Chemometric Analysis of the Volatile Compounds Generated by Aspergillus carbonarius Strains Isolated from Grapes and Dried Vine Fruits
Abstract
:1. Introduction
2. Results and Discussion
2.1. Toxigenic Investigation of A. carbonarius Strains
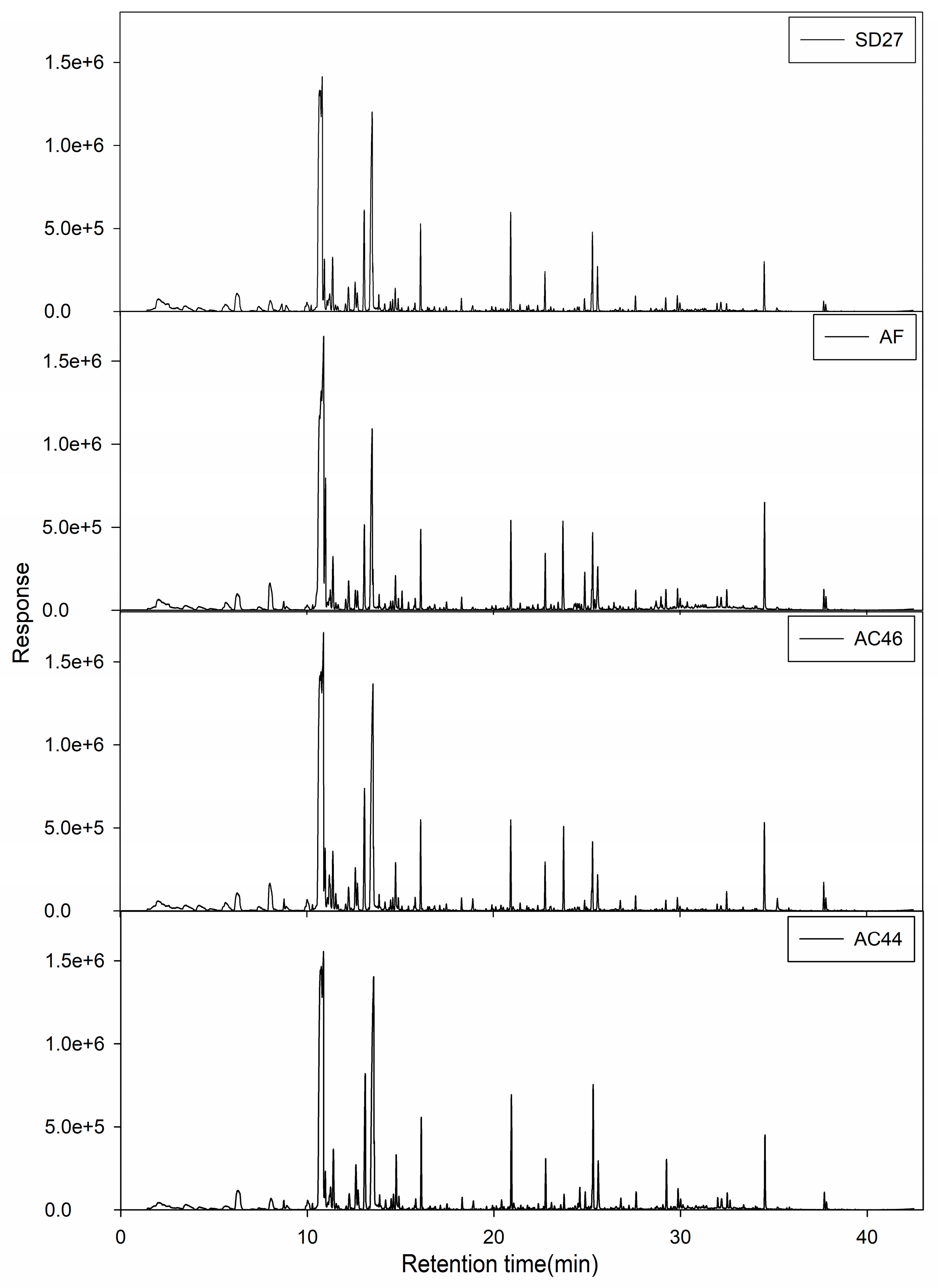
2.2. GC-MS Profiles of Different Toxigenic Strains
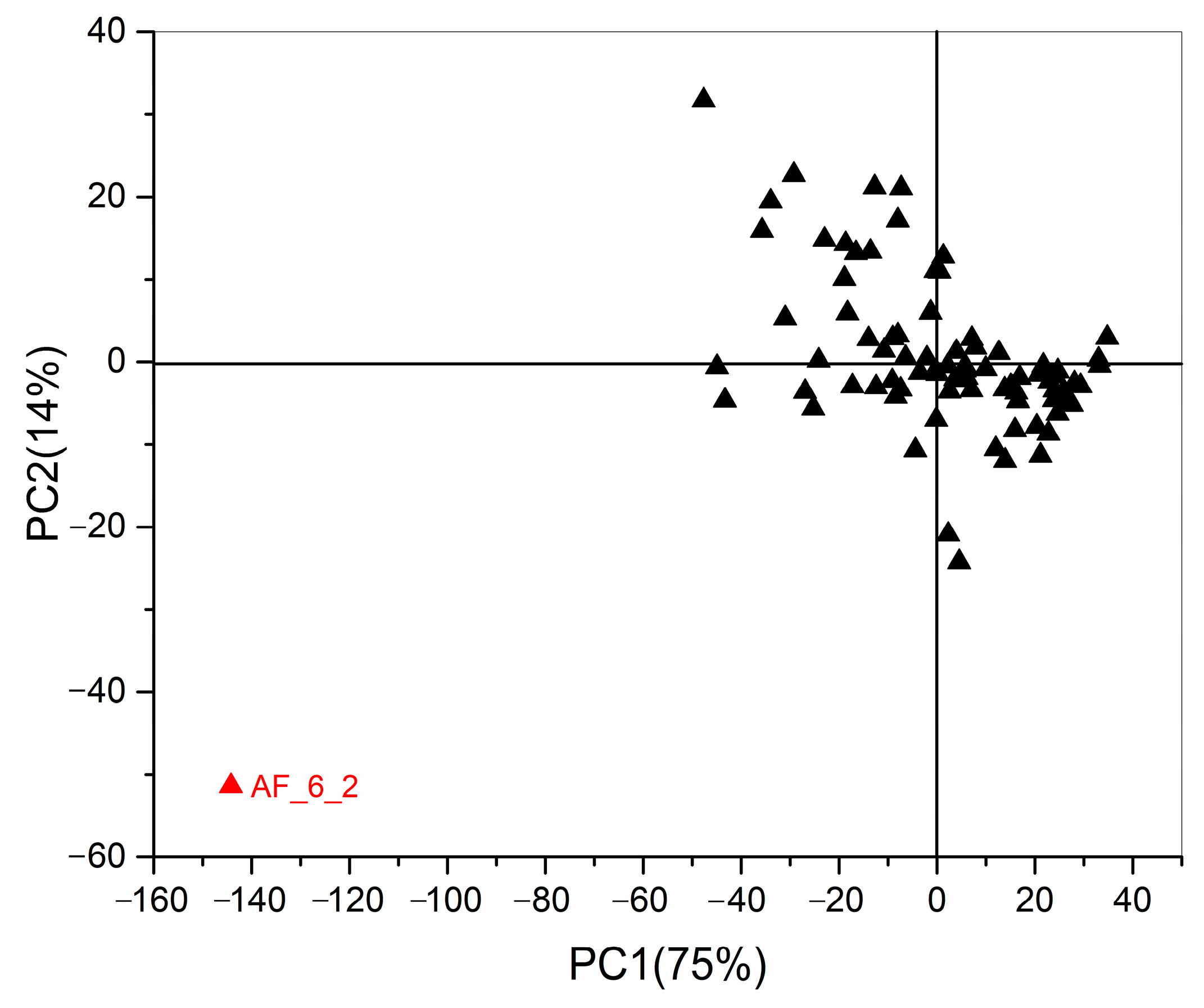
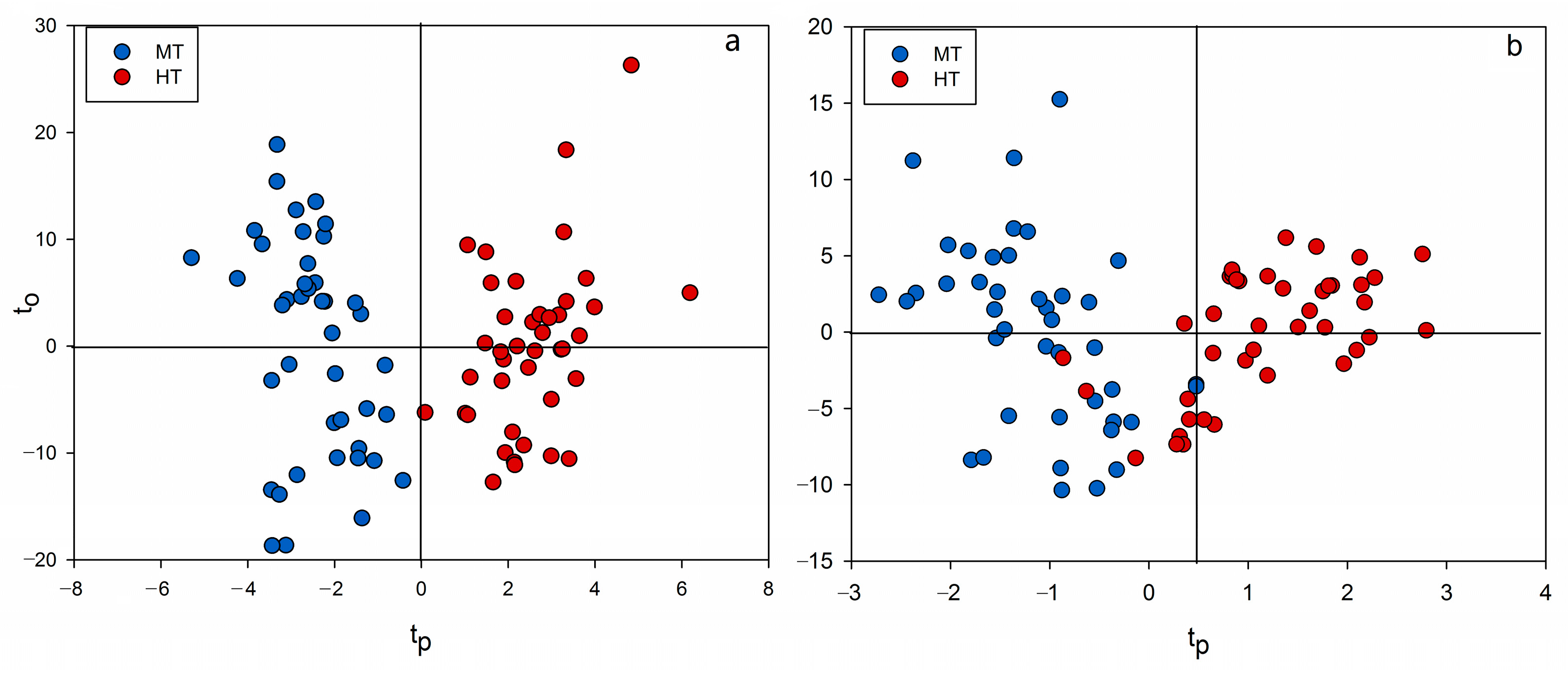
2.3. Chemometrics for Analyzing the Differences of Two Group Strains
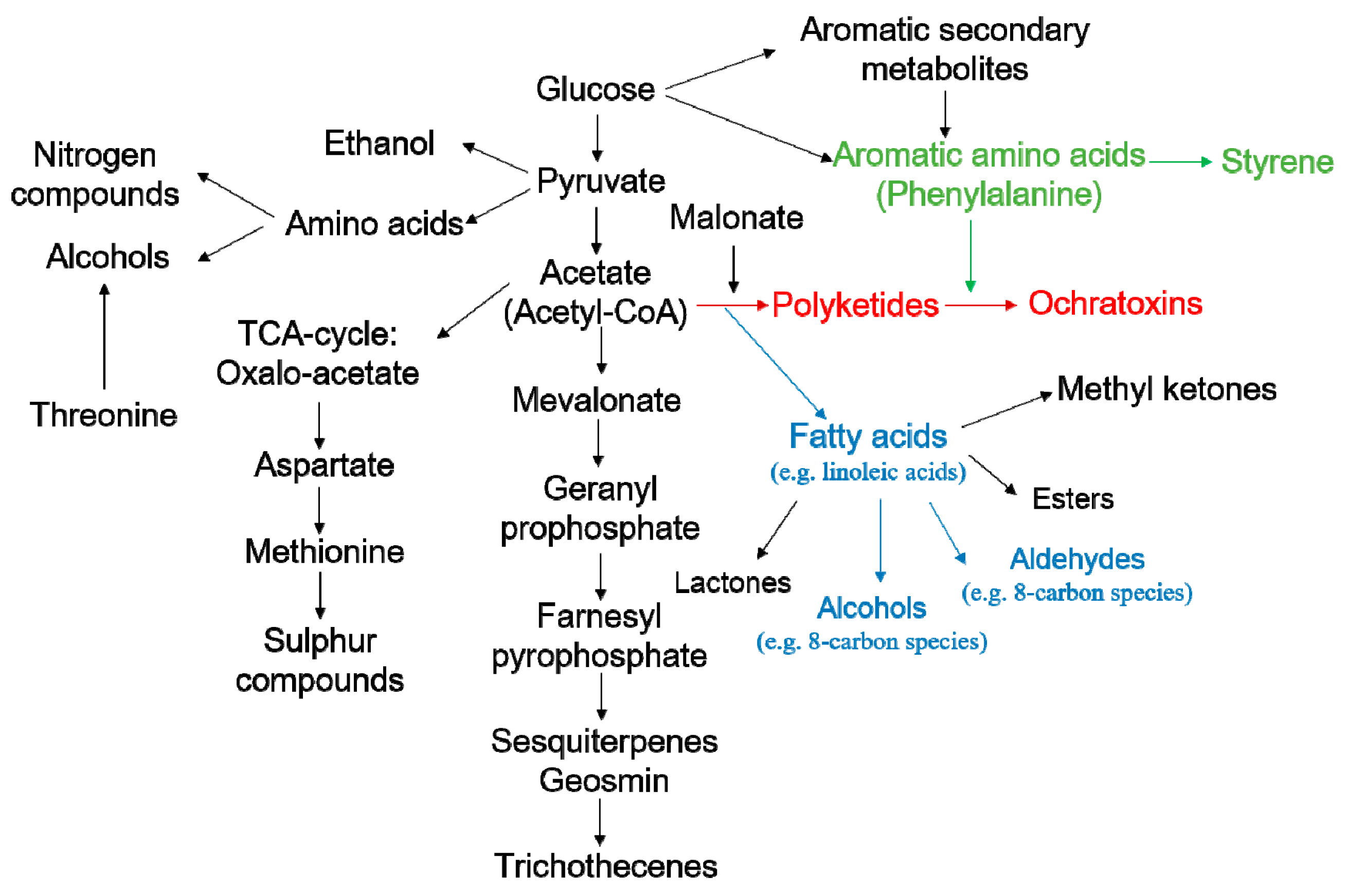
2.4. Discovery of Potential Markers of HT and MT Strains
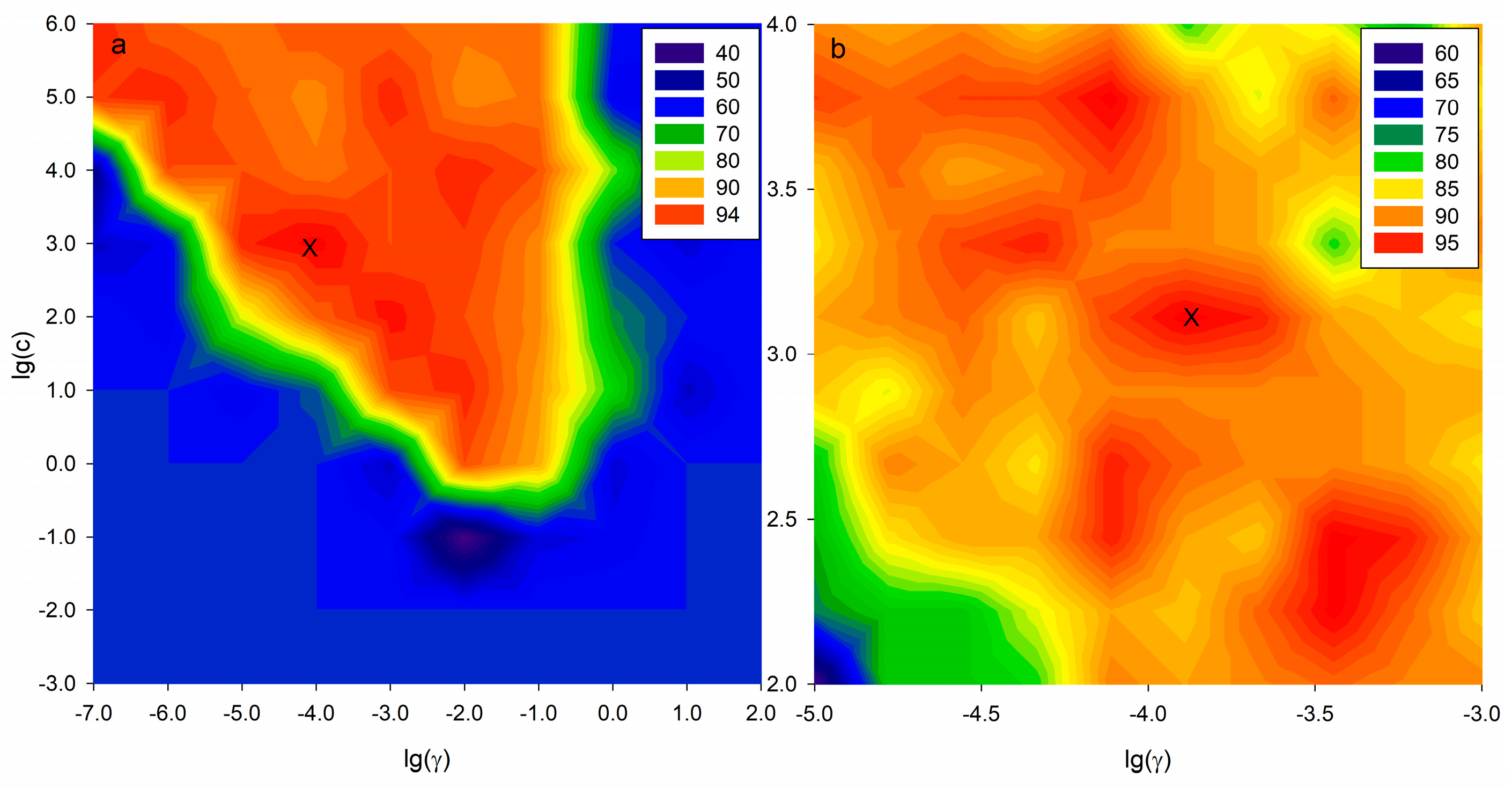
2.5. SVM-C Pattern Recognition Based on Potential Markers
3. Conclusions
4. Materials and Methods
4.1. Chemicals
4.2. Fungi and Cultivation
4.3. GC-MS Analysis
4.4. Ochratoxin A Analysis
4.5. Data Processing
4.6. Chemometrics Analysis
4.7. Support Vector Machine Classification
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bellí, N.; Bau, M.; Marín, S.; Abarca, M.L.; Ramos, A.J.; Bragulat, M.R. Mycobiota and ochratoxin A producing fungi from Spanish wine grapes. Int. J. Food Microbiol. 2006, 111, S40–S45. [Google Scholar] [CrossRef] [PubMed]
- Valero, A.; Marín, S.; Ramos, A.J.; Sanchis, V. Ochratoxin A-producing species in grapes and sun-dried grapes and their relation to ecophysiological factors. Lett. Appl. Microbiol. 2005, 41, 196–201. [Google Scholar] [CrossRef] [PubMed]
- Logrieco, A.; Moretti, A.; Perrone, G.; Mulè, G. Biodiversity of complexes of mycotoxigenic fungal species associated with Fusarium ear rot of maize and Aspergillus rot of grape. Int. J. Food Microbiol. 2007, 119, 11–16. [Google Scholar] [CrossRef] [PubMed]
- Magnoli, C.; Astoreca, A.; Ponsone, L.; Combina, M.; Palacio, G.; Rosa, C.A.; Dalcero, A.M. Survey of mycoflora and ochratoxin A in dried vine fruits from Argentina markets. Lett. Appl. Microbiol. 2004, 39, 326–331. [Google Scholar] [CrossRef] [PubMed]
- Jeleń, H.H.; Grabarkiewicz-Szczȩsna, J. Volatile compounds of aspergillus strains with different abilities to produce ochratoxin A. J. Agric. Food Chem. 2005, 53, 1678–1683. [Google Scholar] [CrossRef] [PubMed]
- Demyttenaere, J.C.R.; Moriña, R.M.; De Kimpe, N.; Sandra, P. Use of headspace solid-phase microextraction and headspace sorptive extraction for the detection of the volatile metabolites produced by toxigenic Fusarium species. J. Chromatogr. A 2004, 1027, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Zeringue, H.J.; Bhatnagar, D.; Cleveland, T.E. C15H24 Volatile compounds unique to aflatoxigenic strains of aspergillus flavus. Appl. Environ. Microbiol. 1993, 59, 2264–2270. [Google Scholar] [PubMed]
- Karlshøj, K.; Larsen, T.O. Differentiation of species from the Penicillium roqueforti group by volatile metabolite profiling. J. Agric. Food Chem. 2005, 53, 708–715. [Google Scholar] [CrossRef] [PubMed]
- Fischer, G.; Schwalbe, R.; Möller, M.; Ostrowski, R.; Dott, W. Species-specific production of microbial volatile organic compounds (MVOC) by airborne fungi from a compost facility. Chemosphere 1999, 39, 795–810. [Google Scholar] [CrossRef]
- Szponar, B.; Larsson, L. Determination of microbial colonisation in water-damaged buildings using chemical marker analysis by gas chromatography-mass spectrometry. Indoor Air. 2000, 10, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Elke, K.; Begerow, J.; Oppermann, H.; Krämer, U.; Jermann, E.; Dunemann, L. Determination of selected microbial volatile organic compounds by diffusive sampling and dual-column capillary GC-FID—A new feasible approach for the detection of an exposure to indoor mould fungi? J. Environ. Monit. 1999, 1, 445–452. [Google Scholar] [CrossRef] [PubMed]
- Demyttenaere, J.C.R.; Moriña, R.M.; Sandra, P. Monitoring and fast detection of mycotoxin-producing fungi based on headspace solid-phase microextraction and headspace sorptive extraction of the volatile metabolites. J. Chromatogr. A 2003, 985, 127–135. [Google Scholar] [CrossRef]
- Jelén, H.H.; Mirocha, C.J.; Wasowicz, E.; Kamiński, E. Production of volatile sesquiterpenes by Fusarium sambucinum strains with different abilities to synthesize trichothecenes. Appl. Environ. Microbiol. 1995, 61, 3815–3820. [Google Scholar] [PubMed]
- Christensen, J.H.; Mortensen, J.; Hansen, A.B.; Andersen, O. Chromatographic preprocessing of GC–MS data for analysis of complex chemical mixtures. J. Chromatogr. A 2005, 1062, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Yi, L.; Dong, N.; Yun, Y.; Deng, B.; Ren, D.; Liu, S.; Liang, Y. Chemometric methods in data processing of mass spectrometry-based metabolomics: A review. Anal. Chim. Acta 2016, 914, 17–34. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, D.R.; Farag, A. Volatiles and primary metabolites profiling in two Hibiscus sabdariffa (Roselle) cutlivars via headspace SPME-GC-MS and chemometrics. Food Res. Int. 2015, 78, 327–335. [Google Scholar] [CrossRef]
- Aliakbarzadeh, G.; Parastar, H.; Sereshti, H. Classification of gas chromatographic fingerprints of saffron using partial least squares discriminant analysis together with different variable selection methods. Chemometr. Intell. Lab. 2016, 158, 165–173. [Google Scholar] [CrossRef]
- Fu, H.Y.; Guo, J.W.; Yu, Y.J.; Li, H.D.; Cui, H.P.; Liu, P.P.; Wang, B.; Wang, S.; Lu, P. A simple multi-scale Gaussian smoothing-based strategy for automatic chromatographic peak extraction. J. Chromatogr. A 2016, 1452, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Fu, H.Y.; Li, H.D.; Yu, Y.J.; Wang, B.; Lu, P.; Cui, H.P.; Liu, P.P.; She, Y.B. Simple automatic strategy for background drift correction in chromatographic data analysis. J. Chromatogr. A 2016, 1449, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Q.X.; Fu, H.Y.; Li, H.D.; Wang, B.; Peng, C.H.; Wang, S.; Cai, J.L.; Liu, S.F.; Zhang, X.B.; Yu, Y.J. Automatic time-shift alignment method for chromatographic data analysis. Sci. Rep. 2017, 7, 256. [Google Scholar] [CrossRef] [PubMed]
- González-Domínguez, R.; García-Barrera, T.; Vitorica, J.; Gómez-Ariza, J.L. Region-specific metabolic alterations in the brain of the APP/PS1 transgenic mice of Alzheimer’s disease. Biochim. Biophys. Acta 2014, 1842, 2395–2402. [Google Scholar] [CrossRef] [PubMed]
- González-Domínguez, R.; García-Barrera, T.; Gómez-Ariza, J.L. Metabolite profiling for the identification of altered metabolic pathways in Alzheimer’s disease. J. Pharm. Biomed. Anal. 2015, 107, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Gromski, P.S.; Muhamadali, H.; Ellis, D.I.; Xu, Y.; Correa, E.; Turner, M.L.; Goodacre, R. A tutorial review: Metabolomics and partial least squares-discriminant analysis-a marriage of convenience or a shotgun wedding. Anal. Chim. Acta 2015, 879, 10–23. [Google Scholar] [CrossRef] [PubMed]
- Wold, S.; Sjöström, M.; Eriksson, L. PLS-regression: A basic tool of chemometrics. Chemometr. Intell. Lab. 2001, 58, 109–130. [Google Scholar] [CrossRef]
- Trygg, J.; Wold, S. Orthogonal projections to latent structures (O-PLS). J. Chemom. 2002, 16, 119–128. [Google Scholar] [CrossRef]
- Boser, B.E.; Guyon, I.M.; Vapnik, V.N. A training algorithm for optimal margin classifiers. Workshop Comput. Learn. Theory 1992, 5, 144–152. [Google Scholar] [CrossRef]
- Vapnik, V.N. Statistical Learning Theory; Wiley: New York, NY, USA, 1998; ISBN 978-0-471-03003-4. [Google Scholar]
- Zhou, X.; Wang, Y.; Yun, Y.; Xia, Z.; Lu, H.; Luo, J.; Liang, Y. A potential tool for diagnosis of male infertility: Plasma metabolomics based on GC–MS. Talanta 2016, 147, 82–89. [Google Scholar] [CrossRef] [PubMed]
- Jonsson, P.; Gullberg, J.; Nordström, A.; Kusano, M.; Kowalczyk, M.; Sjöström, M.; Moritz, T. A strategy for identifying differences in large series of metabolomic samples analyzed by GC/MS. Anal. Chem. 2004, 76, 1738–1745. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Lin, S.S.; Yang, J.T.; Tang, L.J.; Yu, R.Q. Detection of inborn errors of metabolism utilizing GC-MS urinary metabolomics coupled with a modified orthogonal partial least squares discriminant analysis. Talanta 2017, 165, 545–552. [Google Scholar] [CrossRef] [PubMed]
- Westerhuis, J.A.; van Velzen, E.J.J.; Hoefsloot, H.C.J.; Smilde, A.K. Multivariate paired data analysis: Multilevel PLSDA versus OPLSDA. Metabolomics 2010, 6, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Kamal, G.M.; Wang, X.; Yuan, B.; Wang, J.; Sun, P.; Zhang, X.; Liu, M. Compositional differences among Chinese soy sauce types studied by 13C NMR spectroscopy coupled with multivariate statistical analysis. Talanta 2016, 158, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Ledauphin, J.; Le Milbeau, C.; Barillier, D.; Hennequin, D. Differences in the volatile compositions of French labeled brandies (Armagnac, Calvados, Cognac, and Mirabelle) using GC-MS and PLS-DA. J. Agric. Food Chem. 2010, 58, 7782–7793. [Google Scholar] [CrossRef] [PubMed]
- Rajalahti, T.; Arneberg, R.; Berven, F.S.; Myhr, K.M.; Ulvik, R.J.; Kvalheim, O.M. Biomarker discovery in mass spectral profiles by means of selectivity ratio plot. Chemometr. Intell. Lab. 2009, 95, 35–48. [Google Scholar] [CrossRef]
- Furey, T.S.; Cristianini, N.; Duffy, N.; Bednarski, D.W.; Schummer, M.; Haussler, D. Support vector machine classification and validation of cancer tissue samples using microarray expression data. Bioinformatics 2000, 16, 906–914. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.J.; Du, W.; Fu, H.Y.; Jiang, J.H.; Wu, H.L.; Shen, G.L.; Yu, R.Q. New variable selection method using interval segmentation purity with application to blockwise kernel transform support vector machine classification of high-dimensional microarray data. J. Chem. Inf. Model. 2009, 49, 2002–2009. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Cheng, Z.; Ma, L.; Li, J. A study on accumulation of volatile organic compounds during ochratoxin a biosynthesis and characterization of the correlation in Aspergillus carbonarius isolated from grape and dried vine fruit. Food Chem. 2017, 227, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Combet, E.; Henderson, J.; Eastwood, D.C.; Burton, K.S. Eight-carbon volatiles in mushrooms and fungi: Properties, analysis, and biosynthesis. Mycoscience 2006, 47, 317–326. [Google Scholar] [CrossRef]
- Kaminski, E.; Stawicki, S.; Wasowicz, E. Volatile flavor compounds produced by molds of Aspergillus, Penicillium, and Fungi imperfecti. Appl. Microbiol. 1974, 27, 1001–1004. [Google Scholar] [PubMed]
- Kamiński, E.; Libbey, L.M.; Stawicki, S.; Wasowicz, E. Identification of the predominant volatile compounds produced by Aspergillus flavus. Appl. Microbiol. 1972, 24, 721–726. [Google Scholar] [PubMed]
- Magan, N.; Evans, P. Volatiles as an indicator of fungal activity and differentiation between species, and the potential use of electronic nose technology for early detection of grain spoilage. J. Stored Prod. Res. 2000, 36, 319–340. [Google Scholar] [CrossRef]
- Viana, F.; Gil, J.V.; Vallés, S.; Manzanares, P. Increasing the levels of 2-phenylethyl acetate in wine through the use of a mixed culture of Hanseniaspora osmophila and Saccharomyces cerevisiae. Int. J. Food Microbiol. 2009, 135, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Herbert, R.B. The Biosynthesis of Secondary Metabolites; Chapman & Hall: London, UK, 1981; Volume 261, p. 212. [Google Scholar]
- Adda, J.; Dekimpe, J.; Vassal, L.; Spinnler, H.E. Production de styrène par Penicillium camemberti Thom. Le Lait 1989, 69, 115–120. [Google Scholar] [CrossRef]
- Larsen, T.O.; Frisvad, J.C. Characterization of volatile metabolites from 47 Penicillium taxa. Mycol. Res. 1995, 99, 1153–1166. [Google Scholar] [CrossRef]
- Sinha, M.; Sørensen, A.; Ahamed, A.; Ahring, B.K. Production of hydrocarbons by Aspergillus carbonarius ITEM 5010. Fungal Biol. 2015, 119, 274–282. [Google Scholar] [CrossRef] [PubMed]
- Börjesson, T.; Stöllman, U.; Schnürer, J. Volatile metabolites produced by six fungal species compared with other indicators of fungal growth on cereal grains. Appl. Environ. Microbiol. 1992, 58, 2599–2605. [Google Scholar] [PubMed]
- Gallo, A.; Bruno, K.S.; Solfrizzo, M.; Perrone, G.; Mulè, G.; Visconti, A.; Baker, S.E. New Insight into the Ochratoxin A Biosynthetic Pathway through Deletion of a Nonribosomal Peptide Synthetase Gene in Aspergillus carbonarius. Appl. Environ. Microbiol. 2012, 78, 8208–8218. [Google Scholar] [CrossRef] [PubMed]
- Assaf, S.; Hadar, Y.; Dosoretz, C.G. 1-Octen-3-ol and 13-hydroperoxylinoleate are products of distinct pathways in the oxidative breakdown of linoleic acid by Pleurotus pulmonarius. Enzyme Microb. Tech. 1997, 21, 484–490. [Google Scholar] [CrossRef]
- Assaf, S.; Hadar, Y.; Dosoretz, C.G. Biosynthesis of 13-hydroperoxylinoleate, 10-oxo-8-decenoic acid and 1-octen-3-ol from linoleic acid by a mycelial-pellet homogenate of Pleurotus pulmonarius. J. Agric. Food Chem. 1995, 43, 2173–2178. [Google Scholar] [CrossRef]
- Buśko, M.; Kulik, T.; Ostrowska, A.; Góral, T.; Perkowski, J. Quantitative volatile compound profiles in fungal cultures of three different Fusarium graminearum chemotypes. FEMS Microbiol. Lett. 2014, 359, 85–93. [Google Scholar] [CrossRef] [PubMed]
- Kizis, D.; Natskoulis, P.; Nychas, G.J.E.; Panagou, E.Z. Biodiversity and ITS-RFLP characterisation of Aspergillus section Nigri isolates in grapes from four traditional grape-producing areas in Greece. PLoS ONE 2014, 9, e93923. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Li, J.; Cheng, Z.; Zhou, Z.; Ma, L. High-performance liquid chromatography-tandem mass spectrometry method for simultaneous detection of ochratoxin A and relative metabolites in Aspergillus species and dried vine fruits. Food Addit. Contam. A Chem. Anal. Control Expos. Risk Assess. 2016, 33, 1355–1366. [Google Scholar] [CrossRef]
- Zhang, X.; Li, J.; Zong, N.; Zhou, Z.; Ma, L. Ochratoxin A in dried vine fruits from Chinese markets. Food Addit. Contam. B Surveill. 2014, 7, 157–161. [Google Scholar] [CrossRef] [PubMed]
- Katajamaa, M.; Orešič, M. Data processing for mass spectrometry-based metabolomics. J. Chromatogr. A 2007, 1158, 318–328. [Google Scholar] [CrossRef] [PubMed]
- Tautenhahn, R.; Patti, G.J.; Rinehart, D.; Siuzdak, G. XCMS Online: A web-based platform to process untargeted metabolomic data. Anal. Chem. 2012, 84, 5035–5039. [Google Scholar] [CrossRef] [PubMed]
- Du, P.; Kibbe, W.A.; Lin, S.M. Improved peak detection in mass spectrum by incorporating continuous wavelet transform-based pattern matching. Bioinformatics 2006, 22, 2059–2065. [Google Scholar] [CrossRef] [PubMed]
- Mastrangelo, A.; Ferrarini, A.; Rey-Stolle, F.; García, A.; Barbas, C. From sample treatment to biomarker discovery: A tutorial for untargeted metabolomics based on GC-(EI)-Q-MS. Anal. Chim. Acta 2015, 900, 21–35. [Google Scholar] [CrossRef] [PubMed]
- Lippolis, V.; Ferrara, M.; Cervellieri, S.; Damascelli, A.; Epifani, F.; Pascale, M.; Perrone, G. Rapid prediction of ochratoxin A-producing strains of Penicillium on dry-cured meat by MOS-based electronic nose. Int. J. Food Microbiol. 2016, 218, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Wiklund, S.; Johansson, E.; Sjöström, L.; Mellerowicz, E.J.; Edlund, U.; Shockcor, J.P.; Gottfries, J.; Moritz, T.; Trygg, J. Visualization of GC/TOF-MS-based metabolomics data for identification of biochemically interesting compounds using OPLS class models. Anal. Chem. 2008, 80, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Chong, I.G.; Jun, C.H. Performance of some variable selection methods when multicollinearity is present. Chemometr. Intell. Lab. 2015, 78, 103–112. [Google Scholar] [CrossRef]
- Balabin, R.M.; Lomakina, E.I. Support vector machine regression (SVR/LS-SVM)—An alternative to neural networks (ANN) for analytical chemistry? Comparison of nonlinear methods on near infrared (NIR) spectroscopy data. Analyst 2011, 136, 1703. [Google Scholar] [CrossRef] [PubMed]
- Sanaeifar, A.; Bakhshipour, A.; de la Guardia, M. Prediction of banana quality indices from color features using support vector regression. Talanta 2016, 148, 54–61. [Google Scholar] [CrossRef] [PubMed]
- Yao, X.J.; Panaye, A.; Doucet, J.P.; Chen, H.F.; Zhang, R.S.; Fan, B.T.; Liu, M.C.; Hu, Z.D. Comparative classification study of toxicity mechanisms using support vector machines and radial basis function neural networks. Anal. Chim. Acta 2005, 535, 259–273. [Google Scholar] [CrossRef]
- Kavzoglu, T.; Colkesen, I. A kernel functions analysis for support vector machines for land cover classification. Int. J. Appl. Earth Obs. Geoinf. 2009, 11, 352–359. [Google Scholar] [CrossRef]
- Porto-Figueira, P.; Freitas, A.; Cruz, C.J.; Figueira, J.; Câmara, J.S. Profiling of passion fruit volatiles: An effective tool to discriminate between species and varieties. Food Res. Int. 2015, 77, 408–418. [Google Scholar] [CrossRef]
- Hsu, C.W.; Chang, C.C.; Lin, C.J. A Practical Guide to Support Vector Classification; Technical Report; Department of Computer Science and Information Engineering, National Taiwan University: Taipei, Taiwan, 2003; Volume 67, pp. 1–29. Available online: http://www.csie.ntu.edu.tw/~cjlin/papers/guide/guide.pdf (accessed on 19 May 2016).


| NO. | REF.RI 1 | RI | Name | Identification Methods 2 | Ion 3 | CYA Culture Medium |
|---|---|---|---|---|---|---|
| Alcohols | ||||||
| 1 | 979 | 980 | 1-Octen-3-ol | Std, MS, RI | 57 | AC44, AC46, AF, SD27 |
| 2 | 1069 | 1067 | (E)-2-Octen-1-ol | Std, MS, RI | 57 | AC44, AC46, AF, SD27 |
| 3 | 1069 | 1070 | 1-Octanol | MS, RI | 56 | AC44, AC46, AF, SD27 |
| Aldehydes | ||||||
| 4 | 1001 | 1001 | Octanal | MS, RI | 43 | AC44, AC46, AF, SD27 |
| 5 | 1057 | 1055 | (E)-2-Octenal | Std, MS, RI | 41 | AC44, AC46, AF, SD27 |
| 6 | 1102 | 1103 | Nonanal | MS, RI | 57 | AC44, AC46, AF, SD27 |
| 7 | 1115 | 1107 | (E,E)-2,4-Octadienal | MS, RI | 81 | AC44, AC46, AF, SD27 |
| 8 | 1313 | 1314 | (E,E)-2,4-Decadienal | MS, RI | 81 | AC44, AC46, AF, SD27 |
| Ketones | ||||||
| 9 | 978 | 976 | 1-Octen-3-one | MS, RI | 55 | AC44, AC46, AF, SD27 |
| 10 | 984 | 985 | 3-Octanone | Std, MS, RI | 43 | AC44, AC46, AF, SD27 |
| 11 | 1290 | 1291 | 2-Undecanone | MS, RI | 43 | AC44, AC46, AF, SD27 |
| Esters | ||||||
| 12 | 1092 | 1093 | Methyl benzoate | MS, RI | 105 | AC44, AC46, AF, SD27 |
| 13 | 1120 | 1123 | Methyl octanoate | MS, RI | 74 | AC44, AC46, AF, SD27 |
| 14 | 1255 | 1254 | Methyl-2-phenylacetate | MS, RI | 104 | AC44, AC46, AF, SD27 |
| 15 | 1326 | 1322 | Methyl decanoate | MS, RI | 74 | AC44, AC46, AF, SD27 |
| 16 | 1723 | 1723 | Methyl tetradecanoate | MS, RI | 74 | AC44, AC46, AF, SD27 |
| 17 | 1823 | 1825 | Methyl pentadecanoate | MS, RI | 74 | AC44, AC46, AF, SD27 |
| 18 | 1927 | 1926 | Methyl hexadecanoate | Std, MS, RI | 74 | AC44, AC46, AF, SD27 |
| 19 | 2096 | 2095 | Methyl linoleate | Std, MS, RI | 67 | AC44, AC46, AF, SD27 |
| 20 | 2100 | 2102 | Methyl oleate | MS, RI | 55 | AC44, AC46, AF, SD27 |
| Terpenoids | ||||||
| 21 | 1024 | 1024 | p-Cymene | MS, RI | 119 | AC44, AC46, AF, SD27 |
| 22 | 1028 | 1028 | Limonene | MS, RI | 68 | AC44, AC46, AF, SD27 |
| 23 | 1412 | 1411 | Longifolene | MS, RI | 161 | AC44, AC46, AF, SD27 |
| 24 | 1416 | 1417 | α-Cedrene | Std, MS, RI | 119 | AC44, AC46, AF, SD27 |
| 25 | 1428 | 1426 | β-Cedrene | MS, RI | 161 | AF |
| 26 | 1435 | 1436 | (Z)-Thujopsene | MS, RI | 119 | AC44, AC46, AF, SD27 |
| 27 | 1435 | 1438 | α-Bergamotene | MS, RI | 93 | AC44, AC46, AF, SD27 |
| 28 | 1458 | 1457 | β-Farnesene | Std, MS, RI | 41 | AC44, AC46, SD27 |
| 29 | 1481 | 1481 | β-Chamigrene | Std, MS, RI | 189 | AF |
| 30 | 1505 | 1505 | β-Himachalene | MS, RI | 119 | AF |
| 31 | 1509 | 1510 | Cuparene | Std, MS, RI | 132 | AF |
| 32 | 1563 | 1563 | (E)-Nerolidol | Std, MS, RI | 41 | AC44, AC46, AF, SD27 |
| Hydrocarbons | ||||||
| 33 | 893 | 889 | Styrene | Std, MS, RI | 104 | AC44, AC46, AF, SD27 |
| 34 | 1100 | 1100 | Undecane | Std, MS, RI | 57 | AC44, AC46, AF, SD27 |
| 35 | 1200 | 1199 | Dodecane | Std, MS, RI | 57 | AC44, AC46, AF, SD27 |
| 36 | 1300 | 1299 | Tridecane | Std, MS, RI | 57 | AC44, AC46, AF, SD27 |
| 37 | 1318 | 1326 | Decane, 2,3,5,8-tetramethyl- | MS, RI | 57 | AC44, AC46, AF, SD27 |
| 38 | 1400 | 1400 | Tetradecane | Std, MS, RI | 57 | Internal standard |
| 39 | 1460 | 1462 | Tetradecane, 4-methyl- | MS, RI | 43 | AC44, AC46, AF, SD27 |
| 40 | 1500 | 1499 | Pentadecane | Std, MS, RI | 57 | AC44, AC46, AF, SD27 |
| 41 | 1564 | 1562 | Pentadecane, 2-methyl- | MS, RI | 43 | AC44, AC46, AF, SD27 |
| 42 | 1570 | 1569 | Pentadecane, 3-methyl- | MS, RI | 57 | AC44, AC46, AF, SD27 |
| 43 | 1600 | 1600 | Hexadecane | Std, MS, RI | 57 | AC44, AC46, AF, SD27 |
| 44 | 1649 | 1648 | Pentadecane, 2,6,10-trimethyl- | MS, RI | 57 | AC44, AC46, AF, SD27 |
| 45 | 1666 | 1663 | Hexadecane, 2-methyl- | MS, RI | 57 | AC44, AC46, AF, SD27 |
| 46 | 1700 | 1700 | Heptadecane | Std, MS, RI | 57 | AC44, AC46, AF, SD27 |
| 47 | 1703 | 1706 | Pristan | MS, RI | 57 | AC44, AC46, AF, SD27 |
| 48 | 1765 | 1763 | Heptadecane, 2-methyl- | MS, RI | 57 | AC44, AC46, AF, SD27 |
| 49 | 1770 | 1771 | Heptadecane, 3-methyl- | MS, RI | 57 | AC44, AC46, AF, SD27 |
| 50 | 1800 | 1800 | Octadecane | Std, MS, RI | 57 | AC44, AC46, AF, SD27 |
| 51 | 1806 | 1810 | Phytane | MS, RI | 57 | AC44, AC46, AF, SD27 |
| Others | ||||||
| 52 | 1181 | 1182 | Naphthalene | MS, RI | 128 | AC44, AC46, AF, SD27 |
| 53 | - | 1484 | 3-Furanacetic acid, 4-hexyl-2,5-dihydro-2,5-dioxo- | MS | 126 | AC44, AC46, AF, SD27 |
| NO. | Potential Markers | Retention Time/Min | Ion Information | Relative Content 1 | |
|---|---|---|---|---|---|
| MT | HT | ||||
| 1 | Styrene | 8.001–8.004 | 103, 78, 77, 104, 51, 105 | 0.13–29.77 * | 0.08–13.21 |
| 2 | 1-Octen-3-one | 10.627–10.672 | 97, 70, 111, 98, 83, 55 | 1.46–114.32 * | 4.00–86.63 |
| 3 | Octanal | 11.378 | 55 | 0.03–2.95 * | 0.04–1.43 |
| 4 | Limonene | 12.232 | 91 | 0.17–9.21 * | 0.04–0.63 |
| 5 | 2-Octen-1-ol | 13.408–13.466 | 68, 95, 58, 81, 54, 110, 82, 41, 39, 57, 55, 69, 67, 56 | 0.51–75.13 * | 2.00–57.36 |
| 6 | Methyl octanoate | 15.091 | 74 | 0.03–0.14 | 0.03–0.56 * |
| 7 | Unknown | 15.438–15.446 | 69, 84, 55 | 0.02–0.44 | 0.03–1.08 * |
| 8 | Unknown | 20.402 | 91 | 0–0.82 * | 0–0.25 |
| 9 | Unknown | 21.057 | 91 | 0–0.27 * | 0–0.07 |
| 10 | Thujopsene | 23.718–23.756 | 204, 121, 105 | 0–0.67 | 0–4.1 * |
| 11 | Unknown | 24.599 | 165 | 0.05–0.47 * | 0.02–0.23 |
| 12 | Cuparene | 25.542 | 132 | 0–0.01 | 0–1.3 * |
| Variable Selection | Optimized Parameters | No. Variables | Data Sets | Accuracy (%) |
|---|---|---|---|---|
| Full variables | C = 4.64 × 102 | 829 | Cross-Validation | 77.59 |
| γ = 1.67 × 10−4 | Test | 84.00 | ||
| VIP method | C = 1.29 × 103 | 39 | Cross-Validation | 87.93 |
| γ = 1.29 × 10−4 | Test | 92.00 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cheng, Z.; Li, M.; Marriott, P.J.; Zhang, X.; Wang, S.; Li, J.; Ma, L. Chemometric Analysis of the Volatile Compounds Generated by Aspergillus carbonarius Strains Isolated from Grapes and Dried Vine Fruits. Toxins 2018, 10, 71. https://doi.org/10.3390/toxins10020071
Cheng Z, Li M, Marriott PJ, Zhang X, Wang S, Li J, Ma L. Chemometric Analysis of the Volatile Compounds Generated by Aspergillus carbonarius Strains Isolated from Grapes and Dried Vine Fruits. Toxins. 2018; 10(2):71. https://doi.org/10.3390/toxins10020071
Chicago/Turabian StyleCheng, Zhan, Menghua Li, Philip J. Marriott, Xiaoxu Zhang, Shiping Wang, Jiangui Li, and Liyan Ma. 2018. "Chemometric Analysis of the Volatile Compounds Generated by Aspergillus carbonarius Strains Isolated from Grapes and Dried Vine Fruits" Toxins 10, no. 2: 71. https://doi.org/10.3390/toxins10020071
APA StyleCheng, Z., Li, M., Marriott, P. J., Zhang, X., Wang, S., Li, J., & Ma, L. (2018). Chemometric Analysis of the Volatile Compounds Generated by Aspergillus carbonarius Strains Isolated from Grapes and Dried Vine Fruits. Toxins, 10(2), 71. https://doi.org/10.3390/toxins10020071







