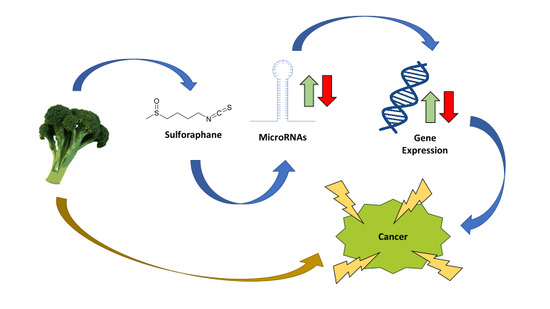
The Role of MicroRNAs in the Chemopreventive Activity of Sulforaphane from Cruciferous Vegetables
Abstract
1. Colorectal Cancer
2. Cruciferous Vegetables
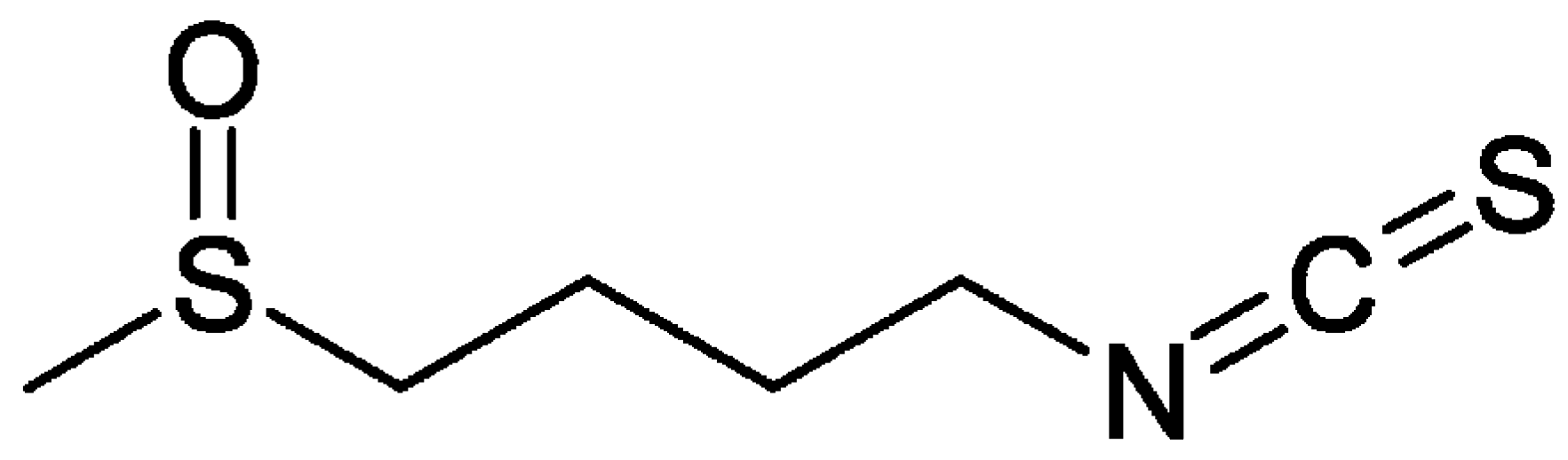
3. Sulforaphane
4. MicroRNAs
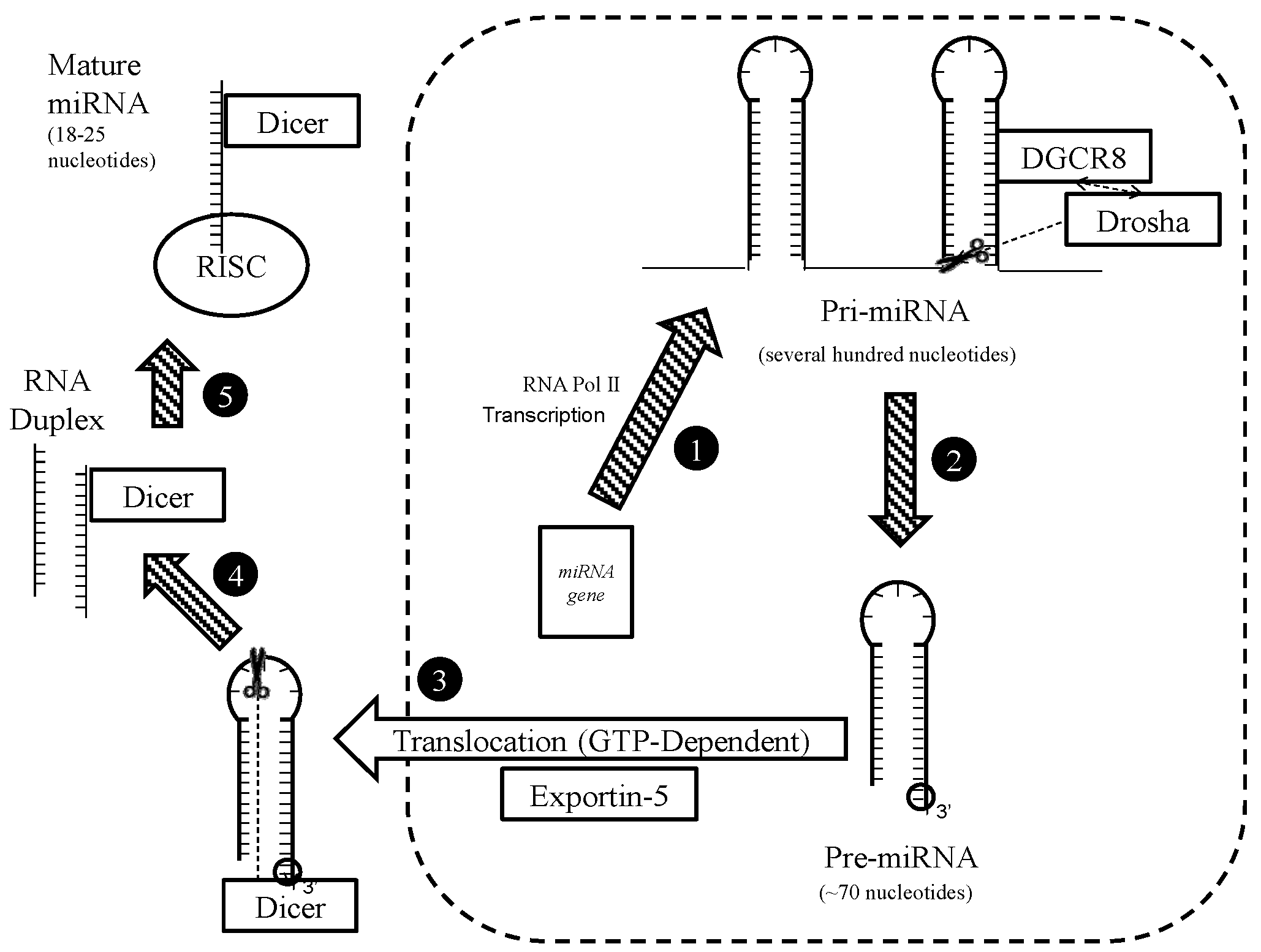
4.1. Biogenesis
4.2. Notes on Nomenclature
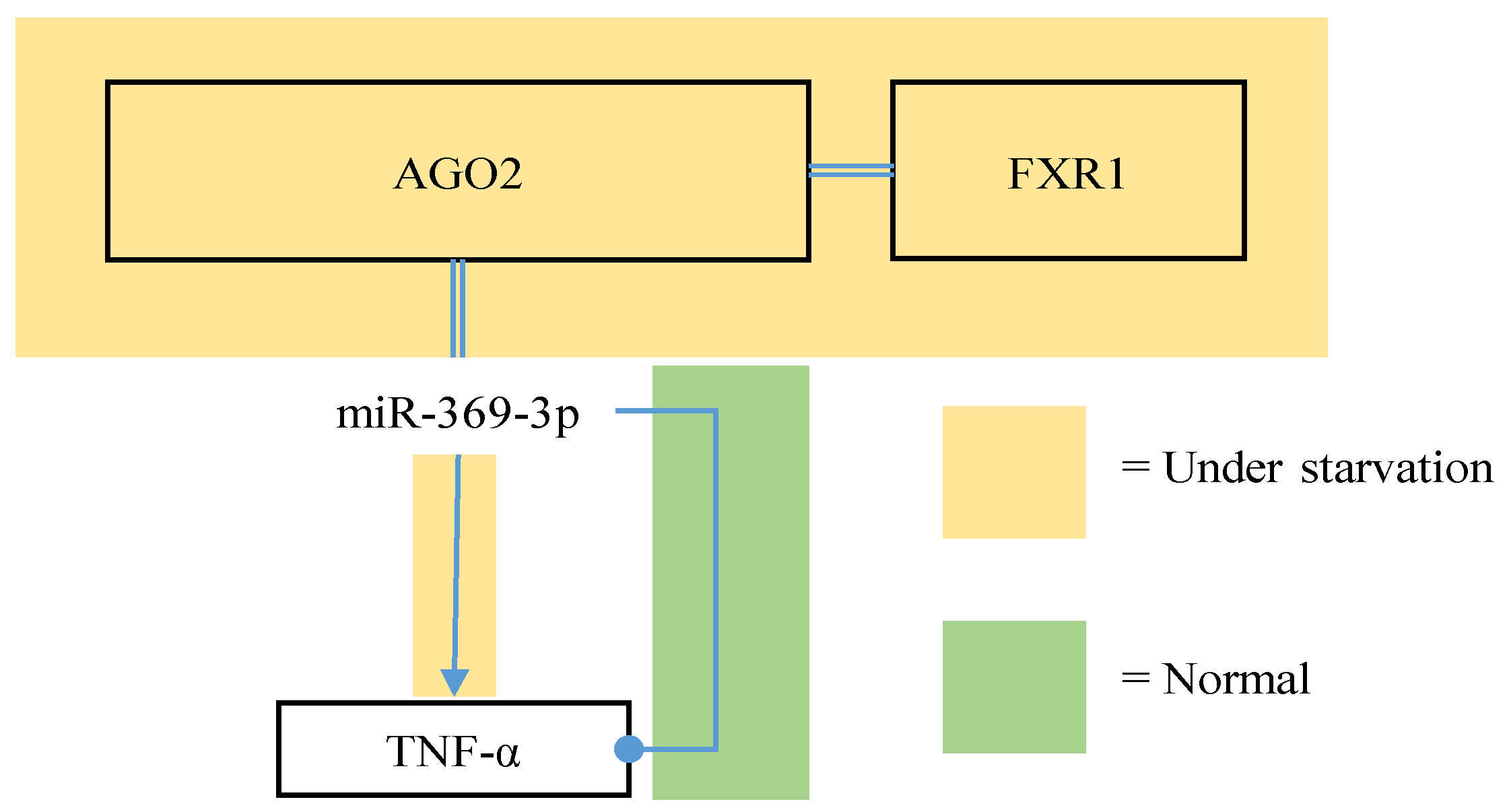
4.3. Activities
4.4. IsomiRs
5. Linking Sulforaphane, MicroRNAs, and Colorectal Cancer
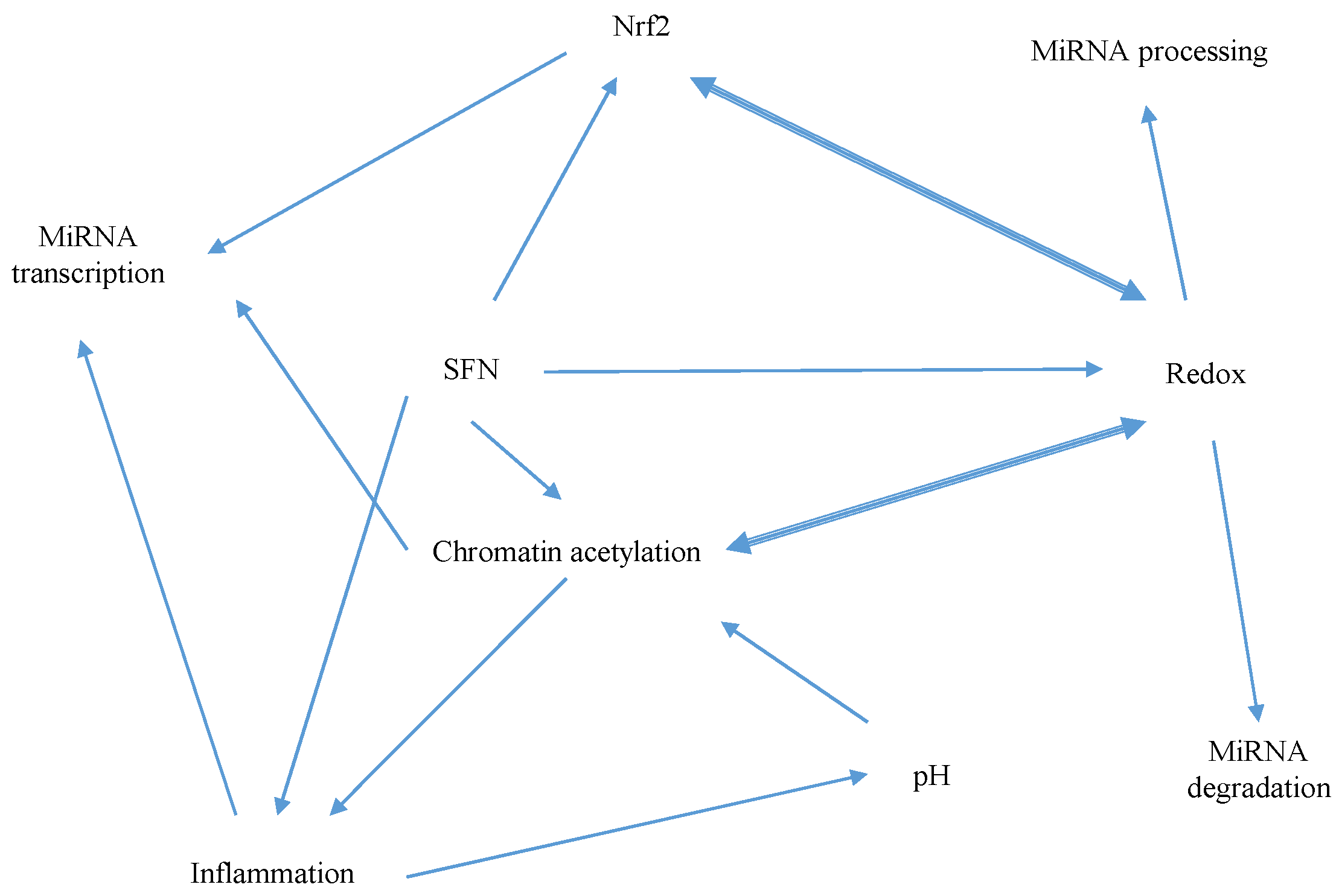
5.1. Mechanisms of Sulforaphane-Mediated Modulation
5.2. Interaction of MicroRNAs with Pathogenesis
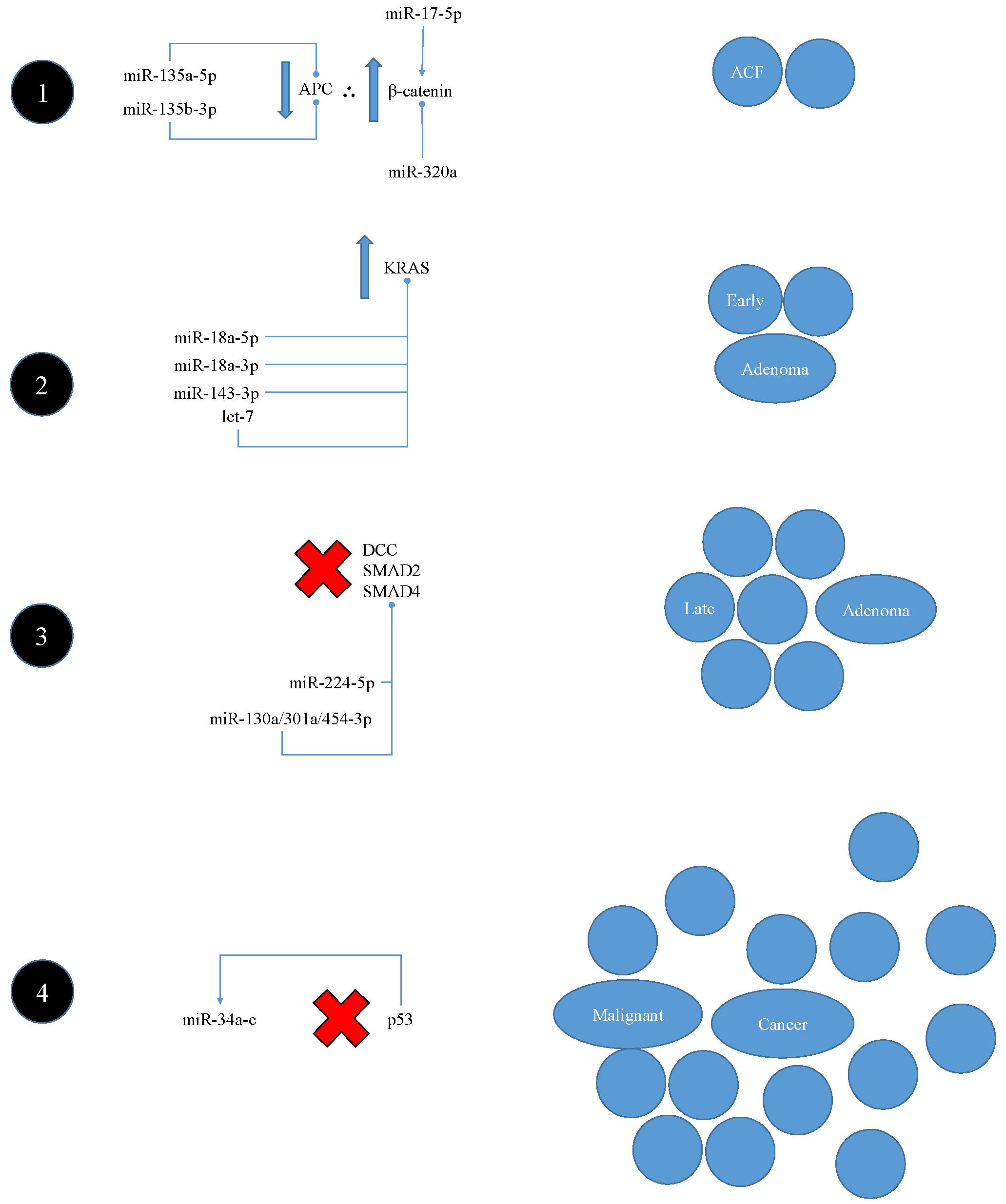
5.2.1. Interaction with Classic Vogelstein-Model Pathogenesis
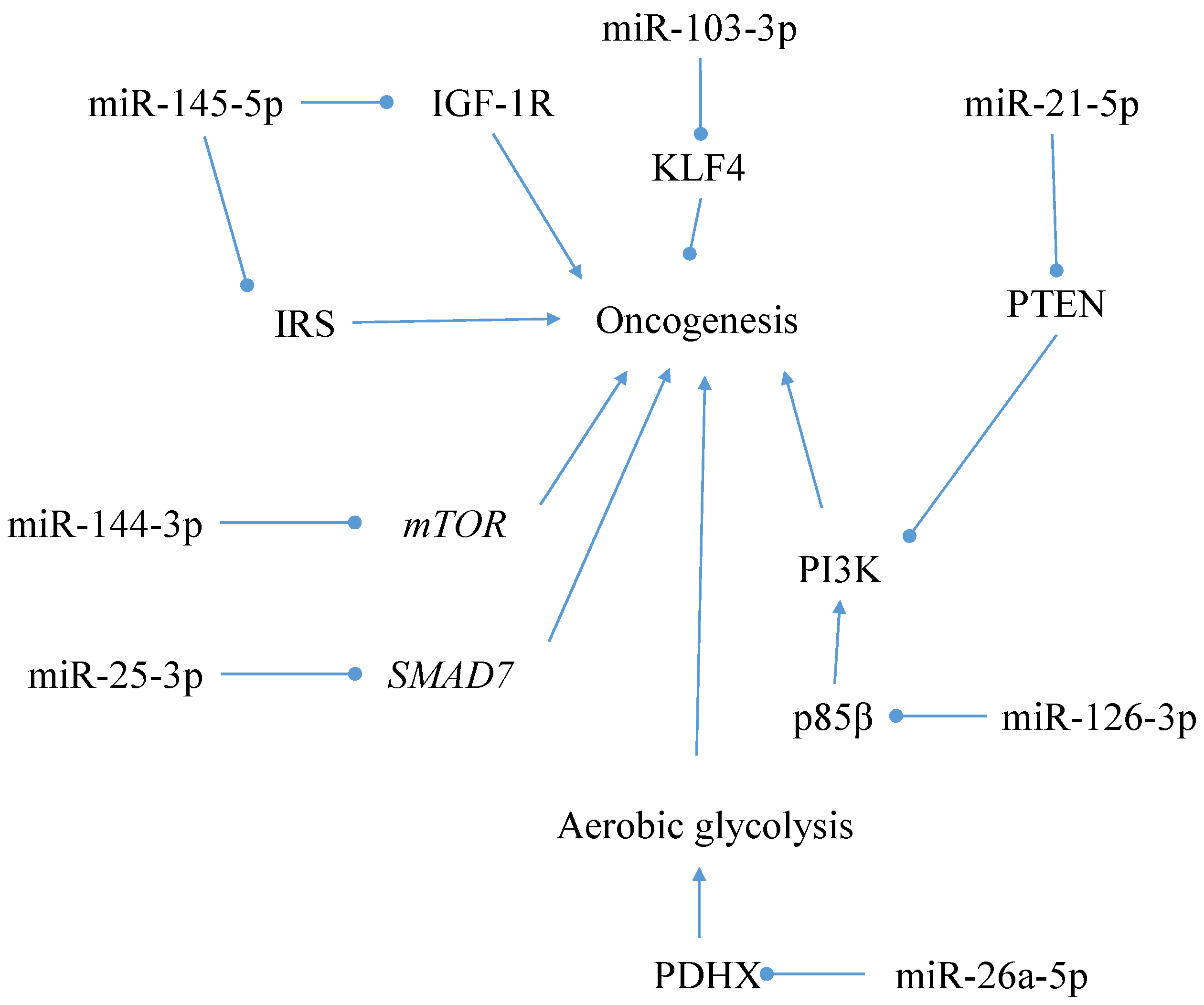
5.2.2. Alternative Pathogenesis Model Interaction
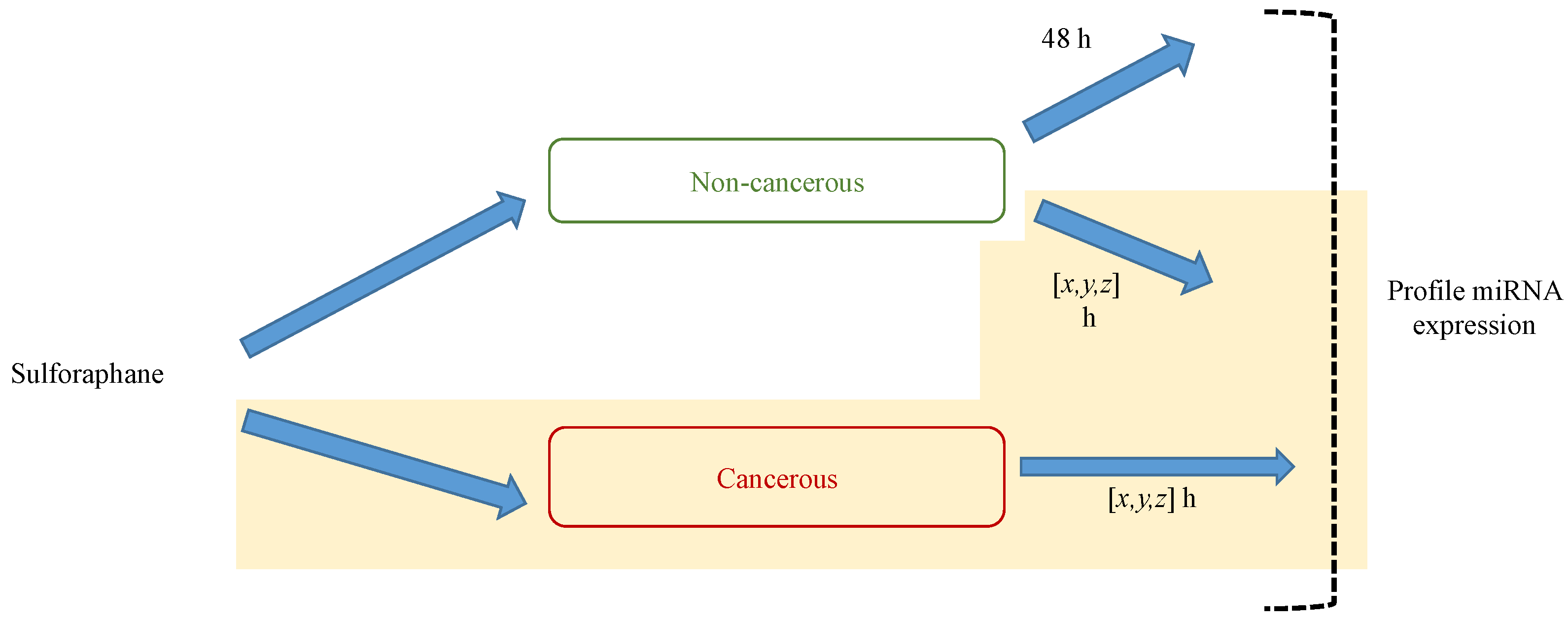
6. MicroRNA Assay Methods
6.1. Array-Based Methods
6.2. Cloning and Deep Sequencing
6.3. Comparison of Approaches to MicroRNA Profiling
7. Summary
Acknowledgments
Author Contributions
Conflicts of Interest
References
- World Health Organization: International Agency for Research on Cancer. Colorectal cancer estimated incidence, mortality and prevalence worldwide in 2012. Available online: http://globocan.iarc.fr/Pages/fact_sheets_cancer.aspx (accessed on 17 October 2016).
- Lozano, R.; Naghavi, M.; Foreman, K.; Lim, S.; Shibuya, K.; Aboyans, V.; Abraham, J.; Adair, T.; Aggarwal, R.; et al. Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: A systematic analysis for the Global Burden of Disease Study 2010. Lancet 2012, 380, 2095–2128. [Google Scholar] [CrossRef]
- Smith, D.; Ballal, M.; Hodder, R.; Soin, G.; Selvachandran, S.N.; Cade, D. Symptomatic presentation of early colorectal cancer. Ann. R. Coll. Surg. Engl. 2006, 88, 185–190. [Google Scholar] [CrossRef] [PubMed]
- Bener, A. Colon cancer in rapidly developing countries: Review of the lifestyle, dietary, consanguinity and hereditary risk factors. Oncol. Rev. 2011, 5, 5–11. [Google Scholar] [CrossRef]
- Phatak, S.; Larson, D.; Hunter, A.; Neal, C.; Contreras, K.; Pontsler, K.; Armbrust, T.; Rodriguez, D.; Abercrombie, B.; Hintze, K.; et al. Ancestral and multi-generational consumption of the total Western diet in mice promotes colitis-associated colorectal cancer in third-generation offspring. FASEB J. 2017, 31. [Google Scholar] [CrossRef]
- Bowel Cancer UK. Understanding Bowel Cancer: Diet & Exercise. Available online: http://www.bowelcanceruk.org.uk/understanding-bowel-cancer/diet-exercise-(1)/ (accessed on 2 September 2016).
- Tse, G.; Eslick, G.D. Cruciferous vegetables and risk of colorectal neoplasms: a systematic review and meta-analysis. Nutr. Cancer 2014, 66, 128–139. [Google Scholar] [CrossRef] [PubMed]
- Chan, R.; Lok, K.; Woo, J. Prostate cancer and vegetable consumption. Mol. Nutr. Food Res. 2009, 53, 201–216. [Google Scholar] [CrossRef] [PubMed]
- Koroleva, O.A.; Davies, A.; Deeken, R.; Thorpe, M.R.; Tomos, A.D.; Hedrich, R. Identification of a New Glucosinolate-Rich Cell Type in Arabidopsis Flower Stalk. Plant Physiol. 2000, 124, 599–608. [Google Scholar] [CrossRef] [PubMed]
- Andréasson, E.; Jørgensen, L.B.; Höglund, A.; Rask, L.; Meijer, J. Different Myrosinase and Idioblast Distribution in Arabidopsis and Brassica napus. Plant Physiol. 2001, 127, 1750–1763. [Google Scholar] [CrossRef] [PubMed]
- Burow, M.; Bergner, A.; Gershenzon, J.; Wittstock, U. Glucosinolate hydrolysis in Lepidium sativum—Identification of the thiocyanate-forming protein. Plant Mol. Biol. 2007, 63, 49–61. [Google Scholar] [CrossRef] [PubMed]
- Clarke, J.D.; Dashwood, R.H.; Ho, E. Multi-targeted prevention of cancer by sulforaphane. Cancer Lett. 2008, 269, 291–304. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y.; Wang, X.; Zhao, S.; Ma, C.; Cui, J.; Zheng, Y. Sulforaphane Protects against Cardiovascular Disease via Nrf2 Activation. Oxid. Med. Cell Longev. 2015, 2015, 13. [Google Scholar] [CrossRef] [PubMed]
- Davidson, R.K.; Jupp, O.; Bao, Y.; MacGregor, A.J.; Donell, S.T.; Cassidy, A.; Clark, I.M. Can sulforaphane prevent the onset or slow the progression of osteoarthritis? Nutr. Bull. 2016, 41, 175–179. [Google Scholar] [CrossRef]
- Davidson, R.; Gardner, S.; Jupp, O.; Bullough, A.; Butters, S.; Watts, L.; Donell, S.; Traka, M.; Saha, S.; Mithen, R.; et al. Isothiocyanates are detected in human synovial fluid following broccoli consumption and can affect the tissues of the knee joint. Sci. Rep. 2017, 7, 3398. [Google Scholar] [CrossRef] [PubMed]
- Ramos-Gomez, M.; Kwak, M.K.; Dolan, P.M.; Itoh, K.; Yamamoto, M.; Talalay, P.; Kensler, T.W. Sensitivity to carcinogenesis is increased and chemoprotective efficacy of enzyme inducers is lost in nrf2 transcription factor-deficient mice. Proc. Natl. Acad. Sci. 2001, 98, 3410–3415. [Google Scholar] [CrossRef] [PubMed]
- Yanaka, A.; Mutoh, M. Su2056 Dietary Intake of Sulforaphane Glucosinolates Inhibits Colon Tumorigenesis in Mice Treated With Azoxymethane and Radioactive Cesium. Gastroenterology 2016, 150, S623. [Google Scholar] [CrossRef]
- Pereira, L.; Silva, P.; Duarte, M.; Rodrigues, L.; Duarte, C.; Albuquerque, C.; Serra, A. Targeting Colorectal Cancer Proliferation, Stemness and Metastatic Potential Using Brassicaceae Extracts Enriched in Isothiocyanates: A 3D Cell Model-Based Study. Nutrients 2017, 9, 368. [Google Scholar] [CrossRef] [PubMed]
- Gupta, P.; Kim, B.; Kim, S.H.; Srivastava, S.K. Molecular targets of isothiocyanates in cancer: Recent advances. Mol. Nutr. Food Res. 2014, 58, 1685–1707. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.C.; Shih, T.Y.; Kuo, C.L.; Ma, Y.S.; Yang, J.L.; Wu, P.P.; Huang, Y.P.; Lai, K.C.; Chung, J.G. Sulforaphane Induces Cell Death Through G2/M Phase Arrest and Triggers Apoptosis in HCT 116 Human Colon Cancer Cells. Am. J. Chin. Med. 2016, 44, 1289–1310. [Google Scholar] [CrossRef] [PubMed]
- Samantha, L.M.; Rishabh, K.; Trygve, O.T. Mechanisms for the inhibition of colon cancer cells by sulforaphane through epigenetic modulation of microRNA-21 and human telomerase reverse transcriptase (hTERT) down-regulation. Curr. Cancer Drug Targets 2017, 17, 1–10. [Google Scholar]
- Wang, Y.; Dacosta, C.; Wang, W.; Zhou, Z.; Liu, M.; Bao, Y. Synergy between sulforaphane and selenium in protection against oxidative damage in colonic CCD841 cells. Nutr. Res. 2015, 35, 610–617. [Google Scholar] [CrossRef] [PubMed]
- Yueh, M.F.; Tukey, R.H. Nrf2-Keap1 signaling pathway regulates human UGT1A1 expression in vitro and in transgenic UGT1 mice. J. Biol. Chem. 2007, 282, 8749–8758. [Google Scholar] [CrossRef] [PubMed]
- Harman, D. Aging: a theory based on free radical and radiation chemistry. J. Gerontol. 1956, 11, 298–300. [Google Scholar] [CrossRef] [PubMed]
- Pi, J.; Zhang, Q.; Fu, J.; Woods, C.G.; Hou, Y.; Corkey, B.E.; Collins, S.; Andersen, M.E. ROS signaling, oxidative stress and Nrf2 in pancreatic beta-cell function. Toxicol. Appl. Pharmacol. 2010, 244, 77–83. [Google Scholar] [CrossRef] [PubMed]
- Gopalakrishna, R.; Gundimeda, U. Antioxidant regulation of protein kinase C in cancer prevention. J. Nutr. 2002, 132, 3819S–3823S. [Google Scholar] [PubMed]
- Guerrero-Beltrán, C.E.; Calderón-Oliver, M.; Pedraza-Chaverri, J.; Chirino, Y.I. Protective effect of sulforaphane against oxidative stress: Recent advances. Exp. Toxicol. Pathol. 2012, 64, 503–508. [Google Scholar] [CrossRef] [PubMed]
- Higgins, L.G.; Kelleher, M.O.; Eggleston, I.M.; Itoh, K.; Yamamoto, M.; Hayes, J.D. Transcription factor Nrf2 mediates an adaptive response to sulforaphane that protects fibroblasts in vitro against the cytotoxic effects of electrophiles, peroxides and redox–cycling agents. Toxicol. Appl. Pharmacol. 2009, 237, 267–280. [Google Scholar] [CrossRef] [PubMed]
- Sestili, P.; Paolillo, M.; Lenzi, M.; Colombo, E.; Vallorani, L.; Casadei, L.; Martinelli, C.; Fimognari, C. Sulforaphane induces DNA single strand breaks in cultured human cells. Mutat. Res. 2010, 689, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Keum, Y.S.; Yu, S.; Chang, P.P.; Yuan, X.; Kim, J.H.; Xu, C.; Han, J.; Agarwal, A.; Kong, A.N. Mechanism of Action of Sulforaphane: Inhibition of p38 Mitogen-Activated Protein Kinase Isoforms Contributing to the Induction of Antioxidant Response Element-Mediated Heme Oxygenase-1 in Human Hepatoma HepG2 Cells. Cancer Res. 2006, 66, 8804–8813. [Google Scholar] [CrossRef] [PubMed]
- Pal, S.; Badireenath Konkimalla, V. Hormetic Potential of Sulforaphane (SFN) in Switching Cells’ Fate Towards Survival or Death. Mini. Rev. Med. Chem. 2016, 16, 980–995. [Google Scholar] [CrossRef] [PubMed]
- Tierens, K.F.M.J.; Thomma, B.P.H.J.; Brouwer, M.; Schmidt, J.; Kistner, K.; Porzel, A.; Mauch-Mani, B.; Cammue, B.P.A.; Broekaert, W.F. Study of the Role of Antimicrobial Glucosinolate-Derived Isothiocyanates in Resistance of Arabidopsis to Microbial Pathogens. Plant Physiol. 2001, 125, 1688–1699. [Google Scholar] [CrossRef] [PubMed]
- Mattson, M.P.; Calabrese, E.J. Hormesis; Mattson, M.P., Calabrese, E.J.B., Eds.; Humana Press: New York, NY, USA, 2010; Volume 1, p. 59. [Google Scholar]
- Myzak, M.C.; Hardin, K.; Wang, R.; Dashwood, R.H.; Ho, E. Sulforaphane inhibits histone deacetylase activity in BPH-1, LnCaP and PC-3 prostate epithelial cells. Carcinogenesis 2006, 27, 811–819. [Google Scholar] [CrossRef] [PubMed]
- Lau, A.; Villeneuve, N.F.; Sun, Z.; Wong, P.K.; Zhang, D.D. Dual roles of Nrf2 in cancer. Pharmacol. Res. 2008, 58, 262–270. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Wang, W.; Zhou, Z.; Sun, C. Benefits and Risks of the Hormetic Effects of Dietary Isothiocyanates on Cancer Prevention. PLoS ONE 2014, 9, e114764. [Google Scholar] [CrossRef] [PubMed]
- Murray-Stewart, T.; Hanigan, C.L.; Woster, P.M.; Marton, L.J.; Casero, R.A., Jr. Histone Deacetylase Inhibition Overcomes Drug Resistance through a miRNA-Dependent Mechanism. Mol. Cancer Ther. 2013, 12, 2088–2099. [Google Scholar] [CrossRef] [PubMed]
- Bat-Chen, W.; Golan, T.; Peri, I.; Ludmer, Z.; Schwartz, B. Allicin Purified From Fresh Garlic Cloves Induces Apoptosis in Colon Cancer Cells Via Nrf2. Nutr. Cancer 2010, 62, 947–957. [Google Scholar] [CrossRef] [PubMed]
- Saddawi-Konefka, R.; Seelige, R.; Gross, E.T.; Levy, E.; Searles, S.C.; Washington, A., Jr.; Santosa, E.K.; Liu, B.; O’Sullivan, T.E.; Harismendy, O.; et al. Nrf2 Induces Il-17D to Mediate Tumor and Virus Surveillance. Cell Rep. 2016, 16, 2348–2358. [Google Scholar] [CrossRef] [PubMed]
- Nian, H.; Delage, B.; Ho, E.; Dashwood, R.H. Modulation of histone deacetylase activity by dietary isothiocyanates and allyl sulfides: Studies with sulforaphane and garlic organosulfur compounds. Environ. Mol. Mutagen 2009, 50, 213–221. [Google Scholar] [CrossRef] [PubMed]
- Rajendran, P.; Kidane, A.I.; Yu, T.W.; Dashwood, W.M.; Bisson, W.H.; Lohr, C.V.; Ho, E.; Williams, D.E.; Dashwood, R.H. HDAC turnover, CtIP acetylation and dysregulated DNA damage signaling in colon cancer cells treated with sulforaphane and related dietary isothiocyanates. Epigenetics 2013, 8, 612–623. [Google Scholar] [CrossRef] [PubMed]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Del Carmen Martínez-Jiménez, V.; Méndez-Mancilla, A.; Portales-Pérez, D.P. miRNAs in nutrition, obesity and cancer: The biology of miRNAs in metabolic disorders and its relationship with cancer development. Mol. Nutr. Food Res. 2017. [Google Scholar] [CrossRef] [PubMed]
- Slaby, O.; Sachlova, M.; Brezkova, V.; Hezova, R.; Kovarikova, A.; Bischofova, S.; Sevcikova, S.; Bienertova-Vasku, J.; Vasku, A.; Svoboda, M.; et al. Identification of microRNAs regulated by isothiocyanates and association of polymorphisms inside their target sites with risk of sporadic colorectal cancer. Nutr. Cancer 2013, 65, 247–254. [Google Scholar] [CrossRef] [PubMed]
- Gregory, R.I.; Shiekhattar, R. MicroRNA Biogenesis and Cancer. Cancer Res. 2005, 65, 3509–3512. [Google Scholar] [CrossRef] [PubMed]
- Westholm, J.O.; Lai, E.C. Mirtrons: microRNA biogenesis via splicing. Biochimie 2011, 93, 1897–1904. [Google Scholar] [CrossRef] [PubMed]
- Cifuentes, D.; Xue, H.; Taylor, D.W.; Patnode, H.; Mishima, Y.; Cheloufi, S.; Ma, E.; Mane, S.; Hannon, G.J.; Lawson, N.D.; et al. A Novel miRNA Processing Pathway Independent of Dicer Requires Argonaute2 Catalytic Activity. Science 2010, 328, 1694–1698. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, Biogenesis, Mechanism, and Function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Liu, J.; Rivas, F.V.; Wohlschlegel, J.; Yates, J.R.; Parker, R.; Hannon, G.J. A role for the P-body component GW182 in microRNA function. Nat. Cell Biol. 2005, 7, 1261–1266. [Google Scholar] [CrossRef] [PubMed]
- Vasudevan, S.; Tong, Y.; Steitz, J.A. Switching from Repression to Activation: MicroRNAs Can Up-Regulate Translation. Science 2007, 318, 1931–1934. [Google Scholar] [CrossRef] [PubMed]
- Ørom, U.A.; Nielsen, F.C.; Lund, A.H. MicroRNA-10a Binds the 5′UTR of Ribosomal Protein mRNAs and Enhances Their Translation. Mol. Cell 2008, 30, 460–471. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Liu, M.F. Mechanisms of microRNA-mediated gene regulation. Sci. China C. Life Sci. 2009, 52, 1111–1116. [Google Scholar] [CrossRef] [PubMed]
- Eiring, A.M.; Harb, J.G.; Neviani, P.; Garton, C.; Oaks, J.J.; Spizzo, R.; Liu, S.; Schwind, S.; Santhanam, R.; Hickey, C.J.; et al. miR-328 Functions as an RNA Decoy to Modulate hnRNP E2 Regulation of mRNA Translation in Leukemic Blasts. Cell 2010, 140, 652–665. [Google Scholar] [CrossRef] [PubMed]
- Wilczynska, A.; Bushell, M. The complexity of mirna-mediated repression. Cell Death Differ. 2015, 22, 22–33. [Google Scholar] [CrossRef] [PubMed]
- Neilsen, C.T.; Goodall, G.J.; Bracken, C.P. IsomiRs – the overlooked repertoire in the dynamic microRNAome. Trends in Genetics 2012, 28, 544–549. [Google Scholar] [CrossRef] [PubMed]
- Cloonan, N.; Wani, S.; Xu, Q.; Gu, J.; Lea, K.; Heater, S.; Barbacioru, C.; Steptoe, A.L.; Martin, H.C.; Nourbakhsh, E.; et al. MicroRNAs and their isomiRs function cooperatively to target common biological pathways. Genome Biol. 2011, 12, R126. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Zhang, P.; Ma, Y.; Yang, J.; Moyer, M.P.; Shi, C.; Peng, J.; Qin, H. NIRF is frequently upregulated in colorectal cancer and its oncogenicity can be suppressed by let-7a microRNA. Cancer Lett. 2012, 314, 223–231. [Google Scholar] [CrossRef] [PubMed]
- Tazawa, H.; Tsuchiya, N.; Izumiya, M.; Nakagama, H. Tumor-suppressive mir-34a induces senescence-like growth arrest through modulation of the E2F pathway in human colon cancer cells. Proc. Natl. Acad. Sci. 2007, 104, 15472–15477. [Google Scholar] [CrossRef] [PubMed]
- Strillacci, A.; Griffoni, C.; Sansone, P.; Paterini, P.; Piazzi, G.; Lazzarini, G.; Spisni, E.; Pantaleo, M.A.; Biasco, G.; Tomasi, V. MiR-101 downregulation is involved in cyclooxygenase-2 overexpression in human colon cancer cells. Exp. Cell Res. 2009, 315, 1439–1447. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Sah, J.F.; Beard, L.; Willson, J.K.V.; Markowitz, S.D.; Guda, K. The noncoding RNA, miR-126, suppresses the growth of neoplastic cells by targeting phosphatidylinositol 3-kinase signaling and is frequently lost in colon cancers. Genes Chromosomes Cancer 2008, 47, 939–946. [Google Scholar] [CrossRef] [PubMed]
- Pagliuca, A.; Valvo, C.; Fabrizi, E.; di Martino, S.; Biffoni, M.; Runci, D.; Forte, S.; De Maria, R.; Ricci-Vitiani, L. Analysis of the combined action of mir-143 and mir-145 on oncogenic pathways in colorectal cancer cells reveals a coordinate program of gene repression. Oncogene 2013, 32, 4806–4813. [Google Scholar] [CrossRef] [PubMed]
- Akao, Y.; Nakagawa, Y.; Hirata, I.; Iio, A.; Itoh, T.; Kojima, K.; Nakashima, R.; Kitade, Y.; Naoe, T. Role of anti-oncomirs miR-143 and -145 in human colorectal tumors. Cancer Gene Ther. 2010, 17, 398–408. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Chen, L.; Xu, Y.; Li, R.; Du, X. MicroRNA-195 promotes apoptosis and suppresses tumorigenicity of human colorectal cancer cells. Biochem. Biophys. Res. Commun. 2010, 400, 236–240. [Google Scholar] [CrossRef] [PubMed]
- Paterson, E.L.; Kazenwadel, J.; Bert, A.G.; Khew-Goodall, Y.; Ruszkiewicz, A.; Goodall, G.J. Down-Regulation of the miRNA-200 Family at the Invasive Front of Colorectal Cancers with Degraded Basement Membrane Indicates EMT is Involved in Cancer Progression. Neoplasia 2013, 15, 180–191. [Google Scholar] [CrossRef]
- Wang, W.H.; Chen, J.; Zhao, F.; Zhang, B.R.; Yu, H.S.; Jin, H.Y.; Dai, J.H. Mir-150-5p Suppresses Colorectal Cancer Cell Migration and Invasion through Targeting MUC4. Asian Pac. J. Cancer Prev. 2014, 15, 6269–6273. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Yang, Y.; Zhang, P.; Wang, F.; Ma, Y.; Qin, H.; Wang, Y. miR-150 functions as a tumour suppressor in human colorectal cancer by targeting c-Myb. J. Cell Mol. Med. 2014, 18, 2125–2134. [Google Scholar] [CrossRef] [PubMed]
- Bitarte, N.; Bandres, E.; Boni, V.; Zarate, R.; Rodriguez, J.; Gonzalez-Huarriz, M.; Lopez, I.; Javier Sola, J.; Alonso, M.M.; Fortes, P.; et al. MicroRNA-451 is Involved in the Self-renewal, Tumorigenicity, and Chemoresistance of Colorectal Cancer Stem Cells. Stem Cells 2011, 29, 1661–1671. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Zhang, P.; Wang, F.; Zhang, H.; Yang, Y.; Shi, C.; Xia, Y.; Peng, J.; Liu, W.; Yang, Z.; et al. Elevated oncofoetal miR-17-5p expression regulates colorectal cancer progression by repressing its target gene P130. Nat. Commun. 2012, 3, 1291. [Google Scholar] [CrossRef] [PubMed]
- Tsuchida, A.; Ohno, S.; Wu, W.; Borjigin, N.; Fujita, K.; Aoki, T.; Ueda, S.; Takanashi, M.; Kuroda, M. Mir-92 is a key oncogenic component of the mir-17–92 cluster in colon cancer. Cancer Sci. 2011, 102, 2264–2271. [Google Scholar] [CrossRef] [PubMed]
- Jahid, S.; Sun, J.; Edwards, R.A.; Dizon, D.; Panarelli, N.C.; Milsom, J.W.; Sikandar, S.S.; Gümüş, Z.H.; Lipkin, S.M. miR-23a Promotes the Transition from Indolent to Invasive Colorectal Cancer. Cancer Discov. 2012, 2, 540–553. [Google Scholar] [CrossRef] [PubMed]
- Chintharlapalli, S.; Papineni, S.; Abdelrahim, M.; Abudayyeh, A.; Jutooru, I.; Chadalapaka, G.; Wu, F.; Mertens-Talcott, S.; Vanderlaag, K.; Cho, S.D.; et al. Oncogenic MicroRNA-27a is a Target For Anticancer Agent Methyl 2-Cyano-3,11-dioxo-18β-olean-1,12-dien-30-oate in Colon Cancer Cells. Int. J. Cancer 2009, 125, 1965–1974. [Google Scholar] [CrossRef] [PubMed]
- Valeri, N.; Braconi, C.; Gasparini, P.; Murgia, C.; Lampis, A.; Paulus-Hock, V.; Hart, J.R.; Ueno, L.; Grivennikov, S.I.; Lovat, F.; et al. Microrna-135b Promotes Cancer Progression by Acting as a Downstream Effector of Oncogenic Pathways in Colon Cancer. Cancer Cell 2014, 25, 469–483. [Google Scholar] [PubMed]
- Lu, M.H.; Huang, C.C.; Pan, M.R.; Chen, H.H.; Hung, W.C. Prospero Homeobox 1 Promotes Epithelial–Mesenchymal Transition in Colon Cancer Cells by Inhibiting E-cadherin via miR-9. Clin. Cancer Res. 2012, 18, 6416–6425. [Google Scholar] [CrossRef] [PubMed]
- Park, Y.R.; Lee, S.T.; Kim, S.L.; Liu, Y.C.; Lee, M.R.; Shin, J.H.; Seo, S.Y.; Kim, S.H.; Kim, I.H.; Lee, S.O.; et al. MicroRNA-9 suppresses cell migration and invasion through downregulation of TM4SF1 in colorectal cancer. Int. J. Oncol. 2016, 48, 2135–2143. [Google Scholar] [PubMed]
- Fassan, M.; Pizzi, M.; Giacomelli, L.; Mescoli, C.; Ludwig, K.; Pucciarelli, S.; Rugge, M. PDCD4 nuclear loss inversely correlates with miR-21 levels in colon carcinogenesis. Virchows Arch. 2011, 458, 413–419. [Google Scholar] [CrossRef] [PubMed]
- Xiong, B.; Cheng, Y.; Ma, L.; Zhang, C. Mir-21 Regulates Biological Behavior Through the PTEN/PI-3 K/Akt Signaling Pathway in Human Colorectal Cancer Cells. Int. J. Oncol. 2013, 42, 219–228. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Kanwar, S.S.; Patel, B.B.; Oh, P.S.; Nautiyal, J.; Sarkar, F.H.; Majumdar, A.P. MicroRNA-21 induces stemness by downregulating transforming growth factor beta receptor 2 (TGFβR2) in colon cancer cells. Carcinogenesis 2012, 33, 68–76. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.H.; Abernathy, B.; Kim, Y.J.; Lee, J.H.; Kim, J.H.; Shin, E.C.; Kim, J.K. Cruciferous vegetables and colorectal cancer prevention through microRNA regulation: A review. Crit. Rev. Food Sci. Nutr. 2017, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Chen, X.; Lin, J.; Lwin, T.; Wright, G.; Moscinski, L.C.; Dalton, W.S.; Seto, E.; Wright, K.; Sotomayor, E.; et al. Myc represses miR-15a/miR-16-1 expression through recruitment of HDAC3 in mantle cell and other non-Hodgkin B-cell lymphomas. Oncogene 2012, 31, 3002–3008. [Google Scholar] [CrossRef] [PubMed]
- Wiesen, J.L.; Tomasi, T.B. Dicer is regulated by cellular stresses and interferons. Mol. Immunol. 2009, 46, 1222–1228. [Google Scholar] [CrossRef] [PubMed]
- Cheng, X.; Ku, C.H.; Siow, R.C.M. Regulation of the Nrf2 antioxidant pathway by microRNAs: New players in micromanaging redox homeostasis. Free Radic. Biol. Med. 2013, 64, 4–11. [Google Scholar] [CrossRef] [PubMed]
- Leung, A.K.; Vyas, S.; Rood, J.E.; Bhutkar, A.; Sharp, P.A.; Chang, P. Poly(ADP-Ribose) Regulates Stress Responses and MicroRNA Activity in the Cytoplasm. Mol. Cell 2011, 42, 489–499. [Google Scholar] [CrossRef] [PubMed]
- Shah, N.M.; Rushworth, S.A.; Murray, M.Y.; Bowles, K.M.; MacEwan, D.J. Understanding the role of NRF2-regulated miRNAs in human malignancies. Oncotarget 2013, 4, 1130–1142. [Google Scholar] [CrossRef] [PubMed]
- Kurinna, S.; Werner, S. Nrf2 and microRNAs: new but awaited relations. Biochem. Soc. Trans. 2015, 43, 595–601. [Google Scholar] [CrossRef] [PubMed]
- Tili, E.; Michaille, J.J.; Wernicke, D.; Alder, H.; Costinean, S.; Volinia, S.; Croce, C.M. Mutator activity induced by microRNA-155 (miR-155) links inflammation and cancer. Proc. Natl. Acad. Sci. 2011, 108, 4908–4913. [Google Scholar] [CrossRef] [PubMed]
- Rahman, I.; Marwick, J.; Kirkham, P. Redox modulation of chromatin remodeling: impact on histone acetylation and deacetylation, NF-κB and pro-inflammatory gene expression. Biochem. Pharmacol. 2004, 68, 1255–1267. [Google Scholar] [CrossRef] [PubMed]
- Luxen, S.; Belinsky, S.A.; Knaus, U.G. Silencing of DUOX NADPH Oxidases by Promoter Hypermethylation in Lung Cancer. Cancer Res. 2008, 68, 1037–1045. [Google Scholar] [CrossRef] [PubMed]
- Bäckdahl, L.; Bushell, A.; Beck, S. Inflammatory signalling as mediator of epigenetic modulation in tissue-specific chronic inflammation. Int. J. Biochem. Cell Biol. 2009, 41, 176–184. [Google Scholar] [CrossRef] [PubMed]
- McBrian, M.A.; Behbahan, I.S.; Ferrari, R.; Su, T.; Huang, T.W.; Li, K. Global Histone Acetylation is Linked to pH. Cancer Discov. 2013, 3, 12. [Google Scholar]
- Leoni, F.; Fossati, G.; Lewis, E.C.; Lee, J.K.; Porro, G.; Pagani, P.; Modena, D.; Moras, M.L.; Pozzi, P.; Reznikov, L.L.; et al. The Histone Deacetylase Inhibitor ITF2357 Reduces Production of Pro-Inflammatory Cytokines In Vitro and Systemic Inflammation In Vivo. Mol. Med. 2005, 11, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Fearon, E.R.; Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 1990, 61, 759–767. [Google Scholar] [CrossRef]
- Slaby, O.; Svoboda, M.; Michalek, J.; Vyzula, R. MicroRNAs in colorectal cancer: Translation of molecular biology into clinical application. Mol. Cancer 2009, 8, 102. [Google Scholar] [CrossRef] [PubMed]
- Amirkhah, R.; Schmitz, U.; Linnebacher, M.; Wolkenhauer, O.; Farazmand, A. MicroRNA-mRNA interactions in colorectal cancer and their role in tumor progression. Genes Chromosomes Cancer 2015, 54, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Liu, Y.; Jin, X.; Lu, W.; Liu, J.; Xia, Z.; Yuan, Q.; Zhao, X.; Xu, N.; Liang, S. MicroRNA-26a regulates glucose metabolism by direct targeting PDHX in colorectal cancer cells. BMC Cancer 2014, 14, 443. [Google Scholar] [CrossRef] [PubMed]
- Vander Heiden, M.G.; Cantley, L.C.; Thompson, C.B. Understanding the Warburg Effect: The Metabolic Requirements of Cell Proliferation. Science 2009, 324, 1029–1033. [Google Scholar] [CrossRef] [PubMed]
- Thomson, J.M.; Parker, J.; Perou, C.M.; Hammond, S.M. A custom microarray platform for analysis of microRNA gene expression. Nat. Meth. 2004, 1, 47–53. [Google Scholar] [CrossRef] [PubMed]
- miRBase. miRBase blog. Available online: http://www.mirbase.org/blog/ (accessed on 8 September 2016).
- Sorefan, K.; Pais, H.; Hall, A.E.; Kozomara, A.; Griffiths-Jones, S.; Moulton, V.; Dalmay, T. Reducing ligation bias of small RNAs in libraries for next generation sequencing. Silence 2012, 3, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Billmeier, M.; Mohorianu, I.; Green, D.; Fraser, W.D.; Dalmay, T. An improved protocol for small RNA library construction using High Definition adapters. Methods Next Gener. Seq. 2015, 2. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Huntley, J.; Fierer, N.; Owens, S.M.; Betley, J.; Fraser, L.; Bauer, M.; et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 2012, 6, 1621–1624. [Google Scholar] [CrossRef] [PubMed]
| MicroRNA | Reported Role in Colorectal Cancer | Reported Target Genes |
|---|---|---|
| let-7a-5p | Tumour Suppressive | NIRF [57] |
| miR-34a-5p | E2F3 [58] | |
| miR-101-3p | PTGS2 [59] | |
| miR-126-3p | PIK3R2 [60] | |
| miR-143-5p | DNMT3A [61]; ERK5 [62] | |
| miR-195-5p | BCL2 [63] | |
| miR-200a-3p | CTNNB1 [64] | |
| miR-150-5p | MUC4 [65]; MYB [66] | |
| miR-451a | MIF [67] | |
| miR-17-5p | Oncogenic | P130 [68] |
| miR-92a-3p | BIM [69] | |
| miR-23a-3p | MTSS1 [70] | |
| miR-27a-3p | ZBTB10 [71] | |
| miR-135b-5p | TGFβR2, DAPK1, APC [72] | |
| miR-9-5p | Ambiguous | CDH [73]; TM4SF1 [74] |
| miR-21-5p | Pdcd4 [75]; CDC25A [76]; TGFβR2 [77] |
| Type of Method | Cost | Relative Time Required | MiRNA Detection |
|---|---|---|---|
| Array | Medium | Low | Limited to miRNAs known at the time of array development. |
| Library-Sequencing | High | High | Can identify and assay novel miRNAs, and retain data for re-mapping against updated miRNA databases. |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dacosta, C.; Bao, Y. The Role of MicroRNAs in the Chemopreventive Activity of Sulforaphane from Cruciferous Vegetables. Nutrients 2017, 9, 902. https://doi.org/10.3390/nu9080902
Dacosta C, Bao Y. The Role of MicroRNAs in the Chemopreventive Activity of Sulforaphane from Cruciferous Vegetables. Nutrients. 2017; 9(8):902. https://doi.org/10.3390/nu9080902
Chicago/Turabian StyleDacosta, Christopher, and Yongping Bao. 2017. "The Role of MicroRNAs in the Chemopreventive Activity of Sulforaphane from Cruciferous Vegetables" Nutrients 9, no. 8: 902. https://doi.org/10.3390/nu9080902
APA StyleDacosta, C., & Bao, Y. (2017). The Role of MicroRNAs in the Chemopreventive Activity of Sulforaphane from Cruciferous Vegetables. Nutrients, 9(8), 902. https://doi.org/10.3390/nu9080902