Identification of a Killer Toxin from Wickerhamomyces anomalus with β-Glucanase Activity
Abstract
1. Introduction
2. Results
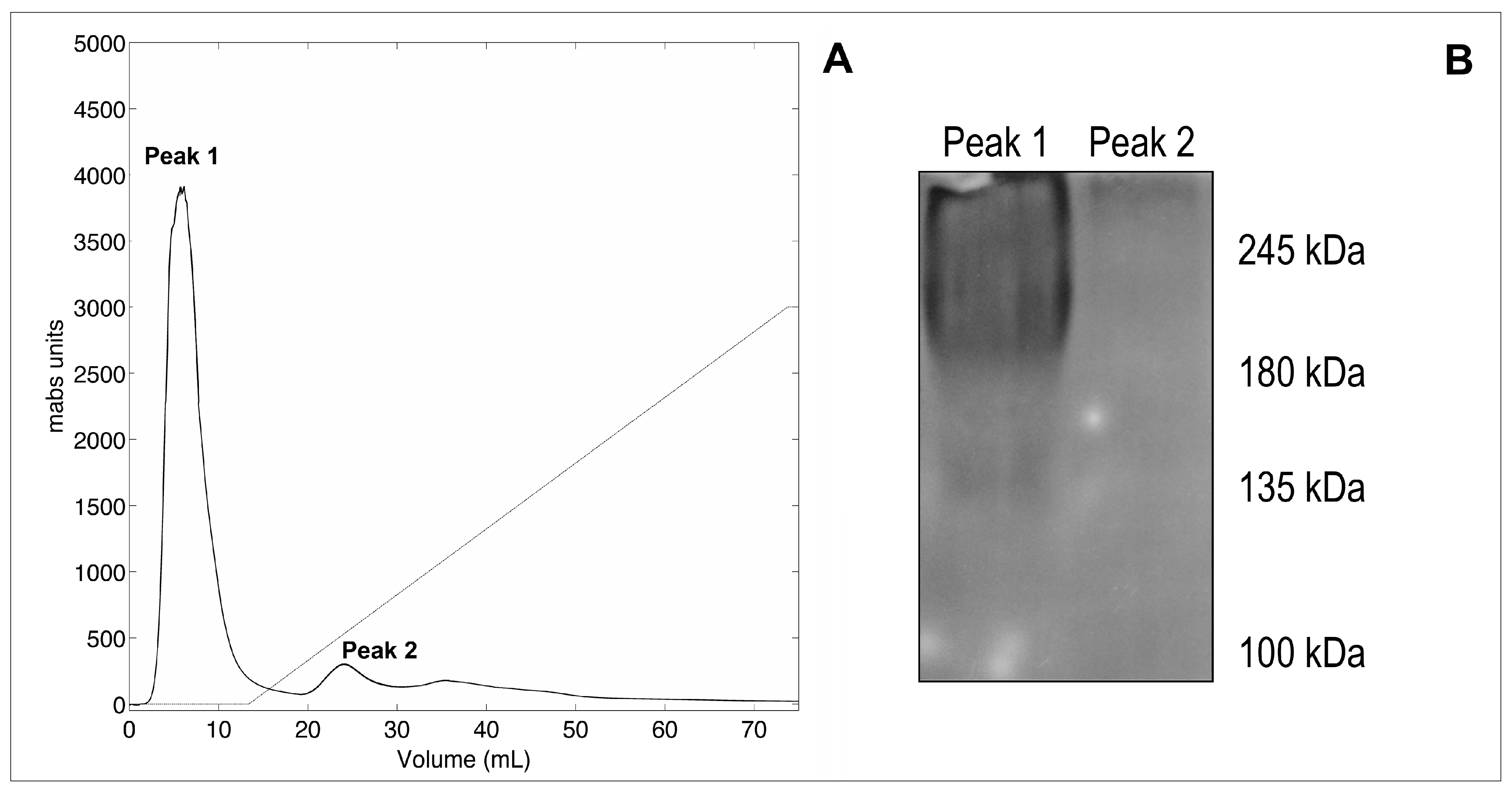
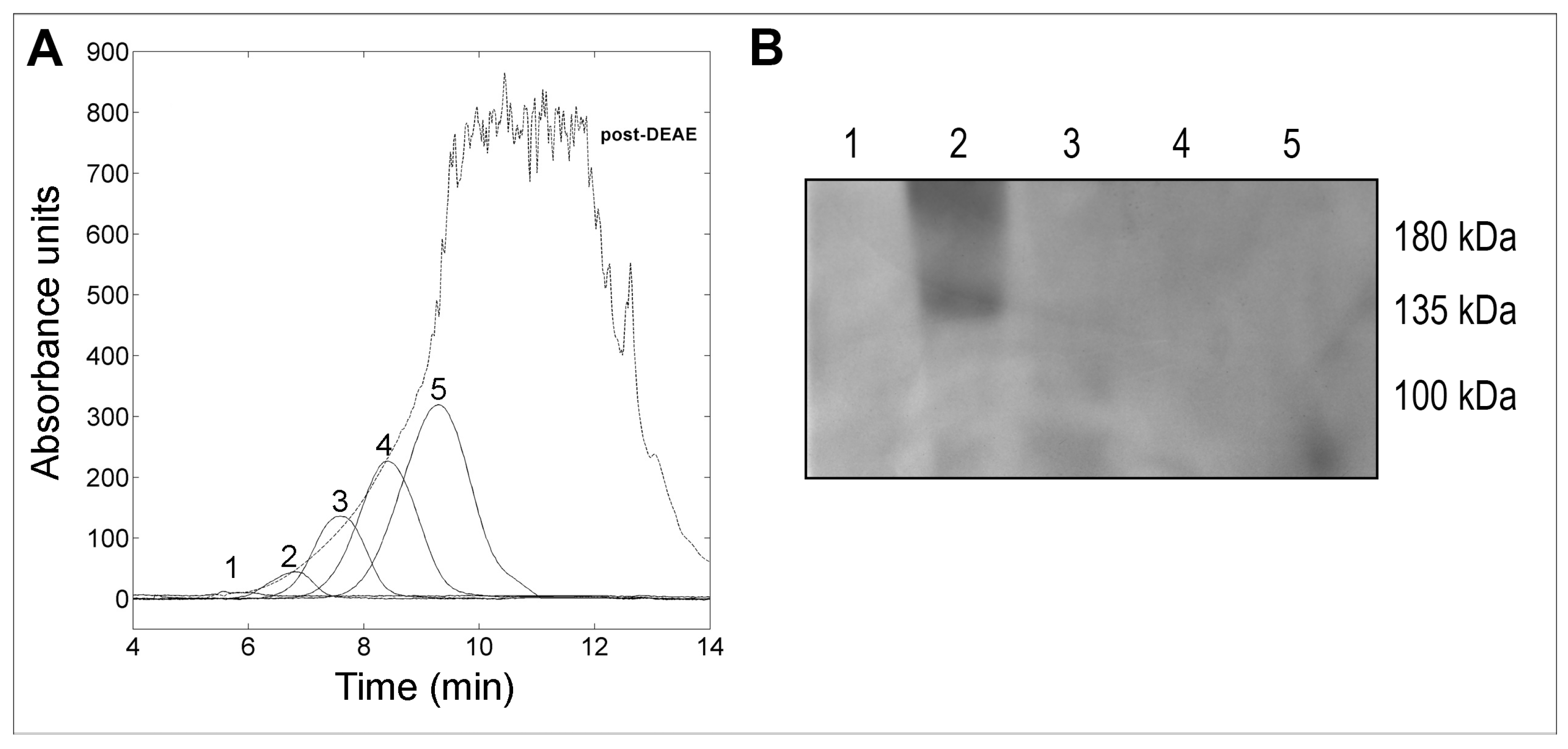
2.1. Purification of the WaF17.12 Killer Toxin
2.2. Identification of the Protein Toxin with Yeast Killing Activity
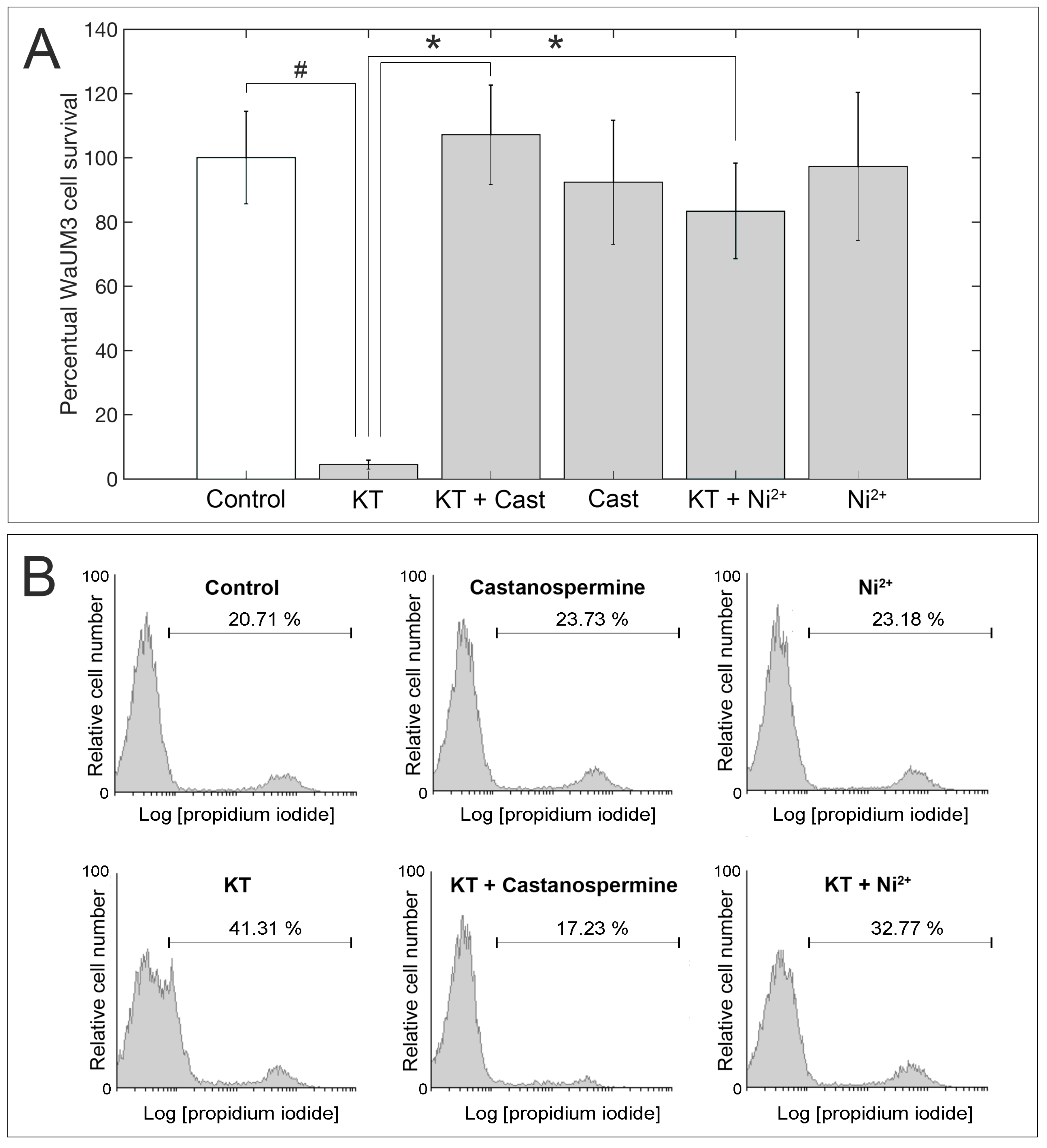
2.3. Evaluation of KT Activity
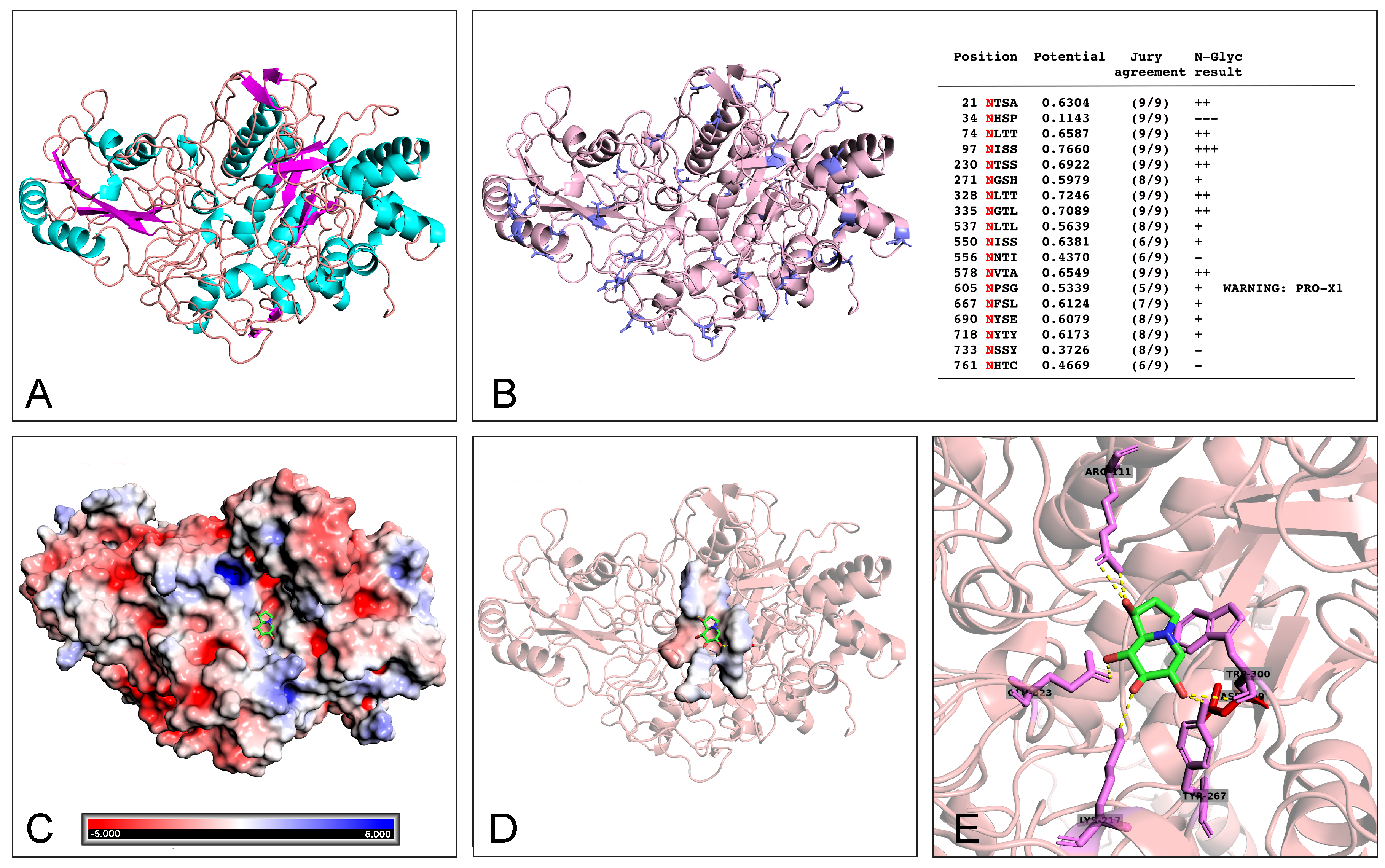
2.4. Docking Analysis of Castanospermine to W. anomalus β-Glucanase
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Yeast Strains and KT Production
4.3. Sample Processing and Concentration
4.4. Anion Exchange Chromatography
4.5. Gel Filtration Chromatography
4.6. Western Blotting
4.7. Sample Preparation for LC−MS/MS
4.8. LC–MS/MS Analysis
4.9. Activity Assays
4.10. Prediction of Three-Dimensional Structures of WaF17.12 β-Glucanase
4.11. β-Glucanase-Castanospermine Docking Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Boynton, P.J. The ecology of killer yeasts: Interference competition in natural habitats. Yeast 2019. [Google Scholar] [CrossRef]
- Schmitt, M.J.; Breinig, F. The viral killer system in yeast: From molecular biology to application. FEMS Microbiol. Rev. 2002, 26, 257–276. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Chi, Z.M.; Wang, X.H.; Li, J. Purification and molecular characterization of exo-beta-1,3-glucanases from the marine yeast Williopsis saturnus WC91-2. Appl. Microbiol. Biotechnol. 2009, 85, 85–94. [Google Scholar] [CrossRef] [PubMed]
- Izgu, F.; Altinbay, D.; Tureli, A.E. In vitro activity of panomycocin, a novel exo-beta-1,3-glucanase isolated from Pichia anomala NCYC 434, against dermatophytes. Mycoses 2007, 50, 31–34. [Google Scholar] [CrossRef] [PubMed]
- De Ullivarri, M.F.; Mendoza, L.M.; Raya, R.R. Killer activity of Saccharomyces cerevisiae strains: Partial characterization and strategies to improve the biocontrol efficacy in winemaking. Antonie van Leeuwenhoek 2014, 106, 865–878. [Google Scholar] [CrossRef] [PubMed]
- Comitini, F.; De Ingeniis, J.; Pepe, L.; Mannazzu, I.; Ciani, M. Pichia anomala and Kluyveromyces wickerhamii killer toxins as new tools against Dekkera/Brettanomyces spoilage yeasts. FEMS Microbiol. Lett. 2004, 238, 235–240. [Google Scholar] [CrossRef] [PubMed]
- Walker, G.M. Pichia anomala: Cell physiology and biotechnology relative to other yeasts. Antonie van Leeuwenhoek 2011, 99, 25–34. [Google Scholar] [CrossRef] [PubMed]
- Ricci, I.; Damiani, C.; Scuppa, P.; Mosca, M.; Crotti, E.; Rossi, P.; Rizzi, A.; Capone, A.; Gonella, E.; Ballarini, P.; et al. The yeast Wickerhamomyces anomalus (Pichia anomala) inhabits the midgut and reproductive system of the Asian malaria vector Anopheles stephensi. Environ. Microbiol. 2011, 13, 911–921. [Google Scholar] [CrossRef] [PubMed]
- Cappelli, A.; Ulissi, U.; Valzano, M.; Damiani, C.; Epis, S.; Gabrielli, M.G.; Conti, S.; Polonelli, L.; Bandi, C.; Favia, G.; et al. A Wickerhamomyces anomalus killer strain in the malaria vector Anopheles stephensi. PLoS ONE 2014, 9, e95988. [Google Scholar] [CrossRef]
- Valzano, M.; Cecarini, V.; Cappelli, A.; Capone, A.; Bozic, J.; Cuccioloni, M.; Epis, S.; Petrelli, D.; Angeletti, M.; Eleuteri, A.M.; et al. A yeast strain associated to Anopheles mosquitoes produces a toxin able to kill malaria parasites. Malar. J. 2016, 15, 21. [Google Scholar] [CrossRef] [PubMed]
- Cappelli, A.; Valzano, M.; Cecarini, V.; Bozic, J.; Rossi, P.; Mensah, P.; Amantini, C.; Favia, G.; Ricci, I. Killer yeasts exert anti-plasmodial activities against the malaria parasite Plasmodium berghei in the vector mosquito Anopheles stephensi and in mice. Parasit Vectors 2019, 12, 329. [Google Scholar] [CrossRef] [PubMed]
- Polonelli, L.; Magliani, W.; Ciociola, T.; Giovati, L.; Conti, S. From Pichia anomala killer toxin through killer antibodies to killer peptides for a comprehensive anti-infective strategy. Antonie van Leeuwenhoek 2011, 99, 35–41. [Google Scholar] [CrossRef] [PubMed]
- Bause, E.; Legler, G. Isolation and structure of a tryptic glycopeptide from the active site of beta-glucosidase A3 from Aspergillus wentii. Biochim. Biophys. Acta 1980, 626, 459–465. [Google Scholar] [CrossRef]
- Mannazzu, I.; Domizio, P.; Carboni, G.; Zara, S.; Zara, G.; Comitini, F.; Budroni, M.; Ciani, M. Yeast killer toxins: From ecological significance to application. Crit. Rev. Biotechnol. 2019, 39, 603–617. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.L.; Chi, Z.; Wang, G.Y.; Wang, Z.P.; Li, Y.; Chi, Z.M. Yeast killer toxins, molecular mechanisms of their action and their applications. Crit. Rev. Biotechnol. 2015, 35, 222–234. [Google Scholar] [CrossRef] [PubMed]
- Giovati, L.; Santinoli, C.; Ferrari, E.; Ciociola, T.; Martin, E.; Bandi, C.; Ricci, I.; Epis, S.; Conti, S. Candidacidal Activity of a Novel Killer Toxin from Wickerhamomyces anomalus against Fluconazole-Susceptible and -Resistant Strains. Toxins 2018, 10. [Google Scholar] [CrossRef]
- Guo, F.J.; Ma, Y.; Xu, H.M.; Wang, X.H.; Chi, Z.M. A novel killer toxin produced by the marine-derived yeast Wickerhamomyces anomalus YF07b. Antonie van Leeuwenhoek 2013, 103, 737–746. [Google Scholar] [CrossRef]
- Westermeier, R. Electrophoresis in Practice: A Guide to Methods and Applications of DNA and Protein Separations, 4th ed.; John Wiley & Sons: Hoboken, NJ, USA, 2004. [Google Scholar]
- Hames, B.D. Gel Electrophoresis of Proteins. A Practical Approach, 3rd ed.; Oxford University Press: Oxford, UK, 1998. [Google Scholar]
- Heinig, M.; Frishman, D. STRIDE: A web server for secondary structure assignment from known atomic coordinates of proteins. Nucleic Acids Res. 2004, 32, W500–W502. [Google Scholar] [CrossRef]
- Saul, R.; Molyneux, R.J.; Elbein, A.D. Studies on the mechanism of castanospermine inhibition of alpha- and beta-glucosidases. Arch. Biochem. Biophys. 1984, 230, 668–675. [Google Scholar] [CrossRef]
- Guyard, C.; Dehecq, E.; Tissier, J.P.; Polonelli, L.; Dei-Cas, E.; Cailliez, J.C.; Menozzi, F.D. Involvement of [beta]-glucans in the wide-spectrum antimicrobial activity of Williopsis saturnus var. mrakii MUCL 41968 killer toxin. Mol. Med. 2002, 8, 686–694. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Cecarini, V.; Bonfili, L.; Amici, M.; Angeletti, M.; Keller, J.N.; Eleuteri, A.M. Amyloid peptides in different assembly states and related effects on isolated and cellular proteasomes. Brain Res. 2008, 1209, 8–18. [Google Scholar] [CrossRef] [PubMed]
- Geisslitz, S.; Ludwig, C.; Scherf, K.A.; Koehler, P. Targeted LC-MS/MS Reveals Similar Contents of alpha-Amylase/Trypsin-Inhibitors as Putative Triggers of Nonceliac Gluten Sensitivity in All Wheat Species except Einkorn. J. Agric. Food Chem. 2018, 66, 12395–12403. [Google Scholar] [CrossRef] [PubMed]
- Perkins, D.N.; Pappin, D.J.; Creasy, D.M.; Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 1999, 20, 3551–3567. [Google Scholar] [CrossRef]
- Kara, H.E.; Sinan, S.; Turan, Y. Purification of beta-glucosidase from olive (Olea europaea L.) fruit tissue with specifically designed hydrophobic interaction chromatography and characterization of the purified enzyme. J. Chromatogr. B 2011, 879, 1507–1512. [Google Scholar] [CrossRef] [PubMed]
- Hayase, M.; Maekawa, A.; Yubisui, T.; Minami, Y. Properties, intracellular localization, and stage-specific expression of membrane-bound beta-glucosidase, BglM1, from Physarum polycephalum. Int. J. Biochem. Cell Biol. 2008, 40, 2141–2150. [Google Scholar] [CrossRef]
- Comitini, F.; Mannazzu, I.; Ciani, M. Tetrapisispora phaffii killer toxin is a highly specific beta-glucanase that disrupts the integrity of the yeast cell wall. Microb. Cell Factories 2009, 8, 55. [Google Scholar] [CrossRef]
- Cuccioloni, M.; Mozzicafreddo, M.; Bonfili, L.; Cecarini, V.; Giangrossi, M.; Falconi, M.; Saitoh, S.I.; Eleuteri, A.M.; Angeletti, M. Interfering with the high-affinity interaction between wheat amylase trypsin inhibitor CM3 and toll-like receptor 4: In silico and biosensor-based studies. Sci. Rep. 2017, 7, 13169. [Google Scholar] [CrossRef]
- Petersen, T.N.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER Suite: Protein structure and function prediction. Nat. Methods 2015, 12, 7–8. [Google Scholar] [CrossRef]
- Xu, D.; Zhang, Y. Improving the physical realism and structural accuracy of protein models by a two-step atomic-level energy minimization. Biophys. J. 2011, 101, 2525–2534. [Google Scholar] [CrossRef] [PubMed]
- Van Gunsteren, W.F.; Billeter, S.R.; Eising, A.A.; Hünenberger, P.H.; Krüger, P.K.; Mark, A.E.; Scott, W.R.; Tironi, I.G. Biomolecular Simulation: The GROMOS 96 Manual and User Guide; Vdf Hochschulverlag AG an der ETH Zürich: Zürich, Switzerland, 1996. [Google Scholar]
- Laskowski, R.A.; MacArthur, M.W.; Thornton, J.M. PROCHECK: Validation of Protein Structure Coordinates; Rossmann, M.G., Arnold, E.D., Eds.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2001. [Google Scholar]
- Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; Shoemaker, B.A.; Thiessen, P.A.; Yu, B.; et al. PubChem 2019 update: Improved access to chemical data. Nucleic Acids Res. 2019, 47, D1102–D1109. [Google Scholar] [CrossRef] [PubMed]
- Schneidman-Duhovny, D.; Inbar, Y.; Nussinov, R.; Wolfson, H.J. PatchDock and SymmDock: Servers for rigid and symmetric docking. Nucleic Acids Res. 2005, 33, W363–W367. [Google Scholar] [CrossRef]
- Andrusier, N.; Nussinov, R.; Wolfson, H.J. FireDock: Fast interaction refinement in molecular docking. Proteins 2007, 69, 139–159. [Google Scholar] [CrossRef] [PubMed]
| Observed (M2+) | Mr (Expected) | Mr (Calculated) | Missed Cleavage | Peptide |
|---|---|---|---|---|
| 753.39 | 1,504.7654 | 1,504.7555 | 1 | ARELVDQMSIAEK + Oxidation (M) |
| 987.02 | 1,972.0254 | 1,972.0200 | 1 | GADAILGPVYGPMGVKAAGGR + Oxidation (M) |
| 637.83 | 1,273.6454 | 1,273.6514 | 0 | ISILGQAAGDDSK |
| 1,088.02 | 2,174.0254 | 2,174.0386 | 0 | VNLTTGVGSASGPCSGNTGSVPR |
| 1,456.16 | 2,910.3054 | 2,910.3243 | 0 | GCGSGAIGTGYGSGAGTFSYFVTPADGIGAR |
| Global Energy | aVdW | rVdW | ACE | Inside |
|---|---|---|---|---|
| −19.49 | −11.04 | 1.14 | −4.42 | 4.62 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cecarini, V.; Cuccioloni, M.; Bonfili, L.; Ricciutelli, M.; Valzano, M.; Cappelli, A.; Amantini, C.; Favia, G.; Eleuteri, A.M.; Angeletti, M.; et al. Identification of a Killer Toxin from Wickerhamomyces anomalus with β-Glucanase Activity. Toxins 2019, 11, 568. https://doi.org/10.3390/toxins11100568
Cecarini V, Cuccioloni M, Bonfili L, Ricciutelli M, Valzano M, Cappelli A, Amantini C, Favia G, Eleuteri AM, Angeletti M, et al. Identification of a Killer Toxin from Wickerhamomyces anomalus with β-Glucanase Activity. Toxins. 2019; 11(10):568. https://doi.org/10.3390/toxins11100568
Chicago/Turabian StyleCecarini, Valentina, Massimiliano Cuccioloni, Laura Bonfili, Massimo Ricciutelli, Matteo Valzano, Alessia Cappelli, Consuelo Amantini, Guido Favia, Anna Maria Eleuteri, Mauro Angeletti, and et al. 2019. "Identification of a Killer Toxin from Wickerhamomyces anomalus with β-Glucanase Activity" Toxins 11, no. 10: 568. https://doi.org/10.3390/toxins11100568
APA StyleCecarini, V., Cuccioloni, M., Bonfili, L., Ricciutelli, M., Valzano, M., Cappelli, A., Amantini, C., Favia, G., Eleuteri, A. M., Angeletti, M., & Ricci, I. (2019). Identification of a Killer Toxin from Wickerhamomyces anomalus with β-Glucanase Activity. Toxins, 11(10), 568. https://doi.org/10.3390/toxins11100568










