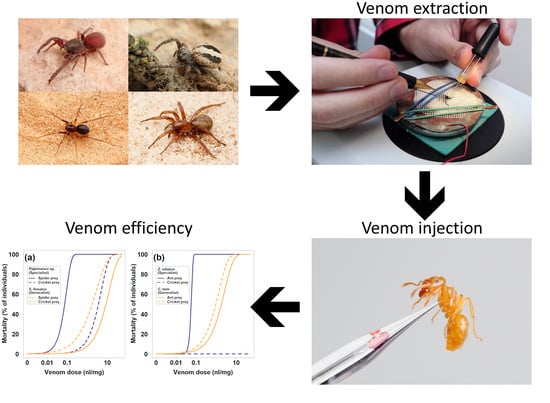
High Specific Efficiency of Venom of Two Prey-Specialized Spiders
Abstract
1. Introduction
2. Results
2.1. Efficiency of Crude Venoms
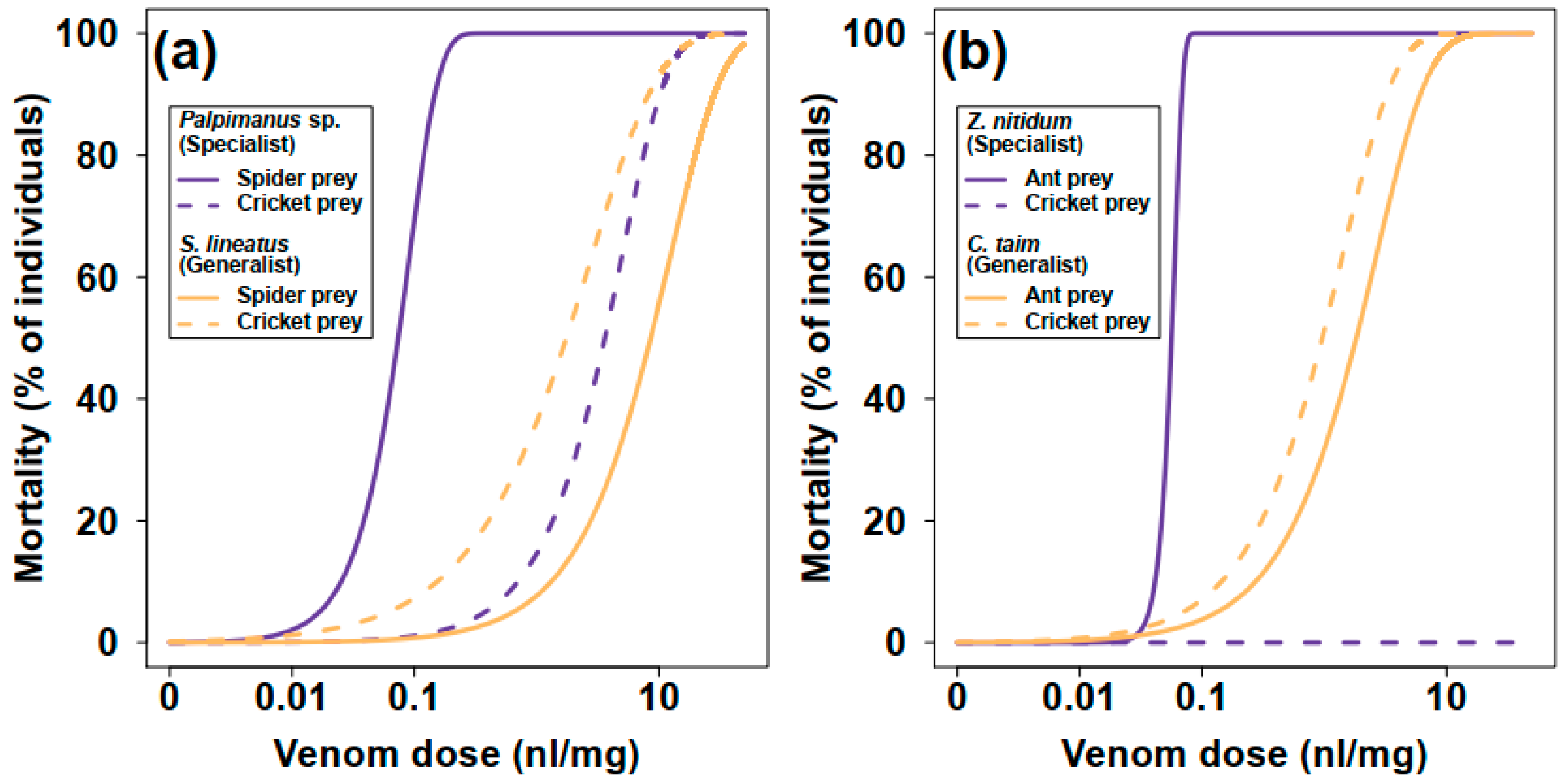
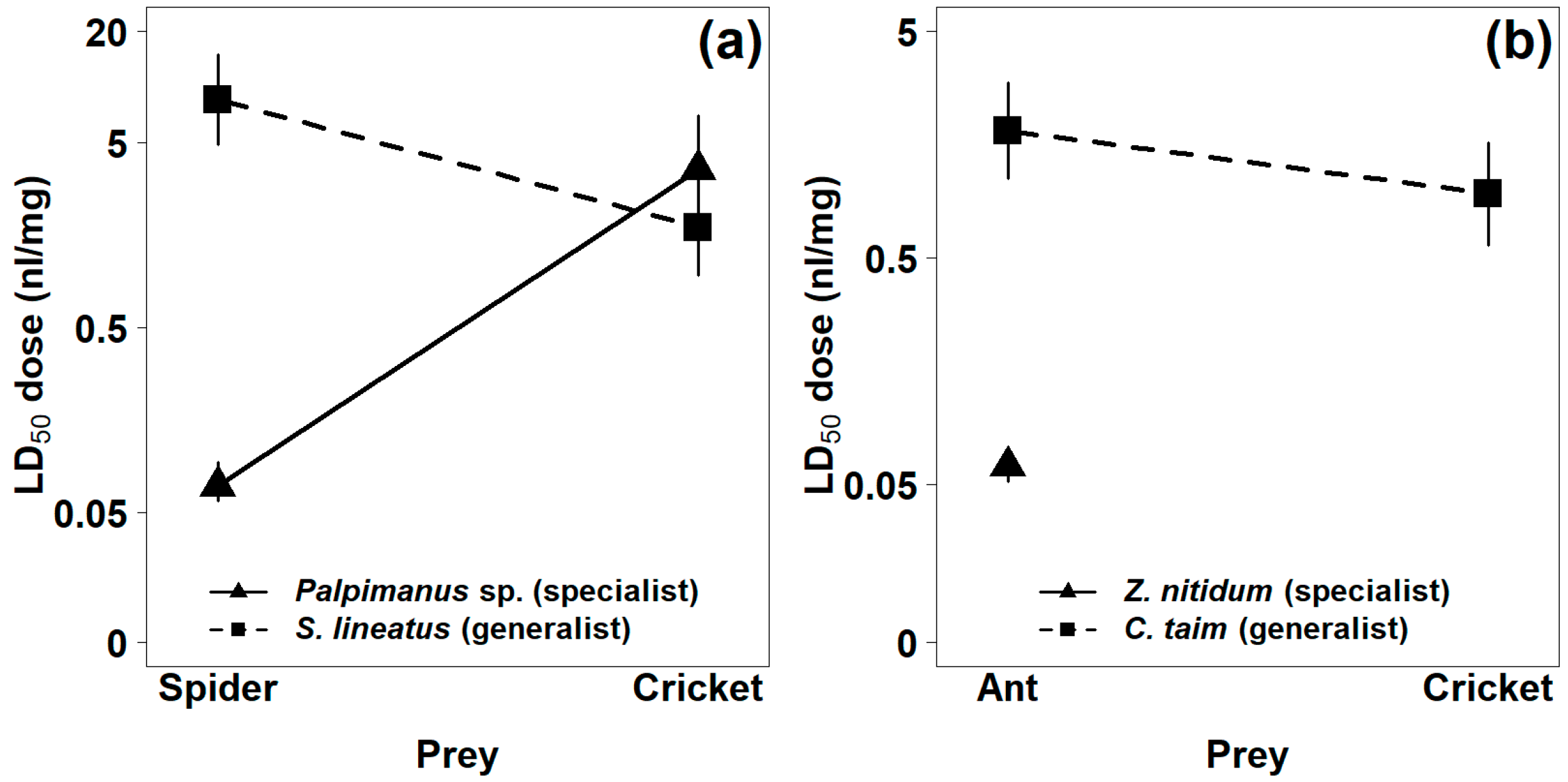
2.1.1. Mortality
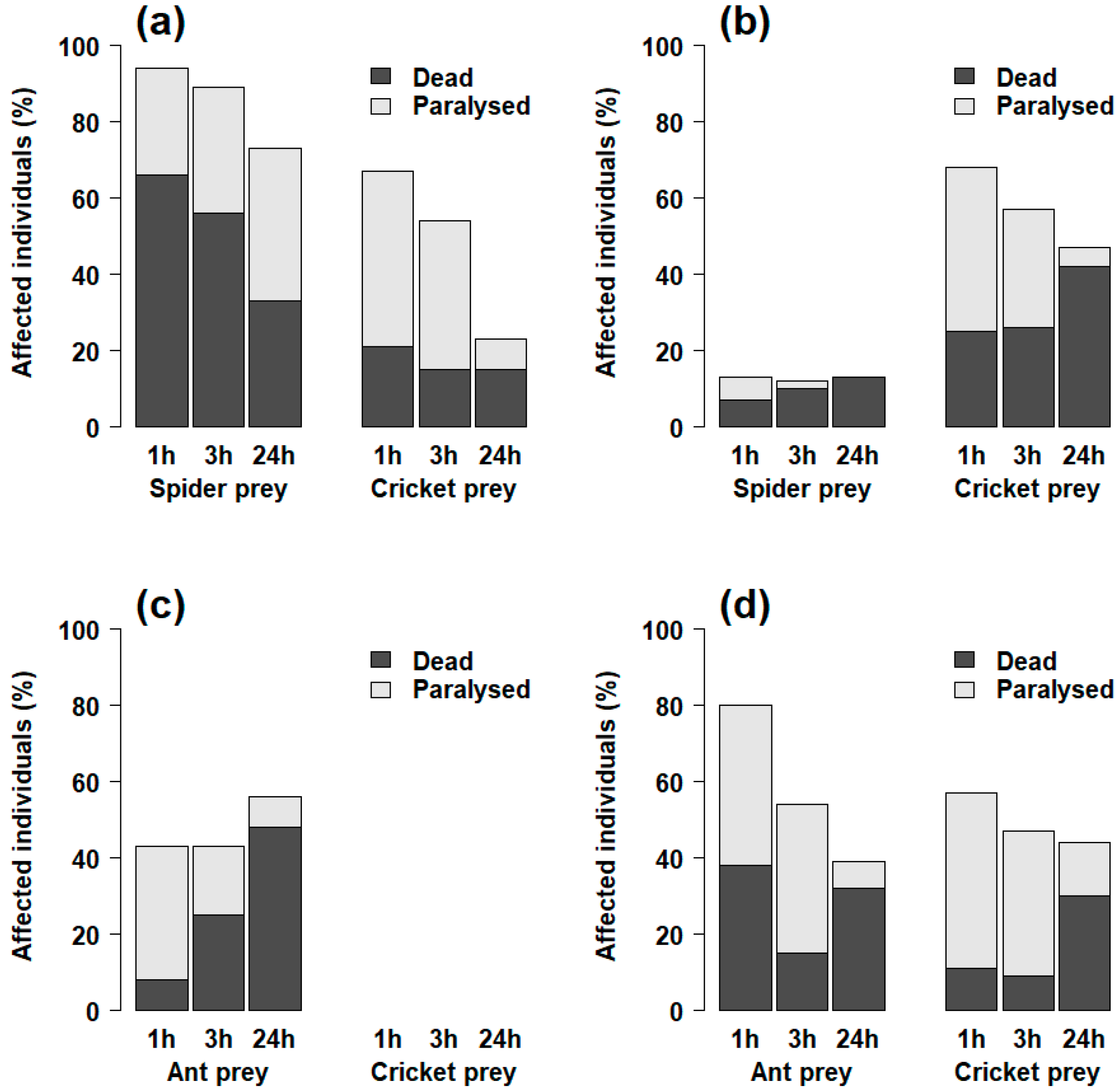
2.1.2. Paralysis
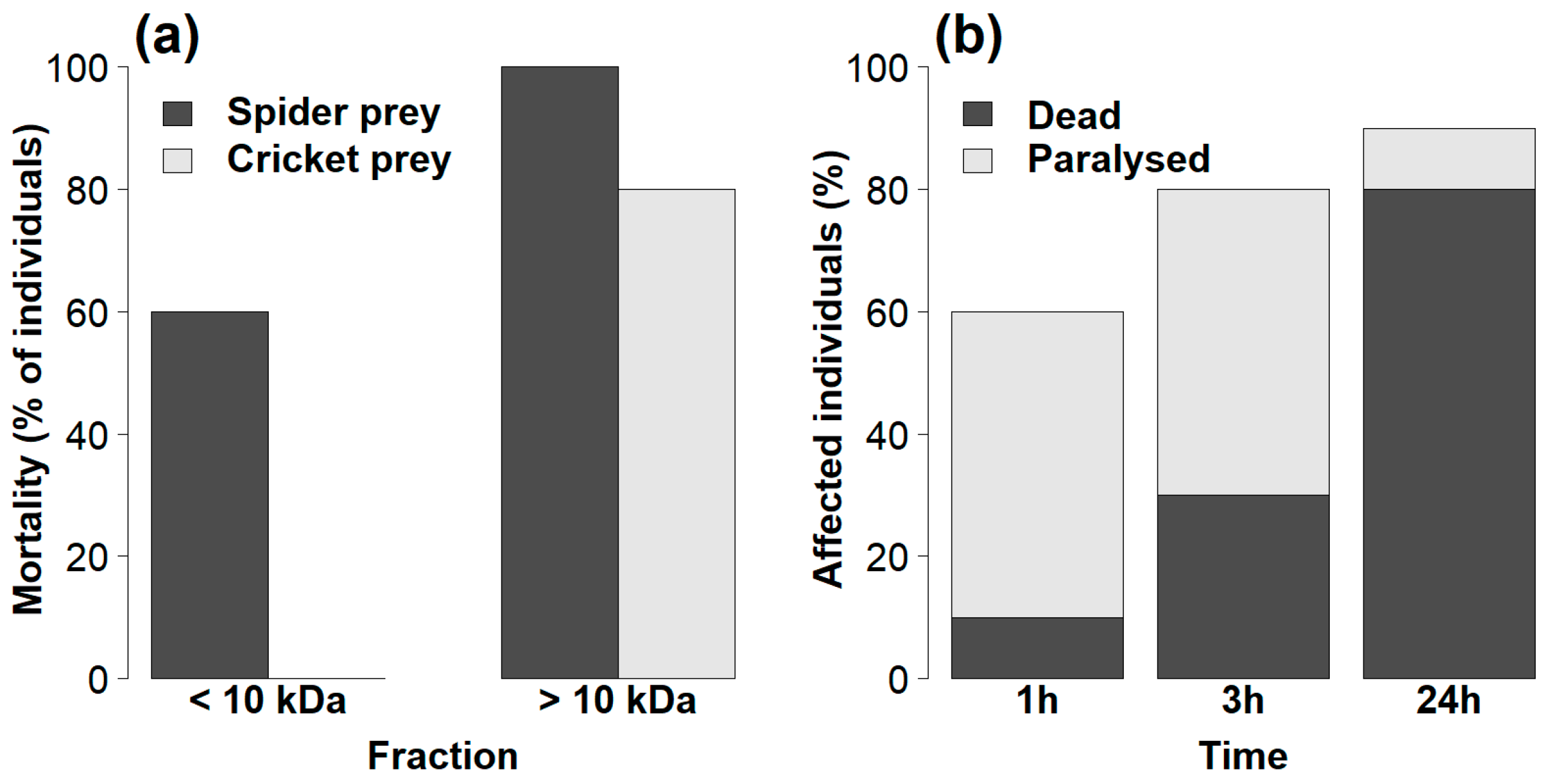
2.2. Efficiency of Venom Fractions
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Obtaining the Venom
4.3. Bioassays with Crude Venom
4.4. Bioassays with Venom Fractions of Palpimanus sp.
4.5. Data Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Fry, B.G.; Roelants, K.; Champagne, D.E.; Scheib, H.; Tyndall, J.D.; King, G.F.; Nevalainen, T.J.; Norman, J.A.; Lewis, R.J.; Norton, R.S.; et al. The toxicogenomic multiverse: Convergent recruitment of proteins into animal venoms. Annu. Rev. Genom. Hum. Genet. 2009, 10, 483–511. [Google Scholar] [CrossRef]
- Morgenstern, D.; King, G.F. The venom optimization hypothesis revisited. Toxicon 2013, 63, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.A.; Robinson, S.D.; Yeates, D.K.; Jin, J.; Baumann, K.; Dobson, J.; Fry, B.G.; King, G.F. Entomo-venomics: The evolution, biology and biochemistry of insect venoms. Toxicon 2018, 154, 5–27. [Google Scholar] [CrossRef] [PubMed]
- Casewell, N.R.; Wüster, W.; Vonk, F.J.; Harrison, R.A.; Fry, B.G. Complex cocktails: The evolutionary novelty of venoms. Trends Ecol. Evol. 2013, 28, 219–229. [Google Scholar] [CrossRef] [PubMed]
- Von Reumont, B.; Campbell, L.; Jenner, R. Quo vadis venomics? A roadmap to neglected venomous invertebrates. Toxins 2014, 6, 3488–3551. [Google Scholar] [CrossRef] [PubMed]
- Wilson, D.; Daly, N. Venomics: A mini-review. High Throughput 2018, 7, 19. [Google Scholar] [CrossRef] [PubMed]
- Chippaux, J.P.; Williams, V.; White, J. Snake venom variability: Methods of study, results and interpretation. Toxicon 1991, 29, 1279–1303. [Google Scholar] [CrossRef]
- Minton, S.A. A note on the venom of an aged rattlesnake. Toxicon 1975, 13, 73–74. [Google Scholar] [CrossRef]
- Fiero, M.K.; Seifert, M.W.; Weaver, T.J.; Bonilla, C.A. Comparative study of juvenile and adult prairie rattlesnake (Crotalus viridis viridis) venoms. Toxicon 1972, 10, 81–82. [Google Scholar] [CrossRef]
- Gubenšek, F.; Sket, D.; Turk, V.; Lebez, D. Fractionation of Vipera ammodytes venom and seasonal variation of its composition. Toxicon 1974, 12, 167–168. [Google Scholar] [CrossRef]
- Jiménez-Porras, J.M. Intraspecific variations in composition of venom of the jumping viper, Bothrops nummifera. Toxicon 1964, 2, 187–195. [Google Scholar] [CrossRef]
- Glenn, J.L.; Straight, R. Mojave rattlesnake Crotalus scutulatus scutulatus venom: Variation in toxicity with geographical origin. Toxicon 1978, 16, 81–84. [Google Scholar] [CrossRef]
- Menezes, M.C.; Furtado, M.F.; Travaglia-Cardoso, S.R.; Camargo, A.C.; Serrano, S.M. Sex-based individual variation of snake venom proteome among eighteen Bothrops jararaca siblings. Toxicon 2006, 47, 304–312. [Google Scholar] [CrossRef] [PubMed]
- Daltry, J.C.; Wüster, W.; Thorpe, R.S. Diet and snake venom evolution. Nature 1996, 379, 537–540. [Google Scholar] [CrossRef]
- Barlow, A.; Pook, C.E.; Harrison, R.A.; Wüster, W. Coevolution of diet and prey-specific venom activity supports the role of selection in snake venom evolution. Proc. R. Soc. B 2009, 276, 2443–2449. [Google Scholar] [CrossRef]
- Healy, K.; Carbone, C.; Jackson, A.L. Snake venom potency and yield are associated with prey-evolution, predator metabolism and habitat structure. Ecol. Lett. 2019, 22, 527–537. [Google Scholar] [CrossRef]
- McCue, M.D. Cost of producing venom in three North American pitviper species. Copeia 2006, 2006, 818–825. [Google Scholar] [CrossRef]
- Nisani, Z.; Dunbar, S.G.; Hayes, W.K. Cost of venom regeneration in Parabuthus transvaalicus (Arachnida: Buthidae). Comp. Biochem. Physiol. A 2007, 147, 509–513. [Google Scholar] [CrossRef]
- Pintor, A.F.; Krockenberger, A.K.; Seymour, J.E. Costs of venom production in the common death adder (Acanthophis antarcticus). Toxicon 2010, 56, 1035–1042. [Google Scholar] [CrossRef]
- Smith, M.T.; Ortega, J.; Beaupre, S.J. Metabolic cost of venom replenishment by Prairie Rattlesnakes (Crotalus viridis viridis). Toxicon 2014, 86, 1–7. [Google Scholar] [CrossRef]
- Arbuckle, K.; de la Vega, R.C.R.; Casewell, N.R. Coevolution takes the sting out of it: Evolutionary biology and mechanisms of toxin resistance in animals. Toxicon 2017, 140, 118–131. [Google Scholar] [CrossRef] [PubMed]
- Church, J.E.; Hodgson, W.C. The pharmacological activity of fish venoms. Toxicon 2002, 40, 1083–1093. [Google Scholar] [CrossRef]
- Peiren, N.; Vanrobaeys, F.; de Graaf, D.C.; Devreese, B.; Van Beeumen, J.; Jacobs, F.J. The protein composition of honeybee venom reconsidered by a proteomic approach. Biochim. Biophys. Acta 2005, 1752, 1–5. [Google Scholar] [CrossRef] [PubMed]
- De Graaf, D.C.; Aerts, M.; Danneels, E.; Devreese, B. Bee, wasp and ant venomics pave the way for a component-resolved diagnosis of sting allergy. J. Proteom. 2009, 72, 145–154. [Google Scholar] [CrossRef]
- Da Silva, N.J.; Aird, S.D. Prey specificity, comparative lethality and compositional differences of coral snake venoms. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2001, 128, 425–456. [Google Scholar] [CrossRef]
- Gibbs, H.L.; Mackessy, S.P. Functional basis of a molecular adaptation: Prey-specific toxic effects of venom from Sistrurus rattlesnakes. Toxicon 2009, 53, 672–679. [Google Scholar] [CrossRef]
- Pawlak, J.; Mackessy, S.P.; Fry, B.G.; Bhatia, M.; Mourier, G.; Fruchart-Gaillard, C.; Servent, D.; Ménez, R.; Stura, E.; Ménez, A.; et al. Denmotoxin, a three-finger toxin from the colubrid snake Boiga dendrophila (Mangrove Catsnake) with bird-specific activity. J. Biol. Chem. 2006, 281, 29030–29041. [Google Scholar] [CrossRef]
- Fainzilber, M.; Gordon, D.; Hasson, A.; Spira, M.E.; Zlotkin, E. Mollusc-specific toxins from the venom of Conus textile neovicarius. Eur. J. Biochem. 1991, 202, 589–595. [Google Scholar] [CrossRef]
- Krasnoperov, V.G.; Shamotienko, O.G.; Grishin, E.V. A crustacean-specific neurotoxin from the venom of the black widow spider Latrodectus mactans tredecimguttatus. Bioorg. Khim. 1990, 16, 1567–1569. [Google Scholar]
- Phuong, M.A.; Mahardika, G.N.; Alfaro, M.E. Dietary breadth is positively correlated with venom complexity in cone snails. BMC Genom. 2016, 17, 401. [Google Scholar] [CrossRef]
- Pekár, S.; Bočánek, O.; Michálek, O.; Petráková, L.; Haddad, C.R.; Šedo, O.; Zdráhal, Z. Venom gland size and venom complexity—essential trophic adaptations of venomous predators: A case study using spiders. Mol. Ecol. 2018, 27, 4257–4269. [Google Scholar] [CrossRef] [PubMed]
- Pekár, S.; Líznarová, E.; Bočánek, O.; Zdráhal, Z. Venom of prey-specialized spiders is more toxic to their preferred prey: A result of prey-specific toxins. J. Anim. Ecol. 2018, 87, 1639–1652. [Google Scholar] [CrossRef] [PubMed]
- World Spider Catalog. Version 20.5. Natural History Museum Bern. Available online: http://wsc.nmbe.ch (accessed on 26 September 2019). [CrossRef]
- King, G.F.; Hardy, M.C. Spider-venom peptides: Structure, pharmacology, and potential for control of insect pests. Annu. Rev. Entomol. 2013, 58, 475–496. [Google Scholar] [CrossRef] [PubMed]
- Michálek, O.; Řezáč, M.; Líznarová, E.; Symondson, W.O.; Pekár, S. Silk versus venom: Alternative capture strategies employed by closely related myrmecophagous spiders. Biol. J. Linn. Soc. 2019, 126, 545–554. [Google Scholar] [CrossRef]
- Malli, H.; Kuhn-Nentwig, L.; Imboden, H.; Nentwig, W. Effects of size, motility and paralysation time of prey on the quantity of venom injected by the hunting spider Cupiennius salei. J. Exp. Biol. 1999, 202, 2083–2089. [Google Scholar] [PubMed]
- Kuhn-Nentwig, L.; Schaller, J.; Nentwig, W. Biochemistry, toxicology and ecology of the venom of the spider Cupiennius salei (Ctenidae). Toxicon 2004, 43, 543–553. [Google Scholar] [CrossRef]
- Petráková, L.; Líznarová, E.; Pekár, S.; Haddad, C.R.; Sentenská, L.; Symondson, W.O.C. Discovery of a monophagous true predator, a specialist termite-eating spider (Araneae: Ammoxenidae). Sci. Rep. 2015, 5, 14013. [Google Scholar] [CrossRef]
- Pekár, S.; Coddington, J.A.; Blackledge, T.A. Evolution of stenophagy in spiders (Araneae): Evidence based on the comparative analysis of spider diets. Evolution 2012, 66, 776–806. [Google Scholar] [CrossRef]
- Pekár, S.; Šobotník, J.; Lubin, Y. Armoured spiderman: Morphological and behavioural adaptations of a specialised araneophagous predator (Araneae: Palpimanidae). Naturwissenschaften 2011, 98, 593–603. [Google Scholar] [CrossRef]
- Pekár, S.; Král, J.; Lubin, Y. Natural history and karyotype of some ant-eating zodariid spiders (Araneae, Zodariidae) from Israel. J. Arachnol. 2005, 33, 50–63. [Google Scholar] [CrossRef]
- Pekár, S.; Toft, S.; Hrušková, M.; Mayntz, D. Dietary and prey-capture adaptations by which Zodarion germanicum, an ant-eating spider (Araneae: Zodariidae), specialises on the Formicinae. Naturwissenschaften 2008, 95, 233–239. [Google Scholar] [CrossRef] [PubMed]
- Poran, N.S.; Coss, R.G.; Benjami, E.L.I. Resistance of California ground squirrels (Spermophilus beecheyi) to the venom of the northern Pacific rattlesnake (Crotalus viridis oreganus): A study of adaptive variation. Toxicon 1987, 25, 767–777. [Google Scholar] [CrossRef]
- Smiley-Walters, S.A.; Farrell, T.M.; Gibbs, H.L. The importance of species: Pygmy rattlesnake venom toxicity differs between native prey and related non-native species. Toxicon 2018, 144, 42–47. [Google Scholar] [CrossRef] [PubMed]
- Pekár, S.; Šedo, O.; Líznarová, E.; Korenko, S.; Zdráhal, Z. David and Goliath: Potent venom of an ant-eating spider (Araneae) enables capture of a giant prey. Naturwissenschaften 2014, 101, 533–540. [Google Scholar] [CrossRef]
- Kuhn-Nentwig, L.; Bücheler, A.; Studer, A.; Nentwig, W. Taurine and histamine: Low molecular compounds in prey hemolymph increase the killing power of spider venom. Naturwissenschaften 1998, 85, 136–138. [Google Scholar] [CrossRef]
- Pekár, S.; Toft, S. Can ant-eating Zodarion spiders (Araneae: Zodariidae) develop on a diet optimal for euryphagous arthropod predators? Physiol. Entomol. 2009, 34, 195–201. [Google Scholar] [CrossRef]
- Adams, M.E. Agatoxins: Ion channel specific toxins from the American funnel web spider, Agelenopsis aperta. Toxicon 2004, 43, 509–525. [Google Scholar] [CrossRef]
- Pekár, S. Predatory behavior of two European ant-eating spiders (Araneae, Zodariidae). J. Arachnol. 2004, 32, 31–42. [Google Scholar] [CrossRef]
- Kuhn-Nentwig, L.; Stöcklin, R.; Nentwig, W. Venom composition and strategies in spiders: Is everything possible. In Spider Physiology and Behaviour; Elsevier: London, UK, 2011; Volume 1, pp. 2–86. [Google Scholar]
- Kuzmenkov, A.I.; Sachkova, M.Y.; Kovalchuk, S.I.; Grishin, E.V.; Vassilevski, A.A. Lachesana tarabaevi, an expert in membrane-active toxins. Biochem. J. 2016, 473, 2495–2506. [Google Scholar] [CrossRef]
- Herzig, V.; King, G.F.; Undheim, E.A. Can we resolve the taxonomic bias in spider venom research? Toxicon X 2019, 1, 100005. [Google Scholar] [CrossRef]
- Skejic, J.; Steer, D.L.; Dunstan, N.; Hodgson, W.C. Venoms of related mammal-eating species of taipans (Oxyuranus) and brown snakes (Pseudonaja) differ in composition of toxins involved in mammal poisoning. bioRxiv 2018, 378141. [Google Scholar] [CrossRef]
- Jenner, R.A.; von Reumont, B.M.; Campbell, L.I.; Undheim, E.A. Parallel evolution of complex centipede venoms revealed by comparative proteotranscriptomic analyses. Mol. Biol. Evol. 2019, msz181. [Google Scholar] [CrossRef] [PubMed]
- Chandler, D.; Bailey, A.S.; Tatchell, G.M.; Davidson, G.; Greaves, J.; Grant, W.P. The development, regulation and use of biopesticides for integrated pest management. Philos. Trans. R. Soc. B 2011, 366, 1987–1998. [Google Scholar] [CrossRef] [PubMed]
- Schultz, T.R. In search of ant ancestors. Proc. Natl. Acad. Sci. USA 2000, 97, 14028–14029. [Google Scholar] [CrossRef]
- Vander Meer, R.K.; Jaffe, K.; Cedeno, A. Applied Myrmecology: A World Perspective; Westview Press: Boulder, CO, USA, 1990; pp. 3–70. ISBN 0813377854. [Google Scholar]
- Ward, D.; Lubin, Y. Habitat selection and the life history of a desert spider, Stegodyphus lineatus (Eresidae). J. Anim. Ecol. 1993, 62, 353–363. [Google Scholar] [CrossRef]
- Pompozzi, G.; García, L.F.; Petráková, L.; Pekár, S. Distinct feeding strategies of generalist and specialist spiders. Ecol. Entomol. 2019, 44, 129–139. [Google Scholar] [CrossRef]
- Kuhn-Nentwig, L.; Schaller, J.; Nentwig, W. Purification of toxic peptides and the amino acid sequence of CSTX-1 from the multicomponent venom of Cupiennius salei (Araneae: Ctenidae). Toxicon 1994, 32, 287–302. [Google Scholar] [CrossRef]
- Friedel, T.; Nentwig, W. Immobilizing and lethal effects of spider venoms on the cockroach and the common mealbeetle. Toxicon 1989, 27, 305–316. [Google Scholar] [CrossRef]
- Eggs, B.; Wolff, J.O.; Kuhn-Nentwig, L.; Gorb, S.N.; Nentwig, W. Hunting without a web: How lycosoid spiders subdue their prey. Ethology 2015, 121, 1166–1177. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018; Available online: https://www.R-project.org/ (accessed on 15 March 2018).
- Venables, W.N.; Ripley, B.D. Modern Applied Statistics with S, 4th ed.; Springer: New York, NY, USA, 2002; ISBN 0-387-95457-0. [Google Scholar]
- Smiley-Walters, S.A.; Farrell, T.M.; Gibbs, H.L. Evaluating local adaptation of a complex phenotype: Reciprocal tests of pigmy rattlesnake venoms on treefrog prey. Oecologia 2017, 184, 739–748. [Google Scholar] [CrossRef]
- Halekoh, U.; Højsgaard, S.; Yan, J. The R package geepack for generalized estimating equations. J. Stat. Softw. 2006, 15, 1–11. [Google Scholar] [CrossRef]
- Pekár, S.; Brabec, M. Generalized estimating equations: A pragmatic and flexible approach to the marginal GLM modelling of correlated data in the behavioural sciences. Ethology 2018, 124, 86–93. [Google Scholar] [CrossRef]

| Spider Species | Preferred Prey (Spider/Ant) | Alternative Prey (Cricket) | ||
|---|---|---|---|---|
| LD50 (nL/mg) | ED50 (nL/mg) | LD50 (nL/mg) | ED50 (nL/mg) | |
| Palpimanus sp. (specialist) | 0.07 (0.06, 0.09) | 0.01 (0.00, 0.02) | 3.60 (1.84, 7.02) | 0.13 (0.01, 0.17) |
| Stegodyphus lineatus (generalist) | 8.52 (4.86, 14.95) | 8.66 (4.82, 15.54) | 1.74 (0.96, 3.17) | 0.27 (0.14, 0.53) |
| Zodarion nitidum (specialist) | 0.06 (0.05, 0.06) | 0.07 (0.06, 0.08) | no effect | no effect |
| Cybaeodamus taim (generalist) | 1.82 (1.11, 2.96) | 0.17 (0.07, 0.40) | 0.96 (0.57, 1.62) | 0.25 (0.15, 0.44) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Michálek, O.; Kuhn-Nentwig, L.; Pekár, S. High Specific Efficiency of Venom of Two Prey-Specialized Spiders. Toxins 2019, 11, 687. https://doi.org/10.3390/toxins11120687
Michálek O, Kuhn-Nentwig L, Pekár S. High Specific Efficiency of Venom of Two Prey-Specialized Spiders. Toxins. 2019; 11(12):687. https://doi.org/10.3390/toxins11120687
Chicago/Turabian StyleMichálek, Ondřej, Lucia Kuhn-Nentwig, and Stano Pekár. 2019. "High Specific Efficiency of Venom of Two Prey-Specialized Spiders" Toxins 11, no. 12: 687. https://doi.org/10.3390/toxins11120687
APA StyleMichálek, O., Kuhn-Nentwig, L., & Pekár, S. (2019). High Specific Efficiency of Venom of Two Prey-Specialized Spiders. Toxins, 11(12), 687. https://doi.org/10.3390/toxins11120687