Plant-Derived Epi-Nutraceuticals as Potential Broad-Spectrum Anti-Viral Agents
Abstract
1. Introduction
1.1. The Viral Life Cycle
1.2. Host Signaling and Virus Life Cycles
1.3. Epigenetic Modifications
1.3.1. DNA Methylation
1.3.2. Histone Modifications
1.3.3. RNA Modifications
1.3.4. miRNA
1.4. Viral Infections and Epigenetic Modifications
1.5. Virus Infection Promotes Epigenetic Modifications in the Immune System and Host Genes
1.6. Viral Infection Affects Host miRNA Expression
2. Plant-Derived Anti-Viral Activity
| Plant | Plant Part | Main Bioactive Constituents | Virus | IC50 a | SI b | Mode of Action | References |
|---|---|---|---|---|---|---|---|
| Aloysia citriodora Palau (Verbenaceae) | nr | geranial (18.9%), neral (15.6%), limonene (10.7%), 1,8-cineole (5.0%), spathulenol (4.7%), geraniol (2.7%) | Yellow fever virus | 19.4 μg/mL | 2.6 | nr c | [140] |
| Artemisia kermanensis Podlech (Compositae) | aerial parts | α-thujone (13.8%), camphore (10.2%), β-thujone (6.2%), p-Mentha-1,5-dien-8-ol (4.4%) | HSV-1 | 0.004% | 66.4 | nr | [128] |
| Ayapana triplinervis (Vahl) R.M.King & H.Rob. (Compositae) | aerial parts | thymohydroquinone dimethyl ether (87.1%), α-phellandrene (2.0%), β-selinene (1.9%) | ZIKV | 38 μg/mL | 12.5 | inhibitor of internalization process | [141] |
| Cananga odorata (Lam.) Hook.f. & Thomson (Annonaceae) | nr | benzyl salicylate (49.3%), benzyl benzoate (18.7%), linalool (16.6%), α-gurjunene (7.1%) | HIV-1 | 0.60 µg/mL | nr | nr | [129] |
| Cinnamomum zeylanicum Blume (Lauraceae) | leaves | eugenol (nr) | H1N1 | <3.1 µL/mL | >4 | intercellular | [142] |
| Citrus × bergamia Risso & Poit. (Rutaceae) | fruit peel | (–)-linalyl acetate (nr), (–)-linalool (nr), (+)-limonene (nr), γ-terpinene (nr), β-pinene (nr), α-pinene (nr), α-terpinene (nr) | H1N1 | <3.1 µL/mL | >5 | intercellular | [142] |
| Cymbopogon citratus (DC.) Stapf (Poaceae) | nr | cis-citral (59.2%), β-pinene (22.5%), cis-verbenol (6.1%), nerol (5.0%) | HIV-1 | 0.61 µg/mL | 1.1 | nr | [129] |
| Cymbopogon flexuosus (Nees ex Steud.) W.Watson (Poaceae) | grass | geranial (nr), neral (nr) | H1N1 | <3.1 µL/mL | >4 | nr | [142] |
| Cymbopogon nardus (L.) Rendle (Poaceae) | nr | nr | HIV-1 | 1.2 mg/mL | nr | nr | [130] |
| Dysphania ambrosioides (L.) Mosyakin & Clemants (Amaranthaceae) | aerial parts | cis-ascaridole (60.3%), m-cymene (22.2%), α-terpinene (1.8%), thymol (1.1%) | Coxsackie virus B4 | 21.7 μg/mL | 74.3 | nr | [143] |
| Eucalyptus caesia Benth. (Myrtaceae) | aerial parts | 1,8-cineole (40.2%), p-cymene (14.1%), γ-terpinene (12.4%), α-pinene (7.7%), terpinen-4-ol (5.6%) | HSV-1 | 0.007% | 38.8 | nr | [128] |
| Eucalyptus globulus Labill. (Myrtaceae) | leaves | 1,8-cineole (nr), α-pinene (nr) | H1N1 | <50 µL/mL | >0.5 | intercellular | [129] |
| 1,8-cineole (68.0%), globulol (5.4%), trans-pinocarveol (4.6%), α-pinene (3.7%) | Coxsackie virus B3 | 0.7 mg/mL | 22.8 | intercellular | [144] | ||
| Fortunella margarita Lour. Swingle (Rutaceae) | fruits | terpineol (55.5%), τ-carveol (5.5%), limonene (1.7%), muurolene (5.5%), cadinene (2.0%) | H5N1 | 6.8 µg/mL | nr | nr | [145] |
| Illicium verum Hooker f. (Schisandraceae) | fruits | (E)-anethole (80.0%) | HSV-1 | 1 µg/mL | 160 | intercellular | [127] |
| Lallemantia royleana (Benth.) Benth. (Lamiaceae) | aerial parts | (E)-pinocarvyl acetate (26.0%), pinocarvone (20.0%), verbenone (7.1%), (E)-β-ocimene (4.1%), (E)-carveol (5.3%), 3-thujen-2-one (5.1%), pulegone (4.4%), (Z)-carveol (3.5%), linalool (3.4%) | HSV-1 | 0.011% | 6.4 | intercellular | [146] |
| Lavandula officinalis Chaix (Lamiaceae) | flowers | linalyl acetate (nr), linalool (nr) | H1N1 | <3.1 µL/mL | >8 | intercellular | [129] |
| Lippia alba (Mill.) N.E.Br. ex Britton & P.Wilson (Verbenaceae) | nr | carvone (39.7%), limonene (30.6%), bicyclosesquiphellandrene (8.9%), piperitenone (4.5%), piperitone (2.8%), β-bourbonene (1.7%) | Yellow fever virus | 4.3 μg/mL | 30.6 | intercellular and intracellular | [140] |
| Lippia graveolens Kunth (Verbenaceae) | nr | carvacrol (56.8%), o-cymene (32.2%), γ-terpinene (3.7%) | HSV-1 | 99.6 µg/mL | 7.4 | intercellular | [147] |
| ACVR-HHV-1 | 55.9 µg/mL | 13.1 | intercellular | ||||
| Bovine viral diarrhoea virus | 78 μg/mL | 7.2 | intracellular | ||||
| Respiratory syncytial virus | 68 μg/mL | 10.8 | intercellular | ||||
| Bovine herpes virus 2 | 58.4 μg/mL | 9.7 | intercellular and intracellular | ||||
| Melaleuca alternifolia (Maiden & Betche) Cheel (Myrtaceae) | leaves | nr | HSV-1 | 13.2 µg/mL | 43 | intracellular | [148] |
| Mentha suaveolens Ehrh. (Lamiaceae) | leaves | piperitenone oxide (86.8%), α-cubebene (2.1%), pulegone (1.4%), limonene (1.4%), caryophyllene (1.3%) | HSV-1 | 5.1 µg/mL | 67 | intracellular | [149] |
| Osmunda regalis L. (Osmundaceae) | aerial parts | hexahydrofarnesyl acetone (11.8%), 2,4-di-t-butylphenol (6.8%), phytol (6.5%), neophytadiene (4.6%), 1-octadecene (4.4%), 1-eicosene (4.4%), 1-hexadecene (4.1%) | Coxsackie virus B4 | 2.2 μg/mL | 789.8 | nr | [150] |
| Pelargonium graveolens L’Hér. (Geraniaceae) | flowering aerial parts | citronellol (nr), geraniol (nr) | H1N1 | <3.1 µL/mL | >21 | intercellular | [129] |
| Pulicaria vulgaris Gaertn. (Compositae) | aerial parts | thymol (50.2%), p-menth-6-en-2-one (carvotanacetone, 20.2%), thymol isobutyrate (16.9%), menthan-2-one (4.3%) | HSV-1 | 0.001% | 1 | intercellular | [146] |
| Rosmarinus officinalis L. (Lamiaceae) | aerial parts | α-pinene (23.9%), verbenon (15.4%), camphor (11.0%), p-cymene (7.5%), 3-octanone (5.6%) | HSV-1 | 0.006% | 46.1 | nr | [128] |
| eucaliptol (50.6%), camphor (13.3%), α-pinene (10.1%), β-pinene (7.7%), camphene (4.6%) | HIV-1 | 0.18 µg/mL | 3.6 | nr | [129] | ||
| Salvia desoleana Atzei & V.Picci (Lamiaceae) | aerial parts | linalyl acetate (30.2%), germacrene D (18.7%), α-terpinyl acetate (16.8%), 1,8-cineole (10.2%), linalool (5.1%) | Acyclovir-resistant HSV-2 | 28.6 µg/mL | 55.2 | intercellular and intracellular | [151] |
| Satureja hortensis L. (Lamiaceae) | aerial parts | carvacrol (32.4%), γ-terpinene (32.0%), thymol (10.0%), p-cymene (6.6%), α-pinene (4.3%) | HSV-1 | 0.008% | 32.2 | nr | [128] |
| Sinapis arvensis L. (Brassicaceae) | flowers | cubenol (14.3%), 2-phyenyl isothiocyanate (7.5), dimethyl trisulfide (5.2%), thymol (4.6%), δ-cadinene (3.4%) | HSV-1 | 0.035% | 1.5 | intercellular | [146] |
| Thymbra capitata (L.) Cav. (Lamiaceae) | aerial parts | cinnamaldehyde (nr), carvacrol (nr) | HSV-1 | 17.6 µg/mL | 6.0 | intercellular | [152] |
| HSV-2 | 18.6 µg/mL | 6.9 | intercellular | ||||
| Thymus vulgaris L. (Lamiaceae) | aerial parts | 1,8-cineole (nr), terpenyl acetate (nr), borneol (nr) | H1N1 | <3.1 µL/mL | >4 | intercellular | [142] |
| thymol (37.8%), iso-thymol (23.2%), γ-terpinene (13.2%), β-caryophyllene (4.2%), linalool (3.3%) | HIV-1 | 1.30 | 1.6 | nr | [129] | ||
| Zataria multiflora Boiss. (Lamiaceae) | aerial parts | thymol (33.1%), carvacrol (25.9%), p-cymene (11.3%), α-pinene (3.9%) | HSV-1 | 0.003% | 55.4 | nr | [128] |
| Plant | Plant Part | Extract | Main Bioactive Constituents | Virus | IC50 a | SI b | Mode of Action | References |
|---|---|---|---|---|---|---|---|---|
| Agrimonia pilosa Ledeb. (Rosaceae) | whole plant | ethanol extract | nr c | H1N1, HV A-B | 0.5–1 μg/mL | nr | block uncoating process | [153] |
| Aloe vera (L.) Burm.f. (Xanthorrhoeaceae) | leaf | Gel | nr | HSV-1 | 5% | nr | replication inhibitor | [154] |
| Arachis hypogaea L. (Leguminosae) | peanut skin | ethanol extract | nr | H1N1 | 1.3 μg/mL | 5.2 | early stages of infection inhib. | [155] |
| Avicennia marina (Forssk.) Vierh. (Acanthaceae) | leaf | methanol extract | nr | HSV-1 | 9 μg/mL | 9.1 | viral replication inhib. | [156] |
| HIV | 15 μg/mL | 5.2 | interference with replication cycle | |||||
| Centella asiatica (L.) Urb. (Apiaceae) | leaf | water extract | nr | HIV | 36 μg/mL | nr | immunomodulatory effect | [157] |
| alcoholic extract | HIV | 8 μg/mL | ||||||
| Combretum adenogonium Steud. ex A.Rich. (Combretaceae) | root | water/ethanol extract | nr | HIV-1 | 24.7 μg/mL | nr | protease inhibitor | [158] |
| Copaifera reticulate Ducke (Leguminosae) | stem, bark and leaf | water/ethanol extract | phenolics, alcohols, organic acids | HSV-2 | 50 μg/mL | nr | block virus attachment | [159] |
| Cornus canadensis L. (Cornaceae) | leaf | water/ethanol extract | tellimagrandin I and other hydrolysable tannins | HSV-1 | 9 μg/mL | nr | virus absorption inhibitor | [160] |
| Embelia ribes Burm.f. (Primulaceae) | fruit | ethylacetate extract | embelin | H1N1 | 0.2 μg/mL | 32 | block virus entry | [161] |
| Epimedium koreanum Nakai (Berberidaceae) | bark | water extract | nr | HSV | 0.20 μg/mL | 23.5 | immunomodulatory effect | [162] |
| Equisetum giganteum L. (Equisetaceae) | root and stem | water/ethanol extract | phenolics, alcohols, organic acids | HSV-2 | 18 μg/mL | nr | block virus attachment | [159] |
| Eupatorium perfoliatum L. (Compositae) | aerial part | hydroalcoholic extract | nr | H1N1 | 7 μg/mL | 52 | viral attachment inhibitor | [163] |
| Euphorbia hirta L. (Euphorbiaceae) | aerial part | methanol extract | nr | HIV-1 | 38 μg/mL | nr | RT inhib. | [164] |
| HIV-2 | 22 μg/mL | |||||||
| Ficus religiosa L. (Moraceae) | bark | methanol extract | nr | HSV-2 | 5.2 μg/mL | 31.1 | nr | [165] |
| Hemidesmus indicus L. Br. ex Schult. (Apocynaceae) | root | water/methanol extract | 2-hydroxy-4-methoxybenzaldehyde (0.41 mg/g), 3-hydroxy-4- methoxybenzaldehyde (0.16 mg/g) | HSV-1 | 66.8 μg/mL | nr | anti-ER α-glucosidase inhib. | [166] |
| 2-hydroxy-4-methoxybenzaldehyde (0.41 mg/g), 3-hydroxy-4- methoxybenzaldehyde (0.16 mg/g) | HSV-2 | 70.6 μg/mL | ||||||
| Jatropha multifida L. (Euphorbiaceae) | stem | water extract | nr | H1N1 | 25 μg/mL | nr | block virus entry | [167] |
| Paeonia lactiflora Pall. (Paeoniaceae) | root | ethanol extract | nr | H1N1 | 0.016 mg/mL | 13.5 | block several stages of infection | [168] |
| Pedilanthus tithymaloides (L.) Poit. (Euphorbiaceae) | leaf | methanol extract | 2-(3,4-dihydroxy-phenyl)-5,7-dihydroxy-chromen-4-one or luteolin | HSV-2 | 48.5 μg/mL | 9.0 | NF-κB signalling pathway modulation | [169] |
| Prunella vulgaris L. (Lamiaceae) | flowers | water extract | nr | HIV | 0.8 μg/mL | nr | early post-virus binding interference | [170] |
| Prunus dulcis (Mill.) DA Webb (Rosaceae) | almond skin | methanol/HCl extract | nr | HSV-1 | 0.04 mg/mL | nr | block virus entry | [171] |
| Quercus brantii Lindl. (Fagaceae) | fruit | chloroform extract | nr | HSV-1 | 2.9 μg/mL | nr | block virus entry | [172] |
| Quercus persica Jaub. & Spach (Fagaceae) | fruit | water/ethanol extract | nr | HSV-1 | 0.26 μg/mL | nr | attachment inhib. | [162] |
| Rhus aromatica Aiton (Anacardiaceae) | root/stem bark | water extract | gallic acid | HSV-1 | 0.0005% | nr | nr | [173] |
| Rhus aromatica Aiton (Anacardiaceae) | root/stem bark | water extract | gallic acid | HSV-2 | 0.0043% | nr | nr | [173] |
| Schinus terebinthifolia Raddi (Anacardiaceae) | bark | water/ethanol extract | condensed tannins (catechin, 5.4 mg/L) | HSV-1 | 0.21 μg/mL | <49.0 | virucidal effect | [174] |
| Solanum melongena L. (Solenaceae) | peel | ethanol/HCl extract | delphinidin-3-rutinoside (90.3–115.0 μg/mg), chlorogenic acid (24.5–60.7 μg/mg) | HSV-1 | 83.4 μg/mL | nr | reduction of viral protein expression | [175] |
| Strychnos pseudoquina A. St.-Hil. (Loganiaceae) | stem bark | ethyl acetate extract | nr | HSV-1 | 5.29 μg/mL | nr | interference with various step of virus cycle | [176] |
| Strychnos pseudoquina A. St.-Hil. (Loganiaceae) | stem bark | ethyl acetate extract | nr | HSV-2 | 6.55 μg/mL | nr | interference with various step of virus cycle | [176] |
| Tanacetum parthenium (L.) Sch.Bip. (Compositae) | aerial parts | water/ethanol extract | chlorogenic acid, flavonoids (aglycones and glycosylated flavonoids), parthenolide | HSV-1 | 3.1 μg/mL | nr | viral replication inhib. | [177] |
| Taxodium distichum (L.) Rich. (Cupressaceae) | cone, leaf, stem | water extract | nr | H1N1 | 0.05 mg/mL | 5.6 | block virus entry | [178] |
| Vachellia nilotica (L.) P.J.H. Hurter & Mabb. (Leguminosae) | bark | methanol extract | nr | HSV-2 | 4.71 μg/mL | 30.6 | block virus attachment | [179] |
| Vachellia nilotica (L.) P.J.H. Hurter & Mabb. (Leguminosae) | bark | methanol extract | nr | HPV-16 | 1.80 μg/mL | 32.6 | block virus attachment | [179] |
| Vachellia nilotica (L.) P.J.H. Hurter & Mabb. (Leguminosae) | bark | methanol extract | nr | HSV-2 acyclovir resistant | 6.71 µg/mL | 21.5 | block virus attachment | [179] |
| Vigna radiata (L.) R.Wilczek (Leguminosae) | sprout | methanol/HCl extract | nr | HSV-1 | 7.62 μg/mL | nr | virucidal effect | [180] |
2.1. Andrographolide
2.2. Apigenin
2.3. Baicalein
2.4. Berberine
2.5. Betulinic Acid
2.6. Butyric Acid
2.7. Cardamonin
2.8. Cordycepin
2.9. Corosolic Acid
2.10. Curcumin
2.11. Ellagic Acid
2.12. Epigallocatechin Gallate
2.13. Galangin
2.14. Garcinol
2.15. Genistein
2.16. Ginkgolic Acid
2.17. Glycyrrhizic Acid
2.18. Grifolin
2.19. Oleacein
2.20. Organosulfur Chemicals
2.21. Orobol 7-O-d-Glucoside
2.22. Orsaponin
2.23. Plitidepsin
2.24. Pterostilbene
2.25. Quercetin
2.26. Raoulic Acid
2.27. Resveratrol
2.28. Silibinin
2.29. Silvestrol
2.30. Sulforaphane
2.31. Tanshinone IIA
2.32. Ursolic Acid
2.33. Withaferin A
3. Conclusions and Future Directions
Author Contributions
Funding
Conflicts of Interest
References
- Forterre, P. Defining life: The virus viewpoint. Orig. Life Evol. Biosph. 2010, 40, 151–160. [Google Scholar] [CrossRef]
- Caspar, D.L.; Klug, A. Physical principles in the construction of regular viruses. Cold Spring Harb. Symp. Quant. Biol. 1962, 27, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Wilen, C.B.; Tilton, J.C.; Doms, R.W. HIV: Cell binding and entry. Cold Spring Harb. Perspect. Med. 2012, 2, a006866. [Google Scholar] [CrossRef]
- Zlotnick, A. Theoretical aspects of virus capsid assembly. J. Mol. Recognit. 2005, 18, 479–490. [Google Scholar] [CrossRef]
- Palmer, K.J.; Tichelaar, W.; Myers, N.; Burns, N.R.; Butcher, S.J.; Kingsman, A.J.; Fuller, S.D.; Saibil, H.R. Cryo-electron microscopy structure of yeast Ty retrotransposon virus-like particles. J. Virol. 1997, 71, 6863–6868. [Google Scholar] [CrossRef]
- Wilson, D.P. Protruding Features of Viral Capsids Are Clustered on Icosahedral Great Circles. PLoS ONE 2016, 11, e0152319. [Google Scholar] [CrossRef] [PubMed]
- Scheiffele, P.; Rietveld, A.; Wilk, T.; Simons, K. Influenza viruses select ordered lipid domains during budding from the plasma membrane. J. Biol. Chem. 1999, 274, 2038–2044. [Google Scholar] [CrossRef]
- Louis, J.M.; Weber, I.T.; Tozser, J.; Clore, G.M.; Gronenborn, A.M. HIV-1 protease: Maturation, enzyme specificity, and drug resistance. Adv. Pharmacol. 2000, 49, 111–146. [Google Scholar] [CrossRef] [PubMed]
- Subissi, L.; Posthuma, C.C.; Collet, A.; Zevenhoven-Dobbe, J.C.; Gorbalenya, A.E.; Decroly, E.; Snijder, E.J.; Canard, B.; Imbert, I. One severe acute respiratory syndrome coronavirus protein complex integrates processive RNA polymerase and exonuclease activities. Proc. Natl. Acad. Sci. USA 2014, 111, E3900–E3909. [Google Scholar] [CrossRef]
- Garcia-Sastre, A.; Egorov, A.; Matassov, D.; Brandt, S.; Levy, D.E.; Durbin, J.E.; Palese, P.; Muster, T. Influenza A virus lacking the NS1 gene replicates in interferon-deficient systems. Virology 1998, 252, 324–330. [Google Scholar] [CrossRef]
- Neuman, B.W.; Angelini, M.M.; Buchmeier, M.J. Does form meet function in the coronavirus replicative organelle? Trends Microbiol. 2014, 22, 642–647. [Google Scholar] [CrossRef] [PubMed]
- Schubert, K.; Karousis, E.D.; Jomaa, A.; Scaiola, A.; Echeverria, B.; Gurzeler, L.A.; Leibundgut, M.; Thiel, V.; Muhlemann, O.; Ban, N. SARS-CoV-2 Nsp1 binds the ribosomal mRNA channel to inhibit translation. Nat. Struct. Mol. Biol. 2020, 27, 959–966. [Google Scholar] [CrossRef]
- Drosten, C.; Gunther, S.; Preiser, W.; van der Werf, S.; Brodt, H.R.; Becker, S.; Rabenau, H.; Panning, M.; Kolesnikova, L.; Fouchier, R.A.; et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 2003, 348, 1967–1976. [Google Scholar] [CrossRef] [PubMed]
- Feldmann, H.; Geisbert, T.W. Ebola haemorrhagic fever. Lancet 2011, 377, 849–862. [Google Scholar] [CrossRef]
- Taylor, L.H.; Latham, S.M.; Woolhouse, M.E. Risk factors for human disease emergence. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2001, 356, 983–989. [Google Scholar] [CrossRef] [PubMed]
- Tellier, R.; Li, Y.; Cowling, B.J.; Tang, J.W. Recognition of aerosol transmission of infectious agents: A commentary. BMC Infect. Dis. 2019, 19, 101. [Google Scholar] [CrossRef] [PubMed]
- Morawska, L.; Cao, J. Airborne transmission of SARS-CoV-2: The world should face the reality. Environ. Int. 2020, 139, 105730. [Google Scholar] [CrossRef] [PubMed]
- Tate, J.E.; Burton, A.H.; Boschi-Pinto, C.; Parashar, U.D.; World Health Organization–Coordinated Global Rotavirus Surveillance Network. Global, Regional, and National Estimates of Rotavirus Mortality in Children < 5 Years of Age, 2000–2013. Clin Infect. Dis 2016, 62 (Suppl. S2), S96–S105. [Google Scholar] [CrossRef]
- Cohen, M.S.; Chen, Y.Q.; McCauley, M.; Gamble, T.; Hosseinipour, M.C.; Kumarasamy, N.; Hakim, J.G.; Kumwenda, J.; Grinsztejn, B.; Pilotto, J.H.; et al. Antiretroviral Therapy for the Prevention of HIV-1 Transmission. N. Engl. J. Med. 2016, 375, 830–839. [Google Scholar] [CrossRef]
- Schillie, S.; Vellozzi, C.; Reingold, A.; Harris, A.; Haber, P.; Ward, J.W.; Nelson, N.P. Prevention of Hepatitis B Virus Infection in the United States: Recommendations of the Advisory Committee on Immunization Practices. MMWR Recomm. Rep. 2018, 67, 1–31. [Google Scholar] [CrossRef]
- Jackson, A.C. Current and future approaches to the therapy of human rabies. Antivir. Res. 2013, 99, 61–67. [Google Scholar] [CrossRef] [PubMed]
- Guzman, M.G.; Harris, E. Dengue. Lancet 2015, 385, 453–465. [Google Scholar] [CrossRef]
- Bhatt, S.; Gething, P.W.; Brady, O.J.; Messina, J.P.; Farlow, A.W.; Moyes, C.L.; Drake, J.M.; Brownstein, J.S.; Hoen, A.G.; Sankoh, O.; et al. The global distribution and burden of dengue. Nature 2013, 496, 504–507. [Google Scholar] [CrossRef] [PubMed]
- Saiz, J.C.; Vazquez-Calvo, A.; Blazquez, A.B.; Merino-Ramos, T.; Escribano-Romero, E.; Martin-Acebes, M.A. Zika Virus: The Latest Newcomer. Front. Microbiol. 2016, 7, 496. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef]
- Gambaryan, A.S.; Tuzikov, A.B.; Piskarev, V.E.; Yamnikova, S.S.; Lvov, D.K.; Robertson, J.S.; Bovin, N.V.; Matrosovich, M.N. Specification of receptor-binding phenotypes of influenza virus isolates from different hosts using synthetic sialylglycopolymers: Non-egg-adapted human H1 and H3 influenza A and influenza B viruses share a common high binding affinity for 6′-sialyl(N-acetyllactosamine). Virology 1997, 232, 345–350. [Google Scholar] [CrossRef]
- Matrosovich, M.; Tuzikov, A.; Bovin, N.; Gambaryan, A.; Klimov, A.; Castrucci, M.R.; Donatelli, I.; Kawaoka, Y. Early alterations of the receptor-binding properties of H1, H2, and H3 avian influenza virus hemagglutinins after their introduction into mammals. J. Virol. 2000, 74, 8502–8512. [Google Scholar] [CrossRef]
- White, J.; Kartenbeck, J.; Helenius, A. Membrane fusion activity of influenza virus. EMBO J. 1982, 1, 217–222. [Google Scholar] [CrossRef]
- Klenk, H.D.; Garten, W. Host cell proteases controlling virus pathogenicity. Trends Microbiol. 1994, 2, 39–43. [Google Scholar] [CrossRef]
- Hogle, J.M. Poliovirus cell entry: Common structural themes in viral cell entry pathways. Annu. Rev. Microbiol. 2002, 56, 677–702. [Google Scholar] [CrossRef]
- Lakadamyali, M.; Rust, M.J.; Zhuang, X. Endocytosis of influenza viruses. Microbes Infect. 2004, 6, 929–936. [Google Scholar] [CrossRef]
- Hirose, T.; Fujita, K.; Kusumoto, S.; Oki, Y.; Murata, Y.; Sugiyama, T.; Ishida, H.; Shirai, T.; Nakashima, M.; Yamaoka, T.; et al. Association of pharmacokinetics and pharmacogenomics with safety and efficacy of gefitinib in patients with EGFR mutation positive advanced non-small cell lung cancer. Lung Cancer 2016, 93, 69–76. [Google Scholar] [CrossRef]
- Hojjat-Farsangi, M. Small-molecule inhibitors of the receptor tyrosine kinases: Promising tools for targeted cancer therapies. Int. J. Mol. Sci. 2014, 15, 13768–13801. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Qian, W.; Sun, X.; Jiang, S. Small-molecule inhibitors, immune checkpoint inhibitors, and more: FDA-approved novel therapeutic drugs for solid tumors from 1991 to 2021. J. Hematol. Oncol. 2022, 15, 143. [Google Scholar] [CrossRef] [PubMed]
- Haas, K.M.; McGregor, M.J.; Bouhaddou, M.; Polacco, B.J.; Kim, E.Y.; Nguyen, T.T.; Newton, B.W.; Urbanowski, M.; Kim, H.; Williams, M.A.P.; et al. Proteomic and genetic analyses of influenza A viruses identify pan-viral host targets. Nat. Commun. 2023, 14, 6030. [Google Scholar] [CrossRef] [PubMed]
- Knight, Z.A. Small molecule inhibitors of the PI3-kinase family. Curr. Top. Microbiol. Immunol. 2010, 347, 263–278. [Google Scholar] [CrossRef]
- Bekerman, E.; Neveu, G.; Shulla, A.; Brannan, J.; Pu, S.Y.; Wang, S.; Xiao, F.; Barouch-Bentov, R.; Bakken, R.R.; Mateo, R.; et al. Anticancer kinase inhibitors impair intracellular viral trafficking and exert broad-spectrum antiviral effects. J. Clin. Investig. 2017, 127, 1338–1352. [Google Scholar] [CrossRef]
- Eaaswarkhanth, M.; Al Madhoun, A.; Al-Mulla, F. Could the D614G substitution in the SARS-CoV-2 spike (S) protein be associated with higher COVID-19 mortality? Int. J. Infect. Dis. 2020, 96, 459–460. [Google Scholar] [CrossRef] [PubMed]
- Keck, F.; Ataey, P.; Amaya, M.; Bailey, C.; Narayanan, A. Phosphorylation of Single Stranded RNA Virus Proteins and Potential for Novel Therapeutic Strategies. Viruses 2015, 7, 5257–5273. [Google Scholar] [CrossRef]
- Kumar, R.; Barua, S.; Tripathi, B.N.; Kumar, N. Role of ROCK signaling in virus replication. Virus Res. 2023, 329, 199105. [Google Scholar] [CrossRef]
- Chander, Y.; Kumar, R.; Khandelwal, N.; Singh, N.; Shringi, B.N.; Barua, S.; Kumar, N. Role of p38 mitogen-activated protein kinase signalling in virus replication and potential for developing broad spectrum antiviral drugs. Rev. Med. Virol. 2021, 31, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Adamson, A.L.; Darr, D.; Holley-Guthrie, E.; Johnson, R.A.; Mauser, A.; Swenson, J.; Kenney, S. Epstein-Barr virus immediate-early proteins BZLF1 and BRLF1 activate the ATF2 transcription factor by increasing the levels of phosphorylated p38 and c-Jun N-terminal kinases. J. Virol. 2000, 74, 1224–1233. [Google Scholar] [CrossRef]
- Sreekanth, G.P.; Yenchitsomanus, P.T.; Limjindaporn, T. Role of mitogen-activated protein kinase signaling in the pathogenesis of dengue virus infection. Cell Signal 2018, 48, 64–68. [Google Scholar] [CrossRef]
- Fung, T.S.; Liu, D.X. Activation of the c-Jun NH(2)-terminal kinase pathway by coronavirus infectious bronchitis virus promotes apoptosis independently of c-Jun. Cell Death Dis. 2017, 8, 3215. [Google Scholar] [CrossRef] [PubMed]
- Voss, K.; Amaya, M.; Mueller, C.; Roberts, B.; Kehn-Hall, K.; Bailey, C.; Petricoin, E., 3rd; Narayanan, A. Inhibition of host extracellular signal-regulated kinase (ERK) activation decreases new world alphavirus multiplication in infected cells. Virology 2014, 468–470, 490–503. [Google Scholar] [CrossRef]
- Shi, W.; Hou, X.; Peng, H.; Zhang, L.; Li, Y.; Gu, Z.; Jiang, Q.; Shi, M.; Ji, Y.; Jiang, J. MEK/ERK signaling pathway is required for enterovirus 71 replication in immature dendritic cells. Virol. J. 2014, 11, 227. [Google Scholar] [CrossRef]
- Cheng, Y.; Sun, F.; Wang, L.; Gao, M.; Xie, Y.; Sun, Y.; Liu, H.; Yuan, Y.; Yi, W.; Huang, Z.; et al. Virus-induced p38 MAPK activation facilitates viral infection. Theranostics 2020, 10, 12223–12240. [Google Scholar] [CrossRef]
- Kumar, N.; Sharma, S.; Kumar, R.; Tripathi, B.N.; Barua, S.; Ly, H.; Rouse, B.T. Host-Directed Antiviral Therapy. Clin. Microbiol. Rev. 2020, 33, e00168-19. [Google Scholar] [CrossRef] [PubMed]
- Keating, J.A.; Striker, R. Phosphorylation events during viral infections provide potential therapeutic targets. Rev. Med. Virol. 2012, 22, 166–181. [Google Scholar] [CrossRef]
- Chen, J.; Ye, C.; Wan, C.; Li, G.; Peng, L.; Peng, Y.; Fang, R. The Roles of c-Jun N-Terminal Kinase (JNK) in Infectious Diseases. Int. J. Mol. Sci. 2021, 22, 9640. [Google Scholar] [CrossRef]
- Shimojima, M.; Ikeda, Y.; Kawaoka, Y. The mechanism of Axl-mediated Ebola virus infection. J. Infect. Dis. 2007, 196 (Suppl. S2), S259–S263. [Google Scholar] [CrossRef]
- Zheng, K.; Kitazato, K.; Wang, Y. Viruses exploit the function of epidermal growth factor receptor. Rev. Med. Virol. 2014, 24, 274–286. [Google Scholar] [CrossRef]
- Wang, X.; Huong, S.M.; Chiu, M.L.; Raab-Traub, N.; Huang, E.S. Epidermal growth factor receptor is a cellular receptor for human cytomegalovirus. Nature 2003, 424, 456–461. [Google Scholar] [CrossRef]
- Weller, M.L.; Amornphimoltham, P.; Schmidt, M.; Wilson, P.A.; Gutkind, J.S.; Chiorini, J.A. Epidermal growth factor receptor is a co-receptor for adeno-associated virus serotype 6. Nat. Med. 2010, 16, 662–664. [Google Scholar] [CrossRef] [PubMed]
- Kumar, N.; Liang, Y.; Parslow, T.G.; Liang, Y. Receptor tyrosine kinase inhibitors block multiple steps of influenza a virus replication. J. Virol. 2011, 85, 2818–2827. [Google Scholar] [CrossRef]
- Karim, M.; Saul, S.; Ghita, L.; Sahoo, M.K.; Ye, C.; Bhalla, N.; Lo, C.-W.; Jin, J.; Park, J.-G.; Martinez-Gualda, B.; et al. Numb-associated kinases are required for SARS-CoV-2 infection and are cellular targets for antiviral strategies. Antivir. Res. 2022, 204, 105367. [Google Scholar] [CrossRef] [PubMed]
- Neveu, G.; Ziv-Av, A.; Barouch-Bentov, R.; Berkerman, E.; Mulholland, J.; Einav, S. AP-2-Associated Protein Kinase 1 and Cyclin G-Associated Kinase Regulate Hepatitis C Virus Entry and Are Potential Drug Targets. J. Virol. 2015, 89, 4387–4404. [Google Scholar] [CrossRef]
- Urits, I.; Israel, J.; Hakobyan, H.; Yusin, G.; Lassiter, G.; Fackler, N.; Berger, A.A.; Kassem, H.; Kaye, A.; Viswanath, O. Baricitinib for the treatment of rheumatoid arthritis. Reumatologia 2020, 58, 407–415. [Google Scholar] [CrossRef]
- Richardson, P.; Griffin, I.; Tucker, C.; Smith, D.; Oechsle, O.; Phelan, A.; Stebbing, J. Baricitinib as potential treatment for 2019-nCoV acute respiratory disease. Lancet 2020, 395, e30–e31. [Google Scholar] [CrossRef] [PubMed]
- Richardson, P.J.; Stebbing, J. Baricitinib as the treatment of choice for hospitalised individuals with COVID-19. EClinicalMedicine 2022, 49, 101493. [Google Scholar] [CrossRef]
- Newman, D.J.; Cragg, G.M. Natural Products as Sources of New Drugs over the Nearly Four Decades from 01/1981 to 09/2019. J. Nat. Prod. 2020, 83, 770–803. [Google Scholar] [CrossRef] [PubMed]
- Lu, A.T.; Fei, Z.; Haghani, A.; Robeck, T.R.; Zoller, J.A.; Li, C.Z.; Lowe, R.; Yan, Q.; Zhang, J.; Vu, H.; et al. Universal DNA methylation age across mammalian tissues. Nat. Aging 2023, 3, 1144–1166. [Google Scholar] [CrossRef] [PubMed]
- Bird, A. Perceptions of epigenetics. Nature 2007, 447, 396–398. [Google Scholar] [CrossRef]
- Musselman, C.A.; Lalonde, M.E.; Côté, J.; Kutateladze, T.G. Perceiving the epigenetic landscape through histone readers. Nat. Struct. Mol. Biol. 2012, 19, 1218–1227. [Google Scholar] [CrossRef] [PubMed]
- Lister, R.; Pelizzola, M.; Dowen, R.H.; Hawkins, R.D.; Hon, G.; Tonti-Filippini, J.; Nery, J.R.; Lee, L.; Ye, Z.; Ngo, Q.M.; et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 2009, 462, 315–322. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.K. Active DNA demethylation mediated by DNA glycosylases. Annu. Rev. Genet. 2009, 43, 143–166. [Google Scholar] [CrossRef]
- Gopalakrishnan, S.; Van Emburgh, B.O.; Robertson, K.D. DNA methylation in development and human disease. Mutat. Res. 2008, 647, 30–38. [Google Scholar] [CrossRef]
- Jin, B.; Tao, Q.; Peng, J.; Soo, H.M.; Wu, W.; Ying, J.; Fields, C.R.; Delmas, A.L.; Liu, X.; Qiu, J.; et al. DNA methyltransferase 3B (DNMT3B) mutations in ICF syndrome lead to altered epigenetic modifications and aberrant expression of genes regulating development, neurogenesis and immune function. Hum. Mol. Genet. 2008, 17, 690–709. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, Y.; Wang, C.; Wang, X. TET (Ten-eleven translocation) family proteins: Structure, biological functions and applications. Signal Transduct. Target. Ther. 2023, 8, 297. [Google Scholar] [CrossRef]
- Horvath, S.; Raj, K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat. Rev. Genet. 2018, 19, 371–384. [Google Scholar] [CrossRef]
- Feinberg, A.P.; Ohlsson, R.; Henikoff, S. The epigenetic progenitor origin of human cancer. Nat. Rev. Genet. 2006, 7, 21–33. [Google Scholar] [CrossRef]
- Kuo, M.H.; Allis, C.D. Roles of histone acetyltransferases and deacetylases in gene regulation. Bioessays 1998, 20, 615–626. [Google Scholar] [CrossRef]
- Dokmanovic, M.; Clarke, C.; Marks, P.A. Histone deacetylase inhibitors: Overview and perspectives. Mol. Cancer Res. 2007, 5, 981–989. [Google Scholar] [CrossRef] [PubMed]
- Blackledge, N.P.; Klose, R.J. The molecular principles of gene regulation by Polycomb repressive complexes. Nat. Rev. Mol. Cell Biol. 2021, 22, 815–833. [Google Scholar] [CrossRef] [PubMed]
- Healey, M.A.; Hu, R.; Beck, A.H.; Collins, L.C.; Schnitt, S.J.; Tamimi, R.M.; Hazra, A. Association of H3K9me3 and H3K27me3 repressive histone marks with breast cancer subtypes in the Nurses’ Health Study. Breast Cancer Res. Treat. 2014, 147, 639–651. [Google Scholar] [CrossRef] [PubMed]
- Handy, D.E.; Castro, R.; Loscalzo, J. Epigenetic modifications: Basic mechanisms and role in cardiovascular disease. Circulation 2011, 123, 2145–2156. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Mason, C.E. The pivotal regulatory landscape of RNA modifications. Annu. Rev. Genom. Hum. Genet. 2014, 15, 127–150. [Google Scholar] [CrossRef] [PubMed]
- Hombach, S.; Kretz, M. Non-coding RNAs: Classification, Biology and Functioning. Adv. Exp. Med. Biol. 2016, 937, 3–17. [Google Scholar] [CrossRef]
- Lujambio, A.; Lowe, S.W. The microcosmos of cancer. Nature 2012, 482, 347–355. [Google Scholar] [CrossRef]
- Kwa, F.A.A.; Jackson, D.E. Manipulating the epigenome for the treatment of disorders with thrombotic complications. Drug Discov. Today 2018, 23, 719–726. [Google Scholar] [CrossRef]
- Silva, J.; Tavares, V.; Afonso, A.; Garcia, J.; Cerqueira, F.; Medeiros, R. Plasmatic MicroRNAs and Treatment Outcomes of Patients with Metastatic Castration-Resistant Prostate Cancer: A Hospital-Based Cohort Study and In Silico Analysis. Int. J. Mol. Sci. 2023, 24, 9101. [Google Scholar] [CrossRef]
- Li, Y.; He, Q.; Wen, X.; Hong, X.; Yang, X.; Tang, X.; Zhang, P.; Lei, Y.; Sun, Y.; Zhang, J.; et al. EZH2-DNMT1-mediated epigenetic silencing of miR-142-3p promotes metastasis through targeting ZEB2 in nasopharyngeal carcinoma. Cell Death Differ. 2019, 26, 1089–1106. [Google Scholar] [CrossRef]
- Chhabra, R. miRNA and methylation: A multifaceted liaison. Chembiochem 2015, 16, 195–203. [Google Scholar] [CrossRef]
- Duursma, A.M.; Kedde, M.; Schrier, M.; le Sage, C.; Agami, R. miR-148 targets human DNMT3b protein coding region. Rna 2008, 14, 872–877. [Google Scholar] [CrossRef]
- Liao, X.; Zhou, Z.; Zhang, X. Effects of miR-195-5p on cell proliferation and apoptosis in gestational diabetes mellitus via targeting EZH2. Mol. Med. Rep. 2020, 22, 803–809. [Google Scholar] [CrossRef] [PubMed]
- Portela, A.; Esteller, M. Epigenetic modifications and human disease. Nat. Biotechnol. 2010, 28, 1057–1068. [Google Scholar] [CrossRef] [PubMed]
- Obata, Y.; Furusawa, Y.; Hase, K. Epigenetic modifications of the immune system in health and disease. Immunol. Cell Biol. 2015, 93, 226–232. [Google Scholar] [CrossRef] [PubMed]
- Smale, S.T.; Tarakhovsky, A.; Natoli, G. Chromatin contributions to the regulation of innate immunity. Annu. Rev. Immunol. 2014, 32, 489–511. [Google Scholar] [CrossRef] [PubMed]
- Tsai, K.; Cullen, B.R. Epigenetic and epitranscriptomic regulation of viral replication. Nat. Rev. Microbiol. 2020, 18, 559–570. [Google Scholar] [CrossRef]
- Balakrishnan, L.; Milavetz, B. Epigenetic Regulation of Viral Biological Processes. Viruses 2017, 9, 346. [Google Scholar] [CrossRef] [PubMed]
- Knipe, D.M. Nuclear sensing of viral DNA, epigenetic regulation of herpes simplex virus infection, and innate immunity. Virology 2015, 479–480, 153–159. [Google Scholar] [CrossRef]
- Knipe, D.M.; Raja, P.; Lee, J. Viral gene products actively promote latent infection by epigenetic silencing mechanisms. Curr. Opin. Virol. 2017, 23, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Tsai, K.; Jaguva Vasudevan, A.A.; Martinez Campos, C.; Emery, A.; Swanstrom, R.; Cullen, B.R. Acetylation of Cytidine Residues Boosts HIV-1 Gene Expression by Increasing Viral RNA Stability. Cell Host Microbe 2020, 28, 306–312. [Google Scholar] [CrossRef]
- Kee, J.; Thudium, S.; Renner, D.M.; Glastad, K.; Palozola, K.; Zhang, Z.; Li, Y.; Lan, Y.; Cesare, J.; Poleshko, A.; et al. SARS-CoV-2 disrupts host epigenetic regulation via histone mimicry. Nature 2022, 610, 381–388. [Google Scholar] [CrossRef]
- Liang, Y.; Vogel, J.L.; Narayanan, A.; Peng, H.; Kristie, T.M. Inhibition of the histone demethylase LSD1 blocks alpha-herpesvirus lytic replication and reactivation from latency. Nat. Med. 2009, 15, 1312–1317. [Google Scholar] [CrossRef]
- Schneider, W.M.; Chevillotte, M.D.; Rice, C.M. Interferon-stimulated genes: A complex web of host defenses. Annu. Rev. Immunol. 2014, 32, 513–545. [Google Scholar] [CrossRef] [PubMed]
- García-Sastre, A.; Biron, C.A. Type 1 interferons and the virus-host relationship: A lesson in détente. Science 2006, 312, 879–882. [Google Scholar] [CrossRef] [PubMed]
- Menachery, V.D.; Eisfeld, A.J.; Schafer, A.; Josset, L.; Sims, A.C.; Proll, S.; Fan, S.; Li, C.; Neumann, G.; Tilton, S.C.; et al. Pathogenic influenza viruses and coronaviruses utilize similar and contrasting approaches to control interferon-stimulated gene responses. mBio 2014, 5, e01174-14. [Google Scholar] [CrossRef]
- Chen, Y.; Liu, Q.; Guo, D. Emerging coronaviruses: Genome structure, replication, and pathogenesis. J. Med. Virol. 2020, 92, 418–423. [Google Scholar] [CrossRef]
- Cole, J.; Morris, P.; Dickman, M.J.; Dockrell, D.H. The therapeutic potential of epigenetic manipulation during infectious diseases. Pharmacol. Ther. 2016, 167, 85–99. [Google Scholar] [CrossRef]
- Laurson, J.; Khan, S.; Chung, R.; Cross, K.; Raj, K. Epigenetic repression of E-cadherin by human papillomavirus 16 E7 protein. Carcinogenesis 2010, 31, 918–926. [Google Scholar] [CrossRef] [PubMed]
- Burgers, W.A.; Blanchon, L.; Pradhan, S.; de Launoit, Y.; Kouzarides, T.; Fuks, F. Viral oncoproteins target the DNA methyltransferases. Oncogene 2007, 26, 1650–1655. [Google Scholar] [CrossRef] [PubMed]
- Holland, D.; Hoppe-Seyler, K.; Schuller, B.; Lohrey, C.; Maroldt, J.; Dürst, M.; Hoppe-Seyler, F. Activation of the enhancer of zeste homologue 2 gene by the human papillomavirus E7 oncoprotein. Cancer Res. 2008, 68, 9964–9972. [Google Scholar] [CrossRef] [PubMed]
- Gil, J.; Peters, G. Regulation of the INK4b-ARF-INK4a tumour suppressor locus: All for one or one for all. Nat. Rev. Mol. Cell Biol. 2006, 7, 667–677. [Google Scholar] [CrossRef] [PubMed]
- Toyooka, S.; Pass, H.I.; Shivapurkar, N.; Fukuyama, Y.; Maruyama, R.; Toyooka, K.O.; Gilcrease, M.; Farinas, A.; Minna, J.D.; Gazdar, A.F. Aberrant methylation and simian virus 40 tag sequences in malignant mesothelioma. Cancer Res. 2001, 61, 5727–5730. [Google Scholar] [PubMed]
- Slack, A.; Cervoni, N.; Pinard, M.; Szyf, M. DNA methyltransferase is a downstream effector of cellular transformation triggered by simian virus 40 large T antigen. J. Biol. Chem. 1999, 274, 10105–10112. [Google Scholar] [CrossRef]
- Ferrari, R.; Pellegrini, M.; Horwitz, G.A.; Xie, W.; Berk, A.J.; Kurdistani, S.K. Epigenetic reprogramming by adenovirus e1a. Science 2008, 321, 1086–1088. [Google Scholar] [CrossRef]
- Li, J.; Hao, D.; Wang, L.; Wang, H.; Wang, Y.; Zhao, Z.; Li, P.; Deng, C.; Di, L.J. Epigenetic targeting drugs potentiate chemotherapeutic effects in solid tumor therapy. Sci. Rep. 2017, 7, 4035. [Google Scholar] [CrossRef]
- El Baba, R.; Herbein, G. Management of epigenomic networks entailed in coronavirus infections and COVID-19. Clin. Epigenetics 2020, 12, 118. [Google Scholar] [CrossRef]
- Zheng, B.; Zhou, J.; Wang, H. Host microRNAs and exosomes that modulate influenza virus infection. Virus Res. 2020, 279, 197885. [Google Scholar] [CrossRef]
- Moghoofei, M.; Najafipour, S.; Mostafaei, S.; Tavakoli, A.; Bokharaei-Salim, F.; Ghorbani, S.; Javanmard, D.; Ghaffari, H.; Monavari, S.H. MicroRNAs Profiling in HIV, HCV, and HIV/HCV Co-Infected Patients. Curr. HIV Res. 2021, 19, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Kunden, R.D.; Khan, J.Q.; Ghezelbash, S.; Wilson, J.A. The Role of the Liver-Specific microRNA, miRNA-122 in the HCV Replication Cycle. Int. J. Mol. Sci. 2020, 21, 5677. [Google Scholar] [CrossRef] [PubMed]
- Jopling, C.L.; Yi, M.; Lancaster, A.M.; Lemon, S.M.; Sarnow, P. Modulation of hepatitis C virus RNA abundance by a liver-specific MicroRNA. Science 2005, 309, 1577–1581. [Google Scholar] [CrossRef] [PubMed]
- Panigrahi, M.; Thibault, P.A.; Wilson, J.A. MicroRNA 122 Affects both the Initiation and the Maintenance of Hepatitis C Virus Infections. J. Virol. 2022, 96, e0190321. [Google Scholar] [CrossRef] [PubMed]
- Yousefpouran, S.; Mostafaei, S.; Manesh, P.V.; Iranifar, E.; Bokharaei-Salim, F.; Nahand, J.S.; Mirzaei, H.; Taran, M.; Babaei, F.; Sayad, B.; et al. The assessment of selected MiRNAs profile in HIV, HBV, HCV, HIV/HCV, HIV/HBV Co-infection and elite controllers for determination of biomarker. Microb. Pathog. 2020, 147, 104355. [Google Scholar] [CrossRef]
- Haneklaus, M.; Gerlic, M.; O’Neill, L.A.; Masters, S.L. miR-223: Infection, inflammation and cancer. J. Intern. Med. 2013, 274, 215–226. [Google Scholar] [CrossRef]
- Morales, L.; Oliveros, J.C.; Enjuanes, L.; Sola, I. Contribution of Host miRNA-223-3p to SARS-CoV-Induced Lung Inflammatory Pathology. mBio 2022, 13, e0313521. [Google Scholar] [CrossRef]
- Tycowski, K.T.; Guo, Y.E.; Lee, N.; Moss, W.N.; Vallery, T.K.; Xie, M.; Steitz, J.A. Viral noncoding RNAs: More surprises. Genes. Dev. 2015, 29, 567–584. [Google Scholar] [CrossRef]
- Cazalla, D.; Yario, T.; Steitz, J.A. Down-regulation of a host microRNA by a Herpesvirus saimiri noncoding RNA. Science 2010, 328, 1563–1566. [Google Scholar] [CrossRef]
- Guo, Y.E.; Riley, K.J.; Iwasaki, A.; Steitz, J.A. Alternative capture of noncoding RNAs or protein-coding genes by herpesviruses to alter host T cell function. Mol. Cell 2014, 54, 67–79. [Google Scholar] [CrossRef]
- Abedini, M.; Zhang, C. Performance assessment of concrete and steel material models in ls-dyna for enhanced numerical simulation, a state of the art review. Arch. Comput. Methods Eng. 2021, 28, 2921–2942. [Google Scholar] [CrossRef]
- Kitazato, K.; Wang, Y.; Kobayashi, N. Viral infectious disease and natural products with antiviral activity. Drug Discov. Ther. 2007, 1, 14–22. [Google Scholar] [PubMed]
- Abookleesh, F.L.; Al-Anzi, B.S.; Ullah, A. Potential Antiviral Action of Alkaloids. Molecules 2022, 27, 903. [Google Scholar] [CrossRef]
- Denaro, M.; Smeriglio, A.; Barreca, D.; De Francesco, C.; Occhiuto, C.; Milano, G.; Trombetta, D. Antiviral activity of plants and their isolated bioactive compounds: An update. Phytother. Res. 2020, 34, 742–768. [Google Scholar] [CrossRef] [PubMed]
- Behl, T.; Rocchetti, G.; Chadha, S.; Zengin, G.; Bungau, S.; Kumar, A.; Mehta, V.; Uddin, M.S.; Khullar, G.; Setia, D.; et al. Phytochemicals from Plant Foods as Potential Source of Antiviral Agents: An Overview. Pharm. 2021, 14, 381. [Google Scholar] [CrossRef] [PubMed]
- Jassim, S.A.; Naji, M.A. Novel antiviral agents: A medicinal plant perspective. J. Appl. Microbiol. 2003, 95, 412–427. [Google Scholar] [CrossRef] [PubMed]
- Astani, A.; Reichling, J.; Schnitzler, P. Screening for antiviral activities of isolated compounds from essential oils. Evid. Based Complement. Altern. Med. 2011, 2011, 253643. [Google Scholar] [CrossRef]
- Gavanji, S.; Sayedipour, S.S.; Larki, B.; Bakhtari, A. Antiviral activity of some plant oils against herpes simplex virus type 1 in Vero cell culture. J. Acute Med. 2015, 5, 62–68. [Google Scholar] [CrossRef]
- Feriotto, G.; Marchetti, N.; Costa, V.; Beninati, S.; Tagliati, F.; Mischiati, C. Chemical Composition of Essential Oils from Thymus vulgaris, Cymbopogon citratus, and Rosmarinus officinalis, and Their Effects on the HIV-1 Tat Protein Function. Chem. Biodivers. 2018, 15, e1700436. [Google Scholar] [CrossRef]
- Mori, K.; Obossou, E.K.; Suwa, S.; Miura, S.; Oh, S.-H.; Jinbo, N.; Ishibashi, Y.; Shikamoto, Y.; Hosono, T.; Toda, T. Human Immunodeficiency Virus Type 1 (HIV-1) Reverse Transcriptase Inhibitory Effect of Cymbopogon Nardus Essential Oil. Int. J. Adv. Res. 2016, 2, 7–13. [Google Scholar]
- Ma, L.; Yao, L. Antiviral Effects of Plant-Derived Essential Oils and Their Components: An Updated Review. Molecules 2020, 25, 2627. [Google Scholar] [CrossRef]
- Naithani, R.; Huma, L.C.; Holland, L.E.; Shukla, D.; McCormick, D.L.; Mehta, R.G.; Moriarty, R.M. Antiviral activity of phytochemicals: A comprehensive review. Mini Rev. Med. Chem. 2008, 8, 1106–1133. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.J.; Lim, C.H.; Song, J.H.; Baek, S.H.; Kwon, D.H. Antiviral activity of raoulic acid from Raoulia australis against Picornaviruses. Phytomedicine 2009, 16, 35–39. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.T.; Chen, T.Y.; Lin, S.C.; Chung, C.Y.; Lin, T.C.; Wang, G.H.; Anderson, R.; Lin, C.C.; Richardson, C.D. Broad-spectrum antiviral activity of chebulagic acid and punicalagin against viruses that use glycosaminoglycans for entry. BMC Microbiol. 2013, 13, 187. [Google Scholar] [CrossRef] [PubMed]
- Cotin, S.; Calliste, C.A.; Mazeron, M.C.; Hantz, S.; Duroux, J.L.; Rawlinson, W.D.; Ploy, M.C.; Alain, S. Eight flavonoids and their potential as inhibitors of human cytomegalovirus replication. Antivir. Res. 2012, 96, 181–186. [Google Scholar] [CrossRef]
- Lyu, S.Y.; Rhim, J.Y.; Park, W.B. Antiherpetic activities of flavonoids against herpes simplex virus type 1 (HSV-1) and type 2 (HSV-2) in vitro. Arch. Pharm. Res. 2005, 28, 1293–1301. [Google Scholar] [CrossRef]
- Kim, D.E.; Min, J.S.; Jang, M.S.; Lee, J.Y.; Shin, Y.S.; Song, J.H.; Kim, H.R.; Kim, S.; Jin, Y.H.; Kwon, S. Natural Bis-Benzylisoquinoline Alkaloids-Tetrandrine, Fangchinoline, and Cepharanthine, Inhibit Human Coronavirus OC43 Infection of MRC-5 Human Lung Cells. Biomolecules 2019, 9, 696. [Google Scholar] [CrossRef]
- Luganini, A.; Mercorelli, B.; Messa, L.; Palù, G.; Gribaudo, G.; Loregian, A. The isoquinoline alkaloid berberine inhibits human cytomegalovirus replication by interfering with the viral Immediate Early-2 (IE2) protein transactivating activity. Antivir. Res. 2019, 164, 52–60. [Google Scholar] [CrossRef]
- Song, S.; Qiu, M.; Chu, Y.; Chen, D.; Wang, X.; Su, A.; Wu, Z. Downregulation of cellular c-Jun N-terminal protein kinase and NF-κB activation by berberine may result in inhibition of herpes simplex virus replication. Antimicrob. Agents Chemother. 2014, 58, 5068–5078. [Google Scholar] [CrossRef]
- Gómez, L.A.; Stashenko, E.; Ocazionez, R.E. Comparative study on in vitro activities of citral, limonene and essential oils from Lippia citriodora and L. alba on yellow fever virus. Nat. Prod. Commun. 2013, 8, 249–252. [Google Scholar] [CrossRef]
- Haddad, J.G.; Picard, M.; Bénard, S.; Desvignes, C.; Desprès, P.; Diotel, N.; El Kalamouni, C. Ayapana triplinervis Essential Oil and Its Main Component Thymohydroquinone Dimethyl Ether Inhibit Zika Virus at Doses Devoid of Toxicity in Zebrafish. Molecules 2019, 24, 3447. [Google Scholar] [CrossRef] [PubMed]
- Vimalanathan, S.; Hudson, J.B. Anti-influenza virus activity of essential oils and vapors. Am. J. Essent. Oils Nat. Prod. 2014, 2, 47–53. [Google Scholar]
- Mokni, R.E.; Youssef, F.S.; Jmii, H.; Khmiri, A.; Bouazzi, S.; Jlassi, I.; Jaidane, H.; Dhaouadi, H.; Ashour, M.L.; Hammami, S. The Essential Oil of Tunisian Dysphania ambrosioides and its Antimicrobial and Antiviral Properties. J. Essent. Oil Bear. Plants 2019, 22, 282–294. [Google Scholar] [CrossRef]
- Elaissi, A.; Rouis, Z.; Salem, N.A.; Mabrouk, S.; ben Salem, Y.; Salah, K.B.; Aouni, M.; Farhat, F.; Chemli, R.; Harzallah-Skhiri, F.; et al. Chemical composition of 8 eucalyptus species’ essential oils and the evaluation of their antibacterial, antifungal and antiviral activities. BMC Complement. Altern. Med. 2012, 12, 81. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, N.A.; El-Hawary, S.S.; Mohammed, M.M.; Farid, M.A.; AbdelWahed, N.A.; Ali, M.A.; El-Abd, E.A. Chemical Composition, Antiviral against avian Influenza (H5N1) Virus and Antimicrobial activities of the Essential Oils of the Leaves and Fruits of Fortunella margarita, Lour. Swingle, Growing in Egypt. J. Appl. Pharm. Sci. 2015, 5, 006–012. [Google Scholar]
- Sharifi-Rad, J.; Salehi, B.; Schnitzler, P.; Ayatollahi, S.A.; Kobarfard, F.; Fathi, M.; Eisazadeh, M.; Sharifi-Rad, M. Susceptibility of herpes simplex virus type 1 to monoterpenes thymol, carvacrol, p-cymene and essential oils of Sinapis arvensis L., Lallemantia royleana Benth. and Pulicaria vulgaris Gaertn. Cell. Mol. Biol. 2017, 63, 42–47. [Google Scholar] [CrossRef] [PubMed]
- Pilau, M.R.; Alves, S.H.; Weiblen, R.; Arenhart, S.; Cueto, A.P.; Lovato, L.T. Antiviral activity of the Lippia graveolens (Mexican oregano) essential oil and its main compound carvacrol against human and animal viruses. Braz. J. Microbiol. 2011, 42, 1616–1624. [Google Scholar] [CrossRef]
- Garozzo, A.; Timpanaro, R.; Stivala, A.; Bisignano, G.; Castro, A. Activity of Melaleuca alternifolia (tea tree) oil on Influenza virus A/PR/8: Study on the mechanism of action. Antivir. Res. 2011, 89, 83–88. [Google Scholar] [CrossRef]
- Civitelli, L.; Panella, S.; Marcocci, M.E.; De Petris, A.; Garzoli, S.; Pepi, F.; Vavala, E.; Ragno, R.; Nencioni, L.; Palamara, A.T.; et al. In vitro inhibition of herpes simplex virus type 1 replication by Mentha suaveolens essential oil and its main component piperitenone oxide. Phytomedicine 2014, 21, 857–865. [Google Scholar] [CrossRef]
- da Silva, J.K.R.; Figueiredo, P.L.B.; Byler, K.G.; Setzer, W.N. Essential Oils as Antiviral Agents, Potential of Essential Oils to Treat SARS-CoV-2 Infection: An In-Silico Investigation. Int. J. Mol. Sci. 2020, 21, 3426. [Google Scholar]
- Cagno, V.; Sgorbini, B.; Sanna, C.; Cagliero, C.; Ballero, M.; Civra, A.; Donalisio, M.; Bicchi, C.; Lembo, D.; Rubiolo, P. In vitro anti-herpes simplex virus-2 activity of Salvia desoleana Atzei & V. Picci essential oil. PLoS ONE 2017, 12, e0172322. [Google Scholar] [CrossRef]
- Toujani, M.M.; Rittà, M.; Civra, A.; Genovese, S.; Epifano, F.; Ghram, A.; Lembo, D.; Donalisio, M. Inhibition of HSV-2 infection by pure compounds from Thymus capitatus extract in vitro. Phytother. Res. 2018, 32, 1555–1563. [Google Scholar] [CrossRef] [PubMed]
- Shin, W.J.; Lee, K.H.; Park, M.H.; Seong, B.L. Broad-spectrum antiviral effect of Agrimonia pilosa extract on influenza viruses. Microbiol. Immunol. 2010, 54, 11–19. [Google Scholar] [CrossRef]
- Rezazadeh, F.; Moshaverinia, M.; Motamedifar, M.; Alyaseri, M. Assessment of Anti HSV-1 Activity of Aloe Vera Gel Extract: An In Vitro Study. J. Dent. 2016, 17, 49–54. [Google Scholar]
- Makau, J.N.; Watanabe, K.; Mohammed, M.M.D.; Nishida, N. Antiviral Activity of Peanut (Arachis hypogaea L.) Skin Extract Against Human Influenza Viruses. J. Med. Food 2018, 21, 777–784. [Google Scholar] [CrossRef] [PubMed]
- Namazi, R.; Zabihollahi, R.; Behbahani, M.; Rezaei, A. Inhibitory Activity of Avicennia marina, a Medicinal Plant in Persian Folk Medicine, against HIV and HSV. Iran. J. Pharm. Res. 2013, 12, 435–443. [Google Scholar]
- Prinsloo, G.; Marokane, C.K.; Street, R.A. Anti-HIV activity of southern African plants: Current developments, phytochemistry and future research. J. Ethnopharmacol. 2018, 210, 133–155. [Google Scholar] [CrossRef]
- Mushi, N.F.; Mbwambo, Z.H.; Innocent, E.; Tewtrakul, S. Antibacterial, anti-HIV-1 protease and cytotoxic activities of aqueous ethanolic extracts from Combretum adenogonium Steud. Ex A. Rich (Combretaceae). BMC Complement. Altern. Med. 2012, 12, 163. [Google Scholar] [CrossRef]
- Churqui, M.P.; Lind, L.; Thörn, K.; Svensson, A.; Savolainen, O.; Aranda, K.T.; Eriksson, K. Extracts of Equisetum giganteum L and Copaifera reticulate Ducke show strong antiviral activity against the sexually transmitted pathogen herpes simplex virus type 2. J. Ethnopharmacol. 2018, 210, 192–197. [Google Scholar] [CrossRef]
- Lavoie, S.; Côté, I.; Pichette, A.; Gauthier, C.; Ouellet, M.; Nagau-Lavoie, F.; Mshvildadze, V.; Legault, J. Chemical composition and anti-herpes simplex virus type 1 (HSV-1) activity of extracts from Cornus canadensis. BMC Complement. Altern. Med. 2017, 17, 123. [Google Scholar] [CrossRef]
- Hossan, M.S.; Fatima, A.; Rahmatullah, M.; Khoo, T.J.; Nissapatorn, V.; Galochkina, A.V.; Slita, A.V.; Shtro, A.A.; Nikolaeva, Y.; Zarubaev, V.V.; et al. Antiviral activity of Embelia ribes Burm. f. against influenza virus in vitro. Arch. Virol. 2018, 163, 2121–2131. [Google Scholar] [CrossRef] [PubMed]
- Cho, W.K.; Weeratunga, P.; Lee, B.H.; Park, J.S.; Kim, C.J.; Ma, J.Y.; Lee, J.S. Epimedium koreanum Nakai displays broad spectrum of antiviral activity in vitro and in vivo by inducing cellular antiviral state. Viruses 2015, 7, 352–377. [Google Scholar] [CrossRef]
- Derksen, A.; Kühn, J.; Hafezi, W.; Sendker, J.; Ehrhardt, C.; Ludwig, S.; Hensel, A. Antiviral activity of hydroalcoholic extract from Eupatorium perfoliatum L. against the attachment of influenza A virus. J. Ethnopharmacol. 2016, 188, 144–152. [Google Scholar] [CrossRef] [PubMed]
- Gyuris, A.; Szlávik, L.; Minárovits, J.; Vasas, A.; Molnár, J.; Hohmann, J. Antiviral activities of extracts of Euphorbia hirta L. against HIV-1, HIV-2 and SIVmac251. Vivo 2009, 23, 429–432. [Google Scholar]
- Ghosh, M.; Civra, A.; Rittà, M.; Cagno, V.; Mavuduru, S.G.; Awasthi, P.; Lembo, D.; Donalisio, M. Ficus religiosa L. bark extracts inhibit infection by herpes simplex virus type 2 in vitro. Arch. Virol. 2016, 161, 3509–3514. [Google Scholar] [CrossRef]
- Bonvicini, F.; Lianza, M.; Mandrone, M.; Poli, F.; Gentilomi, G.A.; Antognoni, F. Hemidesmus indicus (L.) R. Br. extract inhibits the early step of herpes simplex type 1 and type 2 replication. New Microbiol. 2018, 41, 187–194. [Google Scholar] [PubMed]
- Shoji, M.; Woo, S.Y.; Masuda, A.; Win, N.N.; Ngwe, H.; Takahashi, E.; Kido, H.; Morita, H.; Ito, T.; Kuzuhara, T. Anti-influenza virus activity of extracts from the stems of Jatropha multifida Linn. collected in Myanmar. BMC Complement. Altern. Med. 2017, 17, 96. [Google Scholar] [CrossRef]
- Ho, J.Y.; Chang, H.W.; Lin, C.F.; Liu, C.J.; Hsieh, C.F.; Horng, J.T. Characterization of the anti-influenza activity of the Chinese herbal plant Paeonia lactiflora. Viruses 2014, 6, 1861–1875. [Google Scholar] [CrossRef]
- Ojha, D.; Das, R.; Sobia, P.; Dwivedi, V.; Ghosh, S.; Samanta, A.; Chattopadhyay, D. Pedilanthus tithymaloides Inhibits HSV Infection by Modulating NF-κB Signaling. PLoS ONE 2015, 10, e0139338. [Google Scholar] [CrossRef]
- Oh, C.; Price, J.; Brindley, M.A.; Widrlechner, M.P.; Qu, L.; McCoy, J.A.; Murphy, P.; Hauck, C.; Maury, W. Inhibition of HIV-1 infection by aqueous extracts of Prunella vulgaris L. Virol. J. 2011, 8, 188. [Google Scholar] [CrossRef]
- Bisignano, C.; Mandalari, G.; Smeriglio, A.; Trombetta, D.; Pizzo, M.M.; Pennisi, R.; Sciortino, M.T. Almond Skin Extracts Abrogate HSV-1 Replication by Blocking Virus Binding to the Cell. Viruses 2017, 9, 178. [Google Scholar] [CrossRef] [PubMed]
- Karimi, A.; Rafieian-Kopaei, M.; Moradi, M.T.; Alidadi, S. Anti-Herpes Simplex Virus Type-1 Activity and Phenolic Content of Crude Ethanol Extract and Four Corresponding Fractions of Quercus brantii L Acorn. J. Evid. Based Complement. Altern. Med. 2017, 22, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Reichling, J.; Neuner, A.; Sharaf, M.; Harkenthal, M.; Schnitzler, P. Antiviral activity of Rhus aromatica (fragrant sumac) extract against two types of herpes simplex viruses in cell culture. Pharmazie 2009, 64, 538–541. [Google Scholar] [PubMed]
- Nocchi, S.R.; Companhoni, M.V.; de Mello, J.C.; Dias Filho, B.P.; Nakamura, C.V.; Carollo, C.A.; Silva, D.B.; Ueda-Nakamura, T. Antiviral Activity of Crude Hydroethanolic Extract from Schinus terebinthifolia against Herpes simplex Virus Type 1. Planta Med. 2017, 83, 509–518. [Google Scholar] [CrossRef] [PubMed]
- Di Sotto, A.; Di Giacomo, S.; Amatore, D.; Locatelli, M.; Vitalone, A.; Toniolo, C.; Rotino, G.L.; Lo Scalzo, R.; Palamara, A.T.; Marcocci, M.E.; et al. A Polyphenol Rich Extract from Solanum melongena L. DR2 Peel Exhibits Antioxidant Properties and Anti-Herpes Simplex Virus Type 1 Activity In Vitro. Molecules 2018, 23, 2066. [Google Scholar] [CrossRef]
- Boff, L.; Silva, I.T.; Argenta, D.F.; Farias, L.M.; Alvarenga, L.F.; Pádua, R.M.; Braga, F.C.; Leite, J.P.; Kratz, J.M.; Simões, C.M. Strychnos pseudoquina A. St. Hil.: A Brazilian medicinal plant with promising in vitro antiherpes activity. J. Appl. Microbiol. 2016, 121, 1519–1529. [Google Scholar] [CrossRef]
- Benassi-Zanqueta, É.; Marques, C.F.; Valone, L.M.; Pellegrini, B.L.; Bauermeister, A.; Ferreira, I.C.P.; Lopes, N.P.; Nakamura, C.V.; Dias Filho, B.P.; Natali, M.R.M.; et al. Evaluation of anti-HSV-1 activity and toxicity of hydroethanolic extract of Tanacetum parthenium (L.) Sch.Bip. (Asteraceae). Phytomedicine 2019, 55, 249–254. [Google Scholar] [CrossRef]
- Hsieh, C.F.; Chen, Y.L.; Lin, C.F.; Ho, J.Y.; Huang, C.H.; Chiu, C.H.; Hsieh, P.W.; Horng, J.T. An extract from Taxodium distichum targets hemagglutinin- and neuraminidase-related activities of influenza virus in vitro. Sci. Rep. 2016, 6, 36015. [Google Scholar] [CrossRef]
- Donalisio, M.; Cagno, V.; Civra, A.; Gibellini, D.; Musumeci, G.; Rittà, M.; Ghosh, M.; Lembo, D. The traditional use of Vachellia nilotica for sexually transmitted diseases is substantiated by the antiviral activity of its bark extract against sexually transmitted viruses. J. Ethnopharmacol. 2018, 213, 403–408. [Google Scholar] [CrossRef]
- Hafidh, R.R.; Abdulamir, A.S.; Abu Bakar, F.; Sekawi, Z.; Jahansheri, F.; Jalilian, F.A. Novel antiviral activity of mung bean sprouts against respiratory syncytial virus and herpes simplex virus -1: An in vitro study on virally infected Vero and MRC-5 cell lines. BMC Complement. Altern. Med. 2015, 15, 179. [Google Scholar] [CrossRef]
- Latif, R.; Wang, C.Y. Andrographolide as a potent and promising antiviral agent. Chin. J. Nat. Med. 2020, 18, 760–769. [Google Scholar] [CrossRef]
- Gupta, S.; Mishra, K.P.; Ganju, L. Broad-spectrum antiviral properties of andrographolide. Arch. Virol. 2017, 162, 611–623. [Google Scholar] [CrossRef]
- Mishra, A.; Shaik, H.A.; Sinha, R.K.; Shah, B.R. Andrographolide: A Herbal-Chemosynthetic Approach for Enhancing Immunity, Combating Viral Infections, and Its Implication on Human Health. Molecules 2021, 26, 7036. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Khanom, W.; Sun, X.; Paemanee, A.; Roytrakul, S.; Wang, D.; Smith, D.R.; Zhou, G.C. Andrographolide and Its 14-Aryloxy Analogues Inhibit Zika and Dengue Virus Infection. Molecules 2020, 25, 5037. [Google Scholar] [CrossRef] [PubMed]
- Theerawatanasirikul, S.; Lueangaramkul, V.; Thangthamniyom, N.; Chankeeree, P.; Semkum, P.; Lekcharoensuk, P. Andrographolide and Deoxyandrographolide Inhibit Protease and IFN-Antagonist Activities of Foot-and-Mouth Disease Virus 3C(pro). Animals 2022, 12, 1995. [Google Scholar] [CrossRef] [PubMed]
- Malat, P.; Ekalaksananan, T.; Heawchaiyaphum, C.; Suebsasana, S.; Roytrakul, S.; Yingchutrakul, Y.; Pientong, C. Andrographolide Inhibits Epstein-Barr Virus Lytic Reactivation in EBV-Positive Cancer Cell Lines through the Modulation of Epigenetic-Related Proteins. Molecules 2022, 27, 4666. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Lee, E.M.; Sun, X.; Wang, D.; Tang, H.; Zhou, G.C. Design, synthesis and discovery of andrographolide derivatives against Zika virus infection. Eur. J. Med. Chem. 2020, 187, 111925. [Google Scholar] [CrossRef]
- Chen, J.X.; Xue, H.J.; Ye, W.C.; Fang, B.H.; Liu, Y.H.; Yuan, S.H.; Yu, P.; Wang, Y.Q. Activity of andrographolide and its derivatives against influenza virus in vivo and in vitro. Biol. Pharm. Bull. 2009, 32, 1385–1391. [Google Scholar] [CrossRef]
- Ding, Y.; Chen, L.; Wu, W.; Yang, J.; Yang, Z.; Liu, S. Andrographolide inhibits influenza A virus-induced inflammation in a murine model through NF-κB and JAK-STAT signaling pathway. Microbes Infect. 2017, 19, 605–615. [Google Scholar] [CrossRef]
- Khole, S.; Mittal, S.; Jagadish, N.; Ghosh, D.; Gadgil, V.; Sinkar, V.; Ghaskadbi, S. Andrographolide enhances redox status of liver cells by regulating microRNA expression. Free Radic. Biol. Med. 2019, 130, 397–407. [Google Scholar] [CrossRef]
- Shukla, S.; Gupta, S. Apigenin: A promising molecule for cancer prevention. Pharm. Res. 2010, 27, 962–978. [Google Scholar] [CrossRef] [PubMed]
- Pandey, M.; Kaur, P.; Shukla, S.; Abbas, A.; Fu, P.; Gupta, S. Plant flavone apigenin inhibits HDAC and remodels chromatin to induce growth arrest and apoptosis in human prostate cancer cells: In vitro and in vivo study. Mol. Carcinog. 2012, 51, 952–962. [Google Scholar] [CrossRef]
- Tseng, T.H.; Chien, M.H.; Lin, W.L.; Wen, Y.C.; Chow, J.M.; Chen, C.K.; Kuo, T.C.; Lee, W.J. Inhibition of MDA-MB-231 breast cancer cell proliferation and tumor growth by apigenin through induction of G2/M arrest and histone H3 acetylation-mediated p21(WAF1/CIP1) expression. Environ. Toxicol. 2017, 32, 434–444. [Google Scholar] [CrossRef]
- Husain, K.; Villalobos-Ayala, K.; Laverde, V.; Vazquez, O.A.; Miller, B.; Kazim, S.; Blanck, G.; Hibbs, M.L.; Krystal, G.; Elhussin, I.; et al. Apigenin Targets MicroRNA-155, Enhances SHIP-1 Expression, and Augments Anti-Tumor Responses in Pancreatic Cancer. Cancers 2022, 14, 3613. [Google Scholar] [CrossRef] [PubMed]
- Zakaryan, H.; Arabyan, E.; Oo, A.; Zandi, K. Flavonoids: Promising natural compounds against viral infections. Arch. Virol. 2017, 162, 2539–2551. [Google Scholar] [CrossRef] [PubMed]
- Khan, T.; Khan, M.A.; Mashwani, Z.U.; Ullah, N.; Nadhman, A. Therapeutic potential of medicinal plants against COVID-19: The role of antiviral medicinal metabolites. Biocatal. Agric. Biotechnol. 2021, 31, 101890. [Google Scholar] [CrossRef]
- Sharma, R.; Bhattu, M.; Tripathi, A.; Verma, M.; Acevedo, R.; Kumar, P.; Rajput, V.D.; Singh, J. Potential medicinal plants to combat viral infections: A way forward to environmental biotechnology. Environ. Res. 2023, 227, 115725. [Google Scholar] [CrossRef]
- Xu, X.; Miao, J.; Shao, Q.; Gao, Y.; Hong, L. Apigenin suppresses influenza A virus-induced RIG-I activation and viral replication. J. Med. Virol. 2020, 92, 3057–3066. [Google Scholar] [CrossRef]
- Ożarowski, M.; Karpiński, T.M. The Effects of Propolis on Viral Respiratory Diseases. Molecules 2023, 28, 359. [Google Scholar] [CrossRef]
- Chiang, L.C.; Ng, L.T.; Cheng, P.W.; Chiang, W.; Lin, C.C. Antiviral activities of extracts and selected pure constituents of Ocimum basilicum. Clin. Exp. Pharmacol. Physiol. 2005, 32, 811–816. [Google Scholar] [CrossRef]
- Ohno, M.; Shibata, C.; Kishikawa, T.; Yoshikawa, T.; Takata, A.; Kojima, K.; Akanuma, M.; Kang, Y.J.; Yoshida, H.; Otsuka, M.; et al. The flavonoid apigenin improves glucose tolerance through inhibition of microRNA maturation in miRNA103 transgenic mice. Sci. Rep. 2013, 3, 2553. [Google Scholar] [CrossRef] [PubMed]
- Meerson, A.; Khatib, S.; Mahajna, J. Natural Products Targeting Cancer Stem Cells for Augmenting Cancer Therapeutics. Int. J. Mol. Sci. 2021, 22, 13044. [Google Scholar] [CrossRef] [PubMed]
- Tao, Y.; Zhan, S.; Wang, Y.; Zhou, G.; Liang, H.; Chen, X.; Shen, H. Baicalin, the major component of traditional Chinese medicine Scutellaria baicalensis induces colon cancer cell apoptosis through inhibition of oncomiRNAs. Sci. Rep. 2018, 8, 14477. [Google Scholar] [CrossRef]
- Huang, H.; Zhou, W.; Zhu, H.; Zhou, P.; Shi, X. Baicalin benefits the anti-HBV therapy via inhibiting HBV viral RNAs. Toxicol. Appl. Pharmacol. 2017, 323, 36–43. [Google Scholar] [CrossRef]
- Li, B.Q.; Fu, T.; Yan, Y.D.; Baylor, N.W.; Ruscetti, F.W.; Kung, H.F. Inhibition of HIV infection by baicalin--a flavonoid compound purified from Chinese herbal medicine. Cell Mol. Biol. Res. 1993, 39, 119–124. [Google Scholar] [PubMed]
- Zandi, K.; Teoh, B.-T.; Sam, S.-S.; Wong, P.-F.; Mustafa, M.R.; AbuBakar, S. Novel antiviral activity of baicalein against dengue virus. BMC Complement. Altern. Med. 2012, 12, 214. [Google Scholar] [CrossRef]
- Luo, Z.; Kuang, X.-P.; Zhou, Q.-Q.; Yan, C.-Y.; Li, W.; Gong, H.-B.; Kurihara, H.; Li, W.-X.; Li, Y.-F.; He, R.-R. Inhibitory effects of baicalein against herpes simplex virus type 1. Acta Pharm. Sin. B 2020, 10, 2323–2338. [Google Scholar] [CrossRef]
- Yu, X.; Li, H.; Hu, P.; Qing, Y.; Wang, X.; Zhu, M.; Wang, H.; Wang, Z.; Xu, J.; Guo, Q.; et al. Natural HDAC-1/8 inhibitor baicalein exerts therapeutic effect in CBF-AML. Clin. Transl. Med. 2020, 10, e154. [Google Scholar] [CrossRef]
- Jiang, H.; Yao, Q.; An, Y.; Fan, L.; Wang, J.; Li, H. Baicalin suppresses the progression of Type 2 diabetes-induced liver tumor through regulating METTL3/m(6)A/HKDC1 axis and downstream p-JAK2/STAT1/clevaged Capase3 pathway. Phytomedicine 2022, 94, 153823. [Google Scholar] [CrossRef]
- Qi, J.; Li, J.; Bie, B.; Shi, M.; Zhu, M.; Tian, J.; Zhu, K.; Sun, J.; Mu, Y.; Li, Z.; et al. miR-3,178 contributes to the therapeutic action of baicalein against hepatocellular carcinoma cells via modulating HDAC10. Phytother. Res. 2023, 37, 295–309. [Google Scholar] [CrossRef]
- Cicero, A.F.; Baggioni, A. Berberine and Its Role in Chronic Disease. Adv. Exp. Med. Biol. 2016, 928, 27–45. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Liu, S.; Ma, Q.; Xiao, D.; Chen, L. Berberine enhances the AMPK activation and autophagy and mitigates high glucose-induced apoptosis of mouse podocytes. Eur. J. Pharmacol. 2017, 794, 106–114. [Google Scholar] [CrossRef] [PubMed]
- Kedhari Sundaram, M.; Hussain, A.; Haque, S.; Raina, R.; Afroze, N. Quercetin modifies 5′CpG promoter methylation and reactivates various tumor suppressor genes by modulating epigenetic marks in human cervical cancer cells. J. Cell Biochem. 2019, 120, 18357–18369. [Google Scholar] [CrossRef] [PubMed]
- Lou, G.; Liu, Y.; Wu, S.; Xue, J.; Yang, F.; Fu, H.; Zheng, M.; Chen, Z. The p53/miR-34a/SIRT1 Positive Feedback Loop in Quercetin-Induced Apoptosis. Cell Physiol. Biochem. 2015, 35, 2192–2202. [Google Scholar] [CrossRef]
- Lü, Y.; Han, B.; Yu, H.; Cui, Z.; Li, Z.; Wang, J. Berberine regulates the microRNA-21-ITGΒ4-PDCD4 axis and inhibits colon cancer viability. Oncol. Lett. 2018, 15, 5971–5976. [Google Scholar] [CrossRef] [PubMed]
- Warowicka, A.; Nawrot, R.; Goździcka-Józefiak, A. Antiviral activity of berberine. Arch. Virol. 2020, 165, 1935–1945. [Google Scholar] [CrossRef]
- Cui, Y.; Zhang, L.; Hu, D.; Yang, Y. Berberine Inhibits Herpes Simplex Virus 1 Replication in HEK293T Cells. Comput. Math. Methods Med. 2022, 2022, 7137401. [Google Scholar] [CrossRef]
- Hayashi, K.; Minoda, K.; Nagaoka, Y.; Hayashi, T.; Uesato, S. Antiviral activity of berberine and related compounds against human cytomegalovirus. Bioorg Med. Chem. Lett. 2007, 17, 1562–1564. [Google Scholar] [CrossRef]
- Mahata, S.; Bharti, A.C.; Shukla, S.; Tyagi, A.; Husain, S.A.; Das, B.C. Berberine modulates AP-1 activity to suppress HPV transcription and downstream signaling to induce growth arrest and apoptosis in cervical cancer cells. Mol. Cancer 2011, 10, 39. [Google Scholar] [CrossRef]
- Ratanakomol, T.; Roytrakul, S.; Wikan, N.; Smith, D.R. Berberine Inhibits Dengue Virus through Dual Mechanisms. Molecules 2021, 26, 5501. [Google Scholar] [CrossRef]
- Zha, W.; Liang, G.; Xiao, J.; Studer, E.J.; Hylemon, P.B.; Pandak, W.M., Jr.; Wang, G.; Li, X.; Zhou, H. Berberine inhibits HIV protease inhibitor-induced inflammatory response by modulating ER stress signaling pathways in murine macrophages. PLoS ONE 2010, 5, e9069. [Google Scholar] [CrossRef] [PubMed]
- Hung, T.C.; Jassey, A.; Liu, C.H.; Lin, C.J.; Lin, C.C.; Wong, S.H.; Wang, J.Y.; Yen, M.H.; Lin, L.T. Berberine inhibits hepatitis C virus entry by targeting the viral E2 glycoprotein. Phytomedicine 2019, 53, 62–69. [Google Scholar] [CrossRef] [PubMed]
- Pizzorno, A.; Padey, B.; Dubois, J.; Julien, T.; Traversier, A.; Dulière, V.; Brun, P.; Lina, B.; Rosa-Calatrava, M.; Terrier, O. In vitro evaluation of antiviral activity of single and combined repurposable drugs against SARS-CoV-2. Antivir. Res. 2020, 181, 104878. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Li, J.Q.; Kim, Y.J.; Wu, J.; Wang, Q.; Hao, Y. In vivo and in vitro antiviral effects of berberine on influenza virus. Chin. J. Integr. Med. 2011, 17, 444–452. [Google Scholar] [CrossRef] [PubMed]
- Varghese, F.S.; Thaa, B.; Amrun, S.N.; Simarmata, D.; Rausalu, K.; Nyman, T.A.; Merits, A.; McInerney, G.M.; Ng, L.F.P.; Ahola, T. The Antiviral Alkaloid Berberine Reduces Chikungunya Virus-Induced Mitogen-Activated Protein Kinase Signaling. J. Virol. 2016, 90, 9743–9757. [Google Scholar] [CrossRef]
- Kim, S.Y.; Kim, H.; Kim, S.W.; Lee, N.R.; Yi, C.M.; Heo, J.; Kim, B.J.; Kim, N.J.; Inn, K.S. An Effective Antiviral Approach Targeting Hepatitis B Virus with NJK14047, a Novel and Selective Biphenyl Amide p38 Mitogen-Activated Protein Kinase Inhibitor. Antimicrob. Agents Chemother. 2017, 61, e00214-17. [Google Scholar] [CrossRef]
- Zuco, V.; Supino, R.; Righetti, S.C.; Cleris, L.; Marchesi, E.; Gambacorti-Passerini, C.; Formelli, F. Selective cytotoxicity of betulinic acid on tumor cell lines, but not on normal cells. Cancer Lett. 2002, 175, 17–25. [Google Scholar] [CrossRef]
- Fulda, S. Betulinic Acid for cancer treatment and prevention. Int. J. Mol. Sci. 2008, 9, 1096–1107. [Google Scholar] [CrossRef]
- Adewole, K.E.; Ishola, A.A. A Computational Approach to Investigate the HDAC6 and HDAC10 Binding Propensity of Psidium guajava-derived Compounds as Potential Anticancer Agents. Curr. Drug Discov. Technol. 2021, 18, 423–436. [Google Scholar] [CrossRef]
- Liu, X.; Jutooru, I.; Lei, P.; Kim, K.; Lee, S.O.; Brents, L.K.; Prather, P.L.; Safe, S. Betulinic acid targets YY1 and ErbB2 through cannabinoid receptor-dependent disruption of microRNA-27a:ZBTB10 in breast cancer. Mol. Cancer Ther. 2012, 11, 1421–1431. [Google Scholar] [CrossRef]
- Peyrat, L.A.; Eparvier, V.; Eydoux, C.; Guillemot, J.C.; Litaudon, M.; Stien, D. Betulinic Acid, The First Lupane-Type Triterpenoid Isolated from Both a Phomopsis sp. and Its Host Plant Diospyros carbonaria Benoist. Chem. Biodivers. 2017, 14, e1600171. [Google Scholar] [CrossRef] [PubMed]
- Yao, D.; Li, H.; Gou, Y.; Zhang, H.; Vlessidis, A.G.; Zhou, H.; Evmiridis, N.P.; Liu, Z. Betulinic acid-mediated inhibitory effect on hepatitis B virus by suppression of manganese superoxide dismutase expression. Febs J. 2009, 276, 2599–2614. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, P.; Gupta, K.P. Modulation of miR-203 and its regulators as a function of time during the development of 7, 12 dimethylbenz [a] anthracene induced mouse skin tumors in presence or absence of the antitumor agents. Toxicol. Appl. Pharmacol. 2014, 278, 148–158. [Google Scholar] [CrossRef]
- Pant, K.; Mishra, A.K.; Pradhan, S.M.; Nayak, B.; Das, P.; Shalimar, D.; Saraya, A.; Venugopal, S.K. Butyrate inhibits HBV replication and HBV-induced hepatoma cell proliferation via modulating SIRT-1/Ac-p53 regulatory axis. Mol. Carcinog. 2019, 58, 524–532. [Google Scholar] [CrossRef] [PubMed]
- Voon, F.L.; Sulaiman, M.R.; Akhtar, M.N.; Idris, M.F.; Akira, A.; Perimal, E.K.; Israf, D.A.; Ming-Tatt, L. Cardamonin (2′,4′-dihydroxy-6′-methoxychalcone) isolated from Boesenbergia rotunda (L.) Mansf. inhibits CFA-induced rheumatoid arthritis in rats. Eur. J. Pharmacol. 2017, 794, 127–134. [Google Scholar] [CrossRef]
- Jin, Y.H.; Min, J.S.; Kwon, S. Cardamonin as a p38 MAPK Signaling Pathway Activator Inhibits Human Coronavirus OC43 Infection in Human Lung Cells. Nutrients 2023, 15, 1335. [Google Scholar] [CrossRef]
- Das, S.K.; Masuda, M.; Sakurai, A.; Sakakibara, M. Medicinal uses of the mushroom Cordyceps militaris: Current state and prospects. Fitoterapia 2010, 81, 961–968. [Google Scholar] [CrossRef]
- Ryu, E.; Son, M.; Lee, M.; Lee, K.; Cho, J.Y.; Cho, S.; Lee, S.K.; Lee, Y.M.; Cho, H.; Sung, G.H.; et al. Cordycepin is a novel chemical suppressor of Epstein-Barr virus replication. Oncoscience 2014, 1, 866–881. [Google Scholar] [CrossRef]
- Rabie, A.M. Potent Inhibitory Activities of the Adenosine Analogue Cordycepin on SARS-CoV-2 Replication. ACS Omega 2022, 7, 2960–2969. [Google Scholar] [CrossRef]
- Ling, J.Y.; Sun, Y.J.; Zhang, H.; Lv, P.; Zhang, C.K. Measurement of cordycepin and adenosine in stroma of Cordyceps sp. by capillary zone electrophoresis (CZE). J. Biosci. Bioeng. 2002, 94, 371–374. [Google Scholar] [CrossRef]
- Rose, K.M.; Bell, L.E.; Jacob, S.T. Specific inhibition of chromatin-associated poly(A) synthesis in vitro by cordycepin 5′-triphosphate. Nature 1977, 267, 178–180. [Google Scholar] [CrossRef] [PubMed]
- Mahy, B.W.; Cox, N.J.; Armstrong, S.J.; Barry, R.D. Multiplication of influenza virus in the presence of cordycepin, an inhibitor of cellular RNA synthesis. Nat. New Biol. 1973, 243, 172–174. [Google Scholar] [CrossRef]
- Müller, W.E.; Weiler, B.E.; Charubala, R.; Pfleiderer, W.; Leserman, L.; Sobol, R.W.; Suhadolnik, R.J.; Schröder, H.C. Cordycepin analogues of 2′,5′-oligoadenylate inhibit human immunodeficiency virus infection via inhibition of reverse transcriptase. Biochemistry 1991, 30, 2027–2033. [Google Scholar] [CrossRef]
- Lonai, P.; Declève, A.; Kaplan, H.S. Spontaneous induction of endogenous murine leukemia virus-related antigen expression during short-term in vitro incubation of mouse lymphocytes. Proc. Natl. Acad. Sci. USA 1974, 71, 2008–2012. [Google Scholar] [CrossRef]
- Doetsch, P.W.; Suhadolnik, R.J.; Sawada, Y.; Mosca, J.D.; Flick, M.B.; Reichenbach, N.L.; Dang, A.Q.; Wu, J.M.; Charubala, R.; Pfleiderer, W.; et al. Core (2′–5′)oligoadenylate and the cordycepin analog: Inhibitors of Epstein--Barr virus-induced transformation of human lymphocytes in the absence of interferon. Proc. Natl. Acad. Sci. USA 1981, 78, 6699–6703. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, N.; Yang, L.; Song, X.Q. Bioactive natural products in COVID-19 therapy. Front. Pharmacol. 2022, 13, 926507. [Google Scholar] [CrossRef] [PubMed]
- Hou, W.; Li, Y.; Zhang, Q.; Wei, X.; Peng, A.; Chen, L.; Wei, Y. Triterpene acids isolated from Lagerstroemia speciosa leaves as alpha-glucosidase inhibitors. Phytother. Res. 2009, 23, 614–618. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhou, H.; An, Y.; Shen, K.; Yu, L. Biological effects of corosolic acid as an anti-inflammatory, anti-metabolic syndrome and anti-neoplasic natural compound. Oncol. Lett. 2021, 21, 84. [Google Scholar] [CrossRef]
- Hudlikar, R.R.; Sargsyan, D.; Wu, R.; Su, S.; Zheng, M.; Kong, A.N. Triterpenoid corosolic acid modulates global CpG methylation and transcriptome of tumor promotor TPA induced mouse epidermal JB6 P+ cells. Chem. Biol. Interact. 2020, 321, 109025. [Google Scholar] [CrossRef] [PubMed]
- Ratovitski, E.A. Anticancer Natural Compounds as Epigenetic Modulators of Gene Expression. Curr. Genom. 2017, 18, 175–205. [Google Scholar] [CrossRef]
- Yang, J.; Wu, R.; Li, W.; Gao, L.; Yang, Y.; Li, P.; Kong, A.N. The triterpenoid corosolic acid blocks transformation and epigenetically reactivates Nrf2 in TRAMP-C1 prostate cells. Mol. Carcinog. 2018, 57, 512–521. [Google Scholar] [CrossRef] [PubMed]
- Hewlings, S.J.; Kalman, D.S. Curcumin: A Review of Its Effects on Human Health. Foods 2017, 6, 92. [Google Scholar] [CrossRef]
- Liu, H.L.; Chen, Y.; Cui, G.H.; Zhou, J.F. Curcumin, a potent anti-tumor reagent, is a novel histone deacetylase inhibitor regulating B-NHL cell line Raji proliferation. Acta Pharmacol. Sin. 2005, 26, 603–609. [Google Scholar] [CrossRef] [PubMed]
- Shu, L.; Khor, T.O.; Lee, J.H.; Boyanapalli, S.S.; Huang, Y.; Wu, T.Y.; Saw, C.L.; Cheung, K.L.; Kong, A.N. Epigenetic CpG demethylation of the promoter and reactivation of the expression of Neurog1 by curcumin in prostate LNCaP cells. Aaps J. 2011, 13, 606–614. [Google Scholar] [CrossRef]
- Wei, Z.Q.; Zhang, Y.H.; Ke, C.Z.; Chen, H.X.; Ren, P.; He, Y.L.; Hu, P.; Ma, D.Q.; Luo, J.; Meng, Z.J. Curcumin inhibits hepatitis B virus infection by down-regulating cccDNA-bound histone acetylation. World J. Gastroenterol. 2017, 23, 6252–6260. [Google Scholar] [CrossRef]
- Sohn, E.J.; Bak, K.M.; Nam, Y.K.; Park, H.T. Upregulation of microRNA 344a-3p is involved in curcumin induced apoptosis in RT4 schwannoma cells. Cancer Cell Int. 2018, 18, 199. [Google Scholar] [CrossRef] [PubMed]
- Hassan, F.U.; Rehman, M.S.; Khan, M.S.; Ali, M.A.; Javed, A.; Nawaz, A.; Yang, C. Curcumin as an Alternative Epigenetic Modulator: Mechanism of Action and Potential Effects. Front. Genet. 2019, 10, 514. [Google Scholar] [CrossRef] [PubMed]
- Butnariu, M.; Quispe, C.; Koirala, N.; Khadka, S.; Salgado-Castillo, C.M.; Akram, M.; Anum, R.; Yeskaliyeva, B.; Cruz-Martins, N.; Martorell, M.; et al. Bioactive Effects of Curcumin in Human Immunodeficiency Virus Infection Along with the Most Effective Isolation Techniques and Type of Nanoformulations. Int. J. Nanomed. 2022, 17, 3619–3632. [Google Scholar] [CrossRef]
- Chen, D.-Y.; Shien, J.-H.; Tiley, L.; Chiou, S.-S.; Wang, S.-Y.; Chang, T.-J.; Lee, Y.-J.; Chan, K.-W.; Hsu, W.-L. Curcumin inhibits influenza virus infection and haemagglutination activity. Food Chem. 2010, 119, 1346–1351. [Google Scholar] [CrossRef]
- Kim, H.J.; Yoo, H.S.; Kim, J.C.; Park, C.S.; Choi, M.S.; Kim, M.; Choi, H.; Min, J.S.; Kim, Y.S.; Yoon, S.W.; et al. Antiviral effect of Curcuma longa Linn extract against hepatitis B virus replication. J. Ethnopharmacol. 2009, 124, 189–196. [Google Scholar] [CrossRef]
- Prusty, B.K.; Das, B.C. Constitutive activation of transcription factor AP-1 in cervical cancer and suppression of human papillomavirus (HPV) transcription and AP-1 activity in HeLa cells by curcumin. Int. J. Cancer 2005, 113, 951–960. [Google Scholar] [CrossRef]
- Zahedipour, F.; Hosseini, S.A.; Sathyapalan, T.; Majeed, M.; Jamialahmadi, T.; Al-Rasadi, K.; Banach, M.; Sahebkar, A. Potential effects of curcumin in the treatment of COVID-19 infection. Phytother. Res. 2020, 34, 2911–2920. [Google Scholar] [CrossRef]
- Jennings, M.R.; Parks, R.J. Curcumin as an Antiviral Agent. Viruses 2020, 12, 1242. [Google Scholar] [CrossRef] [PubMed]
- Sordillo, P.P.; Helson, L. Curcumin suppression of cytokine release and cytokine storm. A potential therapy for patients with Ebola and other severe viral infections. Vivo 2015, 29, 1–4. [Google Scholar]
- Wen, C.C.; Kuo, Y.H.; Jan, J.T.; Liang, P.H.; Wang, S.Y.; Liu, H.G.; Lee, C.K.; Chang, S.T.; Kuo, C.J.; Lee, S.S.; et al. Specific plant terpenoids and lignoids possess potent antiviral activities against severe acute respiratory syndrome coronavirus. J. Med. Chem. 2007, 50, 4087–4095. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Song, X.Q.; Xu, W.; Lei, S.; Zhang, H.; Yang, L. Stand Up to Stand Out: Natural Dietary Polyphenols Curcumin, Resveratrol, and Gossypol as Potential Therapeutic Candidates against Severe Acute Respiratory Syndrome Coronavirus 2 Infection. Nutrients 2023, 15, 3885. [Google Scholar] [CrossRef] [PubMed]
- Sharifi-Rad, J.; Quispe, C.; Castillo, C.M.S.; Caroca, R.; Lazo-Vélez, M.A.; Antonyak, H.; Polishchuk, A.; Lysiuk, R.; Oliinyk, P.; De Masi, L.; et al. Ellagic Acid: A Review on Its Natural Sources, Chemical Stability, and Therapeutic Potential. Oxid. Med. Cell Longev. 2022, 2022, 3848084. [Google Scholar] [CrossRef]
- Kang, I.; Okla, M.; Chung, S. Ellagic acid inhibits adipocyte differentiation through coactivator-associated arginine methyltransferase 1-mediated chromatin modification. J. Nutr. Biochem. 2014, 25, 946–953. [Google Scholar] [CrossRef]
- Dinata, R.; Nisa, N.; Arati, C.; Rasmita, B.; Uditraj, C.; Siddhartha, R.; Bhanushree, B.; Saeed-Ahmed, L.; Manikandan, B.; Bidanchi, R.M.; et al. Repurposing immune boosting and anti-viral efficacy of Parkia bioactive entities as multi-target directed therapeutic approach for SARS-CoV-2: Exploration of lead drugs by drug likeness, molecular docking and molecular dynamics simulation methods. J. Biomol. Struct. Dyn. 2023, 1–39. [Google Scholar] [CrossRef]
- Prasanth, M.I.; Sivamaruthi, B.S.; Chaiyasut, C.; Tencomnao, T. A Review of the Role of Green Tea (Camellia sinensis) in Antiphotoaging, Stress Resistance, Neuroprotection, and Autophagy. Nutrients 2019, 11, 474. [Google Scholar] [CrossRef]
- Ngwe Tun, M.M.; Luvai, E.; Nwe, K.M.; Toume, K.; Mizukami, S.; Hirayama, K.; Komatsu, K.; Morita, K. Anti-SARS-CoV-2 activity of various PET-bottled Japanese green teas and tea compounds in vitro. Arch. Virol. 2022, 167, 1547–1557. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Z.; Zhang, Z.; Han, Y.; Wang, J.; Wang, Y.; Chen, X.; Shao, Y.; Cheng, Y.; Zhou, W.; Lu, X.; et al. A review on anti-cancer effect of green tea catechins. J. Funct. Foods 2020, 74, 104172. [Google Scholar] [CrossRef]
- Li, F.; Qasim, S.; Li, D.; Dou, Q.P. Updated review on green tea polyphenol epigallocatechin-3-gallate as a cancer epigenetic regulator. Semin. Cancer Biol. 2022, 83, 335–352. [Google Scholar] [CrossRef] [PubMed]
- Bag, A.; Bag, N. Tea Polyphenols and Prevention of Epigenetic Aberrations in Cancer. J. Nat. Sci. Biol. Med. 2018, 9, 2–5. [Google Scholar] [CrossRef] [PubMed]
- Henning, S.M.; Wang, P.; Carpenter, C.L.; Heber, D. Epigenetic effects of green tea polyphenols in cancer. Epigenomics 2013, 5, 729–741. [Google Scholar] [CrossRef]
- Sun, Q.; Huang, M.; Wei, Y. Diversity of the reaction mechanisms of SAM-dependent enzymes. Acta Pharm. Sin. B 2021, 11, 632–650. [Google Scholar] [CrossRef]
- Moseley, V.R.; Morris, J.; Knackstedt, R.W.; Wargovich, M.J. Green tea polyphenol epigallocatechin 3-gallate, contributes to the degradation of DNMT3A and HDAC3 in HCT 116 human colon cancer cells. Anticancer. Res. 2013, 33, 5325–5333. [Google Scholar]
- Meeran, S.M.; Patel, S.N.; Chan, T.H.; Tollefsbol, T.O. A novel prodrug of epigallocatechin-3-gallate: Differential epigenetic hTERT repression in human breast cancer cells. Cancer Prev. Res. 2011, 4, 1243–1254. [Google Scholar] [CrossRef]
- Nandakumar, V.; Vaid, M.; Katiyar, S.K. (−)-Epigallocatechin-3-gallate reactivates silenced tumor suppressor genes, Cip1/p21 and p16INK4a, by reducing DNA methylation and increasing histones acetylation in human skin cancer cells. Carcinogenesis 2011, 32, 537–544. [Google Scholar] [CrossRef]
- Ciesielski, O.; Biesiekierska, M.; Balcerczyk, A. Epigallocatechin-3-gallate (EGCG) Alters Histone Acetylation and Methylation and Impacts Chromatin Architecture Profile in Human Endothelial Cells. Molecules 2020, 25, 2326. [Google Scholar] [CrossRef]
- Fang, M.Z.; Wang, Y.; Ai, N.; Hou, Z.; Sun, Y.; Lu, H.; Welsh, W.; Yang, C.S. Tea polyphenol (−)-epigallocatechin-3-gallate inhibits DNA methyltransferase and reactivates methylation-silenced genes in cancer cell lines. Cancer Res. 2003, 63, 7563–7570. [Google Scholar] [PubMed]
- Guo, C.; Huang, Q.; Wang, Y.; Yao, Y.; Li, J.; Chen, J.; Wu, M.; Zhang, Z.; E, M.; Qi, H.; et al. Therapeutic application of natural products: NAD+ metabolism as potential target. Phytomedicine 2023, 114, 154768. [Google Scholar] [CrossRef]
- Rasheed, Z.; Rasheed, N.; Al-Shaya, O. Epigallocatechin-3-O-gallate modulates global microRNA expression in interleukin-1β-stimulated human osteoarthritis chondrocytes: Potential role of EGCG on negative co-regulation of microRNA-140-3p and ADAMTS5. Eur. J. Nutr. 2018, 57, 917–928. [Google Scholar] [CrossRef]
- Shin, S.; Kim, K.; Lee, M.J.; Lee, J.; Choi, S.; Kim, K.S.; Ko, J.M.; Han, H.; Kim, S.Y.; Youn, H.J.; et al. Epigallocatechin Gallate-Mediated Alteration of the MicroRNA Expression Profile in 5α-Dihydrotestosterone-Treated Human Dermal Papilla Cells. Ann. Dermatol. 2016, 28, 327–334. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Chang, L.; Qu, Y.; Liang, J.; Jin, W.; Xia, X. Tea polyphenols inhibit the proliferation, migration, and invasion of melanoma cells through the down-regulation of TLR4. Int. J. Immunopathol. Pharmacol. 2018, 32, 394632017739531. [Google Scholar] [CrossRef] [PubMed]
- Arffa, M.L.; Zapf, M.A.; Kothari, A.N.; Chang, V.; Gupta, G.N.; Ding, X.; Al-Gayyar, M.M.; Syn, W.; Elsherbiny, N.M.; Kuo, P.C.; et al. Epigallocatechin-3-Gallate Upregulates miR-221 to Inhibit Osteopontin-Dependent Hepatic Fibrosis. PLoS ONE 2016, 11, e0167435. [Google Scholar] [CrossRef] [PubMed]
- Steinmann, J.; Buer, J.; Pietschmann, T.; Steinmann, E. Anti-infective properties of epigallocatechin-3-gallate (EGCG), a component of green tea. Br. J. Pharmacol. 2013, 168, 1059–1073. [Google Scholar] [CrossRef]
- Tsvetkov, V.; Varizhuk, A.; Kozlovskaya, L.; Shtro, A.; Lebedeva, O.; Komissarov, A.; Vedekhina, T.; Manuvera, V.; Zubkova, O.; Eremeev, A.; et al. EGCG as an anti-SARS-CoV-2 agent: Preventive versus therapeutic potential against original and mutant virus. Biochimie 2021, 191, 27–32. [Google Scholar] [CrossRef]
- da Silva-Júnior, E.F.; Silva, L.R. Multi-target Approaches of Epigallocatechin-3-O-gallate (EGCG) and its Derivatives against Influenza Viruses. Curr. Top. Med. Chem. 2022, 22, 1485–1500. [Google Scholar] [CrossRef]
- Miyoshi, N.; Tanabe, H.; Suzuki, T.; Saeki, K.; Hara, Y. Applications of a Standardized Green Tea Catechin Preparation for Viral Warts and Human Papilloma Virus-Related and Unrelated Cancers. Molecules 2020, 25, 2588. [Google Scholar] [CrossRef]
- Mekky, R.Y.; El-Ekiaby, N.; El Sobky, S.A.; Elemam, N.M.; Youness, R.A.; El-Sayed, M.; Hamza, M.T.; Esmat, G.; Abdelaziz, A.I. Epigallocatechin gallate (EGCG) and miR-548m reduce HCV entry through repression of CD81 receptor in HCV cell models. Arch. Virol. 2019, 164, 1587–1595. [Google Scholar] [CrossRef] [PubMed]
- Date, A.A.; Destache, C.J. Natural polyphenols: Potential in the prevention of sexually transmitted viral infections. Drug Discov. Today 2016, 21, 333–341. [Google Scholar] [CrossRef]
- Shorobi, F.M.; Nisa, F.Y.; Saha, S.; Chowdhury, M.A.H.; Srisuphanunt, M.; Hossain, K.H.; Rahman, M.A. Quercetin: A Functional Food-Flavonoid Incredibly Attenuates Emerging and Re-Emerging Viral Infections through Immunomodulatory Actions. Molecules 2023, 28, 938. [Google Scholar] [CrossRef] [PubMed]
- Carneiro, B.M.; Batista, M.N.; Braga, A.C.S.; Nogueira, M.L.; Rahal, P. The green tea molecule EGCG inhibits Zika virus entry. Virology 2016, 496, 215–218. [Google Scholar] [CrossRef]
- Ahn, W.S.; Huh, S.W.; Bae, S.M.; Lee, I.P.; Lee, J.M.; Namkoong, S.E.; Kim, C.K.; Sin, J.I. A major constituent of green tea, EGCG, inhibits the growth of a human cervical cancer cell line, CaSki cells, through apoptosis, G(1) arrest, and regulation of gene expression. DNA Cell Biol. 2003, 22, 217–224. [Google Scholar] [CrossRef] [PubMed]
- Stockfleth, E.; Meyer, T. Sinecatechins (Polyphenon E) ointment for treatment of external genital warts and possible future indications. Expert. Opin. Biol. Ther. 2014, 14, 1033–1043. [Google Scholar] [CrossRef] [PubMed]
- Mekky, R.Y.; El-Ekiaby, N.M.; Hamza, M.T.; Elemam, N.M.; El-Sayed, M.; Esmat, G.; Abdelaziz, A.I. Mir-194 is a hepatocyte gate keeper hindering HCV entry through targeting CD81 receptor. J. Infect. 2015, 70, 78–87. [Google Scholar] [CrossRef]
- Kong, Y.; Feng, Z.; Chen, A.; Qi, Q.; Han, M.; Wang, S.; Zhang, Y.; Zhang, X.; Yang, N.; Wang, J.; et al. The Natural Flavonoid Galangin Elicits Apoptosis, Pyroptosis, and Autophagy in Glioblastoma. Front. Oncol. 2019, 9, 942. [Google Scholar] [CrossRef]
- Zeng, H.; Huang, P.; Wang, X.; Wu, J.; Wu, M.; Huang, J. Galangin-induced down-regulation of BACE1 by epigenetic mechanisms in SH-SY5Y cells. Neuroscience 2015, 294, 172–181. [Google Scholar] [CrossRef]
- Deng, X.; Zuo, M.; Pei, Z.; Xie, Y.; Yang, Z.; Zhang, Z.; Jiang, M.; Kuang, D. MicroRNA-455-5p Contributes to Cholangiocarcinoma Growth and Mediates Galangin’s Anti-Tumor Effects. J. Cancer 2021, 12, 4710–4721. [Google Scholar] [CrossRef]
- Meyer, J.J.; Afolayan, A.J.; Taylor, M.B.; Erasmus, D. Antiviral activity of galangin isolated from the aerial parts of Helichrysum aureonitens. J. Ethnopharmacol. 1997, 56, 165–169. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, V.; Tuli, H.S.; Kaur, J.; Aggarwal, D.; Parashar, G.; Chaturvedi Parashar, N.; Kulkarni, S.; Kaur, G.; Sak, K.; Kumar, M.; et al. Garcinol Exhibits Anti-Neoplastic Effects by Targeting Diverse Oncogenic Factors in Tumor Cells. Biomedicines 2020, 8, 103. [Google Scholar] [CrossRef]
- Kopytko, P.; Piotrowska, K.; Janisiak, J.; Tarnowski, M. Garcinol-A Natural Histone Acetyltransferase Inhibitor and New Anti-Cancer Epigenetic Drug. Int. J. Mol. Sci. 2021, 22, 2828. [Google Scholar] [CrossRef] [PubMed]
- Behera, A.K.; Swamy, M.M.; Natesh, N.; Kundu, T.K. Garcinol and Its Role in Chronic Diseases. Adv. Exp. Med. Biol. 2016, 928, 435–452. [Google Scholar] [PubMed]
- Kim, J.-Y.; Jo, J.; Leem, J.; Park, K.-K. Inhibition of p300 by Garcinol Protects against Cisplatin-Induced Acute Kidney Injury through Suppression of Oxidative Stress, Inflammation, and Tubular Cell Death in Mice. Antioxidants 2020, 9, 1271. [Google Scholar] [CrossRef] [PubMed]
- Balasubramanyam, K.; Altaf, M.; Varier, R.A.; Swaminathan, V.; Ravindran, A.; Sadhale, P.P.; Kundu, T.K. Polyisoprenylated benzophenone, garcinol, a natural histone acetyltransferase inhibitor, represses chromatin transcription and alters global gene expression. J. Biol. Chem. 2004, 279, 33716–33726. [Google Scholar] [CrossRef]
- Padhye, S.; Ahmad, A.; Oswal, N.; Sarkar, F.H. Emerging role of Garcinol, the antioxidant chalcone from Garcinia indica Choisy and its synthetic analogs. J. Hematol. Oncol. 2009, 2, 38. [Google Scholar] [CrossRef]
- Saadat, N.; Gupta, S.V. Potential role of garcinol as an anticancer agent. J. Oncol. 2012, 2012, 647206. [Google Scholar] [CrossRef]
- Parasramka, M.A.; Ali, S.; Banerjee, S.; Deryavoush, T.; Sarkar, F.H.; Gupta, S. Garcinol sensitizes human pancreatic adenocarcinoma cells to gemcitabine in association with microRNA signatures. Mol. Nutr. Food Res. 2013, 57, 235–248. [Google Scholar] [CrossRef]
- Biersack, B. Current state of phenolic and terpenoidal dietary factors and natural products as non-coding RNA/microRNA modulators for improved cancer therapy and prevention. Noncoding RNA Res. 2016, 1, 12–34. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, A.; Sarkar, S.H.; Bitar, B.; Ali, S.; Aboukameel, A.; Sethi, S.; Li, Y.; Bao, B.; Kong, D.; Banerjee, S.; et al. Garcinol regulates EMT and Wnt signaling pathways in vitro and in vivo, leading to anticancer activity against breast cancer cells. Mol. Cancer Ther. 2012, 11, 2193–2201. [Google Scholar] [CrossRef] [PubMed]
- Corona, A.; Seibt, S.; Schaller, D.; Schobert, R.; Volkamer, A.; Biersack, B.; Tramontano, E. Garcinol from Garcinia indica inhibits HIV-1 reverse transcriptase-associated ribonuclease H. Arch. Pharm. 2021, 354, e2100123. [Google Scholar] [CrossRef] [PubMed]
- Sharifi-Rad, J.; Quispe, C.; Imran, M.; Rauf, A.; Nadeem, M.; Gondal, T.A.; Ahmad, B.; Atif, M.; Mubarak, M.S.; Sytar, O.; et al. Genistein: An Integrative Overview of Its Mode of Action, Pharmacological Properties, and Health Benefits. Oxid. Med. Cell Longev. 2021, 2021, 3268136. [Google Scholar] [CrossRef] [PubMed]
- Fang, M.Z.; Chen, D.; Sun, Y.; Jin, Z.; Christman, J.K.; Yang, C.S. Reversal of hypermethylation and reactivation of p16INK4a, RARbeta, and MGMT genes by genistein and other isoflavones from soy. Clin. Cancer Res. 2005, 11, 7033–7041. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, Q.; Chen, H. DNA methylation and histone modifications of Wnt genes by genistein during colon cancer development. Carcinogenesis 2013, 34, 1756–1763. [Google Scholar] [CrossRef]
- Ma, J.; Cheng, L.; Liu, H.; Zhang, J.; Shi, Y.; Zeng, F.; Miele, L.; Sarkar, F.H.; Xia, J.; Wang, Z. Genistein down-regulates miR-223 expression in pancreatic cancer cells. Curr. Drug Targets 2013, 14, 1150–1156. [Google Scholar] [CrossRef]
- Gerstmeier, J.; Seegers, J.; Witt, F.; Waltenberger, B.; Temml, V.; Rollinger, J.M.; Stuppner, H.; Koeberle, A.; Schuster, D.; Werz, O. Ginkgolic Acid is a Multi-Target Inhibitor of Key Enzymes in Pro-Inflammatory Lipid Mediator Biosynthesis. Front. Pharmacol. 2019, 10, 797. [Google Scholar] [CrossRef]
- Fukuda, I.; Ito, A.; Hirai, G.; Nishimura, S.; Kawasaki, H.; Saitoh, H.; Kimura, K.; Sodeoka, M.; Yoshida, M. Ginkgolic acid inhibits protein SUMOylation by blocking formation of the E1-SUMO intermediate. Chem. Biol. 2009, 16, 133–140. [Google Scholar] [CrossRef]
- Flotho, A.; Melchior, F. Sumoylation: A regulatory protein modification in health and disease. Annu. Rev. Biochem. 2013, 82, 357–385. [Google Scholar] [CrossRef]
- Yang, W.S.; Campbell, M.; Chang, P.C. SUMO modification of a heterochromatin histone demethylase JMJD2A enables viral gene transactivation and viral replication. PLoS Pathog. 2017, 13, e1006216. [Google Scholar] [CrossRef]
- Bhutta, M.S.; Shechter, O.; Gallo, E.S.; Martin, S.D.; Jones, E.; Doncel, G.F.; Borenstein, R. Ginkgolic Acid Inhibits Herpes Simplex Virus Type 1 Skin Infection and Prevents Zosteriform Spread in Mice. Viruses 2021, 13, 86. [Google Scholar] [CrossRef] [PubMed]
- Lü, J.M.; Yan, S.; Jamaluddin, S.; Weakley, S.M.; Liang, Z.; Siwak, E.B.; Yao, Q.; Chen, C. Ginkgolic acid inhibits HIV protease activity and HIV infection in vitro. Med. Sci. Monit. 2012, 18, Br293–Br298. [Google Scholar] [CrossRef] [PubMed]
- Pastorino, G.; Cornara, L.; Soares, S.; Rodrigues, F.; Oliveira, M. Liquorice (Glycyrrhiza glabra): A phytochemical and pharmacological review. Phytother. Res. 2018, 32, 2323–2339. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.G.; Zhao, T.T.; Lu, N.; Yang, Y.A.; Zhu, H.L. Research Progress of Glycyrrhizic Acid on Antiviral Activity. Mini Rev. Med. Chem. 2019, 19, 826–832. [Google Scholar] [CrossRef]
- Cao, L.; Ding, W.; Jia, R.; Du, J.; Wang, T.; Zhang, C.; Gu, Z.; Yin, G. Anti-inflammatory and hepatoprotective effects of glycyrrhetinic acid on CCl4-induced damage in precision-cut liver slices from Jian carp (Cyprinus carpio var. jian) through inhibition of the nf-kB pathway. Fish Shellfish Immunol. 2017, 64, 234–242. [Google Scholar] [CrossRef]
- Zhong, L.L.D.; Lam, W.C.; Yang, W.; Chan, K.W.; Sze, S.C.W.; Miao, J.; Yung, K.K.L.; Bian, Z.; Wong, V.T. Potential Targets for Treatment of Coronavirus Disease 2019 (COVID-19): A Review of Qing-Fei-Pai-Du-Tang and Its Major Herbs. Am. J. Chin. Med. 2020, 48, 1051–1071. [Google Scholar] [CrossRef]
- Omer, M.O.; Almalki, W.H.; Shahid, I.; Khuram, S.; Altaf, I.; Imran, S. Comparative study to evaluate the anti-viral efficacy of Glycyrrhiza glabra extract and ribavirin against the Newcastle disease virus. Pharmacogn. Res. 2014, 6, 6–11. [Google Scholar] [CrossRef]
- Cinatl, J.; Morgenstern, B.; Bauer, G.; Chandra, P.; Rabenau, H.; Doerr, H.W. Glycyrrhizin, an active component of liquorice roots, and replication of SARS-associated coronavirus. Lancet 2003, 361, 2045–2046. [Google Scholar] [CrossRef]
- Fiore, C.; Eisenhut, M.; Krausse, R.; Ragazzi, E.; Pellati, D.; Armanini, D.; Bielenberg, J. Antiviral effects of Glycyrrhiza species. Phytother. Res. 2008, 22, 141–148. [Google Scholar] [CrossRef]
- Chen, F.; Chan, K.H.; Jiang, Y.; Kao, R.Y.; Lu, H.T.; Fan, K.W.; Cheng, V.C.; Tsui, W.H.; Hung, I.F.; Lee, T.S.; et al. In vitro susceptibility of 10 clinical isolates of SARS coronavirus to selected antiviral compounds. J. Clin. Virol. 2004, 31, 69–75. [Google Scholar] [CrossRef]
- Yang, R.; Liu, H.; Bai, C.; Wang, Y.; Zhang, X.; Guo, R.; Wu, S.; Wang, J.; Leung, E.; Chang, H.; et al. Chemical composition and pharmacological mechanism of Qingfei Paidu Decoction and Ma Xing Shi Gan Decoction against Coronavirus Disease 2019 (COVID-19): In silico and experimental study. Pharmacol. Res. 2020, 157, 104820. [Google Scholar] [CrossRef]
- Yue, G.G.; Chan, B.C.; Kwok, H.F.; To, M.H.; Hon, K.L.; Fung, K.P.; Lau, C.B.; Leung, P.C. Screening for anti-inflammatory and bronchorelaxant activities of 12 commonly used Chinese herbal medicines. Phytother. Res. 2012, 26, 915–925. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Huang, S.; Li, Y.; Li, Y.; Li, D.; Wu, P.; Wang, Q.; Zheng, X.; Zhang, K. Glycyrrhizic acid from licorice down-regulates inflammatory responses via blocking MAPK and PI3K/Akt-dependent NF-κB signalling pathways in TPA-induced skin inflammation. Medchemcomm 2018, 9, 1502–1510. [Google Scholar] [CrossRef] [PubMed]
- Kwon, H.S.; Oh, S.M.; Kim, J.K. Glabridin, a functional compound of liquorice, attenuates colonic inflammation in mice with dextran sulphate sodium-induced colitis. Clin. Exp. Immunol. 2008, 151, 165–173. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.Q.; Xue, C.C.; Zhou, Z.W.; Li, C.G.; Du, Y.M.; Liang, J.; Zhou, S.F. In vitro and in vivo neuroprotective effect and mechanisms of glabridin, a major active isoflavan from Glycyrrhiza glabra (licorice). Life Sci. 2008, 82, 68–78. [Google Scholar] [CrossRef]
- Hsu, Y.L.; Wu, L.Y.; Hou, M.F.; Tsai, E.M.; Lee, J.N.; Liang, H.L.; Jong, Y.J.; Hung, C.H.; Kuo, P.L. Glabridin, an isoflavan from licorice root, inhibits migration, invasion and angiogenesis of MDA-MB-231 human breast adenocarcinoma cells by inhibiting focal adhesion kinase/Rho signaling pathway. Mol. Nutr. Food Res. 2011, 55, 318–327. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Mu, J.; Wang, X.; Ye, X.; Si, L.; Ning, S.; Li, Z.; Li, Y. The repressive effect of miR-148a on TGF beta-SMADs signal pathway is involved in the glabridin-induced inhibition of the cancer stem cells-like properties in hepatocellular carcinoma cells. PLoS ONE 2014, 9, e96698. [Google Scholar] [CrossRef]
- Jiang, F.; Li, Y.; Mu, J.; Hu, C.; Zhou, M.; Wang, X.; Si, L.; Ning, S.; Li, Z. Glabridin inhibits cancer stem cell-like properties of human breast cancer cells: An epigenetic regulation of miR-148a/SMAd2 signaling. Mol. Carcinog. 2016, 55, 929–940. [Google Scholar] [CrossRef] [PubMed]
- Mu, J.; Ning, S.; Wang, X.; Si, L.; Jiang, F.; Li, Y.; Li, Z. The repressive effect of miR-520a on NF-κB/IL-6/STAT-3 signal involved in the glabridin-induced anti-angiogenesis in human breast cancer cells. RSC Adv. 2015, 5, 34257–34264. [Google Scholar] [CrossRef]
- Zhou, C.; Li, X.; Du, W.; Feng, Y.; Kong, X.; Li, Y.; Xiao, L.; Zhang, P. Antitumor effects of ginkgolic acid in human cancer cell occur via cell cycle arrest and decrease the Bcl-2/Bax ratio to induce apoptosis. Chemotherapy 2010, 56, 393–402. [Google Scholar] [CrossRef]
- Borenstein, R.; Hanson, B.A.; Markosyan, R.M.; Gallo, E.S.; Narasipura, S.D.; Bhutta, M.; Nicholson, D.A. Ginkgolic acid inhibits fusion of enveloped viruses. Scientific Reports 2020, 10, 4746. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, M.A.; Ramadan, H.H.; Mohammed, R.N. Evidence that Ginkgo Biloba could use in the influenza and coronavirus COVID-19 infections. J. Basic. Clin. Physiol. Pharmacol. 2021, 32, 131–143. [Google Scholar] [CrossRef] [PubMed]
- Ye, M.; Liu, J.K.; Lu, Z.X.; Zhao, Y.; Liu, S.F.; Li, L.L.; Tan, M.; Weng, X.X.; Li, W.; Cao, Y. Grifolin, a potential antitumor natural product from the mushroom Albatrellus confluens, inhibits tumor cell growth by inducing apoptosis in vitro. FEBS Lett. 2005, 579, 3437–3443. [Google Scholar] [CrossRef]
- Luo, X.; Yang, L.; Xiao, L.; Xia, X.; Dong, X.; Zhong, J.; Liu, Y.; Li, N.; Chen, L.; Li, H.; et al. Grifolin directly targets ERK1/2 to epigenetically suppress cancer cell metastasis. Oncotarget 2015, 6, 42704–42716. [Google Scholar] [CrossRef] [PubMed]
- Nediani, C.; Ruzzolini, J.; Romani, A.; Calorini, L. Oleuropein, a Bioactive Compound from Olea europaea L., as a Potential Preventive and Therapeutic Agent in Non-Communicable Diseases. Antioxidants 2019, 8, 578. [Google Scholar] [CrossRef] [PubMed]
- Fabiani, R.; Vella, N.; Rosignoli, P. Epigenetic Modifications Induced by Olive Oil and Its Phenolic Compounds: A Systematic Review. Molecules 2021, 26, 273. [Google Scholar] [CrossRef]
- Juli, G.; Oliverio, M.; Bellizzi, D.; Gallo Cantafio, M.E.; Grillone, K.; Passarino, G.; Colica, C.; Nardi, M.; Rossi, M.; Procopio, A.; et al. Anti-tumor Activity and Epigenetic Impact of the Polyphenol Oleacein in Multiple Myeloma. Cancers 2019, 11, 990. [Google Scholar] [CrossRef]
- Cuyàs, E.; Gumuzio, J.; Lozano-Sánchez, J.; Carreras, D.; Verdura, S.; Llorach-Parés, L.; Sanchez-Martinez, M.; Selga, E.; Pérez, G.J.; Scornik, F.S.; et al. Extra Virgin Olive Oil Contains a Phenolic Inhibitor of the Histone Demethylase LSD1/KDM1A. Nutrients 2019, 11, 1656. [Google Scholar] [CrossRef]
- Lee-Huang, S.; Huang, P.L.; Zhang, D.; Lee, J.W.; Bao, J.; Sun, Y.; Chang, Y.T.; Zhang, J.; Huang, P.L. Discovery of small-molecule HIV-1 fusion and integrase inhibitors oleuropein and hydroxytyrosol: Part I. fusion [corrected] inhibition. Biochem. Biophys. Res. Commun. 2007, 354, 872–878. [Google Scholar] [CrossRef]
- Yamaguchi, Y.; Kumagai, H. Characteristics, biosynthesis, decomposition, metabolism and functions of the garlic odour precursor, S-allyl-L-cysteine sulfoxide. Exp. Ther. Med. 2020, 19, 1528–1535. [Google Scholar] [CrossRef]
- Tolo, F.M.; Rukunga, G.M.; Muli, F.W.; Njagi, E.N.; Njue, W.; Kumon, K.; Mungai, G.M.; Muthaura, C.N.; Muli, J.M.; Keter, L.K.; et al. Anti-viral activity of the extracts of a Kenyan medicinal plant Carissa edulis against herpes simplex virus. J. Ethnopharmacol. 2006, 104, 92–99. [Google Scholar] [CrossRef]
- Lin, L.T.; Hsu, W.C.; Lin, C.C. Antiviral natural products and herbal medicines. J. Tradit. Complement. Med. 2014, 4, 24–35. [Google Scholar] [CrossRef] [PubMed]
- Tsai, Y.; Cole, L.L.; Davis, L.E.; Lockwood, S.J.; Simmons, V.; Wild, G.C. Antiviral properties of garlic: In vitro effects on influenza B, herpes simplex and coxsackie viruses. Planta Med. 1985, 51, 460–461. [Google Scholar] [CrossRef] [PubMed]
- Abiy, E.; Berhe, A. Anti-bacterial effect of garlic (Allium sativum) against clinical isolates of Staphylococcus aureus and Escherichia coli from patients attending Hawassa Referral Hospital, Ethiopia. J. Infect. Dis. Treat. 2016, 2, 1–5. [Google Scholar] [CrossRef]
- Gebreyohannes, G.; Gebreyohannes, M. Medicinal values of garlic: A review. Int. J. Med. Med. Sci. 2013, 5, 401–408. [Google Scholar]
- Lawson, L.D.; Bauer, R. Phytomedicines of Europe: Chemistry and Biological Activity; ACS Publications: Washington, DC, USA, 1998. [Google Scholar]
- Borlinghaus, J.; Albrecht, F.; Gruhlke, M.C.; Nwachukwu, I.D.; Slusarenko, A.J. Allicin: Chemistry and biological properties. Molecules 2014, 19, 12591–12618. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Jiao, H.; Zhao, J.; Wang, X.; Sun, S.; Lin, H. Allicin Alleviates Reticuloendotheliosis Virus-Induced Immunosuppression via ERK/Mitogen-Activated Protein Kinase Pathway in Specific Pathogen-Free Chickens. Front. Immunol. 2017, 8, 1856. [Google Scholar] [CrossRef]
- Weber, N.D.; Andersen, D.O.; North, J.A.; Murray, B.K.; Lawson, L.D.; Hughes, B.G. In vitro virucidal effects of Allium sativum (garlic) extract and compounds. Planta Med. 1992, 58, 417–423. [Google Scholar] [CrossRef]
- Arreola, R.; Quintero-Fabián, S.; López-Roa, R.I.; Flores-Gutiérrez, E.O.; Reyes-Grajeda, J.P.; Carrera-Quintanar, L.; Ortuño-Sahagún, D. Immunomodulation and anti-inflammatory effects of garlic compounds. J. Immunol. Res. 2015, 2015, 401630. [Google Scholar] [CrossRef]
- Metwally, D.M.; Al-Olayan, E.M.; Alanazi, M.; Alzahrany, S.B.; Semlali, A. Antischistosomal and anti-inflammatory activity of garlic and allicin compared with that of praziquantel in vivo. BMC Complement. Altern. Med. 2018, 18, 135. [Google Scholar] [CrossRef]
- Prasad, K.; Laxdal, V.A.; Yu, M.; Raney, B.L. Antioxidant activity of allicin, an active principle in garlic. Mol. Cell Biochem. 1995, 148, 183–189. [Google Scholar] [CrossRef] [PubMed]
- El-Saber Batiha, G.; Magdy Beshbishy, A.; Wasef, L.G.; Elewa, Y.H.; Al-Sagan, A.A.; Abd El-Hack, M.E.; Taha, A.E.; Abd-Elhakim, Y.M.; Prasad Devkota, H. Chemical Constituents and Pharmacological Activities of Garlic (Allium sativum L.): A Review. Nutrients 2020, 12, 872. [Google Scholar] [CrossRef] [PubMed]
- Trio, P.Z.; You, S.; He, X.; He, J.; Sakao, K.; Hou, D.X. Chemopreventive functions and molecular mechanisms of garlic organosulfur compounds. Food Funct. 2014, 5, 833–844. [Google Scholar] [CrossRef]
- Fang, F.; Li, H.; Cui, W.; Dong, Y. Treatment of hepatitis caused by cytomegalovirus with allitridin injection--an experimental study. J. Tongji Med. Univ. 1999, 19, 271–274. [Google Scholar] [CrossRef] [PubMed]
- Keyaerts, E.; Vijgen, L.; Pannecouque, C.; Van Damme, E.; Peumans, W.; Egberink, H.; Balzarini, J.; Van Ranst, M. Plant lectins are potent inhibitors of coronaviruses by interfering with two targets in the viral replication cycle. Antivir. Res. 2007, 75, 179–187. [Google Scholar] [CrossRef]
- Liu, Z.F.; Fang, F.; Dong, Y.S.; Li, G.; Zhen, H. Experimental study on the prevention and treatment of murine cytomegalovirus hepatitis by using allitridin. Antivir. Res. 2004, 61, 125–128. [Google Scholar] [CrossRef] [PubMed]
- Terrasson, J.; Xu, B.; Li, M.; Allart, S.; Davignon, J.L.; Zhang, L.H.; Wang, K.; Davrinche, C. Activities of Z-ajoene against tumour and viral spreading in vitro. Fundam. Clin. Pharmacol. 2007, 21, 281–289. [Google Scholar] [CrossRef]
- Walder, R.; Kalvatchev, Z.; Garzaro, D.; Barrios, M.; Apitz-Castro, R. In vitro suppression of HIV-1 replication by ajoene [(e)-(z)-4,5,9-trithiadodeca-1,6,11-triene-9 oxide]. Biomed. Pharmacother. 1997, 51, 397–403. [Google Scholar] [CrossRef]
- Hall, A.; Troupin, A.; Londono-Renteria, B.; Colpitts, T.M. Garlic Organosulfur Compounds Reduce Inflammation and Oxidative Stress during Dengue Virus Infection. Viruses 2017, 9, 159. [Google Scholar] [CrossRef]
- Li, Y.N.; Huang, F.; Liu, X.L.; Shu, S.N.; Huang, Y.J.; Cheng, H.J.; Fang, F. Allium sativum-derived allitridin inhibits Treg amplification in cytomegalovirus infection. J. Med. Virol. 2013, 85, 493–500. [Google Scholar] [CrossRef]
- Yi, X.; Feng, F.; Xiang, Z.; Ge, L. The effects of allitridin on the expression of transcription factors T-bet and GATA-3 in mice infected by murine cytomegalovirus. J. Med. Food 2005, 8, 332–336. [Google Scholar] [CrossRef] [PubMed]
- Kaschula, C.H.; Hunter, R.; Parker, M.I. Garlic-derived anticancer agents: Structure and biological activity of ajoene. Biofactors 2010, 36, 78–85. [Google Scholar] [CrossRef]
- Yi, L.; Su, Q. Molecular mechanisms for the anti-cancer effects of diallyl disulfide. Food Chem. Toxicol. 2013, 57, 362–370. [Google Scholar] [CrossRef]
- Choi, H.J.; Bae, E.Y.; Song, J.H.; Baek, S.H.; Kwon, D.H. Inhibitory effects of orobol 7-O-D-glucoside from banaba (Lagerstroemia speciosa L.) on human rhinoviruses replication. Lett. Appl. Microbiol. 2010, 51, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Pevear, D.C.; Tull, T.M.; Seipel, M.E.; Groarke, J.M. Activity of pleconaril against enteroviruses. Antimicrob. Agents Chemother. 1999, 43, 2109–2115. [Google Scholar] [CrossRef] [PubMed]
- Burgett, A.W.; Poulsen, T.B.; Wangkanont, K.; Anderson, D.R.; Kikuchi, C.; Shimada, K.; Okubo, S.; Fortner, K.C.; Mimaki, Y.; Kuroda, M.; et al. Natural products reveal cancer cell dependence on oxysterol-binding proteins. Nat. Chem. Biol. 2011, 7, 639–647. [Google Scholar] [CrossRef] [PubMed]
- Albulescu, L.; Bigay, J.; Biswas, B.; Weber-Boyvat, M.; Dorobantu, C.M.; Delang, L.; van der Schaar, H.M.; Jung, Y.S.; Neyts, J.; Olkkonen, V.M.; et al. Uncovering oxysterol-binding protein (OSBP) as a target of the anti-enteroviral compound TTP-8307. Antivir. Res. 2017, 140, 37–44. [Google Scholar] [CrossRef] [PubMed]
- Roberts, B.L.; Severance, Z.C.; Bensen, R.C.; Le-McClain, A.T.; Malinky, C.A.; Mettenbrink, E.M.; Nuñez, J.I.; Reddig, W.J.; Blewett, E.L.; Burgett, A.W.G. Differing activities of oxysterol-binding protein (OSBP) targeting anti-viral compounds. Antivir. Res. 2019, 170, 104548. [Google Scholar] [CrossRef] [PubMed]
- Albulescu, L.; Strating, J.R.; Thibaut, H.J.; van der Linden, L.; Shair, M.D.; Neyts, J.; van Kuppeveld, F.J. Broad-range inhibition of enterovirus replication by OSW-1, a natural compound targeting OSBP. Antivir. Res. 2015, 117, 110–114. [Google Scholar] [CrossRef] [PubMed]
- Alonso-Álvarez, S.; Pardal, E.; Sánchez-Nieto, D.; Navarro, M.; Caballero, M.D.; Mateos, M.V.; Martín, A. Plitidepsin: Design, development, and potential place in therapy. Drug Des. Devel Ther. 2017, 11, 253–264. [Google Scholar] [CrossRef]
- Gomes, N.G.M.; Valentão, P.; Andrade, P.B.; Pereira, R.B. Plitidepsin to treat multiple myeloma. Drugs Today 2020, 56, 337–347. [Google Scholar] [CrossRef] [PubMed]
- Wei, T.; Li, D.; Marcial, D.; Khan, M.; Lin, M.H.; Snape, N.; Ghildyal, R.; Harrich, D.; Spann, K. The eukaryotic elongation factor 1A is critical for genome replication of the paramyxovirus respiratory syncytial virus. PLoS ONE 2014, 9, e114447. [Google Scholar] [CrossRef] [PubMed]
- White, K.M.; Rosales, R.; Yildiz, S.; Kehrer, T.; Miorin, L.; Moreno, E.; Jangra, S.; Uccellini, M.B.; Rathnasinghe, R.; Coughlan, L.; et al. Plitidepsin has potent preclinical efficacy against SARS-CoV-2 by targeting the host protein eEF1A. Science 2021, 371, 926–931. [Google Scholar] [CrossRef]
- Reuschl, A.-K.; Thorne, L.G.; Zuliani-Alvarez, L.; Bouhaddou, M.; Obernier, K.; Hiatt, J.; Soucheray, M.; Turner, J.; Fabius, J.M.; Nguyen, G.T. Host-directed therapies against early-lineage SARS-CoV-2 retain efficacy against B. 1.1. 7 variant. bioRxiv 2021. [Google Scholar] [CrossRef]
- Duke, S.O. Benefits of Resveratrol and Pterostilbene to Crops and Their Potential Nutraceutical Value to Mammals. Agriculture 2022, 12, 368. [Google Scholar] [CrossRef]
- Kala, R.; Shah, H.N.; Martin, S.L.; Tollefsbol, T.O. Epigenetic-based combinatorial resveratrol and pterostilbene alters DNA damage response by affecting SIRT1 and DNMT enzyme expression, including SIRT1-dependent γ-H2AX and telomerase regulation in triple-negative breast cancer. BMC Cancer 2015, 15, 672. [Google Scholar] [CrossRef]
- Liu, T.; Liu, P.Y.; Marshall, G.M. The critical role of the class III histone deacetylase SIRT1 in cancer. Cancer Res. 2009, 69, 1702–1705. [Google Scholar] [CrossRef]
- Ter Ellen, B.M.; Dinesh Kumar, N.; Bouma, E.M.; Troost, B.; van de Pol, D.P.I.; van der Ende-Metselaar, H.H.; Apperloo, L.; van Gosliga, D.; van den Berge, M.; Nawijn, M.C.; et al. Resveratrol and Pterostilbene Inhibit SARS-CoV-2 Replication in Air-Liquid Interface Cultured Human Primary Bronchial Epithelial Cells. Viruses 2021, 13, 1335. [Google Scholar] [CrossRef]
- Anand David, A.V.; Arulmoli, R.; Parasuraman, S. Overviews of Biological Importance of Quercetin: A Bioactive Flavonoid. Pharmacogn. Rev. 2016, 10, 84–89. [Google Scholar] [CrossRef]
- Lee, W.J.; Chen, Y.R.; Tseng, T.H. Quercetin induces FasL-related apoptosis, in part, through promotion of histone H3 acetylation in human leukemia HL-60 cells. Oncol. Rep. 2011, 25, 583–591. [Google Scholar] [CrossRef]
- Ahmed, F.; Ijaz, B.; Ahmad, Z.; Farooq, N.; Sarwar, M.B.; Husnain, T. Modification of miRNA Expression through plant extracts and compounds against breast cancer: Mechanism and translational significance. Phytomedicine 2020, 68, 153168. [Google Scholar] [CrossRef]
- Huang, Q.; Chen, L.; Luo, M.; Lv, H.; Luo, D.; Li, T.; Huang, S.; Xie, L.; Teng, Y.; Liu, Z.; et al. HIV-1-Induced miR-146a Attenuates Monocyte Migration by Targeting CCL5 in Human Primary Macrophages. AIDS Res. Hum. Retroviruses 2018, 34, 580–589. [Google Scholar] [CrossRef] [PubMed]
- Noratto, G.D.; Kim, Y.; Talcott, S.T.; Mertens-Talcott, S.U. Flavonol-rich fractions of yaupon holly leaves (Ilex vomitoria, Aquifoliaceae) induce microRNA-146a and have anti-inflammatory and chemopreventive effects in intestinal myofibroblast CCD-18Co cells. Fitoterapia 2011, 82, 557–569. [Google Scholar] [CrossRef] [PubMed]
- Sonoki, H.; Sato, T.; Endo, S.; Matsunaga, T.; Yamaguchi, M.; Yamazaki, Y.; Sugatani, J.; Ikari, A. Quercetin Decreases Claudin-2 Expression Mediated by Up-Regulation of microRNA miR-16 in Lung Adenocarcinoma A549 Cells. Nutrients 2015, 7, 4578–4592. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Guo, Q.; Chen, J.; Chen, Z. Quercetin Enhances Cisplatin Sensitivity of Human Osteosarcoma Cells by Modulating microRNA-217-KRAS Axis. Mol. Cells 2015, 38, 638–642. [Google Scholar] [CrossRef]
- Zhou, J.; Gong, J.; Ding, C.; Chen, G. Quercetin induces the apoptosis of human ovarian carcinoma cells by upregulating the expression of microRNA-145. Mol. Med. Rep. 2015, 12, 3127–3131. [Google Scholar] [CrossRef]
- Di Petrillo, A.; Orrù, G.; Fais, A.; Fantini, M.C. Quercetin and its derivates as antiviral potentials: A comprehensive review. Phytother. Res. 2022, 36, 266–278. [Google Scholar] [CrossRef]
- Zarenezhad, E.; Abdulabbas, H.T.; Kareem, A.S.; Kouhpayeh, S.A.; Barbaresi, S.; Najafipour, S.; Mazarzaei, A.; Sotoudeh, M.; Ghasemian, A. Protective role of flavonoids quercetin and silymarin in the viral-associated inflammatory bowel disease: An updated review. Arch. Microbiol. 2023, 205, 252. [Google Scholar] [CrossRef]
- Brito, J.C.M.; Lima, W.G.; Cordeiro, L.P.B.; da Cruz Nizer, W.S. Effectiveness of supplementation with quercetin-type flavonols for treatment of viral lower respiratory tract infections: Systematic review and meta-analysis of preclinical studies. Phytother. Res. 2021, 35, 4930–4942. [Google Scholar] [CrossRef]
- Pop, S.; Enciu, A.M.; Tarcomnicu, I.; Gille, E.; Tanase, C. Phytochemicals in cancer prevention: Modulating epigenetic alterations of DNA methylation. Phytochem. Rev. 2019, 18, 1005–1024. [Google Scholar] [CrossRef]
- Alway, S.E.; McCrory, J.L.; Kearcher, K.; Vickers, A.; Frear, B.; Gilleland, D.L.; Bonner, D.E.; Thomas, J.M.; Donley, D.A.; Lively, M.W.; et al. Resveratrol Enhances Exercise-Induced Cellular and Functional Adaptations of Skeletal Muscle in Older Men and Women. J. Gerontol. A Biol. Sci. Med. Sci. 2017, 72, 1595–1606. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, O.; Fontanes, V.; Raychaudhuri, S.; Loo, R.; Loo, J.; Arumugaswami, V.; Sun, R.; Dasgupta, A.; French, S.W. The heat shock protein inhibitor Quercetin attenuates hepatitis C virus production. Hepatology 2009, 50, 1756–1764. [Google Scholar] [CrossRef] [PubMed]
- Saiko, P.; Pemberger, M.; Horvath, Z.; Savinc, I.; Grusch, M.; Handler, N.; Erker, T.; Jaeger, W.; Fritzer-Szekeres, M.; Szekeres, T. Novel resveratrol analogs induce apoptosis and cause cell cycle arrest in HT29 human colon cancer cells: Inhibition of ribonucleotide reductase activity. Oncol. Rep. 2008, 19, 1621–1626. [Google Scholar]
- Koushki, M.; Amiri-Dashatan, N.; Ahmadi, N.; Abbaszadeh, H.A.; Rezaei-Tavirani, M. Resveratrol: A miraculous natural compound for diseases treatment. Food Sci. Nutr. 2018, 6, 2473–2490. [Google Scholar] [CrossRef]
- Parashar, G.; Capalash, N. Promoter methylation-independent reactivation of PAX1 by curcumin and resveratrol is mediated by UHRF1. Clin. Exp. Med. 2016, 16, 471–478. [Google Scholar] [CrossRef] [PubMed]
- Izquierdo-Torres, E.; Hernández-Oliveras, A.; Meneses-Morales, I.; Rodríguez, G.; Fuentes-García, G.; Zarain-Herzberg, Á. Resveratrol up-regulates ATP2A3 gene expression in breast cancer cell lines through epigenetic mechanisms. Int. J. Biochem. Cell Biol. 2019, 113, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Kala, R.; Tollefsbol, T.; Li, Y. Potential of resveratrol in inhibiting cancer and slowing aging. J. Nutr. Food Sci. S 2012, S5, 001. [Google Scholar] [CrossRef]
- Borra, M.T.; Smith, B.C.; Denu, J.M. Mechanism of human SIRT1 activation by resveratrol. J. Biol. Chem. 2005, 280, 17187–17195. [Google Scholar] [CrossRef]
- Tili, E.; Michaille, J.J. Resveratrol, MicroRNAs, Inflammation, and Cancer. J. Nucleic Acids 2011, 2011, 102431. [Google Scholar] [CrossRef]
- Bai, T.; Dong, D.S.; Pei, L. Synergistic antitumor activity of resveratrol and miR-200c in human lung cancer. Oncol. Rep. 2014, 31, 2293–2297. [Google Scholar] [CrossRef]
- Smoliga, J.M.; Blanchard, O.L. Recent data do not provide evidence that resveratrol causes ‘mainly negative’ or ‘adverse’ effects on exercise training in humans. J. Physiol. 2013, 591, 5251–5252. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Song, X.; Zhao, X.; Zhang, Y.; Wang, Y.; Jia, R.; Zou, Y.; Li, L.; Yin, Z. Insights into the Anti-inflammatory and Antiviral Mechanisms of Resveratrol. Mediat. Inflamm. 2022, 2022, 7138756. [Google Scholar] [CrossRef] [PubMed]
- Docherty, J.J.; Smith, J.S.; Fu, M.M.; Stoner, T.; Booth, T. Effect of topically applied resveratrol on cutaneous herpes simplex virus infections in hairless mice. Antivir. Res. 2004, 61, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Ayipo, Y.O.; Yahaya, S.N.; Alananzeh, W.A.; Babamale, H.F.; Mordi, M.N. Pathomechanisms, therapeutic targets and potent inhibitors of some beta-coronaviruses from bench-to-bedside. Infect. Genet. Evol. 2021, 93, 104944. [Google Scholar] [CrossRef]
- Yang, M.; Wei, J.; Huang, T.; Lei, L.; Shen, C.; Lai, J.; Yang, M.; Liu, L.; Yang, Y.; Liu, G.; et al. Resveratrol inhibits the replication of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in cultured Vero cells. Phytother. Res. 2021, 35, 1127–1129. [Google Scholar] [CrossRef] [PubMed]
- Shariq, M.; Malik, A.A.; Sheikh, J.A.; Hasnain, S.E.; Ehtesham, N.Z. Regulation of autophagy by SARS-CoV-2: The multifunctional contributions of ORF3a. J. Med. Virol. 2023, 95, e28959. [Google Scholar] [CrossRef]
- Ding, L.; Jiang, P.; Xu, X.; Lu, W.; Yang, C.; Zhou, P.; Liu, S. Resveratrol promotes HSV-2 replication by increasing histone acetylation and activating NF-κB. Biochem. Pharmacol. 2020, 171, 113691. [Google Scholar] [CrossRef]
- Docherty, J.J.; Sweet, T.J.; Bailey, E.; Faith, S.A.; Booth, T. Resveratrol inhibition of varicella-zoster virus replication in vitro. Antivir. Res. 2006, 72, 171–177. [Google Scholar] [CrossRef]
- Sasivimolphan, P.; Lipipun, V.; Likhitwitayawuid, K.; Takemoto, M.; Pramyothin, P.; Hattori, M.; Shiraki, K. Inhibitory activity of oxyresveratrol on wild-type and drug-resistant varicella-zoster virus replication in vitro. Antivir. Res. 2009, 84, 95–97. [Google Scholar] [CrossRef]
- Lehman, C.W.; Kehn-Hall, K.; Aggarwal, M.; Bracci, N.R.; Pan, H.C.; Panny, L.; Lamb, R.A.; Lin, S.C. Resveratrol Inhibits Venezuelan Equine Encephalitis Virus Infection by Interfering with the AKT/GSK Pathway. Plants 2021, 10, 346. [Google Scholar] [CrossRef]
- Eladwy, R.A.; Vu, H.T.; Shah, R.; Li, C.G.; Chang, D.; Bhuyan, D.J. The Fight against the Carcinogenic Epstein-Barr Virus: Gut Microbiota, Natural Medicines, and Beyond. Int. J. Mol. Sci. 2023, 24, 1716. [Google Scholar] [CrossRef]
- Yu, K.; Zhou, L.; Wang, Y.; Yu, C.; Wang, Z.; Liu, H.; Wei, H.; Han, L.; Cheng, J.; Wang, F.; et al. Mechanisms and Therapeutic Strategies of Viral Myocarditis Targeting Autophagy. Front. Pharmacol. 2022, 13, 843103. [Google Scholar] [CrossRef]
- Xie, X.H.; Zang, N.; Li, S.M.; Wang, L.J.; Deng, Y.; He, Y.; Yang, X.Q.; Liu, E.M. Resveratrol Inhibits respiratory syncytial virus-induced IL-6 production, decreases viral replication, and downregulates TRIF expression in airway epithelial cells. Inflammation 2012, 35, 1392–1401. [Google Scholar] [CrossRef]
- Kong, F.; Li, Q.; Zhang, F.; Li, X.; You, H.; Pan, X.; Zheng, K.; Tang, R. Sirtuins as Potential Therapeutic Targets for Hepatitis B Virus Infection. Front. Med. 2021, 8, 751516. [Google Scholar] [CrossRef]
- Bijak, M. Silybin, a Major Bioactive Component of Milk Thistle (Silybum marianum L. Gaernt.)-Chemistry, Bioavailability, and Metabolism. Molecules 2017, 22, 1942. [Google Scholar] [CrossRef] [PubMed]
- Anestopoulos, I.; Sfakianos, A.P.; Franco, R.; Chlichlia, K.; Panayiotidis, M.I.; Kroll, D.J.; Pappa, A. A novel role of silibinin as a putative epigenetic modulator in human prostate carcinoma. Molecules 2016, 22, 62. [Google Scholar] [CrossRef] [PubMed]
- Barros, T.M.B.; Lima, A.P.B.; Almeida, T.C.; da Silva, G.N. Inhibition of urinary bladder cancer cell proliferation by silibinin. Environ. Mol. Mutagen. 2020, 61, 445–455. [Google Scholar] [CrossRef]
- Kauntz, H.; Bousserouel, S.; Gossé, F.; Raul, F. Epigenetic effects of the natural flavonolignan silibinin on colon adenocarcinoma cells and their derived metastatic cells. Oncol. Lett. 2013, 5, 1273–1277. [Google Scholar] [CrossRef] [PubMed]
- Bouyahya, A.; Mechchate, H.; Oumeslakht, L.; Zeouk, I.; Aboulaghras, S.; Balahbib, A.; Zengin, G.; Kamal, M.A.; Gallo, M.; Montesano, D.; et al. The Role of Epigenetic Modifications in Human Cancers and the Use of Natural Compounds as Epidrugs: Mechanistic Pathways and Pharmacodynamic Actions. Biomolecules 2022, 12, 367. [Google Scholar] [CrossRef]
- Zadeh, M.M.; Motamed, N.; Ranji, N.; Majidi, M.; Falahi, F. Silibinin-Induced Apoptosis and Downregulation of MicroRNA-21 and MicroRNA-155 in MCF-7 Human Breast Cancer Cells. J. Breast Cancer 2016, 19, 45–52. [Google Scholar] [CrossRef]
- Liu, C.H.; Jassey, A.; Hsu, H.Y.; Lin, L.T. Antiviral Activities of Silymarin and Derivatives. Molecules 2019, 24, 1552. [Google Scholar] [CrossRef]
- Edwards, R.B.; Lucas, D.M.; Lozanski, G.; Johnson, A.J.; Su, B.-N.; Lin, T.S.; Byrd, J.C.; Kinghorn, A.D.; Grever, M.R. Silvestrol, a Rocaglate Derivative from the Indonesian Plant Aglaia foveolata, Has Significant Bcl-2- and p53-Independent Anti-Tumor Activity against Chronic Lymphocytic Leukemia Cells. Blood 2006, 108, 2600. [Google Scholar] [CrossRef]
- Chu, J.; Galicia-Vázquez, G.; Cencic, R.; Mills, J.R.; Katigbak, A.; Porco, J.A., Jr.; Pelletier, J. CRISPR-Mediated Drug-Target Validation Reveals Selective Pharmacological Inhibition of the RNA Helicase, eIF4A. Cell Rep. 2016, 15, 2340–2347. [Google Scholar] [CrossRef]
- Biedenkopf, N.; Lange-Grünweller, K.; Schulte, F.W.; Weißer, A.; Müller, C.; Becker, D.; Becker, S.; Hartmann, R.K.; Grünweller, A. The natural compound silvestrol is a potent inhibitor of Ebola virus replication. Antivir. Res. 2017, 137, 76–81. [Google Scholar] [CrossRef] [PubMed]
- Elgner, F.; Sabino, C.; Basic, M.; Ploen, D.; Grünweller, A.; Hildt, E. Inhibition of Zika Virus Replication by Silvestrol. Viruses 2018, 10, 149. [Google Scholar] [CrossRef]
- Henss, L.; Scholz, T.; Grünweller, A.; Schnierle, B.S. Silvestrol Inhibits Chikungunya Virus Replication. Viruses 2018, 10, 592. [Google Scholar] [CrossRef]
- Müller, C.; Schulte, F.W.; Lange-Grünweller, K.; Obermann, W.; Madhugiri, R.; Pleschka, S.; Ziebuhr, J.; Hartmann, R.K.; Grünweller, A. Broad-spectrum antiviral activity of the eIF4A inhibitor silvestrol against corona- and picornaviruses. Antivir. Res. 2018, 150, 123–129. [Google Scholar] [CrossRef]
- Müller, C.; Obermann, W.; Schulte, F.W.; Lange-Grünweller, K.; Oestereich, L.; Elgner, F.; Glitscher, M.; Hildt, E.; Singh, K.; Wendel, H.G.; et al. Comparison of broad-spectrum antiviral activities of the synthetic rocaglate CR-31-B (−) and the eIF4A-inhibitor Silvestrol. Antivir. Res. 2020, 175, 104706. [Google Scholar] [CrossRef]
- Müller, C.; Obermann, W.; Karl, N.; Wendel, H.G.; Taroncher-Oldenburg, G.; Pleschka, S.; Hartmann, R.K.; Grünweller, A.; Ziebuhr, J. The rocaglate CR-31-B (−) inhibits SARS-CoV-2 replication at non-cytotoxic, low nanomolar concentrations in vitro and ex vivo. Antivir. Res. 2021, 186, 105012. [Google Scholar] [CrossRef] [PubMed]
- M de Figueiredo, S.; S Binda, N.; A Nogueira-Machado, J.; A Vieira-Filho, S.; B Caligiorne, R. The antioxidant properties of organosulfur compounds (sulforaphane). Recent. Pat. Endocr. Metab. Immune Drug Discov. 2015, 9, 24–39. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.-L.; Zhou, S.-J.; Zhang, X.-M.; Chen, H.-Q.; Liu, W. Sulforaphane suppresses in vitro and in vivo lung tumorigenesis through downregulation of HDAC activity. Biomed. Pharmacother. 2016, 78, 74–80. [Google Scholar] [CrossRef]
- Dos Santos, P.W.d.S.; Machado, A.R.T.; De Grandis, R.A.; Ribeiro, D.L.; Tuttis, K.; Morselli, M.; Aissa, A.F.; Pellegrini, M.; Antunes, L.M.G. Transcriptome and DNA methylation changes modulated by sulforaphane induce cell cycle arrest, apoptosis, DNA damage, and suppression of proliferation in human liver cancer cells. Food Chem. Toxicol. 2020, 136, 111047. [Google Scholar] [CrossRef]
- Yu, J.-S.; Chen, W.-C.; Tseng, C.-K.; Lin, C.-K.; Hsu, Y.-C.; Chen, Y.-H.; Lee, J.-C. Sulforaphane suppresses hepatitis C virus replication by up-regulating heme oxygenase-1 expression through PI3K/Nrf2 pathway. PLoS ONE 2016, 11, e0152236. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.-C.; Huang, C.-H.; Liu, W.; Lee, J.-C. Sulforaphane suppresses dengue virus replication by inhibition of dengue protease and enhancement of antiviral interferon response through Nrf2-mediated heme oxygenase-1 induction. Antivir. Res. 2022, 207, 105400. [Google Scholar] [CrossRef] [PubMed]
- Ordonez, A.A.; Bullen, C.K.; Villabona-Rueda, A.F.; Thompson, E.A.; Turner, M.L.; Merino, V.F.; Yan, Y.; Kim, J.; Davis, S.L.; Komm, O.; et al. Sulforaphane exhibits antiviral activity against pandemic SARS-CoV-2 and seasonal HCoV-OC43 coronaviruses in vitro and in mice. Commun. Biol. 2022, 5, 242. [Google Scholar] [CrossRef]
- Fields, N.J.; Palmer, K.R.; Rolnik, D.L.; Yo, J.; Nold, M.F.; Giles, M.L.; Marshall, S.A. CO-Sprout-A Pilot Double-Blinded Placebo-Controlled Randomised Trial of Broccoli Sprout Powder Supplementation for Pregnant Women with COVID-19 on the Duration of COVID-19-Associated Symptoms: Study Protocol. Nutrients 2023, 15, 3980. [Google Scholar] [CrossRef]
- Zeng, A.; Wang, D.; Wu, N.; Yang, R.; Nan, G.; Bian, X.; Yang, G. Extraction of Tanshinone IIA and Cryptotanshinone from the Rhizome of Salvia miltiorrhiza Bunge: Kinetics and Modeling. Sep. Sci. Technol. 2014, 49, 2330–2337. [Google Scholar] [CrossRef]
- Wang, L.; Zhang, C.; Guo, Y.; Su, Z.-Y.; Yang, Y.; Shu, L.; Kong, A.-N.T. Blocking of JB6 cell transformation by tanshinone IIA: Epigenetic reactivation of Nrf2 antioxidative stress pathway. AAPS J. 2014, 16, 1214–1225. [Google Scholar] [CrossRef]
- Valipour, M. Therapeutic prospects of naturally occurring p38 MAPK inhibitors tanshinone IIA and pinocembrin for the treatment of SARS-CoV-2-induced CNS complications. Phytother. Res. 2023, 37, 3724–3743. [Google Scholar] [CrossRef]
- Machado, D.G.; Neis, V.B.; Balen, G.O.; Colla, A.; Cunha, M.P.; Dalmarco, J.B.; Pizzolatti, M.G.; Prediger, R.D.; Rodrigues, A.L. Antidepressant-like effect of ursolic acid isolated from Rosmarinus officinalis L. in mice: Evidence for the involvement of the dopaminergic system. Pharmacol. Biochem. Behav. 2012, 103, 204–211. [Google Scholar] [CrossRef]
- Zhang, C.; Wang, C.; Li, W.; Wu, R.; Guo, Y.; Cheng, D.; Yang, Y.; Androulakis, I.P.; Kong, A.N. Pharmacokinetics and Pharmacodynamics of the Triterpenoid Ursolic Acid in Regulating the Antioxidant, Anti-inflammatory, and Epigenetic Gene Responses in Rat Leukocytes. Mol. Pharm. 2017, 14, 3709–3717. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Chen, J.; Liu, T.; Fang, J.; Wan, J.; Zhao, J.; Li, W.; Liu, J.; Zhao, X.; Chen, S. Anti-viral effects of urosolic acid on guinea pig cytomegalovirus in vitro. J. Huazhong Univ. Sci. Technol. Med. Sci. 2012, 32, 883–887. [Google Scholar] [CrossRef] [PubMed]
- Bungau, S.; Vesa, C.M.; Abid, A.; Behl, T.; Tit, D.M.; Purza, A.L.; Pasca, B.; Todan, L.M.; Endres, L. Withaferin A-A Promising Phytochemical Compound with Multiple Results in Dermatological Diseases. Molecules 2021, 26, 2407. [Google Scholar] [CrossRef]
- Royston, K.J.; Udayakumar, N.; Lewis, K.; Tollefsbol, T.O. A novel combination of withaferin A and sulforaphane inhibits epigenetic machinery, cellular viability and induces apoptosis of breast cancer cells. Int. J. Mol. Sci. 2017, 18, 1092. [Google Scholar] [CrossRef]
- Choe, J.; Har Yong, P.; Xiang Ng, Z. The Efficacy of Traditional Medicinal Plants in Modulating the Main Protease of SARS-CoV-2 and Cytokine Storm. Chem. Biodivers. 2022, 19, e202200655. [Google Scholar] [CrossRef]
- Cai, Z.; Zhang, G.; Tang, B.; Liu, Y.; Fu, X.; Zhang, X. Promising Anti-influenza Properties of Active Constituent of Withania somnifera Ayurvedic Herb in Targeting Neuraminidase of H1N1 Influenza: Computational Study. Cell Biochem. Biophys. 2015, 72, 727–739. [Google Scholar] [CrossRef] [PubMed]
- Huan, C.; Xu, Y.; Zhang, W.; Guo, T.; Pan, H.; Gao, S. Research Progress on the Antiviral Activity of Glycyrrhizin and its Derivatives in Liquorice. Front. Pharmacol. 2021, 12, 680674. [Google Scholar] [CrossRef]
- Wolkerstorfer, A.; Kurz, H.; Bachhofner, N.; Szolar, O.H. Glycyrrhizin inhibits influenza A virus uptake into the cell. Antivir. Res. 2009, 83, 171–178. [Google Scholar] [CrossRef]
- Zuo, J.; Meng, T.; Wang, Y.; Tang, W. A Review of the Antiviral Activities of Glycyrrhizic Acid, Glycyrrhetinic Acid and Glycyrrhetinic Acid Monoglucuronide. Pharmaceuticals 2023, 16, 641. [Google Scholar] [CrossRef]
- El Omari, N.; Bakrim, S.; Bakha, M.; Lorenzo, J.M.; Rebezov, M.; Shariati, M.A.; Aboulaghras, S.; Balahbib, A.; Khayrullin, M.; Bouyahya, A. Natural Bioactive Compounds Targeting Epigenetic Pathways in Cancer: A Review on Alkaloids, Terpenoids, Quinones, and Isothiocyanates. Nutrients 2021, 13, 3714. [Google Scholar] [CrossRef]
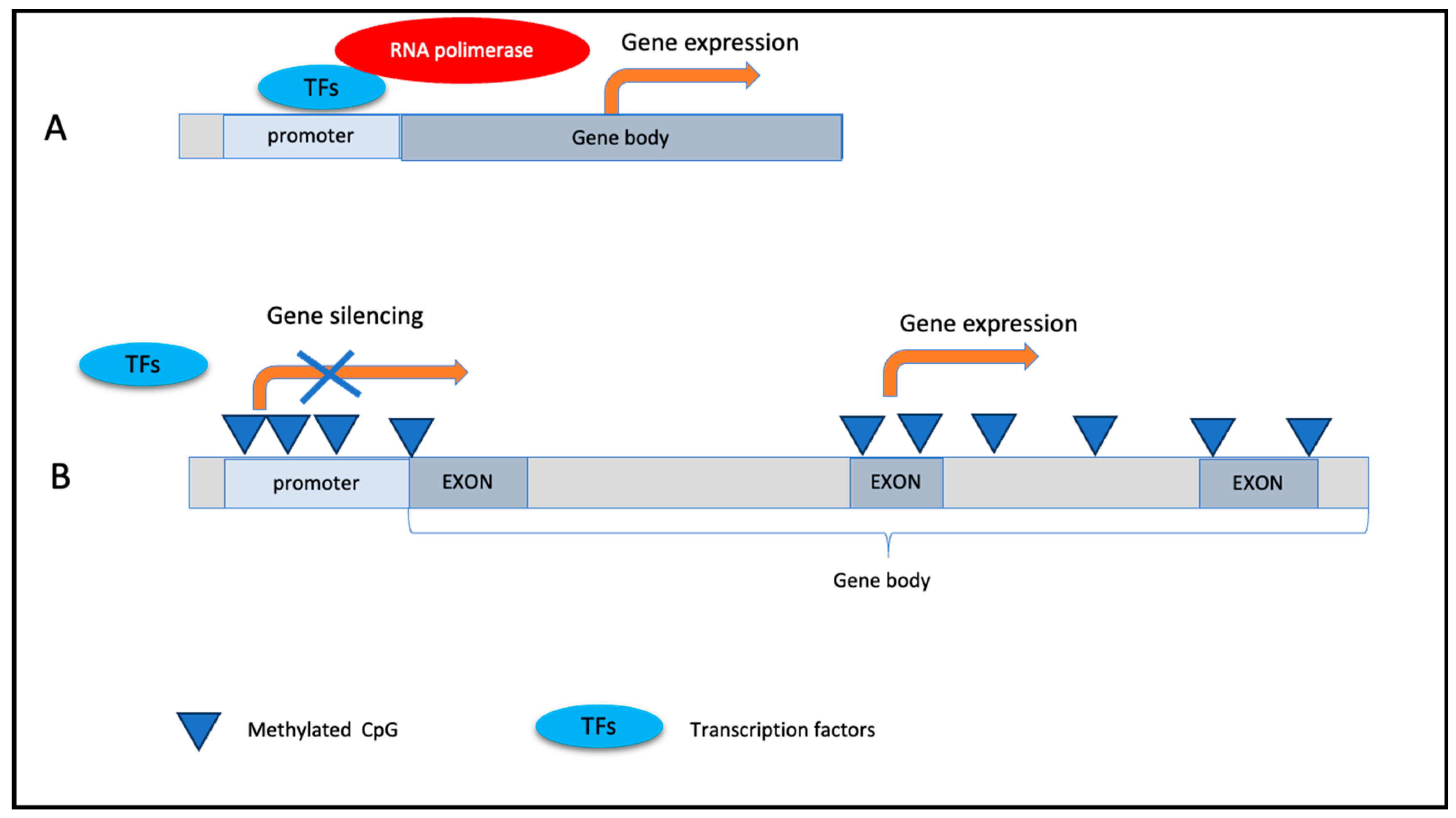
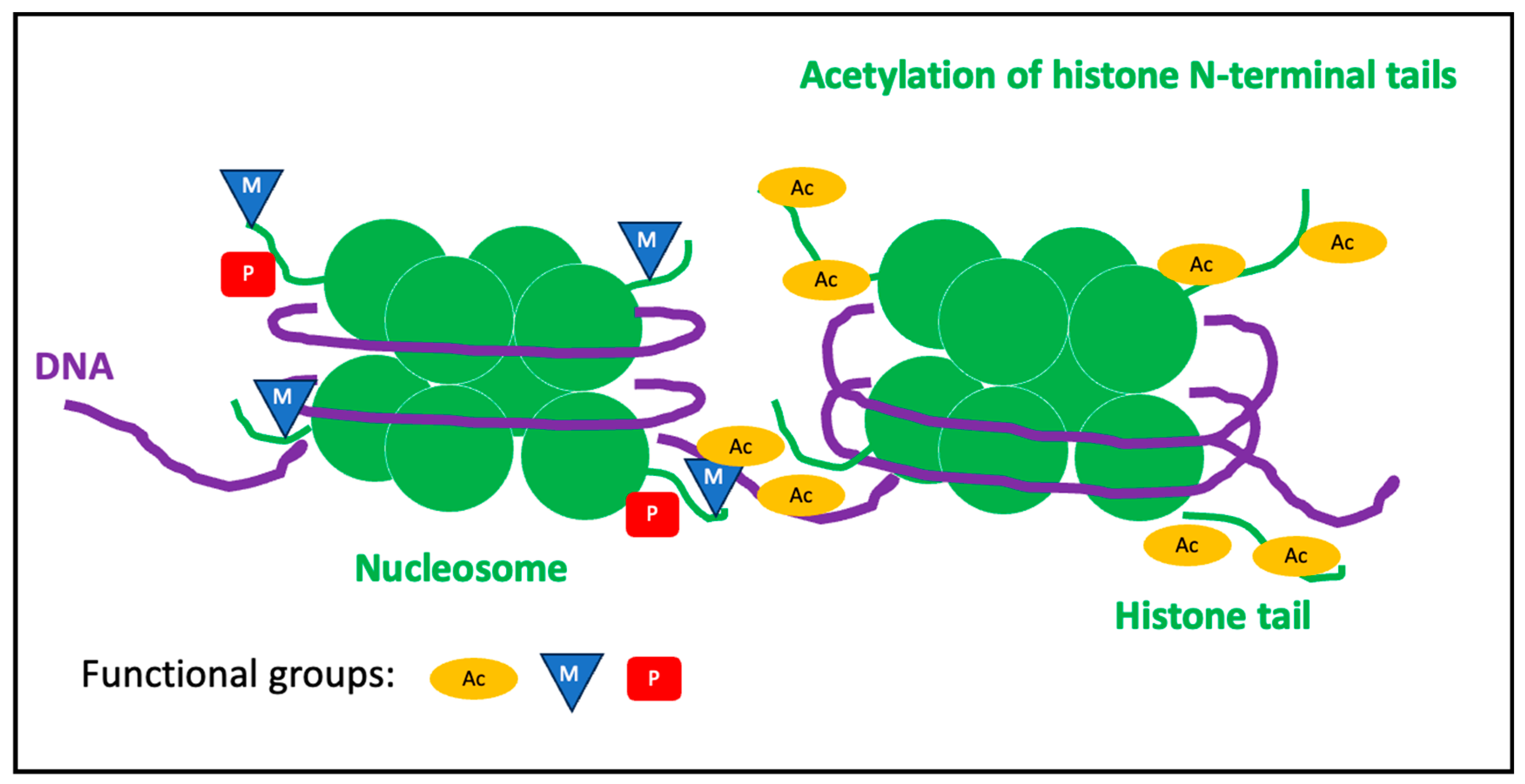
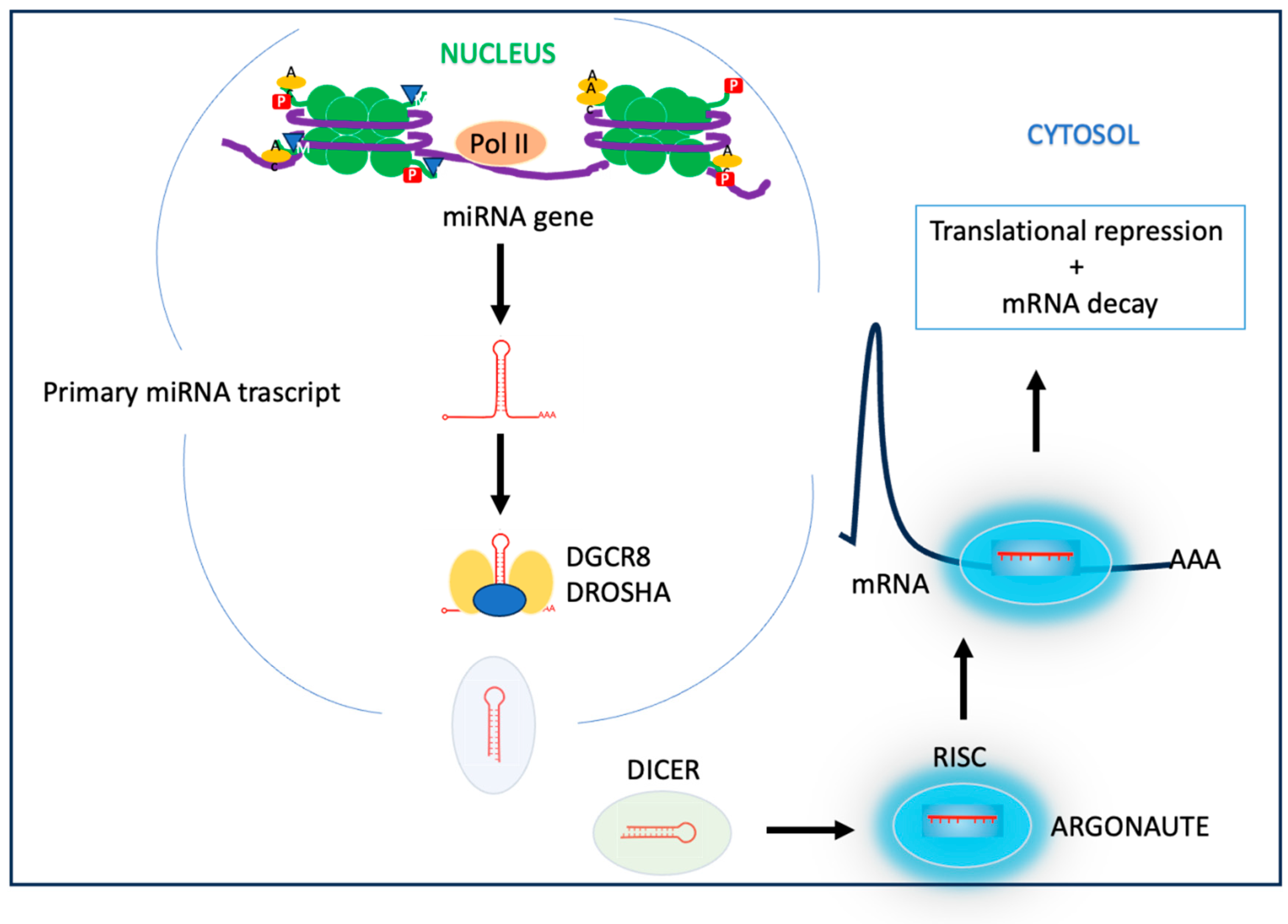

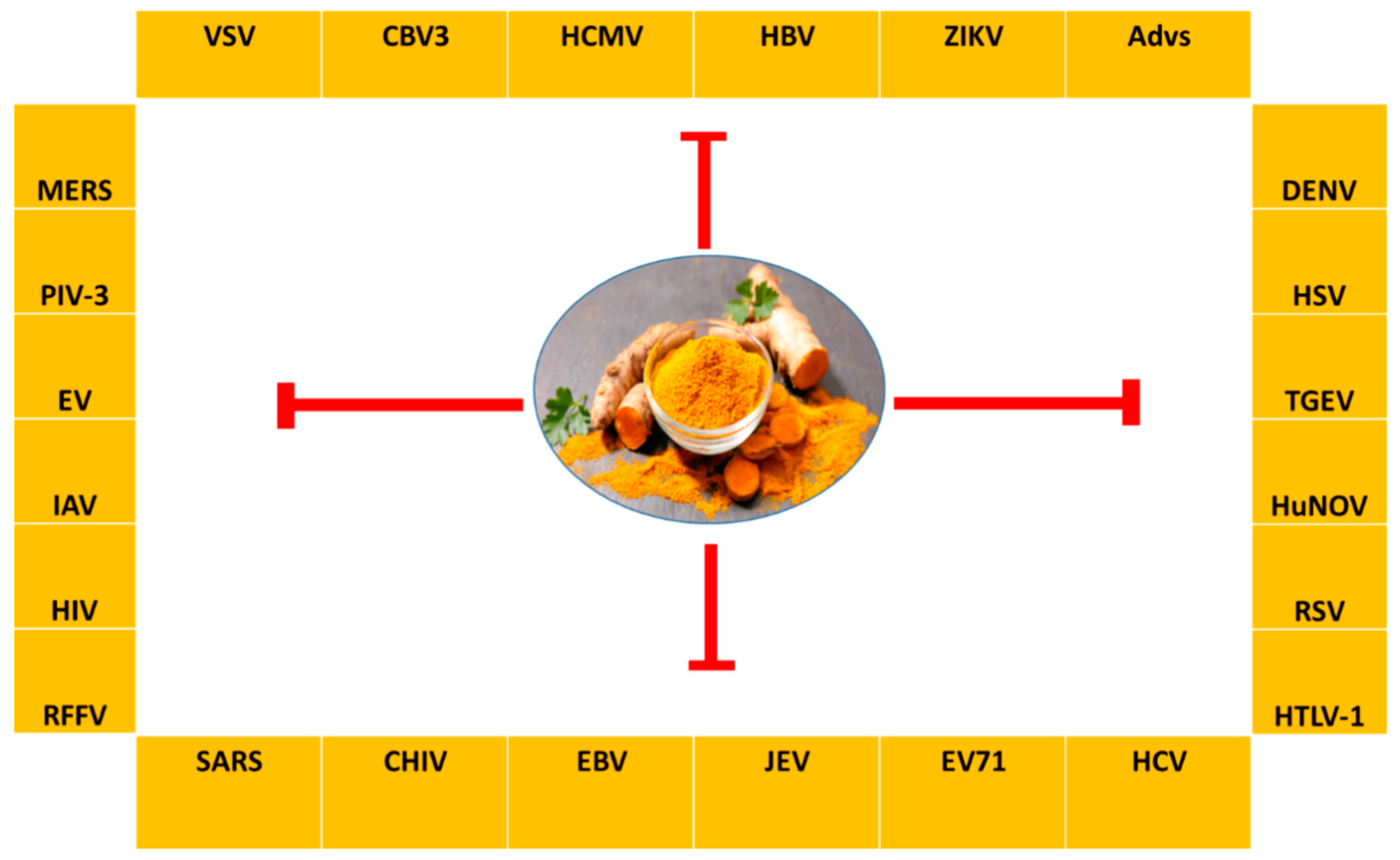
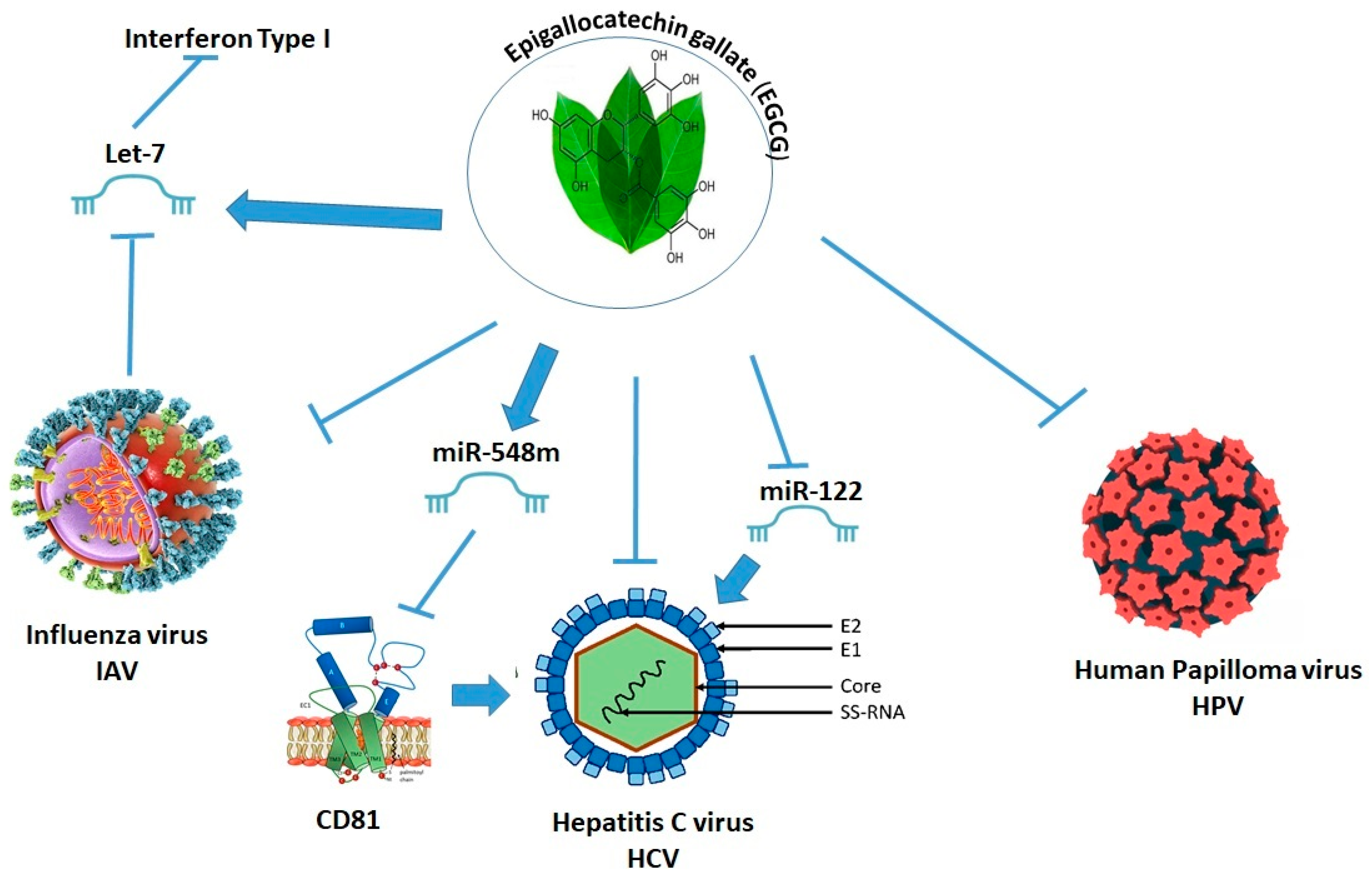
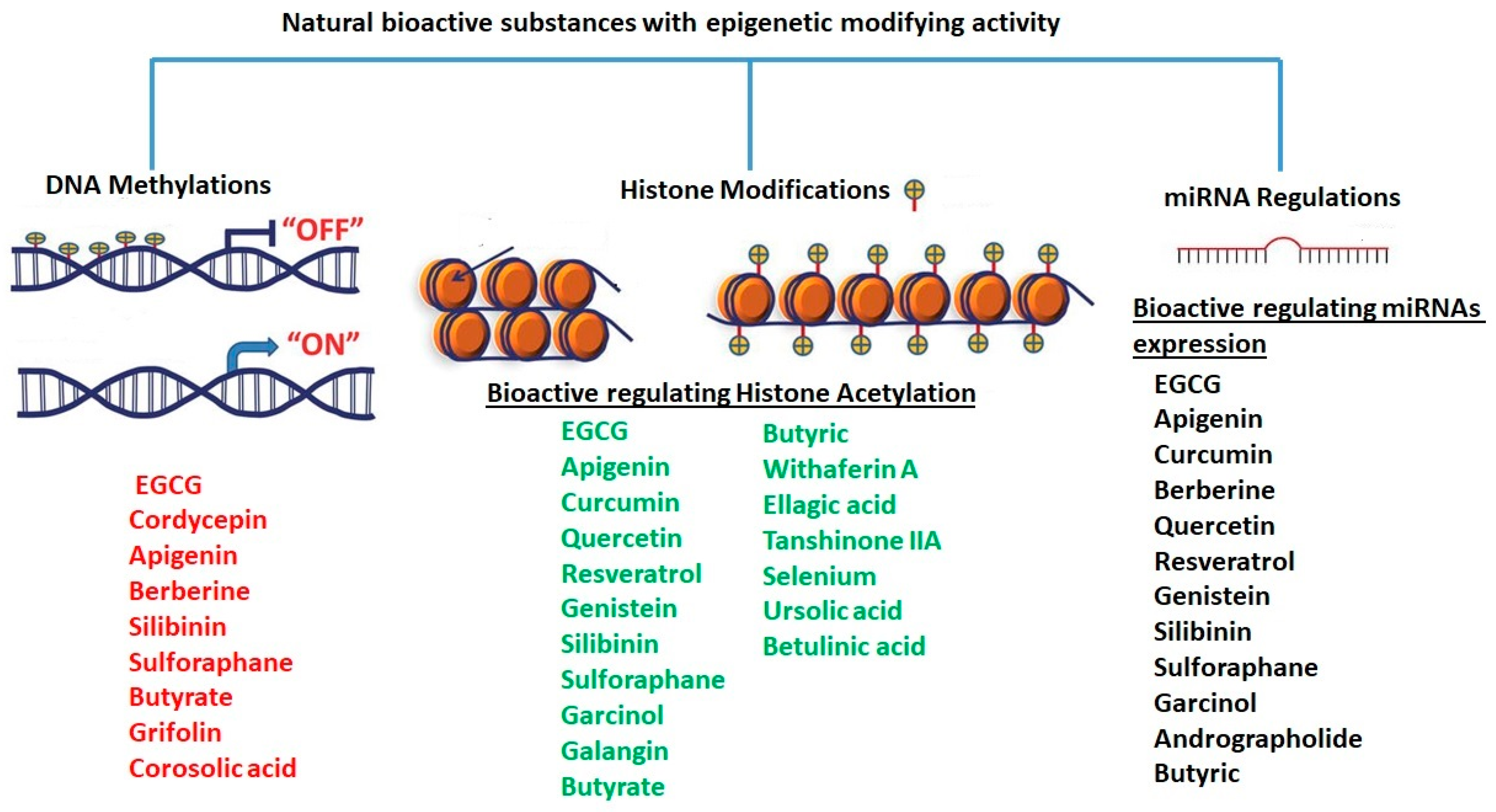
| Name | Structure | Anti-Viral Activity and Mode of Action |
|---|---|---|
| Andrographolide |  | Modulates miR-377 to regulate HO-1. Downregulates miR-433 to regulate glutathione cysteine ligase. Upregulates miR-17, miR-224, and miR-181a [190]. Inhibits virus replications such HIV, HSV-1, HBV, HCV, ZIKV, DENV, CHIKV, FMDV, and IAV [181,182,184]. Prevents EBV reactivation by suppressing EBV lytic genes via histone modifications [186]. Also prevents IVA-induced inflammation through modulation of NF-κB and JAK-STAT signaling pathways [189]. |
| Apigenin |  | A potent inhibitor of HDAC1, 3 [192]. Exhibits antiviral properties against IAV, HRV, HSV, enterovirus, HBV, HCV, EV71, CVB1, and SARS-CoV-2 [195,196,197,198,200]. Anti-viral activity is attributed to the inhibition of HDAC activity and chromatin remodeling [192,199]. Downregulates miR-103 [201], miR-155 [111], and miR-122 [202] to affect HCV and HIV replication [113,114,115]. |
| Baicalein |  | Inhibits HBV [204], HIV [205], DENV [206], and HSV-1 [207]. Inhibits DNMT and HDAC and affects influence epigenetic modifications [208,209]. Upregulates miR-3,178 [210] which targets HDAC10 [209]. Exhibits potent antiviral activity against DENV [206] |
| Berberine | 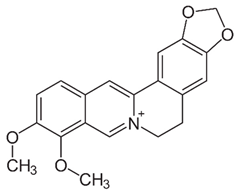 | Regulates AMPK activity [212] and suppresses SIRT1 deacetylases [214]. Inhibits miR-21 expression [215]. Inhibits replication of HSV [217], HCMV [218], HPV [219], DENV [220], HIV [221], HCV [222], IAV [224], and SARS-CoV-2 [223]. Also inhibits p38 MAPK activity [39] which might explain its anti-viral activity [42,43,44,45,46,226]. |
| Betulinic acid | 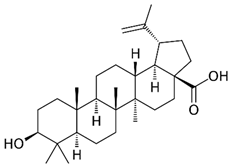 | A computational approach predicted the capacity to alter HDAC6 and HDAC10 activity [229]. Inhibits DENV-2 NS5 polymerase activity [231] and HBV replication [232]. C-3 esterification of lead resulted in the discovery of Bevirimat, an HIV-1 maturation inhibitor. |
| Butyric acid |  | Induces HDAC expression and activity by upregulation of miR-203 promoter methylation [233]. Inhibits HBV replication by reducing HBx protein expression, HBV-DNA, and HBsAg [234]. |
| Cardamonin |  | Exhibits antiviral action against the human coronavirus HCoV-OC43 which was mediated by induction of p38 MAPK signaling pathway [236]. |
| Cordycepin |  | Promotes methylation of EBV genomic sites near Fp/Qp promoters. Increases DNMT3 [238] and reduces EBV replication [239]. As adenosine derivative, it exhibits antiviral activity against several viruses, including IAV, plant viruses, HIV, murine leukemia virus, EBV [241,242,243,244,245], and COVID-19 [246]. |
| Corosolic acid | 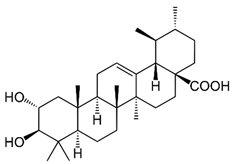 | Alters CpG methylation sites, resulting in altered gene expression [249]. Increases the expression of acetylated histone H3 lysine 27 (H3K27ac), while decreasing histone H3 lysine 27 trimethylation (H3K27Me3) [251]. Exhibits anti-viral activity against a number of viruses [248]. |
| Curcumin |  | Decreases the expression of HDAC1, HDAC3, HDAC8, and histone acetyltransferase p300 while enhancing the expression of Ac-histone H4 protein [253]. Reduces HAT activity and inhibits DNMT [254]. Inhibition of HBV replication was attributed to a decrease in the acetylation level of cccDNA-bound histone H3 and H4 [255]. Downregulates miR-350, miR-17-2-3p, let 7e-3p, miR-1224, miR-466b-1-3p, miR-18a-5p, and miR-322-5p. Upregulates miR-122-5p, miR-3473, miR-182, and miR-344a-3p [256]. Inhibits the replication of various viruses, including HBV [255], HIV [258], IAV [259], HPV-18 [261], ZIKV, CHIKV, VSV, CVB3, EV71, RSV, HSV-2, KSHV, and HAdV [263]. |
| Ellagic acid | 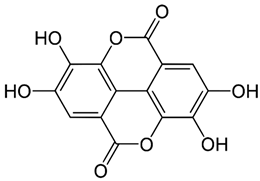 | Increases HDACs’ gene expression and histone arginine methylation [268]. Decreases H3K9 acetylation and HDAC9 dissociation [268]. Inhibits SARS-CoV-2 viral entry and replication [269]. |
| Epigallocatechin gallate (EGCG) | 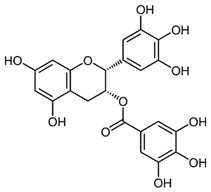 | Inhibits the activity of DNMT 1, DNMT 3a, DNMT 3b [273], and HDACs [275] and downregulates the expression of HDAC1, HDAC2, and HDAC3 [277]. Inhibits HAT activity [278]. Decreases the levels of let-7e-5p, miR-103a-3p, miR-151a-5p, miR-195-5p, miR-222-3p, miR-23a-3p, miR-23b-3p, miR-26a-5p, miR-27a-3p, miR-29b-3p, miR-3195, miR-3651, miR-4281, miR-4459, miR-4516, miR-762, and miR-125b-5p [283]. Induces the expression of miR-3663-3p, miR-1181, miR-3613-3p, miR1281, miR-1539, miR-221-5p, miR-374b, miR-4306, miR-500a-5p, miR590-5p miR-140-3p, and miR-221 [284,285,286]. Inhibits the replication of IAV, HBV, HCV, HSV-1 and HSV-2, HPV, ZIKV, and SARS-CoV-2 [288,289,290,291,292,293,294]. Upregulates miR-548m and inhibits miR-122 expression, which modulates HCV infectivity [291]. Upregulates let-7 to increase interferon expression and inhibit IAV infection [202]. |
| Galangin |  | Inhibits HDAC activity [299] and upregulates miR-455-5p [300]. Exhibits antiviral activity against HSV-1 and CoxB1 [301]. |
| Garcinol | 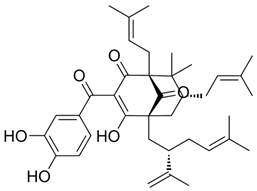 | Decreases HAT activity of p300 and pCAF [303]. Downregulates miR-21, miR-494, miR-495, and miR-1977 [309]. Upregulates miR-453, miR-128, miR-1280 and miR-720, let-7a, let-7e, let-7f, miR-200b, and miR-200c [311]. Inhibits HIV-1 reverse-transcriptase-associated ribonuclease H [312]. |
| Genistein | 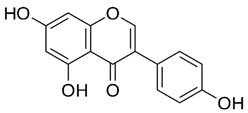 | Reduces HDAC while increasing HAT activity [313]. Inhibits miR-223 and miR-223 expression [316] which is involved in regulation of immune response and viral infections [115,116,117]. |
| Ginkgolic acid | 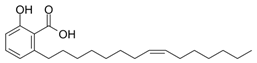 | Impairs protein SUMOylation [318]. Inhibits HSV-1, HSV-2, VZV, HCMV, ZIKV, IAV, EBV, HIV, EBOV, and Coronavirus COVID-19 [342]. |
| Glycyrrhizic acid | 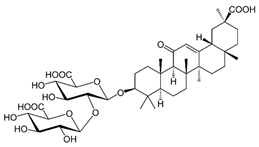 | Inhibits replications of various viruses including HBV, HCV, IAV H1N1, HIV [326], NDV [327], SARS-CoV-2 [458,459,460], RSV, VACV, HSV [329], and VSV [329,330]. Exhibits anti-inflammatory effects, decreasing IL-6 release [331] by regulating NF-κB and PI3K signaling pathways [333]. Inhibits viral replication of various viruses including HBV, HCV, IAV H1N1, and HIV [326]. |
| Grifolin | 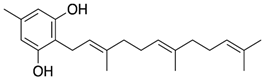 | Reduces Elk1 transcription as well as its binding to the DNMT1 promoter region [461]. Modulates ERK1/2-Elk1-DNMT1 signaling [344]. |
| Oleacein | 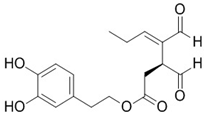 | Downregulates several class I/II HDACs [346,347]. Exhibits antiviral effect against HIV-1 [349]. |
| Plitidepsin | 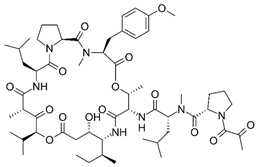 | Targets the eukaryotic translation elongation factor 1A (eEF1A) [382]. Exhibits anti-viral activity against RSV, gastroenteritis coronavirus [383], and SARS-CoV-2 [384,385]. |
| Pterostilbene | 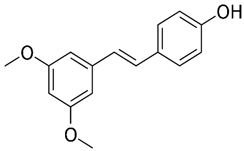 | Modulates HDAC activity and inhibits SIRT1 [387,388]. Inhibits SARS-CoV-2 replication [389] |
| Quercetin | 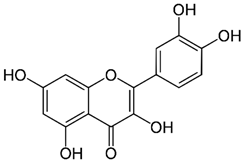 | Enhances histone H3 acetylation, activates HAT, and inhibits HDAC activities [391]. Inhibits HMT [213]. Inhibits miR-146a expression [392], a regulator of HIV replication [393], and miR-16, miR-217, and miR-145 [395,396,397]. Inhibits replication of IAV H1N1, IVA H3N2, HBV, HCV, DENV, poliovirus, rhinovirus, CHIKV, MERS-CoV, HSV 1/2, EBV, RSV, Arbovirus, EBOV, HIV, Japanese encephalitis virus, hAdV, enterovirus, ZIKV, NDV, MAYV, and SARS-CoV-2 [197,293,398,399,400]. Activates SIRT1 which resulted in inhibition of HCV [403] Upregulates let-7 which restores anti-viral immune response and thus exhibits anti-IVA activity [202]. |
| Resveratrol | 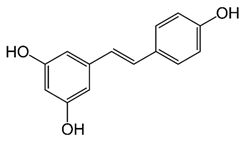 | Inhibits HDAC [406,407] and activates SIRT1 [409]. Decreases the levels of miR-17, miR-21, miR-25, miR-92a-2, miR-103-1, and miR-103-2 [410] and upregulates miR-200c [411]. In a human study, it increased miR-21, miR-181b, miR-663, and miR-30c, while reducing inflammatory cytokines like IL-6, CCL3, IL-1β, and TNF-α [412]. Inhibits HSV infection [414], beta-corona viruses such as MERS-COV and SARS-CoV-2 [415], Varicella-zoster virus (VZV) wild-type and DNA polymerase mutants with acyclovir-resistant VZV [419,420], VEEV [421], EBV [422], CV [423], and RSV. SIRT proteins regulate HBV replication and thus SIRT modulators such as resveratrol are suitable as to be used against HBV and RSV infections [402,425]. |
| Silibinin | 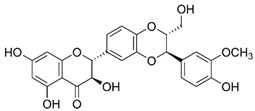 | Inhibits the expression of HDAC1, HDAC2, HDAC3, HDAC6, SET domain proteins (SETD1A, D4, D6), and lysine-specific demethylases (KDM 5B, 5C, 4A) [315]. Inhibits DNMTs [428,429,430]. Downregulates expression of miR-21 and miR-155 [431]. Exhibits anti-viral activity against HBV, DENV, CHIKV, MAYV, IVA, HIV, and HBV [432]. |
| Silvestrol | 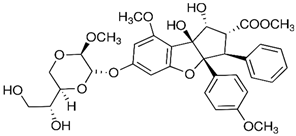 | Targets the eukaryotic initiation factor-4A (eIF4A) [434]. Exhibits activity against EBOV, ZIKV, CHIKV and coronaviruses, MERS-CoV, HCoV-229E, and SARS-CoV-2 [435,436,437,438,439,440]. |
| Sulforaphane | 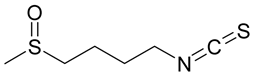 | Reduces HDAC activity [442] while increases the expression of acetylated histones H3 and H4 [442]. Upregulates let-7 expression, exhibits anti-IVA activity [202], and diminishes viral-induced immune cell activation in the lungs [446]. Inhibits replications of HCV and DENV [444] |
| Tanshinone IIA | 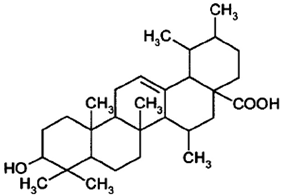 | Decreases the expression and activity of HDACs [449]. Inhibits MAPK p38 which resulted in, lowering the replications of a number of viruses including DENV [43], coronavirus [44], VEEV [45], EV71 [46], SFTSV, HSV-1, and SARS-CoV-2 [47,450]. |
| Ursolic acid | 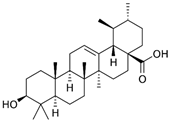 | Reduces the expression of HDAC1, HDAC2, HDAC3, HDAC8 (Class I), HDAC6, and HDAC7 (Class II) [452]. Exhibits anti-CMV activity [453]. |
| Withaferin A |  | Downregulates HDAC1 [454,455]. Decreases HMT activity, but enhances HAT activity [455]. Inhibits Mpro main protease of SARS-CoV-2 [456]. Attenuates H1N1 IVA [457] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gabbianelli, R.; Shahar, E.; de Simone, G.; Rucci, C.; Bordoni, L.; Feliziani, G.; Zhao, F.; Ferrati, M.; Maggi, F.; Spinozzi, E.; et al. Plant-Derived Epi-Nutraceuticals as Potential Broad-Spectrum Anti-Viral Agents. Nutrients 2023, 15, 4719. https://doi.org/10.3390/nu15224719
Gabbianelli R, Shahar E, de Simone G, Rucci C, Bordoni L, Feliziani G, Zhao F, Ferrati M, Maggi F, Spinozzi E, et al. Plant-Derived Epi-Nutraceuticals as Potential Broad-Spectrum Anti-Viral Agents. Nutrients. 2023; 15(22):4719. https://doi.org/10.3390/nu15224719
Chicago/Turabian StyleGabbianelli, Rosita, Ehud Shahar, Gaia de Simone, Chiara Rucci, Laura Bordoni, Giulia Feliziani, Fanrui Zhao, Marta Ferrati, Filippo Maggi, Eleonora Spinozzi, and et al. 2023. "Plant-Derived Epi-Nutraceuticals as Potential Broad-Spectrum Anti-Viral Agents" Nutrients 15, no. 22: 4719. https://doi.org/10.3390/nu15224719
APA StyleGabbianelli, R., Shahar, E., de Simone, G., Rucci, C., Bordoni, L., Feliziani, G., Zhao, F., Ferrati, M., Maggi, F., Spinozzi, E., & Mahajna, J. (2023). Plant-Derived Epi-Nutraceuticals as Potential Broad-Spectrum Anti-Viral Agents. Nutrients, 15(22), 4719. https://doi.org/10.3390/nu15224719









