The Associations between Diet and Socioeconomic Disparities and the Intestinal Microbiome in Preadolescence
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Population and Design
2.2. Data Collection and Definition of the Study Variables
2.3. Dietary Intake Questionnaire
2.4. The Dependent Variable
2.5. The Main Independent Variable
2.6. Co-Variates
2.7. Data Management
2.8. Sample Collection, DNA Extraction and Bacterial DNA Amplification
2.9. Statistical Analyses
2.10. Ethical Aspects
3. Results
3.1. Demographic Characteristics and Dietary Intake
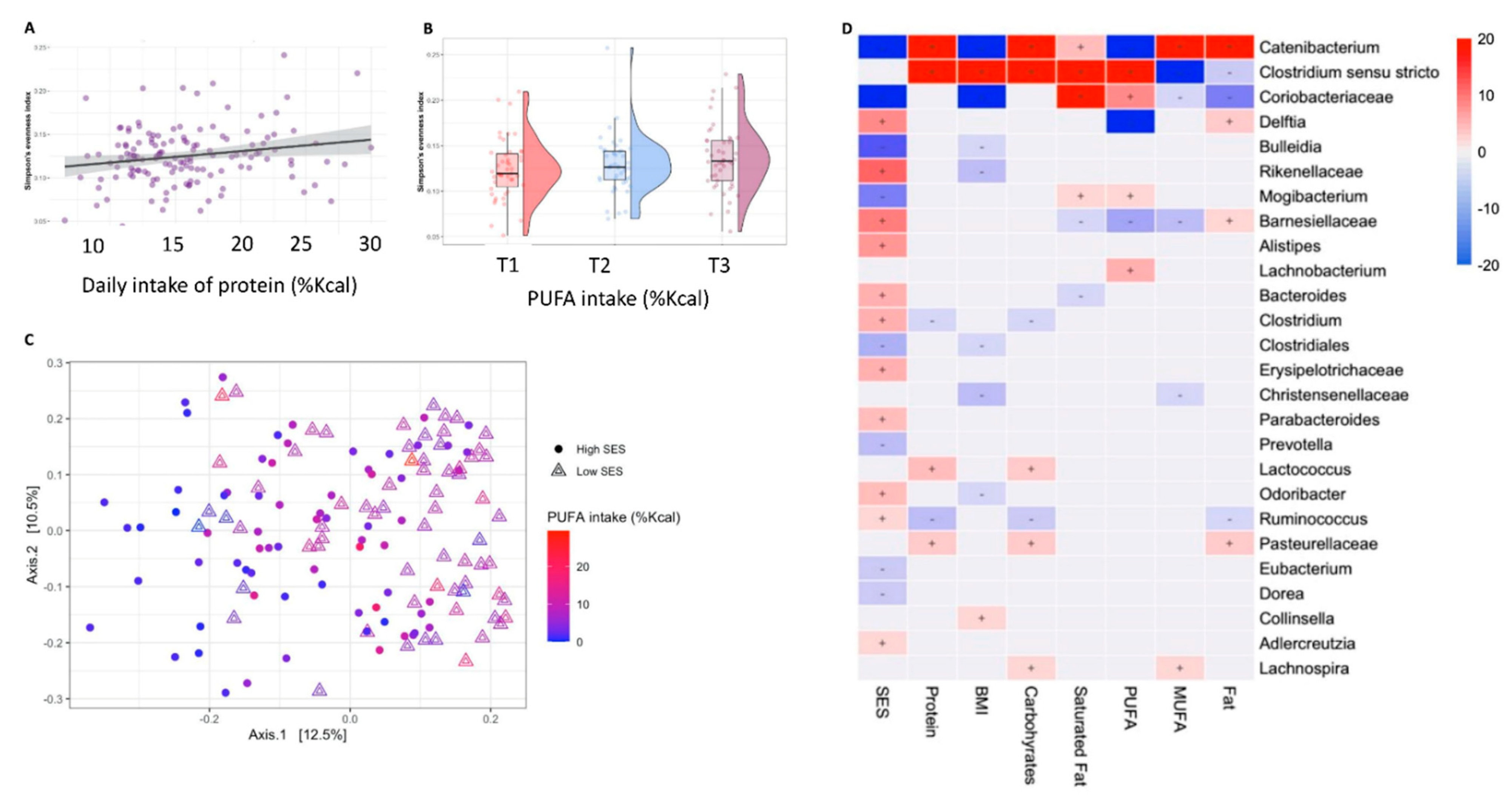
3.2. Microbial Diversity and Composition
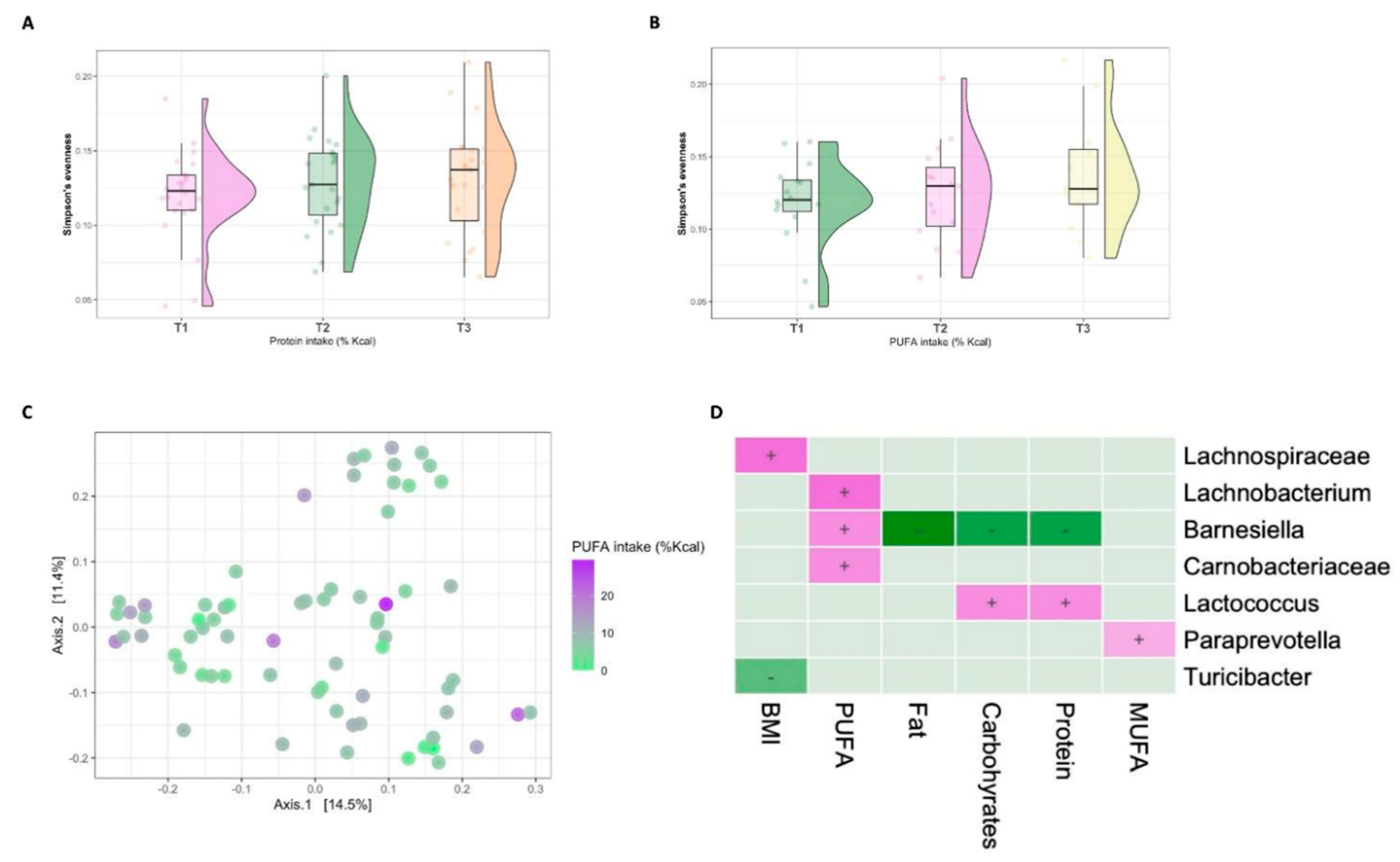
3.3. Diet–Microbiome Relationships within the Low-SES Subgroup
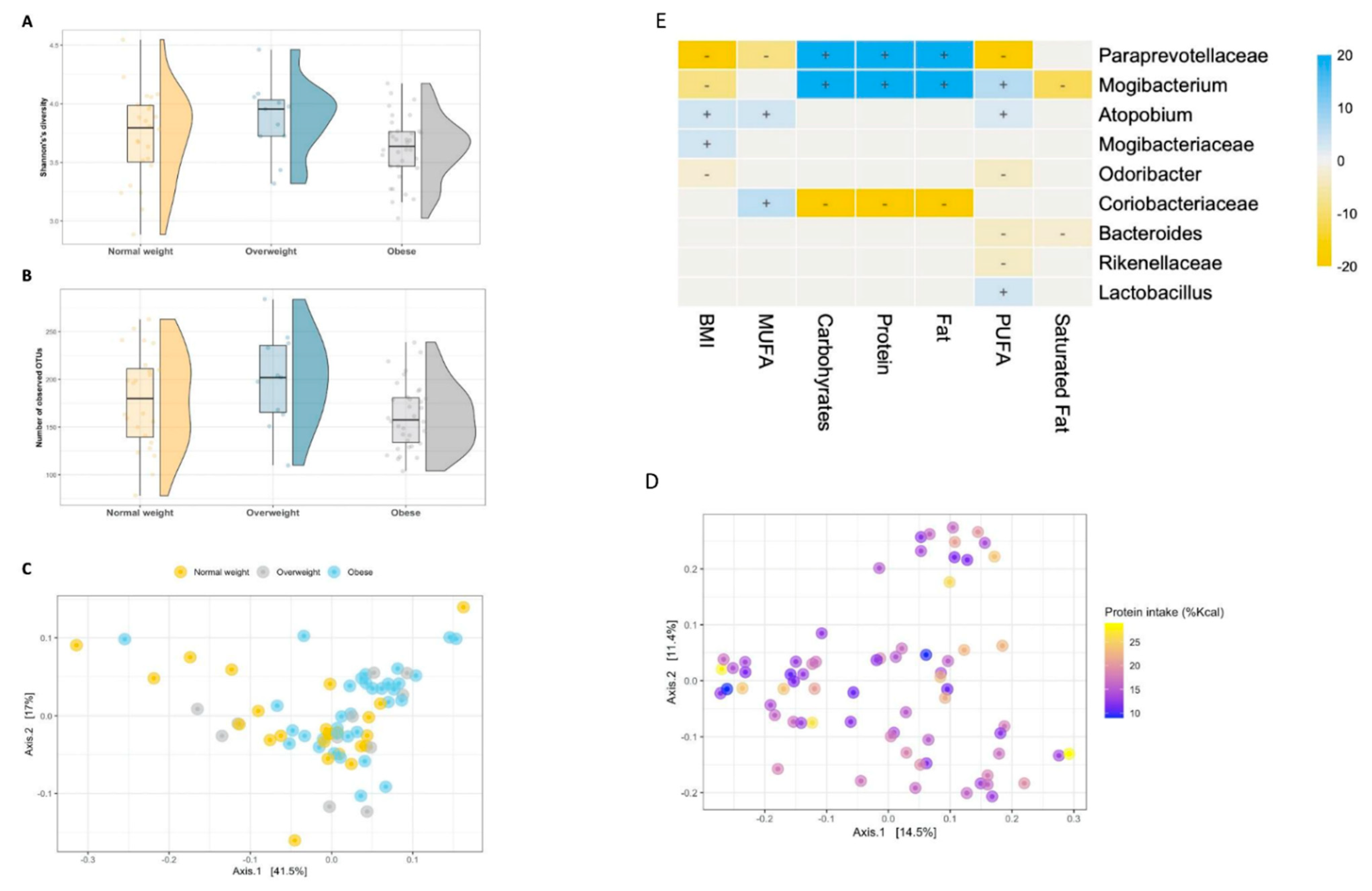
3.4. Diet–Microbiome Relationships within the High-SES Subgroup
3.5. The Intestinal Microbiome of Overweight/Obese Children
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Stewart, C.J.; Ajami, N.J.; O’Brien, J.L.; Hutchinson, D.S.; Smith, D.P.; Wong, M.C.; Ross, M.C.; Lloyd, R.E.; Doddapaneni, H.; Metcalf, G.A.; et al. Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 2018, 562, 583–588. [Google Scholar] [CrossRef] [PubMed]
- Hollister, E.B.; Riehle, K.; Luna, R.A.; Weidler, E.M.; Rubio-Gonzales, M.; Mistretta, T.-A.; Raza, S.; Doddapaneni, H.V.; Metcalf, G.A.; Muzny, D.M.; et al. Structure and function of the healthy pre-adolescent pediatric gut microbiome. Microbiome 2015, 3, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Derrien, M.; Alvarez, A.-S.; de Vos, W.M. The Gut Microbiota in the First Decade of Life. Trends Microbiol. 2019, 27, 997–1010. [Google Scholar] [CrossRef] [Green Version]
- Carlson, A.; Xia, K.; Azcarate-Peril, M.A.; Goldman, B.D.; Ahn, M.; Styner, M.A.; Thompson, A.L.; Geng, X.; Gilmore, J.H.; Knickmeyer, R.C. Infant Gut Microbiome Associated with Cognitive Development. Biol. Psychiatry 2018, 83, 148–159. [Google Scholar] [CrossRef]
- Long-Smith, C.; O’Riordan, K.J.; Clarke, G.; Stanton, C.; Dinan, T.G.; Cryan, J.F. Microbiota-Gut-Brain Axis: New Therapeutic Opportunities. Annu. Rev. Pharmacol. Toxicol. 2020, 60, 477–502. [Google Scholar] [CrossRef] [Green Version]
- Li, B.; Selmi, C.; Tang, R.; Gershwin, M.E.; Ma, X. The microbiome and autoimmunity: A paradigm from the gut–liver axis. Cell. Mol. Immunol. 2018, 15, 595–609. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tripathi, A.; Debelius, J.; Brenner, D.A.; Karin, M.; Loomba, R.; Schnabl, B.; Knight, R. The gut–liver axis and the intersection with the microbiome. Nat. Rev. Gastroenterol. Hepatol. 2018, 15, 397–411. [Google Scholar] [CrossRef]
- Stokholm, J.; Blaser, M.J.; Thorsen, J.; Rasmussen, M.A.; Waage, J.; Vinding, R.K.; Schoos, A.-M.M.; Kunøe, A.; Fink, N.R.; Chawes, B.; et al. Maturation of the gut microbiome and risk of asthma in childhood. Nat. Commun. 2018, 9, 141. [Google Scholar] [CrossRef] [PubMed]
- Mohammadkhah, A.I.; Simpson, E.B.; Patterson, S.G.; Ferguson, J.F. Development of the Gut Microbiome in Children, and Lifetime Implications for Obesity and Cardiometabolic Disease. Children 2018, 5, 160. [Google Scholar] [CrossRef] [Green Version]
- Rampelli, S.; Guenther, K.; Turroni, S.; Wolters, M.; Veidebaum, T.; Kourides, Y.; Molnár, D.; Lissner, L.; Benitez-Paez, A.; Sanz, Y.; et al. Pre-obese children’s dysbiotic gut microbiome and unhealthy diets may predict the development of obesity. Commun. Biol. 2018, 1, 222. [Google Scholar] [CrossRef] [PubMed]
- David, L.A.; Maurice, C.F.; Carmody, R.N.; Gootenberg, D.B.; Button, J.E.; Wolfe, B.E.; Ling, A.V.; Devlin, A.S.; Varma, Y.; Fischbach, M.A.; et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 2014, 505, 559–563. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davis, E.C.; Dinsmoor, A.M.; Wang, M.; Donovan, S.M. Microbiome Composition in Pediatric Populations from Birth to Adolescence: Impact of Diet and Prebiotic and Probiotic Interventions. Dig. Dis. Sci. 2020, 65, 706–722. [Google Scholar] [CrossRef] [Green Version]
- Chen, E.; Martin, A.; Matthews, K.A. Understanding Health Disparities: The Role of Race and Socioeconomic Status in Children’s Health. Am. J. Public Health 2006, 96, 702–708. [Google Scholar] [CrossRef] [PubMed]
- Stormacq, C.; Van Den Broucke, S.; Wosinski, J. Does health literacy mediate the relationship between socioeconomic status and health disparities? Integrative review. Health Promot. Int. 2019, 34, e1–e17. [Google Scholar] [CrossRef]
- Lapidot, Y. Socioeconomic disparities and household crowding affect the fecal microbiome of school-age children. NPJ Biofilms Microbiomes 2021. under review. [Google Scholar]
- Bowyer, R.C.E.; Jackson, M.A.; Le Roy, C.I.; Ni Lochlainn, M.; Spector, T.D.; Dowd, J.B.; Steves, C.J. Socioeconomic Status and the Gut Microbiome: A TwinsUK Cohort Study. Microorganisms 2019, 7, 17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miller, G.E.; Engen, P.A.; Gillevet, P.M.; Shaikh, M.; Sikaroodi, M.; Forsyth, C.B.; Mutlu, E.; Keshavarzian, A. Lower Neighborhood Socioeconomic Status Associated with Reduced Diversity of the Colonic Microbiota in Healthy Adults. PLoS ONE 2016, 11, e0148952. [Google Scholar] [CrossRef]
- Israel Central Bureau of Statistics. Localities, Their Population and Additional Information; Israel Central Bureau of Statistics: Jerusalem, Israel, 2017.
- Burck, L.; Feinstein, Y. Characterization and Classification of Geographical Units by the Socioeconomic Level of the Population; Israel Central Bureau of Statistics: Jerusalem, Israel, 2000. Available online: https://www.cbs.gov.il/en/publications/Pages/2017/Characterization-and-Classification-of-Geographical-Units-by-the-Socio-Economic-Level-of-the-Population-2013.aspx (accessed on 10 June 2021).
- Israel Central Bureau of Statistics. Characterization and Classification of Local Authorities by the Socio-Economic Level of the Population 2006; Israel Central Bureau of Statistics: Jerusalem, Israel, 2009. Available online: Http://Www.Cbs.Gov.Il/Www/Publications/Local (accessed on 6 January 2021).
- Abu-Saad, K.; Murad, H.; Lubin, F.; Freedman, L.S.; Ziv, A.; Alpert, G.; Atamna, A.; Kalter-Leibovici, O. Jews and Arabs in the Same Region in Israel Exhibit Major Differences in Dietary Patterns. J. Nutr. 2012, 142, 2175–2181. [Google Scholar] [CrossRef] [PubMed]
- Israel Center for Disease Control. Mabat Youth Second National Health and Nutrition Survey of 7th–12th Grade Students 2015–2016; Ministry of Health: Tel Hashomer, Israel, 2017; Volume 373, pp. 16E–19E.
- Muhsen, K.; Jurban, M.; Goren, S.; Cohen, D. Incidence, Age of Acquisition and Risk Factors of Helicobacter pylori Infection among Israeli Arab Infants. J. Trop. Pediatr. 2011, 58, 208–213. [Google Scholar] [CrossRef]
- World Health Organization. BMI-for-Age (5–19 years). 2016. Available online: https://www.who.int/tools/growth-reference-data-for-5to19-years/indicators/bmi-for-age (accessed on 10 June 2021).
- De Onis, M.; Onyango, A.W.; Borghi, E.; Siyam, A.; Nishida, C.; Siekmann, J. Development of a WHO growth reference for school-aged children and adolescents. Bull. World Health Organ. 2007, 85, 660–667. [Google Scholar] [CrossRef]
- Onis, M. WHO Child Growth Standards: Length/Height-for-Age, Weight-for-Age, Weight-for-Length, Weight-for-Height and Body Mass Index-for-Age: Methods and Development; WHO: Geneva, Switzerland, 2006; Available online: https://www.who.int/publications/i/item/924154693X (accessed on 10 June 2021).
- Amir, A.; McDonald, D.; Navas-Molina, J.A.; Kopylova, E.; Morton, J.T.; Xu, Z.Z.; Kightley, E.P.; Thompson, L.R.; Hyde, E.R.; Gonzalez, A.; et al. Deblur Rapidly Resolves Single-Nucleotide Community Sequence Patterns. mSystems 2017, 2, e00191-16. [Google Scholar] [CrossRef] [Green Version]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- McDonald, D.; Price, M.N.; Goodrich, J.; Nawrocki, E.P.; DeSantis, T.Z.; Probst, A.; Andersen, G.L.; Knight, R.; Hugenholtz, P. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2011, 6, 610–618. [Google Scholar] [CrossRef]
- Households and Families: Demographic Characteristics 2018 Based on Labour Force Surveys. Available online: https://www.cbs.gov.il/en/publications/Pages/2020/HOUSEHOLDS-FAMILIES-LabourForce-2018.aspx (accessed on 18 August 2020).
- Oksanen, J.; Blanchet, F.G.; Friendly, M. Vegan: Community Ecology Package | McGlinn Lab. 2019. Available online: https://www.mcglinnlab.org/publication/2019-01-01_oksanen_vegan_2019/ (accessed on 10 November 2020).
- Mallick, H.; Rahnavard, A.; McIver, L.J.; Ma, S.; Zhang, Y.; Nguyen, L.H.; Tickle, T.L.; Weingart, G.; Ren, B.; Schwager, E.H. Multivariable association discovery in population-scale meta-omics studies. bioRxiv 2021. [Google Scholar] [CrossRef]
- Robinson, M.D.; Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010, 11, R25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lkhagva, E.; Chung, H.-J.; Hong, J.; Tang, W.H.W.; Lee, S.-I.; Hong, S.-T.; Lee, S. The regional diversity of gut microbiome along the GI tract of male C57BL/6 mice. BMC Microbiol. 2021, 21, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Clemente, J.C.; Ursell, L.K.; Parfrey, L.W.; Knight, R. The Impact of the Gut Microbiota on Human Health: An Integrative View. Cell 2012, 148, 1258–1270. [Google Scholar] [CrossRef] [Green Version]
- Thaiss, C.A.; Zmora, N.; Levy, M.; Elinav, E. The microbiome and innate immunity. Nature 2016, 535, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Komaroff, A.L. The Microbiome and Risk for Obesity and Diabetes. JAMA 2017, 317, 355–356. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef]
- Sanchez-Rodriguez, E.; Egea-Zorrilla, A.; Plaza-Díaz, J.; Aragón-Vela, J.; Muñoz-Quezada, S.; Tercedor-Sánchez, L.; Abadia-Molina, F. The Gut Microbiota and Its Implication in the Development of Atherosclerosis and Related Cardiovascular Diseases. Nutrients 2020, 12, 605. [Google Scholar] [CrossRef] [Green Version]
- Karaduta, O.; Glazko, G.; Dvanajscak, Z.; Arthur, J.; Mackintosh, S.; Orr, L.; Rahmatallah, Y.; Yeruva, L.; Tackett, A.; Zybailov, B. Resistant starch slows the progression of CKD in the 5/6 nephrectomy mouse model. Physiol. Rep. 2020, 8. [Google Scholar] [CrossRef] [PubMed]
- Osadchiy, V.; Martin, C.R.; Mayer, E.A. The gut-brain axis and the microbiome: Mechanisms and clinical implications. Clin. Gastroenterol. Hepatol. 2019, 17, 322–332. [Google Scholar] [CrossRef] [PubMed]
- Jiang, T.; Wu, H.; Yang, X.; Li, Y.; Zhang, Z.; Chen, F.; Zhao, L.; Zhang, C. Lactobacillus Mucosae Strain Promoted by a High-Fiber Diet in Genetic Obese Child Alleviates Lipid Metabolism and Modifies Gut Microbiota in ApoE−/− Mice on a Western Diet. Microorganisms 2020, 8, 1225. [Google Scholar] [CrossRef] [PubMed]
- Scher, J.U.; Sczesnak, A.; Longman, R.S.; Segata, N.; Ubeda, C.; Bielski, C.; Rostron, T.; Cerundolo, V.; Pamer, E.G.; Abramson, S.B.; et al. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. eLife 2013, 2, e01202. [Google Scholar] [CrossRef]
- Dillon, S.; Lee, E.J.; Kotter, C.V.; Austin, G.; Dong, Z.; Hecht, D.K.; Gianella, S.; Siewe, B.; Smith, D.; Landay, A.L.; et al. An altered intestinal mucosal microbiome in HIV-1 infection is associated with mucosal and systemic immune activation and endotoxemia. Mucosal Immunol. 2014, 7, 983–994. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wen, C.; Zheng, Z.; Shao, T.; Liu, L.; Xie, Z.; Le Chatelier, E.; He, Z.; Zhong, W.; Fan, Y.; Zhang, L.; et al. Quantitative metagenomics reveals unique gut microbiome biomarkers in ankylosing spondylitis. Genome Biol. 2017, 18, 142. [Google Scholar] [CrossRef]
- Pedersen, H.K.; Gudmundsdottir, V.; Nielsen, H.B.; Hyotylainen, T.; Nielsen, T.; Jensen, B.A.H.; Forslund, K.; Hildebrand, F.; Prifti, E.; Falony, G.; et al. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature 2016, 535, 376–381. [Google Scholar] [CrossRef]
- Kovatcheva-Datchary, P.; Nilsson, A.; Akrami, R.; Lee, Y.S.; De Vadder, F.; Arora, T.; Hallen, A.; Martens, E.; Björck, I.; Bäckhed, F. Dietary Fiber-Induced Improvement in Glucose Metabolism Is Associated with Increased Abundance of Prevotella. Cell Metab. 2015, 22, 971–982. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Vadder, F.; Kovatcheva-Datchary, P.; Zitoun, C.; Duchampt, A.; Bäckhed, F.; Mithieux, G. Microbiota-Produced Succinate Improves Glucose Homeostasis via Intestinal Gluconeogenesis. Cell Metab. 2016, 24, 151–157. [Google Scholar] [CrossRef] [Green Version]
- Pasolli, E.; Asnicar, F.; Manara, S.; Zolfo, M.; Karcher, N.; Armanini, F.; Beghini, F.; Manghi, P.; Tett, A.; Ghensi, P.; et al. Extensive Unexplored Human Microbiome Diversity Revealed by Over 150,000 Genomes from Metagenomes Spanning Age, Geography, and Lifestyle. Cell 2019, 176, 649–662. [Google Scholar] [CrossRef] [Green Version]
- Brewster, R.; Tamburini, F.B.; Asiimwe, E.; Achinike-Oduaran, O.; Hazelhurst, S.; Bhatt, A.S. Surveying Gut Microbiome Research in Africans: Toward Improved Diversity and Representation. Trends Microbiol. 2019, 27, 824–835. [Google Scholar] [CrossRef]
- Del Chierico, F.; Abbatini, F.; Russo, A.; Quagliariello, A.; Reddel, S.; Capoccia, D.; Caccamo, R.; Corradini, S.G.; Nobili, V.; De Peppo, F.; et al. Gut Microbiota Markers in Obese Adolescent and Adult Patients: Age-Dependent Differential Patterns. Front. Microbiol. 2018, 9, 1210. [Google Scholar] [CrossRef] [PubMed]
- Yun, Y.; Kim, H.-N.; Kim, S.E.; Heo, S.G.; Chang, Y.; Ryu, S.; Shin, H.; Kim, H.-L. Comparative analysis of gut microbiota associated with body mass index in a large Korean cohort. BMC Microbiol. 2017, 17, 151. [Google Scholar] [CrossRef] [Green Version]
- Oh, T.J.; Sul, W.J.; Na Oh, H.; Lee, Y.-K.; Lim, H.L.; Choi, S.H.; Park, K.S.; Jang, H.C. Butyrate attenuated fat gain through gut microbiota modulation in db/db mice following dapagliflozin treatment. Sci. Rep. 2019, 9, 20300. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Q.; Chang, Y.; Zhang, K.; Chen, H.; Tao, S.; Zhang, Z. Implication of the gut microbiome composition of type 2 diabetic patients from northern China. Sci. Rep. 2020, 10, 5450. [Google Scholar] [CrossRef] [PubMed]
- Shah, V.O.; Lambeth, S.M.; Carson, T.; Lowe, J.; Ramaraj, T.; Leff, J.W.; Luo, L.; Bell, C.J. Composition Diversity and Abundance of Gut Microbiome in Prediabetes and Type 2 Diabetes. J. Diabetes Obes. 2015, 2, 108–114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, J.; Wright, K.; Davis, J.M.; Jeraldo, P.; Marietta, E.V.; Murray, J.; Nelson, H.; Matteson, E.L.; Taneja, V. An expansion of rare lineage intestinal microbes characterizes rheumatoid arthritis. Genome Med. 2016, 8, 43. [Google Scholar] [CrossRef] [Green Version]
- Ridlon, J.M.; Kang, D.-J.; Hylemon, P.B. Bile salt biotransformations by human intestinal bacteria. J. Lipid Res. 2006, 47, 241–259. [Google Scholar] [CrossRef] [Green Version]
- Cani, P.D.; Possemiers, S.; Van De Wiele, T.; Guiot, Y.; Everard, A.; Rottier, O.; Geurts, L.; Naslain, D.; Neyrinck, A.; Lambert, D.M.; et al. Changes in gut microbiota control inflammation in obese mice through a mechanism involving GLP-2-driven improvement of gut permeability. Gut 2009, 58, 1091–1103. [Google Scholar] [CrossRef] [Green Version]
- Gomez-Arango, L.F.; Barrett, H.L.; Wilkinson, S.; Callaway, L.K.; McIntyre, H.D.; Morrison, M.; Nitert, M.D. Low dietary fiber intake increases Collinsella abundance in the gut microbiota of overweight and obese pregnant women. Gut Microbes 2018, 9, 189–201. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scudiero, O.; Pero, R.; Ranieri, A.; Terracciano, D.; Fimiani, F.; Cesaro, A.; Gentile, L.; Leggiero, E.; Laneri, S.; Moscarella, E.; et al. Childhood obesity: An overview of laboratory medicine, exercise and microbiome. Clin. Chem. Lab. Med. 2020, 58, 1385–1406. [Google Scholar] [CrossRef]
- Ley, R.; Turnbaugh, P.J.; Klein, S.; Gordon, J.I. Microbial ecology: Human gut microbes associated with obesity. Nature 2006, 444, 1022–1023. [Google Scholar] [CrossRef]
- Dalby, M.J.; Ross, A.; Walker, A.; Morgan, P.J. Dietary Uncoupling of Gut Microbiota and Energy Harvesting from Obesity and Glucose Tolerance in Mice. Cell Rep. 2017, 21, 1521–1533. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ahrens, W.; Iacoviello, L.; Lissner, L.; Veidebaum, T.; Pohlabeln, H.; Pigeot, I.; Moreno, L.A.; Marild, S.; Molnar, D.; Siani, A.; et al. Metabolic syndrome in young children: Definitions and results of the IDEFICS study. Int. J. Obes. 2014, 38, S4–S14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miyamoto, J.; Igarashi, M.; Watanabe, K.; Karaki, S.-I.; Mukouyama, H.; Kishino, S.; Li, X.; Ichimura, A.; Irie, J.; Sugimoto, Y.; et al. Gut microbiota confers host resistance to obesity by metabolizing dietary polyunsaturated fatty acids. Nat. Commun. 2019, 10, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Muhsen, K.; Green, M.S.; Soskolne, V.; Neumark, Y. Inequalities in non-communicable diseases between the major population groups in Israel: Achievements and challenges. Lancet 2017, 389, 2531–2541. [Google Scholar] [CrossRef]
- Kishino, S.; Ogawa, J.; Yokozeki, K.; Shimizu, S. Metabolic diversity in biohydrogenation of polyunsaturated fatty acids by lactic acid bacteria involving conjugated fatty acid production. Appl. Microbiol. Biotechnol. 2009, 84, 87–97. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kishino, S.; Takeuchi, M.; Park, S.-B.; Hirata, A.; Kitamura, N.; Kunisawa, J.; Kiyono, H.; Iwamoto, R.; Isobe, Y.; Arita, M.; et al. Polyunsaturated fatty acid saturation by gut lactic acid bacteria affecting host lipid composition. Proc. Natl. Acad. Sci. USA 2013, 110, 17808–17813. [Google Scholar] [CrossRef] [Green Version]
- Alderman, H.; Behrman, J.R.; Glewwe, P.; Fernald, L.; Walker, S. Evidence of Impact of Interventions on Growth and De-velopment during Early and Middle Childhood. In Disease Control Priorities, Third Edition (Volume 8): Child and Adolescent Health and Development, 3rd ed.; Bundy, D.A.P., Silva, N.D., Horton, S., Jamison, D.T., Patton, G.C., Eds.; The International Bank for Reconstruction and Development/The World Bank: Washington, DC, USA, 2017; Chapter 7. [Google Scholar]

| Socioeconomic Status | ||||
|---|---|---|---|---|
| Characteristic | Overall, N = 139 a | High SES, N = 70 a | Low SES, N = 69 a | p-Value b,c |
| Age | 11.42 (10.83, 11.67) | 11.04 (10.67, 11.50) | 11.58 (11.42, 11.75) | <0.001 |
| Sex | 0.9 | |||
| Female | 57 (41%) | 29 (41%) | 28 (41%) | |
| Male | 82 (59%) | 41 (59%) | 41 (59%) | |
| Socioeconomic score | 6.45 (4.69, 7.53) | 7.53 (6.89, 8.13) | 4.66 (3.83, 5.49) | <0.001 |
| Village | <0.001 | |||
| Village A | 46 (33%) | 37 (53%) | 9 (13%) | |
| Village B | 93 (67%) | 33 (47%) | 60 (87%) | |
| Household crowding | 1.50 (1.15, 2.00) | 1.20 (1.00, 1.50) | 2.00 (1.50, 2.33) | <0.001 |
| Body Mass Index Z (BMIZ) score | 1.61 (0.22, 2.59) | 0.92 (−0.13, 2.27) | 1.98 (0.68, 2.59) | 0.024 |
| BMIZ score classification | 0.14 | |||
| Normal weight | 60 (43%) | 36 (51%) | 24 (35%) | |
| Overweight | 19 (14%) | 8 (11%) | 11 (16%) | |
| Obese | 60 (43%) | 26 (37%) | 34 (49%) | |
| Carbohydrates intake (%Kcal) | 53 (48, 57) | 53 (49, 57) | 51 (47, 57) | 0.3 |
| Protein intake (%Kcal) | 15.0 (12.2, 18.6) | 14.4 (11.8, 18.6) | 16.0 (13.0, 18.5) | 0.14 |
| Fat intake (%Kcal) | 31 (27, 35) | 30 (26, 34) | 31 (27, 35) | 0.4 |
| Saturated fat intake (%Kcal) | 9.5 (7.5, 12.0) | 9.5 (7.4, 12.3) | 9.8 (7.7, 11.7) | 0.8 |
| Mono-unsaturated fat intake (%Kcal) | 11.4 (9.2, 13.3) | 11.0 (9.1, 13.2) | 11.7 (9.4, 13.3) | 0.5 |
| Poly-unsaturated fat intake (%Kcal) | 6.3 (3.2, 8.6) | 4.1 (1.6, 7.9) | 7.1 (5.5, 9.0) | 0.001 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lapidot, Y.; Reshef, L.; Goldsmith, R.; Na’amnih, W.; Kassem, E.; Ornoy, A.; Gophna, U.; Muhsen, K. The Associations between Diet and Socioeconomic Disparities and the Intestinal Microbiome in Preadolescence. Nutrients 2021, 13, 2645. https://doi.org/10.3390/nu13082645
Lapidot Y, Reshef L, Goldsmith R, Na’amnih W, Kassem E, Ornoy A, Gophna U, Muhsen K. The Associations between Diet and Socioeconomic Disparities and the Intestinal Microbiome in Preadolescence. Nutrients. 2021; 13(8):2645. https://doi.org/10.3390/nu13082645
Chicago/Turabian StyleLapidot, Yelena, Leah Reshef, Rebecca Goldsmith, Wasef Na’amnih, Eias Kassem, Asher Ornoy, Uri Gophna, and Khitam Muhsen. 2021. "The Associations between Diet and Socioeconomic Disparities and the Intestinal Microbiome in Preadolescence" Nutrients 13, no. 8: 2645. https://doi.org/10.3390/nu13082645
APA StyleLapidot, Y., Reshef, L., Goldsmith, R., Na’amnih, W., Kassem, E., Ornoy, A., Gophna, U., & Muhsen, K. (2021). The Associations between Diet and Socioeconomic Disparities and the Intestinal Microbiome in Preadolescence. Nutrients, 13(8), 2645. https://doi.org/10.3390/nu13082645







