Inhibition of Aflatoxin Production by Paraquat and External Superoxide Dismutase in Aspergillus flavus
Abstract
:1. Introduction
2. Results
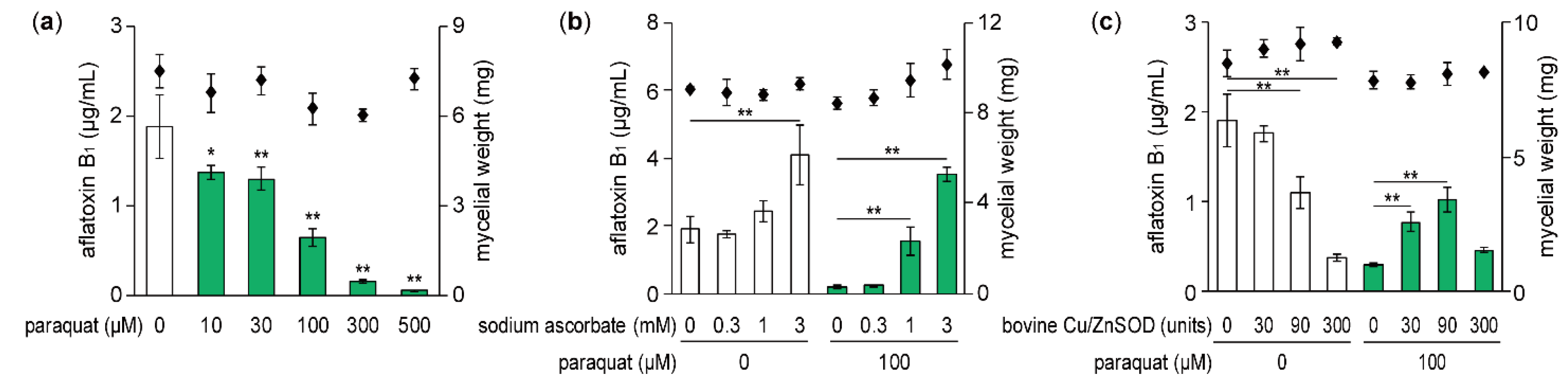
2.1. Effect of Paraquat on Aflatoxin Production
2.2. Effect of External SOD on Aflatoxin Production
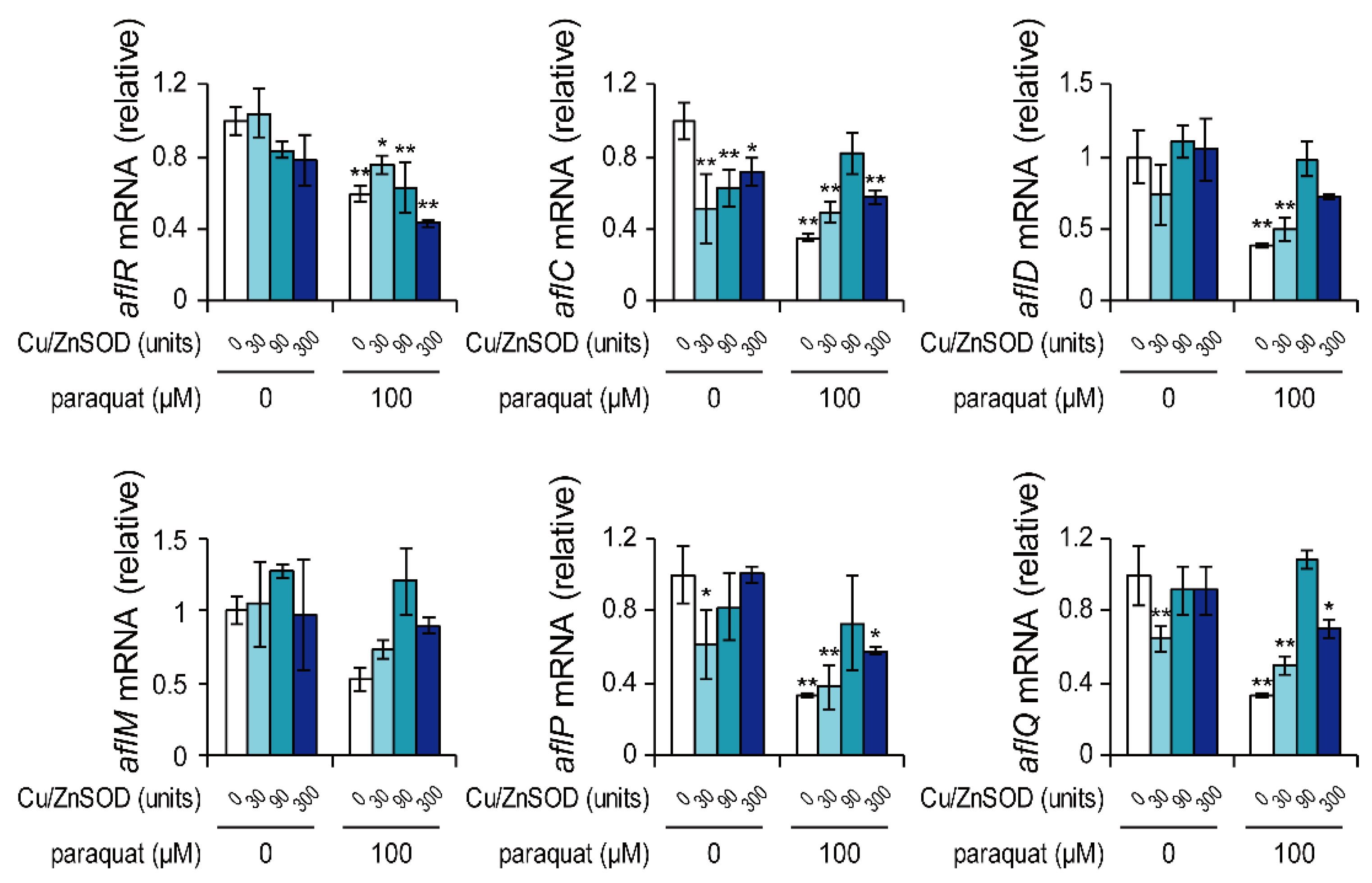
2.3. Effects of Paraquat and External SOD on mRNA Levels of Genes Responsible for Aflatoxin Biosynthesis
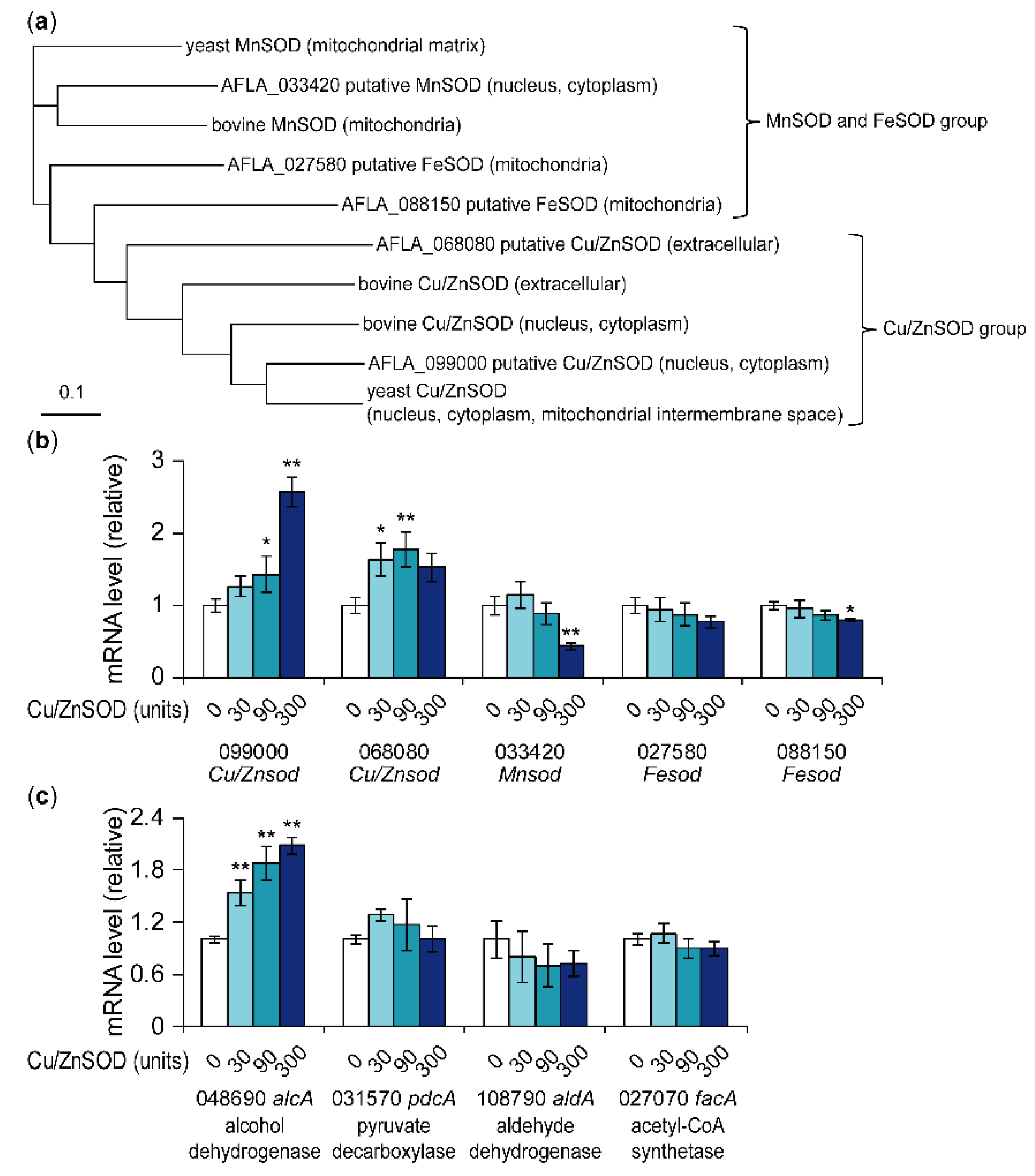
2.4. Effects of External SOD on mRNA Levels of Genes Encoding SOD and Acetyl-CoA Metabolic Enzymes
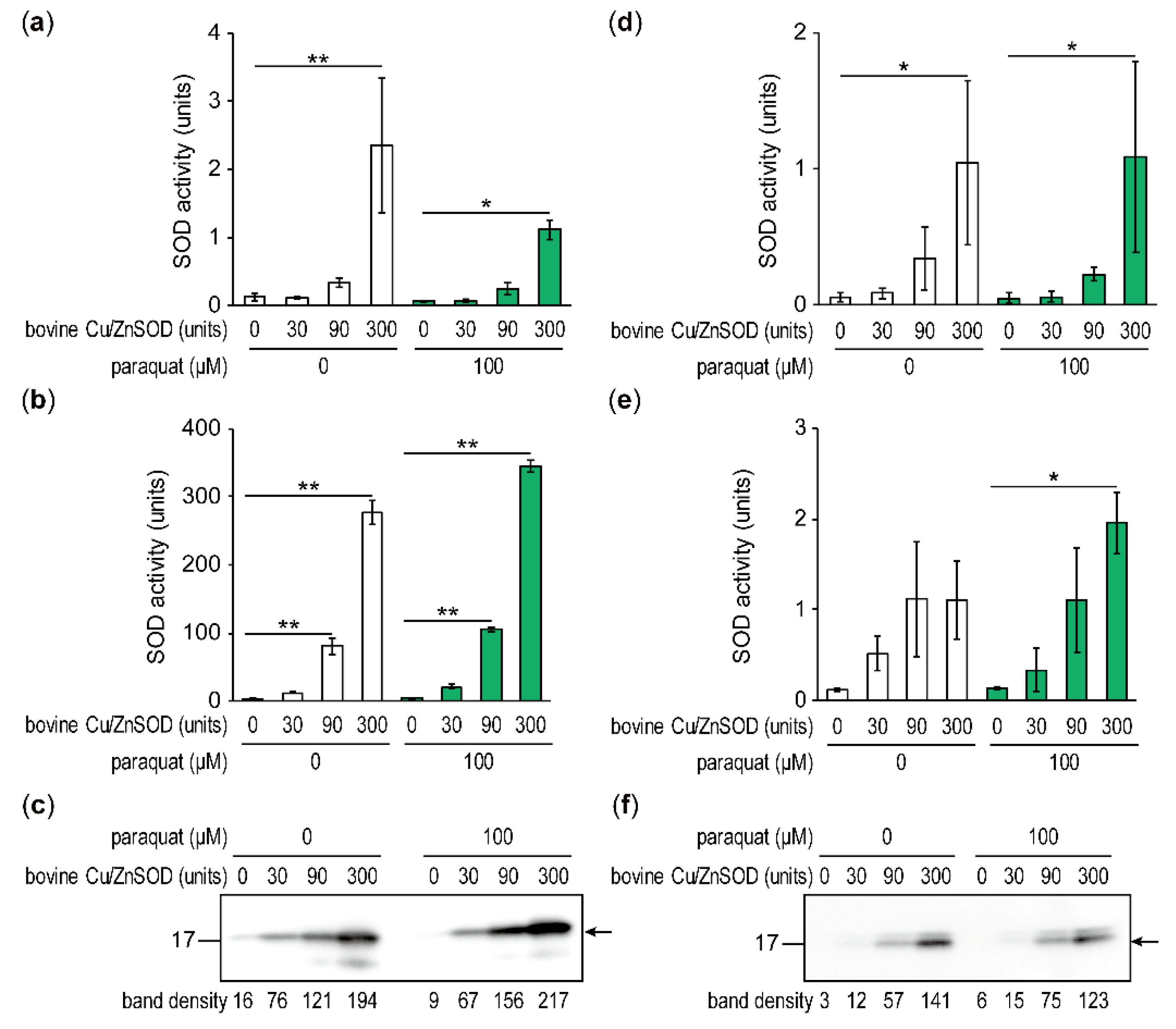
2.5. SOD Activities in Fungal Cells
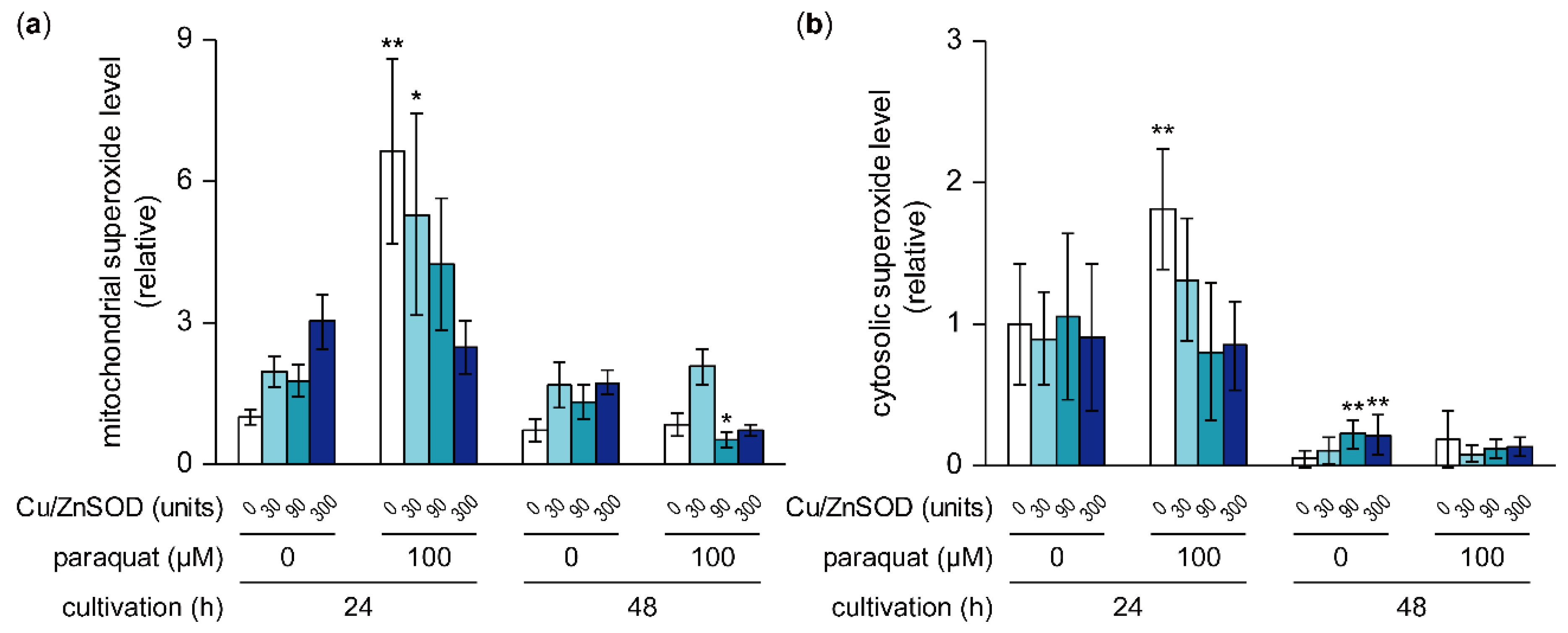
2.6. Effects of Paraquat and External SOD on Mitochondrial and Cytosolic Superoxide Levels
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Chemicals
5.2. Strain and Culture Conditions
5.3. Analysis of Aflatoxin B1 Production and Mycelial Weight
5.4. Determination of SOD Activity
5.5. Western Blotting
5.6. RT-qPCR Analysis
5.7. Determination of Superoxide Level
5.8. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Bennett, J.W.; Klich, M. Mycotoxins. Clin. Microbiol. Rev. 2003, 16, 497–516. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wu, F. Global burden of Aflatoxin-induced hepatocellular carcinoma: A risk assessment. Environ. Health Perspect. 2010, 118, 818–824. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, N.J.; Bowers, E.; Hurburgh, C.; Wu, F. Potential economic losses to the US corn industry from aflatoxin contamination. Food Addit. Contam. Part A 2016, 33, 540–550. [Google Scholar] [CrossRef] [PubMed]
- Ehrlich, K.C. Non-aflatoxigenic Aspergillus flavus to prevent aflatoxin contamination in crops: Advantages and limitations. Front. Microbiol. 2014, 5, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Villers, P. Aflatoxins and safe storage. Front. Microbiol. 2014, 5, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Cary, J.W.; Rajasekaran, K.; Brown, R.L.; Luo, M.; Chen, Z.-Y.; Bhatnagar, D. Developing resistance to aflatoxin in maize and cottonseed. Toxins (Basel) 2011, 3, 678–696. [Google Scholar] [CrossRef] [PubMed]
- Yabe, K.; Nakajima, H. Enzyme reactions and genes in aflatoxin biosynthesis. Appl. Microbiol. Biotechnol. 2004, 64, 745–755. [Google Scholar] [CrossRef] [PubMed]
- Ehrlich, K.C.; Montalbano, B.G.; Cary, J.W. Binding of the C6-zinc cluster protein, AFLR, to the promoters of aflatoxin pathway biosynthesis genes in Aspergillus parasiticus. Gene 1999, 230, 249–257. [Google Scholar] [CrossRef]
- Bayram, O.; Krappmann, S.; Ni, M.; Bok, J.W.; Helmstaedt, K.; Valerius, O.; Braus-Stromeyer, S.; Kwon, N.-J.; Keller, N.P.; Yu, J.-H.; et al. VelB/VeA/LaeA complex coordinates light signal with fungal development and secondary metabolism. Science 2008, 320, 1504–1506. [Google Scholar] [CrossRef]
- Tilburn, J.; Sarkar, S.; Widdick, D.A.; Espeso, E.A.; Orejas, M.; Mungroo, J.; Peñalva, M.; Arst, H.N., Jr. The Aspergillus PacC zinc finger transcription factor mediates regulation of both acid- and alkaline-expressed genes by ambient pH. EMBO J. 1995, 14, 779–790. [Google Scholar] [CrossRef]
- Ehrlich, K.C.; Montalbano, B.G.; Cary, J.W.; Cotty, P.J. Promoter elements in the aflatoxin pathway polyketide synthase gene. Biochim. Biophys. Acta 2002, 1576, 171–175. [Google Scholar] [CrossRef]
- Reverberi, M.; Zjalic, S.; Ricelli, A.; Punelli, F.; Camera, E.; Fabbri, C.; Picardo, M.; Fanelli, C.; Fabbri, A.A. Modulation of antioxidant defense in Aspergillus parasiticus is involved in aflatoxin biosynthesis: A role for the ApyapA gene. Eukaryot. Cell 2008, 7, 988–1000. [Google Scholar] [CrossRef]
- Narasaiah, K.V.; Sashidhar, R.B.; Subramanyam, C. Biochemical analysis of oxidative stress in the production of aflatoxin and its precursor intermediates. Mycopathologia 2006, 162, 179–189. [Google Scholar] [CrossRef]
- Fountain, J.C.; Scully, B.T.; Chen, Z.-Y.; Gold, S.E.; Glenn, A.E.; Abbas, H.K.; Lee, R.D.; Kemerait, R.C.; Guo, B. Effects of hydrogen peroxide on different toxigenic and atoxigenic isolates of Aspergillus flavus. Toxins (Basel) 2015, 7, 2985–2999. [Google Scholar] [CrossRef] [PubMed]
- Fountain, J.C.; Koh, J.; Yang, L.; Pandey, M.K.; Nayak, S.N.; Bajaj, P.; Zhuang, W.-J.; Chen, Z.-Y.; Kemerait, R.C.; Lee, R.D.; et al. Proteome analysis of Aspergillus flavus isolate-specific responses to oxidative stress in relationship to aflatoxin production capability. Sci. Rep. 2018, 8, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Apel, K.; Hirt, H. Reactive oxygen species: Metabolism, oxidative stress, and signal transduction. Annu. Rev. Plant Biol. 2004, 55, 373–399. [Google Scholar] [CrossRef] [PubMed]
- Lustgarten, M.S.; Bhattacharya, A.; Muller, F.L.; Jang, Y.C.; Shimizu, T.; Shirasawa, T.; Richardson, A.; Van Remmen, H. Complex I generated, mitochondrial matrix-directed superoxide is released from the mitochondria through voltage dependent anion channels. Biochem. Biophys. Res. Commun. 2012, 422, 515–521. [Google Scholar] [CrossRef]
- Liochev, S.I.; Fridovich, I. Superoxide and iron: Partners in crime. IUBMB Life 1999, 48, 157–161. [Google Scholar] [CrossRef]
- D’Autréaux, B.; Toledano, M.B. ROS as signalling molecules: Mechanisms that generate specificity in ROS homeostasis. Nat. Rev. Mol. Cell Biol. 2007, 8, 813–824. [Google Scholar] [CrossRef]
- Jayashree, T.; Subramanyam, C. Oxidative stress as a prerequisite for aflatoxin production by Aspergillus parasiticus. Free Radic. Biol. Med. 2000, 29, 981–985. [Google Scholar] [CrossRef]
- Grintzalis, K.; Vernardis, S.I.; Klapa, M.I.; Georgiou, C.D. Role of oxidative stress in sclerotial differentiation and aflatoxin B1 biosynthesis in Aspergillus flavus. Appl. Environ. Microbiol. 2014, 80, 5561–5571. [Google Scholar] [CrossRef] [PubMed]
- Zaccaria, M.; Ludovici, M.; Sanzani, S.M.; Ippolito, A.; Cigliano, R.A.; Sanseverino, W.; Scarpari, M.; Scala, V.; Fanelli, C.; Reverberi, M. Menadione-induced oxidative stress re-shapes the oxylipin profile of Aspergillus flavus and its lifestyle. Toxins (Basel) 2015, 7, 4315–4329. [Google Scholar] [CrossRef] [PubMed]
- Ponts, N.; Pinson-Gadais, L.; Verdal-Bonnin, M.-N.; Barreau, C.; Richard-Forget, F. Accumulation of deoxynivalenol and its 15-acetylated form is significantly modulated by oxidative stress in liquid cultures of Fusarium graminearum. FEMS Microbiol. Lett. 2006, 258, 102–107. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, T.; Sakamoto, N.; Suzuki, M.; Kimura, M.; Nagasawa, H.; Sakuda, S. Precocene II, a trichothecene production inhibitor, binds to voltage-dependent anion channel and increases the superoxide level in mitochondria of Fusarium graminearum. PLoS ONE 2015, 10, e0135031. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, T.; Yoshinari, T.; Sakuda, S. Intracellular superoxide level controlled by manganese superoxide dismutases affects trichothecene production in Fusarium graminearum. FEMS Microbiol. Lett. 2017, 364, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Chung, W.H. Unraveling new functions of superoxide dismutase using yeast model system: Beyond its conventional role in superoxide radical scavenging. J. Microbiol. 2017, 55, 409–416. [Google Scholar] [CrossRef] [PubMed]
- Emanuelsson, O.; Nielsen, H.; Brunak, S.; von Heijne, G. Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J. Mol. Biol. 2000, 300, 1005–1016. [Google Scholar] [CrossRef] [PubMed]
- Miller, A.F. Superoxide dismutases: Ancient enzymes and new insights. FEBS Lett. 2012, 586, 585–595. [Google Scholar] [CrossRef]
- Chanda, A.; Roze, L.V.; Kang, S.; Artymovich, K.A.; Hicks, G.R.; Raikhel, N.V.; Calvo, A.M.; Linz, J.E. A key role for vesicles in fungal secondary metabolism. Proc. Natl. Acad. Sci. USA 2009, 106, 19533–19538. [Google Scholar] [CrossRef] [PubMed]
- European Bioinformatics Institute. Available online: https://www.ebi.ac.uk/Tools/msa/clustalo/ (accessed on 15 January 2019).
- UniProtKB. Available online: https://www.uniprot.org/ (accessed on 15 January 2019).
- Peñalva, M.A. Tracing the endocytic pathway of Aspergillus nidulans with FM4-64. Fungal Genet. Biol. 2005, 42, 963–975. [Google Scholar] [CrossRef]
- Higuchi, Y.; Nakahama, T.; Shoji, J.; Arioka, M.; Kitamoto, K. Visualization of the endocytic pathway in the filamentous fungus Aspergillus oryzae using an EGFP-fused plasma membrane protein. Biochem. Biophys. Res. Commun. 2006, 340, 784–791. [Google Scholar] [CrossRef] [PubMed]
- Nikiforova, A.B.; Saris, N.-E.L.; Kruglov, A.G. External mitochondrial NADH-dependent reductase of redox cyclers: VDAC1 or Cyb5R3? Free Radic. Biol. Med. 2014, 74, 74–84. [Google Scholar] [CrossRef] [PubMed]
- Flint, D.H.; Tuminello, J.F.; Emptage, M.H. The inactivation of Fe-S cluster containing hydro-lyases by superoxide. J. Biol. Chem. 1993, 268, 22369–22376. [Google Scholar] [PubMed]
- Breitenbach, M.; Rinnerthaler, M.; Hartl, J.; Stincone, A.; Vowinckel, J.; Breitenbach-Koller, H.; Ralser, M. Mitochondria in ageing: There is metabolism beyond the ROS. FEMS Yeast Res. 2014, 14, 198–212. [Google Scholar] [CrossRef] [PubMed]
- Sakuda, S.; Prabowo, D.F.; Takagi, K.; Shiomi, K.; Mori, M.; Omura, S.; Nagasawa, H. Inhibitory effects of respiration inhibitors on aflatoxin production. Toxins (Basel) 2014, 6, 1193–1200. [Google Scholar] [CrossRef] [PubMed]
- Tsang, C.K.; Liu, Y.; Thomas, J.; Zhang, Y.; Zheng, X.F.S. Superoxide dismutase 1 acts as a nuclear transcription factor to regulate oxidative stress resistance. Nat. Commun. 2014, 5, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Magrì, A.; Rosa, M.C.D.; Tomasello, M.F.; Guarino, F.; Reina, S.; Messina, A.; Pinto, V.D. Overexpression of human SOD1 in VDAC1-less yeast restores mitochondrial functionality modulating beta-barrel outer membrane protein genes. Biochim. Biophys. Acta 2016, 1857, 789–798. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Furukawa, T.; Sakuda, S. Inhibition of Aflatoxin Production by Paraquat and External Superoxide Dismutase in Aspergillus flavus. Toxins 2019, 11, 107. https://doi.org/10.3390/toxins11020107
Furukawa T, Sakuda S. Inhibition of Aflatoxin Production by Paraquat and External Superoxide Dismutase in Aspergillus flavus. Toxins. 2019; 11(2):107. https://doi.org/10.3390/toxins11020107
Chicago/Turabian StyleFurukawa, Tomohiro, and Shohei Sakuda. 2019. "Inhibition of Aflatoxin Production by Paraquat and External Superoxide Dismutase in Aspergillus flavus" Toxins 11, no. 2: 107. https://doi.org/10.3390/toxins11020107
APA StyleFurukawa, T., & Sakuda, S. (2019). Inhibition of Aflatoxin Production by Paraquat and External Superoxide Dismutase in Aspergillus flavus. Toxins, 11(2), 107. https://doi.org/10.3390/toxins11020107




