The Association between Cardiorespiratory Fitness and Gut Microbiota Composition in Premenopausal Women
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design and Subjects
2.2. Background Information
2.3. Anthropometrical and Body Composition Assessments
2.4. Fitness Test
2.5. Blood and Biochemicla Measurements
2.6. Fecal Samples
2.7. Statistical Analyses
3. Results
3.1. Descriptive Statistics
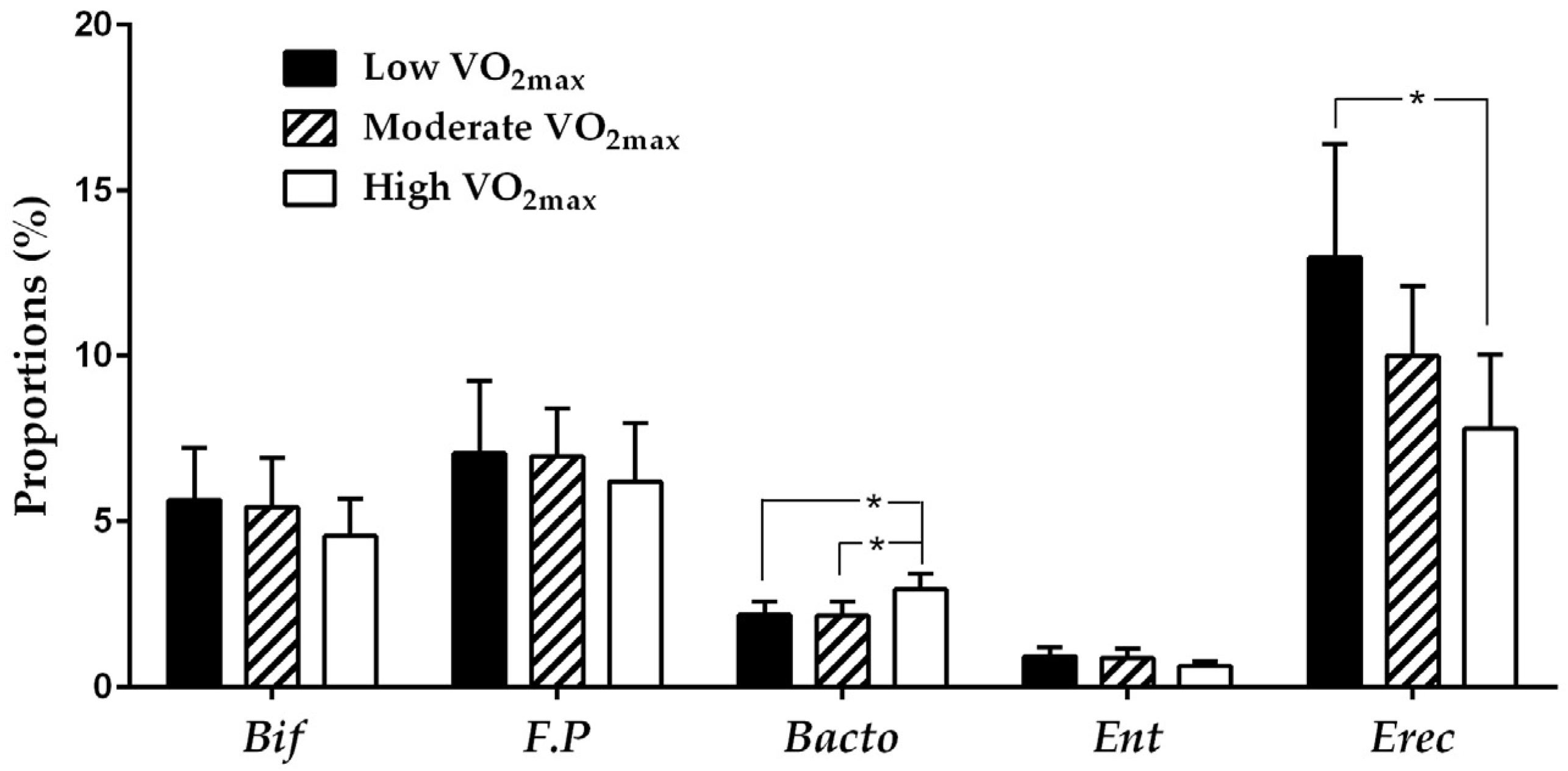
3.2. Comparison of Microbial Composition in Different Fitness Groups
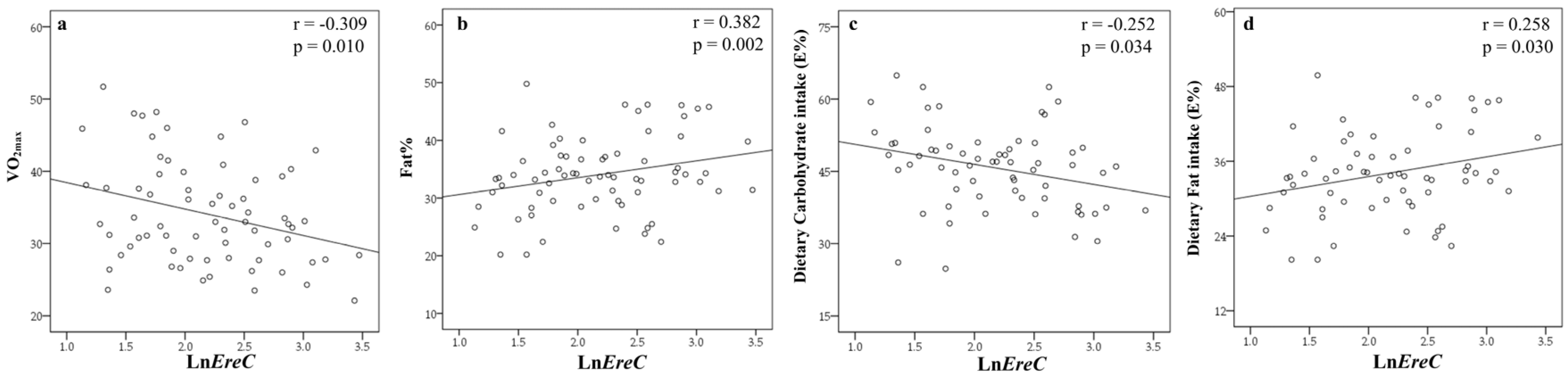
3.3. Associations of Certain Bacterial Groups with Different Variables
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lynch, S.V.; Pedersen, O. The human intestinal microbiome in health and disease. New Engl. J. Med. 2016, 375, 2369–2379. [Google Scholar] [CrossRef] [PubMed]
- Clemente, J.C.; Ursell, L.K.; Parfrey, L.W.; Knight, R. The impact of the gut microbiota on human health: An integrative view. Cell 2012, 148, 1258–1270. [Google Scholar] [CrossRef] [PubMed]
- Gerard, P. Gut microbiota and obesity. Cell Mol. Life Sci. 2016, 73, 147–162. [Google Scholar] [CrossRef] [PubMed]
- Larsen, N.; Vogensen, F.K.; van den Berg, F.W.; Nielsen, D.S.; Andreasen, A.S.; Pedersen, B.K.; Al-Soud, W.A.; Sorensen, S.J.; Hansen, L.H.; Jakobsen, M. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS ONE 2010, 5, e9085. [Google Scholar] [CrossRef] [PubMed]
- Kelly, T.N.; Bazzano, L.A.; Ajami, N.J.; He, H.; Zhao, J.; Petrosino, J.F.; Correa, A.; He, J. Gut microbiome associates with lifetime cardiovascular disease risk profile among bogalusa heart study participants. Circ. Res. 2016, 119, 956–964. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Wu, N.; Wang, X.; Chi, Y.; Zhang, Y.; Qiu, X.; Hu, Y.; Li, J.; Liu, Y. Dysbiosis gut microbiota associated with inflammation and impaired mucosal immune function in intestine of humans with non-alcoholic fatty liver disease. Sci. Rep. 2015, 5, 8096. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, M.; Inoue, R.; Tsukahara, T.; Ushida, K.; CHIJI, H.; Matsubara, N.; Hara, H. Voluntary running exercise alters microbiota composition and increases n-butyrate concentration in the rat cecum. Biosci. Biotechnol. Biochem. 2008, 72, 572–576. [Google Scholar] [CrossRef] [PubMed]
- Duncan, S.H.; Belenguer, A.; Holtrop, G.; Johnstone, A.M.; Flint, H.J.; Lobley, G.E. Reduced dietary intake of carbohydrates by obese subjects results in decreased concentrations of butyrate and butyrate-producing bacteria in feces. Appl. Environ. Microbiol. 2007, 73, 1073–1078. [Google Scholar] [CrossRef] [PubMed]
- Clarke, S.F.; Murphy, E.F.; O'Sullivan, O.; Lucey, A.J.; Humphreys, M.; Hogan, A.; Hayes, P.; O'Reilly, M.; Jeffery, I.B.; Wood-Martin, R.; et al. Exercise and associated dietary extremes impact on gut microbial diversity. Gut 2014, 63, 1913–1920. [Google Scholar] [CrossRef] [PubMed]
- Estaki, M.; Pither, J.; Baumeister, P.; Little, J.P.; Gill, S.K.; Ghosh, S.; Ahmadi-Vand, Z.; Marsden, K.R.; Gibson, D.L. Cardiorespiratory fitness as a predictor of intestinal microbial diversity and distinct metagenomic functions. Microbiome 2016, 4. [Google Scholar] [CrossRef] [PubMed]
- Baker, J.M.; Al-Nakkash, L.; Herbst-Kralovetz, M.M. Estrogen-gut microbiome axis: Physiological and clinical implications. Maturitas 2017, 103, 45–53. [Google Scholar] [CrossRef]
- Ortega, F.B.; Ruiz, J.R.; Castillo, M.J.; Sjostrom, M. Physical fitness in childhood and adolescence: A powerful marker of health. Int. J. Obes. (Lond) 2008, 32, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Shephard, R.J.; Allen, C.; Benade, A.J.; Davies, C.T.; Di Prampero, P.E.; Hedman, R.; Merriman, J.E.; Myhre, K.; Simmons, R. The maximum oxygen intake. An international reference standard of cardiorespiratory fitness. Bull. World Health Organ. 1968, 38, 757–764. [Google Scholar] [PubMed]
- Wiklund, P.K.; Pekkala, S.; Autio, R.; Munukka, E.; Xu, L.; Saltevo, J.; Cheng, S.; Kujala, U.M.; Alen, M.; Cheng, S. Serum metabolic profiles in overweight and obese women with and without metabolic syndrome. Diabetol. Metab. Syndr. 2014, 6, 40. [Google Scholar] [CrossRef] [PubMed]
- Tan, X.; Alen, M.; Cheng, S.M.; Mikkola, T.M.; Tenhunen, J.; Lyytikainen, A.; Wiklund, P.; Cong, F.; Saarinen, A.; Tarkka, I.; et al. Associations of disordered sleep with body fat distribution, physical activity and diet among overweight middle-aged men. J. Sleep Res. 2015, 24, 414–424. [Google Scholar] [CrossRef] [PubMed]
- Munukka, E.; Pekkala, S.; Wiklund, P.; Rasool, O.; Borra, R.; Kong, L.; Ojanen, X.; Cheng, S.M.; Roos, C.; Tuomela, S.; et al. Gut-adipose tissue axis in hepatic fat accumulation in humans. J. Hepatol. 2014, 61, 132–138. [Google Scholar] [CrossRef] [PubMed]
- Lyytikainen, A.; Lamberg-Allardt, C.; Kannas, L.; Cheng, S. Food consumption and nutrient intakes with a special focus on milk product consumption in early pubertal girls in Central Finland. Public Health Nutr. 2005, 8, 284–289. [Google Scholar] [CrossRef] [PubMed]
- Cheng, S.; Wiklund, P.; Autio, R.; Borra, R.; Ojanen, X.; Xu, L.; Tormakangas, T.; Alen, M. Adipose tissue dysfunction and altered systemic amino acid metabolism are associated with non-alcoholic fatty liver disease. PLoS ONE. 2015, 10, e138889. [Google Scholar] [CrossRef] [PubMed]
- Vaahtovuo, J.; Korkeamaki, M.; Munukka, E.; Viljanen, M.K.; Toivanen, P. Quantification of bacteria in human feces using 16S rRNA-hybridization, DNA-staining and flow cytometry. J. Microbiol. Methods 2005, 63, 276–286. [Google Scholar] [CrossRef] [PubMed]
- Vaahtovuo, J.; Munukka, E.; Korkeamaki, M.; Luukkainen, R.; Toivanen, P. Fecal microbiota in early rheumatoid arthritis. J. Rheumatol. 2008, 35, 1500–1505. [Google Scholar] [PubMed]
- Ley, R.E.; Backhed, F.; Turnbaugh, P.; Lozupone, C.A.; Knight, R.D.; Gordon, J.I. Obesity alters gut microbial ecology. Proc. Natl. Acad. Sci. USA. 2005, 102, 11070–11075. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Hamady, M.; Yatsunenko, T.; Cantarel, B.L.; Duncan, A.; Ley, R.E.; Sogin, M.L.; Jones, W.J.; Roe, B.A.; Affourtit, J.P.; et al. A core gut microbiome in obese and lean twins. Nature 2009, 457, 480–484. [Google Scholar] [CrossRef] [PubMed]
- Cho, I.; Yamanishi, S.; Cox, L.; Methe, B.A.; Zavadil, J.; Li, K.; Gao, Z.; Mahana, D.; Raju, K.; Teitler, I.; et al. Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature 2012, 488, 621–626. [Google Scholar] [CrossRef] [PubMed]
- Munukka, E.; Wiklund, P.; Pekkala, S.; Volgyi, E.; Xu, L.; Cheng, S.; Lyytikainen, A.; Marjomaki, V.; Alen, M.; Vaahtovuo, J.; et al. Women with and without metabolic disorder differ in their gut microbiota composition. Obesity 2012, 20, 1082–1087. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef] [PubMed]
- Guinane, C.M.; Cotter, P.D. Role of the gut microbiota in health and chronic gastrointestinal disease: understanding a hidden metabolic organ. Therap. Adv. Gastroenterol. 2013, 6, 295–308. [Google Scholar] [CrossRef] [PubMed]
- Shoaie, S.; Karlsson, F.; Mardinoglu, A.; Nookaew, I.; Bordel, S.; Nielsen, J. Understanding the interactions between bacteria in the human gut through metabolic modeling. Sci. Rep. 2013, 3, 2532. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Bonder, M.J.; Cenit, M.C.; Tigchelaar, E.F.; Maatman, A.; Dekens, J.A.; Brandsma, E.; Marczynska, J.; Imhann, F.; Weersma, R.K.; et al. The gut microbiome contributes to a substantial proportion of the variation in blood lipids. Circ. Res. 2015, 117, 817–824. [Google Scholar] [CrossRef] [PubMed]
- Duncan, S.H.; Lobley, G.E.; Holtrop, G.; Ince, J.; Johnstone, A.M.; Louis, P.; Flint, H.J. Human colonic microbiota associated with diet, obesity and weight loss. Int. J. Obes. 2008, 32, 1720–1724. [Google Scholar] [CrossRef] [PubMed]
- Simoes, C.D.; Maukonen, J.; Scott, K.P.; Virtanen, K.A.; Pietilainen, K.H.; Saarela, M. Impact of a very low-energy diet on the fecal microbiota of obese individuals. Eur. J. Nutr. 2014, 53, 1421–1429. [Google Scholar] [CrossRef] [PubMed]
- Ley, R.E.; Turnbaugh, P.J.; Klein, S.; Gordon, J.I. Microbial ecology: Human gut microbes associated with obesity. Nature 2006, 444, 1022–1023. [Google Scholar] [CrossRef] [PubMed]
- Donohoe, D.R.; Garge, N.; Zhang, X.; Sun, W.; O'Connell, T.M.; Bunger, M.K.; Bultman, S.J. The microbiome and butyrate regulate energy metabolism and autophagy in the mammalian colon. Cell Metab. 2011, 13, 517–526. [Google Scholar] [CrossRef] [PubMed]
- Schwiertz, A.; Taras, D.; Schafer, K.; Beijer, S.; Bos, N.A.; Donus, C.; Hardt, P.D. Microbiota and SCFA in lean and overweight healthy subjects. Obesity. 2010, 18, 190–195. [Google Scholar] [CrossRef] [PubMed]
- De Schryver, A.M.; Keulemans, Y.C.; Peters, H.P.; Akkermans, L.M.; Smout, A.J.; De Vries, W.R.; van Berge-Henegouwen, G.P. Effects of regular physical activity on defecation pattern in middle-aged patients complaining of chronic constipation. Scand. J. Gastroenterol. 2005, 40, 422–429. [Google Scholar] [CrossRef] [PubMed]
- Peters, H.P.; De Vries, W.R.; Vanberge-Henegouwen, G.P.; Akkermans, L.M. Potential benefits and hazards of physical activity and exercise on the gastrointestinal tract. Gut 2001, 48, 435–439. [Google Scholar] [CrossRef] [PubMed]
- Liou, A.P.; Paziuk, M.; Luevano, J.J.; Machineni, S.; Turnbaugh, P.J.; Kaplan, L.M. Conserved shifts in the gut microbiota due to gastric bypass reduce host weight and adiposity. Sci. Transl. Med. 2013, 5, 141r–178r. [Google Scholar] [CrossRef] [PubMed]
- Gisolfi, C.V. Is the GI System Built For Exercise? News Physiol. Sci. 2000, 15, 114–119. [Google Scholar] [PubMed]
- Rosa, E.F.; Silva, A.C.; Ihara, S.S.; Mora, O.A.; Aboulafia, J.; Nouailhetas, V.L. Habitual exercise program protects murine intestinal, skeletal, and cardiac muscles against aging. J. Appl. Physiol. 2005, 99, 1569–1575. [Google Scholar] [CrossRef] [PubMed]
- Le Chatelier, E.; Nielsen, T.; Qin, J.; Prifti, E.; Hildebrand, F.; Falony, G.; Almeida, M.; Arumugam, M.; Batto, J.; Kennedy, S.; et al. Richness of human gut microbiome correlates with metabolic markers. Nature 2013, 500, 541–546. [Google Scholar] [CrossRef] [PubMed]
- Caspi, R.; Altman, T.; Billington, R.; Dreher, K.; Foerster, H.; Fulcher, C.A.; Holland, T.A.; Keseler, I.M.; Kothari, A.; Kubo, A.; et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases. Nucleic Acids Res. 2014, 42, D459–D471. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, F.H.; Tremaroli, V.; Nookaew, I.; Bergstrom, G.; Behre, C.J.; Fagerberg, B.; Nielsen, J.; Backhed, F. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 2013, 498, 99–103. [Google Scholar] [CrossRef] [PubMed]
- Amann, R.I.; Krumholz, L.; Stahl, D.A. Fluorescent-oligonucleotide probing of whole cells for determinative, phylogenetic, and environmental studies in microbiology. J. Bacteriol. 1990, 172, 762–770. [Google Scholar] [CrossRef] [PubMed]
- Rigottier-Gois, L.; Bourhis, A.G.; Gramet, G.; Rochet, V.; Dore, J. Fluorescent hybridisation combined with flow cytometry and hybridisation of total RNA to analyse the composition of microbial communities in human faeces using 16S rRNA probes. Fems Microbiol. Ecol. 2003, 43, 237–245. [Google Scholar] [CrossRef] [PubMed]
- Rigottier-Gois, L.; Rochet, V.; Garrec, N.; Suau, A.; Dore, J. Enumeration of Bacteroides species in human faeces by fluorescent in situ hybridisation combined with flow cytometry using 16S rRNA probes. Syst. Appl. Microbiol. 2003, 26, 110–118. [Google Scholar] [CrossRef] [PubMed]
- Amann, R.I.; Binder, B.J.; Olson, R.J.; Chisholm, S.W.; Devereux, R.; Stahl, D.A. Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl. Environ. Microbiol. 1990, 56, 1919–1925. [Google Scholar] [PubMed]
- Amann, R.I.; Ludwig, W.; Schleifer, K.H. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Rev. 1995, 59, 143–169. [Google Scholar] [PubMed]
- Fraher, M.H.; O'Toole, P.W.; Quigley, E.M. Techniques used to characterize the gut microbiota: A guide for the clinician. Nat. Rev. Gastroenterol. Hepatol. 2012, 9, 312–322. [Google Scholar] [CrossRef] [PubMed]
| Variables | Low VO2max Mean 95% CI | Moderate VO2max Mean 95% CI | High VO2max Mean 95% CI | ANOVA (p-Value) | ||
|---|---|---|---|---|---|---|
| Number | 24 | 23 | 24 | L-M | L-H | M-H |
| Age (years) | 40.4 (36.9–44.0) | 39.7 (35.5–43.8) | 30.6 (25.6–35.6) | 0.991 | 0.003 | 0.009 |
| Height (cm) | 166.1 (164.0–168.3) | 166.7 (163.9–168.9) | 164.0 (161.7–166.3) | 0.984 | 0.490 | 0.307 |
| Weight (kg) | 87.3 (83.2–91.5) | 77.6 (73.0–82.1) | 66.1 (62.1–70.1) | 0.004 | <0.001 | 0.001 |
| BMI | 31.7 (30.2–33.1) | 27.9 (26.7–29.1) | 24.6 (23.0–26.2) | 0.001 | <0.001 | 0.004 |
| Fat% | 40.6 (38.1–43.0) | 35.5 (33.2–37.8) | 28.0 (25.0–31.0) | 0.018 | <0.001 | <0.001 |
| PA (time/week) | 1.39 (0.95–1.83) | 1.80 (1.22–2.38) | 3.44 (2.61–4.27) | 0.718 | <0.001 | 0.001 |
| PA (h/week) | 1.17 (0.69–1.64) | 2.11 (1.27–2.95) | 3.69 (2.82–4.56) | 0.195 | <0.001 | 0.009 |
| Fiber (g) | 20.4 (17.4–23.5) | 24.1 (20.6–27.6) | 21.0 (17.3–24.6) | 0.323 | 0.995 | 0.450 |
| Carbohydrate (E%) | 47.1 (43.7–50.5) | 45.6 (41.7–49.5) | 44.7 (41.5–48.0) | 0.901 | 0.698 | 0.979 |
| Protein (E%) | 18.4 (17.0–19.8) | 18.0 (17.0–19.0) | 18.0 (16.9–19.1) | 0.937 | 0.957 | 1.000 |
| Fat (E%) | 32.8 (30.3–35.4) | 34.1 (30.9–37.2) | 35.2 (32.5–38.0) | 0.887 | 0.508 | 0.910 |
| Alcohol (E%) | 1.73 (−54–3.99) | 2.38 (0.45–4.31) | 0.63 (−37–1.62) | 0.958 | 0.955 | 0.769 |
| TG (mmol/L) | 1.56 (1.17–1.95) | 1.22 (1.02–1.42) | 1.01 (0.82–1.20) | 0.608 | 0.041 | 0.434 |
| Cholestol (mmol/L) | 5.06 (4.55–5.56) | 5.03 (4.68–5.38) | 4.44 (4.14–4.74) | 0.999 | 0.072 | 0.098 |
| HDL (mmol/L) | 1.42 (1.29–1.55) | 1.59 (1.47–1.72) | 1.57 (1.42–1.73) | 0.198 | 0.279 | 0.996 |
| Leptin (ng/mL) | 56.8 (45.8–67.8) | 41.7 (34.5–48.8) | 26.9 (18.2–35.6) | 0.054 | <0.001 | 0.062 |
| EreC | Erec Adjusted for Age and Energy Yield Macronutrients Intakes of Carbohydrate, Protein, Fat and Alcohol | EreC Adjusted for Age, Fat% and Energy Yield Macronutrients Intakes of Carbohydrate, Protein, Fat and Alcohol | ||||
|---|---|---|---|---|---|---|
| r | p | r | p | r | p | |
| VO2max | −0.309 | 0.010 | −0.339 | 0.006 | −0.113 | 0.369 |
| Fat% | 0.382 | 0.002 | 0.364 | 0.005 | ||
| Leptin | 0.424 | 0.001 | 0.390 | 0.003 | 0.178 | 0.208 |
| HDL | −0.260 | 0.028 | −0.313 | 0.010 | −0.233 | 0.124 |
| TG | 0.390 | 0.002 | 0.393 | 0.003 | 0.300 | 0.060 |
| Y | VO2max | Leptin | HDL | TG | |
|---|---|---|---|---|---|
| X | |||||
| EreC | −0.0711 | 0.1206 | −0.2494 | 0.3277 | |
| Age | −0.3369 | −0.0917 | 0.1579 | 0.0618 | |
| Fat% | −0.6163 | 0.7887 | −0.2130 | 0.3008 | |
| Carbohydrates (E%) | −0.2886 | −0.2693 | 0.2631 | −0.1922 | |
| Protein (E%) | −0.0680 | −0.1145 | −0.0252 | −0.0104 | |
| Fat (E%) | −0.0520 | −0.1298 | 0.3869 | −0.1923 | |
| Alcohol (E%) | −0.1851 | −0.1049 | 0.1990 | −0.3277 | |
| Y | VO2max | Fat% | Leptin | HDL | TG | |
|---|---|---|---|---|---|---|
| X | ||||||
| Bif | 0.0348 | 0.1092 | 0.0892 | −0.0679 | 0.1539 | |
| Bacto | 0.1536 | −0.1479 | −0.0939 | 0.2540 | −0.1184 | |
| F.p | 0.0113 | 0.0929 | −0.0080 | 0.1721 | −0.1377 | |
| Ent | −0.0723 | −0.0129 | −0.0016 | 0.1995 | −0.0139 | |
| EreC | −0.2568 | 0.3276 | 0.3942 | −0.3129 | 0.3894 | |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Y.; Shi, Y.; Wiklund, P.; Tan, X.; Wu, N.; Zhang, X.; Tikkanen, O.; Zhang, C.; Munukka, E.; Cheng, S. The Association between Cardiorespiratory Fitness and Gut Microbiota Composition in Premenopausal Women. Nutrients 2017, 9, 792. https://doi.org/10.3390/nu9080792
Yang Y, Shi Y, Wiklund P, Tan X, Wu N, Zhang X, Tikkanen O, Zhang C, Munukka E, Cheng S. The Association between Cardiorespiratory Fitness and Gut Microbiota Composition in Premenopausal Women. Nutrients. 2017; 9(8):792. https://doi.org/10.3390/nu9080792
Chicago/Turabian StyleYang, Yifan, Yi Shi, Petri Wiklund, Xiao Tan, Na Wu, Xiaobo Zhang, Olli Tikkanen, Chenhong Zhang, Eveliina Munukka, and Sulin Cheng. 2017. "The Association between Cardiorespiratory Fitness and Gut Microbiota Composition in Premenopausal Women" Nutrients 9, no. 8: 792. https://doi.org/10.3390/nu9080792
APA StyleYang, Y., Shi, Y., Wiklund, P., Tan, X., Wu, N., Zhang, X., Tikkanen, O., Zhang, C., Munukka, E., & Cheng, S. (2017). The Association between Cardiorespiratory Fitness and Gut Microbiota Composition in Premenopausal Women. Nutrients, 9(8), 792. https://doi.org/10.3390/nu9080792





