholipase A2 isolated from the venom of Crotalus durissus terrificus inactivates dengue virus and other enveloped viruses by disrupting the viral envelope. PLoS ONE 2014, 9, e112351. [Google Scholar] [CrossRef] [PubMed]
- Mi, H.; Muruganujan, A.; Casagrande, J.T.; Thomas, P.D. Large-scale gene function analysis with the PANTHER classification system. Nat. Protoc. 2013, 8, 1551. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).