Detection of Abrin-Like and Prepropulchellin-Like Toxin Genes and Transcripts Using Whole Genome Sequencing and Full-Length Transcript Sequencing of Abrus precatorius
Abstract
:1. Introduction
2. Results
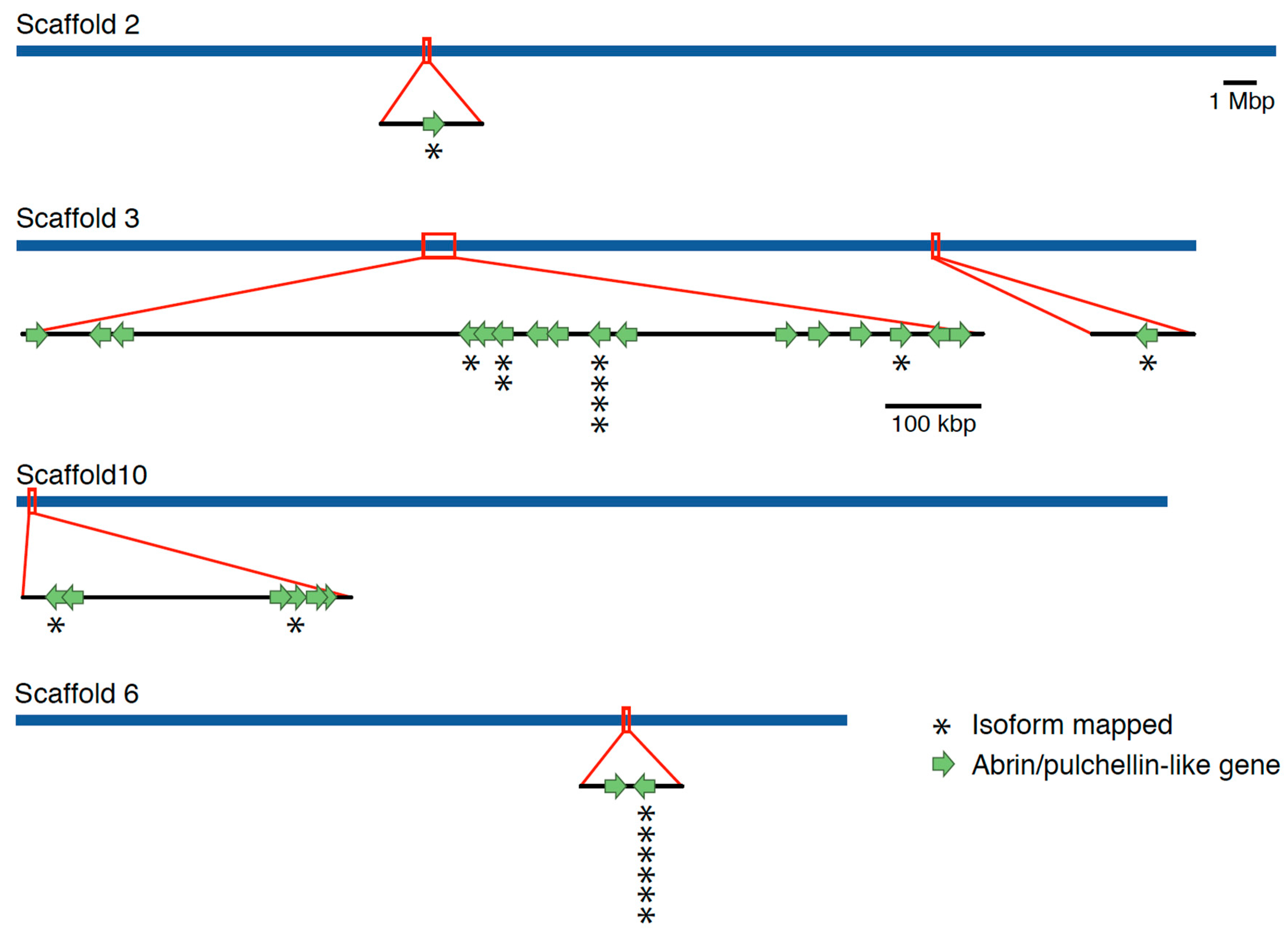
Genome Sequencing, Assembly, and Annotation
3. Discussion and Conclusions
4. Materials and Methods
4.1. Abrus Precatorius DNA Extraction and Preparation
4.2. Genome Sequencing
4.3. Genome Assembly and Polishing
4.4. Full-Length Isoform Transcriptomic Sequencing and Analysis (Iso-Seq)
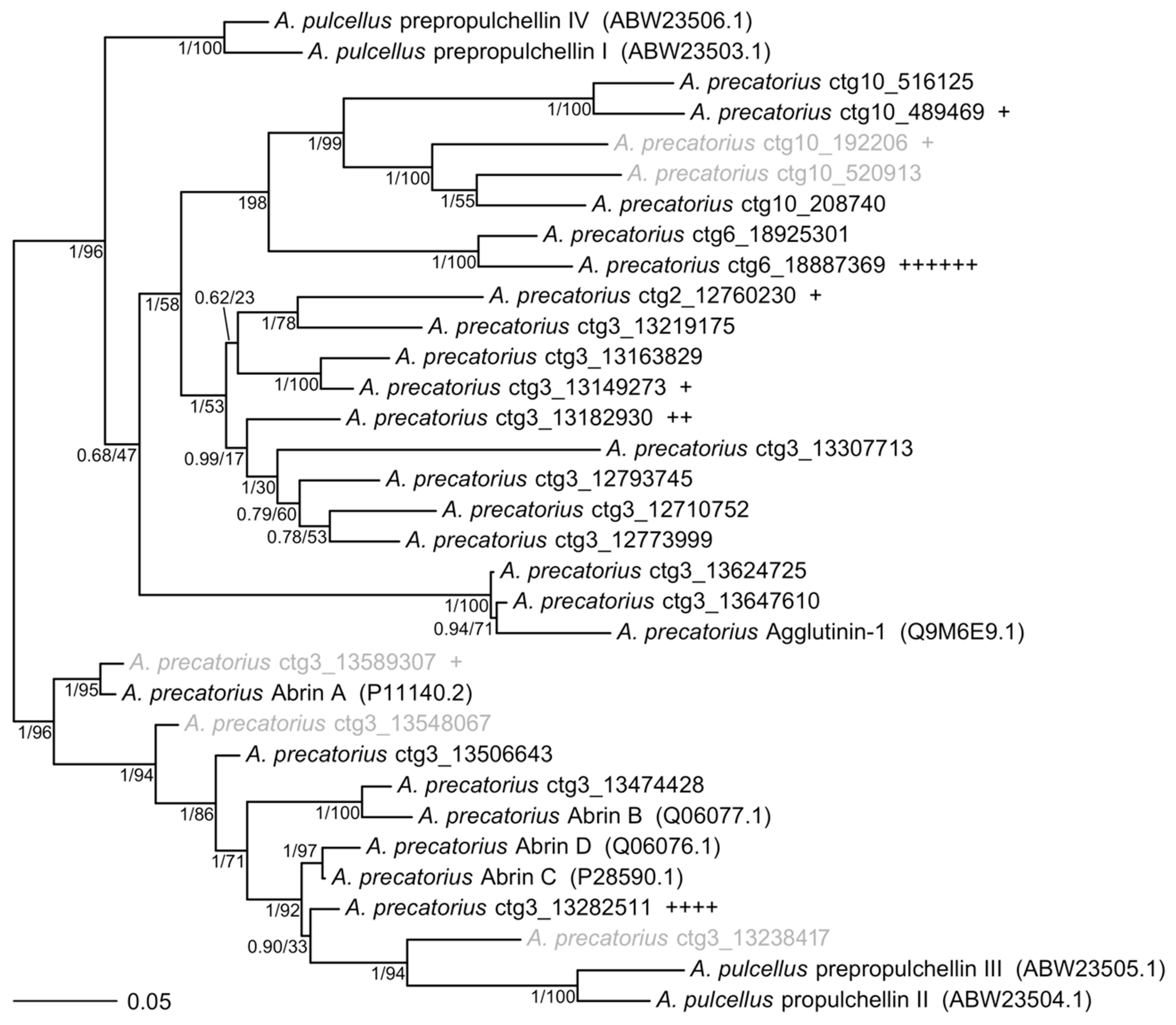
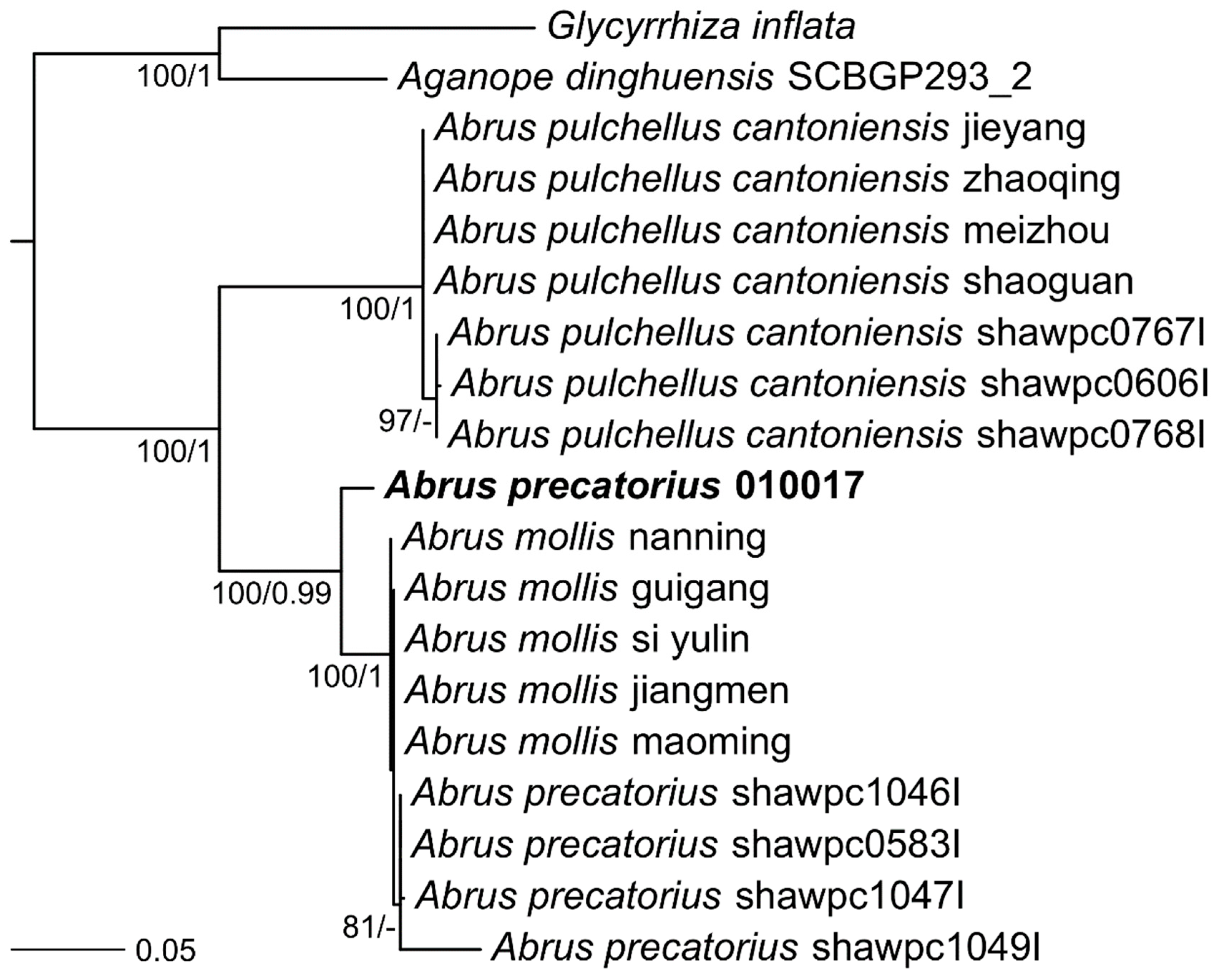
4.5. Phylogenetic Trees
4.6. Description of the Cultivar
4.6.1. Abrus Adanson. Familles des Plantes 2: 327, 511. 1763
4.6.2. Abrus precatorius Linnaeus. Systema Naturae, ed. 12 2: 472. 1767
4.6.3. Basionym: Glycine abrus Linnaeus. Species Plantarum 2: 753-754. 1753.
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Centers for Disease Control and Prevention. HHS and USDA Select Agents and Toxins 7CFR Part 331, 9 CFR Part 121, and 42 CFR Part 73; Centers for Disease Control and Prevention: Atlanta, GA, USA, 2011.
- Reyes, L.F.; Nobre, T.M.; Pavinatto, F.J.; Zaniquelli, M.E.; Caseli, L.; Oliveira, O.N., Jr.; Araujo, A.P. The role of the C-terminal region of pulchellin A-chain in the interaction with membrane model systems. Biochim. Biophys. Acta 2012, 1818, 82–89. [Google Scholar] [CrossRef] [PubMed]
- Mishra, R.; Kumar, M.S.; Karande, A.A. Inhibition of protein synthesis leading to unfolded protein response is the major event in abrin-mediated apoptosis. Mol. Cell Biochem. 2015, 403, 255–265. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Yang, R.; Zhao, X.; Qin, D.; Liu, Z.; Liu, F.; Song, X.; Li, L.; Feng, R.; Gao, N. Abrin P2 suppresses proliferation and induces apoptosis of colon cancer cells via mitochondrial membrane depolarization and caspase activation. Acta Biochim. Biophys. Sin. (Shanghai) 2016, 48, 420–429. [Google Scholar] [CrossRef] [PubMed]
- Tahirov, T.H.; Lu, T.H.; Liaw, Y.C.; Chen, Y.L.; Lin, J.Y. Crystal structure of abrin-a at 2.14 A. J. Mol. Biol. 1995, 250, 354–367. [Google Scholar] [CrossRef] [PubMed]
- Sandvig, K.; van Deurs, B. Entry of ricin and Shiga toxin into cells: Molecular mechanisms and medical perspectives. EMBO J. 2000, 19, 5943–5950. [Google Scholar] [CrossRef] [PubMed]
- Tam, C.C.; Henderson, T.D.; Stanker, L.H.; He, X.H.; Cheng, L.W. Abrin Toxicity and Bioavailability after Temperature and pH Treatment. Toxins 2017, 9, 320. [Google Scholar] [CrossRef]
- Castilho, P.V.; Goto, L.S.; Roberts, L.M.; Araujo, A.P. Isolation and characterization of four type 2 ribosome inactivating pulchellin isoforms from Abrus pulchellus seeds. FEBS J. 2008, 275, 948–959. [Google Scholar] [CrossRef]
- Cheng, J.; Lu, T.H.; Liu, C.L.; Lin, J.Y. A biophysical elucidation for less toxicity of agglutinin than abrin-a from the seeds of Abrus precatorius in consequence of crystal structure. J. Biomed. Sci. 2010, 17, 34. [Google Scholar] [CrossRef]
- Lin, J.Y.; Lee, T.C.; Tung, T.C. Isolation of antitumor proteins abrin-A and abrin-B from Abrus precatorius. Int. J. Pept. Protein Res. 1978, 12, 311–317. [Google Scholar] [CrossRef]
- Lin, J.Y.; Lee, T.C.; Hu, S.T.; Tung, T.C. Isolation of four isotoxic proteins and one agglutinin from jequiriti bean (Abrus precatorius). Toxicons 1981, 19, 41–51. [Google Scholar] [CrossRef]
- Wood, K.A.; Lord, J.M.; Wawrzynczak, E.J.; Piatak, M. Preproabrin: Genomic cloning, characterisation and the expression of the A-chain in Escherichia coli. Eur J Biochem 1991, 198, 723–732. [Google Scholar] [CrossRef] [PubMed]
- Hung, C.H.; Lee, M.C.; Lee, T.C.; Lin, J.W. Primary Structure of Three Distinct Isoabrins Determined by cDNA Sequencing. J. Mol. Biol. 1992, 229, 263–267. [Google Scholar] [CrossRef] [PubMed]
- Goto, L.S.; Beltramini, L.M.; de Moraes, D.I.; Moreira, R.A.; de Araújo, A.P.U. Abrus pulchellus type-2 RIP, pulchellin: Heterologous expression and refolding of the sugar-binding B chain. Protein Expr. Purif. 2003, 31, 12–18. [Google Scholar] [CrossRef]
- Silva, A.L.; Goto, L.S.; Dinarte, A.R.; Hansen, D.; Moreira, R.A.; Beltramini, L.M.; Araujo, A.P. Pulchellin, a highly toxic type 2 ribosome-inactivating protein from Abrus pulchellus. Cloning heterologous expression of A-chain and structural studies. FEBS J. 2005, 272, 1201–1210. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Garay, M.L. Introduction to Isoform Sequencing Using Pacific Biosciences Technology (Iso-Seq). In Transcriptomics and Gene Regulation; Wu, J., Ed.; Springer Netherlands: Berlin/Heidelberg, Germany, 2016; pp. 141–160. [Google Scholar] [CrossRef]
- Françoise, T.-N.; Alexander, S.; Terence, M.; Michael, D.; Cuccio, M.D.; Paul, K. Eukaryotic Genome Annotation Pipeline. In The NCBI Handbook; National Center for Biotechnology Information: Bethesda, MD, USA, 2013. [Google Scholar]
- Lo, C.-C.; Chain Patrick, S.G. Rapid evaluation and quality control of next generation sequencing data with FaQCs. BMC Bioinform. 2014, 15, 366. [Google Scholar] [CrossRef] [PubMed]
- Chin, C.S.; Peluso, P.; Sedlazeck, F.J.; Nattestad, M.; Concepcion, G.T.; Clum, A.; Dunn, C.; O’Malley, R.; Figueroa-Balderas, R.; Morales-Cruz, A.; et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat. Methods 2016, 13, 1050–1054. [Google Scholar] [CrossRef]
- Burton, J.N.; Adey, A.; Patwardhan, R.P.; Qiu, R.; Kitzman, J.O.; Shendure, J. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechnol. 2013, 31, 1119–1125. [Google Scholar] [CrossRef]
- Pacific Biosciences. Procedure-Checklist-Iso-Seq-Template-Preparation-for-Sequel-Systems. 2018. Available online: https://www.pacb.com/wp-content/uploads/Procedure-Checklist-Iso-Seq-Template-Preparation-for-Sequel-Systems.pdf (accessed on 27 July 2017).
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Kosiol, C.; Goldman, N. Different Versions of the Dayhoff Rate Matrix. Mol. Biol. Evol. 2005, 22, 193–199. [Google Scholar] [CrossRef] [PubMed]
| Genome Size | 347,231,436 bp |
|---|---|
| GC% | 31.8% |
| Number of contigs | 344 |
| Contig N50/L50 | 11,837,218/12 |
| Number of scaffolds | 160 |
| Scaffold N50/L50 | 35,860,869/5 |
| Predicted genes | 29,216 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hovde, B.T.; Daligault, H.E.; Hanschen, E.R.; Kunde, Y.A.; Johnson, M.B.; Starkenburg, S.R.; Johnson, S.L. Detection of Abrin-Like and Prepropulchellin-Like Toxin Genes and Transcripts Using Whole Genome Sequencing and Full-Length Transcript Sequencing of Abrus precatorius. Toxins 2019, 11, 691. https://doi.org/10.3390/toxins11120691
Hovde BT, Daligault HE, Hanschen ER, Kunde YA, Johnson MB, Starkenburg SR, Johnson SL. Detection of Abrin-Like and Prepropulchellin-Like Toxin Genes and Transcripts Using Whole Genome Sequencing and Full-Length Transcript Sequencing of Abrus precatorius. Toxins. 2019; 11(12):691. https://doi.org/10.3390/toxins11120691
Chicago/Turabian StyleHovde, Blake T., Hajnalka E. Daligault, Erik R. Hanschen, Yuliya A. Kunde, Matthew B. Johnson, Shawn R. Starkenburg, and Shannon L. Johnson. 2019. "Detection of Abrin-Like and Prepropulchellin-Like Toxin Genes and Transcripts Using Whole Genome Sequencing and Full-Length Transcript Sequencing of Abrus precatorius" Toxins 11, no. 12: 691. https://doi.org/10.3390/toxins11120691
APA StyleHovde, B. T., Daligault, H. E., Hanschen, E. R., Kunde, Y. A., Johnson, M. B., Starkenburg, S. R., & Johnson, S. L. (2019). Detection of Abrin-Like and Prepropulchellin-Like Toxin Genes and Transcripts Using Whole Genome Sequencing and Full-Length Transcript Sequencing of Abrus precatorius. Toxins, 11(12), 691. https://doi.org/10.3390/toxins11120691





