Current Status and Future Opportunities of Omics Tools in Mycotoxin Research
Abstract
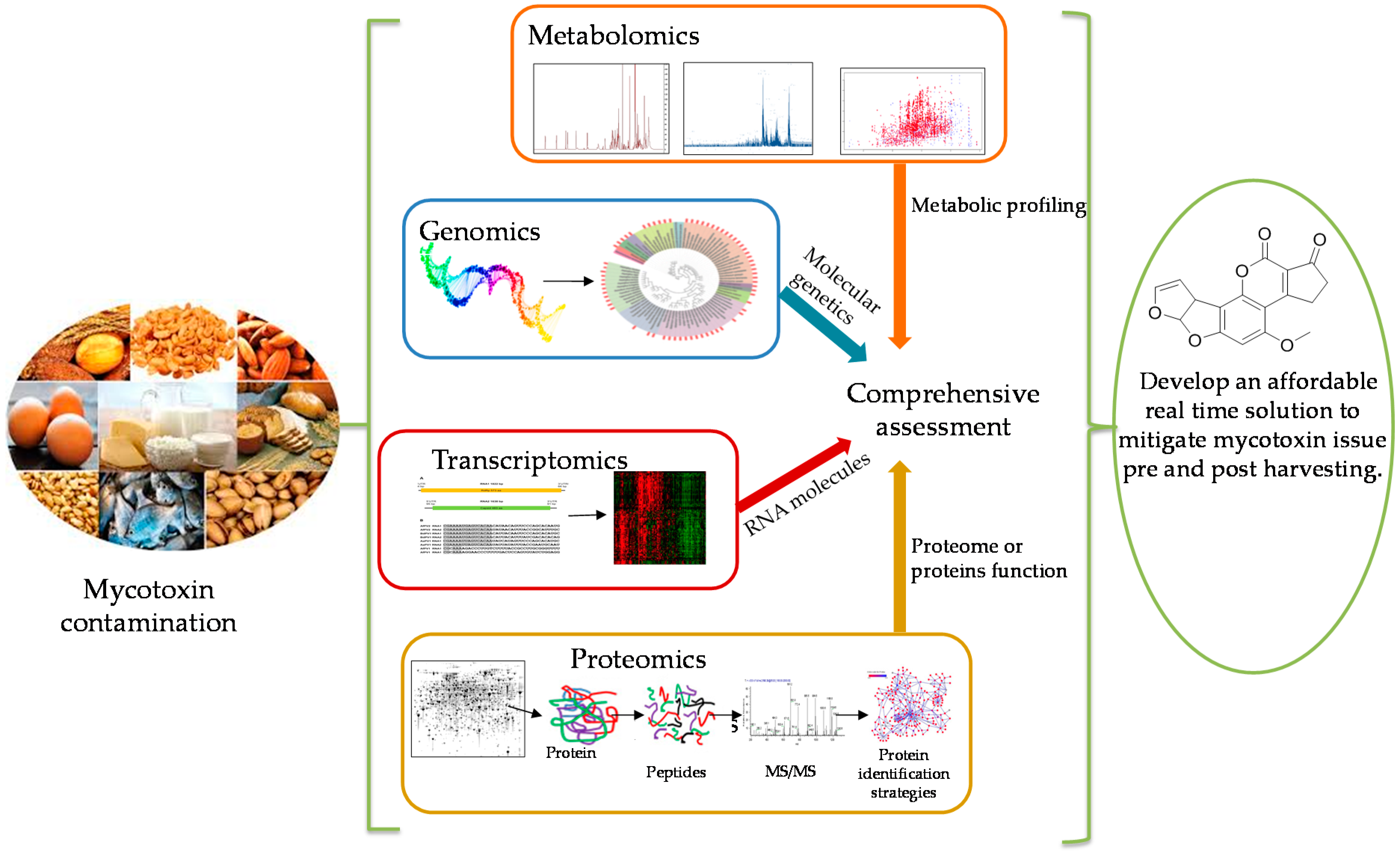
1. Introduction
2. Metabolomics Approach
Analytical Techniques Used in Mycotoxin Metabolomics Studies
3. Genomics Approach
Genomics Analysis for Mycotoxin Producing Fungi
4. Transcriptomics Approach
Transcriptional Profiling
5. Proteomics Approach
6. The Current Status of Omics Studies and Future Opportunities
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Coppock, R.W.; Christian, R.G.; Jacobsen, B.J. Aflatoxins. In Veterinary Toxicology; Elsevier: London, UK, 2018; pp. 983–994. ISBN 9780128114100. [Google Scholar]
- Bhat, R.; Rai, R.V.; Karim, A.A. Mycotoxins in Food and Feed: Present Status and Future Concerns. Compr. Rev. Food Sci. Food Saf. 2010, 9, 57–81. [Google Scholar] [CrossRef]
- Adekoya, I.; Njobeh, P.; Obadina, A.; Chilaka, C.; Okoth, S.; De Boevre, M.; De Saeger, S. Awareness and prevalence of mycotoxin contamination in selected nigerian fermented foods. Toxins 2017, 9, 363. [Google Scholar] [CrossRef] [PubMed]
- Dellafiora, L.; Dall’Asta, C. Forthcoming Challenges in Mycotoxins Toxicology Research for Safer Food-A Need for Multi-Omics Approach. Toxins 2017, 9, 18. [Google Scholar] [CrossRef] [PubMed]
- Tola, M.; Kebede, B. Occurrence, importance and control of mycotoxins: A review. Cogent Food Agric. 2016, 2. [Google Scholar] [CrossRef]
- Bui-Klimke, T.R.; Wu, F. Ochratoxin A and human health risk: A review of the evidence. Crit. Rev. Food Sci. Nutr. 2015, 55, 1860–1869. [Google Scholar] [CrossRef] [PubMed]
- Adeyeye, S.A.O. Fungal mycotoxins in foods: A review. Cogent Food Agric. 2016, 2, 1213127. [Google Scholar] [CrossRef]
- Rieswijk, L.; Claessen, S.M.H.; Bekers, O.; van Herwijnen, M.; Theunissen, D.H.J.; Jennen, D.G.J.; de Kok, T.M.C.M.; Kleinjans, J.C.S.; van Breda, S.G.J. Aflatoxin B1 induces persistent epigenomic effects in primary human hepatocytes associated with hepatocellular carcinoma. Toxicology 2016, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Reddy, K.; Salleh, B.; Saad, B.; Abbas, H.; Abel, C.; Shier, W. An overview of mycotoxin contamination in foods and its implications for human health. Toxin Rev. 2010, 29, 3–26. [Google Scholar] [CrossRef]
- Kumar, P.; Mahato, D.K.; Kamle, M.; Mohanta, T.K.; Kang, S.G. Aflatoxins: A Global Concern for Food Safety, Human Health and Their Management. Front. Microbiol. 2016, 7, 2170. [Google Scholar] [CrossRef] [PubMed]
- Krishnamachari, K.A.; Bhat, R.V.; Nagarajan, V.; Tilak, T.B. Hepatitis due to aflatoxicosis. An outbreak in Western India. Lancet 1975, 1, 1061–1063. [Google Scholar] [CrossRef]
- Bennett, J.W.; Klich, M. Mycotoxins. Clin. Microbiol. Rev. 2003, 16, 497–516. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.D.; Su, P.; Shan, H. Mycotoxin Contamination of Maize in China. Compr. Rev. Food Sci. Food Saf. 2017, 16, 835–849. [Google Scholar] [CrossRef]
- Bhatnagar, D.; Rajasekaran, K.; Payne, G.; Brown, R.; Yu, J.; Cleveland, T. The “omics” tools: Genomics, proteomics, metabolomics and their potential for solving the aflatoxin contamination problem. World Mycotoxin J. 2008, 1, 3–12. [Google Scholar] [CrossRef]
- Rezaei, M.; Karimi, F.; Parviz, M.; Behzadi, A.A.; Javadzadeh, M.; Mohammadpourfard, I.; Fallahzadeh, R.A.; Aghamirlou, H.M.; Malekirad, A.A. An Empirical Study on Aflatoxin Occurrence in Nuts Consumed in Tehran, Iran 2013. Health (Irvine Calif) 2014, 6, 649–653. [Google Scholar] [CrossRef]
- Klich, M.A. Aspergillus flavus: The major producer of aflatoxin. Mol. Plant Pathol. 2007, 8, 713–722. [Google Scholar] [CrossRef] [PubMed]
- Ji, C.; Fan, Y.; Zhao, L. Review on biological degradation of mycotoxins. Anim. Nutr. 2016, 2, 127–133. [Google Scholar] [CrossRef] [PubMed]
- Cabañes, F.J.; Sanseverino, W.; Castellá, G.; Bragulat, M.R.; Cigliano, R.A.; Sánchez, A. Rapid genome resequencing of an atoxigenic strain of Aspergillus carbonarius. Sci. Rep. 2015, 5, 9086. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Li, H.; Li, X.; Zhang, Q. Determination of trace patulin in apple-based food matrices. Food Chem. 2017, 233, 290–301. [Google Scholar] [CrossRef] [PubMed]
- Soler, L.; Oswald, I.P.P. The importance of accounting for sex in the search of proteomic signatures of mycotoxin exposure. J. Proteom. 2018, 178, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Sweeney, M.J.; Dobson, A.D. Mycotoxin production by Aspergillus, Fusarium and Penicillium species. Int. J. Food Microbiol. 1998, 43, 141–158. [Google Scholar] [CrossRef]
- Malir, F.; Ostry, V.; Pfohl-Leszkowicz, A.; Malir, J.; Toman, J. Ochratoxin A: 50 Years of Research. Toxins 2016, 8, 191. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Koutchma, T.; Warriner, K.; Zhou, T. Reduction of Patulin in Apple Juice Products by UV Light of Different Wavelengths in the UVC Range. J. Food Prot. 2014, 77, 963–971. [Google Scholar] [CrossRef] [PubMed]
- Zbyňovská, K.; Petruška, P.; Kalafová, A.; Capcarová, M. Patulin—A contaminant of food and feed: A review. Acta Fytotechnica et Zootechnica 2016, 19, 62–67. [Google Scholar] [CrossRef]
- IARC Working Group on the Evaluation of Carcinogenic Risks to Humans, World Health Organization. Overall Evaluations of Carcinogenicity: An Updating of IARC Monographs Volumes 1 to 42; WHO Press: Geneva, Switzerland, 1987; Volume 7, pp. 1–440. [Google Scholar]
- Luo, Y.; Liu, X.; Li, J. Updating techniques on controlling mycotoxins—A review. Food Control. 2018, 89, 123–132. [Google Scholar] [CrossRef]
- Zain, M.E. Impact of mycotoxins on humans and animals. J. Saudi Chem. Soc. 2011, 15, 129–144. [Google Scholar] [CrossRef]
- Yu, J. Current understanding on aflatoxin biosynthesis and future perspective in reducing aflatoxin contamination. Toxins 2012, 4, 1024–1057. [Google Scholar] [CrossRef] [PubMed]
- Cardwell, K.F.; Desjardins, A.; Henry, S.H.; Munkvold, G. Mycotoxins: The Cost of Achieving Food Securty and Food Quality. APSnet Featur. Artic. 2001. [Google Scholar] [CrossRef]
- Mazumder, P.M.; Sasmal, D. Mycotoxins—Limits and regulations. Anc. Sci. Life 2001, 20, 1–19. [Google Scholar] [PubMed]
- Garcia-Cela, E.; Verheecke-Vaessen, C.; Magan, N.; Medina, A. The “-omics” contributions to the understanding of mycotoxin production under diverse environmental conditions. Curr. Opin. Food Sci. 2018, 23, 97–104. [Google Scholar] [CrossRef]
- Bhatnagar-Mathur, P.; Sunkara, S.; Bhatnagar-Panwar, M.; Waliyar, F.; Sharma, K.K. Biotechnological advances for combating Aspergillus flavus and aflatoxin contamination in crops. Plant Sci. 2015, 234, 119–132. [Google Scholar] [CrossRef] [PubMed]
- Raja, K.; Patrick, M.; Gao, Y.; Madu, D.; Yang, Y.; Tsoi, L.C. A Review of Recent Advancement in Integrating Omics Data with Literature Mining towards Biomedical Discoveries. Int. J. Genom. 2017, 2017, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Righetti, L.; Dall’Asta, C.; Hajslova, J.; Rubert, J. Metabolomics Approaches and their Hidden Potential for Explaining the Mycotoxin Contamination Problem. In Metabolomics-Fundamentals and Applications; InTech: London, UK, 2016. [Google Scholar]
- Yin, Y.-N.; Yan, L.-Y.; Jiang, J.-H.; Ma, Z.-H. Biological control of aflatoxin contamination of crops. J. Zhejiang Univ. Sci. B 2008, 9, 787–792. [Google Scholar] [CrossRef] [PubMed]
- Caceres, I.; El-Khoury, R.; Bailly, S.; Oswald, I.P.; Puel, O.; Bailly, J.D. Piperine inhibits aflatoxin B1 production in Aspergillus flavus by modulating fungal oxidative stress response. Fungal Genet. Biol. 2017, 107, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Bergholz, T.M.; Moreno Switt, A.I.; Wiedmann, M. Omics approaches in food safety: Fulfilling the promise? Trends Microbiol. 2014, 22, 275–281. [Google Scholar] [CrossRef] [PubMed]
- Leng, Y.; Sun, K.; Chen, X.; Li, W. Suspension arrays based on nanoparticle-encoded microspheres for high-throughput multiplexed detection. Chem. Soc. Rev. 2015, 44, 5552–5595. [Google Scholar] [CrossRef] [PubMed]
- Hasin, Y.; Seldin, M.; Lusis, A. Multi-omics approaches to disease. Genome Biol. 2017, 18, 83. [Google Scholar] [CrossRef] [PubMed]
- Riekeberg, E.; Powers, R. New frontiers in metabolomics: From measurement to insight. F1000Research 2017, 6, 1148. [Google Scholar] [CrossRef] [PubMed]
- Zhang, A.; Sun, H.; Wang, P.; Han, Y.; Wang, X. Modern analytical techniques in metabolomics analysis. Analyst 2012, 137, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Zwickel, T.; Kahl, S.; Klaffke, H.; Rychlik, M.; Müller, M. Spotlight on the Underdogs—An Analysis of Underrepresented Alternaria Mycotoxins Formed Depending on Varying Substrate, Time and Temperature Conditions. Toxins 2016, 8, 344. [Google Scholar] [CrossRef] [PubMed]
- Eshelli, M.; Harvey, L.; Edrada-Ebel, R.; McNeil, B. Metabolomics of the bio-degradation process of aflatoxin B1 by actinomycetes at an initial pH of 6.0. Toxins 2015, 7, 439–456. [Google Scholar] [CrossRef] [PubMed]
- Smedsgaard, J.; Nielsen, J. Metabolite profiling of fungi and yeast: From phenotype to metabolome by MS and informatics. J. Exp. Bot. 2005, 56, 273–286. [Google Scholar] [CrossRef] [PubMed]
- Horgan, R.P.; Kenny, L.C. ‘Omic’ technologies: Genomics, transcriptomics, proteomics and metabolomics. Obstet. Gynaecol. 2011, 13, 189–195. [Google Scholar] [CrossRef]
- Iriti, M.; Faoro, F. Chemical Diversity and Defence Metabolism: How Plants Cope with Pathogens and Ozone Pollution. Int. J. Mol. Sci. 2009, 10, 3371–3399. [Google Scholar] [CrossRef] [PubMed]
- Rychlik, M.; Kanawati, B.; Schmitt-Kopplin, P. Foodomics as a promising tool to investigate the mycobolome. TrAC Trends Anal. Chem. 2017, 96, 22–30. [Google Scholar] [CrossRef]
- Castro-Puyana, M.; Pérez-Míguez, R.; Montero, L.; Herrero, M. Application of mass spectrometry-based metabolomics approaches for food safety, quality and traceability. TrAC Trends Anal. Chem. 2017, 93, 102–118. [Google Scholar] [CrossRef]
- Falade, T.; Chrysanthopoulos, P.; Hodson, M.; Sultanbawa, Y.; Fletcher, M.; Darnell, R.; Korie, S.; Fox, G.; Falade, T.D.O.; Chrysanthopoulos, P.K.; et al. Metabolites Identified during Varied Doses of Aspergillus Species in Zea mays Grains, and Their Correlation with Aflatoxin Levels. Toxins 2018, 10, 187. [Google Scholar] [CrossRef] [PubMed]
- Saldan, N.C.; Almeida, R.T.R.; Avíncola, A.; Porto, C.; Galuch, M.B.; Magon, T.F.S.; Pilau, E.J.; Svidzinski, T.I.E.; Oliveira, C.C. Development of an analytical method for identification of Aspergillus flavus based on chemical markers using HPLC-MS. Food Chem. 2018, 241, 113–121. [Google Scholar] [CrossRef] [PubMed]
- Tan, K.-C.; Trengove, R.D.; Maker, G.L.; Oliver, R.P.; Solomon, P.S. Metabolite profiling identifies the mycotoxin alternariol in the pathogen Stagonospora nodorum. Metabolomics 2009, 5, 330–335. [Google Scholar] [CrossRef]
- Fiehn, O. Combining Genomics, Metabolome Analysis, and Biochemical Modelling to Understand Metabolic Networks. Comp. Funct. Genom. 2001, 2, 155–168. [Google Scholar] [CrossRef] [PubMed]
- Varga, E.; Glauner, T.; Berthiller, F.; Krska, R.; Schuhmacher, R.; Sulyok, M. Development and validation of a (semi-)quantitative UHPLC-MS/MS method for the determination of 191 mycotoxins and other fungal metabolites in almonds, hazelnuts, peanuts and pistachios. Anal. Bioanal. Chem. 2013, 405, 5087–5104. [Google Scholar] [CrossRef] [PubMed]
- Hird, S.J.; Lau, B.P.-Y.; Schuhmacher, R.; Krska, R. Liquid chromatography-mass spectrometry for the determination of chemical contaminants in food. TrAC Trends Anal. Chem. 2014, 59, 59–72. [Google Scholar] [CrossRef]
- Rodríguez-Carrasco, Y.; Mañes, J.; Berrada, H.; Juan, C. Development and Validation of a LC-ESI-MS/MS Method for the Determination of Alternaria Toxins Alternariol, Alternariol Methyl-Ether and Tentoxin in Tomato and Tomato-Based Products. Toxins 2016, 8, 328. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Carrasco, Y.; Moltó, J.C.; Mañes, J.; Berrada, H. Development of a GC-MS/MS strategy to determine 15 mycotoxins and metabolites in human urine. Talanta 2014, 128, 125–131. [Google Scholar] [CrossRef] [PubMed]
- Meng-Reiterer, J.; Bueschl, C.; Rechthaler, J.; Berthiller, F.; Lemmens, M.; Schuhmacher, R. Metabolism of HT-2 Toxin and T-2 Toxin in Oats. Toxins 2016, 8, 364. [Google Scholar] [CrossRef] [PubMed]
- Juan, C.; Berrada, H.; Mañes, J.; Oueslati, S. Multi-mycotoxin determination in barley and derived products from Tunisia and estimation of their dietary intake. Food Chem. Toxicol. 2017, 103, 148–156. [Google Scholar] [CrossRef] [PubMed]
- Liao, C.D.; Wong, J.W.; Zhang, K.; Yang, P.; Wittenberg, J.B.; Trucksess, M.W.; Hayward, D.G.; Lee, N.S.; Chang, J.S. Multi-mycotoxin Analysis of Finished Grain and Nut Products Using Ultrahigh-Performance Liquid Chromatography and Positive Electrospray Ionization-Quadrupole Orbital Ion Trap High-Resolution Mass Spectrometry. J. Agric. Food Chem. 2015, 63, 8314–8332. [Google Scholar] [CrossRef] [PubMed]
- Rubert, J.; Righetti, L.; Stranska-Zachariasova, M.; Dzuman, Z.; Chrpova, J.; Dall’Asta, C.; Hajslova, J. Untargeted metabolomics based on ultra-high-performance liquid chromatography–high-resolution mass spectrometry merged with chemometrics: A new predictable tool for an early detection of mycotoxins. Food Chem. 2017, 224, 423–431. [Google Scholar] [CrossRef] [PubMed]
- Oplatowska-Stachowiak, M.; Haughey, S.A.; Chevallier, O.P.; Galvin-King, P.; Campbell, K.; Magowan, E.; Adam, G.; Berthiller, F.; Krska, R.; Elliott, C.T. Determination of the Mycotoxin Content in Distiller’s Dried Grain with Solubles Using a Multianalyte UHPLC-MS/MS Method. J. Agric. Food Chem. 2015, 63, 9441–9451. [Google Scholar] [CrossRef] [PubMed]
- Malachová, A.; Stránská, M.; Václavíková, M.; Elliott, C.T.; Black, C.; Meneely, J.; Hajšlová, J.; Ezekiel, C.N.; Schuhmacher, R.; Krska, R. Advanced LC–MS-based methods to study the co-occurrence and metabolization of multiple mycotoxins in cereals and cereal-based food. Anal. Bioanal. Chem. 2018, 410, 801–825. [Google Scholar] [CrossRef] [PubMed]
- Hopfgartner, G.; Varesio, E.; Tschäppät, V.; Grivet, C.; Bourgogne, E.; Leuthold, L.A. Triple quadrupole linear ion trap mass spectrometer for the analysis of small molecules and macromolecules. J. Mass Spectrom. 2004, 39, 845–855. [Google Scholar] [CrossRef] [PubMed]
- Righetti, L.; Paglia, G.; Galaverna, G.; Dall’Asta, C. Recent Advances and Future Challenges in Modified Mycotoxin Analysis: Why HRMS Has Become a Key Instrument in Food Contaminant Research. Toxins 2016, 8, 361. [Google Scholar] [CrossRef] [PubMed]
- Malachová, A.; Sulyok, M.; Beltrán, E.; Berthiller, F.; Krska, R. Optimization and validation of a quantitative liquid chromatography–tandem mass spectrometric method covering 295 bacterial and fungal metabolites including all regulated mycotoxins in four model food matrices. J. Chromatogr. A 2014, 1362, 145–156. [Google Scholar] [CrossRef] [PubMed]
- Croley, T.R.; White, K.D.; Callahan, J.H.; Musser, S.M. The Chromatographic Role in High Resolution Mass Spectrometry for Non-Targeted Analysis. J. Am. Soc. Mass Spectrom. 2012, 23, 1569–1578. [Google Scholar] [CrossRef] [PubMed]
- Aretz, I.; Meierhofer, D. Advantages and Pitfalls of Mass Spectrometry Based Metabolome Profiling in Systems Biology. Int. J. Mol. Sci. 2016, 17, 632. [Google Scholar] [CrossRef] [PubMed]
- Kos, G.; Sieger, M.; McMullin, D.; Zahradnik, C.; Sulyok, M.; Öner, T.; Mizaikoff, B.; Krska, R. A novel chemometric classification for FTIR spectra of mycotoxin-contaminated maize and peanuts at regulatory limits. Food Addit. Contam. Part. A 2016, 33, 1596–1607. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Wu, H.-L.; Xie, L.-X.; Hu, Y.; Fang, H.; Sun, X.-D.; Wang, T.; Xiao, R.; Yu, R.-Q. Chemometrics-enhanced liquid chromatography-full scan-mass spectrometry for interference-free analysis of multi-class mycotoxins in complex cereal samples. Chemom. Intell. Lab. Syst. 2017, 160, 125–138. [Google Scholar] [CrossRef]
- Chernushevich, I.V.; Loboda, A.V.; Thomson, B.A. An introduction to quadrupole-time-of-flight mass spectrometry. J. Mass Spectrom. 2001, 36, 849–865. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Zhang, Y.; Shao, S.; Cai, Z.; Feng, L.; Pan, H.; Wang, Z. Simultaneous determination of multi-component mycotoxin contaminants in foods and feeds by ultra-performance liquid chromatography tandem mass spectrometry. J. Chromatogr. A 2007, 1143, 48–64. [Google Scholar] [CrossRef] [PubMed]
- Streit, E.; Schwab, C.; Sulyok, M.; Naehrer, K.; Krska, R.; Schatzmayr, G. Multi-Mycotoxin Screening Reveals the Occurrence of 139 Different Secondary Metabolites in Feed and Feed Ingredients. Toxins 2013, 5, 504–523. [Google Scholar] [CrossRef] [PubMed]
- Ok, H.; Tian, F.; Hong, E.; Paek, O.; Kim, S.-H.; Kim, D.; Chun, H. Harmonized Collaborative Validation of Aflatoxins and Sterigmatocystin in White Rice and Sorghum by Liquid Chromatography Coupled to Tandem Mass Spectrometry. Toxins 2016, 8, 371. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Li, W.; Zhang, Y.; Hu, X.; Wu, L.; Wang, B. QuEChERS Purification Combined with Ultrahigh-Performance Liquid Chromatography Tandem Mass Spectrometry for Simultaneous Quantification of 25 Mycotoxins in Cereals. Toxins 2016, 8, 375. [Google Scholar] [CrossRef] [PubMed]
- Mylroie, J.E.; Ozkan, S.; Shivaji, R.; Windham, G.L.; Alpe, M.N.; Williams, W.P. Identification and Quantification of a Toxigenic and Non-Toxigenic Aspergillus flavus Strain in Contaminated Maize Using Quantitative Real-Time PCR. Toxins 2016, 8, 15. [Google Scholar] [CrossRef] [PubMed]
- Lattanzio, V.M.T.; Solfrizzo, M.; Powers, S.; Visconti, A. Simultaneous determination of aflatoxins, ochratoxin A andFusarium toxins in maize by liquid chromatography/tandem mass spectrometry after multitoxin immunoaffinity cleanup. Rapid Commun. Mass Spectrom. 2007, 21, 3253–3261. [Google Scholar] [CrossRef] [PubMed]
- Vaclavikova, M.; MacMahon, S.; Zhang, K.; Begley, T.H. Application of single immunoaffinity clean-up for simultaneous determination of regulated mycotoxins in cereals and nuts. Talanta 2013, 117, 345–351. [Google Scholar] [CrossRef] [PubMed]
- Lattanzio, V.M.T.; Ciasca, B.; Powers, S.; Visconti, A. Improved method for the simultaneous determination of aflatoxins, ochratoxin A and Fusarium toxins in cereals and derived products by liquid chromatography–tandem mass spectrometry after multi-toxin immunoaffinity clean up. J. Chromatogr. A 2014, 1354, 139–143. [Google Scholar] [CrossRef] [PubMed]
- Wilcox, J.; Donnelly, C.; Leeman, D.; Marley, E. The use of immunoaffinity columns connected in tandem for selective and cost-effective mycotoxin clean-up prior to multi-mycotoxin liquid chromatographic–tandem mass spectrometric analysis in food matrices. J. Chromatogr. A 2015, 1400, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Desmarchelier, A.; Oberson, J.-M.; Tella, P.; Gremaud, E.; Seefelder, W.; Mottier, P. Development and Comparison of Two Multiresidue Methods for the Analysis of 17 Mycotoxins in Cereals by Liquid Chromatography Electrospray Ionization Tandem Mass Spectrometry. J. Agric. Food Chem. 2010, 58, 7510–7519. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira Filho, J.W.G.; Islam, M.T.; Ali, E.S.; Uddin, S.J.; Santos, J.V.d.O.; de Alencar, M.V.O.B.; Júnior, A.L.G.; Paz, M.F.C.J.; de Brito, M.d.R.M.; E Sousa, J.M.d.C.; et al. A comprehensive review on biological properties of citrinin. Food Chem. Toxicol. 2017, 110, 130–141. [Google Scholar] [CrossRef] [PubMed]
- Lacina, O.; Zachariasova, M.; Urbanova, J.; Vaclavikova, M.; Cajka, T.; Hajslova, J. Critical assessment of extraction methods for the simultaneous determination of pesticide residues and mycotoxins in fruits, cereals, spices and oil seeds employing ultra-high performance liquid chromatography–tandem mass spectrometry. J. Chromatogr. A 2012, 1262, 8–18. [Google Scholar] [CrossRef] [PubMed]
- Sulyok, M.; Berthiller, F.; Krska, R.; Schuhmacher, R. Development and validation of a liquid chromatography/tandem mass spectrometric method for the determination of 39 mycotoxins in wheat and maize. Rapid Commun. Mass Spectrom. 2006, 20, 2649–2659. [Google Scholar] [CrossRef] [PubMed]
- Rubert, J.; Dzuman, Z.; Vaclavikova, M.; Zachariasova, M.; Soler, C.; Hajslova, J. Analysis of mycotoxins in barley using ultra high liquid chromatography high resolution mass spectrometry: Comparison of efficiency and efficacy of different extraction procedures. Talanta 2012, 99, 712–719. [Google Scholar] [CrossRef] [PubMed]
- Beccari, G.; Caproni, L.; Tini, F.; Uhlig, S.; Covarelli, L. Presence of Fusarium Species and Other Toxigenic Fungi in Malting Barley and Multi-Mycotoxin Analysis by Liquid Chromatography–High-Resolution Mass Spectrometry. J. Agric. Food Chem. 2016, 64, 4390–4399. [Google Scholar] [CrossRef] [PubMed]
- Diana Di Mavungu, J.; Monbaliu, S.; Scippo, M.-L.; Maghuin-Rogister, G.; Schneider, Y.-J.; Larondelle, Y.; Callebaut, A.; Robbens, J.; Van Peteghem, C.; De Saeger, S. LC-MS/MS multi-analyte method for mycotoxin determination in food supplements. Food Addit. Contam. Part A 2009, 26, 885–895. [Google Scholar] [CrossRef] [PubMed]
- Warth, B.; Parich, A.; Atehnkeng, J.; Bandyopadhyay, R.; Schuhmacher, R.; Sulyok, M.; Krska, R. Quantitation of Mycotoxins in Food and Feed from Burkina Faso and Mozambique Using a Modern LC-MS/MS Multitoxin Method. J. Agric. Food Chem. 2012, 60, 9352–9363. [Google Scholar] [CrossRef] [PubMed]
- Martins, C.; Assunção, R.; Cunha, S.C.; Fernandes, J.O.; Jager, A.; Petta, T.; Oliveira, C.A.; Alvito, P. Assessment of multiple mycotoxins in breakfast cereals available in the Portuguese market. Food Chem. 2018, 239, 132–140. [Google Scholar] [CrossRef] [PubMed]
- Lindenmeier, M.; Schieberle, P.; Rychlik, M. Quantification of ochratoxin A in foods by a stable isotope dilution assay using high-performance liquid chromatography-tandem mass spectrometry. J. Chromatogr. A 2004, 1023, 57–66. [Google Scholar] [CrossRef] [PubMed]
- Zöllner, P.; Mayer-Helm, B. Trace mycotoxin analysis in complex biological and food matrices by liquid chromatography–atmospheric pressure ionisation mass spectrometry. J. Chromatogr. A 2006, 1136, 123–169. [Google Scholar] [CrossRef] [PubMed]
- Human Genome Sequencing Consortium, I. Finishing the euchromatic sequence of the human genome. Nature 2004, 431, 931–945. [Google Scholar] [CrossRef] [PubMed]
- Cary, J.W.; OBrian, G.R.; Nielsen, D.M.; Nierman, W.; Harris-Coward, P.; Yu, J.; Bhatnagar, D.; Cleveland, T.E.; Payne, G.A.; Calvo, A.M. Elucidation of veA-dependent genes associated with aflatoxin and sclerotial production in Aspergillus flavus by functional genomics. Appl. Microbiol. Biotechnol. 2007, 76, 1107–1118. [Google Scholar] [CrossRef] [PubMed]
- Amaike, S.; Keller, N.P. Aspergillus flavus. Annu. Rev. Phytopathol. 2011, 49, 107–133. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.-Y.; Brown, R.L.; Damann, K.E.; Cleveland, T.E. Identification of Unique or Elevated Levels of Kernel Proteins in Aflatoxin-Resistant Maize Genotypes Through Proteome Analysis. Phytopathology 2002, 92, 1084–1094. [Google Scholar] [CrossRef] [PubMed]
- Cleveland, T.E.; Yu, J.; Fedorova, N.; Bhatnagar, D.; Payne, G.A.; Nierman, W.C.; Bennett, J.W. Potential of Aspergillus flavus genomics for applications in biotechnology. Trends Biotechnol. 2009, 27, 151–157. [Google Scholar] [CrossRef] [PubMed]
- Mahmoud, M.A. Detection of Aspergillus flavus in Stored Peanuts Using Real-Time PCR and the Expression of Aflatoxin Genes in Toxigenic and Atoxigenic A. flavus Isolates. Foodborne Pathog. Dis. 2015, 12, 289–296. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Chiodini, R.; Badr, A.; Zhang, G. The impact of next-generation sequencing on genomics. J. Genet. Genom. 2011, 38, 95–109. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Ronning, C.M.; Wilkinson, J.R.; Campbell, B.C.; Payne, G.A.; Bhatnagar, D.; Cleveland, T.E.; Nierman, W.C. Gene profiling for studying the mechanism of aflatoxin biosynthesis in Aspergillus flavus and A. parasiticus. Food Addit. Contam. 2007, 24, 1035–1042. [Google Scholar] [CrossRef] [PubMed]
- Moore, G.G.; Mack, B.M.; Beltz, S.B. Genomic sequence of the aflatoxigenic filamentous fungus Aspergillus nomius. BMC Genom. 2015, 16, 551. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Yu, J.; Mahoney, N.; Chan, K.L.; Molyneux, R.J.; Varga, J.; Bhatnagar, D.; Cleveland, T.E.; Nierman, W.C.; Campbell, B.C. Elucidation of the functional genomics of antioxidant-based inhibition of aflatoxin biosynthesis. Int. J. Food Microbiol. 2008, 122, 49–60. [Google Scholar] [CrossRef] [PubMed]
- Cleveland, T.E.; Bhatnagar, D.; Yu, J. Elimination and control of aflatoxin contamination in agricultural crops through Aspergillus flavus genomics. In Mycotoxin Prevention and Control in Agriculture; Appell, M., Kendra, F.D.F., Trucksess, M.W., Eds.; ACS eBooks Publications: Washington, DC, USA, 2010; pp. 93–106. [Google Scholar]
- Yu, J.; Bhatnagar, D.; Cleveland, T.E. Chapter eleven Genetics and biochemistry of aflatoxin formation and genomics approach for preventing aflatoxin contamination. Recent Adv. Phytochem. 2004, 38, 223–255. [Google Scholar] [CrossRef]
- Niessen, L.; Bechtner, J.; Fodil, S.; Taniwaki, M.H.; Vogel, R.F. LAMP-based group specific detection of aflatoxin producers within Aspergillus section Flavi in food raw materials, spices, and dried fruit using neutral red for visible-light signal detection. Int. J. Food Microbiol. 2018, 266, 241–250. [Google Scholar] [CrossRef] [PubMed]
- Sadhasivam, S.; Britzi, M.; Zakin, V.; Kostyukovsky, M.; Trostanetsky, A.; Quinn, E.; Sionov, E. Rapid Detection and Identification of Mycotoxigenic Fungi and Mycotoxins in Stored Wheat Grain. Toxins 2017, 9, 302. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Payne, G.A.; Nierman, W.C.; Machida, M.; Bennett, J.W.; Campbell, B.C.; Robens, J.F.; Bhatnagar, D.; Dean, R.A.; Cleveland, T.E. Aspergillus flavus genomics as a tool for studying the mechanism of aflatoxin formation. Food Addit. Contam. Part A Chem. Anal. Control. Expos. Risk Assess. 2008, 25, 1152–1157. [Google Scholar] [CrossRef]
- Chancharoonpong, C.; Hsieh, P.-C.; Sheu, S.-C. Enzyme Production and Growth of Aspergillus oryzae S. on Soybean Koji Fermentation. APCBEE Procedia 2012, 2, 57–61. [Google Scholar] [CrossRef]
- Faustinelli, P.C.; Palencia, E.R.; Sobolev, V.S.; Horn, B.W.; Sheppard, H.T.; Lamb, M.C.; Wang, X.M.; Scheffler, B.E.; Martinez Castillo, J.; Arias, R.S. Study of the genetic diversity of the aflatoxin biosynthesis cluster in Aspergillus section Flavi using insertion/deletion markers in peanut seeds from Georgia, USA. Mycologia 2017, 109, 200–209. [Google Scholar] [CrossRef] [PubMed]
- Faustinelli, P.C.; Wang, X.M.; Palencia, E.R.; Arias, R.S. Genome Sequences of Eight Aspergillus flavus spp. and One A. parasiticus sp., Isolated from Peanut Seeds in Georgia. Genome Announc. 2016, 4. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, M.K.; Mack, B.M.; Payne, G.A.; Bhatnagar, D. Use of functional genomics to assess the climate change impact on Aspergillus flavus and aflatoxin production. World Mycotoxin J. 2016, 9, 665–672. [Google Scholar] [CrossRef]
- Gallo, A.; Bruno, K.S.; Solfrizzo, M.; Perrone, G.; Mulè, G.; Visconti, A.; Baker, S.E. New insight into the ochratoxin A biosynthetic pathway through deletion of a nonribosomal peptide synthetase gene in Aspergillus carbonarius. Appl. Environ. Microbiol. 2012, 78, 8208–8218. [Google Scholar] [CrossRef] [PubMed]
- Chakrabortti, A.; Li, J.; Liang, Z.-X. Complete Genome Sequence of the Filamentous Fungus Aspergillus westerdijkiae Reveals the Putative Biosynthetic Gene Cluster of Ochratoxin A. Genome Announc. 2016, 4. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Chakrabortti, A.; Zhu, J.; Liang, Z.-X.; Li, J. Sequencing and functional annotation of the whole genome of the filamentous fungus Aspergillus westerdijkiae. BMC Genom. 2016, 17, 633. [Google Scholar] [CrossRef] [PubMed]
- Puel, O.; Galtier, P.; Oswald, I.P. Biosynthesis and toxicological effects of patulin. Toxins 2010, 2, 613–631. [Google Scholar] [CrossRef] [PubMed]
- Snini, S.P.; Tadrist, S.; Laffitte, J.; Jamin, E.L.; Oswald, I.P.; Puel, O. The gene PatG involved in the biosynthesis pathway of patulin, a food-borne mycotoxin, encodes a 6-methylsalicylic acid decarboxylase. Int. J. Food Microbiol. 2014, 171, 77–83. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zong, Y.; Du, Z.; Chen, Y.; Zhang, Z.; Qin, G.; Zhao, W.; Tian, S. Genomic Characterization Reveals Insights Into Patulin Biosynthesis and Pathogenicity in Penicillium Species. Mol. Plant.-Microbe Interact. 2015, 28, 635–647. [Google Scholar] [CrossRef] [PubMed]
- Carvajal-Campos, A.; Manizan, A.; Tadrist, S.; Akaki, D.; Koffi-Nevry, R.; Moore, G.; Fapohunda, S.; Bailly, S.; Montet, D.; Oswald, I.; et al. Aspergillus korhogoensis, a Novel Aflatoxin Producing Species from the Côte d’Ivoire. Toxins 2017, 9, 353. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.; Chen, Y. Transcriptomics: Advances and approaches. Sci. China Life Sci. 2013, 56, 960–967. [Google Scholar] [CrossRef] [PubMed]
- Scheper, G.C.; van der Knaap, M.S.; Proud, C.G. Translation matters: Protein synthesis defects in inherited disease. Nat. Rev. Genet. 2007, 8, 711–723. [Google Scholar] [CrossRef] [PubMed]
- Pedrotty, D.M.; Morley, M.P.; Cappola, T.P. Transcriptomic biomarkers of cardiovascular disease. Prog. Cardiovasc. Dis. 2012, 55, 64–69. [Google Scholar] [CrossRef] [PubMed]
- Macaulay, I.C.; Carr, P.; Gusnanto, A.; Ouwehand, W.; Fitzgerald, D.; Watkins, N. Platelet genomics and proteomics in human health and disease. J. Clin. Investig. 2005, 115, 3370–3377. [Google Scholar] [CrossRef] [PubMed]
- Sayanthooran, S.; Magana-Arachchi, D.N.; Gunerathne, L.; Abeysekera, T. Potential diagnostic biomarkers for chronic kidney disease of unknown etiology (CKDu) in Sri Lanka: A pilot study. BMC Nephrol. 2017, 18, 31. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Hu, B.; Shao, L.; Tian, Y.; Jin, T.; Jin, Y.; Ji, S.; Fan, X. Integrated analysis of transcriptomics and metabonomics profiles in aflatoxin B1-induced hepatotoxicity in rat. Food Chem. Toxicol. 2013, 55, 444–455. [Google Scholar] [CrossRef] [PubMed]
- Alwine, J.C.; Kemp, D.J.; Stark, G.R. Method for detection of specific RNAs in agarose gels by transfer to diazobenzyloxymethyl-paper and hybridization with DNA probes. Proc. Natl. Acad. Sci. USA 1977, 74, 5350–5354. [Google Scholar] [CrossRef] [PubMed]
- Malone, J.H.; Oliver, B. Microarrays, deep sequencing and the true measure of the transcriptome. BMC Biol. 2011, 9, 34. [Google Scholar] [CrossRef] [PubMed]
- Lowe, R.; Shirley, N.; Bleackley, M.; Dolan, S.; Shafee, T. Transcriptomics technologies. PLOS Comput. Biol. 2017, 13, e1005457. [Google Scholar] [CrossRef] [PubMed]
- Castellá, G.; Bragulat, M.R.; Puig, L.; Sanseverino, W.; Cabañes, F.J. Genomic diversity in ochratoxigenic and non ochratoxigenic strains of Aspergillus carbonarius. Sci. Rep. 2018, 8, 5439. [Google Scholar] [CrossRef] [PubMed]
- Musungu, B.M.; Bhatnagar, D.; Brown, R.L.; Payne, G.A.; OBrian, G.; Fakhoury, A.M.; Geisler, M. A Network Approach of Gene Co-expression in the Zea mays/Aspergillus flavus Pathosystem to Map Host/Pathogen Interaction Pathways. Front. Genet. 2016, 7, 206. [Google Scholar] [CrossRef] [PubMed]
- Cunha, S.C.; Faria, M.A.; Fernandes, J.O. Determination of patulin in apple and quince products by GC–MS using 13C5–7 patulin as internal standard. Food Chem. 2009, 115, 352–359. [Google Scholar] [CrossRef]
- Nayak, S.N.; Agarwal, G.; Pandey, M.K.; Sudini, H.K.; Jayale, A.S.; Purohit, S.; Desai, A.; Wan, L.; Guo, B.; Liao, B.; et al. Aspergillus flavus infection triggered immune responses and host-pathogen cross-talks in groundnut during in-vitro seed colonization. Sci. Rep. 2017, 7, 9659. [Google Scholar] [CrossRef] [PubMed]
- Bhatnagar, D.; Rajasekaran, K.; Gilbert, M.; Cary, J.W.; Magan, N. Advances in molecular and genomic research to safeguard food and feed supply from aflatoxin contamination. World Mycotoxin J. 2018, 11, 47–72. [Google Scholar] [CrossRef]
- Vettorazzi, A.; van Delft, J.; López de Cerain, A. A review on ochratoxin A transcriptomic studies. Food Chem. Toxicol. 2013, 59, 766–783. [Google Scholar] [CrossRef] [PubMed]
- Smit, E.; Souza, T.; Jennen, D.G.J.; Kleinjans, J.C.S.; van den Beucken, T. Identification of essential transcription factors for adequate DNA damage response after benzo(a)pyrene and aflatoxin B1 exposure by combining transcriptomics with functional genomics. Toxicology 2017, 390, 74–82. [Google Scholar] [CrossRef] [PubMed]
- Hoek-van den Hil, E.F.; van Schothorst, E.M.; van der Stelt, I.; Hollman, P.C.H.; Keijer, J.; Rietjens, I.M.C.M. Quercetin tests negative for genotoxicity in transcriptome analyses of liver and small intestine of mice. Food Chem. Toxicol. 2015, 81, 34–39. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Sang, Y.; Zhang, G. Combined cytotoxicity of aflatoxin B1 and deoxynivalenol to hepatoma HepG2/C3A cells. World Mycotoxin J. 2017, 10, 387–399. [Google Scholar] [CrossRef]
- Josse, R.; Dumont, J.; Fautrel, A.; Robin, M.-A.; Guillouzo, A. Identification of early target genes of aflatoxin B1 in human hepatocytes, inter-individual variability and comparison with other genotoxic compounds. Toxicol. Appl. Pharmacol. 2012, 258, 176–187. [Google Scholar] [CrossRef] [PubMed]
- Vanacloig-Pedros, E.; Proft, M.; Pascual-Ahuir, A. Different Toxicity Mechanisms for Citrinin and Ochratoxin A Revealed by Transcriptomic Analysis in Yeast. Toxins 2016, 8, 273. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.-S.; Lin, Y.-T.; Huang, Y.-T.; Cheng, Y.-C.; Yu, F.-Y.; Liu, B.-H. Disruption of liver development and coagulation pathway by ochratoxin A in embryonic zebrafish. Toxicol. Appl. Pharmacol. 2018, 340, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Jennings, P.; Weiland, C.; Limonciel, A.; Bloch, K.M.; Radford, R.; Aschauer, L.; McMorrow, T.; Wilmes, A.; Pfaller, W.; Ahr, H.J.; et al. Transcriptomic alterations induced by Ochratoxin A in rat and human renal proximal tubular in vitro models and comparison to a rat in vivo model. Arch. Toxicol. 2012, 86, 571–589. [Google Scholar] [CrossRef] [PubMed]
- Ianiri, G.; Idnurm, A.; Castoria, R. Transcriptomic responses of the basidiomycete yeast Sporobolomyces sp. to the mycotoxin patulin. BMC Genom. 2016, 17, 210. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, M.R.; Pasquali, C.; Appel, R.D.; Ou, K.; Golaz, O.; Sanchez, J.C.; Yan, J.X.; Gooley, A.A.; Hughes, G.; Humphery-Smith, I.; et al. From proteins to proteomes: Large scale protein identification by two-dimensional electrophoresis and amino acid analysis. Nat. Biotechnol. 1996, 14, 61. [Google Scholar] [CrossRef]
- Gupta, A.; Govila, V.; Saini, A. Proteomics—The research frontier in periodontics. J. Oral Biol. Craniofacial Res. 2015, 5, 46–52. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, C.M.; Emanuelli, B.; Kahn, C.R. Critical nodes in signalling pathways: Insights into insulin action. Nat. Rev. Mol. Cell Biol. 2006, 7, 85–96. [Google Scholar] [CrossRef] [PubMed]
- Giacometti, J.; Tomljanović, A.B.; Josić, D. Application of proteomics and metabolomics for investigation of food toxins. Food Res. Int. 2013, 54, 1042–1051. [Google Scholar] [CrossRef]
- Liebler, D.C. Introduction to Proteomics: Tools for the New Biology; Humana Press: New York, NY, USA, 2002; ISBN 9781592591305. [Google Scholar]
- Matthiesen, R.; Mutenda, K.E. Introduction to Proteomics. In Mass Spectrometry Data Analysis in Proteomics; Humana Press: New York, NY, USA, 2007; pp. 1–36. [Google Scholar]
- Gaspari, M.; Cuda, G. Nano LC–MS/MS: A Robust Setup for Proteomic Analysis. In Methods in Molecular Biology (Clifton, N.J.); Humana Press: New York, NY, USA, 2011; Volume 790, pp. 115–126. [Google Scholar]
- Ishihama, Y. Proteomic LC-MS systems using nanoscale liquid chromatography with tandem mass spectrometry. J. Chromatogr. A 2005, 1067, 73–83. [Google Scholar] [CrossRef] [PubMed]
- Issaq, H.; Chan, K.; Janini, G.; Conrads, T.; Veenstra, T. Multidimensional separation of peptides for effective proteomic analysis. J. Chromatogr. B 2005, 817, 35–47. [Google Scholar] [CrossRef] [PubMed]
- O’Farrell, P.H. High resolution two-dimensional electrophoresis of proteins. J. Biol. Chem. 1975, 250, 4007–4021. [Google Scholar] [PubMed]
- Mohammadi, M.; Anoop, V.; Gleddie, S.; Harris, L.J. Proteomic profiling of two maize inbreds during early gibberella ear rot infection. Proteomics 2011, 11, 3675–3684. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.-Y.; Brown, R.L.; Guo, B.Z.; Menkir, A.; Cleveland, T.E. Identifying Aflatoxin Resistance-related Proteins/Genes through Proteomics and RNAi Gene Silencing. Peanut Sci. 2009, 36, 35–41. [Google Scholar] [CrossRef]
- Campo, S.; Carrascal, M.; Coca, M.; Abián, J.; San Segundo, B. The defense response of germinating maize embryos against fungal infection: A proteomics approach. Proteomics 2004, 4, 383–396. [Google Scholar] [CrossRef] [PubMed]
- Chivasa, S.; Simon, W.J.; Yu, X.-L.; Yalpani, N.; Slabas, A.R. Pathogen elicitor-induced changes in the maize extracellular matrix proteome. Proteomics 2005, 5, 4894–4904. [Google Scholar] [CrossRef] [PubMed]
- Medina, M.L.; Francisco, W.A. Isolation and Enrichment of Secreted Proteins from Filamentous Fungi. In Methods in Molecular Biology (Clifton, N.J.); Humana Press: New York, NY, USA, 2008; Volume 425, pp. 275–285. [Google Scholar]
- Crespo-Sempere, A.; Gil, J.V.V.; Martínez-Culebras, P.V. V Proteome analysis of the fungus Aspergillus carbonarius under ochratoxin A producing conditions. Int. J. Food Microbiol. 2011, 147, 162–169. [Google Scholar] [CrossRef] [PubMed]
- Pechanova, O.; Pechan, T.; Williams, W.P.; Luthe, D.S. Proteomic analysis of the maize rachis: Potential roles of constitutive and induced proteins in resistance to Aspergillus flavus infection and aflatoxin accumulation. Proteomics 2011, 11, 114–127. [Google Scholar] [CrossRef] [PubMed]
- Stoll, D.A.; Link, S.; Kulling, S.; Geisen, R.; Schmidt-Heydt, M. Comparative proteome analysis of Penicillium verrucosum grown under light of short wavelength shows an induction of stress-related proteins associated with modified mycotoxin biosynthesis. Int. J. Food Microbiol. 2014, 175, 20–29. [Google Scholar] [CrossRef] [PubMed]
- Degola, F.; Bisceglie, F.; Pioli, M.; Palmano, S.; Elviri, L.; Pelosi, G.; Lodi, T.; Restivo, F.M. Structural modification of cuminaldehyde thiosemicarbazone increases inhibition specificity toward aflatoxin biosynthesis and sclerotia development in Aspergillus flavus. Appl. Microbiol. Biotechnol. 2017, 101, 6683–6696. [Google Scholar] [CrossRef] [PubMed]
- Razzazi-Fazeli, E.; Rizwan, M.; Mayrhofer, C.; Nobauer, K. The Use of Proteomics as a Novel Tool in Aflatoxin Research. In Aflatoxins-Biochemistry and Molecular Biology; InTech: London, UK, 2011. [Google Scholar]
- Nöbauer, K.; Hummel, K.; Mayrhofer, C.; Ahrens, M.; Setyabudi, F.M.C.; Schmidt-Heydt, M.; Eisenacher, M.; Razzazi-Fazeli, E. Comprehensive proteomic analysis of Penicillium verrucosum. Proteomics 2017, 17, 1600467. [Google Scholar] [CrossRef] [PubMed]
- Jeyaramraja, P.R.; Meenakshi, S.N.; Woldesenbet, F. Relationship between drought and preharvest aflatoxin contamination in groundnut (Arachis hypogaea L.). World Mycotoxin J. 2018, 11, 187–199. [Google Scholar] [CrossRef]
- Wang, Z.; Yan, S.; Liu, C.; Chen, F.; Wang, T. Proteomic Analysis Reveals an Aflatoxin-Triggered Immune Response in Cotyledons of Arachis hypogaea Infected with Aspergillus flavus. J. Proteome Res. 2012, 11, 2739–2753. [Google Scholar] [CrossRef] [PubMed]
- Fountain, J.C.; Koh, J.; Yang, L.; Pandey, M.K.; Nayak, S.N.; Bajaj, P.; Zhuang, W.-J.; Chen, Z.-Y.; Kemerait, R.C.; Lee, R.D.; et al. Proteome analysis of Aspergillus flavus isolate-specific responses to oxidative stress in relationship to aflatoxin production capability. Sci. Rep. 2018, 8, 3430. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Wang, X.; Luo, J.; Ye, B.; Zhou, Y.; Zhou, L.; Lai, T. Identification of differentially expressed genes involved in spore germination of Penicillium expansum by comparative transcriptome and proteome approaches. Microbiologyopen 2018, 7, e00562. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.; Li, H.; Liu, F.; Zhang, Y.; Zhang, Q.; Wang, Y.; Li, P. Proteome Changes in Penicillium expansum Grown in a Medium Derived from Host Plant. J. Microbiol. Biotechnol. 2017, 27, 624–632. [Google Scholar] [CrossRef] [PubMed]
- Lai, T.; Wang, Y.; Fan, Y.; Zhou, Y.; Bao, Y.; Zhou, T. The response of growth and patulin production of postharvest pathogen Penicillium expansum to exogenous potassium phosphite treatment. Int. J. Food Microbiol. 2017, 244, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Katam, R.; Sakata, K.; Suravajhala, P.; Pechan, T.; Kambiranda, D.M.; Naik, K.S.; Guo, B.; Basha, S.M. Comparative leaf proteomics of drought-tolerant and -susceptible peanut in response to water stress. J. Proteom. 2016, 143, 209–226. [Google Scholar] [CrossRef] [PubMed]
- Lv, Y.; Lv, A.; Zhai, H.; Zhang, S.; Li, L.; Cai, J.; Hu, Y. Insight into the global regulation of laeA in Aspergillus flavus based on proteomic profiling. Int. J. Food Microbiol. 2018, 284, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Peng, X.; Yang, Z.; Zhao, W.; Xu, W.; Hao, J.; Wu, W.; Shen, X.L.; Luo, Y.; Huang, K. iTRAQ Mitoproteome Analysis Reveals Mechanisms of Programmed Cell Death in Arabidopsis thaliana Induced by Ochratoxin A. Toxins 2017, 9, 167. [Google Scholar] [CrossRef] [PubMed]
| Mycotoxin | Structure | Fungal Species | IARC Classification | Ref. |
|---|---|---|---|---|
| Aflatoxin B1 | 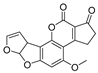 | Aspergillus flavus, Aspergillus parasiticus, Aspergillus bombycis, Aspergillus A. coracles, Aspergillus nomius, Aspergillus pseudotamari | Group A carcinogen | [1,13,15,16,17] |
| Ochratoxin A | 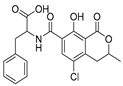 | Aspergillus alliaceus, Aspergillus melleus, Aspergillus cabonarius, Aspergillus glaucus, Aspergillus niger, Penicillium viridicatum | Group 2B possible human carcinogen | [13,18,26] |
| Patulin | 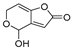 | Penicillium expansum, Penicillium patulum, Penicillium crustosum | Group 3 | [19,25] |
| Toxin | Crops | Detection Techniques (Targeted or Non-Targeted) | LoD * | LoQ * | Ref. |
|---|---|---|---|---|---|
| AFB1, G1 | Peanuts, corn, soy beans | Targeted and Non-targeted HPLC-ESI-MS-qTOF, ESI+ UHPLC-ESI-MS/MS, sSRM | 0.1–0.3 µg/kg | 0.2–0.9 µg/kg | [50] |
| AFB1, B2, G1, G2, M1, OTA | Feed and feed raw materials (silage, maize, wheat, wheat by-products, barley, soy beans, sunflower seeds) | Targeted and Non-targeted LC-ESI-MS/MS (QTRAP) ESI+, ESI−, sSRM | n/a | n/a | [72] |
| AFB1, B2, G1, G2, M1, M2, OTA, OTB, Patulin | Almonds, hazelnuts, peanuts, pistachio | Targeted UHPLC-ESI-MS/MS (qQq), ESI+, ESI−, sSRM | n/a | AFB1 3.0 µg/kg AFB2 10.0 µg/kg AFG1 10.0 µg/kg AFG2 8.2 µg/kg AFM1 7.9 µg/kg OTA 15.0 µg/kg OTB 9.9 µg/kg PAT n/a | [53] |
| AFB1, B2, G1, G2, OTA | Barley | Targeted GC-MS/MS (qQq), EI, derivatizied, LC-ESI-MS/MS (QTRAP), ESI | AFs 2.0 ng/kg OTA 2.0 ng/kg | AFs 3.5 ng/kg OTA 3.5 ng/kg | [58] |
| AFB1, B2, G1, G2 | Rice, sorghum | Targeted LC-ESI-MS/MS or UHPLC-ESI-MS/MS (tandem quadrupole), ESI+, sSRM | 0.1–1.0 µg/kg | 0.28–0.9 µg/kg | [73] |
| AFB1, B2, G1, G2, OTA | Wheat, corn and rice cereals | Targeted UHPLC-ESI-MS/MS (tandem quadrupole) ESI+, sSRM | 0.1–5.0 µg/kg, (AFB1 0.03 µg/kg) | 0.1–25.0 µg/kg | [74] |
| AFB1, B2, G1, G2, M1, OTA | Various foods and feed (24 types of corn feeds, peanut butter) | Targeted UHPLC-ESI-MS/MS (qQq tandem) ESI+, ESI−, sSRM | AFs 0.003 µg/kg AFG2 0.006 µg/kg OTA 0.064 µg/kg | AFs 0.01 µg/kg AFG2 0.02 µg/kg OTA 0.21 µg/kg | [75] |
| AFB1, B2, G1, G2, OTA | Maize | Targeted LC-ESI-MS/MS (QTRAP qQq) ESI+, ESI−, sSRM | AFB1 0.6 µg/kg AFB2 0.3 µg/kg AFG1 0.4 µg/kg AFG2 0.8 µg/kg OTA 0.6 µg/kg | n/a | [76] |
| AFB1, B2, G1, G2, OTA | Barley based breakfast cereals, maize, peanuts | Targeted UHPLC-ESI-MS/MS (QTRAP qQq) ESI+ ESI− (in single run), sSRM | AFs 0.05 µg/kg OTA 0.1 µg/kg | AFs 0.1 µg/kg OTA 0.25 µg/kg | [77] |
| AFB1, B2, G1, G2, OTA | Durum wheat, corn flakes, maize and maize crackers | Targeted LC-ESI-MS/MS (QTRAP qQq) ESI+ ESI−, sSRM | n/a | AFs 1.0 µg/kg OTA 1.0 µg/mg | [78] |
| AFB1, B2, G1, G2, OTA | Muesli, wheat flakes, oats, raisins, sultanas, whey powder, hazelnuts, whole meal bread | Targeted LC-ESI-MS/MS (tandem quadrupole) ESI+, sSRM | AFB1 0.05 ng/g AFB2 0.03 ng/g AFG1 0.03 ng/g AFG2 0.03 ng/g OTA 0.03 ng/g | AFB1 0.1 ng/g AFB2 0.05 ng/g AFG1 0.05 ng/g AFG2 0.05 ng/g OTA 0.4 ng/g | [79] |
| AFB1, B2, G1, G2, OTA | Barley, corn, corn gluten, infant cereals, oat, rice, rye, wheat | Targeted LC-ESI-MS/MS (QTRAP qQq tandem mass) ESI+, ESI−, sSRM | n/a | AFs 1.0–10.0 µg/kg OTA 0.5–2.5 µg/kg | [80] |
| AFB1, B2, G1, G2, OTA | Maize, rice, wheat | Targeted LC-ESI-MS/MS (qQq tandem) ESI+, sSRM | AFB1 0.12–0.21 g/kg AFB2 0.06–0.7 µg/kg AFG1 0.07–2.3 µg/kg AFG2 0.11–2.2 µg/kg OTA 0.18–3.2 µg/kg | AFB1 0.12–0.21 µg/kg AFB2 0.06–0.7 µg/kg AFG1 0.07–2.3 µg/kg AFG2 0.11–2.2 µg/kg OTA 0.18–3.2 µg/kg | [81] |
| AFs, OTA | Black pepper, infant food (apple baby food), paprika, sunflower seed, wheat flour | Targeted UHPLC-ESI-MS/MS (QTRAP tandem) ESI+, ESI−, sSRM, Non-targeted UHPLC-ESI-HRMS (TOF) ESI+ ESI− | n/a | n/a | [82] |
| AFB1, B2, G1, G2, OTA, OTB, OTC, Patulin | Maize, wheat | Targeted HPLC-ESI-MS/MS (QTRAP qQq), ESI+, ESI−, sSRM | 0.03–220 µg/kg | n/a | [83] |
| AFB1, B2, G1, G2, M1, OTA, Patulin | Apple puree, green pepper, hazelnut, maize | Targeted UHPLC-ESI-MS/MS (QTRAP) ESI+, ESI−, sSRM | AFB1 0.6 µg/kg AFB2 0.6 µg/kg AFG1 1.2 µg/kg AFG2 2.3 µg/kg AFM1 0.6 µg/kg OTA 1.2 µg/kg PAT 35.9 µg/kg | AFB1 1.9 µg/kg AFB2 4.0 µg/kg AFG1 7.6 µg/kg AFG2 8.7 µg/kg AFM1 2.1 µg/kg OTA 3.7 µg/kg PAT 119.7 µg/kg | [65] |
| AFB1, B2, G1, G2, OTA | Barley | Targeted UHPLC-HRMS (Orbitrap) Heated EPI (HEPI), HEPI+, HEPI− | n/a | n/a | [84] |
| OTA | Barley | Targeted UHPLC-FTHRMS HEPI, HEPI+, HEPI− | n/a | n/a | [85] |
| AFB1, B2, G1, G2, OTA | Black radish, Ginkgo biloba, garlic, soy | Targeted UHPLC-ESI-MS/MS (qQq), ESI+, sSRM | AFs 6.0 ng/g OTA 1.0 ng/g | AFs 2.0 ng/g OTA 0.3 ng/g | [86] |
| AFB1, B2, G1, G2, M1, OTA, OTB | Maize, groundnut, sorghum, millet, rice, wheat, soy, dried fruits, infant foods, other processed food, animal feed | Targeted HPLC-ESI-MS/MS (QTRAP) ESI+, ESI−, sSRM | AFB1 3.0 µg/kg AFB2 6.0 µg/kg AFG1 8.0 µg/kg AFG2 8.0 µg/kg AFM1 4.0 µg/kg OTA, OTB 5.0 µg/kg | n/a | [87] |
| AFB1, B2, G1, G2, M1, OTA | Breakfast cereals (maize, wheat, rice, multigrain, chocolate) | Targeted HPLC-fluorescence detector-EI-MS/MS, sSRM | AFB1 0.003 µg/kg AFB2 0.001 µg/kg AFG1 0.006 µg/kg AFG2 n/a AFM1 0.011 µg/kg OTA 0.006 µg/kg | AFB1 0.009 µg/kg AFB2 0.004 µg/kg AFG1 0.018 µg/kg AFG2 n/a AFM1 0.032 µg/kg OTA 0.019 µg/kg | [88] |
| OTA | Wheat flour, coffee, spices, wine, beer | Targeted HPLC-MS/MS (ion trap), (1) ESI+ (2) APCI, sSRM | 0.5 µg/kg | 1.4 µg/kg | [89] |
| AFB1, B2, G1, G2 | Peanut, peanut butter, spices, figs | Targeted LC-APCI-MS/MS (qQq), APCI+, sSRM, targeted | 0.1 µg/kg | n/a | [90] |
| Patulin | Wheat, rice, spelt, oat, soy, tapioca based cereals (cassava), pasta, infant food | Targeted GC-MS/MS (qQq), electron impact ion source (EI), SRM, derivatizied, targeted | n/a | 5–10 µg/kg | [56] |
| Genomic Tools | Mycotoxins | Crops | Comments | Ref. |
|---|---|---|---|---|
| Ion Torrent Personal Genome Machine (PGM) | Aflatoxins | — | Whole genome sequencing | [99] |
| Microarray analysis, quantitative reverse transcription-PCR (qRT-PCR) | Aflatoxins | — | Aflatoxin biosynthesis | [100] |
| Microarray analysis | Aflatoxins | — | Whole genome sequencing | [101] |
| Microarray analysis | Aflatoxins | — | Gene expression profiles | [102] |
| Whole genome sequencing | — | Identify genes differentially expressed in wild-type veA and veA mutant strains that could be involved in aflatoxin production. | [92] | |
| RT-PCR and reverse-transcription PCR | Peanuts | Develop a screening method | [96] | |
| PCR and LAMP-based group specific | Rice, nuts, raisins, dried figs | Develop a screening method to detect several aflatoxin producing species in a single analysis | [103] | |
| Microarray | Aflatoxins, ochratoxin A | Wheat grain | Rapid detection for mycotoxins | [104] |
| Fungal Strains | Genomic Size (Mbp *) | Mycotoxin | Mycotoxigenic | Ref. |
|---|---|---|---|---|
| LOAM00000000 flavus | 36.0 | Aflatoxin | Yes | [108] |
| LIZI00000000 flavus | 36.4 | Aflatoxin | Yes | |
| LIZJ00000000 flavus | 36.3 | Aflatoxin | Yes | |
| LOAK00000000 flavus | 35.9 | Aflatoxin | Yes | |
| LOAL00000000 flavus | 35.8 | Aflatoxin | Yes | |
| LOAP00000000 parasiticus | 30.1 | Aflatoxin | Yes | |
| NRRL 13137 nominus | 36.1 | Aflatoxin | Yes | [99] |
| Aspergillus korhogoensis | N/a | Aflatoxin | Yes | [116] |
| Aspergillus westerdijkiae | 36.1 | Ochratoxin A | Yes | [111] |
| Aspergillus carbonarius | 36 | Ochratoxin A | Yes | [110] |
| Penicillium expansum | 33.52 | Patulin | Yes | [115] |
| Penicillium italicum | 28.99 | Patulin | Yes | [115] |
| Mycotoxin | Studies | Outcomes | Ref. |
|---|---|---|---|
| Aflatoxin B1 | Identification of essential transcription factors for adequate DNA damage response after benzo (a) pyrene and aflatoxin B1 exposure by combining transcriptomics with functional genomics. | Transcriptomics and functional genomics tools used to investigate the genotoxicity of aflatoxin B1. | [132] |
| Aflatoxin B1 induces persistent epigenomic effects in primary human hepatocytes associated with hepatocellular carcinoma. | Transcriptomics and epigenome studies used to understand the mechanisms of hepatocellular carcinoma development. | [8] | |
| Quercetin tests negative for genotoxicity in transcriptome analyses of liver and small intestine of mice. | Genotoxicity related pathways in mice liver and small intestine. | [133] | |
| Combined cytotoxicity of aflatoxin B1 and deoxynivalenol to hepatoma HepG2/C3A cells. | Different cytotoxicity pathways and their apoptotic process might be the mechanism of the synergistic cytotoxicity of HepG2/C3A carcinoma cells. | [134] | |
| Integrated analysis of transcriptomics and metabolomics profiles in aflatoxin B1-induced hepatotoxicity in rat. | Gluconeogenesis, lipid metabolism disorder, and induced hepatotoxicity affect majorly after the acute AFB1 exposure. | [122] | |
| Identification of early target genes of aflatoxin B1 in human hepatocytes, inter-individual variability and comparison with other genotoxic compounds. | Gene subset from AFB1 induced human hepatocytes identified several genes which are potential biomarkers of genotoxic compounds. | [135] | |
| Aflatoxins | Use of functional genomics to assess the climate change impact on Aspergillus flavus and aflatoxin production. | Global temperature, water availability and rising CO2 levels affect the expression of the aflatoxin biosynthetic regulatory gene aflR. | [109] |
| Ochratoxin A | Different toxicity mechanisms for citrinin and ochratoxin A revealed by transcriptomic analysis in yeast. | OTA deregulates developmental genes. | [136] |
| Disruption of liver development and coagulation pathway by ochratoxin A in embryonic zebrafish. | OTA exposure led to a deficiency of coagulation factors. | [137] | |
| Transcriptomic alterations induced by OTA in rat and human renal proximal tubular in vitro models and comparison to rat in vivo model. | The study provided a non-genotoxic mechanism of OTA-induced carcinogenicity. | [138] | |
| Patulin | Transcriptomic responses of the basidiomycete Sporobolomyces sp. to the mycotoxin patulin. | Exposure to PAT directed the changes in gene expression in Sporobolomyces sp. This finding may lead to develop a bio-detoxification process. | [139] |
| Mycotoxin | Fungal Strains | Study | Analysis Techniques | Outcome | Ref. |
|---|---|---|---|---|---|
| Aflatoxin B1 | Aspergillus flavus | Proteomic analysis reveals an aflatoxin-triggered immune response in cotyledons of Arachis hypogaea infected with Aspergillus flavus. | 2-D gel electrophoresis and MALDI-TOF/TOF mass spectrometer. | Three grades of the immune response in A. hypogaea during infection with toxigenic A. flavus were identified. PAMP-triggered immunity, effector-triggered immunity and metabolite-triggered immunity. | [162] |
| Aspergillus flavus | Comparative leaf proteomics of drought-tolerant and-susceptible peanut in response to water stress. | 2-D gel electrophoresis and MALDI-TOF/TOF mass spectrometer. | 42 unique proteins showed interactions in the tolerant cultivar. | [167] | |
| Aspergillus flavus | Insight into the global regulation of laeA in Aspergillus flavus based on proteomic profiling | Protein extraction, trypsin digestion, TMT-labelling and HPLC fractionation and LC-MS/MS | laeA gene affects cell morphology and contributes to the production of aflatoxin production. | [168] | |
| Aspergillus flavus | Proteome analysis of A. flavus isolate-specific responses to oxidative stress in relationship to aflatoxin production capability. | Protein digestion and iTRAQ * labelling | 1173 proteins were identified, and 220 were differentially expressed. | [163] | |
| Ochratoxin A | Aspergillus carbonarius | Proteome analysis of the fungus Aspergillus carbonarius under ochratoxin A producing conditions. | 2-D gel electrophoresis and MALDI-TOF/TOF mass spectrometer. | Nine differential proteins were identified by MALDI-MS/MS and MASCOT. Identified proteins were involved in regulation, amino acid metabolism, oxidative stress and sporulation. A protein with 126.5 fold higher abundance in high OTA-producing strain showed homology with CipC. | [155] |
| Arabidopsis thaliana | iTRAQ mitoproteome Analysis reveals mechanisms of programmed cell death in Arabidopsis thaliana induced by ochratoxin A | iTRAQ * Analysis | The study investigated the toxicity mechanism of OTA on the host plant; their results indicated that OTA induced PCD in A. thaliana. 42 and 43 proteins were identified within 8 and 24 h. those proteins were mainly involved in perturbation of the mitochondrial electron transport chain, interfering with ATP synthesis and inducing PCD | [169] | |
| Patulin | Penicillium expansum | Identification of differentially expressed genes involved in spore germination of Penicillium expansum by comparative transcriptome and proteome approaches. | RNA-seq (RNA sequencing) and iTRAQ * (isobaric tags for relative and absolute quantitation) approaches. | A total of 3026 differentially expressed genes, 77 differentially expressed predicted transcription factors and 489 differentially expressed proteins identified. Posttranscriptional regulation and modification serve essential roles in the management of fungal germination. | [164] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Eshelli, M.; Qader, M.M.; Jambi, E.J.; Hursthouse, A.S.; Rateb, M.E. Current Status and Future Opportunities of Omics Tools in Mycotoxin Research. Toxins 2018, 10, 433. https://doi.org/10.3390/toxins10110433
Eshelli M, Qader MM, Jambi EJ, Hursthouse AS, Rateb ME. Current Status and Future Opportunities of Omics Tools in Mycotoxin Research. Toxins. 2018; 10(11):433. https://doi.org/10.3390/toxins10110433
Chicago/Turabian StyleEshelli, Manal, M. Mallique Qader, Ebtihaj J. Jambi, Andrew S. Hursthouse, and Mostafa E. Rateb. 2018. "Current Status and Future Opportunities of Omics Tools in Mycotoxin Research" Toxins 10, no. 11: 433. https://doi.org/10.3390/toxins10110433
APA StyleEshelli, M., Qader, M. M., Jambi, E. J., Hursthouse, A. S., & Rateb, M. E. (2018). Current Status and Future Opportunities of Omics Tools in Mycotoxin Research. Toxins, 10(11), 433. https://doi.org/10.3390/toxins10110433






