Zearalenone Biodegradation by the Combination of Probiotics with Cell-Free Extracts of Aspergillus oryzae and Its Mycotoxin-Alleviating Effect on Pig Production Performance
Abstract
1. Introduction
2. Results
2.1. The Probiotics for ZEA Degradation
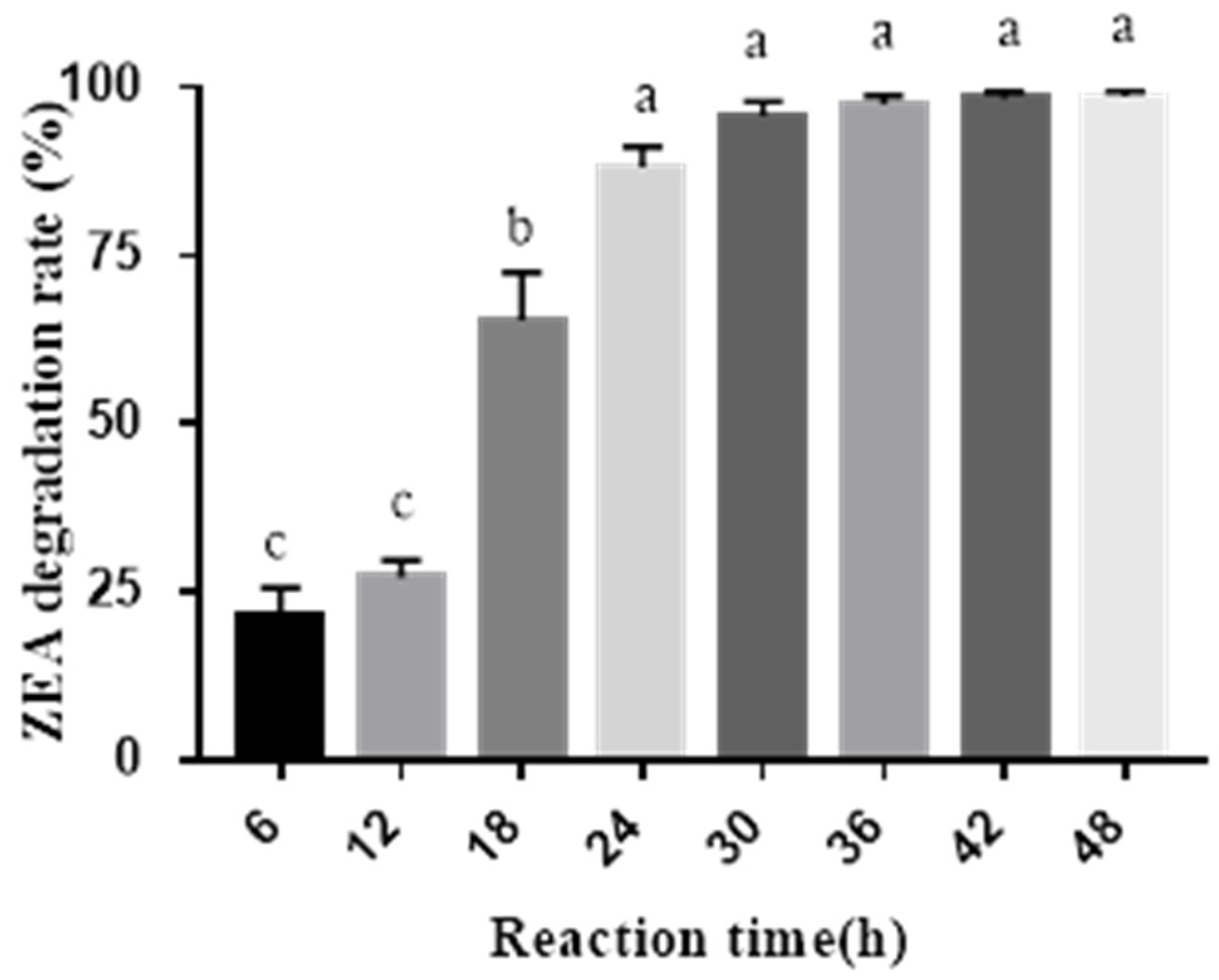
2.2. ZEA Degradation by the Combined Probiotics and Cell-Free Extracts of A. oryzae
2.3. Effect of MBP on Pig Growth Performance and Nutrient Digestibility
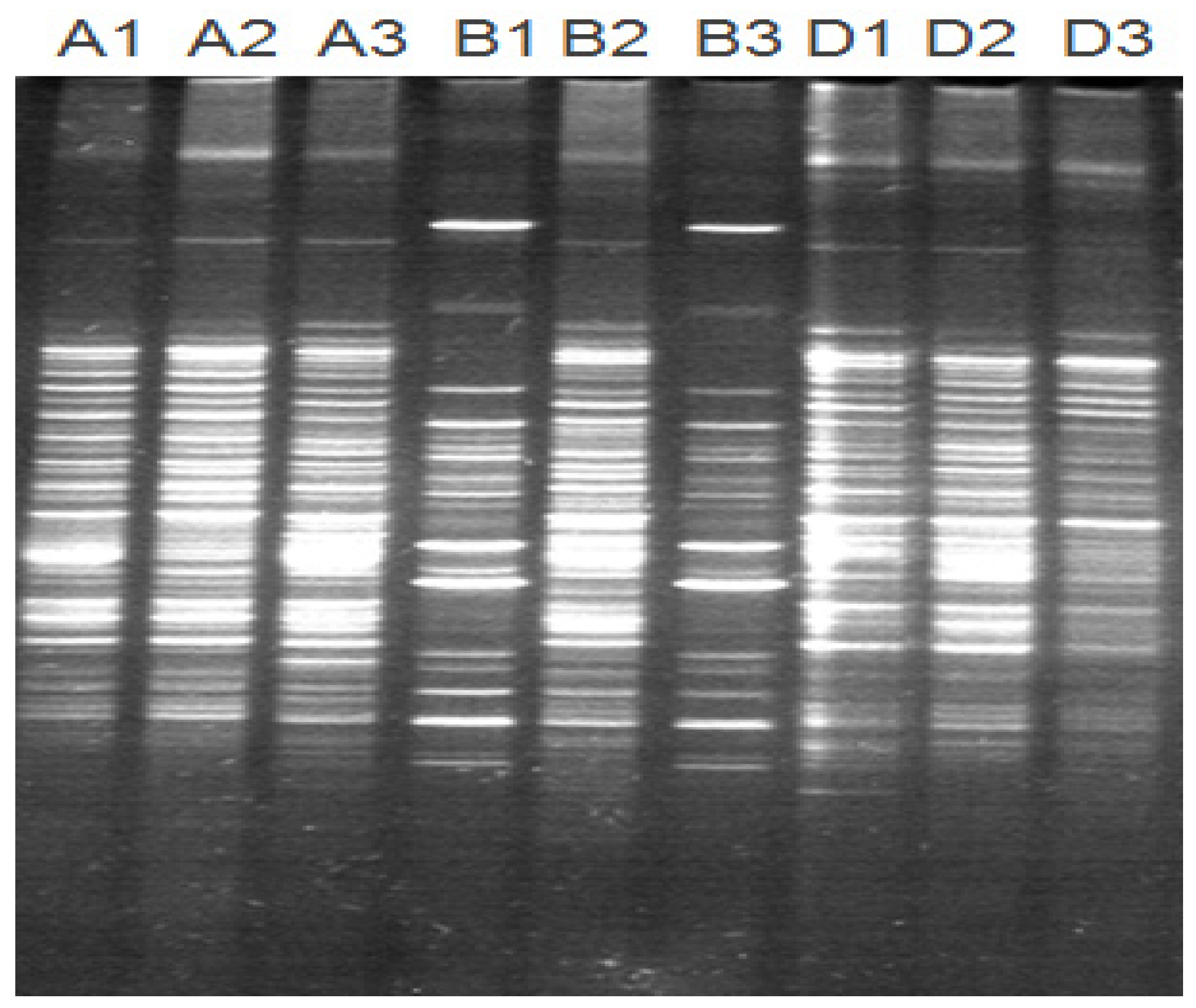
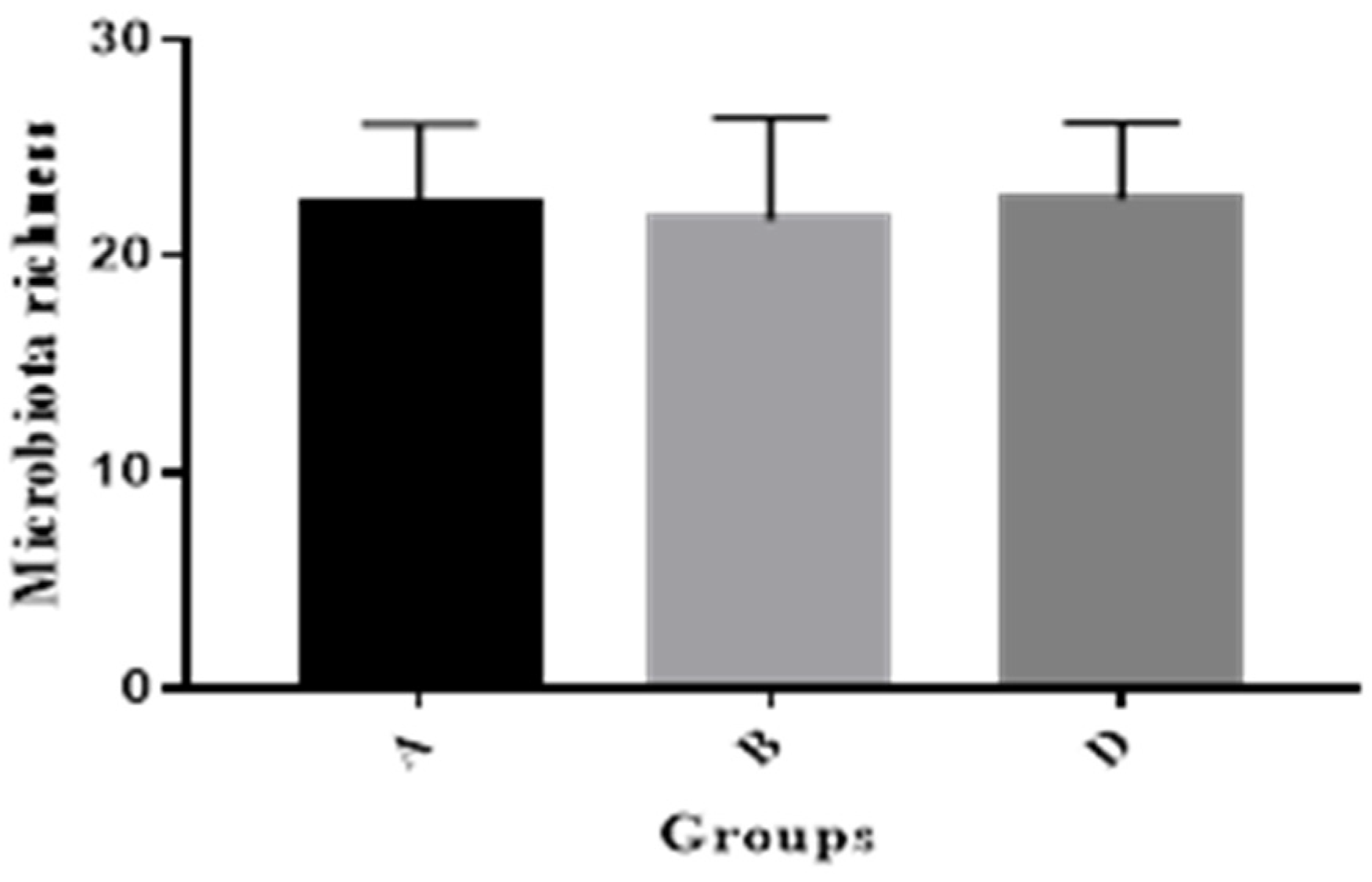
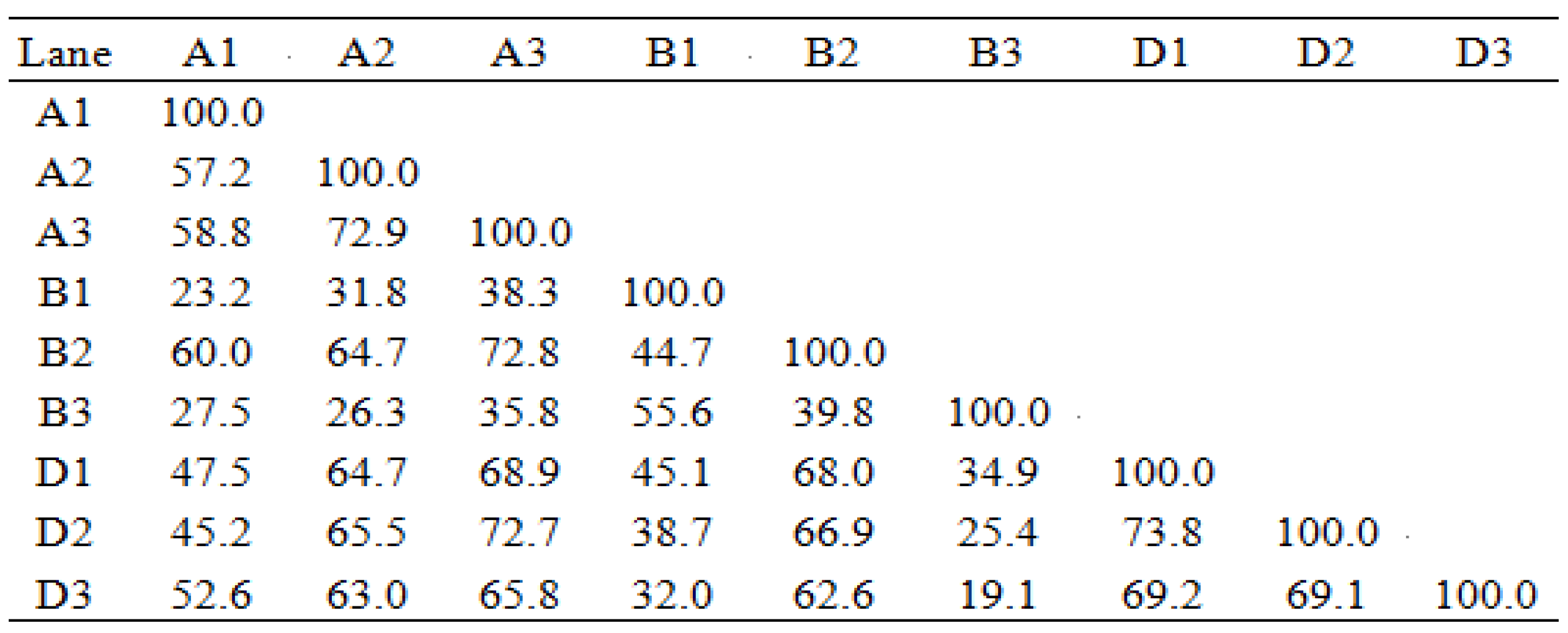
2.4. Pig Gut Microbiota Affected by MBP
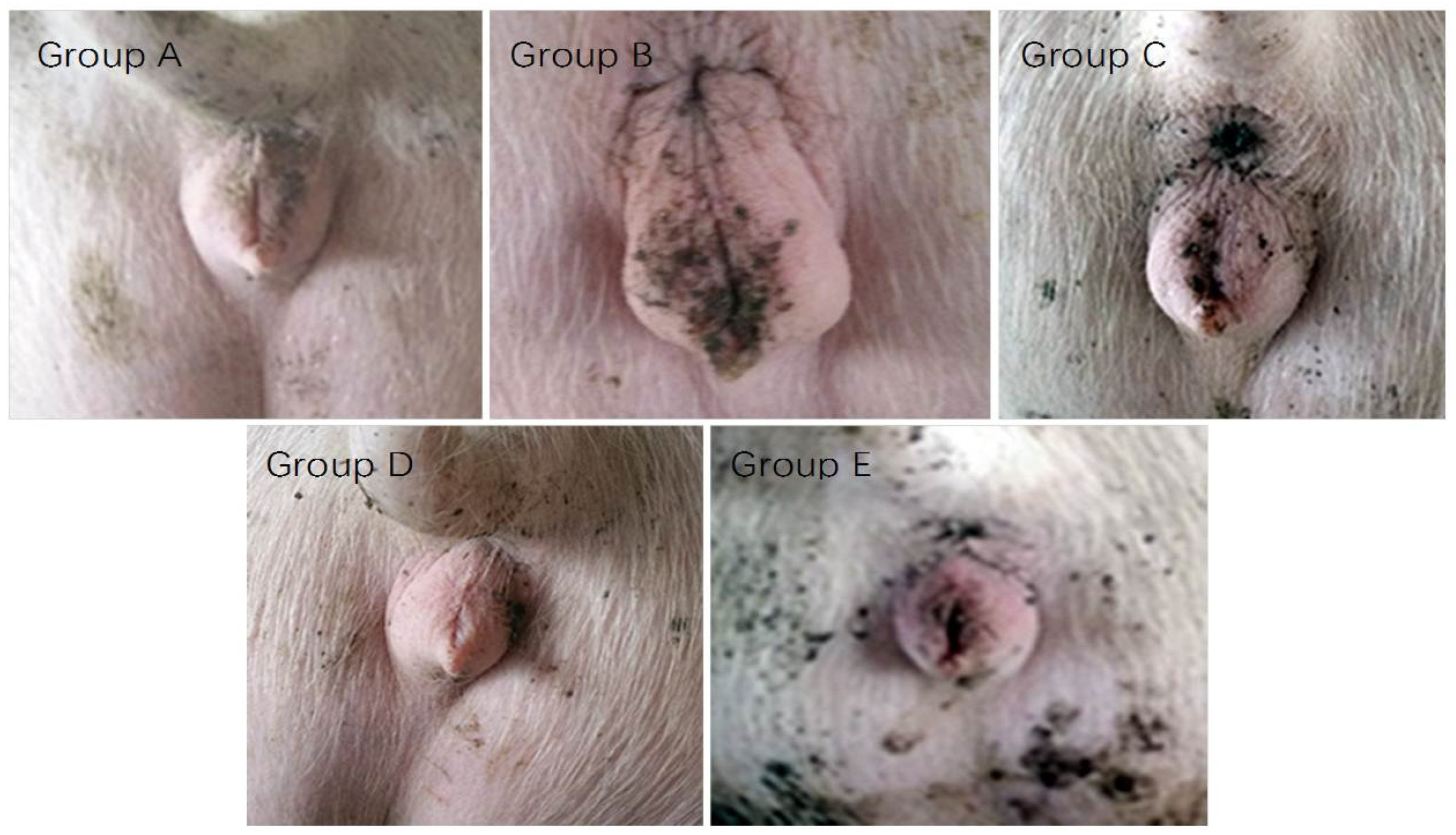
2.5. The Vulvar Area and Serum Parameters
2.6. The Relative Organ Weight and ERα mRNA Abundance in Ovaries and the Uterus
2.7. ZEA Concentrations in Serum, Relative Tissues, and Gut
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Probiotics and Experimental Materials
5.2. Degradation of ZEA by Probiotic Incubation
5.3. Preparation of Cell-Free Extracts from A. oryzae
5.4. ZEA Degradation by Probiotics Combined with Cell-Free Extracts of A. oryzae
5.5. Animals and Management
- Group A: Basal diet with 86.19 μg/kg ZEA
- Group B: Basal diet containing 300.00 μg/kg ZEA
- Group C: Basal diet containing 300.00 μg/kg ZEA and 0.05% MBP
- Group D: Basal diet containing 300.00 μg/kg ZEA and 0.10% MBP
- Group E: Basal diet containing 300.00 μg/kg ZEA and 0.15% MBP
5.6. Nutrient Digestibility Measurement
5.7. Vulvar Area Measurement of Gilts
5.8. Serum Parameter Determination
5.9. Relative Organ Weight and ERα mRNA Abundance in Ovaries and the Uterus
5.10. DNA Extraction of Gut Bacteria and PCR-DGGE
5.11. Determination of ZEA Content in Serum and Tissues
5.12. Statistical Analyses
Author Contributions
Funding
Conflicts of Interest
References
- Caldwell, R.W.; Tuite, J.; Stob, M.; Baldwin, R. Zearalenone production by Fusarium species. Appl. Microbiol. 1970, 20, 31–34. [Google Scholar] [PubMed]
- Takemura, H.; Shim, J.Y.; Sayama, K.; Tsubura, A.; Zhu, B.T.; Shimoi, K. Characterization of the estrogenic activities of zearalenone and zeranol in vivo and in vitro. J. Steroid Biochem. Mol. Biol. 2007, 103, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Gajecka, M. The effect of low-dose experimental zearalenone intoxication on the immunoexpression of estrogen receptors in the ovaries of pre-pubertal bitches. Pol. J. Vet. Sci. 2012, 15, 685–691. [Google Scholar] [CrossRef] [PubMed]
- Benzoni, E.; Minervini, F.; Giannoccaro, A.; Fornelli, F.; Vigo, D.; Visconti, A. Influence of in vitro exposure to mycotoxin zearalenone and its derivatives on swine sperm quality. Reprod. Toxicol. 2008, 25, 461–467. [Google Scholar] [CrossRef] [PubMed]
- Kuiper, G.G.J.M.; Lemmen, J.G.; Carlsson, B.; Corton, J.C.; Safe, S.H.; van der Saag, P.T.; van der Burg, B.; Gustafsson, J.A. Interaction of estrogenic chemicals and phytoestrogens with estrogen receptor β. Endocrinology 1998, 139, 4252–4263. [Google Scholar] [CrossRef]
- Aiko, V.; Mehta, A. Occurrence, detection and detoxification of mycotoxins. Toxicol. Appl. Pharm. 2015, 50, 943–954. [Google Scholar] [CrossRef] [PubMed]
- Kosicki, R.; Blajet-Kosicka, A.; Grajewski, J.; Twaruzek, M. Multiannual mycotoxin survey in feed materials and feedstuffs. Anim. Feed Sci. Technol. 2016, 215, 165–180. [Google Scholar] [CrossRef]
- Phuong, N.H.; Thieu, N.Q.; Ogle, B.; Pettersson, H. Aflatoxins, fumonisins and zearalenone contamination of maize in the southeastern and central highlands provinces of Vietnam. Agriculture 2015, 5, 1195–1203. [Google Scholar] [CrossRef]
- Oliveiraa, M.S.; Rochaa, A.; Sulyok, M.; Krskab, R.; Mallmann, C.A. Natural mycotoxin contamination of maize (Zea mays L.) in the south region of Brazil. Food Control 2017, 73, 127–132. [Google Scholar] [CrossRef]
- Lee, H.J.; Ryu, D. Worldwide occurrence of mycotoxins in cereals and cereal-derived food products: Public health perspectives of their co-occurrence. J. Agric. Food Chem. 2017, 65, 7034–7051. [Google Scholar] [CrossRef]
- FAO. Worldwide Regulations for Mycotoxins in Food and Feed in 2003; FAO Food and Nutrition Paper No. 81; Food and Agriculture Organization of the United Nations: Rome, Italy, 2004. [Google Scholar]
- Zinedine, A.; Soriano, J.M.; Molto, J.C.; Manes, J. Review on the toxicity, occurrence, metabolism, detoxification, regulations and intake of zearalenone: An oestrogenic mycotoxin. Food Chem. Toxicol. 2007, 45, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Bordini, J.G.; Borsato, D.; Oliveira, A.S.; Ono, M.A.; Zaninelli, T.H.; Hirooka, E.Y.; Ono, E.Y.S. In vitro zearalenone adsorption by a mixture of organic and inorganic adsorbents: Application of the Box Behnken approach. World Mycotoxin J. 2015, 8, 291–299. [Google Scholar] [CrossRef]
- Markovic, M.; Dakovic, A.; Rottinghaus, G.E.; Kragovic, M.; Petkovic, A.; Krajisnik, D.; Milic, J.; Mercurio, M.; de Gennaro, B. Adsorption of the mycotoxin zearalenone by clinoptilolite and phillipsite zeolites treated with cetylpyridinium surfactant. Colloid Surf. B 2017, 151, 324–332. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wang, Y.; Shao, H.L.; Luo, X.H.; Wang, R.; Li, Y.F.; Li, Y.N.; Luo, Y.P.; Zhang, D.J.; Chen, Z.X. In vivo toxicity assessment of deoxynivalenol-contaminated wheat after ozone degradation. Food Addit. Contam. A 2017, 34, 103–112. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.S.; Qiu, L.P.; Wu, H.; Tang, Y.Q.; Yu, Y.G.; Li, X.F.; Liu, D.M. Degradation of zearalenone by the extracellular extracts of Acinetobacter sp. SM04 liquid cultures. Mycotoxin Biodegr. 2011, 22, 613–622. [Google Scholar] [CrossRef] [PubMed]
- Kriszt, R.; Krifaton, C.; Szoboszlay, S.; Cserhati, M.; Kriszt, B.; Kukolya, J.; Czeh, A.; Feher-Toth, S.; Torok, L.; Szoke, Z.; et al. A new zearalenone mycotoxin biodegradation strategy using non-pathogenic Rhodococcus pyridinivorans K408 strain. PLoS ONE 2012, 7, e43608. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.Q.; Xiao, J.M.; Chen, Y.; Yu, Y.G.; Xiao, X.L.; Yu, Y.S.; Wu, H. Secretory expression and characterization of a novel peroxiredoxin for zearalenone detoxification in Saccharomyces cerevisiae. Microbiol. Res. 2013, 168, 6–11. [Google Scholar] [CrossRef]
- Lei, Y.P.; Zhao, L.H.; Ma, Q.G.; Zhang, J.Y.; Zhou, T.; Gao, C.Q.; Ji, C. Degradation of zearalenone in swine feed and feed ingredients by Bacillus subtilis ANSB01G. World Mycotoxin J. 2014, 7, 143–151. [Google Scholar] [CrossRef]
- Zhao, L.; Jin, H.T.; Lan, J.; Zhang, R.Y.; Ren, H.B.; Zhang, X.B.; Yu, G.P. Detoxification of zearalenone by three strains of Lactobacillus plantarum from fermented food in vitro. Food Control 2015, 54, 158–164. [Google Scholar] [CrossRef]
- Guo, J.; Zhang, L.S.; Wang, Y.M.; Yan, C.H.; Huang, W.P.; Wu, J.; Yuan, H.T.; Lin, B.W.; Shen, J.L.; Peng, S.Q. Study of embryo toxicity of Fusarium mycotoxin butenolide using a whole rat embryo culture model. Toxicol. In Vitro 2011, 25, 1727–1732. [Google Scholar] [CrossRef]
- Wang, M.; Yin, L.; Hu, H.; Selvaraj, J.N.; Zhou, Y.; Zhang, G. Expression, functional analysis and mutation of a novel neutral zearalenone-degrading enzyme. Int. J. Biol. Macromol. 2018, 118, 1284–1292. [Google Scholar] [CrossRef] [PubMed]
- Pereyra, C.M.; Cavaglieri, L.R.; Chiacchiera, S.M.; Dalcero, A. The corn influence on the adsorption levels of aflatoxin B1 and zearalenone by yeast cell wall. J. Appl. Microbiol. 2013, 114, 655–662. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Wu, H.; Tang, Y.; Qiu, L. Cloning, expression of a peroxiredoxin gene from Acinetobacter sp. SM04 and characterization of its recombinant protein for zearalenone detoxification. Microbiol. Res. 2012, 167, 121–126. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Dong, M.; Yang, Q.; Apaliya, M.T.; Li, J.; Zhang, X. Biodegradation of zearalenone by Saccharomyces cerevisiae: Possible involvement of ZEN responsive proteins of the yeast. J. Proteom. 2016, 143, 416–423. [Google Scholar] [CrossRef]
- Cho, K.J.; Kang, J.S.; Cho, W.T.; Lee, C.H.; Ha, J.K.; Bin, S.K. In vitro degradation of zearalenone by Bacillus subtilis. Biotechnol. Lett. 2010, 32, 1921–1924. [Google Scholar] [CrossRef]
- El-Sharkawy, S.; Abul-Hajj, Y.J. Microbial cleavage of zearalenone. Xenobiotica 1988, 18, 365–371. [Google Scholar] [CrossRef] [PubMed]
- Lei, Y.P. Mechanism of Degrading ZEA by ANSB01G Strain and Its Application Effect in Animal Production. Ph.D. Thesis, China Agricultural University, Beijing, China, 2014. [Google Scholar]
- Brodehl, A.; Moller, A.; Kunte, H.J.; Koch, M.; Maul, R. Biotransformation of the mycotoxin zearalenone by fungi of the genera Rhizopus and Aspergillus. FEMS Micobiol. Lett. 2014, 359, 124–130. [Google Scholar] [CrossRef] [PubMed]
- Tran, S.T.; Smith, T.K. Conjugation of deoxynivalenol by Alternaria alternate (54028 NRRL), Rhizopusmicrosporus var. rhizopodiformis (54029 NRRL) and Aspergillus oryzae (5509 NRRL). Mycotoxin Res. 2014, 30, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Zuo, R.Y.; Chang, J.; Yin, Q.Q.; Wang, P.; Yang, Y.R.; Wang, X.; Wang, G.Q.; Zheng, Q.H. Effect of the combined probiotics with aflatoxin B1-degrading enzyme on aflatoxin detoxification, broiler production performance and hepatic enzyme gene expression. Food Chem. Toxicol. 2013, 59, 470–475. [Google Scholar] [CrossRef]
- Jiang, S.; Yang, Z.; Yang, W.; Gao, J.; Liu, F.; Chen, C.; Chi, F. Physiopathological effects of zearalenone in post-weaning female piglets with or without montmorillonite clay adsorbent. Livest. Sci. 2010, 131, 130–136. [Google Scholar] [CrossRef]
- Oliver, W.T.; Miles, J.R.; Diaz, D.E.; Dibner, J.J.; Rottinghaus, G.E.; Harrell, R.J. Zearalenone enhances reproductive tract development, but does not alter skeletal muscle signaling in prepubertal gilts. Anim. Feed Sci. Technol. 2012, 174, 79–85. [Google Scholar] [CrossRef]
- Brousseau, J.P.; Talbot, G.; Beaudoin, F.; Lauzon, K.; Roy, D.; Lessard, M. Effects of probiotics Pediococcus acidilactici strain MA18/5M and Saccharomyces cerevisiae subsp. boulardii strain SB-CNCM I-1079 on fecal and intestinal microbiota of nursing and weanling piglets. J. Anim. Sci. 2015, 93, 5313–5326. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.F.; Zhou, H.L.; Hou, G.Y.; Qi, D.S.; Zhang, N.Y. Soybean isoflavone reduces the residue of zearalenone in the muscle and liver of prepubertal gilts. Animal 2013, 7, 699–703. [Google Scholar] [CrossRef] [PubMed]
- Gajecki, M. Evaluation of selected serum biochemical and haematological parameters in gilts exposed to 100 ppb of zearalenone. Pol. J. Vet. Sci. 2015, 18, 865–872. [Google Scholar]
- Marin, D.E.; Pistol, G.C.; Neagoe, I.V.; Calin, L.; Taranu, I. Effects of zearalenone on oxidative stress and inflammation in weanling piglets. Food Chem. Toxicol. 2013, 58, 401–408. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.H.; Lei, Y.P.; Bao, Y.H.; Jia, R.; Ma, Q.G.; Zhang, J.Y.; Chen, J.; Ji, C. Ameliorative effects of Bacillus subtilis ANSB01G on zearalenone toxicosis in pre-pubertal female gilts. Food Addit. Contam. A 2015, 34, 617–625. [Google Scholar] [CrossRef] [PubMed]
- Niu, Q.S. Effect of Fusarium Toxins on Hematological Values, Development of Reproductive Organs, and Expression of Estrogen Receptor Genes in Post-Weaning Gilts. Ph.D. Thesis, Shandong Agricultural University, Taian, China, 2015. [Google Scholar]
- Liu, X.L.; Wu, R.Y.; Sun, X.F.; Cheng, S.F.; Zhang, R.Q.; Zhang, T.Y.; Zhang, X.F.; Zhao, Y.; Shen, W.; Li, L. Mycotoxin zearalenone exposure impairs genomic stability of swine follicular granulosa cells in vitro. Int. J. Biol. Sci. 2018, 14, 294–305. [Google Scholar] [CrossRef]
- Muthulakshmi, S.; Hamideh, P.F.; Habibi, H.R.; Maharajan, K.; Kadirvelu, K.; Mudili, V. Mycotoxin zearalenone induced gonadal impairment and altered gene expression in the hypothalamic-pituitary-gonadal axis of adult female zebrafish (Danio rerio). J. Appl. Toxicol. 2018, 38, 1388–1397. [Google Scholar] [CrossRef]
- Cao, H.; Liu, D.L.; Mo, X.M.; Xie, C.F.; Yao, D.L. A fungal enzyme with the ability of aflatoxin B1 conversion: Purification and ESI-MS/MS identification. Microbiol. Res. 2011, 166, 475–483. [Google Scholar] [CrossRef]
- Yin, Q.Q.; Fan, G.G.; Chang, J.; Zuo, R.Y.; Zheng, Q.H. Effect of the combined probiotics on inhibiting pathogenic Escherichia coli proliferation. Adv. Mater. Res. 2012, 343, 802–808. [Google Scholar]
- Huang, W.W.; Chang, J.; Wang, P.; Liu, C.Q.; Yin, Q.Q.; Zhu, Q.; Lu, F.S.; Gao, T.Z. Effect of the combined compound probiotics with mycotoxin-degradation enzyme on detoxifying aflatoxin B1 and zearalenone. J. Toxicol. Sci. 2018, 43, 377–385. [Google Scholar] [CrossRef] [PubMed]
- National Research Council (NRC). Nutrient Requirements of Swine, 11th rev. ed.; National Academic Press: Washington, DC, USA, 2012. [Google Scholar]
- Jurgens, M.H. Animal Feeding and Nutrition, 8th ed.; Kendall/Hunt Publishing Company: Dubuque, IA, USA, 1997. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2 −ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Maibach, R.C.; Dutly, F.; Altwegg, M. Detection of Tropheryma whipplei DNA in feces by PCR using a target capture method. J. Clin. Microbiol. 2002, 40, 2466–2471. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.Q.; Zhu, Q.; Chang, J.; Yin, Q.Q.; Song, A.D.; Li, Z.T.; Wang, E.Z.; Lu, F.S. Effects of Lactobacillus casei and Enterococcus faecalis on growth performance, immune function and gut microbiota of suckling piglets. Arch. Anim. Nutr. 2017, 71, 120–133. [Google Scholar] [CrossRef] [PubMed]
- Konstantinov, S.R.; Zhu, W.Y.; Williams, B.A.; Tamminga, S.; de Vos, W.M.; Akkermans, A.D.L. Effect of fermentable carbohydrates on piglet faecal bacterial communities as revealed by denaturing gradient gel electrophoresis analysis of 16S ribosomal DNA. FEMS Microbiol. Ecol. 2003, 43, 225–235. [Google Scholar] [CrossRef]
| Number | C. utilis (%) A | B. subtilis SP1 (%) B | B. subtilis SP2 (%) C | ZEA Content (μg/L) | ZEA Degradation Rate (%) |
|---|---|---|---|---|---|
| 1 | 0.5 | 0.5 | 0.5 | 79.14 ± 24.91e | 85.00 ± 4.98ab |
| 2 | 0.5 | 1 | 1 | 40.80 ± 6.68e | 92.27 ± 1.30a |
| 3 | 0.5 | 1.5 | 1.5 | 40.82 ± 12.25e | 92.26 ± 2.45a |
| 4 | 1 | 0.5 | 1 | 123.53 ± 14.87de | 76.58 ± 2.41bc |
| 5 | 1 | 1 | 1.5 | 163.59 ± 16.09cde | 68.99 ± 0.78cd |
| 6 | 1 | 1.5 | 0.5 | 247.85 ± 42.91c | 53.02 ± 3.63e |
| 7 | 1.5 | 0.5 | 1.5 | 238.45 ± 13.50cd | 54.80 ± 7.96e |
| 8 | 1.5 | 1 | 0.5 | 382.36 ± 102.93b | 27.52 ± 13.24f |
| 9 | 1.5 | 1.5 | 1 | 282.28 ± 20.81bc | 46.49 ± 2.89e |
| Control | 0 | 0 | 0 | 527.52 ± 63.55a | |
| T1 | 269.2 | 215.47 | 166 | ||
| T2 | 198.58 | 189.18 | 214.96 | ||
| T3 | 128.48 | 191.61 | 215.3 | ||
| X1 | 89.73 | 71.82 | 55.33 | ||
| X2 | 66.19 | 63.06 | 71.65 | ||
| X3 | 42.83 | 63.87 | 71.77 | ||
| R | 46.9 | 8.76 | 16.44 | ||
| Impact order | A > C > B | ||||
| Optimal solution | A1B1C3 | ||||
| Sources | Sum of Squares | DF | Mean Square | F | p |
|---|---|---|---|---|---|
| Corrected Model | 2808.72 | 6 | 468.12 | 73.68 | 0.0132 * |
| A | 2545.95 | 2 | 1272.98 | 200.35 | 0.0051 * |
| B | 83.97 | 2 | 41.99 | 6.61 | 0.1312 |
| C | 178.79 | 2 | 89.40 | 14.07 | 0.0661 |
| Error | 12.71 | 2 | 6.35 | ||
| Corrected total | 2821.42 | 8 |
| Groups | Probiotics: Cell-free extracts of A. oryzae | ZEA Content (μg/L) | ZEA Degradation Rate (%) |
|---|---|---|---|
| 1 | 1:1 | 120.57 ± 21.56c | 89.89 ± 1.81a |
| 2 | 1:2 | 128.27 ± 79.74c | 89.24 ± 6.69a |
| 3 | 1:3 | 64.31 ± 83.73c | 94.60 ± 7.02a |
| 4 | 2:1 | 57.81 ± 12.83c | 95.15 ± 1.08a |
| 5 | 2:3 | 108.19 ± 28.55c | 90.92 ± 2.39a |
| 6 | 3:1 | 84.05 ± 73.25c | 92.95 ± 6.14a |
| 7 | 3:2 | 102.72 ± 72.17c | 91.83 ± 6.05a |
| 8 | 1:0 | 349.38 ± 34.13b | 70.69 ± 2.86b |
| 9 | 0:1 | 170.59 ± 64.49c | 85.69 ± 5.41a |
| Control | 0:0 | 1192.10 ± 86.55a |
| Items | Group A | Group B | Group C | Group D | Group E |
|---|---|---|---|---|---|
| Initial weight (Kg) | 34.83 ± 1.06 | 34.92 ± 1.34 | 35.17 ± 2.35 | 34.88 ± 1.27 | 34.83 ± 1.89 |
| Final weight (Kg) | 90.00 ± 4.23 | 83.58 ± 4.47 | 82.63 ± 1.80 | 88.54 ± 2.47 | 88.33 ± 3.70 |
| ADG (Kg) | 0.9211 ± 0.0812 | 0.8088 ± 0.0811 | 0.7886 ± 0.0304 | 0.8913 ± 0.0412 | 0.8908 ± 0.0321 |
| ADFI (Kg) | 2.612 ± 0.151 | 2.402 ± 0.171 | 2.389 ± 0.080 | 2.614 ± 0.140 | 2.542 ± 0.079 |
| F/G | 2.842 ± 0.100 | 2.963 ± 0.121 | 3.019 ± 0.213 | 2.914 ± 0.020 | 2.771 ± 0.169 |
| CP digestibility (%) | 89.73 ± 0.16 | 91.97 ± 0.94 | 89.53 ± 1.67 | 89.23 ± 1.25 | 90.67 ± 1.93 |
| CF digestibility (%) | 76.99 ± 3.94 | 80.87 ± 5.56 | 82.63 ± 4.28 | 78.41 ± 3.11 | 75.32 ± 1.22 |
| P digestibility (%) | 88.19 ± 0.42 | 87.67 ± 0.48 | 86.38 ± 1.58 | 87.21 ± 0.20 | 88.98 ± 0.07 |
| Ca digestibility (%) | 78.83 ± 0.22 | 77.51 ± 0.75 | 77.44 ± 0.09 | 76.48 ± 1.19 | 77.21 ± 0.89 |
| Items | Group A | Group B | Group D |
|---|---|---|---|
| Heart (g/Kg) | 3.241 ± 0.100 | 3.177 ± 0.311 | 3.616 ± 0.194 |
| Liver (g/Kg) | 15.97 ± 1.81 | 16.33 ± 0.80 | 18.67 ± 1.67 |
| Spleen (g/Kg) | 1.365 ± 0.249 | 1.605 ± 0.041 | 1.703 ± 0.164 |
| Kidney (g/Kg) | 2.960 ± 0.172 | 2.967 ± 0.221 | 3.432 ± 0.496 |
| Uterus (g/Kg) | 1.245 ± 0.752 | 1.932 ± 0.587 | 1.673 ± 0.318 |
| Serum E2 (pg/ml) | 109.03 ± 8.29 | 112.48 ± 15.75 | 103.88 ± 5.06 |
| ERα mRNA abundance in ovaries | 0.7899 ± 0.1021 | 1.171 ± 0.141 | 1.171 ± 0.122 |
| ERα mRNA abundance in uterus | 1.010 ± 0.086 | 1.111 ± 0.153 | 1.071 ± 0.191 |
| Items | Group A | Group B | Group D |
|---|---|---|---|
| Serum | — | — | — |
| Longissimus dorsi | — | — | — |
| Uterus | — | — | — |
| Liver | — | — | — |
| Contents in jejunum | 11.64 ± 0.27b | 110.54 ± 16.19a | 22.03 ± 8.20b |
| Contents in large intestine | 43.45 ± 2.44b | 138.23 ± 4.67a | 120.55 ± 18.87a |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, C.; Chang, J.; Wang, P.; Yin, Q.; Huang, W.; Dang, X.; Lu, F.; Gao, T. Zearalenone Biodegradation by the Combination of Probiotics with Cell-Free Extracts of Aspergillus oryzae and Its Mycotoxin-Alleviating Effect on Pig Production Performance. Toxins 2019, 11, 552. https://doi.org/10.3390/toxins11100552
Liu C, Chang J, Wang P, Yin Q, Huang W, Dang X, Lu F, Gao T. Zearalenone Biodegradation by the Combination of Probiotics with Cell-Free Extracts of Aspergillus oryzae and Its Mycotoxin-Alleviating Effect on Pig Production Performance. Toxins. 2019; 11(10):552. https://doi.org/10.3390/toxins11100552
Chicago/Turabian StyleLiu, Chaoqi, Juan Chang, Ping Wang, Qingqiang Yin, Weiwei Huang, Xiaowei Dang, Fushan Lu, and Tianzeng Gao. 2019. "Zearalenone Biodegradation by the Combination of Probiotics with Cell-Free Extracts of Aspergillus oryzae and Its Mycotoxin-Alleviating Effect on Pig Production Performance" Toxins 11, no. 10: 552. https://doi.org/10.3390/toxins11100552
APA StyleLiu, C., Chang, J., Wang, P., Yin, Q., Huang, W., Dang, X., Lu, F., & Gao, T. (2019). Zearalenone Biodegradation by the Combination of Probiotics with Cell-Free Extracts of Aspergillus oryzae and Its Mycotoxin-Alleviating Effect on Pig Production Performance. Toxins, 11(10), 552. https://doi.org/10.3390/toxins11100552




