Positive Effects of a Mediterranean Diet Supplemented with Almonds on Female Adipose Tissue Biology in Severe Obesity
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design and Subjects
2.2. Nutritional Intervention
2.3. Examinations and Calculations
2.4. Mixed Meal Tolerance Test
2.5. Body Composition
2.6. Adipose Tissue Biopsies
2.7. Fat Cell Area and Adipose Tissue Fibrosis
2.8. Gene Expression
2.9. Immunofluorescence
2.10. Immunoblotting
2.11. Circulating Levels
2.12. Statistical Analysis
3. Results
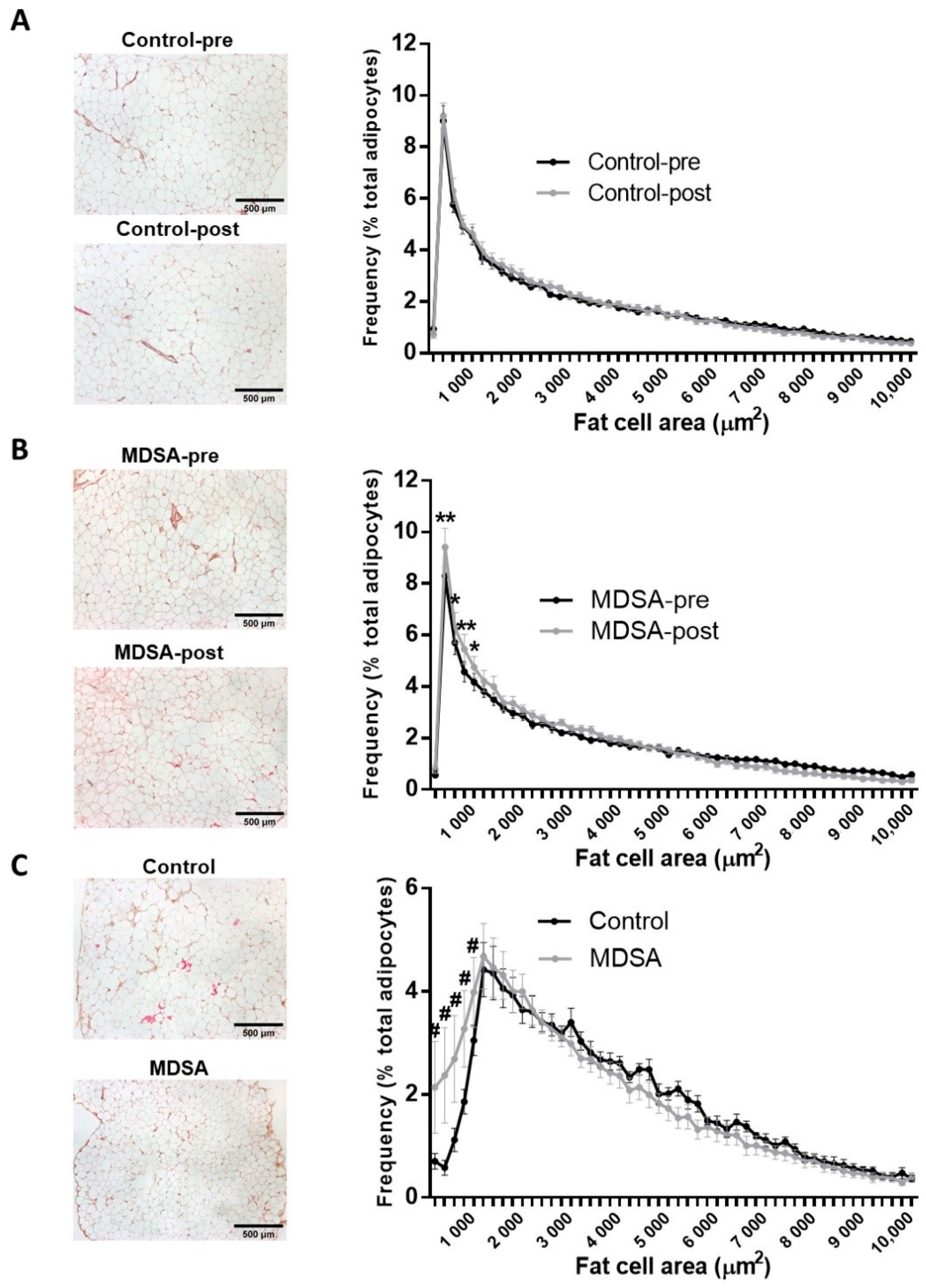
3.1. Adipose Tissue Morphology
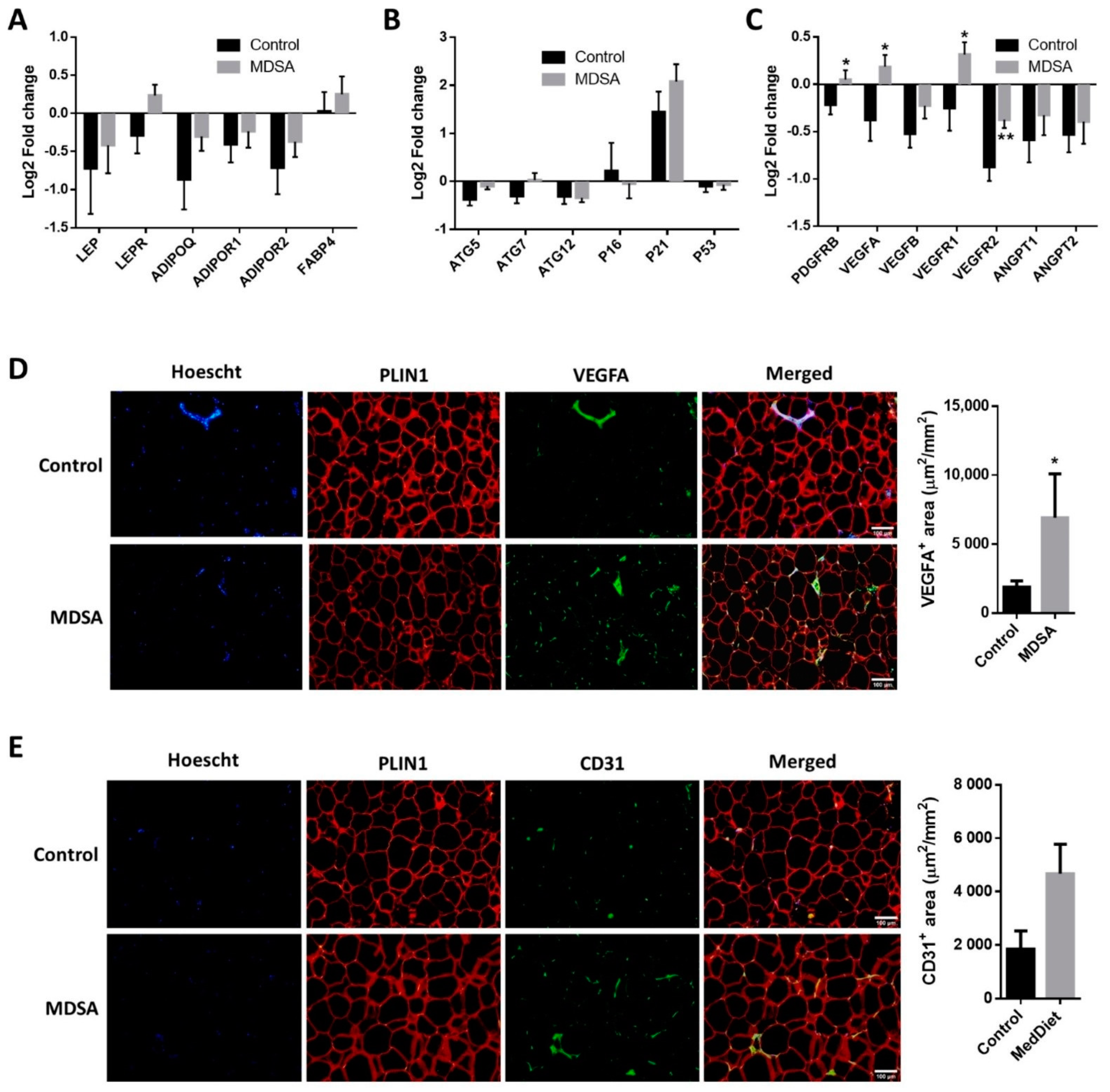
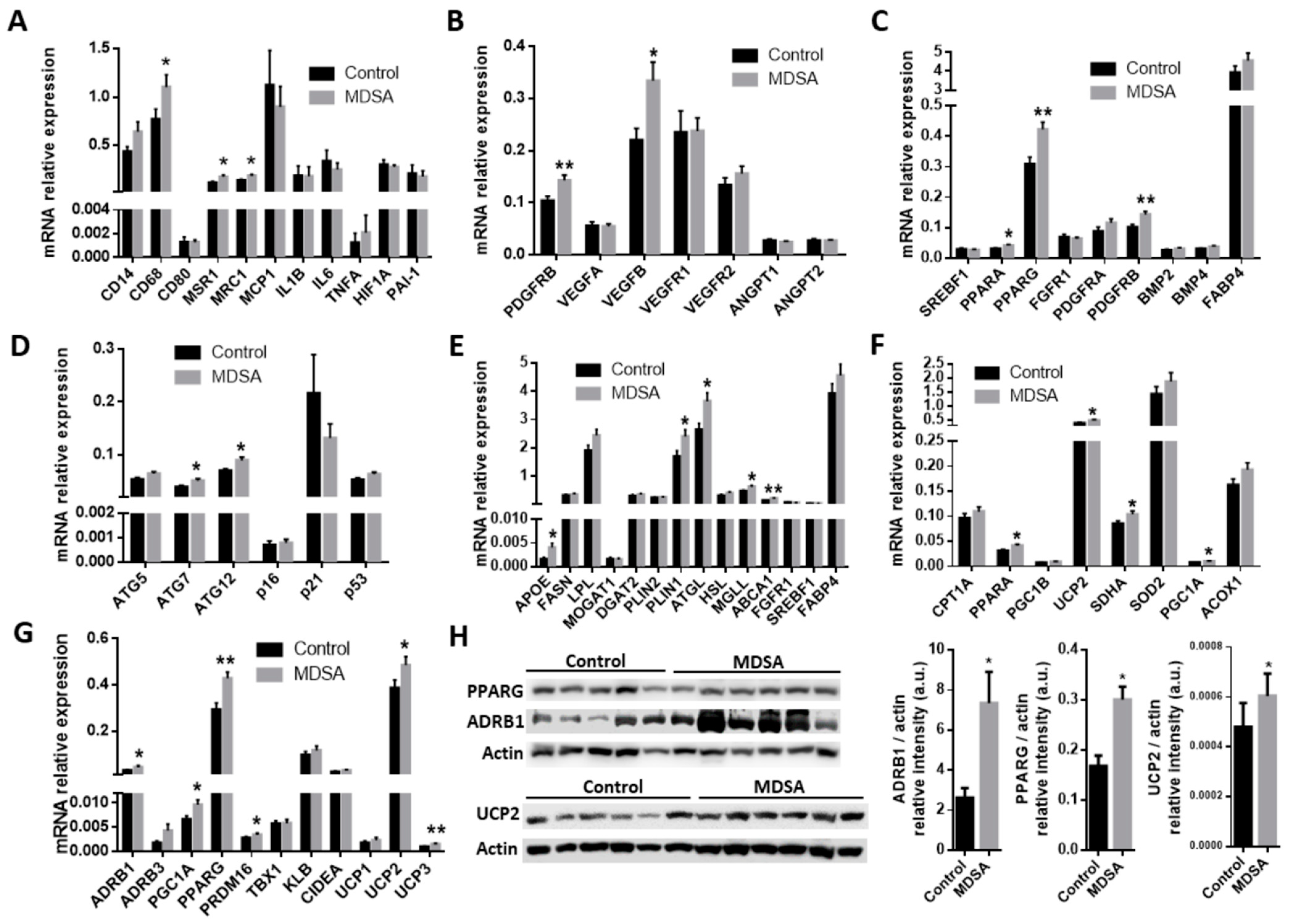
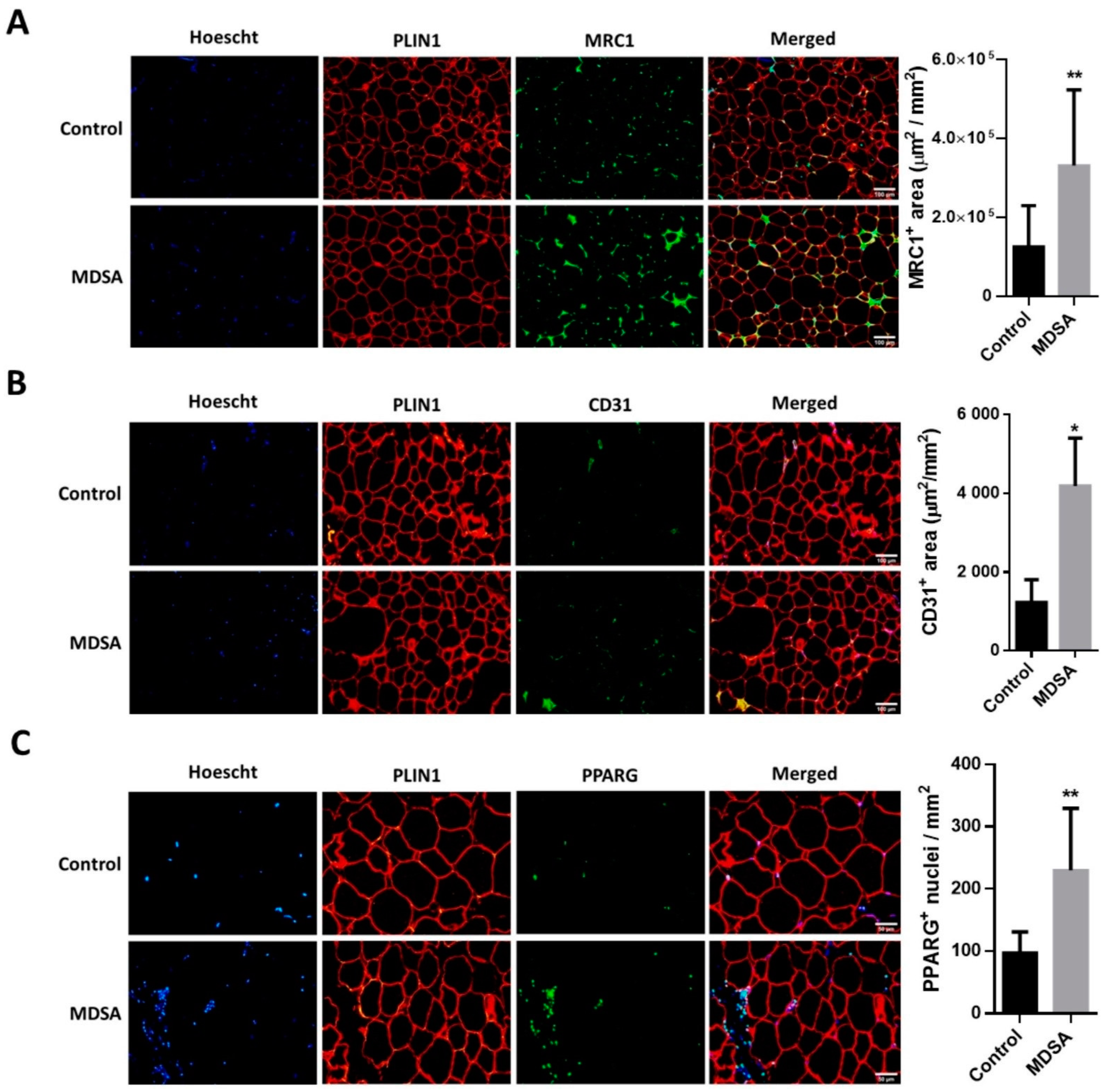
3.2. Changes in Gene Expression in SAT
3.3. Differences in Gene Expression in VAT
3.4. Clinical and Circulating Parameters
3.5. Associations with Mediterranean Diet Adherence Screener (MEDAS)
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Estruch, R.; Ros, E.; Salas-Salvadó, J.; Covas, M.-I.; Corella, D.; Arós, F.; Gómez-Gracia, E.; Ruiz-Gutiérrez, V.; Fiol, M.; Lapetra, J.; et al. Primary Prevention of Cardiovascular Disease with a Mediterranean Diet Supplemented with Extra-Virgin Olive Oil or Nuts. N. Engl. J. Med. 2018, 378, e34. [Google Scholar] [CrossRef]
- Konieczna, J.; Yañez, A.; Moñino, M.; Babio, N.; Toledo, E.; Martínez-González, M.A.; Sorlí, J.V.; Salas-Salvadó, J.; Estruch, R.; Ros, E.; et al. Longitudinal changes in Mediterranean diet and transition between different obesity phenotypes. Clin. Nutr. 2020, 39, 966–975. [Google Scholar] [CrossRef]
- Jayedi, A.; Mirzaei, K.; Rashidy-Pour, A.; Yekaninejad, M.S.; Zargar, M.-S.; Akbari Eidgahi, M.R. Dietary approaches to stop hypertension, mediterranean dietary pattern, and diabetic nephropathy in women with type 2 diabetes: A case-control study. Clin. Nutr. ESPEN 2019, 33, 164–170. [Google Scholar] [CrossRef]
- Jalilpiran, Y.; Darooghegi Mofrad, M.; Mozaffari, H.; Bellissimo, N.; Azadbakht, L. Adherence to dietary approaches to stop hypertension (DASH) and Mediterranean dietary patterns in relation to cardiovascular risk factors in older adults. Clin. Nutr. ESPEN 2020, 39, 87–95. [Google Scholar] [CrossRef]
- Salas-Salvadó, J.; Bulló, M.; Babio, N.; Martínez-González, M.Á.; Ibarrola-Jurado, N.; Basora, J.; Estruch, R.; Covas, M.I.; Corella, D.; Arós, F.; et al. Reduction in the incidence of type 2 diabetes with the mediterranean diet: Results of the PREDIMED-Reus nutrition intervention randomized trial. Diabetes Care 2011, 34, 14–19. [Google Scholar] [CrossRef] [Green Version]
- Ahmad, S.; Demler, O.V.; Sun, Q.; Moorthy, M.V.; Li, C.; Lee, I.-M.; Ridker, P.M.; Manson, J.E.; Hu, F.B.; Fall, T.; et al. Association of the Mediterranean Diet with Onset of Diabetes in the Women’s Health Study. JAMA Netw. open 2020, 3, e2025466. [Google Scholar] [CrossRef]
- Casas, R.; Sacanella, E.; Urpí-Sardà, M.; Chiva-Blanch, G.; Ros, E.; Martínez-González, M.-A.; Covas, M.-I.; Salas-Salvadó, J.; Fiol, M.; Arós, F.; et al. The Effects of the Mediterranean Diet on Biomarkers of Vascular Wall Inflammation and Plaque Vulnerability in Subjects with High Risk for Cardiovascular Disease. A Randomized Trial. PLoS ONE 2014, 9, e100084. [Google Scholar] [CrossRef] [Green Version]
- Ros, E. Contribution of Nuts to the Mediterranean Diet. In The Mediterranean Diet; Elsevier: Amsterdam, The Netherlands, 2015; pp. 175–184. ISBN 9780124079427. [Google Scholar]
- Damasceno, N.R.T.; Sala-Vila, A.; Cofán, M.; Pérez-Heras, A.M.; Fitó, M.; Ruiz-Gutiérrez, V.; Martínez-González, M.-Á.; Corella, D.; Arós, F.; Estruch, R.; et al. Mediterranean diet supplemented with nuts reduces waist circumference and shifts lipoprotein subfractions to a less atherogenic pattern in subjects at high cardiovascular risk. Atherosclerosis 2013, 230, 347–353. [Google Scholar] [CrossRef]
- Salas-Salvadó, J.; Fernández-Ballart, J.; Ros, E.; Martínez-González, M.-A.; Fitó, M.; Estruch, R.; Corella, D.; Fiol, M.; Gómez-Gracia, E.; Arós, F.; et al. Effect of a Mediterranean Diet Supplemented with Nuts on Metabolic Syndrome Status. Arch. Intern. Med. 2008, 168, 2449. [Google Scholar] [CrossRef] [Green Version]
- Tektonidis, T.G.; Åkesson, A.; Gigante, B.; Wolk, A.; Larsson, S.C. A Mediterranean diet and risk of myocardial infarction, heart failure and stroke: A population-based cohort study. Atherosclerosis 2015, 243, 93–98. [Google Scholar] [CrossRef]
- Estruch, R.; Martínez-González, M.A.; Corella, D.; Salas-Salvadó, J.; Ruiz-Gutiérrez, V.; Covas, M.I.; Fiol, M.; Gómez-Gracia, E.; López-Sabater, M.C.; Vinyoles, E.; et al. Effects of a Mediterranean-Style Diet on Cardiovascular Risk Factors a Randomized Trial. Ann. Intern. Med. 2006, 145, 1–11. [Google Scholar] [CrossRef]
- Bolling, B.W. Almond Polyphenols: Methods of Analysis, Contribution to Food Quality, and Health Promotion. Compr. Rev. Food Sci. Food Saf. 2017, 16, 346–368. [Google Scholar] [CrossRef] [Green Version]
- Garrido, I.; Urpi-Sarda, M.; Monagas, M.; Gómez-Cordovés, C.; Martín-Álvarez, P.J.; Llorach, R.; Bartolomé, B.; Andrés-Lacueva, C. Targeted analysis of conjugated and microbial-derived phenolic metabolites in human urine after consumption of an almond skin phenolic extract. J. Nutr. 2010, 140, 1799–1807. [Google Scholar] [CrossRef] [Green Version]
- Kamil, A.; Chen, C.Y.O. Health benefits of almonds beyond cholesterol reduction. J. Agric. Food Chem. 2012, 60, 6694–6702. [Google Scholar] [CrossRef]
- Chen, C.Y.O.; Holbrook, M.; Duess, M.A.; Dohadwala, M.M.; Hamburg, N.M.; Asztalos, B.F.; Milbury, P.E.; Blumberg, J.B.; Vita, J.A. Effect of almond consumption on vascular function in patients with coronary artery disease: A randomized, controlled, cross-over trial. Nutr. J. 2015, 14, 61. [Google Scholar] [CrossRef] [Green Version]
- Waniek, S.; di Giuseppe, R.; Plachta-Danielzik, S.; Ratjen, I.; Jacobs, G.; Koch, M.; Borggrefe, J.; Both, M.; Müller, H.P.; Kassubek, J.; et al. Association of Vitamin E Levels with Metabolic Syndrome, and MRI-Derived Body Fat Volumes and Liver Fat Content. Nutrients 2017, 9, 1143. [Google Scholar] [CrossRef] [Green Version]
- Zorena, K.; Jachimowicz-Duda, O.; Ślęzak, D.; Robakowska, M.; Mrugacz, M. Adipokines and obesity. Potential link to metabolic disorders and chronic complications. Int. J. Mol. Sci. 2020, 21, 3570. [Google Scholar] [CrossRef]
- Crewe, C.; An, Y.A.; Scherer, P.E. The ominous triad of adipose tissue dysfunction: Inflammation, fibrosis, and impaired angiogenesis. J. Clin. Investig. 2017, 127, 74–82. [Google Scholar] [CrossRef] [Green Version]
- Cancello, R.; Henegar, C.; Viguerie, N.; Taleb, S.; Poitou, C.; Rouault, C.; Coupaye, M.; Pelloux, V.; Hugol, D.; Bouillot, J.-L.; et al. Reduction of macrophage infiltration and chemoattractant gene expression changes in white adipose tissue of morbidly obese subjects after surgery-induced weight loss. Diabetes 2005, 54, 2277–2286. [Google Scholar] [CrossRef] [Green Version]
- Hill, A.A.; Reid Bolus, W.; Hasty, A.H. A decade of progress in adipose tissue macrophage biology. Immunol. Rev. 2014, 262, 134–152. [Google Scholar] [CrossRef] [Green Version]
- Mele, L.; Bidault, G.; Mena, P.; Crozier, A.; Brighenti, F.; Vidal-Puig, A.; Del Rio, D. Dietary (Poly)phenols, Brown Adipose Tissue Activation, and Energy Expenditure: A Narrative Review. Adv. Nutr. An Int. Rev. J. 2017, 8, 694–704. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- El Hadi, H.; Di Vincenzo, A.; Vettor, R.; Rossato, M. Food Ingredients Involved in White-to-Brown Adipose Tissue Conversion and in Calorie Burning. Front. Physiol. 2019, 9, 1954. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schröder, H.; Fitó, M.; Estruch, R.; Martínez-González, M.A.; Corella, D.; Salas-Salvadó, J.; Lamuela-Raventós, R.; Ros, E.; Salaverría, I.; Fiol, M.; et al. A short screener is valid for assessing Mediterranean diet adherence among older Spanish men and women. J. Nutr. 2011, 141, 1140–1145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Freitas-Simoes, T.-M.; Cofán, M.; Blasco, M.A.; Soberón, N.; Foronda, M.; Corella, D.; Asensio, E.M.; Serra-Mir, M.; Roth, I.; Calvo, C.; et al. The red blood cell proportion of arachidonic acid relates to shorter leukocyte telomeres in Mediterranean elders: A secondary analysis of a randomized controlled trial. Clin. Nutr. 2019, 38, 958–961. [Google Scholar] [CrossRef] [PubMed]
- Gutch, M.; Kumar, S.; Razi, S.M.; Gupta, K.K.; Gupta, A. Assessment of insulin sensitivity/resistance. Indian J. Endocrinol. Metab. 2015, 19, 160–164. [Google Scholar] [CrossRef] [PubMed]
- Spalding, K.L.; Bernard, S.; Näslund, E.; Salehpour, M.; Possnert, G.; Appelsved, L.; Fu, K.-Y.; Alkass, K.; Druid, H.; Thorell, A.; et al. Impact of fat mass and distribution on lipid turnover in human adipose tissue. Nat. Commun. 2017, 8, 15253. [Google Scholar] [CrossRef]
- Thangavel, N.; Al Bratty, M.; Akhtar Javed, S.; Ahsan, W.; Alhazmi, H.A. Targeting Peroxisome Proliferator-Activated Receptors Using Thiazolidinediones: Strategy for Design of Novel Antidiabetic Drugs. Int. J. Med. Chem. 2017, 2017, 1069718. [Google Scholar] [CrossRef] [Green Version]
- Aprile, M.; Ambrosio, M.R.; D’esposito, V.; Beguinot, F.; Formisano, P.; Costa, V.; Ciccodicola, A. Differential Contribution of Canonical Transcripts and Dominant Negative Isoforms. Ppar Res. 2014, 2014, 537865. [Google Scholar] [CrossRef] [Green Version]
- Li, P.; Zhu, Z.; Lu, Y.; Granneman, J.G. Metabolic and cellular plasticity in white adipose tissue II: Role of peroxisome proliferator-activated receptor-α. Am. J. Physiol. Endocrinol. Metab. 2005, 289, E617–E626. [Google Scholar] [CrossRef]
- Barquissau, V.; Beuzelin, D.; Pisani, D.F.; Beranger, G.E.; Mairal, A.; Montagner, A.; Roussel, B.; Tavernier, G.; Marques, M.-A.; Moro, C.; et al. White-to-brite conversion in human adipocytes promotes metabolic reprogramming towards fatty acid anabolic and catabolic pathways. Mol. Metab. 2016, 5, 352–365. [Google Scholar] [CrossRef]
- Bukowiecki, L.; Collet, A.J.; Follea, N. Brown adipose tissue hyperplasia: A fundamental mechanism of adaptation to cold and hyperphagia. Am. J. Physiol. Endocrinol. Metab. 1982, 5, E353–E359. [Google Scholar] [CrossRef] [PubMed]
- Collins, S.; Surwit, R.S. The β-adrenergic receptors and the control of adipose tissue metabolism and thermogenesis. Recent Prog. Horm. Res. 2001, 56, 309–328. [Google Scholar] [CrossRef] [PubMed]
- Riis-Vestergaard, M.J.; Richelsen, B.; Bruun, J.M.; Li, W.; Hansen, J.B.; Pedersen, S.B. Beta-1 and not Beta-3 adrenergic receptors may be the primary regulator of human brown adipocyte metabolism. J. Clin. Endocrinol. Metab. 2020, 105, e994–e1005. [Google Scholar] [CrossRef] [PubMed]
- Kajimura, S.; Spiegelman, B.M.; Seale, P. Brown and Beige Fat: Physiological Roles beyond Heat Generation. Cell Metab. 2015, 22, 546–559. [Google Scholar] [CrossRef] [Green Version]
- Ikeda, K.; Yamada, T. UCP1 Dependent and Independent Thermogenesis in Brown and Beige Adipocytes. Front. Endocrinol. 2020, 11, 498. [Google Scholar] [CrossRef]
- Puigserver, P.; Wu, Z.; Park, C.W.; Graves, R.; Wright, M.; Spiegelman, B.M. A cold-inducible coactivator of nuclear receptors linked to adaptive thermogenesis. Cell 1998, 92, 829–839. [Google Scholar] [CrossRef] [Green Version]
- Uldry, M.; Yang, W.; St-Pierre, J.; Lin, J.; Seale, P.; Spiegelman, B.M. Complementary action of the PGC-1 coactivators in mitochondrial biogenesis and brown fat differentiation. Cell Metab. 2006, 3, 333–341. [Google Scholar] [CrossRef] [Green Version]
- Vega, R.B.; Huss, J.M.; Kelly, D.P. The Coactivator PGC-1 Cooperates with Peroxisome Proliferator-Activated Receptor α in Transcriptional Control of Nuclear Genes Encoding Mitochondrial Fatty Acid Oxidation Enzymes. Mol. Cell. Biol. 2000, 20, 1868–1876. [Google Scholar] [CrossRef] [Green Version]
- Hondares, E.; Rosell, M.; Díaz-Delfín, J.; Olmos, Y.; Monsalve, M.; Iglesias, R.; Villarroya, F.; Giralt, M. Peroxisome proliferator-activated receptor α (PPARα) induces PPARγ coactivator 1α (PGC-1α) gene expression and contributes to thermogenic activation of brown fat: Involvement of PRDM16. J. Biol. Chem. 2011, 286, 43112–43122. [Google Scholar] [CrossRef] [Green Version]
- Seale, P. Transcriptional Regulatory Circuits Controlling Brown Fat Development and Activation. Diabetes 2015, 64, 2369–2375. [Google Scholar] [CrossRef] [Green Version]
- Vozza, A.; Parisi, G.; De Leonardis, F.; Lasorsa, F.M.; Castegna, A.; Amorese, D.; Marmo, R.; Calcagnile, V.M.; Palmieri, L.; Ricquier, D.; et al. UCP2 transports C4 metabolites out of mitochondria, regulating glucose and glutamine oxidation. Proc. Natl. Acad. Sci. USA 2014, 111, 960–965. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bouillaud, F.; Alves-Guerra, M.-C.; Ricquier, D. UCPs, at the interface between bioenergetics and metabolism. Biochim. Biophys. Acta 2016, 1863, 2443–2456. [Google Scholar] [CrossRef] [PubMed]
- Pecqueur, C.; Alves-Guerra, C.; Ricquier, D.; Bouillaud, F. UCP2, a metabolic sensor coupling glucose oxidation to mitochondrial metabolism? IUBMB Life 2009, 61, 762–767. [Google Scholar] [CrossRef] [PubMed]
- Cioffi, F.; Senese, R.; De Lange, P.; Goglia, F.; Lanni, A.; Lombardi, A. Uncoupling proteins: A complex journey to function discovery. BioFactors 2009, 35, 417–428. [Google Scholar] [CrossRef]
- Dulloo, A.G.; Samec, S. Uncoupling proteins: Their roles in adaptive thermogenesis and substrate metabolism reconsidered. Br. J. Nutr. 2001, 86, 123–139. [Google Scholar] [CrossRef] [Green Version]
- Oliveira, B.A.P.; Pinhel, M.A.S.; Nicoletti, C.F.; Oliveira, C.C.; Quinhoneiro, D.C.G.; Noronha, N.Y.; Marchini, J.S.; Marchry, A.J.; Junior, W.S.; Nonino, C.B. UCP1 and UCP3 expression is associated with lipid and carbohydrate oxidation and body composition. PLoS ONE 2016, 11, e0150811. [Google Scholar] [CrossRef]
- Hilse, K.E.; Rupprecht, A.; Egerbacher, M.; Bardakji, S.; Zimmermann, L.; Wulczyn, A.E.M.S.; Pohl, E.E. The Expression of Uncoupling Protein 3 Coincides with the Fatty Acid Oxidation Type of Metabolism in Adult Murine Heart. Front. Physiol. 2018, 9, 747. [Google Scholar] [CrossRef]
- Busiello, R.A.; Savarese, S.; Lombardi, A. Mitochondrial uncoupling proteins and energy metabolism. Front. Physiol. 2015, 6, 36. [Google Scholar] [CrossRef] [Green Version]
- Pohl, E.E.; Rupprecht, A.; Macher, G.; Hilse, K.E. Important trends in UCP3 investigation. Front. Physiol. 2019, 10, 470. [Google Scholar] [CrossRef]
- Babashamsi, M.M.; Koukhaloo, S.Z.; Halalkhor, S.; Salimi, A.; Babashamsi, M. ABCA1 and metabolic syndrome; a review of the ABCA1 role in HDL-VLDL production, insulin-glucose homeostasis, inflammation and obesity. Diabetes Metab. Syndr. Clin. Res. Rev. 2019, 13, 1529–1534. [Google Scholar] [CrossRef]
- Kypreos, K.E.; Karagiannides, I.; Fotiadou, E.H.; Karavia, E.A.; Brinkmeier, M.S.; Giakoumi, S.M.; Tsompanidi, E.M. Mechanisms of obesity and related pathologies: Role of apolipoprotein E in the development of obesity. FEBS J. 2009, 276, 5720–5728. [Google Scholar] [CrossRef] [PubMed]
- Razani, B.; Feng, C.; Coleman, T.; Emanuel, R.; Wen, H.; Hwang, S.; Ting, J.P.; Virgin, H.W.; Kastan, M.B.; Semenkovich, C.F. Autophagy links inflammasomes to atherosclerotic progression. Cell Metab. 2012, 15, 534–544. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Madrigal-Matute, J.; Cuervo, A.M. Regulation of Liver Metabolism by Autophagy. Gastroenterology 2016, 150, 328–339. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Soussi, H.; Reggio, S.; Alili, R.; Prado, C.; Mutel, S.; Pini, M.; Rouault, C.; Clément, K.; Dugail, I. DAPK2 downregulation associates with attenuated adipocyte autophagic clearance in human obesity. Diabetes 2015, 64, 3452–3463. [Google Scholar] [CrossRef] [Green Version]
- Corella, D.; Coltell, O.; Macian, F.; Ordovás, J.M. Advances in Understanding the Molecular Basis of the Mediterranean Diet Effect. Annu. Rev. Food Sci. Technol. 2018, 9, 227–249. [Google Scholar] [CrossRef] [PubMed]
- Morselli, E.; Mariño, G.; Bennetzen, M.V.; Eisenberg, T.; Megalou, E.; Schroeder, S.; Cabrera, S.; Bénit, P.; Rustin, P.; Criollo, A.; et al. Spermidine and resveratrol induce autophagy by distinct pathways converging on the acetylproteome. J. Cell Biol. 2011, 192, 615–629. [Google Scholar] [CrossRef] [Green Version]
- Knutson, M.D.; Leeuwenburgh, C. Resveratrol and novel potent activators of SIRT1: Effects on aging and age-related diseases. Nutr. Rev. 2008, 66, 591–596. [Google Scholar] [CrossRef]
- Rigacci, S. Olive Oil Phenols as Promising Multi-targeting Agents Against Alzheimer’s Disease. Adv. Exp. Med. Biol. 2015, 863, 1–20. [Google Scholar] [CrossRef]
- Barreca, D.; Nabavi, S.M.; Sureda, A.; Rasekhian, M.; Raciti, R.; Silva, A.S.; Annunziata, G.; Arnone, A.; Tenore, G.C.; Süntar, İ.; et al. Almonds (Prunus Dulcis Mill. D. A. Webb): A Source of Nutrients and Health-Promoting Compounds. Nutrients 2020, 12, 672. [Google Scholar] [CrossRef] [Green Version]
- Koloverou, E.; Panagiotakos, D.B.; Pitsavos, C.; Chrysohoou, C.; Georgousopoulou, E.N.; Grekas, A.; Christou, A.; Chatzigeorgiou, M.; Skoumas, I.; Tousoulis, D.; et al. Adherence to Mediterranean diet and 10-year incidence (2002–2012) of diabetes: Correlations with inflammatory and oxidative stress biomarkers in the ATTICA cohort study. Diabetes. Metab. Res. Rev. 2016, 32, 73–81. [Google Scholar] [CrossRef]
- Barbagallo, C.M.; Cefalù, A.B.; Gallo, S.; Rizzo, M.; Noto, D.; Cavera, G.; Rao Camemi, A.; Marino, G.; Caldarella, R.; Notarbartolo, A.; et al. Effects of Mediterranean diet on lipid levels and cardiovascular risk in renal transplant recipients. Nephron 1999, 82, 199–204. [Google Scholar] [CrossRef] [PubMed]
- Casas, R.; Urpi-Sardà, M.; Sacanella, E.; Arranz, S.; Corella, D.; Castañer, O.; Lamuela-Raventós, R.M.; Salas-Salvadó, J.; Lapetra, J.; Portillo, M.P.; et al. Anti-Inflammatory Effects of the Mediterranean Diet in the Early and Late Stages of Atheroma Plaque Development. Mediat. Inflamm. 2017, 2017, 3674390. [Google Scholar] [CrossRef] [PubMed]
| MDSA Group (n = 19) | Control Group (n = 17) | p-Value | p-Value | |||
|---|---|---|---|---|---|---|
| Baseline | 3 Months | Baseline | 3 Months | Time | Time * Group | |
| Age (years) | 48.9 ± 11.0 | - | 45.3 ± 11.8 | - | ||
| BMI (kg/m2) | 43.8 ± 4.0 | 44.0 ± 3.8 | 45.7 ± 3.0 | 45.8 ± 3.1 | 0.276 | 0.769 |
| Weight (kg) | 110 ± 11.3 | 111 ± 11.7 | 110 ± 11.3 | 110 ± 11.9 | 0.243 | 0.643 |
| S-BP (mmHg) | 135 ± 15 | 132 ± 18 | 133 ± 16 | 131 ± 13 | 0.429 | 0.951 |
| D-BP (mmHg) | 86 ± 9.2 | 83 ± 9.5 | 90.3 ± 13 | 83 ± 8.0 | 0.061 | 0.341 |
| FM (%) | 52.8 ± 4.3 | 53.1 ± 3.2 | 55.1 ± 3.0 | 54.9 ± 3.0 | 0.705 | 0.274 |
| eVAT (gr) | 2351 ± 912 | 2261 ± 693 | 2396 ± 953 | 2262 ± 663 | 0.867 | 0.875 |
| eVAT (cm2) | 2492 ± 966 | 2540 ± 1010 | 2397 ± 734 | 2398 ± 703 | 0.866 | 0.874 |
| Total Cholesterol (mg/dL) | 187 ± 32 | 175 ± 24 | 200 ± 32 | 207 ± 32 | 0.549 | 0.028 |
| HDL-c (mg/dL) | 46.3 ± 8.4 | 46.6 ± 8.0 | 48.2 ± 8.3 | 49.7 ± 7.2 | 0.353 | 0.523 |
| LDL-c (mg/dL) | 120 ± 33 | 105 ± 19 | 125 ± 25 | 132 ± 30 | 0.364 | 0.012 |
| Triglycerides (mg/dL) | 124 ± 47 | 118 ± 30 | 144 ± 42.4 | 151.7 ± 61 | 0.904 | 0.384 |
| FPG (mg/dL) | 106 ± 14.0 | 106 ± 12.4 | 100 ± 10.6 | 100 ± 15.1 | 0.837 | 0.862 |
| Insulin (uU/L) | 20.1 ± 6.5 | 20.3 ± 6.2 | 24.3 ± 13.4 | 24.1 ± 14.1 | 0.998 | 0.894 |
| HOMA-IR | 5.3 ± 2.1 | 5.4 ± 2.0 | 6.2 ± 4.0 | 6.2 ± 4.1 | 0.985 | 0.884 |
| Matsuda Index | 2.3 ± 1.0 | 2.1 ± 1.0 | 2.2 ± 1.8 | 2.1 ± 1.4 | 0.354 | 0.9 |
| Insulinogenic Index | 2.9 ± 1.6 | 3.5 ± 2.1 | 3.3 ± 2.2 | 3.2 ± 1.7 | 0.531 | 0.174 |
| Disposition Index | 7.1 ± 5.1 | 7.4 ± 5.4 | 6.7 ± 5.6 | 6.5 ± 5.7 | 0.938 | 0.652 |
| hsCRP (mg/dL) | 0.8 ± 0.7 | 0.7 ± 0.6 | 0.8 ± 0.6 | 1.2 ± 1.3 | 0.271 | 0.142 |
| GM-CSF (pg/mL) | 26.6 ± 21.0 | 14.9 ± 7.7 | 15.8 ± 9.9 | 17.7 ± 13.7 | 0.089 | 0.018 |
| IFNγ (pg/mL) | 5.7 ± 4.5 | 3.2 ± 1.7 | 3.6 ± 3.0 | 4.3 ± 4.1 | 0.164 | 0.014 |
| IL6 (pg/mL) * | 2.1 ± 1.4 | 1.5 ± 1.0 | 1.2 ± 0.5 | 1.2 ± 0.8 | 0.103 | 0.122 |
| TNFα (pg/mL) | 4.2 ± 2.4 | 3.1 ± 1.2 | 3.6 ± 1.6 | 3.8 ± 1.9 | 0.188 | 0.082 |
| IL1ß (pg/mL) * | 1.2 ± 1.0 | 0.7 ± 0.3 | 0.7 ± 0.3 | 0.7 ± 0.4 | 0.049 | 0.024 |
| Selectin (pg/mL) | 87.2 ± 40.5 | 85.9 ± 45.34 | 86.8 ± 36.3 | 83.9 ± 27.2 | 0.391 | 0.749 |
| Adiponectin (ug/mL) | 19.3 ± 10.5 | 20.4 ± 10.1 | 25.1 ± 13.5 | 24.3 ± 15.9 | 0.841 | 0.22 |
| sICAM-1 (ng/mL) | 176 ± 110 | 163 ± 80.4 | 178 ± 91 | 170 ± 86.7 | 0.332 | 0.831 |
| sVCAM-1 (ng/mL) | 679 ± 162 | 633 ± 148 | 683 ± 141 | 666 ± 115 | 0.303 | 0.618 |
| SAA (ug/mL) | 26.9 ± 26.3 | 38.7 ± 42.7 | 62.3 ± 68.6 | 60.6 ± 62 | 0.489 | 0.351 |
| Correlation Coefficient (95%CI) | p-Value | |
|---|---|---|
| VAT-ABCA1 | ||
| Unadjusted * | 0.636 (0.269, 0.842) | 0.002 |
| Adjusted ** | 36.62 (13.07, 60.16) | 0.004 |
| VAT-UCP3 | ||
| Unadjusted * | 0.579 (0.169, 0.818) | 0.007 |
| Adjusted ** | 1596.6 (−678.3, 3871.6) | 0.160 |
| sICAM-1 (ng/mL) | ||
| Unadjusted * | −0.375 (−0.659, 0.0009) | 0.04 |
| Adjusted ** | −0.008 (−0.024, 0.009) | 0.357 |
| VAT-PPARA | ||
| Unadjusted * | 0.350 (−0.109, 0.686) | 0.119 |
| Adjusted ** | 173.29 (21.41, 325.1) | 0.027 |
| VAT-PGC1A | ||
| Unadjusted * | 0.232 (−0.165, 0.564) | 0.234 |
| Adjusted ** | 539.9 (109.5, 970.3) | 0.016 |
| VAT-ADRB3 | ||
| Unadjusted * | 0.291 (−0.103, 0.606) | 0.132 |
| Adjusted ** | 411.7 (60.24, 763.2) | 0.023 |
| VAT-ADRB1 | ||
| Unadjusted * | 0.236 (−0.161, 0.568) | 0.225 |
| Adjusted ** | 69.17 (9.218, 129.14) | 0.025 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Osorio-Conles, Ó.; Olbeyra, R.; Moizé, V.; Ibarzabal, A.; Giró, O.; Viaplana, J.; Jiménez, A.; Vidal, J.; de Hollanda, A. Positive Effects of a Mediterranean Diet Supplemented with Almonds on Female Adipose Tissue Biology in Severe Obesity. Nutrients 2022, 14, 2617. https://doi.org/10.3390/nu14132617
Osorio-Conles Ó, Olbeyra R, Moizé V, Ibarzabal A, Giró O, Viaplana J, Jiménez A, Vidal J, de Hollanda A. Positive Effects of a Mediterranean Diet Supplemented with Almonds on Female Adipose Tissue Biology in Severe Obesity. Nutrients. 2022; 14(13):2617. https://doi.org/10.3390/nu14132617
Chicago/Turabian StyleOsorio-Conles, Óscar, Romina Olbeyra, Violeta Moizé, Ainitze Ibarzabal, Oriol Giró, Judith Viaplana, Amanda Jiménez, Josep Vidal, and Ana de Hollanda. 2022. "Positive Effects of a Mediterranean Diet Supplemented with Almonds on Female Adipose Tissue Biology in Severe Obesity" Nutrients 14, no. 13: 2617. https://doi.org/10.3390/nu14132617
APA StyleOsorio-Conles, Ó., Olbeyra, R., Moizé, V., Ibarzabal, A., Giró, O., Viaplana, J., Jiménez, A., Vidal, J., & de Hollanda, A. (2022). Positive Effects of a Mediterranean Diet Supplemented with Almonds on Female Adipose Tissue Biology in Severe Obesity. Nutrients, 14(13), 2617. https://doi.org/10.3390/nu14132617







