Aflatoxin B1 and M1: Biological Properties and Their Involvement in Cancer Development
Abstract
1. Introduction
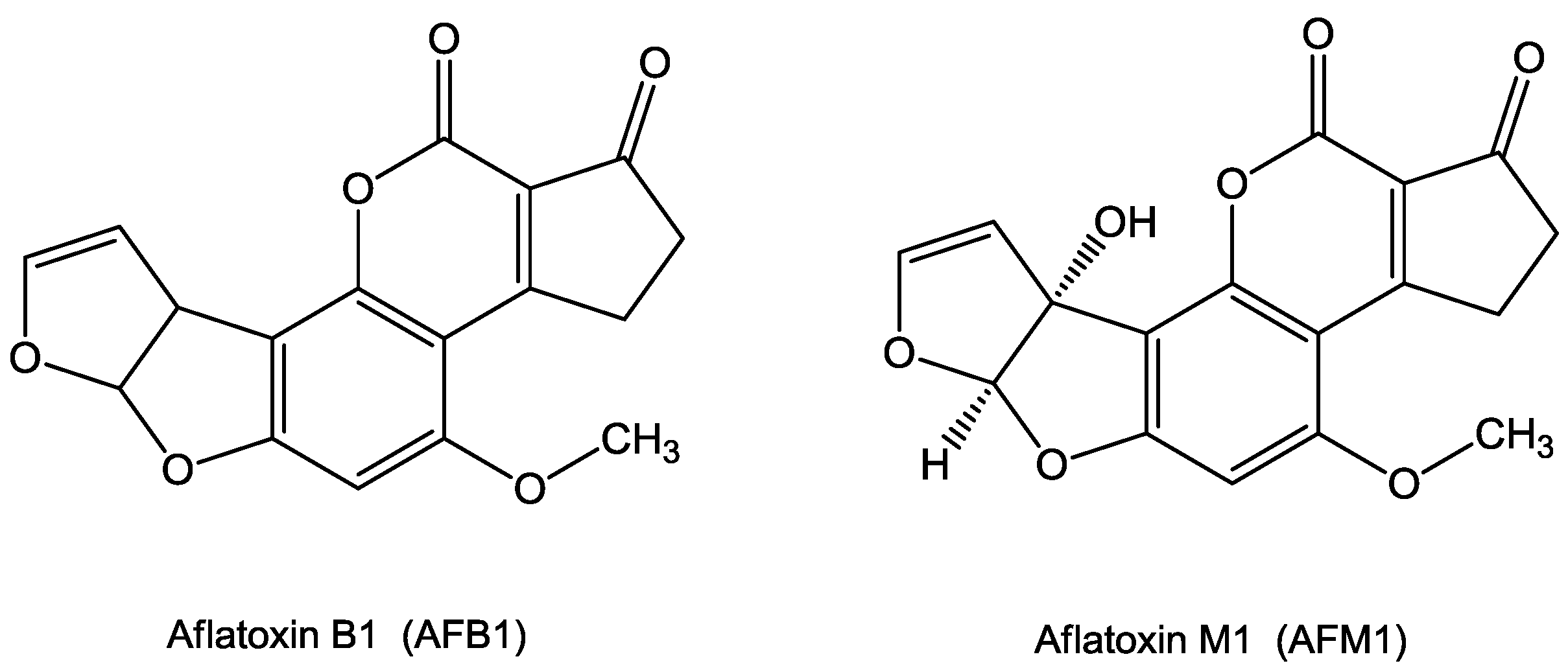
2. Chemical Properties of AFB1 and AFM1
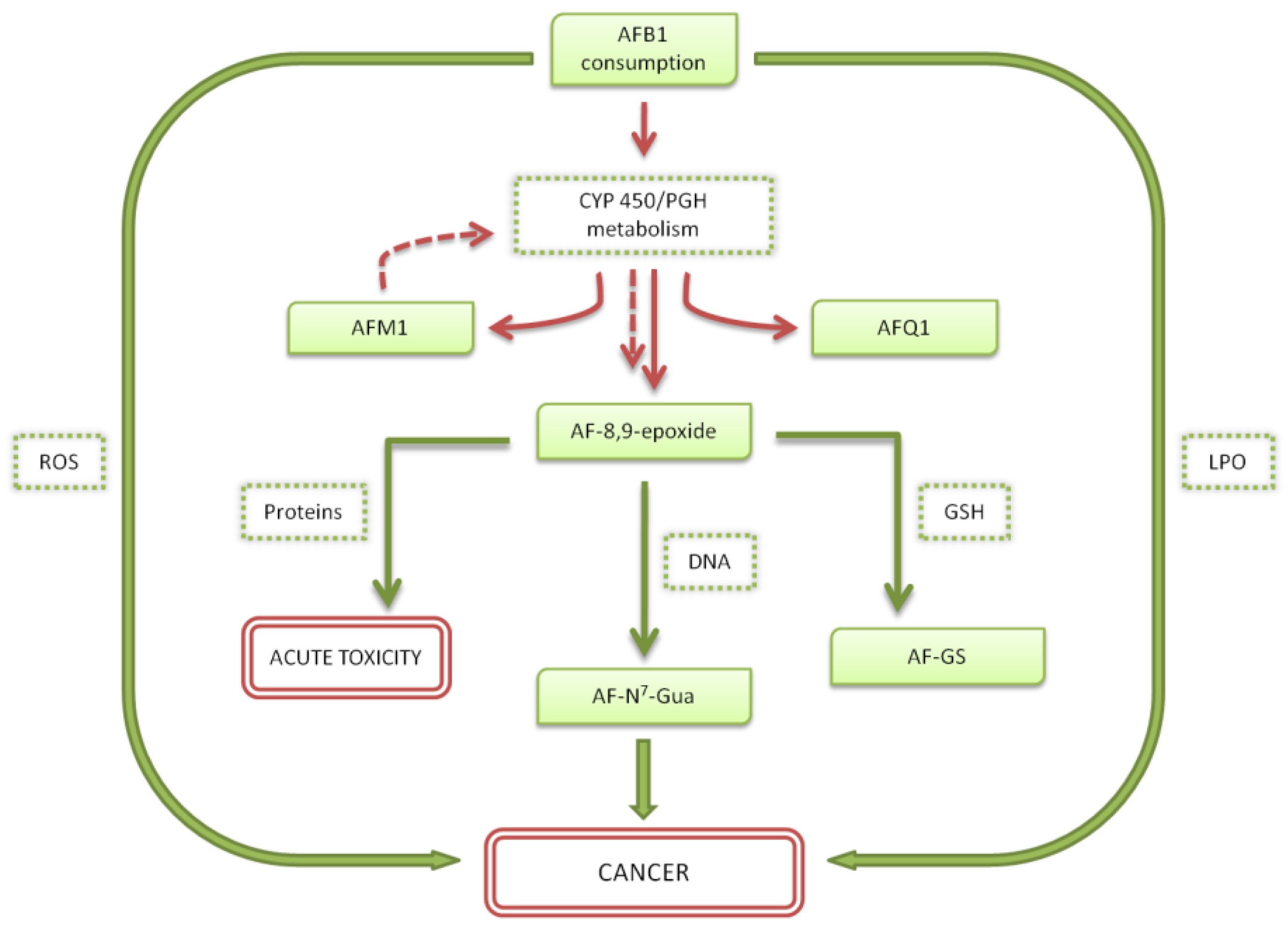
3. Principal Activation Pathways of AFB1 and AFM1
4. Aflatoxins Related Diseases
4.1. Liver Cancer
4.2. Lung Cancer
4.3. Gastrointestinal Cancer and Other Cancers
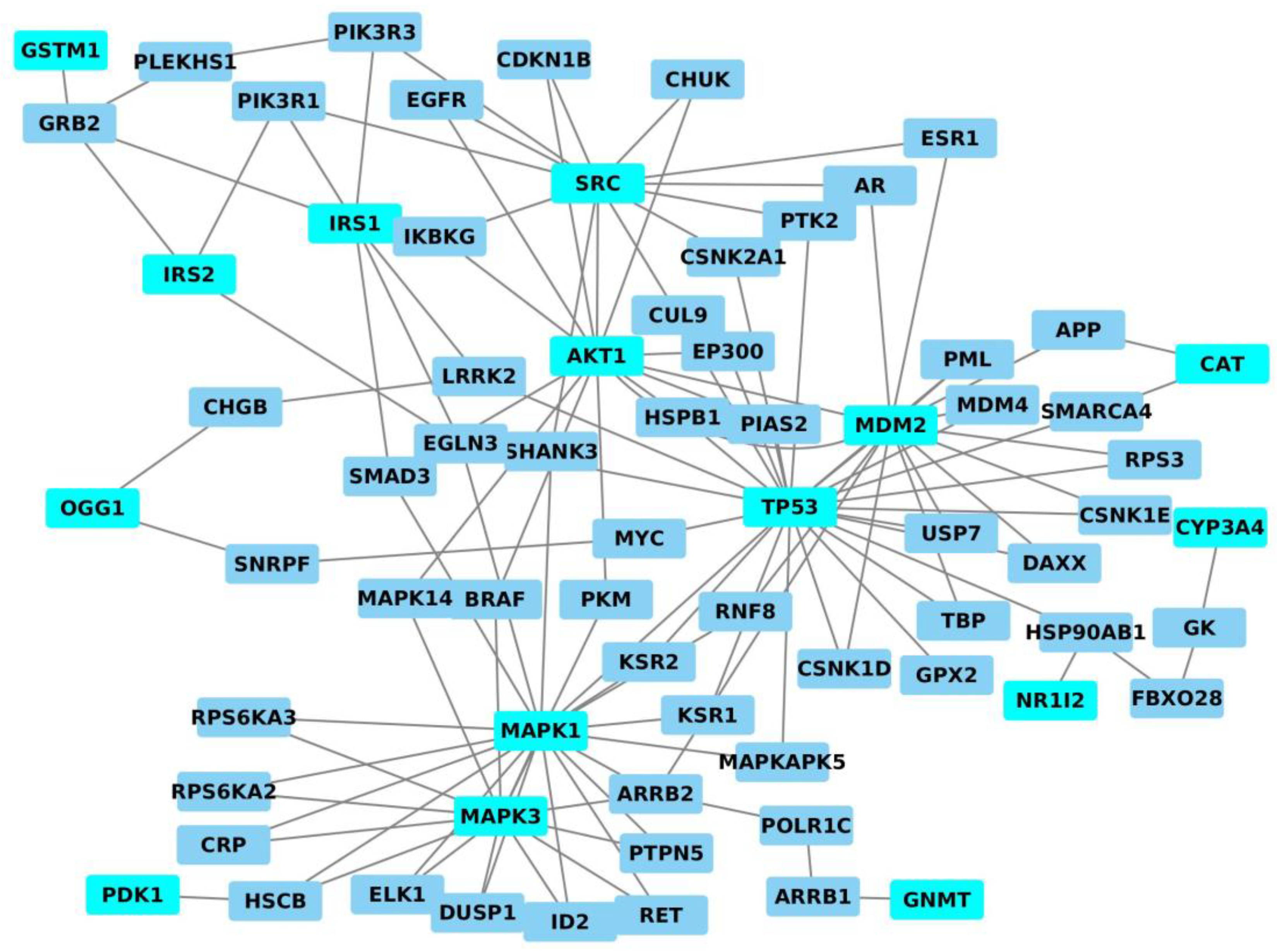
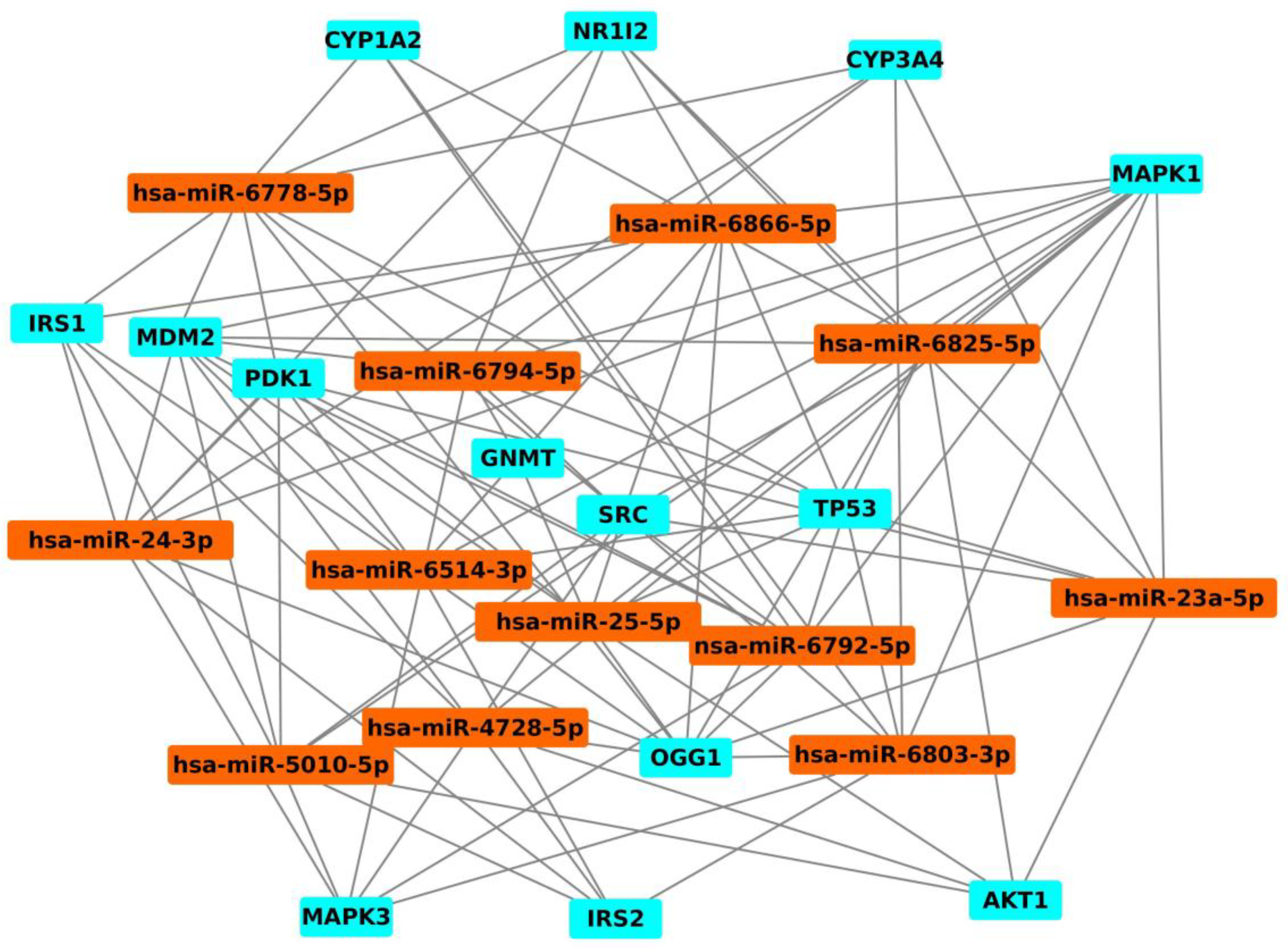
5. Analysis of Genes/Proteins Modulated by AFB1/AFM1 by Systems Biology Approach
6. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Blount, W.P. Turkey “X” disease. Turkeys 1961, 9, 52–54. [Google Scholar]
- World Health Organization; International Agency for Research on Cancer. Aflatoxins. In IARC Monographs on the Evaluation of Carcinogenic Risks to Humans; IARC Press: Lyon, France, 2002; Volume 82, pp. 171–300. [Google Scholar]
- Meissonnier, G.M.; Laffitte, J.; Loiseau, N.; Benoit, E.; Raymond, I.; Pinton, P.; Cossalter, A.M.; Bertin, G.; Oswald, I.P.; Galtier, P. Selective impairment of drug-metabolizing enzymes in pig liver during subchronic dietary exposure to aflatoxin B1. Food Chem. Toxicol. 2007, 45, 2145–2154. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization; International Agency for Research on Cancer. Aflatoxins. In IARC Monographs on the Evaluation of Carcinogenic Risks to Humans; IARC Press: Lyon, France, 1993; Volume 56, pp. 245–395. [Google Scholar]
- Sklan, D.; Klipper, E.; Friedman, A.; Shelly, M.; Makovsky, B. The Effect of Chronic Feeding of Diacetoxyscirpenol, T-2 Toxin, and Aflatoxin on Performance, Health, and Antibody Production in Chicks. J. Appl. Poult. Res. 2001, 10, 79–85. [Google Scholar] [CrossRef]
- Ayres, J.L.; Lee, D.J.; Wales, J.H.; Sinnhuber, R.O. Aflatoxin structure and hepatocarcinogenicity in rainbow trout (Salmogairdneri). J. Natl. Cancer Inst. 1971, 46, 561–564. [Google Scholar] [PubMed]
- Bodine, A.B.; Fisher, S.F.; Gangjee, S. Effect of Aflatoxin B1 and Major Metabolites on Phytohemeagglutinin-Stimulated Lymphoblastogenesis of Bovine Lymphocytes. J. Dairy Sci. 1984, 67, 110–114. [Google Scholar] [CrossRef]
- Wogan, G.N.; Paglialunga, S.; Newberne, P.M. Carcinogenic effects of low dietary levels of aflatoxin B1 in rats. Food Cosmet. Toxicol. 1974, 12, 681–685. [Google Scholar] [CrossRef]
- Cullen, J.M.; Ruebner, B.H.; Hsieh, L.S.; Hyde, D.M.; Hsieh, D.P. Carcinogenicity of Dietary Aflatoxin M1 in Male Fischer Rats Compared to Aflatoxin B1. Cancer Res. 1987, 47, 1913–1917. [Google Scholar] [PubMed]
- Peraica, M.; Radić, B.; Lucić, A.; Pavlović, M. Toxic effects of mycotoxins in humans. Bull. World Health Organ. 1999, 77, 754–766. [Google Scholar] [PubMed]
- Wogan, G.N. Aflatoxins as Risk Factors for Hepatocellular Carcinoma in Humans. Cancer Res. 1992, 52, 2114s–2118s. [Google Scholar] [PubMed]
- Turner, P.C.; Moore, S.E.; Hall, A.J.; Prentice, A.M.; Wild, C.P. Modification of immune function through exposure to dietary aflatoxin in Gambian children. Environ. Health Perspect. 2003, 111, 217–220. [Google Scholar] [CrossRef] [PubMed]
- Neal, G.E.; Eaton, D.L.; Judah, D.J.; Verma, A. Metabolism and toxicity of aflatoxins M1 and B1 in human-derived in vitro systems. Toxicol. Appl. Pharmacol. 1998, 151, 152–158. [Google Scholar] [CrossRef] [PubMed]
- Bianco, G.; Russo, R.; Marzocco, S.; Velotto, S.; Autore, G.; Severino, L. Modulation of macrophage activity by aflatoxins B1 and B2 and their metabolites aflatoxins M1 and M2. Toxicon 2012, 59, 644–650. [Google Scholar] [CrossRef] [PubMed]
- Veldman, A.; Meijs, J.; Borggreve, G.; Heeres-van der Tol, J. Carry-over of aflatoxin from cows’ food to milk. Anim. Sci. 1992, 55, 163–168. [Google Scholar] [CrossRef]
- Jafari, T.; Fallah, A.A.; Kheiri, S.; Fadaei, A.; Amini, S.A. Aflatoxin M1 in human breast milk in Shahrekord, Iran and association with dietary factors. Food Addit. Contam. Part B 2017, 10, 128–136. [Google Scholar] [CrossRef] [PubMed]
- Luongo, D.; Russo, R.; Balestrieri, A.; Marzocco, S.; Bergamo, P.; Severino, L. In vitro study of AFB1 and AFM1 effects on human lymphoblastoidJurkat T-cell model. J. Immunotoxicol. 2013, 11, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Wild, C.P.; Turner, P.C. The toxicology of aflatoxins as a basis for public health decisions. Mutagenesis 2002, 17, 471–481. [Google Scholar] [CrossRef] [PubMed]
- Gallagher, E.P.; Wienkers, L.C.; Stapleton, P.L.; Kunze, K.L.; Eaton, D.L. Role of Human Microsomal and Human Complementary DNA-expressed Cytochromes P4501A2 and P4503A4 in the Bioactivation of Aflatoxin B1. Cancer Res. 1994, 54, 101–108. [Google Scholar] [PubMed]
- MdQuadri, S.H.; Niranjan, M.S.; Chaluvaraju, K.C.; Shantaram, U.; Enamul, H.S. An overview on chemistry, toxicity, analysis and control of aflatoxins. Int. J. Chem. Life Sci. 2013, 2, 1071–1078. [Google Scholar]
- Jard, G.; Liboz, T.; Mathieu, F.; Guyonvarc’h, A.; Lebrihi, A. Review of mycotoxin reduction in food and feed: From prevention in the field to detoxification by adsorption or transformation. Food Addit. Contam. Part A 2011, 28, 1590–1609. [Google Scholar] [CrossRef] [PubMed]
- Prandini, A.; Tansini, G.; Sigolo, S.; Filippi, L.; Laporta, M.; Piva, G. On the occurrence of aflatoxin M1 in milk and dairy products. Food Chem. Toxicol. 2009, 47, 984–991. [Google Scholar] [CrossRef] [PubMed]
- Rustom, I.Y.S. Aflatoxin in food and feed: Occurrence, legislation and inactivation by physical methods. Food Chem. 1997, 59, 57–67. [Google Scholar] [CrossRef]
- Park, D.L.; Price, W.D. Reduction of aflatoxin hazards using ammoniation. Rev. Environ. Contam. Toxicol. 2001, 171, 139–175. [Google Scholar]
- Giovati, L.; Magliani, W.; Ciociola, T.; Santinoli, C.; Conti, S.; Polonelli, L. AFM1in Milk: Physical, Biological, and Prophylactic Methods to Mitigate Contamination. Toxins 2015, 7, 4330–4349. [Google Scholar] [CrossRef] [PubMed]
- Jebali, R.; Abbès, S.; Salah-Abbès, J.B.; Ben Younes, R.; Haous, Z.; Oueslati, R. Ability of Lactobacillus plantarum MON03 to mitigate aflatoxins (B1 and M1) immunotoxicities in mice. J. Immunotoxicol. 2014, 12, 290–299. [Google Scholar] [CrossRef] [PubMed]
- Eaton, D.L.; Gallagher, E.P. Mechanisms of Aflatoxin Carcinogenesis. Annu. Rev. Pharmacol. Toxicol. 1994, 34, 135–172. [Google Scholar] [CrossRef] [PubMed]
- Verma, R.J. Aflatoxin Cause DNA Damage. Int. J. Hum. Genet. 2004, 4, 231–236. [Google Scholar] [CrossRef]
- Bbosa, G.S.; Kitya, D.; Lubega, A.; Ogwal-Okeng, J.; Anokbonggo, W.W.; Kyegombe, D.B. Review of the biological and health effects of aflatoxins on bodyorgans and body systems. In Aflatoxins—Recent Advances and Future Prospects; Razzaghi-Abyaneh, M., Ed.; InTech: Rijeka, Croatia, 2013; pp. 239–265. ISBN 978-953-51-0904-4. [Google Scholar]
- Gallagher, E.P.; Kunze, K.L.; Stapleton, P.L.; Eaton, D.L. The Kinetics of Aflatoxin B1 Oxidation by Human cDNA-Expressed and Human Liver Microsomal Cytochromes P450 1A2 and 3A4. Toxicol. Appl. Pharmacol. 1996, 141, 595–606. [Google Scholar] [CrossRef] [PubMed]
- Bailey, E.A.; Iyer, R.S.; Stone, M.P.; Harris, T.M.; Essigmann, J.M. Mutational properties of the primary aflatoxin B1-DNA adduct. Proc. Natl. Acad. Sci. USA 1996, 93, 1535–1539. [Google Scholar] [CrossRef] [PubMed]
- Macé, K.; Aguilar, F.; Wang, J.S.; Vautravers, P.; Gómez-Lechón, M.; Gonzalez, F.J.; Groopman, J.; Harris, C.C.; Pfeifer, A.M. Aflatoxin B1-induced DNA adduct formation and p53 mutations in CYP450- expressing human liver cell lines. Carcinogenesis 1997, 18, 1291–1297. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Cao, Y.; He, L.; Wang, N.J.; Gu, J.R. Aberrations of p53 gene in human hepatocellular carcinoma from China. Carcinogenesis 1993, 14, 169–173. [Google Scholar] [CrossRef] [PubMed]
- Soman, N.R.; Wogan, G.N. Activation of the c-Ki-ras oncogene in aflatoxin B1-induced hepatocellular carcinoma and adenoma in the rat: Detection by denaturing gradient gel electrophoresis. Proc. Natl. Acad. Sci. USA 1993, 90, 2045–2049. [Google Scholar] [CrossRef] [PubMed]
- Riley, J.; Mandel, H.G.; Sinha, S.; Judah, D.J.; Neal, G.E. In vitro activation of the human Harvey-ras proto-oncogene by aflatoxin B1. Carcinogenesis 1997, 18, 905–910. [Google Scholar] [CrossRef] [PubMed]
- Battista, J.R.; Marnett, L.J. Prostaglandin H synthase-dependent epoxidation of aflatoxin B1. Carcinogenesis 1985, 6, 1227–1229. [Google Scholar] [CrossRef] [PubMed]
- Weng, M.W.; Lee, H.W.; Choi, B.; Wang, H.T.; Hu, Y.; Mehta, M.; Desai, D.; Amin, S.; Zheng, Y.; Tang, M.S. AFB1 hepatocarcinogenesis is via lipid peroxidation that inhibits DNA repair, sensitizes mutation susceptibility and induces aldehyde-DNA adducts at p53 mutational hotspot codon 249. Oncotarget 2017, 8, 18213–18226. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, W. Aflatoxin B1 impairs mitochondrial functions, activates ROS generation, induces apoptosis and involves Nrf2 signal pathway in primary broiler hepatocytes. Anim. Sci. J. 2016, 87, 1490–1500. [Google Scholar] [CrossRef] [PubMed]
- Wogan, G.W.; Paglialunga, S. Carcinogenicity of synthetic aflatoxin M1 in rats. Food Cosmet. Toxicol. 1974, 12, 381–384. [Google Scholar] [CrossRef]
- Nugraha, A.; Khotimah, K.; Rietjens, I.M.C.M. Risk assessment of aflatoxin B1 exposure from maize and peanut consumption in Indonesia using the margin of exposure and liver cancer risk estimation approaches. Food Chem. Toxicol. 2018, 113, 134–144. [Google Scholar] [CrossRef] [PubMed]
- Kucukcakan, B.; Hayrulai-Musliu, Z. Challenging role of dietary aflatoxin B1 exposure and hepatitis B infection on risk of hepatocellular carcinoma. Open Access Maced. J. Med. Sci. 2015, 3, 363–369. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wu, F. Global burden of aflatoxin-induced hepatocellular carcinoma: A risk assessment. Environ. Health Perspect. 2010, 118, 818–824. [Google Scholar] [CrossRef] [PubMed]
- Rapisarda, V.; Loreto, C.; Malaguarnera, M.; Ardiri, A.; Proiti, M.; Rigano, G.; Frazzetto, E.; Ruggeri, M.I.; Malaguarnera, G.; Bertino, N.; et al. Hepatocellular carcinoma and the risk of occupational exposure. World J. Hepatol. 2016, 8, 573–590. [Google Scholar] [CrossRef] [PubMed]
- Xiang, X.; Qin, H.G.; You, X.M.; Wang, Y.Y.; Qi, L.N.; Ma, L.; Xiang, B.D.; Zhong, J.H.; Li, L.Q. Expression of P62 in hepatocellular carcinoma involving hepatitis B virus infection and aflatoxin B1 exposure. Cancer Med. 2017, 6, 2357–2369. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Wang, L.; Zheng, C.; Liu, L.; Wang, J.; Li, D.; Tan, Y.; Zhao, X.; He, L.; Shu, W. Microcystin-LR increases genotoxicity induced by aflatoxin B1 through oxidative stress and DNA base excision repair genes in human hepatic cell lines. Environ. Pollut. 2018, 233, 455–463. [Google Scholar] [CrossRef] [PubMed]
- Fedeles, B.I.; Chawanthayatham, S.; Croy, R.G.; Wogan, G.N.; Essigmann, J.M. Early detection of the aflatoxin B1 mutational fingerprint: A diagnostic tool for liver cancer. Mol. Cell. Oncol. 2017, 4, e1329693:1–e1329693:3. [Google Scholar] [CrossRef] [PubMed]
- Tsai, F.J.; Chen, S.Y.; Liu, Y.C.; Liao, H.Y.; Chen, C.J. The comparison of CHCA solvent compositions for improving LC-MALDI performance and its application to study the impact of aflatoxin B1 on the liver proteome of diabetes mellitus type 1 mice. PLoS ONE 2017, 12, e0181423:1–e0181423:16. [Google Scholar] [CrossRef] [PubMed]
- Chu, Y.J.; Yang, H.I.; Wu, H.C.; Liu, J.; Wang, L.Y.; Lu, S.N.; Lee, M.H.; Jen, C.L.; You, S.L.; Santella, R.M.; et al. Aflatoxin B1 exposure increases the risk of cirrhosis and hepatocellular carcinoma in chronic hepatitis B virus carriers. Int. J. Cancer. 2017, 141, 711–720. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; He, Z.; Li, D.; Zhang, B.; Li, M.; Li, W.; Zhu, W.; Xing, X.; Zeng, X.; Wang, Q.; et al. Aberrant methylation of RUNX3 is present in Aflatoxin B1-induced transformation of the L02R cell line. Toxicology 2017, 385, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; He, H.; Zang, M.; Wu, Q.; Zhao, H.; Lu, L.L.; Ma, P.; Zheng, H.; Wang, N.; Zhang, Y.; et al. Genetic Features of Aflatoxin-Associated Hepatocellular Carcinoma. Gastroenterology 2017, 153, 249–262. [Google Scholar] [CrossRef] [PubMed]
- Livingstone, M.C.; Johnson, N.M.; Roebuck, B.D.; Kensler, T.W.; Groopman, J.D. Profound changes in miRNA expression during cancer initiation by aflatoxin B1 and their abrogation by the chemopreventive triterpenoid CDDO-Im. Mol. Carcinog. 2017, 56, 2382–2390. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; He, J.; Lin, J.; Sun, X.; Sun, F.; Ou, C.; Jiang, C. Distinct response of the hepatic transcriptome to Aflatoxin B1 induced hepatocellular carcinogenesis and resistance in rats. Sci. Rep. 2016, 6, 31898:1–31898:9. [Google Scholar] [CrossRef] [PubMed]
- Rieswijk, L.; Claessen, S.M.; Bekers, O.; van Herwijnen, M.; Theunissen, D.H.; Jennen, D.G.; de Kok, T.M.; Kleinjans, J.C.; van Breda, S.G. Aflatoxin B1 induces persistent epigenomic effects in primary human hepatocytes associated with hepatocellular carcinoma. Toxicology 2016, 350–352, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Ju, H.; Shim, Y.; Arumugam, P.; Song, J.M. Crosstalk-eliminated quantitative determination of aflatoxin B1-induced hepatocellular cancer stem cells based on concurrent monitoring of CD133, CD44, and aldehyde dehydrogenase1. Toxicol. Lett. 2016, 243, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Marrone, A.K.; Tryndyak, V.; Beland, F.A.; Pogribny, I.P. MicroRNA Responses to the Genotoxic Carcinogens Aflatoxin B1 and Benzo[a]pyrene in Human HepaRG Cells. Toxicol. Sci. 2016, 149, 496–502. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Gao, J.; Huang, K.; Luo, Y.; Zhang, B.; Xu, W. miR-34a screened by miRNA profiling negatively regulates Wnt/β-catenin signaling pathway in Aflatoxin B1 induced hepatotoxicity. Sci. Rep. 2015, 5, 16732:1–16732:13. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.W.; Lien, J.P.; Chiu, Y.H.; Santella, R.M.; Liaw, Y.F.; Chen, C.J. Effect of aflatoxin metabolism and DNA adduct formation on hepatocellular carcinoma among chronic hepatitis B carriers in Taiwan. J. Hepatol. 1997, 27, 320–330. [Google Scholar] [CrossRef]
- Sun, Z.; Lu, P.; Gail, M.H.; Pee, D.; Zhang, Q.; Ming, L.; Wang, J.; Wu, Y.; Liu, G.; Wu, Y.; et al. Increased risk of hepatocellular carcinoma in male hepatitis B surface antigen carriers with chronic hepatitis who have detectable urinary aflatoxin metabolite M1. Hepatology 1999, 30, 379–383. [Google Scholar] [CrossRef] [PubMed]
- Mokhles, M.; Abd El Wahhab, M.A.; Tawfik, M.; Ezzat, W.; Gamil, K.; Ibrahim, M. Detection of aflatoxin among hepatocellular carcinoma patients in Egypt. Pak. J. Biol. Sci. 2007, 10, 1422–1429. [Google Scholar] [CrossRef] [PubMed]
- Lu, P.X.; Wang, J.B.; Zhang, Q.N.; Wu, Y.; Sun, Y.; Chen, T.Y. Longitudinal study of aflatoxin exposure in the development of primary liver cancer in patients with chronic hepatitis. Zhonghua Yi Xue Za Zhi 2010, 90, 1665–1669. [Google Scholar] [PubMed]
- Saad-Hussein, A.; Beshir, S.; Moubarz, G.; Elserougy, S.; Ibrahim, M.I. Effect of occupational exposure to aflatoxins on some liver tumor markers in textile workers. Am. J. Ind. Med. 2013, 56, 818–824. [Google Scholar] [CrossRef] [PubMed]
- Hayes, R.B.; Van Nieuwenhuize, J.P.; Raategever, J.W.; Kate, F.J.W. Aflatoxin exposure in the industrial setting: an epidemiological study of mortality. Food Chem. Toxicol. 1984, 22, 39–43. [Google Scholar] [CrossRef]
- Donnelly, P.J.; Stewart, R.K.; Ali, S.L.; Conlan, A.A.; Reid, K.R.; Petsikas, D.; Massey, T.E. Biotransformation of aflatoxin B1 in human lung. Carcinogenesis 1996, 17, 2487–2494. [Google Scholar] [CrossRef] [PubMed]
- Massey, T.E.; Smith, G.B.; Tam, A.S. Mechanisms of aflatoxin B1 lung tumorigenesis. Exp. Lung Res. 2000, 26, 673–683. [Google Scholar] [PubMed]
- Van Vleet, T.R.; Klein, P.J.; Coulombe, R.A., Jr. Metabolism of aflatoxin B1 by normal human bronchial epithelial cells. J. Toxicol. Environ. Health A 2001, 63, 525–540. [Google Scholar] [CrossRef] [PubMed]
- Van Vleet, T.R.; Watterson, T.L.; Klein, P.J.; Coulombe, R.A., Jr. Aflatoxin B1 alters the expression of p53 in cytochrome P450-expressing human lung cells. Toxicol. Sci. 2006, 89, 399–407. [Google Scholar] [CrossRef] [PubMed]
- He, X.Y.; Tang, L.; Wang, S.L.; Cai, Q.S.; Wang, J.S.; Hong, J.Y. Efficient activation of aflatoxin B1 by cytochrome P450 2A13, an enzyme predominantly expressed in human respiratory tract. Int. J. Cancer 2006, 118, 2665–2671. [Google Scholar] [CrossRef] [PubMed]
- Guindon, K.A.; Foley, J.F.; Maronpot, R.R.; Massey, T.E. Failure of catalase to protect against aflatoxin B1-induced mouse lung tumorigenicity. Toxicol. Appl. Pharmacol. 2008, 227, 179–183. [Google Scholar] [CrossRef] [PubMed]
- Mulder, J.E.; Turner, P.V.; Massey, T.E. Effect of 8-oxoguanine glycosylase deficiency on aflatoxin B1 tumourigenicity in mice. Mutagenesis 2015, 30, 401–409. [Google Scholar] [CrossRef] [PubMed]
- Jakšić, D.; Puel, O.; Canlet, C.; Kopjar, N.; Kosalec, I.; Klarić, M.Š. Cytotoxicity and genotoxicity of versicolorins and 5-methoxysterigmatocystin in A549 cells. Arch. Toxicol. 2012, 86, 1583–1591. [Google Scholar] [CrossRef] [PubMed]
- Cui, A.; Hua, H.; Shao, T.; Song, P.; Kong, Q.; Luo, T.; Jiang, Y. Aflatoxin B1 induces Src phosphorylation and stimulates lung cancer cell migration. Tumour Biol. 2015, 36, 6507–6513. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, Z.; Wang, H.; Zhang, Y.; Ji, M.; Xu, H.; Wang, C.; Sun, Z.; Gao, W.; Wang, S.L. miR-138-1* regulates aflatoxin B1-induced malignant transformation of BEAS-2B cells by targeting PDK1. Arch. Toxicol. 2016, 90, 1239–1249. [Google Scholar] [CrossRef] [PubMed]
- Harrison, J.C.; Carvajal, M.; Garner, R.C. Does aflatoxin exposure in the United Kingdom constitute a cancer risk? Environ. Health Perspect. 1993, 99, 99–105. [Google Scholar] [CrossRef] [PubMed]
- Sobral, M.M.C.; Faria, M.A.; Cunha, S.C.; Ferreira, I.M. Toxicological interactions between mycotoxins from ubiquitous fungi: Impact on hepatic and intestinal human epithelial cells. Chemosphere 2018, 202, 538–548. [Google Scholar] [CrossRef] [PubMed]
- Gursoy-Yuzugullu, O.; Yuzugullu, H.; Yilmaz, M.; Ozturk, M. Aflatoxin genotoxicity is associated with a defective DNA damage response bypassing p53 activation. Liver Int. 2011, 31, 561–571. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Park, S.-H.; Do, K.H.; Kim, D.; Moon, Y. Interference with mutagenic aflatoxin B1-induced checkpoints through antagonistic action of ochratoxin A in intestinal cancer cells: A molecular explanation on potential risk of crosstalk between carcinogens. Oncotarget 2016, 7, 39627–39639. [Google Scholar] [CrossRef] [PubMed]
- Caloni, F.; Stammati, A.; Friggè, G.; De Angelis, I. Aflatoxin M1 absorption and cytotoxicity on human intestinal in vitro model. Toxicon 2006, 47, 409–415. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zheng, N.; Liu, J.; Li, F.D.; Li, S.L.; Wang, J.Q. Aflatoxin B1 and aflatoxin M1 induced cytotoxicity and DNA damage in differentiated and undifferentiated Caco-2 cells. Food Chem. Toxicol. 2015, 83, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.N.; Wang, J.Q.; Li, S.L.; Zhang, Y.D.; Zheng, N. Aflatoxin M1 cytotoxicity against human intestinal Caco-2 cells is enhanced in the presence of other mycotoxins. Food Chem. Toxicol. 2016, 96, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Li, S.; Wang, J.; Luo, C.; Zhao, S.; Zheng, N. Modulation of Intestinal Epithelial Permeability in Differentiated Caco-2 Cells Exposed to Aflatoxin M1 and Ochratoxin A Individually or Collectively. Toxins 2018, 10, 13. [Google Scholar] [CrossRef] [PubMed]
- Epstein, S.M.; Bartus, B.; Farber, E. Renal epithelial neoplasms induced in male Wistar rats by oral aflatoxin B1. Cancer Res. 1969, 29, 1045–1050. [Google Scholar] [PubMed]
- Merkow, L.P.; Epstein, S.M.; Slifkin, M.; Pardo, M. The ultrastructure of renal neoplasms induced by aflatoxin B1. Cancer Res. 1973, 33, 1608–1614. [Google Scholar] [PubMed]
- Kitamura, R.; Sato, K.; Sawada, M.; Itoh, S.; Kitada, M.; Komori, M.; Kamataki, T. Stable expression of cytochrome P450IIIA7 cDNA in human breast cancer cell line MCF-7 and its application to cytotoxicity testing. Arch. Biochem. Biophys. 1992, 292, 136–140. [Google Scholar] [CrossRef]
- Yip, K.Y.; Wan, M.L.Y.; Wong, A.S.T.; Korach, K.S.; El-Nezami, H. Combined low-dose zearalenone and aflatoxin B1 on cell growth and cell-cycle progression in breast cancer MCF-7 cells. Toxicol. Lett. 2017, 281, 139–151. [Google Scholar] [CrossRef] [PubMed]
- Faneyte, I.F.; Kristel, P.M.; Maliepaard, M.; Scheffer, G.L.; Scheper, R.J.; Schellens, J.H.; van de Vijver, M.J. Expression of the Breast Cancer Resistance Protein in Breast Cancer. Clin. Cancer Res. 2002, 8, 1068–1074. [Google Scholar] [PubMed]
- Van Herwaarden, A.E.; Wagenaar, E.; Karnekamp, B.; Merino, G.; Jonker, J.W.; Schinkel, A.H. Breast cancer resistance protein (Bcrp1/Abcg2) reduces systemic exposure of the dietary carcinogens aflatoxin B1, IQ and Trp-P-1 but also mediates their secretion into breast milk. Carcinogenesis 2005, 27, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Rastogi, S.; Dogra, R.K.; Khanna, S.K.; Das, M. Skin tumorigenic potential of aflatoxin B1 in mice. Food Chem. Toxicol. 2006, 44, 670–677. [Google Scholar] [CrossRef] [PubMed]
- Koshiol, J.; Gao, Y.T.; Dean, M.; Egner, P.; Nepal, C.; Jones, K.; Wang, B.; Rashid, A.; Luo, W.; Van Dyke, A.L.; et al. Association of Aflatoxin and Gallbladder Cancer. Gastroenterology 2017, 153, 488–494. [Google Scholar] [CrossRef] [PubMed]
- Ghasemi-Kebria, F.; Joshaghani, H.; Taheri, N.S.; Semnani, S.; Aarabi, M.; Salamat, F.; Roshandel, G. Aflatoxin contamination of wheat flour and the risk of esophageal cancer in a high risk area in Iran. Cancer Epidemiol. 2013, 37, 290–293. [Google Scholar] [CrossRef] [PubMed]
- Eom, S.Y.; Yim, D.H.; Zhang, Y.; Yun, J.K.; Moon, S.I.; Yun, H.Y.; Song, Y.J.; Youn, S.J.; Hyun, T.; Park, J.S.; et al. Dietary aflatoxin B1 intake, genetic polymorphisms of CYP1A2, CYP2E1, EPHX1, GSTM1, and GSTT1, and gastric cancer risk in Korean. Cancer Causes Control 2013, 24, 1963–1972. [Google Scholar] [CrossRef] [PubMed]
- Polo, A.; Crispo, A.; Cerino, P.; Falzone, L.; Candido, S.; Giudice, A.; De Petro, G.; Ciliberto, G.; Montella, M.; Budillon, A.; et al. Environment and bladdercancer: Molecularanalysis by interaction networks. Oncotarget 2017, 8, 65240–65252. [Google Scholar] [CrossRef] [PubMed]
- Ratajewski, M.; Walczak-Drzewiecka, A.; Sałkowska, A.; Dastych, J. Aflatoxins upregulate CYP3A4 mRNA expression in a process that involves the PXR transcription factor. Toxicol. Lett. 2011, 205, 146–153. [Google Scholar] [CrossRef] [PubMed]
- Yen, C.H.; Hung, J.H.; Ueng, Y.F.; Liu, S.P.; Chen, S.Y.; Liu, H.H.; Chou, T.Y.; Tsai, T.F.; Darbha, R.; Hsieh, L.L.; et al. Glycine N-methyltransferase affects the metabolism of aflatoxin B1 and blocks its carcinogenic effect. Toxicol. Appl. Pharmacol. 2009, 235, 296–304. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.H.; Chen, K.H.; Shih, Y.P.; Lui, W.Y.; Wong, F.H.; Chen, Y.M. Characterization of reduced expression of glycine N-methyltransferase in cancerous hepatic tissues using two newly developed monoclonal antibodies. J. Biomed. Sci. 2003, 10, 87–97. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.H.; Shiota, M.; Kuroiwa, K.; Naito, S.; Oda, Y. The important role of glycine N-methyltransferase in the carcinogenesis and progression of prostate cancer. Mod. Pathol. 2011, 24, 1272–1280. [Google Scholar] [CrossRef] [PubMed]
- Meyer zu Schwabedissen, H.E.; Tirona, R.G.; Yip, C.S.; Ho, R.H.; Kim, R.B. Interplay between the Nuclear Receptor Pregnane X Receptor and the Uptake Transporter Organic Anion Transporter Polypeptide 1A2 Selectively Enhances Estrogen Effects in Breast Cancer. Cancer Res. 2008, 68, 9338–9347. [Google Scholar] [CrossRef] [PubMed]
- Pondugula, S.R.; Pavek, P.; Mani, S. Pregnane X Receptor and Cancer: Context-Specificity is Key. Nucl. Recept. Res. 2016, 3, 101198:1–101198:12. [Google Scholar] [CrossRef] [PubMed]
- Casey, S.C.; Nelson, E.L.; Turco, G.M.; Janes, M.R.; Fruman, D.A.; Blumberg, B. B-1 Cell Lymphoma in Mice Lacking the Steroid and Xenobiotic Receptor, SXR. Mol. Endocrinol. 2011, 25, 933–943. [Google Scholar] [CrossRef] [PubMed]
- Yue, X.; Akahira, J.; Utsunomiya, H.; Miki, Y.; Takahashi, N.; Niikura, H.; Ito, K.; Sasano, H.; Okamura, K.; Yaegashi, N. Steroid and Xenobiotic Receptor (SXR) as a possible prognostic marker in epithelial ovarian cancer. Pathol. Int. 2010, 60, 400–406. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Venkatesh, M.; Li, H.; Goetz, R.; Mukherjee, S.; Biswas, A.; Zhu, L.; Kaubisch, A.; Wang, L.; Pullman, J.; et al. Pregnane X receptor activation induces FGF19-dependent tumor aggressiveness in humans and mice. J. Clin. Investig. 2011, 121, 3220–3232. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Duan, J.; Qu, Y.; Deng, T.; Liu, R.; Zhang, L.; Bai, M.; Li, J.; Ning, T.; Ge, S.; et al. Onco-miR-24 regulates cell growth and apoptosis by targeting BCL2L11 in gastric cancer. Protein Cell 2016, 7, 141–151. [Google Scholar] [CrossRef] [PubMed]
- Du, W.W.; Fang, L.; Li, M.; Yang, X.; Liang, Y.; Peng, C.; Qian, W.; O’Malley, Y.Q.; Askeland, R.W.; Sugg, S.L.; et al. MicroRNA miR-24 enhances tumor invasion and metastasis by targeting PTPN9 and PTPRF to promote EGF signaling. J. Cell Sci. 2013, 126, 1440–1453. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Wang, Y.; Liang, H.; Fan, Q.; Zhu, R.; Cui, J.; Zhang, W.; Zen, K.; Zhang, C.Y.; Hou, D.; et al. miR-23a/b promote tumor growth and suppress apoptosis by targeting PDCD4 in gastric cancer. Cell. Death Dis. 2017, 8, e3059:1–e3059:10. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wei, W.; Sarkar, F.H. miR-23a, a critical regulator of “migR” ation and metastasis in colorectal cancer. Cancer Discov. 2012, 2, 489–491. [Google Scholar] [CrossRef] [PubMed]
- Roufayel, R.; Kadry, S. Expression of miR-23a by apoptotic regulators in human cancer: A review. Cancer Biol. Ther. 2017, 18, 269–276. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Wang, H.; Yang, Y.; Guo, S.; Zhang, W.; Liu, Y.; Yi, X.; Ma, J.; Zhao, T.; Liu, L.; et al. Down-regulated miR-23a Contributes to the Metastasis of Cutaneous Melanoma by Promoting Autophagy. Theranostics 2017, 7, 2231–2249. [Google Scholar] [CrossRef] [PubMed]
- Xiang, J.; Hang, J.B.; Che, J.M.; Li, H.C. MiR-25 is up-regulated in non-small cell lung cancer and promotes cell proliferation and motility by targeting FBXW7. Int. J. Clin. Exp. Pathol. 2015, 8, 9147–9153. [Google Scholar] [PubMed]
- Ren, C.; Wang, W.; Han, C.; Chen, H.; Fu, D.; Wang, D.; Shen, M. High expression of mir-25 predicting poor prognosis in gastric cancer. Transl. Surg. 2016, 1, 69–74. [Google Scholar] [CrossRef]
- Zhang, H.; Zuo, Z.; Lu, X.; Wang, L.; Wang, H.; Zhu, Z. MiR-25 regulates apoptosis by targeting Bim in human ovarian cancer. Oncol. Rep. 2012, 27, 594–598. [Google Scholar] [PubMed]
- Chen, H.; Pan, H.; Qian, Y.; Zhou, W.; Liu, X. MiR-25-3p promotes the proliferation of triple negative breast cancer by targeting BTG2. Mol. Cancer. 2018, 17, 4. [Google Scholar] [CrossRef] [PubMed]
- Zoni, E.; van der Horst, G.; van de Merbel, A.F.; Chen, L.; Rane, J.K.; Pelger, R.C.; Collins, A.T.; Visakorpi, T.; Snaar-Jagalska, B.E.; Maitland, N.J.; et al. miR-25 Modulates Invasiveness and Dissemination of Human Prostate Cancer Cells via Regulation of αv- and α6-Integrin Expression. Cancer Res. 2015, 75, 2326–2336. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, D.C.; Madeira da Silva, L.; Zhang, W.; Liu, Z.; Arora, R.; Lim, S.; Schuler, A.M.; McClellan, S.; Andrews, J.F.; Kahn, A.G.; et al. ErbB2-intronic MicroRNA-4728: A novel tumor suppressor and antagonist of oncogenic MAPK signaling. Cell Death Dis. 2015, 6, e1742:1–e1742:8. [Google Scholar] [CrossRef] [PubMed]
- Pekow, J.; Hutchison, A.L.; Meckel, K.; Harrington, K.; Deng, Z.; Talasila, N.; Rubin, D.T.; Hanauer, S.B.; Hurst, R.; Umanskiy, K.; et al. miR-4728-3p Functions as a Tumor Suppressor in Ulcerative Colitis-associated Colorectal Neoplasia Through Regulation of Focal Adhesion Signaling. Inflamm. Bowel Dis. 2017, 23, 1328–1337. [Google Scholar] [CrossRef] [PubMed]
- Yan, S.; Jiang, Y.; Liang, C.; Cheng, M.; Jin, C.; Duan, Q.; Xu, D.; Yang, L.; Zhang, X.; Ren, B.; et al. Exosomal miR-6803-5p as potential diagnostic and prognostic marker in colorectal cancer. J. Cell. Biochem. 2018, 119, 4113–4119. [Google Scholar] [CrossRef] [PubMed]
| Pathway | Proteins |
|---|---|
| Linoleic acid metabolism | CYP1A2, CYP3A4 |
| Steroid hormone biosynthesis | CYP1A2, CYP3A4 |
| Retinol metabolism | CYP1A2, CYP3A4 |
| Drug metabolism—cytochrome P450 | CYP1A2, CYP3A4 |
| Metabolism of xenobiotics by cytochrome P450 | CYP1A2, CYP3A4 |
| Chemical carcinogenesis | CYP1A2, CYP3A4 |
| Nuclear Receptors in Lipid Metabolism and Toxicity | CYP3A4, NR1I2 |
| FoxO signalling pathway | AKT1, MDM2, CAT, IRS1, IRS2, MAPK1, MAPK3 |
| Thyroid hormone signalling pathway | AKT1, MDM2, SRC, MAPK1, MAPK3, TP53 |
| Neutrophin signalling pathway | AKT1, IRS1, IRS2, MAPK1, MAPK3, TP53 |
| Insulin signalling pathway | AKT1, IRS1, IRS2, MAPK1 MAPK3 |
| mTOR signalling pathway | AKT1, IRS1, MAPK1, MAPK3 |
| VEGF signalling pathway | AKT1, SRC, MAPK1, MAPK3 |
| cGMP-PKG signalling pathway | AKT1, IRS1, IRS2, MAPK1, MAPK3 |
| Prolactin signalling pathway | AKT1, SRC, MAPK1, MAPK3 |
| PI3K-Akt signalling pathway | AKT1, MDM2, IRS1, IRS2, MAPK1, MAPK3, TP53 |
| ErbB signalling pathway | AKT1, SRC, MAPK1, MAPK3 |
| HIF-1 signalling pathway | AKT1, MAPK1, MAPK3, PDK1 |
| Oestrogensignalling pathway | AKT1, SRC, MAPK1, MAPK3 |
| Sphingolipid signalling pathway | AKT1, MAPK1, MAPK3, TP53 |
| Platelet activation | AKT1, SRC, MAPK1, MAPK3 |
| Aldosterone-regulated sodium reabsorption | IRS1, MAPK1, MAPK3 |
| Chemokine signalling pathway | AKT1, SRC, MAPK1, MAPK3 |
| Regulation of lipolysis in adipocytes | AKT1, IRS1, IRS2 |
| Focal adhesion | AKT1, SRC, MAPK1, MAPK3 |
| Rap1 signalling pathway | AKT1, SRC, MAPK1, MAPK3 |
| Fc gamma R-mediated phagocytosis | AKT1, MAPK1, MAPK3 |
| MAPK signalling pathway | AKT1, MAPK1, MAPK3, TP53 |
| Gap junction | SRC, MAPK1, MAPK3 |
| GnRH signalling pathway | SRC, MAPK1, MAPK3 |
| T cell receptor signalling pathway | AKT1, MAPK1, MAPK3 |
| TNF signalling pathway | AKT1, MAPK1, MAPK3 |
| Toll-like receptor signalling pathway | AKT1, MAPK1, MAPK3 |
| AMPK signalling pathway | AKT1, IRS1, IRS2 |
| cAMP signalling pathway | AKT1, MAPK1, MAPK3 |
| miRNA | Gene |
|---|---|
| hsa-miR24-3p | CYP3A4, IRS1, IRS2, MAPK1, MAPK3, MDM2, NR1I2, OGG1, PDK1 |
| hsa-miR-6778-5p | CYP1A2, CYP3A4, IRS1, MDM2, NR1I2, OGG1, PDK1, SRC, TP53 |
| hsa-miR-6514-3p | GNMT, IRS1, IRS2, MAPK1, MDM2, OGG1, PDK1, TP53 |
| hsa-miR-5010-5p | AKT1, IRS1, IRS2, MAPK1, MAPK3, MDM2, PDK1, SRC |
| hsa-miR-23a-5p | AKT1, CYP3A4, MAPK1, NR1I2, OGG1, PDK1, SRC, TP53 |
| hsa-miR-25-5p | AKT1, GNMT, MAPK1, MDM2, OGG1, PDK1, SRC, TP53 |
| hsa-miR-6792-5p | CYP1A2, MAPK1, MAPK3, MDM2, OGG1, PDK1, SRC, TP53 |
| hsa-miR-6866-5p | GNMT, IRS1, MAPK1, MDM2, NR1I2, OGG1, SRC, TP53 |
| hsa-miR-4728-5p | AKT1, IRS1, IRS2, MAPK1, MAPK3, MDM2, OGG1, SRC |
| hsa-miR-6825-5p | AKT1, CYP1A2, MAPK1, MDM2, NR1I2, OGG1, SRC, TP53 |
| hsa-miR-6803-3p | CYP1A2, CYP3A4, IRS2, MAPK1, MAPK3, OGG1, SRC, TP53 |
| hsa-miR-6794-5p | CYP3A4, GNMT, MAPK1, MAPK3, MDM2, NR1I2, SRC, TP53 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marchese, S.; Polo, A.; Ariano, A.; Velotto, S.; Costantini, S.; Severino, L. Aflatoxin B1 and M1: Biological Properties and Their Involvement in Cancer Development. Toxins 2018, 10, 214. https://doi.org/10.3390/toxins10060214
Marchese S, Polo A, Ariano A, Velotto S, Costantini S, Severino L. Aflatoxin B1 and M1: Biological Properties and Their Involvement in Cancer Development. Toxins. 2018; 10(6):214. https://doi.org/10.3390/toxins10060214
Chicago/Turabian StyleMarchese, Silvia, Andrea Polo, Andrea Ariano, Salvatore Velotto, Susan Costantini, and Lorella Severino. 2018. "Aflatoxin B1 and M1: Biological Properties and Their Involvement in Cancer Development" Toxins 10, no. 6: 214. https://doi.org/10.3390/toxins10060214
APA StyleMarchese, S., Polo, A., Ariano, A., Velotto, S., Costantini, S., & Severino, L. (2018). Aflatoxin B1 and M1: Biological Properties and Their Involvement in Cancer Development. Toxins, 10(6), 214. https://doi.org/10.3390/toxins10060214






