Probiotic and Functional Properties of Limosilactobacillus reuteri INIA P572
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Growth Conditions
2.2. Resistance to Stimulated Gastrointestinal Conditions In Vitro
2.3. Viability of L. reuteri INIA P572 and Reuterin Production in an In Vitro Colonic Model. Modulation of Fecal Bacterial Population
2.4. Immunomodulatory Activity In Vitro
2.4.1. Nitric Oxide (NO) Production
2.4.2. Cytokine Production
2.5. Animal Studies
2.6. Dextran Sodium Sulfate (DSS) Model of Mouse Colitis
2.6.1. Intestinal Inflammatory Process Evaluation
2.6.2. Histological Analysis
2.6.3. Analysis of Gene Expression in Colonic Samples by RT-qPCR
2.6.4. Flow Cytometry
2.7. Statistical Analysis
3. Results
3.1. L. reuteri INIA P572 Resistance to Gastrointestinal Conditions In Vitro
3.2. Viability of L. reuteri INIA P572 and Faecal Microbiota in an In Vitro Colonic Model. Reuterin Production
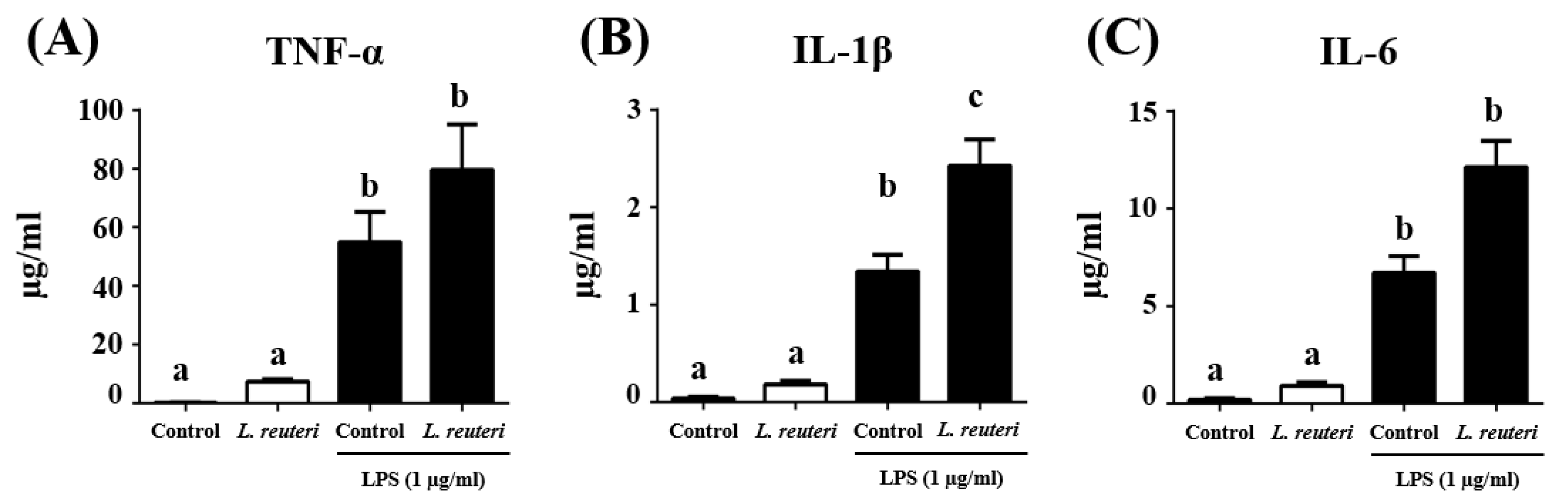
3.3. Immunomodulatory Activity of L. reuteri INIA P572 Using In Vitro Models
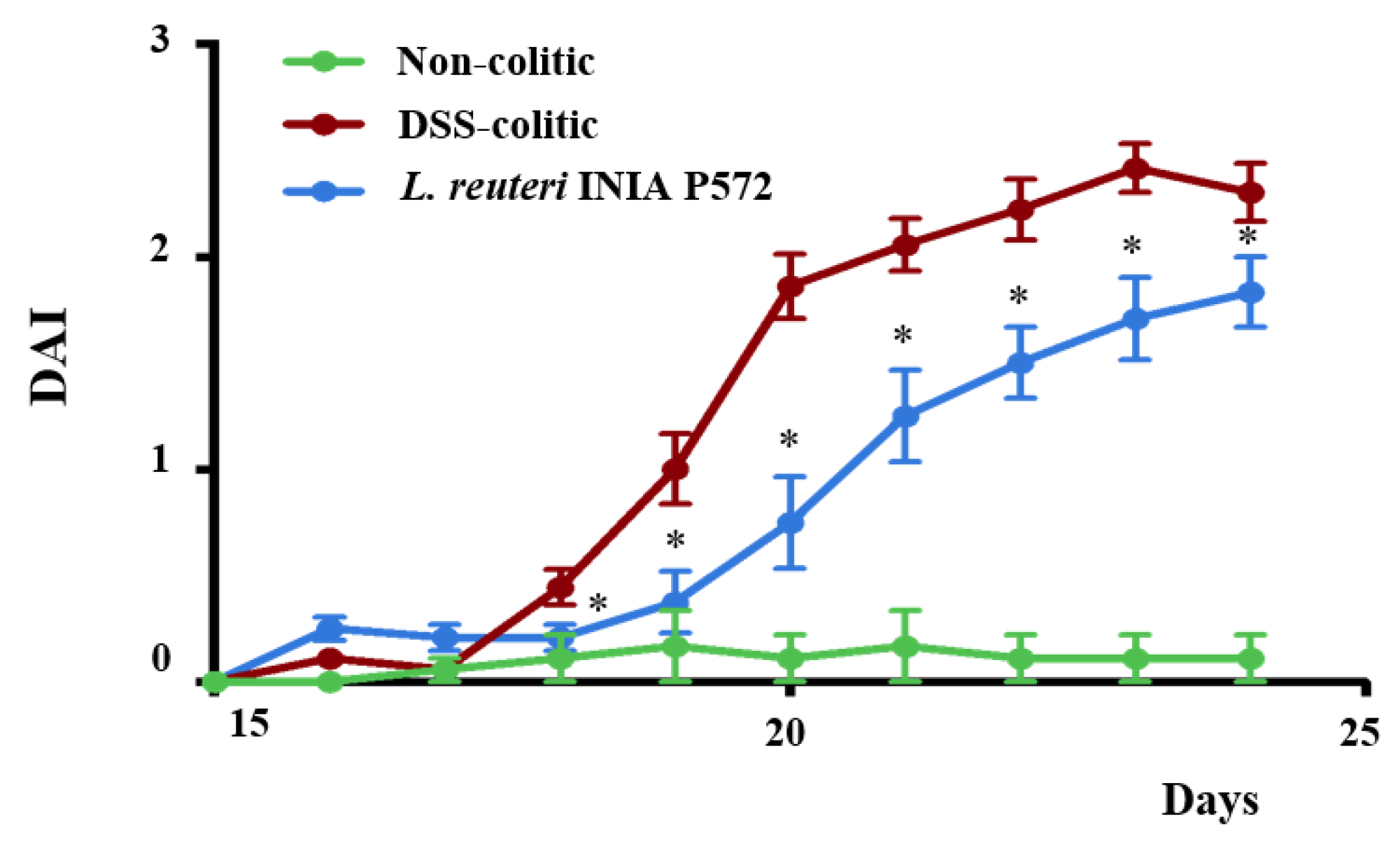
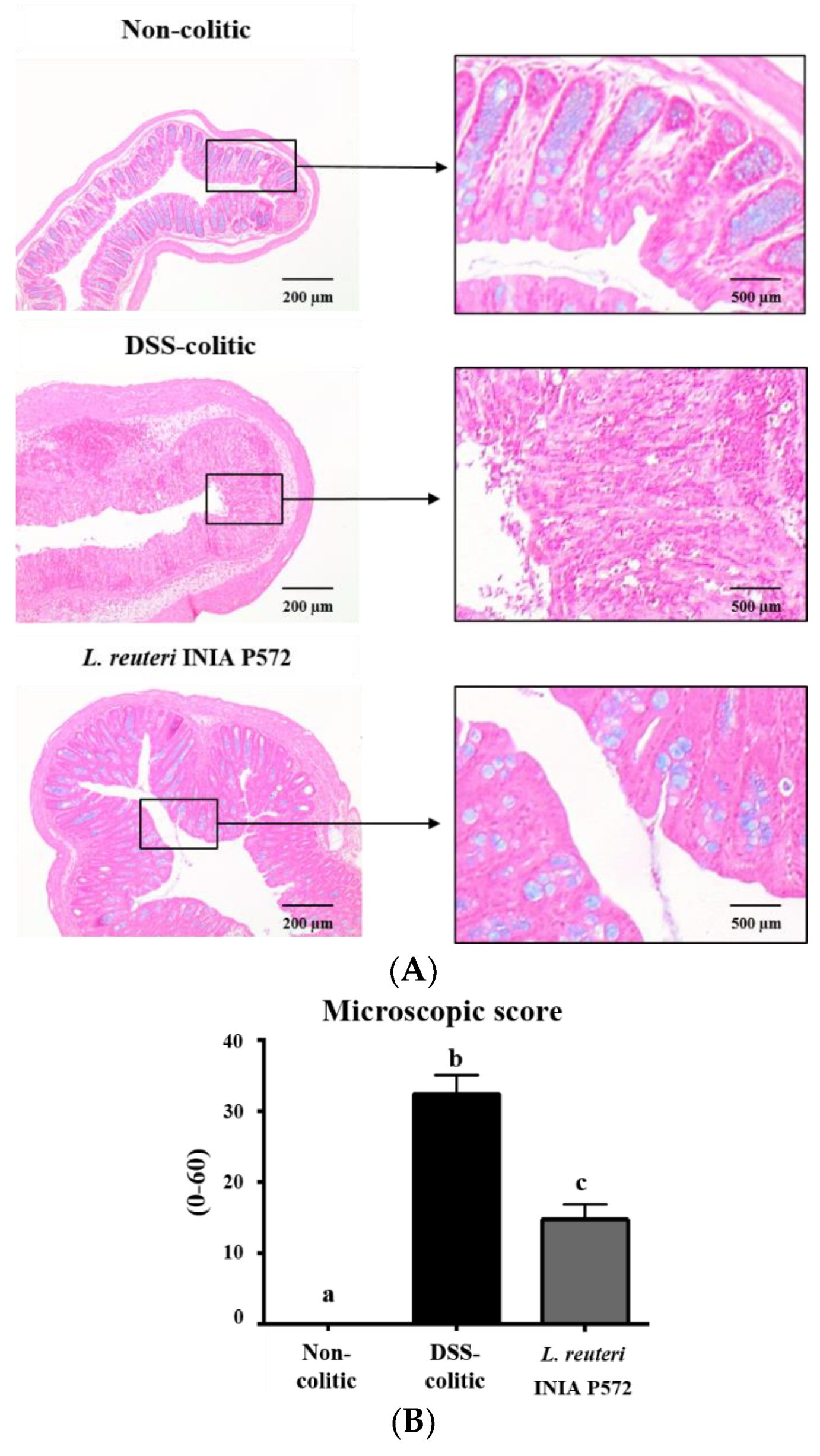
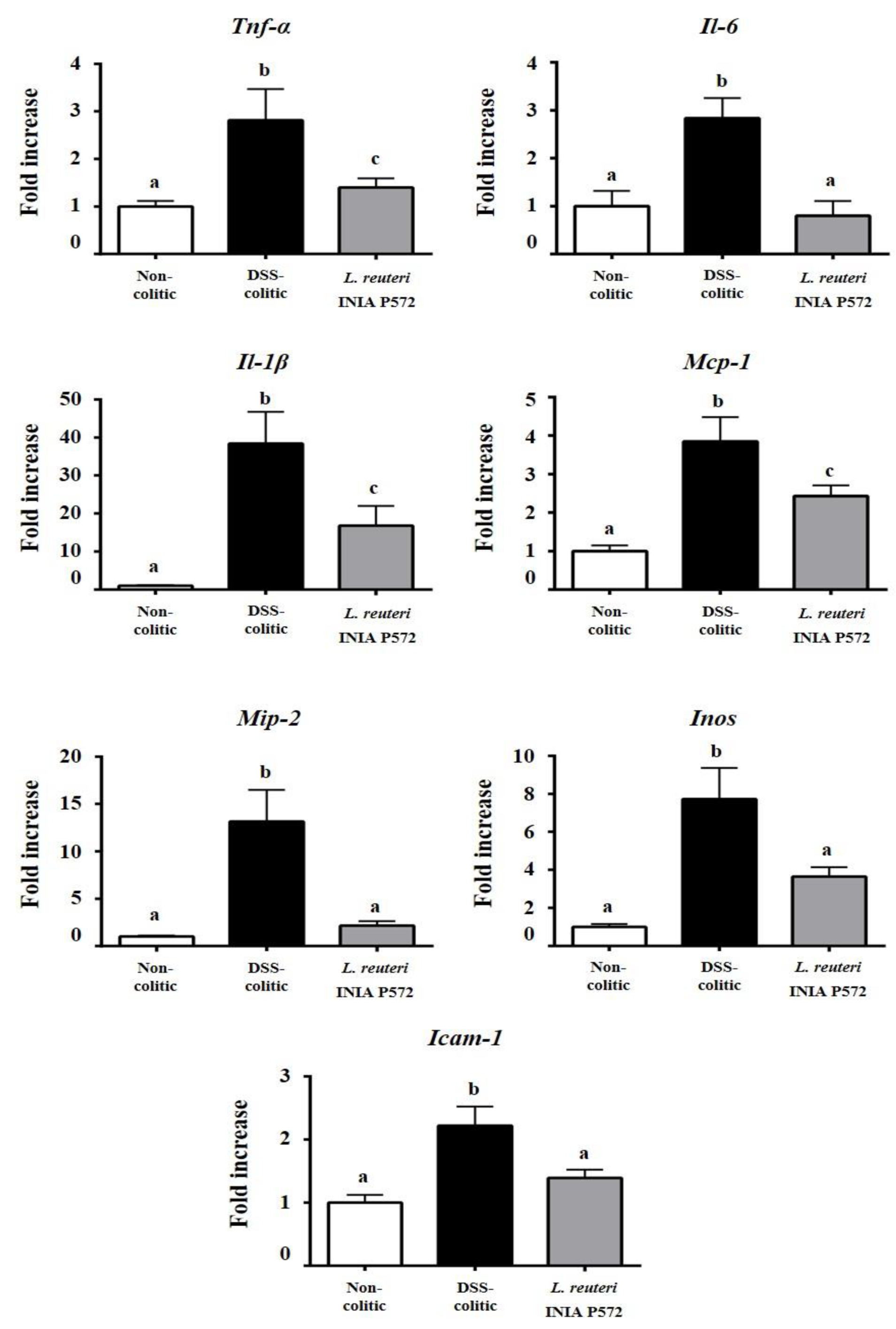
3.4. Intestinal Anti-Inflammatory Effect of L. reuteri INIA P572 in Experimental Colitis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Reuter, G. Probiotics—possibilities and limitations of their application in food, animal feed, and in pharmaceutical preparations for men and animals. Berliner Munchener Tierarztliche Wochenschrift 2001, 114, 410–419. [Google Scholar]
- Walter, J.; Schwab, C.; Loach, D.M.; Gänzle, M.G.; Tannock, G.W. Glucosyltransferase A (GtfA) and inulosucrase (Inu) of Lactobacillus reuteri TMW1.106 contribute to cell aggregation, in vitro biofilm formation, and colonization of the mouse gastrointestinal tract. Microbiology 2008, 154, 72–80. [Google Scholar] [CrossRef] [PubMed]
- Oh, P.L.; Benson, A.K.; Peterson, D.A.; Patil, P.B.; Moriyama, E.N.; Roos, S.; Walter, J. Diversification of the gut symbiont Lactobacillus reuteri as a result of host-driven evolution. ISME J. 2010, 4, 377–387. [Google Scholar] [CrossRef]
- Mishra, S.K.; Malik, R.K.; Manju, G.; Pandey, N.; Singroha, G.; Behare, P.; Kaushik, J.K. Characterization of a Reuterin-Producing Lactobacillus reuteri BPL-36 Strain Isolated from Human Infant Fecal Sample. Probiotics Antimicrob. Proteins 2012, 4, 154–161. [Google Scholar] [CrossRef]
- Stiles, M.E. Biopreservation by lactic acid bacteria. Antonie Leeuwenhoek 1996, 70, 331–345. [Google Scholar] [CrossRef]
- Casas, I.A.; Dobrogosz, W.J. Validation of the probiotic concept: Lactobacillus reuteri confers broad-spectrum protection against disease in humans and animals. Microb. Ecol. Health Dis. 2000, 12, 247–285. [Google Scholar]
- Vollenweider, S.; Lacroix, C. 3-hydroxypropionaldehyde: Applications and perspectives of biotechnological production. Appl. Microbiol. Biotechnol. 2004, 64, 16–27. [Google Scholar] [CrossRef]
- Francavilla, R.; Lionetti, E.; Castellaneta, S.; Ciruzzi, F.; Indrio, F.; Masciale, A.; Fontana, C.; La Rosa, M.M.; Cavallo, L.; Francavilla, A. Randomised clinical trial: Lactobacillus reuteri DSM 17938 vs. placebo in children with acute diarrhoea—A double-blind study. Aliment. Pharmacol. Ther. 2012, 36, 363–369. [Google Scholar] [CrossRef]
- Urbańska, M.; Gieruszczak-Białek, D.; Szajewska, H. Systematic review with meta-analysis: Lactobacillus reuteri DSM 17938 for diarrhoeal diseases in children. Aliment. Pharmacol. Ther. 2016, 43, 1025–1034. [Google Scholar] [CrossRef]
- Singh, T.P.; Malik, R.K.; Katkamwar, S.G.; Kaur, G. Hypocholesterolemic effects of Lactobacillus reuteri LR6 in rats fed on high-cholesterol diet. Int. J. Food Sci. Nutr. 2015, 66, 71–75. [Google Scholar] [CrossRef]
- Francavilla, R.; Lionetti, E.; Castellaneta, S.P.; Magistà, A.M.; Maurogiovanni, G.; Bucci, N.; De Canio, A.; Indrio, F.; Cavallo, L.; Ierardi, E.; et al. Inhibition of Helicobacter pylori infection in humans by Lactobacillus reuteri ATCC 55730 and effect on eradication therapy: A pilot study. Helicobacter 2008, 13, 127–134. [Google Scholar] [CrossRef]
- Madsen, K.L.; Doyle, J.S.; Jewell, L.D.; Tavernini, M.M.; Fedorak, R.N. Lactobacillus species prevents colitis in interleukin 10 gene-deficient mice. Gastroenterology 1999, 116, 1107–1114. [Google Scholar] [CrossRef]
- Holma, R.; Salmenperä, P.; Lohi, J.; Vapaatalo, H.; Korpela, R. Effects of Lactobacillus rhamnosus GG and Lactobacillus reuteri R2LC on acetic acid-induced colitis in rats. Scand. J. Gastroenterol. 2001, 36, 630–635. [Google Scholar] [CrossRef] [PubMed]
- Møller, P.L.; Paerregaard, A.; Gad, M.; Kristensen, N.N.; Claesson, M.H. Colitic scid mice fed Lactobacillus spp. show an ameliorated gut histopathology and an altered cytokine profile by local T cells. Inflamm. Bowel Dis. 2005, 11, 814–819. [Google Scholar] [CrossRef] [PubMed]
- Peran, L.; Sierra, S.; Comalada, M.; Lara-Villoslada, F.; Bailón, E.; Nieto, A.; Concha, A.; Olivares, M.; Zarzuelo, A.; Xaus, J.; et al. A comparative study of the preventative effects exerted by two probiotics, Lactobacillus reuteri and Lactobacillus fermentum, in the trinitrobenzenesulfonic acid model of rat colitis. Br. J. Nutr. 2007, 97, 96–103. [Google Scholar] [CrossRef]
- Schreiber, O.; Petersson, J.; Phillipson, M.; Perry, M.; Roos, S.; Holm, L. Lactobacillus reuteri prevents colitis by reducing P-selectin-associated leukocyte- and platelet-endothelial cell interactions. Am. J. Physiol. Gastrointest. Liver Physiol. 2009, 296, G534–G542. [Google Scholar] [CrossRef] [PubMed]
- Vulevic, J.; Tzortzis, G.; Juric, A.; Gibson, G.R. Effect of a prebiotic galactooligosaccharide mixture (B-GOS®) on gastrointestinal symptoms in adults selected from a general population who suffer with bloating, abdominal pain, or flatulence. Neurogastroenterol. Motil. 2018, 30, e13440. [Google Scholar] [CrossRef] [PubMed]
- Talarico, T.L.; Dobrogosz, W.J. Chemical characterization of an antimicrobial substance produced by Lactobacillus reuteri. Antimicrob. Agents Chemother. 1989, 33, 674–679. [Google Scholar] [CrossRef] [PubMed]
- Vollenweider, S.; Evers, S.; Zurbriggen, K.; Lacroix, C. Unraveling the hydroxypropionaldehyde (HPA) system: An active antimicrobial agent against human pathogens. J. Agric. Food Chem. 2010, 58, 10315–10322. [Google Scholar] [CrossRef]
- Jones, S.E.; Versalovic, J. Probiotic Lactobacillus reuteri biofilms produce antimicrobial and anti-inflammatory factors. BMC Microbiol. 2009, 9, 35. [Google Scholar] [CrossRef]
- Griet, M.; de Valdez, G.F.; Gerez, C.L.; Rodríguez, A.V. Two-step production of anti-inflammatory soluble factor by Lactobacillus reuteri CRL 1098. PLoS ONE 2018, 13, e0200426. [Google Scholar] [CrossRef] [PubMed]
- Das, N.K.; Schwartz, A.J.; Barthel, G.; Inohara, N.; Liu, Q.; Sankar, A.; Hill, D.R.; Ma, X.; Lamberg, O.; Schnizlein, M.K.; et al. Microbial Metabolite Signaling Is Required for Systemic Iron Homeostasis. Cell Metab. 2020, 31, 115–130. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority (EFSA). The maintenance of the list of QPS microorganisms intentionally added to food or feed-Scientific Opinion of the Panel on Biological Hazards. EFSA J. 2008, 6, 923. [Google Scholar] [CrossRef]
- Wolf, B.W.; Wheeler, K.B.; Ataya, D.G.; Garleb, K.A. Safety and tolerance of Lactobacillus reuteri supplementation to a population infected with the human immunodeficiency virus. Food Chem. Toxicol. Int. J. Publ. Br. Ind. Biol. Res. Assoc. 1998, 36, 1085–1094. [Google Scholar] [CrossRef]
- Jones, M.L.; Martoni, C.J.; Di Pietro, E.; Simon, R.R.; Prakash, S. Evaluation of clinical safety and tolerance of a Lactobacillus reuteri NCIMB 30242 supplement capsule: A randomized control trial. Regul. Toxicol. Pharmacol. RTP 2012, 63, 313–320. [Google Scholar] [CrossRef]
- Rodríguez, E.; Arqués, J.L.; Rodríguez, R.; Nuñez, M.; Medina, M. Reuterin production by lactobacilli isolated from pig faeces and evaluation of probiotic traits. Lett. Appl. Microbiol. 2003, 37, 259–263. [Google Scholar] [CrossRef]
- Langa, S.; Landete, J.M.; Martín-Cabrejas, I.; Rodríguez, E.; Arqués, J.L.; Medina, M. In situ reuterin production by Lactobacillus reuteri in dairy products. Food Control 2013, 33, 200–206. [Google Scholar] [CrossRef]
- Langa, S.; Arqués, J.L.; Gaya, P.; Medina, M.; Landete, J.M. Glycerol and cobalamin metabolism in lactobacilli: Relevance of the propanediol dehydrogenase pdh 30. Eur. Food Res. Technol. 2015, 241, 173–184. [Google Scholar] [CrossRef]
- Martín-Cabrejas, I.; Langa, S.; Gaya, P.; Rodríguez, E.; Landete, J.M.; Medina, M.; Arqués, J.L. Optimization of reuterin production in cheese by Lactobacillus reuteri. J. Food Sci. Technol. 2017, 54, 1346–1349. [Google Scholar] [CrossRef]
- Langa, S.; Martín-Cabrejas, I.; Montiel, R.; Peirotén, Á.; Arqués, J.L.; Medina, M. Protective effect of reuterin-producing Lactobacillus reuteri against Listeria monocytogenes and Escherichia coli O157: H7 in semi-hard cheese. Food Control 2018, 84, 284–289. [Google Scholar] [CrossRef]
- Gómez-Torres, N.; Ávila, M.; Gaya, P.; Garde, S. Prevention of late blowing defect by reuterin produced in cheese by a Lactobacillus reuteri adjunct. Food Microbiol. 2014, 42, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.P.; Thibodeaux, C.H.; Peña, J.A.; Ferry, G.D.; Versalovic, J. Probiotic Lactobacillus reuteri suppress proinflammatory cytokines via c-Jun. Inflamm. Bowel Dis. 2008, 14, 1068–1083. [Google Scholar] [CrossRef]
- Brisbin, J.T.; Davidge, L.; Roshdieh, A.; Sharif, S. Characterization of the effects of three Lactobacillus species on the function of chicken macrophages. Res. Vet. Sci. 2015, 100, 39–44. [Google Scholar] [CrossRef]
- Borruel, N.; Carol, M.; Casellas, F.; Antolín, M.; de Lara, F.; Espín, E.; Naval, J.; Guarner, F.; Malagelada, J.R. Increased mucosal tumour necrosis factor alpha production in Crohn’s disease can be downregulated ex vivo by probiotic bacteria. Gut 2002, 51, 659–664. [Google Scholar] [CrossRef]
- Schultz, M.; Schölmerich, J.; Rath, H.C. Rationale for probiotic and antibiotic treatment strategies in inflammatory bowel diseases. Dig. Dis. 2003, 21, 105–128. [Google Scholar] [CrossRef] [PubMed]
- Pathmakanthan, S.; Li, C.K.; Cowie, J.; Hawkey, C.J. Lactobacillus plantarum 299: Beneficial in vitro immunomodulation in cells extracted from inflamed human colon. J. Gastroenterol. Hepatol. 2004, 19, 166–173. [Google Scholar] [CrossRef] [PubMed]
- Madsen, K.; Cornish, A.; Soper, P.; McKaigney, C.; Jijon, H.; Yachimec, C.; Doyle, J.; Jewell, L.; De Simone, C. Probiotic bacteria enhance murine and human intestinal epithelial barrier function. Gastroenterology 2001, 121, 580–591. [Google Scholar] [CrossRef] [PubMed]
- Landete, J.M.; Langa, S.; Revilla, C.; Margolles, A.; Medina, M.; Arqués, J.L. Use of anaerobic green fluorescent protein versus green fluorescent protein as reporter in lactic acid bacteria. Appl. Microbiol. Biotechnol. 2015, 99, 6865–6877. [Google Scholar] [CrossRef] [PubMed]
- Haller, D.; Colbus, H.; Gänzle, M.G.; Scherenbacher, P.; Bode, C.; Hammes, W.P. Metabolic and functional properties of lactic acid bacteria in the gastro-intestinal ecosystem: A comparative in vitro study between bacteria of intestinal and fermented food origin. Syst. Appl. Microbiol. 2001, 24, 218–226. [Google Scholar] [CrossRef]
- Vulevic, J.; Rastall, R.A.; Gibson, G.R. Developing a quantitative approach for determining the in vitro prebiotic potential of dietary oligosaccharides. FEMS Microbiol. Lett. 2004, 236, 153–159. [Google Scholar] [CrossRef]
- Circle, S.J.; Stone, L.; Boruff, C.S. Acrolein determination by means of tryptophane. A colorimetric micromethod. Ind. Eng. Chem. Anal. Ed. 1945, 17, 259–262. [Google Scholar] [CrossRef]
- Ferrari, M.; Fornasiero, M.C.; Isetta, A.M. MTT colorimetric assay for testing macrophage cytotoxic activity in vitro. J. Immunol. Methods 1990, 131, 165–172. [Google Scholar] [CrossRef]
- Green, L.C.; Wagner, D.A.; Glogowski, J.; Skipper, P.L.; Wishnok, J.S.; Tannenbaum, S.R. Analysis of nitrate, nitrite, and [15N]nitrate in biological fluids. Anal. Biochem. 1982, 126, 131–138. [Google Scholar] [CrossRef]
- Kim, H.J.; Lee, S.S.; Choi, B.Y.; Kim, M.K. Nitrate intake relative to antioxidant vitamin intake affects gastric cancer risk: A case-control study in Korea. Nutr. Cancer 2007, 59, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Kim, H.J.; Kim, J.S.; Yoo, K.; Lee, J.C.; Anderson, W.A.; Lee, J.H. Photocatalytic reduction of nitrate in wastewater using ZnO nanopowder synthesized by solution combustion method. J. Nanosci. Nanotechnol. 2007, 7, 4069–4072. [Google Scholar] [CrossRef] [PubMed]
- Cooper, H.S.; Murthy, S.N.; Shah, R.S.; Sedergran, D.J. Clinicopathologic study of dextran sulfate sodium experimental murine colitis. Lab. Investig. J. Tech. Methods Pathol. 1993, 69, 238–249. [Google Scholar]
- Garrido-Mesa, J.; Rodríguez-Nogales, A.; Algieri, F.; Vezza, T.; Hidalgo-Garcia, L.; Garrido-Barros, M.; Utrilla, M.P.; Garcia, F.; Chueca, N.; Rodriguez-Cabezas, M.E.; et al. Immunomodulatory tetracyclines shape the intestinal inflammatory response inducing mucosal healing and resolution. Br. J. Pharmacol. 2018, 175, 4353–4370. [Google Scholar] [CrossRef]
- Garrido-Mesa, J.; Algieri, F.; Rodriguez-Nogales, A.; Utrilla, M.P.; Rodriguez-Cabezas, M.E.; Zarzuelo, A.; Ocete, M.A.; Garrido-Mesa, N.; Galvez, J. A new therapeutic association to manage relapsing experimental colitis: Doxycycline plus Saccharomyces boulardii. Pharmacol. Res. 2015, 97, 48–63. [Google Scholar] [CrossRef]
- Scott, C.L.; Bain, C.C.; Mowat, A.M. Isolation and Identification of Intestinal Myeloid Cells. Methods Mol. Biol. 2017, 1559, 223–239. [Google Scholar] [CrossRef]
- Arribas, B.; Suárez-Pereira, E.; Ortiz Mellet, C.; García Fernández, J.M.; Buttersack, C.; Rodríguez-Cabezas, M.E.; Garrido-Mesa, N.; Bailon, E.; Guerra-Hernández, E.; Zarzuelo, A.; et al. Di-D-fructose dianhydride-enriched caramels: Effect on colon microbiota, inflammation, and tissue damage in trinitrobenzenesulfonic acid-induced colitic rats. J. Agric. Food Chem. 2010, 58, 6476–6484. [Google Scholar] [CrossRef]
- Nomoto, K. Prevention of infections by probiotics. J. Biosci. Bioeng. 2005, 100, 583–592. [Google Scholar] [CrossRef] [PubMed]
- Galdeano, C.M.; Perdigón, G. Role of viability of probiotic strains in their persistence in the gut and in mucosal immune stimulation. J. Appl. Microbiol. 2004, 97, 673–681. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, E.; Arqués, J.L.; Rodríguez, R.; Peirotén, Á.; Landete, J.M.; Medina, M. Antimicrobial properties of probiotic strains isolated from breast-fed infants. J. Funct. Foods 2012, 4, 542–551. [Google Scholar] [CrossRef]
- Fernandez, B.; Hammami, R.; Savard, P.; Jean, J.; Fliss, I. Pediococcus acidilactici UL5 and Lactococcus lactis ATCC 11454 are able to survive and express their bacteriocin genes under simulated gastrointestinal conditions. J. Appl. Microbiol. 2014, 116, 677–688. [Google Scholar] [CrossRef]
- Gasser, F. Safety of lactic acid bacteria and their occurrence in human clinical infections. Bulletin l’Institut Pasteur 1994, 92, 45–67. [Google Scholar]
- Donohue, D.; Salminen, S. Safety of probiotic bacteria. Asia Pacific J. Clin. Nutr. 1996, 5, 25–28. [Google Scholar]
- Cleusix, V.; Lacroix, C.; Vollenweider, S.; Duboux, M.; Le Blay, G. Inhibitory activity spectrum of reuterin produced by Lactobacillus reuteri against intestinal bacteria. BMC Microbiol. 2007, 7, 101. [Google Scholar] [CrossRef]
- De Weirdt, R.; Possemiers, S.; Vermeulen, G.; Moerdijk-Poortvliet, T.C.; Boschker, H.T.; Verstraete, W.; Van de Wiele, T. Human faecal microbiota display variable patterns of glycerol metabolism. FEMS Microbiol. Ecol. 2010, 74, 601–611. [Google Scholar] [CrossRef]
- Biebl, H.; Menzel, K.; Zeng, A.P.; Deckwer, W.D. Microbial production of 1,3-propanediol. Appl. Microbiol. Biotechnol. 1999, 52, 289–297. [Google Scholar] [CrossRef] [PubMed]
- Sauvageot, N.; Gouffi, K.; Laplace, J.M.; Auffray, Y. Glycerol metabolism in Lactobacillus collinoides: Production of 3-hydroxypropionaldehyde, a precursor of acrolein. Int. J. Food Microbiol. 2000, 55, 167–170. [Google Scholar] [CrossRef]
- Martín, R.; Olivares, M.; Marín, M.L.; Fernández, L.; Xaus, J.; Rodríguez, J.M. Probiotic potential of 3 Lactobacilli strains isolated from breast milk. J. Hum. Lact. Off. J. Int. Lact. Consult. Assoc. 2005, 21, 8–17. [Google Scholar] [CrossRef]
- Garai-Ibabe, G.; Ibarburu, I.; Berregi, I.; Claisse, O.; Lonvaud-Funel, A.; Irastorza, A.; Dueñas, M.T. Glycerol metabolism and bitterness producing lactic acid bacteria in cidermaking. Int. J. Food Microbiol. 2008, 121, 253–261. [Google Scholar] [CrossRef] [PubMed]
- Bauer, R.; du Toit, M.; Kossmann, J. Influence of environmental parameters on production of the acrolein precursor 3-hydroxypropionaldehyde by Lactobacillus reuteri DSMZ 20016 and its accumulation by wine lactobacilli. Int. J. Food Microbiol. 2010, 137, 28–31. [Google Scholar] [CrossRef]
- Morita, H.; Toh, H.; Fukuda, S.; Horikawa, H.; Oshima, K.; Suzuki, T.; Murakami, M.; Hisamatsu, S.; Kato, Y.; Takizawa, T.; et al. Comparative genome analysis of Lactobacillus reuteri and Lactobacillus fermentum reveal a genomic island for reuterin and cobalamin production. DNA Res. Int. J. Rapid Publ. Rep. Genes Genomes 2008, 15, 151–161. [Google Scholar] [CrossRef] [PubMed]
- De Weirdt, R.; Crabbé, A.; Roos, S.; Vollenweider, S.; Lacroix, C.; van Pijkeren, J.P.; Britton, R.A.; Sarker, S.; Van de Wiele, T.; Nickerson, C.A. Glycerol supplementation enhances L. reuteri’s protective effect against S. Typhimurium colonization in a 3-D model of colonic epithelium. PLoS ONE 2012, 7, e37116. [Google Scholar] [CrossRef] [PubMed]
- De Boever, P.; Deplancke, B.; Verstraete, W. Fermentation by gut microbiota cultured in a simulator of the human intestinal microbial ecosystem is improved by supplementing a soygerm powder. J. Nutr. 2000, 130, 2599–2606. [Google Scholar] [CrossRef]
- Vardakou, M.; Palop, C.N.; Christakopoulos, P.; Faulds, C.B.; Gasson, M.A.; Narbad, A. Evaluation of the prebiotic properties of wheat arabinoxylan fractions and induction of hydrolase activity in gut microflora. Int. J. Food Microbiol. 2008, 123, 166–170. [Google Scholar] [CrossRef]
- Filocamo, A.; Nueno-Palop, C.; Bisignano, C.; Mandalari, G.; Narbad, A. Effect of garlic powder on the growth of commensal bacteria from the gastrointestinal tract. Phytomed. Int. J. Phytother. Phytopharm. 2012, 19, 707–711. [Google Scholar] [CrossRef]
- Tejada-Simon, M.V.; Pestka, J.J. Proinflammatory cytokine and nitric oxide induction in murine macrophages by cell wall and cytoplasmic extracts of lactic acid bacteria. J. Food Prot. 1999, 62, 1435–1444. [Google Scholar] [CrossRef]
- Chon, H.; Choi, B.; Lee, E.; Lee, S.; Jeong, G. Immunomodulatory effects of specific bacterial components of Lactobacillus plantarum KFCC11389P on the murine macrophage cell line RAW 264.7. J. Appl. Microbiol. 2009, 107, 1588–1597. [Google Scholar] [CrossRef]
- Rodes, L.; Khan, A.; Paul, A.; Coussa-Charley, M.; Marinescu, D.; Tomaro-Duchesneau, C.; Shao, W.; Kahouli, I.; Prakash, S. Effect of probiotics Lactobacillus and Bifidobacterium on gut-derived lipopolysaccharides and inflammatory cytokines: An in vitro study using a human colonic microbiota model. J. Microbiol. Biotechnol. 2013, 23, 518–526. [Google Scholar] [CrossRef] [PubMed]
- Kelsall, B.L.; Leon, F.; Smythies, L.E.; Smith, P.D. Antigen handling and presentation by mucosal dendritic cells and macrophages. In Mucosal Immunology; Elsevier: Amsterdam, The Netherlands, 2005; pp. 451–485. [Google Scholar]
- Laskin, J.D.; Heck, D.E.; Laskin, D.L. Multifunctional role of nitric oxide in inflammation. Trends Endocrinol. Metab. TEM 1994, 5, 377–382. [Google Scholar] [CrossRef]
- Oh, N.S.; Joung, J.Y.; Lee, J.Y.; Kim, Y. Probiotic and anti-inflammatory potential of Lactobacillus rhamnosus 4B15 and Lactobacillus gasseri 4M13 isolated from infant feces. PLoS ONE 2018, 13, e0192021. [Google Scholar] [CrossRef] [PubMed]
- Maitra, U.; Deng, H.; Glaros, T.; Baker, B.; Capelluto, D.G.; Li, Z.; Li, L. Molecular mechanisms responsible for the selective and low-grade induction of proinflammatory mediators in murine macrophages by lipopolysaccharide. J. Immunol. 2012, 189, 1014–1023. [Google Scholar] [CrossRef]
- Neurath, M.F. Cytokines in inflammatory bowel disease. Nat. Rev. Immunol. 2014, 14, 329–342. [Google Scholar] [CrossRef]
- Matsumoto, S.; Hara, T.; Hori, T.; Mitsuyama, K.; Nagaoka, M.; Tomiyasu, N.; Suzuki, A.; Sata, M. Probiotic Lactobacillus-induced improvement in murine chronic inflammatory bowel disease is associated with the down-regulation of pro-inflammatory cytokines in lamina propria mononuclear cells. Clin. Exp. Immunol. 2005, 140, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Marin, M.L.; Lee, J.H.; Murtha, J.; Ustunol, Z.; Pestka, J.J. Differential cytokine production in clonal macrophage and T-cell lines cultured with bifidobacteria. J. Dairy Sci. 1997, 80, 2713–2720. [Google Scholar] [CrossRef]
- Cross, M.L.; Ganner, A.; Teilab, D.; Fray, L.M. Patterns of cytokine induction by gram-positive and gram-negative probiotic bacteria. FEMS Immunol. Med Microbiol. 2004, 42, 173–180. [Google Scholar] [CrossRef]
- Marcinkiewicz, J.; Ciszek, M.; Bobek, M.; Strus, M.; Heczko, P.B.; Kurnyta, M.; Biedroń, R.; Chmielarczyk, A. Differential inflammatory mediator response in vitro from murine macrophages to lactobacilli and pathogenic intestinal bacteria. Int. J. Exp. Pathol. 2007, 88, 155–164. [Google Scholar] [CrossRef]
- Rong, J.; Zheng, H.; Liu, M.; Hu, X.; Wang, T.; Zhang, X.; Jin, F.; Wang, L. Probiotic and anti-inflammatory attributes of an isolate Lactobacillus helveticus NS8 from Mongolian fermented koumiss. BMC Microbiol. 2015, 15, 196. [Google Scholar] [CrossRef]
- Peña, J.A.; Versalovic, J. Lactobacillus rhamnosus GG decreases TNF-alpha production in lipopolysaccharide-activated murine macrophages by a contact-independent mechanism. Cell. Microbiol. 2003, 5, 277–285. [Google Scholar] [CrossRef]
- Wang, H.; Zhou, C.; Huang, J.; Kuai, X.; Shao, X. The potential therapeutic role of Lactobacillus reuteri for treatment of inflammatory bowel disease. Am. J. Transl. Res. 2020, 12, 1569–1583. [Google Scholar] [PubMed]
- Vezza, T.; Algieri, F.; Rodríguez-Nogales, A.; Garrido-Mesa, J.; Utrilla, M.P.; Talhaoui, N.; Gómez-Caravaca, A.M.; Segura-Carretero, A.; Rodríguez-Cabezas, M.E.; Monteleone, G.; et al. Immunomodulatory properties of Olea europaea leaf extract in intestinal inflammation. Mol. Nutr. Food Res. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Nogales, A.; Algieri, F.; Garrido-Mesa, J.; Vezza, T.; Utrilla, M.P.; Chueca, N.; Garcia, F.; Olivares, M.; Rodríguez-Cabezas, M.E.; Gálvez, J. Differential intestinal anti-inflammatory effects of Lactobacillus fermentum and Lactobacillus salivarius in DSS mouse colitis: Impact on microRNAs expression and microbiota composition. Mol. Nutr. Food Res. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Nogales, A.; Algieri, F.; Garrido-Mesa, J.; Vezza, T.; Utrilla, M.P.; Chueca, N.; Fernández-Caballero, J.A.; García, F.; Rodríguez-Cabezas, M.E.; Gálvez, J. The Administration of Escherichia coli Nissle 1917 Ameliorates Development of DSS-Induced Colitis in Mice. Front. Pharmacol. 2018, 9, 468. [Google Scholar] [CrossRef] [PubMed]
- O’Mahony, L.; O’Callaghan, L.; McCarthy, J.; Shilling, D.; Scully, P.; Sibartie, S.; Kavanagh, E.; Kirwan, W.O.; Redmond, H.P.; Collins, J.K.; et al. Differential cytokine response from dendritic cells to commensal and pathogenic bacteria in different lymphoid compartments in humans. Am. J. Physiol. Gastrointest. Liver Physiol. 2006, 290, G839–G845. [Google Scholar] [CrossRef]
- Foligne, B.; Nutten, S.; Grangette, C.; Dennin, V.; Goudercourt, D.; Poiret, S.; Dewulf, J.; Brassart, D.; Mercenier, A.; Pot, B. Correlation between in vitro and in vivo immunomodulatory properties of lactic acid bacteria. World J. Gastroenterol. 2007, 13, 236–243. [Google Scholar] [CrossRef] [PubMed]
- Wells, J.M. Immunomodulatory mechanisms of lactobacilli. Microb. Cell Fact. 2011, 10 (Suppl. 1), S17. [Google Scholar] [CrossRef]
- Pagnini, C.; Corleto, V.D.; Martorelli, M.; Lanini, C.; D’Ambra, G.; Di Giulio, E.; Delle Fave, G. Mucosal adhesion and anti-inflammatory effects of Lactobacillus rhamnosus GG in the human colonic mucosa: A proof-of-concept study. World J. Gastroenterol. 2018, 24, 4652–4662. [Google Scholar] [CrossRef]



| Gene | Sequence (5′-3′) | Annealing Temperature (°C) |
|---|---|---|
| GAPDH | FW:CCATCACCATCTTCCAGGAG RV:CCTGCTTCACCACCTTCTTG | 60 |
| TNF- α | FW:CCATCACCATCTTCCAGGAG RV:CTTCACAGAGCAATGACTCC | 56 |
| IL-6 | FW:TAGTCCTTCCTACCCCAATTTCC RV:TTGGTCCTTAGCCACTCCTTC | 60 |
| IL-1β | FW:TGATGAGAATGACCTCTTCT RV:CTTCTTCAAAGATGAAGGAAA | 55 |
| MCP-1 | FW:CAGCTGGGGACAGAATGGGG RV:GAGCTCTCTGGTACTCTTTTG | 62 |
| MIP-2 | FW:CAGTGAGCTGCGCTGTCCAATG RV:CAGTTAGCCTTGCCTTTGTTCAG | 60 |
| iNOS | FW:GTTGAAGACTGAGACTCTGG RV:GACTAGGCTACTCCGTGGA | 56 |
| ICAM-1 | FW:GAGGAGGTGAATGTATAAGTTATG RV:GGATGTGGAGGAGCAGAG | 60 |
| t (h) | INIA P572 | Glycerol | INIA P572 Glycerol | |
|---|---|---|---|---|
| Total aerobes | 0 | 7.19 ± 0.12 aA | 7.30 ± 0.10 aA | 7.38 ± 0.10 aA |
| 6 | 9.13 ± 0.23 bA | 9.19 ± 0.07 bA | 8.83 ± 0.43 bA | |
| 24 | 9.32 ± 0.05 bA | 9.08 ± 0.26 bA | 9.44 ± 0.16 bA | |
| Anaerobes | 0 | 7.58 ± 0.18 aA | 7.34 ± 0.10 aA | 7.46 ± 0.08 aA |
| 6 | 9.16 ± 0.13 bA | 9.12 ± 0.14 bA | 9.15 ± 0.23 bA | |
| 24 | 9.31 ± 0.09 bA | 9.13 ± 0.31 bA | 9.45 ± 0.10 bA | |
| Bacteroides | 0 | 6.85 ± 0.24 aA | 6.57 ± 0.07 aA | 7.13 ± 0.40 aA |
| 6 | 7.12 ± 0.04 aA | 6.63 ± 0.22 aA | 7.41 ± 0.21 aA | |
| 24 | 8.45 ± 0.10 bA | 8.26 ± 0.21 bA | 8.48 ± 0.21 bA | |
| Bifidobacteria | 0 | 7.34 ± 0.02 aA | 7.19 ± 0.09 aA | 7.30 ± 0.08 aA |
| 6 | 9.19 ± 0.18 bA | 9.13 ± 0.12 bA | 9.19 ± 0.18 bA | |
| 24 | 9.26 ± 0.18 bA | 8.84 ± 0.51 bA | 9.43 ± 0.18 bA | |
| Clostridia | 0 | 6.72 ± 0.87 aA | 6.35 ± 0.06 aA | 6.44 ± 0.31 aA |
| 6 | 7.14 ± 0.79 aA | 6.19 ± 0.19 aA | 7.65 ± 0.38 bA | |
| 24 | 8.37 ± 0.11 aAB | 8.19 ± 0.08 bA | 8.68 ± 0.18 bB | |
| Enterobacteria | 0 | 6.38 ± 0.07 aA | 6.29 ± 0.10 aA | 6.31 ± 0.15 aA |
| 6 | 9.16 ± 0.18 bA | 9.16 ± 0.09 bA | 8.94 ± 0.53 aA | |
| 24 | 9.25 ± 0.16 bA | 8.63 ± 0.51 bA | 8.72 ± 1.18 aA | |
| Lactobacilli | 0 | 6.46 ± 0.56 aA | 5.16 ± 0.20 aA | 6.41 ± 0.55 aA |
| 6 | 7.44 ± 0.21 aA | 6.27 ± 0.30 aB | 7.40 ± 0.22 aA | |
| 24 | 7.28 ± 0.19 aA | 6.68 ± 0.74 aA | 7.40 ± 0,05 aA | |
| L. reuteri INIA P572 | 0 | 6.28 ± 0.62 a | – | 6.27 ± 0.54 a |
| 6 | 7.10 ± 0.43 b | – | 7.17 ± 0.23 b | |
| 24 | 7.00 ± 0.52 ab | – | 7.07 ± 0.26 b |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Diez-Echave, P.; Martín-Cabrejas, I.; Garrido-Mesa, J.; Langa, S.; Vezza, T.; Landete, J.M.; Hidalgo-García, L.; Algieri, F.; Mayer, M.J.; Narbad, A.; et al. Probiotic and Functional Properties of Limosilactobacillus reuteri INIA P572. Nutrients 2021, 13, 1860. https://doi.org/10.3390/nu13061860
Diez-Echave P, Martín-Cabrejas I, Garrido-Mesa J, Langa S, Vezza T, Landete JM, Hidalgo-García L, Algieri F, Mayer MJ, Narbad A, et al. Probiotic and Functional Properties of Limosilactobacillus reuteri INIA P572. Nutrients. 2021; 13(6):1860. https://doi.org/10.3390/nu13061860
Chicago/Turabian StyleDiez-Echave, Patricia, Izaskun Martín-Cabrejas, José Garrido-Mesa, Susana Langa, Teresa Vezza, José M. Landete, Laura Hidalgo-García, Francesca Algieri, Melinda J. Mayer, Arjan Narbad, and et al. 2021. "Probiotic and Functional Properties of Limosilactobacillus reuteri INIA P572" Nutrients 13, no. 6: 1860. https://doi.org/10.3390/nu13061860
APA StyleDiez-Echave, P., Martín-Cabrejas, I., Garrido-Mesa, J., Langa, S., Vezza, T., Landete, J. M., Hidalgo-García, L., Algieri, F., Mayer, M. J., Narbad, A., García-Lafuente, A., Medina, M., Rodríguez-Nogales, A., Rodríguez-Cabezas, M. E., Gálvez, J., & Arqués, J. L. (2021). Probiotic and Functional Properties of Limosilactobacillus reuteri INIA P572. Nutrients, 13(6), 1860. https://doi.org/10.3390/nu13061860









