Metabolic Basis of Creatine in Health and Disease: A Bioinformatics-Assisted Review
Abstract
1. Introduction
2. Methods
3. Findings
3.1. What Happens When Creatine Is Transported into Cells?
3.2. How Is the CK/PCr System Involved in Cellular Bioenergetics?
CK Isozymes
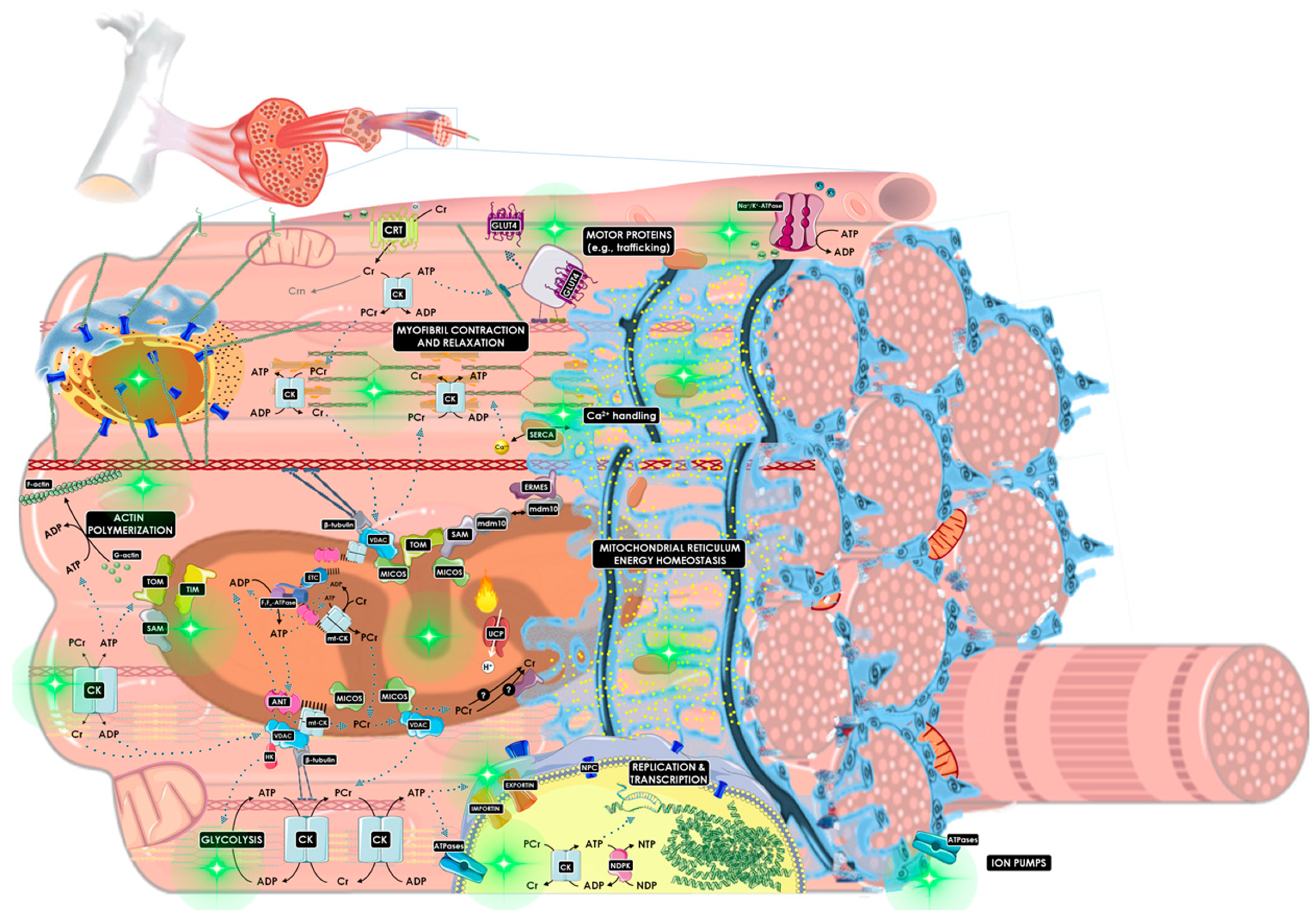
3.3. How Is the CK/PCr System Compartmentalized throughout the Cell?
3.3.1. Mitochondrial Reticulum
3.3.2. Cytosol and Cytoskeleton
3.3.3. Nucleus
3.3.4. Ion Pumps
3.3.5. Motor Proteins
3.4. What Is the Role of Creatine among Tissues?
| Tissue | CK Isozyme | Function |
|---|---|---|
| Brain | BB-CK uMtCK | Supports brain cells energy production and buffers ATP and ion pumping during electrical activity in neurons [50]. Oral Cr supplementation has been shown to improve memory in healthy adults, and potential benefits for aging and stressed individuals have been described [222]. Additionally, Cr supplementation seems beneficial in reducing the severity or enhancing recovery from mild traumatic brain injury, but further studies are needed not only as a post-injury therapy but also as a neuroprotective agent in populations at high risk of mild traumatic brain injury [223]. |
| Heart | MB-CK sMtCK | PCr provides about 80% of the energy needed for contraction and ion pumping, and about 20% of energy is transported into the cytoplasm via adenylate kinase and glycolytic phosphotransfer pathways [133,224]. MB-CK is an acute myocardial infarction marker [225]. |
| Testes | BB-CK uMtCK | Energy production and ATP buffer at axoneme, where microtubules and dynein use direct energy for sperm motility [13,226]. Cr concentrations and CK activity are potential indicators of sperm quality [227]. |
| Uterus | BB-CK uMtCK | Special attention should be paid to the increased Cr demand during pregnancy due to the important role of the PCr/CK system in the uterus and placenta for the maintenance and termination of gestation [34,228,229]. |
| Sensory organs | BB-CK MM-CK MB-CK uMtCK sMtCK | Visual system: important role in phototransduction by providing energy for the visual cycle, maintaining high local ATP/ADP ratios and consuming H+ produced by ATPases located in the outer segment and, thereby, preventing acidification [230]. |
| Auditory system: MM-CK is located in the strial marginal cells and dark cells while BB-CK in the inner hair cells. High levels of CK are also found in the cochlea’s inner and outer phalangeal cells. This provides a source of energy for ion transport and transduction activities in the inner ear [231]. | ||
| Olfactory system: Olfactory sensory neurons express BB-CK in the cilia [232]. In large cells within the olfactory neuroepithelium and ventral spinal cord, differential compartmentation of CK isoforms has been evident, with B-CK localized primarily in cell nuclei, whereas uMtCK is present in the cell body (but not within nuclei). In olfactory bulb neuroepithelium, both isoforms are expressed in the middle zone of the germinal layer associated with DNA synthesis [233]. | ||
| Tactile and skin system: BB-CK co-expresses with low amounts of uMtCK in suprabasal layers of the epidermis (cell of hair follicles, sebaceous glands, and the subcutaneous panniculus carnosus muscle). MM-CK and sMtCK were restricted to panniculus carnosus [234]. Epidermal CK is very important for cellular energy metabolism and might decline under oxidative stress conditions, including skin-aging processes; interestingly, application of Cr to skin cells in vitro and in vivo can refuel these cells energetically, and, thus, protect them against free radical-induced cell damage [235]. | ||
| Gustatory system: crucial for optimal cell and motor development and function [236]. CK is also involved in the control of maturation and maintenance of myofibers in the distal tongue [237,238]. | ||
| Intestines | BB-CK uMtCK | Distributed in the brush border web region, specifically at the contractile-ring myosin, to supply energy for contraction [239,240]. |
| Miscellaneous | BB-CK MB-CK uMtCK | CK has been associated with the clotting cascade by means of thrombin receptor signaling [241]. The CK/PCr system has also been implicated in the function of the immune cells [126]. Finally, Cr metabolism has been implicated in UCP-independent thermogenesis in the brown and beige adipose tissue [129,242], and B-CK has been shown to be a key effector of the futile Cr cycle [243]. |
3.5. What Is the Basis of Creatine Transport?
4. Limitations/Strengths and Future Directions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Brosnan, J.T.; Brosnan, M.E. Creatine: Endogenous Metabolite, Dietary, and Therapeutic Supplement. Annu. Rev. Nutr. 2007, 27, 241–261. [Google Scholar] [CrossRef]
- Béard, E.; Braissant, O. Synthesis and transport of creatine in the CNS: Importance for cerebral functions. J. Neurochem. 2010, 115, 297–313. [Google Scholar] [CrossRef] [PubMed]
- Moore, N.P. The distribution, metabolism and function of creatine in the male mammalian reproductive tract: A review. Int. J. Androl. 2000, 23, 4–12. [Google Scholar] [CrossRef]
- Brosnan, J.T.; da Silva, R.P.; Brosnan, M.E. The metabolic burden of creatine synthesis. Amino Acids 2011, 40, 1325–1331. [Google Scholar] [CrossRef] [PubMed]
- Humm, A.; Fritsche, E.; Steinbacher, S. Structure and reaction mechanism of L-arginine:glycine amidinotransferase. Biol. Chem. 1997, 378, 193–197. [Google Scholar] [PubMed]
- Komoto, J.; Yamada, T.; Takata, Y.; Konishi, K.; Ogawa, H.; Gomi, T.; Fujioka, M.; Takusagawa, F. Catalytic Mechanism of Guanidinoacetate Methyltransferase: Crystal Structures of Guanidinoacetate Methyltransferase Ternary Complexes. Biochemistry 2004, 43, 14385–14394. [Google Scholar] [CrossRef] [PubMed]
- Bonilla, D.A.; Moreno, Y. Molecular and metabolic insights of creatine supplementation on resistance training. Rev. Colomb. Química 2015, 44, 11–18. [Google Scholar] [CrossRef]
- Wyss, M.; Kaddurah-Daouk, R. Creatine and Creatinine Metabolism. Physiol. Rev. 2000, 80, 1107–1213. [Google Scholar] [CrossRef]
- Brosnan, M.E.; Edison, E.E.; da Silva, R.; Brosnan, J.T. New insights into creatine function and synthesis. Adv. Enzyme Regul. 2007, 47, 252–260. [Google Scholar] [CrossRef]
- Bonilla, D.A. A Systems Biology Approach to Creatine Metabolism. In Creatine: Biosynthesis, Health Effects and Clinical Perspectives; Hogan, L., Ed.; Nova Science Publishers Inc.: New York, NY, USA, 2017. [Google Scholar]
- Wallimann, T.; Wyss, M.; Brdiczka, D.; Nicolay, K.; Eppenberger, H.M. Intracellular compartmentation, structure and function of creatine kinase isoenzymes in tissues with high and fluctuating energy demands: The ‘phosphocreatine circuit’ for cellular energy homeostasis. Biochem. J. 1992, 281, 21–40. [Google Scholar] [CrossRef]
- Hemmer, W.; Wallimann, T. Functional Aspects of Creatine Kinase in Brain. Dev. Neurosci. 1993, 15, 249–260. [Google Scholar] [CrossRef] [PubMed]
- Wallimann, T.; Hemmer, W. Creatine kinase in non-muscle tissues and cells. Mol. Cell. Biochem. 1994, 133–134, 193–220. [Google Scholar] [CrossRef] [PubMed]
- Balestrino, M.; Gandolfo, C.; Perasso, L. Controlling the Flow of Energy: Inhibition and Stimulation of the Creatine Transporter. Curr. Enzym. Inhib. 2009, 5, 223–233. [Google Scholar] [CrossRef]
- Speer, O.; Neukomm, L.J.; Murphy, R.M.; Zanolla, E.; Schlattner, U.; Henry, H.; Snow, R.J.; Wallimann, T. Creatine transporters: A reappraisal. Mol. Cell. Biochem. 2004, 256, 407–424. [Google Scholar] [CrossRef]
- Christie, D.L. Functional Insights into the Creatine Transporter. In Creatine and Creatine Kinase in Health and Disease; Salomons, G.S., Wyss, M., Eds.; Springer: Dordrecht, The Netherlands, 2007; pp. 99–118. [Google Scholar] [CrossRef]
- Nash, S.R.; Giros, B.; Kingsmore, S.F.; Rochelle, J.M.; Suter, S.T.; Gregor, P.; Seldin, M.F.; Caron, M.G. Cloning, pharmacological characterization, and genomic localization of the human creatine transporter. Recept Channels 1994, 2, 165–174. [Google Scholar] [PubMed]
- Hanna-El-Daher, L.; Braissant, O. Creatine synthesis and exchanges between brain cells: What can be learned from human creatine deficiencies and various experimental models? Amino Acids 2016, 48, 1877–1895. [Google Scholar] [CrossRef] [PubMed]
- Braissant, O.; Rackayová, V.; Pierzchala, K.; Grosse, J.; McLin, V.A.; Cudalbu, C. Longitudinal neurometabolic changes in the hippocampus of a rat model of chronic hepatic encephalopathy. J. Hepatol. 2019, 71, 505–515. [Google Scholar] [CrossRef]
- Joncquel-Chevalier Curt, M.; Voicu, P.-M.; Fontaine, M.; Dessein, A.-F.; Porchet, N.; Mention-Mulliez, K.; Dobbelaere, D.; Soto-Ares, G.; Cheillan, D.; Vamecq, J. Creatine biosynthesis and transport in health and disease. Biochimie 2015, 119, 146–165. [Google Scholar] [CrossRef]
- Marques, E.P.; Wyse, A.T.S. Creatine as a Neuroprotector: An Actor that Can Play Many Parts. Neurotox. Res. 2019, 36, 411–423. [Google Scholar] [CrossRef]
- Wallimann, T.; Harris, R. Creatine: A miserable life without it. Amino Acids 2016, 48, 1739–1750. [Google Scholar] [CrossRef]
- Frampton, C.S.; Wilson, C.C.; Shankland, N.; Florence, A.J. Single-crystal neutron refinement of creatine monohydrate at 20 K and 123 K. J. Chem. Soc. Faraday Trans. 1997, 93, 1875–1879. [Google Scholar] [CrossRef]
- Arlin, J.-B.; Bhardwaj, R.M.; Johnston, A.; Miller, G.J.; Bardin, J.; MacDougall, F.; Fernandes, P.; Shankland, K.; David, W.I.F.; Florence, A.J. Structure and stability of two polymorphs of creatine and its monohydrate. CrystEngComm 2014, 16. [Google Scholar] [CrossRef]
- Dash, A.K.; Mo, Y.; Pyne, A. Solid-state Properties of Creatine Monohydrate. J. Pharm. Sci. 2002, 91, 708–718. [Google Scholar] [CrossRef]
- Pischel, I.; Gastner, T. Creatine—its Chemical Synthesis, Chemistry, and Legal Status. In Creatine and Creatine Kinase in Health and Disease; Salomons, G.S., Wyss, M., Eds.; Springer: Dordrecht, The Netherlands, 2007; pp. 291–307. [Google Scholar] [CrossRef]
- Kreider, R.B.; Kalman, D.S.; Antonio, J.; Ziegenfuss, T.N.; Wildman, R.; Collins, R.; Candow, D.G.; Kleiner, S.M.; Almada, A.L.; Lopez, H.L. International Society of Sports Nutrition position stand: Safety and efficacy of creatine supplementation in exercise, sport, and medicine. J. Int. Soc. of Sports Nutr. 2017, 14. [Google Scholar] [CrossRef] [PubMed]
- Wallimann, T.; Tokarska-Schlattner, M.; Schlattner, U. The creatine kinase system and pleiotropic effects of creatine. Amino Acids 2011, 40, 1271–1296. [Google Scholar] [CrossRef]
- Bonilla, D.A.; Pérez-Idárraga, A.; Odriozola-Martínez, A.; Kreider, R.B. The 4R’s Framework of Nutritional Strategies for Post-Exercise Recovery: A Review with Emphasis on New Generation of Carbohydrates. Int. J. Environ. Res. Public Health 2020, 18, 103. [Google Scholar] [CrossRef]
- Mielgo-Ayuso, J.; Calleja-Gonzalez, J.; Marqués-Jiménez, D.; Caballero-García, A.; Córdova, A.; Fernández-Lázaro, D. Effects of Creatine Supplementation on Athletic Performance in Soccer Players: A Systematic Review and Meta-Analysis. Nutrients 2019, 11, 757. [Google Scholar] [CrossRef]
- Kaviani, M.; Shaw, K.; Chilibeck, P.D. Benefits of Creatine Supplementation for Vegetarians Compared to Omnivorous Athletes: A Systematic Review. Int. J. Environ. Res. Public Health 2020, 17, 3041. [Google Scholar] [CrossRef]
- Bakian, A.V.; Huber, R.S.; Scholl, L.; Renshaw, P.F.; Kondo, D. Dietary creatine intake and depression risk among U.S. adults. Transl. Psychiatry 2020, 10. [Google Scholar] [CrossRef]
- Forbes, S.C.; Candow, D.G.; Smith-Ryan, A.E.; Hirsch, K.R.; Roberts, M.D.; VanDusseldorp, T.A.; Stratton, M.T.; Kaviani, M.; Little, J.P. Supplements and Nutritional Interventions to Augment High-Intensity Interval Training Physiological and Performance Adaptations—A Narrative Review. Nutrients 2020, 12, 390. [Google Scholar] [CrossRef] [PubMed]
- De Guingand, D.L.; Palmer, K.R.; Bilardi, J.E.; Ellery, S.J. Acceptability of dietary or nutritional supplementation in pregnancy (ADONS)—Exploring the consumer’s perspective on introducing creatine monohydrate as a pregnancy supplement. Midwifery 2020, 82. [Google Scholar] [CrossRef]
- Candow, D.G.; Forbes, S.C.; Chilibeck, P.D.; Cornish, S.M.; Antonio, J.; Kreider, R.B. Effectiveness of Creatine Supplementation on Aging Muscle and Bone: Focus on Falls Prevention and Inflammation. J. Clin. Med. 2019, 8, 488. [Google Scholar] [CrossRef]
- Stares, A.; Bains, M. The Additive Effects of Creatine Supplementation and Exercise Training in an Aging Population: A Systematic Review of Randomized Controlled Trials. J. Geriatr. Phys. Ther. 2020, 43, 99–112. [Google Scholar] [CrossRef] [PubMed]
- Rawson, E.S.; Miles, M.P.; Larson-Meyer, D.E. Dietary Supplements for Health, Adaptation, and Recovery in Athletes. Int. J. Sport Nutr. Exerc. Metab. 2018, 28, 188–199. [Google Scholar] [CrossRef]
- Clarke, H.; Kim, D.-H.; Meza, C.A.; Ormsbee, M.J.; Hickner, R.C. The Evolving Applications of Creatine Supplementation: Could Creatine Improve Vascular Health? Nutrients 2020, 12, 2834. [Google Scholar] [CrossRef] [PubMed]
- Machek, S.B.; Bagley, J.R. Creatine Monohydrate Supplementation: Considerations for Cognitive Performance in Athletes. Strength Cond. J. 2018, 40, 82–93. [Google Scholar] [CrossRef]
- Dolan, E.; Gualano, B.; Rawson, E.S. Beyond muscle: The effects of creatine supplementation on brain creatine, cognitive processing, and traumatic brain injury. Eur. J. Sport Sci. 2018, 19, 1–14. [Google Scholar] [CrossRef]
- Forbes, S.C.; Candow, D.G.; Ferreira, L.H.B.; Souza-Junior, T.P. Effects of Creatine Supplementation on Properties of Muscle, Bone, and Brain Function in Older Adults: A Narrative Review. J. Diet. Suppl. 2021, 1–18. [Google Scholar] [CrossRef]
- De Souza e Silva, A.; Pertille, A.; Reis Barbosa, C.G.; Aparecida de Oliveira Silva, J.; de Jesus, D.V.; Ribeiro, A.G.S.V.; Baganha, R.J.; de Oliveira, J.J. Effects of Creatine Supplementation on Renal Function: A Systematic Review and Meta-Analysis. J. Ren. Nutr. 2019, 29, 480–489. [Google Scholar] [CrossRef] [PubMed]
- Forbes, S.C.; Candow, D.G.; Krentz, J.R.; Roberts, M.D.; Young, K.C. Changes in Fat Mass Following Creatine Supplementation and Resistance Training in Adults ≥50 Years of Age: A Meta-Analysis. J. Funct. Morphol. Kinesiol. 2019, 4, 62. [Google Scholar] [CrossRef] [PubMed]
- Galvan, E.; Walker, D.K.; Simbo, S.Y.; Dalton, R.; Levers, K.; O’Connor, A.; Goodenough, C.; Barringer, N.D.; Greenwood, M.; Rasmussen, C.; et al. Acute and chronic safety and efficacy of dose dependent creatine nitrate supplementation and exercise performance. J. Int. Soc. Sports Nutr. 2016, 13. [Google Scholar] [CrossRef] [PubMed]
- Dalton, R.; Sowinski, R.; Grubic, T.; Collins, P.; Coletta, A.; Reyes, A.; Sanchez, B.; Koozehchian, M.; Jung, Y.; Rasmussen, C.; et al. Hematological and Hemodynamic Responses to Acute and Short-Term Creatine Nitrate Supplementation. Nutrients 2017, 9, 1359. [Google Scholar] [CrossRef] [PubMed]
- Ostojic, S.M.; Stajer, V.; Vranes, M.; Ostojic, J. Searching for a better formulation to enhance muscle bioenergetics: A randomized controlled trial of creatine nitrate plus creatininevs.creatine nitratevs.creatine monohydrate in healthy men. Food Sci. Nutr. 2019, 7, 3766–3773. [Google Scholar] [CrossRef]
- Antonio, J.; Candow, D.G.; Forbes, S.C.; Gualano, B.; Jagim, A.R.; Kreider, R.B.; Rawson, E.S.; Smith-Ryan, A.E.; VanDusseldorp, T.A.; Willoughby, D.S.; et al. Common questions and misconceptions about creatine supplementation: What does the scientific evidence really show? J. Int. Soc. Sports Nutr. 2021, 18. [Google Scholar] [CrossRef] [PubMed]
- Kreider, R.B.; Stout, J.R. Creatine in Health and Disease. Nutrients 2021, 13, 447. [Google Scholar] [CrossRef]
- Patra, S.; Bera, S.; SinhaRoy, S.; Ghoshal, S.; Ray, S.; Basu, A.; Schlattner, U.; Wallimann, T.; Ray, M. Progressive decrease of phosphocreatine, creatine and creatine kinase in skeletal muscle upon transformation to sarcoma. FEBS J. 2008, 275, 3236–3247. [Google Scholar] [CrossRef]
- Bender, A.; Klopstock, T. Creatine for neuroprotection in neurodegenerative disease: End of story? Amino Acids 2016, 48, 1929–1940. [Google Scholar] [CrossRef]
- Cheng, Y.; Chen, Y.; Shang, H. Aberrations of biochemical indicators in amyotrophic lateral sclerosis: A systematic review and meta-analysis. Transl. Neurodegener. 2021, 10. [Google Scholar] [CrossRef]
- Salazar, M.D.; Zelt, N.B.; Saldivar, R.; Kuntz, C.P.; Chen, S.; Penn, W.D.; Bonneau, R.; Koehler Leman, J.; Schlebach, J.P. Classification of the Molecular Defects Associated with Pathogenic Variants of the SLC6A8 Creatine Transporter. Biochemistry 2020, 59, 1367–1377. [Google Scholar] [CrossRef]
- Salomons, G.S.; van Dooren, S.J.; Verhoeven, N.M.; Cecil, K.M.; Ball, W.S.; Degrauw, T.J.; Jakobs, C. X-linked creatine-transporter gene (SLC6A8) defect: A new creatine-deficiency syndrome. Am. J. Hum. Genet. 2001, 68, 1497–1500. [Google Scholar] [CrossRef]
- Shearer, J.; Weljie, A.M. Biomarkers of skeletal muscle regulation, metabolism and dysfunction. In Metabolomics and Systems Biology in Human Health and Medicine; Jones, O., Ed.; CABI: Oxfordshire, UK, 2014; pp. 157–170. [Google Scholar] [CrossRef]
- McLeish, M.J.; Kenyon, G.L. Relating Structure to Mechanism in Creatine Kinase. Crit. Rev. Biochem. Mol. Biol. 2008, 40, 1–20. [Google Scholar] [CrossRef]
- Stout, J.R.; Antonio, J.; Kalman, D. Essentials of Creatine in Sports and Health; Humana Press: New York, USA, 2008. [Google Scholar] [CrossRef]
- Ellington, W.R. Phosphocreatine represents a thermodynamic and functional improvement over other muscle phosphagens. J. Exp. Biol. 1989, 143, 177–194. [Google Scholar]
- Uzzan, M.; Nechrebeki, J.; Zhou, P.; Labuza, T.P. Effect of water activity and temperature on the stability of creatine during storage. Drug Dev. Ind. Pharm. 2009, 35, 1003–1008. [Google Scholar] [CrossRef] [PubMed]
- Harris, R.C.; Söderlund, K.; Hultman, E. Elevation of creatine in resting and exercised muscle of normal subjects by creatine supplementation. Clin. Sci. 1992, 83, 367–374. [Google Scholar] [CrossRef] [PubMed]
- Vermeulen, A.; Wieme, R.; Robbrecht, J.; De Buyzere, M.; De Slypere, J.P.; Delanghe, J. Normal reference values for creatine, creatinine, and carnitine are lower in vegetarians. Clin. Chem. 1989, 35, 1802–1803. [Google Scholar] [CrossRef]
- Blancquaert, L.; Baguet, A.; Bex, T.; Volkaert, A.; Everaert, I.; Delanghe, J.; Petrovic, M.; Vervaet, C.; De Henauw, S.; Constantin-Teodosiu, D.; et al. Changing to a vegetarian diet reduces the body creatine pool in omnivorous women, but appears not to affect carnitine and carnosine homeostasis: A randomised trial. Br. J. Nutr. 2018, 119, 759–770. [Google Scholar] [CrossRef]
- Balsom, P.D.; Söderlund, K.; Ekblom, B. Creatine in Humans with Special Reference to Creatine Supplementation. Sports Med. 1994, 18, 268–280. [Google Scholar] [CrossRef]
- Wu, G. Important roles of dietary taurine, creatine, carnosine, anserine and 4-hydroxyproline in human nutrition and health. Amino Acids 2020, 52, 329–360. [Google Scholar] [CrossRef]
- Casey, A.; Constantin-Teodosiu, D.; Howell, S.; Hultman, E.; Greenhaff, P.L. Creatine ingestion favorably affects performance and muscle metabolism during maximal exercise in humans. Am. J. Physiol. Endocrinol. Metab. 1996, 271, E31–E37. [Google Scholar] [CrossRef]
- Greenhaff, P.L.; Bodin, K.; Soderlund, K.; Hultman, E. Effect of oral creatine supplementation on skeletal muscle phosphocreatine resynthesis. Am. J. Physiol. Endocrinol. Metab. 1994, 266, E725–E730. [Google Scholar] [CrossRef] [PubMed]
- Dechent, P.; Pouwels, P.J.W.; Wilken, B.; Hanefeld, F.; Frahm, J. Increase of total creatine in human brain after oral supplementation of creatine-monohydrate. Am. J. Physiol. Regul. Integr. Comp. Physiol. 1999, 277, R698–R704. [Google Scholar] [CrossRef] [PubMed]
- Kreider, R.; Willoughby, D.; Greenwood, M.; Parise, G.; Payne, E.T. Effects of serum creatine supplementation on muscle creatine and phosphagen levels. J. Exerc. Physiol. Online 2003, 6, 24–33. [Google Scholar]
- Schulze, A. Creatine deficiency syndromes. Mol. Cell. Biochem. 2003, 244, 143–150. [Google Scholar] [CrossRef] [PubMed]
- Stockler-Ipsiroglu, S.; Apatean, D.; Battini, R.; DeBrosse, S.; Dessoffy, K.; Edvardson, S.; Eichler, F.; Johnston, K.; Koeller, D.M.; Nouioua, S.; et al. Arginine: Glycine amidinotransferase (AGAT) deficiency: Clinical features and long term outcomes in 16 patients diagnosed worldwide. Mol. Genet. Metab. 2015, 116, 252–259. [Google Scholar] [CrossRef] [PubMed]
- Stöckler-Ipsiroglu, S.; Battini, R.; DeGrauw, T.; Schulze, A. Disorders of Creatine Metabolism. In Physician’s Guide to the Treatment and Follow-Up of Metabolic Diseases; Blau, N., Leonard, J., Hoffmann, G.F., Clarke, J.T.R., Eds.; Springer: Berlin/Heidelberg, Germany, 2006; pp. 255–265. [Google Scholar] [CrossRef]
- Mesa, J.L.M.; Ruiz, J.R.; Gonzalez-Gross, M.M.; Gutierrez Sainz, A.; Castillo Garzon, M.J. Oral Creatine Supplementation and Skeletal Muscle Metabolism in Physical Exercise. Sports Med. 2002, 32, 903–944. [Google Scholar] [CrossRef] [PubMed]
- Harris, R.C.; Almada, A.L.; Harris, D.B.; Dunnett, M.; Hespel, P. The creatine content of Creatine Serum™ and the change in the plasma concentration with ingestion of a single dose. J. Sports Sci. 2004, 22, 851–857. [Google Scholar] [CrossRef]
- Brault, J.J.; Towse, T.F.; Slade, J.M.; Meyer, R.A. Parallel Increases in Phosphocreatine and Total Creatine in Human Vastus Lateralis Muscle during Creatine Supplementation. Int. J. Sport Nutr. Exerc. Metab. 2007, 17, 624–634. [Google Scholar] [CrossRef]
- Broxterman, R.M.; Layec, G.; Hureau, T.J.; Amann, M.; Richardson, R.S. Skeletal muscle bioenergetics during all-out exercise: Mechanistic insight into the oxygen uptake slow component and neuromuscular fatigue. J. Appl. Physiol. 2017, 122, 1208–1217. [Google Scholar] [CrossRef]
- Burnley, M.; Jones, A.M. Oxygen uptake kinetics as a determinant of sports performance. Eur. J. Sport Sci. 2007, 7, 63–79. [Google Scholar] [CrossRef]
- Sweeney, H.L. The importance of the creatine kinase reaction: The concept of metabolic capacitance. Med. Sci. Sports Exerc. 1994, 26, 30–36. [Google Scholar] [CrossRef] [PubMed]
- Francescato, M.P.; Cettolo, V.; di Prampero, P.E. Influence of phosphagen concentration on phosphocreatine breakdown kinetics. Data from human gastrocnemius muscle. J. Appl. Physiol. 2008, 105, 158–164. [Google Scholar] [CrossRef] [PubMed]
- Meyer, R.A. A linear model of muscle respiration explains monoexponential phosphocreatine changes. Am. J. Physiol. Cell Physiol. 1988, 254, C548–C553. [Google Scholar] [CrossRef] [PubMed]
- Willis, W.T.; Jackman, M.R.; Messer, J.I.; Kuzmiak-Glancy, S.; Glancy, B. A Simple Hydraulic Analog Model of Oxidative Phosphorylation. Med. Sci. Sports Exerc. 2016, 48, 990–1000. [Google Scholar] [CrossRef]
- Gonzalez-Freire, M.; Scalzo, P.; D’Agostino, J.; Moore, Z.A.; Diaz-Ruiz, A.; Fabbri, E.; Zane, A.; Chen, B.; Becker, K.G.; Lehrmann, E.; et al. Skeletal muscle ex vivo mitochondrial respiration parallels decline in vivo oxidative capacity, cardiorespiratory fitness, and muscle strength: The Baltimore Longitudinal Study of Aging. Aging Cell 2018, 17. [Google Scholar] [CrossRef] [PubMed]
- Jones, A.M.; Wilkerson, D.P.; Fulford, J. Influence of dietary creatine supplementation on muscle phosphocreatine kinetics during knee-extensor exercise in humans. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2009, 296, R1078–R1087. [Google Scholar] [CrossRef] [PubMed]
- De Andrade Nemezio, K.M.; Bertuzzi, R.; Correia-Oliveira, C.R.; Gualano, B.; Bishop, D.J.; Lima-Silva, A.E. Effect of Creatine Loading on Oxygen Uptake during a 1-km Cycling Time Trial. Med. Sci. Sports Exerc. 2015, 47, 2660–2668. [Google Scholar] [CrossRef]
- Rigoulet, M.; Bouchez, C.L.; Paumard, P.; Ransac, S.; Cuvellier, S.; Duvezin-Caubet, S.; Mazat, J.P.; Devin, A. Cell energy metabolism: An update. Biochim. Biophys. Acta BBA Bioenerg. 2020, 1861. [Google Scholar] [CrossRef]
- Sumien, N.; Shetty, R.A.; Gonzales, E.B. Creatine, Creatine Kinase, and Aging. In Biochemistry and Cell Biology of Ageing: Part I Biomedical Science; Harris, J., Korolchuk, V., Eds.; Springer: Singapore, 2018; pp. 145–168. [Google Scholar] [CrossRef]
- Schlattner, U.; Kay, L.; Tokarska-Schlattner, M. Mitochondrial Proteolipid Complexes of Creatine Kinase. In Membrane Protein Complexes: Structure and Function; Harris, J., Boekema, E., Eds.; Springer: Singapore, 2018; pp. 365–408. [Google Scholar] [CrossRef]
- Dzeja, P.P.; Terzic, A. Phosphotransfer networks and cellular energetics. J. Exp. Biol. 2003, 206, 2039–2047. [Google Scholar] [CrossRef]
- Bessman, S.P.; Carpenter, C.L. The Creatine-Creatine Phosphate Energy Shuttle. Annu. Rev. Biochem. 1985, 54, 831–862. [Google Scholar] [CrossRef]
- Kongas, O.; van Beek, J. Creatine kinase in energy metabolic signaling in muscle. Nat. Preced. 2007. [Google Scholar] [CrossRef]
- Fiedler, G.B.; Schmid, A.I.; Goluch, S.; Schewzow, K.; Laistler, E.; Niess, F.; Unger, E.; Wolzt, M.; Mirzahosseini, A.; Kemp, G.J.; et al. Skeletal muscle ATP synthesis and cellular H+ handling measured by localized 31P-MRS during exercise and recovery. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef]
- Barclay, C.J. Energy demand and supply in human skeletal muscle. J. Muscle Res. Cell Motil. 2017, 38, 143–155. [Google Scholar] [CrossRef] [PubMed]
- Uda, K.; Ellington, W.R.; Suzuki, T. A diverse array of creatine kinase and arginine kinase isoform genes is present in the starlet sea anemone Nematostella vectensis, a cnidarian model system for studying developmental evolution. Gene 2012, 497, 214–227. [Google Scholar] [CrossRef] [PubMed]
- Bertin, M.; Pomponi, S.M.; Kokuhuta, C.; Iwasaki, N.; Suzuki, T.; Ellington, W.R. Origin of the genes for the isoforms of creatine kinase. Gene 2007, 392, 273–282. [Google Scholar] [CrossRef] [PubMed]
- Eppenberger, H.M.; Dawson, D.M.; Kaplan, N.O. The comparative enzymology of creatine kinases. I. Isolation and characterization from chicken and rabbit tissues. J. Biol. Chem. 1967, 242, 204–209. [Google Scholar] [CrossRef]
- Wallimann, T.; Tokarska-Schlattner, M.; Neumann, D.; Epand, R.M.; Epand, R.F.; Andres, R.H.; Widmer, H.R.; Hornemann, T.; Saks, V.; Agarkova, I.; et al. The Phosphocreatine Circuit: Molecular and Cellular Physiology of Creatine Kinases, Sensitivity to Free Radicals, and Enhancement by Creatine Supplementation. In Molecular System Bioenergetics; Saks, V., Ed.; Wiley-VCH: Weinheim, Germany, 2007; pp. 195–264. [Google Scholar] [CrossRef]
- Ramírez Ríos, S.; Lamarche, F.; Cottet-Rousselle, C.; Klaus, A.; Tuerk, R.; Thali, R.; Auchli, Y.; Brunisholz, R.; Neumann, D.; Barret, L.; et al. Regulation of brain-type creatine kinase by AMP-activated protein kinase: Interaction, phosphorylation and ER localization. Biochim. Biophys. Acta BBA Bioenerg. 2014, 1837, 1271–1283. [Google Scholar] [CrossRef]
- McFarland, E.W.; Kushmerick, M.J.; Moerland, T.S. Activity of creatine kinase in a contracting mammalian muscle of uniform fiber type. Biophys. J. 1994, 67, 1912–1924. [Google Scholar] [CrossRef]
- Wallimann, T.; Schlösser, T.; Eppenberger, H.M. Function of M-line-bound creatine kinase as intramyofibrillar ATP regenerator at the receiving end of the phosphorylcreatine shuttle in muscle. J. Biol. Chem. 1984, 259, 5238–5246. [Google Scholar] [CrossRef]
- Fritz-Wolf, K.; Schnyder, T.; Wallimann, T.; Kabsch, W. Structure of mitochondrial creatine kinase. Nature 1996, 381, 341–345. [Google Scholar] [CrossRef]
- Eder, M.; Fritz-Wolf, K.; Kabsch, W.; Wallimann, T.; Schlattner, U. Crystal structure of human ubiquitous mitochondrial creatine kinase. Proteins 2000, 39, 216–225. [Google Scholar] [CrossRef]
- Guzun, R.; Gonzalez-Granillo, M.; Karu-Varikmaa, M.; Grichine, A.; Usson, Y.; Kaambre, T.; Guerrero-Roesch, K.; Kuznetsov, A.; Schlattner, U.; Saks, V. Regulation of respiration in muscle cells in vivo by VDAC through interaction with the cytoskeleton and MtCK within Mitochondrial Interactosome. Biochim. Biophys. Acta BBA Biomembr. 2012, 1818, 1545–1554. [Google Scholar] [CrossRef]
- Wu, C.; Orozco, C.; Boyer, J.; Leglise, M.; Goodale, J.; Batalov, S.; Hodge, C.L.; Haase, J.; Janes, J.; Huss, J.W.; et al. BioGPS: An extensible and customizable portal for querying and organizing gene annotation resources. Genome Biol. 2009, 10. [Google Scholar] [CrossRef] [PubMed]
- Glancy, B.; Hartnell, L.M.; Malide, D.; Yu, Z.-X.; Combs, C.A.; Connelly, P.S.; Subramaniam, S.; Balaban, R.S. Mitochondrial reticulum for cellular energy distribution in muscle. Nature 2015, 523, 617–620. [Google Scholar] [CrossRef]
- Wallimann, T. The extended, dynamic mitochondrial reticulum in skeletal muscle and the creatine kinase (CK)/phosphocreatine (PCr) shuttle are working hand in hand for optimal energy provision. J. Muscle Res. Cell Motil. 2015, 36, 297–300. [Google Scholar] [CrossRef]
- Glancy, B.; Hartnell, L.M.; Combs, C.A.; Femnou, A.; Sun, J.; Murphy, E.; Subramaniam, S.; Balaban, R.S. Power Grid Protection of the Muscle Mitochondrial Reticulum. Cell Rep. 2017, 19, 487–496. [Google Scholar] [CrossRef]
- Saks, V.; Schlattner, U.; Tokarska-Schlattner, M.; Wallimann, T.; Bagur, R.; Zorman, S.; Pelosse, M.; Santos, P.D.; Boucher, F.; Kaambre, T.; et al. Systems Level Regulation of Cardiac Energy Fluxes Via Metabolic Cycles: Role of Creatine, Phosphotransfer Pathways, and AMPK Signaling. In Systems Biology of Metabolic and Signaling Networks; Aon, M., Saks, V., Schlattner, U., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 261–320. [Google Scholar] [CrossRef]
- Timohhina, N.; Guzun, R.; Tepp, K.; Monge, C.; Varikmaa, M.; Vija, H.; Sikk, P.; Kaambre, T.; Sackett, D.; Saks, V. Direct measurement of energy fluxes from mitochondria into cytoplasm in permeabilized cardiac cells in situ: Some evidence for mitochondrial interactosome. J. Bioenerg. Biomembr. 2009, 41, 259–275. [Google Scholar] [CrossRef]
- Guzun, R.; Saks, V. Application of the Principles of Systems Biology and Wiener’s Cybernetics for Analysis of Regulation of Energy Fluxes in Muscle Cells in Vivo. Int. J. Mol. Sci. 2010, 11, 982–1019. [Google Scholar] [CrossRef] [PubMed]
- Saks, V.; Guzun, R.; Timohhina, N.; Tepp, K.; Varikmaa, M.; Monge, C.; Beraud, N.; Kaambre, T.; Kuznetsov, A.; Kadaja, L.; et al. Structure–function relationships in feedback regulation of energy fluxes in vivo in health and disease: Mitochondrial Interactosome. Biochim. Biophys. Acta BBA Bioenerg. 2010, 1797, 678–697. [Google Scholar] [CrossRef] [PubMed]
- Karo, J.; Peterson, P.; Vendelin, M. Molecular Dynamics Simulations of Creatine Kinase and Adenine Nucleotide Translocase in Mitochondrial Membrane Patch. J. Biol. Chem. 2012, 287, 7467–7476. [Google Scholar] [CrossRef] [PubMed]
- Bonilla, D.A.; Marín, E.; Pérez, A.; Carbone, L.; Kammerer, M.; Vargas, S.; Lozano, J.; Barale, A.; Quiroga, L.; Mata, F.; et al. Thermogenesis and Obesity; A Brief Review and rs104894319 Polymorphism in Venezuelan Population. EC Nutr. 2018, 13, 4–16. [Google Scholar]
- Rousset, S.; Alves-Guerra, M.C.; Mozo, J.; Miroux, B.; Cassard-Doulcier, A.M.; Bouillaud, F.; Ricquier, D. The Biology of Mitochondrial Uncoupling Proteins. Diabetes 2004, 53, S130–S135. [Google Scholar] [CrossRef]
- Brand, M.D.; Esteves, T.C. Physiological functions of the mitochondrial uncoupling proteins UCP2 and UCP3. Cell Metab. 2005, 2, 85–93. [Google Scholar] [CrossRef]
- Ramsden, D.B.; Ho, P.W.L.; Ho, J.W.M.; Liu, H.F.; So, D.H.F.; Tse, H.M.; Chan, K.H.; Ho, S.L. Human neuronal uncoupling proteins 4 and 5 (UCP4 and UCP5): Structural properties, regulation, and physiological role in protection against oxidative stress and mitochondrial dysfunction. Brain Behav. 2012, 2, 468–478. [Google Scholar] [CrossRef]
- Krauss, S.; Zhang, C.-Y.; Lowell, B.B. The mitochondrial uncoupling-protein homologues. Nat. Rev. Mol. Cell Biol. 2005, 6, 248–261. [Google Scholar] [CrossRef] [PubMed]
- Busiello, R.A.; Savarese, S.; Lombardi, A. Mitochondrial uncoupling proteins and energy metabolism. Front. Physiol. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Pohl, E.E.; Rupprecht, A.; Macher, G.; Hilse, K.E. Important Trends in UCP3 Investigation. Front. Physiol. 2019, 10. [Google Scholar] [CrossRef]
- Skulachev, V.P. Fatty acid circuit as a physiological mechanism of uncoupling of oxidative phosphorylation. FEBS Lett. 1991, 294, 158–162. [Google Scholar] [CrossRef]
- Ježek, P.; Engstová, H.; Žáčková, M.; Vercesi, A.E.; Costa, A.D.T.; Arruda, P.; Garlid, K.D. Fatty acid cycling mechanism and mitochondrial uncoupling proteins. Biochim. Biophys. Acta BBA Bioenerg. 1998, 1365, 319–327. [Google Scholar] [CrossRef]
- Klingenberg, M.; Huang, S.-G. Structure and function of the uncoupling protein from brown adipose tissue. Biochim. Biophys. Acta BBA Biomembr. 1999, 1415, 271–296. [Google Scholar] [CrossRef]
- Fedorenko, A.; Lishko, P.V.; Kirichok, Y. Mechanism of Fatty-Acid-Dependent UCP1 Uncoupling in Brown Fat Mitochondria. Cell 2012, 151, 400–413. [Google Scholar] [CrossRef] [PubMed]
- Macher, G.; Koehler, M.; Rupprecht, A.; Kreiter, J.; Hinterdorfer, P.; Pohl, E.E. Inhibition of mitochondrial UCP1 and UCP3 by purine nucleotides and phosphate. Biochim. Biophys. Acta BBA Biomembr. 2018, 1860, 664–672. [Google Scholar] [CrossRef]
- Kazak, L.; Chouchani, E.T.; Jedrychowski, M.P.; Erickson, B.K.; Shinoda, K.; Cohen, P.; Vetrivelan, R.; Lu, G.Z.; Laznik-Bogoslavski, D.; Hasenfuss, S.C.; et al. A Creatine-Driven Substrate Cycle Enhances Energy Expenditure and Thermogenesis in Beige Fat. Cell 2015, 163, 643–655. [Google Scholar] [CrossRef]
- Kazak, L.; Chouchani, E.T.; Lu, G.Z.; Jedrychowski, M.P.; Bare, C.J.; Mina, A.I.; Kumari, M.; Zhang, S.; Vuckovic, I.; Laznik-Bogoslavski, D.; et al. Genetic Depletion of Adipocyte Creatine Metabolism Inhibits Diet-Induced Thermogenesis and Drives Obesity. Cell Metab. 2017, 26, 660–671.e663. [Google Scholar] [CrossRef]
- Bertholet, A.M.; Kazak, L.; Chouchani, E.T.; Bogaczyńska, M.G.; Paranjpe, I.; Wainwright, G.L.; Bétourné, A.; Kajimura, S.; Spiegelman, B.M.; Kirichok, Y. Mitochondrial Patch Clamp of Beige Adipocytes Reveals UCP1-Positive and UCP1-Negative Cells Both Exhibiting Futile Creatine Cycling. Cell Metab. 2017, 25, 811–822.e814. [Google Scholar] [CrossRef] [PubMed]
- Kazak, L.; Roesler, A. UCP1-independent thermogenesis. Biochem. J. 2020, 477, 709–725. [Google Scholar] [CrossRef]
- Kazak, L.; Cohen, P. Creatine metabolism: Energy homeostasis, immunity and cancer biology. Nat. Rev. Endocrinol. 2020, 16, 421–436. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, K.; Yamada, T. UCP1 Dependent and Independent Thermogenesis in Brown and Beige Adipocytes. Front. Endocrinol. (Lausanne) 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Chouchani, E.T.; Kajimura, S. Metabolic adaptation and maladaptation in adipose tissue. Nat. Metab. 2019, 1, 189–200. [Google Scholar] [CrossRef] [PubMed]
- Wallimann, T.; Tokarska-Schlattner, M.; Kay, L.; Schlattner, U. Role of creatine and creatine kinase in UCP1-independent adipocyte thermogenesis. Am. J. Physiol. Endocrinol. Metab. 2020, 319, E944–E946. [Google Scholar] [CrossRef]
- Connell, N.J.; Doligkeit, D.; Andriessen, C.; Kornips-Moonen, E.; Bruls, Y.M.H.; Schrauwen-Hinderling, V.B.; van de Weijer, T.; van Marken-Lichtenbelt, W.D.; Havekes, B.; Kazak, L.; et al. No evidence for brown adipose tissue activation after creatine supplementation in adult vegetarians. Nat. Metab. 2021, 3, 107–117. [Google Scholar] [CrossRef] [PubMed]
- Kraft, T.; Hornemann, T.; Stolz, M.; Nier, V.; Wallimann, T. Coupling of creatine kinase to glycolytic enzymes at the sarcomeric I-band of skeletal muscle: A biochemical study in situ. J. Muscle Res. Cell Motil. 2000, 21, 691–703. [Google Scholar] [CrossRef]
- Westerblad, H.; Allen, D.G.; Lännergren, J. Muscle Fatigue: Lactic Acid or Inorganic Phosphate the Major Cause? Physiology 2002, 17, 17–21. [Google Scholar] [CrossRef]
- Wu, F.; Beard, D.A. Roles of the creatine kinase system and myoglobin in maintaining energetic state in the working heart. BMC Syst. Biol. 2009, 3. [Google Scholar] [CrossRef]
- Gerlach, G.; Hofer, H.W. Interaction of immobilized phosphofructokinase with soluble muscle proteins. Biochim. Biophys. Acta BBA Gen. Subj. 1986, 881, 398–404. [Google Scholar] [CrossRef]
- Mor, I.; Cheung, E.C.; Vousden, K.H. Control of Glycolysis through Regulation of PFK1: Old Friends and Recent Additions. Cold Spring Harb. Symp. Quant. Biol. 2011, 76, 211–216. [Google Scholar] [CrossRef]
- Foucault, G.; Vacher, M.; Merkulova, T.; Keller, A.; Arrio-Dupont, M. Presence of enolase in the M-band of skeletal muscle and possible indirect interaction with the cytosolic muscle isoform of creatine kinase. Biochem. J. 1999, 338, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Brown, K.S.; Hettling, H.; van Beek, J.H.G.M. Analyzing the Functional Properties of the Creatine Kinase System with Multiscale ‘Sloppy’ Modeling. PLoS Comput. Biol. 2011, 7. [Google Scholar] [CrossRef]
- Bose, S.; French, S.; Evans, F.J.; Joubert, F.; Balaban, R.S. Metabolic Network Control of Oxidative Phosphorylation. J. Biol. Chem. 2003, 278, 39155–39165. [Google Scholar] [CrossRef] [PubMed]
- Saks, V.; Monge, C.; Guzun, R. Philosophical Basis and Some Historical Aspects of Systems Biology: From Hegel to Noble—Applications for Bioenergetic Research. Int. J. Mol. Sci. 2009, 10, 1161–1192. [Google Scholar] [CrossRef] [PubMed]
- Guzun, R.; Timohhina, N.; Tepp, K.; Monge, C.; Kaambre, T.; Sikk, P.; Kuznetsov, A.V.; Pison, C.; Saks, V. Regulation of respiration controlled by mitochondrial creatine kinase in permeabilized cardiac cells in situ. Biochim. Biophys. Acta BBA Bioenerg. 2009, 1787, 1089–1105. [Google Scholar] [CrossRef]
- Klepinin, A.; Ounpuu, L.; Mado, K.; Truu, L.; Chekulayev, V.; Puurand, M.; Shevchuk, I.; Tepp, K.; Planken, A.; Kaambre, T. The complexity of mitochondrial outer membrane permeability and VDAC regulation by associated proteins. J. Bioenerg. Biomembr. 2018, 50, 339–354. [Google Scholar] [CrossRef]
- Anflous-Pharayra, K.; Cai, Z.-J.; Craigen, W.J. VDAC1 serves as a mitochondrial binding site for hexokinase in oxidative muscles. Biochim. Biophys. Acta BBA Bioenerg. 2007, 1767, 136–142. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pedersen, P.L. Warburg, me and Hexokinase 2: Multiple discoveries of key molecular events underlying one of cancers’ most common phenotypes, the “Warburg Effect”, i.e., elevated glycolysis in the presence of oxygen. J. Bioenerg. Biomembr. 2007, 39, 211–222. [Google Scholar] [CrossRef] [PubMed]
- Sestili, P.; Martinelli, C.; Colombo, E.; Barbieri, E.; Potenza, L.; Sartini, S.; Fimognari, C. Creatine as an antioxidant. Amino Acids 2011, 40, 1385–1396. [Google Scholar] [CrossRef]
- Hofmann, P. Cancer and Exercise: Warburg Hypothesis, Tumour Metabolism and High-Intensity Anaerobic Exercise. Sports 2018, 6, 10. [Google Scholar] [CrossRef]
- Fadaka, A.; Ajiboye, B.; Ojo, O.; Adewale, O.; Olayide, I.; Emuowhochere, R. Biology of glucose metabolization in cancer cells. J. Oncol. Sci. 2017, 3, 45–51. [Google Scholar] [CrossRef]
- Marchesi, F.; Vignali, D.; Manini, B.; Rigamonti, A.; Monti, P. Manipulation of Glucose Availability to Boost Cancer Immunotherapies. Cancers Basel 2020, 12, 2940. [Google Scholar] [CrossRef]
- Balsom, P.D.; Söderlund, K.; Sjödin, B.; Ekblom, B. Skeletal muscle metabolism during short duration high-intensity exercise: Influence of creatine supplementation. Acta Physiol. Scand. 1995, 154, 303–310. [Google Scholar] [CrossRef]
- Balsom, P.D.; Ekblom, B.; Söerlund, K.; Sjödln, B.; Hultman, E. Creatine supplementation and dynamic high-intensity intermittent exercise. Scand. J. Med. Sci. Sports 2007, 3, 143–149. [Google Scholar] [CrossRef]
- Dos Santos, M.G. Estudio Del Metabolismo Energético Muscular Y De La Composición Corporal De Atletas Por Métodos No Destructivos; Universitat Autònoma de Barcelona: Barcelona, Spain, 2001. [Google Scholar]
- Ceddia, R.B.; Sweeney, G. Creatine supplementation increases glucose oxidation and AMPK phosphorylation and reduces lactate production in L6 rat skeletal muscle cells. J. Physiol. 2004, 555, 409–421. [Google Scholar] [CrossRef] [PubMed]
- Dobgenski, V.; Santos, M.; Kreider, R. Effects of creatine supplementation in the concentrations of creatine kinase, creatinine, urea and lactate on male swimmers. J. Nutr. Health 2016, 2, 1–5. [Google Scholar]
- Oliver, J.M.; Joubert, D.P.; Martin, S.E.; Crouse, S.F. Oral Creatine Supplementation’s Decrease of Blood Lactate During Exhaustive, Incremental Cycling. Int. J. Sport Nutr. Exerc. Metab. 2013, 23, 252–258. [Google Scholar] [CrossRef]
- Storey, K.B.; Hochachka, P.W. Activation of muscle glycolysis: A role for creatine phosphate in phosphofructokinase regulation. FEBS Lett. 1974, 46, 337–339. [Google Scholar] [CrossRef]
- Kemp, R.G. Inhibition of muscle pyruvate kinase by creatine phosphate. J. Biol. Chem. 1973, 248, 3963–3967. [Google Scholar] [CrossRef]
- Fu, J.Y.; Kemp, R.G. Activation of Muscle Fructose 1,6-Diphosphatase by Creatine Phosphate and Citrate. J. Biol. Chem. 1973, 248, 1124–1125. [Google Scholar] [CrossRef]
- Ponticos, M.; Lu, Q.L.; Hardie, D.G.; Partridge, T.A.; Carling, D. Dual regulation of the AMP-activated protein kinase provides a novel mechanism for the control of creatine kinase in skeletal muscle. EMBO J. 1998, 17, 1688–1699. [Google Scholar] [CrossRef] [PubMed]
- Jørgensen, S.B.; Richter, E.A.; Wojtaszewski, J.F.P. Role of AMPK in skeletal muscle metabolic regulation and adaptation in relation to exercise. J. Physiol. 2006, 574, 17–31. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Wang, X.; Li, J.; Zhu, X.; Gao, F.; Zhou, G. Creatine Monohydrate Enhances Energy Status and Reduces Glycolysis via Inhibition of AMPK Pathway in Pectoralis Major Muscle of Transport-Stressed Broilers. J. Agric. Food Chem. 2017, 65, 6991–6999. [Google Scholar] [CrossRef] [PubMed]
- Taylor, E.B.; Ellingson, W.J.; Lamb, J.D.; Chesser, D.G.; Compton, C.L.; Winder, W.W. Evidence against regulation of AMP-activated protein kinase and LKB1/STRAD/MO25 activity by creatine phosphate. Am. J. Physiol. Endocrinol. Metab. 2006, 290, E661–E669. [Google Scholar] [CrossRef] [PubMed]
- Eijnde, B.O.; Derave, W.; Wojtaszewski, J.F.P.; Richter, E.A.; Hespel, P. AMP kinase expression and activity in human skeletal muscle: Effects of immobilization, retraining, and creatine supplementation. J. Appl. Physiol. 2005, 98, 1228–1233. [Google Scholar] [CrossRef] [PubMed]
- Gautel, M.; Djinović-Carugo, K. The sarcomeric cytoskeleton: From molecules to motion. J. Exp. Biol. 2016, 219, 135–145. [Google Scholar] [CrossRef]
- Puurand, M.; Tepp, K.; Timohhina, N.; Aid, J.; Shevchuk, I.; Chekulayev, V.; Kaambre, T. Tubulin βII and βIII Isoforms as the Regulators of VDAC Channel Permeability in Health and Disease. Cells 2019, 8, 239. [Google Scholar] [CrossRef]
- Kuznetsov, A.V.; Javadov, S.; Guzun, R.; Grimm, M.; Saks, V. Cytoskeleton and regulation of mitochondrial function: The role of beta-tubulin II. Front. Physiol. 2013, 4, 82. [Google Scholar] [CrossRef]
- Raskin, A.; Lange, S.; Banares, K.; Lyon, R.C.; Zieseniss, A.; Lee, L.K.; Yamazaki, K.G.; Granzier, H.L.; Gregorio, C.C.; McCulloch, A.D.; et al. A Novel Mechanism Involving Four-and-a-half LIM Domain Protein-1 and Extracellular Signal-regulated Kinase-2 Regulates Titin Phosphorylation and Mechanics. J. Biol. Chem. 2012, 287, 29273–29284. [Google Scholar] [CrossRef]
- Henderson, C.A.; Gomez, C.G.; Novak, S.M.; Mi-Mi, L.; Gregorio, C.C. Overview of the Muscle Cytoskeleton. Compr. Physiol. 2017, 7, 891–944. [Google Scholar] [CrossRef]
- Kaasik, A.; Veksler, V.; Boehm, E.; Novotova, M.; Minajeva, A.; Ventura-Clapier, R.e. Energetic Crosstalk Between Organelles. Circ. Res. 2001, 89, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Piquereau, J.; Veksler, V.; Novotova, M.; Ventura-Clapier, R. Energetic Interactions Between Subcellular Organelles in Striated Muscles. Front. Cell Dev. Biol. 2020, 8. [Google Scholar] [CrossRef] [PubMed]
- Kuznetsov, A.V.; Javadov, S.; Grimm, M.; Margreiter, R.; Ausserlechner, M.J.; Hagenbuchner, J. Crosstalk between Mitochondria and Cytoskeleton in Cardiac Cells. Cells 2020, 9, 222. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-Lasso, D.C.; Romá-Mateo, C.; Pallardó, F.V.; Gonzalez-Cabo, P. Much More Than a Scaffold: Cytoskeletal Proteins in Neurological Disorders. Cells 2020, 9, 358. [Google Scholar] [CrossRef] [PubMed]
- Ross, J.A.; Levy, Y.; Ripolone, M.; Kolb, J.S.; Turmaine, M.; Holt, M.; Lindqvist, J.; Claeys, K.G.; Weis, J.; Monforte, M.; et al. Impairments in contractility and cytoskeletal organisation cause nuclear defects in nemaline myopathy. Acta Neuropathol. 2019, 138, 477–495. [Google Scholar] [CrossRef]
- Dowling, P.; Gargan, S.; Murphy, S.; Zweyer, M.; Sabir, H.; Swandulla, D.; Ohlendieck, K. The Dystrophin Node as Integrator of Cytoskeletal Organization, Lateral Force Transmission, Fiber Stability and Cellular Signaling in Skeletal Muscle. Proteomes 2021, 9, 9. [Google Scholar] [CrossRef] [PubMed]
- Lai, W.-F.; Wong, W.-T. Roles of the actin cytoskeleton in aging and age-associated diseases. Ageing Res. Rev. 2020, 58. [Google Scholar] [CrossRef] [PubMed]
- Perry, R.L.; Rudnick, M.A. Molecular mechanisms regulating myogenic determination and differentiation. Front. Biosci. 2000, 5, D750–D767. [Google Scholar] [CrossRef]
- O’Connor, R.S.; Steeds, C.M.; Wiseman, R.W.; Pavlath, G.K. Phosphocreatine as an energy source for actin cytoskeletal rearrangements during myoblast fusion. J. Physiol. 2008, 586, 2841–2853. [Google Scholar] [CrossRef]
- Simionescu-Bankston, A.; Pichavant, C.; Canner, J.P.; Apponi, L.H.; Wang, Y.; Steeds, C.; Olthoff, J.T.; Belanto, J.J.; Ervasti, J.M.; Pavlath, G.K. Creatine kinase B is necessary to limit myoblast fusion during myogenesis. Am. J. Physiol. Cell Physiol. 2015, 308, C919–C931. [Google Scholar] [CrossRef] [PubMed]
- Lehka, L.; Rędowicz, M.J. Mechanisms regulating myoblast fusion: A multilevel interplay. Semin. Cell Dev. Biol. 2020, 104, 81–92. [Google Scholar] [CrossRef]
- Stroud, M.J.; Banerjee, I.; Veevers, J.; Chen, J. Linker of Nucleoskeleton and Cytoskeleton Complex Proteins in Cardiac Structure, Function, and Disease. Circ. Res. 2014, 114, 538–548. [Google Scholar] [CrossRef]
- Spichal, M.; Fabre, E. The Emerging Role of the Cytoskeleton in Chromosome Dynamics. Front. Genet. 2017, 8. [Google Scholar] [CrossRef]
- Loo, T.H.; Ye, X.; Chai, R.J.; Ito, M.; Bonne, G.; Ferguson-Smith, A.C.; Stewart, C.L. The mammalian LINC complex component SUN1 regulates muscle regeneration by modulating drosha activity. eLife 2019, 8. [Google Scholar] [CrossRef]
- Piccus, R.; Brayson, D. The nuclear envelope: LINCing tissue mechanics to genome regulation in cardiac and skeletal muscle. Biol. Lett. 2020, 16. [Google Scholar] [CrossRef] [PubMed]
- Brull, A.; Morales Rodriguez, B.; Bonne, G.; Muchir, A.; Bertrand, A.T. The Pathogenesis and Therapies of Striated Muscle Laminopathies. Front. Physiol. 2018, 9. [Google Scholar] [CrossRef]
- Starr, D.A.; Rose, L.S. TorsinA regulates the LINC to moving nuclei. J. Cell Biol. 2017, 216, 543–545. [Google Scholar] [CrossRef]
- Dzeja, P.P.; Bortolon, R.; Perez-Terzic, C.; Holmuhamedov, E.L.; Terzic, A. Energetic communication between mitochondria and nucleus directed by catalyzed phosphotransfer. Proc. Natl. Acad. Sci. USA 2002, 99, 10156–10161. [Google Scholar] [CrossRef] [PubMed]
- Dzeja, P.P.; Terzic, A. Adenylate kinase and creatine kinase phosphotransfer in regulation of mitochondrial respiration and cellular energetic efficiency. In Creatine Kinase; Vial, C., Ed.; Nova Science Publishers: London, UK, 2006; pp. 195–221. [Google Scholar]
- Adam, K.; Ning, J.; Reina, J.; Hunter, T. NME/NM23/NDPK and Histidine Phosphorylation. Int. J. Mol. Sci. 2020, 21, 5848. [Google Scholar] [CrossRef]
- Attwood, P.V.; Muimo, R. The actions of NME1/NDPK-A and NME2/NDPK-B as protein kinases. Lab. Invest. 2017, 98, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Macara, I.G. Transport into and out of the Nucleus. Microbiol. Mol. Biol. Rev. 2001, 65, 570–594. [Google Scholar] [CrossRef] [PubMed]
- de Groof, A.J.C.; Fransen, J.A.M.; Errington, R.J.; Willems, P.H.G.M.; Wieringa, B.; Koopman, W.J.H. The Creatine Kinase System Is Essential for Optimal Refill of the Sarcoplasmic Reticulum Ca2+ Store in Skeletal Muscle. J. Biol. Chem. 2002, 277, 5275–5284. [Google Scholar] [CrossRef] [PubMed]
- Duke, A.M.; Steele, D.S. Effects of creatine phosphate on Ca2+regulation by the sarcoplasmic reticulum in mechanically skinned rat skeletal muscle fibres. J. Physiol. 1999, 517, 447–458. [Google Scholar] [CrossRef]
- Sistermans, E.A.; Klaassen, C.H.W.; Peters, W.; Swarts, H.G.P.; Jap, P.H.K.; De Pont, J.J.H.H.M.; Wieringa, B. Co-localization and functional coupling of creatine kinase B and gastric H+/K+-ATPase on the apical membrane and the tubulovesicular system of parietal cells. Biochem. J. 1995, 311, 445–451. [Google Scholar] [CrossRef]
- Grosse, R.; Spitzer, E.; Kupriyanov, V.V.; Saks, V.A.; Repke, K.R.H. Coordinate interplay between (Na+ + K+)-ATPase and creatine phosphokinase optimizes (Na+/K+)-antiport across the membrane of vesicles formed from the plasma membrane of cardiac muscle cell. Biochim. Biophys. Acta BBA Biomembr. 1980, 603, 142–156. [Google Scholar] [CrossRef]
- Yang, Y.-C.; Fann, M.-J.; Chang, W.-H.; Tai, L.-H.; Jiang, J.-H.; Kao, L.-S. Regulation of Sodium-Calcium Exchanger Activity by Creatine Kinase under Energy-compromised Conditions. J. Biol. Chem. 2010, 285, 28275–28285. [Google Scholar] [CrossRef]
- Kato, Y.; Miyakawa, T.; Tanokura, M. Overview of the mechanism of cytoskeletal motors based on structure. Biophys. Rev. 2017, 10, 571–581. [Google Scholar] [CrossRef]
- Jena, B.P. Myosin: Cellular Molecular Motor. In Cellular Nanomachines; Jena, B.P., Ed.; Springer: Cham, Switzerland, 2020; pp. 79–89. [Google Scholar] [CrossRef]
- Wickstead, B. The evolutionary biology of dyneins. In Dyneins; King, S.M., Ed.; Academic Press: New York, NY, USA, 2018; pp. 100–138. [Google Scholar] [CrossRef]
- Ali, I.; Yang, W.-C. The functions of kinesin and kinesin-related proteins in eukaryotes. Cell Adhes. Migr. 2020, 14, 139–152. [Google Scholar] [CrossRef] [PubMed]
- Schlattner, U.; Klaus, A.; Ramirez Rios, S.; Guzun, R.; Kay, L.; Tokarska-Schlattner, M. Cellular compartmentation of energy metabolism: Creatine kinase microcompartments and recruitment of B-type creatine kinase to specific subcellular sites. Amino Acids 2016, 48, 1751–1774. [Google Scholar] [CrossRef] [PubMed]
- Krause, S.M.; Jacobus, W.E. Specific enhancement of the cardiac myofibrillar ATPase by bound creatine kinase. J. Biol. Chem. 1992, 267, 2480–2486. [Google Scholar] [CrossRef]
- Kuiper, J.W.P.; Pluk, H.; Oerlemans, F.; van Leeuwen, F.N.; de Lange, F.; Fransen, J.; Wieringa, B. Creatine Kinase–Mediated ATP Supply Fuels Actin-Based Events in Phagocytosis. PLoS Biol. 2008, 6. [Google Scholar] [CrossRef]
- Aziz, S.A.; Kuiper, J.W.P.; van Horssen, R.; Oerlemans, F.; Peters, W.; van Dommelen, M.M.T.; te Lindert, M.M.; ten Hagen, T.L.M.; Janssen, E.; Fransen, J.A.M.; et al. Local ATP Generation by Brain-Type Creatine Kinase (CK-B) Facilitates Cell Motility. PLoS ONE 2009, 4. [Google Scholar] [CrossRef]
- Duran-Trio, L.; Fernandes-Pires, G.; Simicic, D.; Grosse, J.; Roux-Petronelli, C.; Bruce, S.J.; Binz, P.-A.; Sandi, C.; Cudalbu, C.; Braissant, O. A new rat model of creatine transporter deficiency reveals behavioral disorder and altered brain metabolism. Sci. Rep. 2021, 11. [Google Scholar] [CrossRef]
- Hu, W.-J.; Zhou, S.-M.; Yang, J.S.; Meng, F.-G. Computational Simulations to Predict Creatine Kinase-Associated Factors: Protein-Protein Interaction Studies of Brain and Muscle Types of Creatine Kinases. Enzym. Res. 2011, 2011, 1–12. [Google Scholar] [CrossRef][Green Version]
- Wang, Q.; Qian, W.; Xu, X.; Bajpai, A.; Guan, K.; Zhang, Z.; Chen, R.; Flamini, V.; Chen, W. Energy-Mediated Machinery Drives Cellular Mechanical Allostasis. Adv. Mater. 2019, 31. [Google Scholar] [CrossRef]
- Lee, J.H.; Jin, H.E.; Desai, M.S.; Ren, S.; Kim, S.; Lee, S.W. Biomimetic sensor design. Nanoscale 2015, 7, 18379–18391. [Google Scholar] [CrossRef]
- Deldicque, L.; Theisen, D.; Bertrand, L.; Hespel, P.; Hue, L.; Francaux, M. Creatine enhances differentiation of myogenic C2C12 cells by activating both p38 and Akt/PKB pathways. Am. J. Physiol. Cell Physiol. 2007, 293, C1263–C1271. [Google Scholar] [CrossRef]
- Sestili, P.; Barbieri, E.; Stocchi, V. Effects of Creatine in Skeletal Muscle Cells and in Myoblasts Differentiating Under Normal or Oxidatively Stressing Conditions. Mini Rev. Med. Chem. 2015, 16, 4–11. [Google Scholar] [CrossRef] [PubMed]
- Gyoeva, F.K. The role of motor proteins in signal propagation. Biochem. Mosc. 2014, 79, 849–855. [Google Scholar] [CrossRef] [PubMed]
- Solis, M.Y.; Artioli, G.G.; Gualano, B. Potential of Creatine in Glucose Management and Diabetes. Nutrients 2021, 13, 570. [Google Scholar] [CrossRef] [PubMed]
- Somwar, R.; Kim, D.Y.; Sweeney, G.; Huang, C.; Niu, W.; Lador, C.; Ramlal, T.; Klip, A. GLUT4 translocation precedes the stimulation of glucose uptake by insulin in muscle cells: Potential activation of GLUT4 via p38 mitogen-activated protein kinase. Biochem. J. 2001, 359. [Google Scholar] [CrossRef]
- Niu, W.; Huang, C.; Nawaz, Z.; Levy, M.; Somwar, R.; Li, D.; Bilan, P.J.; Klip, A. Maturation of the Regulation of GLUT4 Activity by p38 MAPK during L6 Cell Myogenesis. J. Biol. Chem. 2003, 278, 17953–17962. [Google Scholar] [CrossRef] [PubMed]
- Kleinert, M.; Parker, B.L.; Fritzen, A.M.; Knudsen, J.R.; Jensen, T.E.; Kjøbsted, R.; Sylow, L.; Ruegg, M.; James, D.E.; Richter, E.A. Mammalian target of rapamycin complex 2 regulates muscle glucose uptake during exercise in mice. J. Physiol. 2017, 595, 4845–4855. [Google Scholar] [CrossRef]
- Sterling, P. Allostasis: A model of predictive regulation. Physiol. Behav. 2012, 106, 5–15. [Google Scholar] [CrossRef] [PubMed]
- Rankin, A.; O’Donavon, C.; Madigan, S.M.; O’Sullivan, O.; Cotter, P.D. ‘Microbes in sport’—The potential role of the gut microbiota in athlete health and performance. Br. J. Sports Med. 2017, 51, 698–699. [Google Scholar] [CrossRef]
- Hiergeist, A.; Gläsner, J.; Reischl, U.; Gessner, A. Analyses of Intestinal Microbiota: Culture versus Sequencing: Figure 1. ILAR J. 2015, 56, 228–240. [Google Scholar] [CrossRef] [PubMed]
- Turer, E.; McAlpine, W.; Wang, K.-w.; Lu, T.; Li, X.; Tang, M.; Zhan, X.; Wang, T.; Zhan, X.; Bu, C.-H.; et al. Creatine maintains intestinal homeostasis and protects against colitis. Proc. Natl. Acad. Sci. USA 2017, 114, E1273–E1281. [Google Scholar] [CrossRef] [PubMed]
- Marcobal, A.; Kashyap, P.C.; Nelson, T.A.; Aronov, P.A.; Donia, M.S.; Spormann, A.; Fischbach, M.A.; Sonnenburg, J.L. A metabolomic view of how the human gut microbiota impacts the host metabolome using humanized and gnotobiotic mice. ISME J. 2013, 7, 1933–1943. [Google Scholar] [CrossRef] [PubMed]
- Savidge, T. Predicting Inflammatory Bowel Disease Symptoms Onset: Nitrous Take on Gut Bacteria Is No Laughing Matter. Cell. Mol. Gastroenterol. Hepatol. 2021, 11, 661–662. [Google Scholar] [CrossRef]
- Langille, M.G.I.; Meehan, C.J.; Koenig, J.E.; Dhanani, A.S.; Rose, R.A.; Howlett, S.E.; Beiko, R.G. Microbial shifts in the aging mouse gut. Microbiome 2014, 2. [Google Scholar] [CrossRef]
- O’Sullivan, O.; Cronin, O.; Clarke, S.F.; Murphy, E.F.; Molloy, M.G.; Shanahan, F.; Cotter, P.D. Exercise and the microbiota. Gut Microbes 2015, 6, 131–136. [Google Scholar] [CrossRef]
- Ostojic, S.M. Human gut microbiota as a source of guanidinoacetic acid. Med. Hypotheses 2020, 142. [Google Scholar] [CrossRef]
- Avgerinos, K.I.; Spyrou, N.; Bougioukas, K.I.; Kapogiannis, D. Effects of creatine supplementation on cognitive function of healthy individuals: A systematic review of randomized controlled trials. Exp. Gerontol. 2018, 108, 166–173. [Google Scholar] [CrossRef]
- Roschel, H.; Gualano, B.; Ostojic, S.M.; Rawson, E.S. Creatine Supplementation and Brain Health. Nutrients 2021, 13, 586. [Google Scholar] [CrossRef] [PubMed]
- Aliev, M.; Guzun, R.; Karu-Varikmaa, M.; Kaambre, T.; Wallimann, T.; Saks, V. Molecular System Bioenergics of the Heart: Experimental Studies of Metabolic Compartmentation and Energy Fluxes versus Computer Modeling. Int. J. Mol. Sci. 2011, 12, 9296–9331. [Google Scholar] [CrossRef]
- Mair, J.; Artner-Dworzak, E.; Dienstl, A.; Lechleitner, P.; Morass, B.; Smidt, J.; Wagner, I.; Wettach, C.; Puschendorf, B. Early detection of acute myocardial infarction by measurement of mass concentration of creatine kinase-MB. Am. J. Cardiol. 1991, 68, 1545–1550. [Google Scholar] [CrossRef]
- Hoag, G.N.; Singh, R.; Franks, C.R.; DeCoteau, W.E. Creatine kinase isoenzymes in testicular tissue of normal subjects and in a case of lymphoblastic lymphosarcoma. Clin. Chem. 1980, 26, 1360–1361. [Google Scholar] [CrossRef]
- Nasrallah, F.; Hammami, M.; Omar, S.; Aribia, H.; Sanhaji, H.; Feki, M. Semen Creatine and Creatine Kinase Activity as an Indicator of Sperm Quality. Clin. Lab. 2020, 66. [Google Scholar] [CrossRef] [PubMed]
- Philip, M.; Snow, R.J.; Gatta, P.A.D.; Bellofiore, N.; Ellery, S.J. Creatine metabolism in the uterus: Potential implications for reproductive biology. Amino Acids 2020, 52, 1275–1283. [Google Scholar] [CrossRef] [PubMed]
- Muccini, A.M.; Tran, N.T.; de Guingand, D.L.; Philip, M.; Della Gatta, P.A.; Galinsky, R.; Sherman, L.S.; Kelleher, M.A.; Palmer, K.R.; Berry, M.J.; et al. Creatine Metabolism in Female Reproduction, Pregnancy and Newborn Health. Nutrients 2021, 13, 490. [Google Scholar] [CrossRef]
- Hemmer, W.; Riesinger, I.; Wallimann, T.; Eppenberger, H.M.; Quest, A.F. Brain-type creatine kinase in photoreceptor cell outer segments: Role of a phosphocreatine circuit in outer segment energy metabolism and phototransduction. J. Cell Sci. 1993, 106, 671–683. [Google Scholar] [PubMed]
- Spicer, S.S.; Schulte, B.A. Creatine kinase in epithelium of the inner ear. J. Histochem. Cytochem. 1992, 40, 185–192. [Google Scholar] [CrossRef]
- Acevedo, C.; Blanchard, K.; Bacigalupo, J.; Vergara, C. Possible ATP trafficking by ATP-shuttles in the olfactory cilia and glucose transfer across the olfactory mucosa. FEBS Lett. 2019, 593, 601–610. [Google Scholar] [CrossRef]
- Chen, L.; Roberts, R.; Friedman, D.L. Expression of brain-type creatine kinase and ubiquitous mitochondrial creatine kinase in the fetal rat brain: Evidence for a nuclear energy shuttle. J. Comp. Neurol. 1995, 363, 389–401. [Google Scholar] [CrossRef]
- Schlattner, U.; Möckli, N.; Speer, O.; Werner, S.; Wallimann, T. Creatine Kinase and Creatine Transporter in Normal, Wounded, and Diseased Skin. J. Invest. Dermatol. 2002, 118, 416–423. [Google Scholar] [CrossRef]
- Lenz, H.; Schmidt, M.; Welge, V.; Schlattner, U.; Wallimann, T.; Elsässer, H.-P.; Wittern, K.-P.; Wenck, H.; Stäb, F.; Blatt, T. The Creatine Kinase System in Human Skin: Protective Effects of Creatine Against Oxidative and UV Damage In Vitro and In Vivo. J. Invest. Dermatol. 2005, 124, 443–452. [Google Scholar] [CrossRef]
- Lyons, G.E.; Muhlebach, S.; Moser, A.; Masood, R.; Paterson, B.M.; Buckingham, M.E.; Perriard, J.C. Developmental regulation of creatine kinase gene expression by myogenic factors in embryonic mouse and chick skeletal muscle. Development 1991, 113, 1017–1029. [Google Scholar]
- Yamane, A.; Mayo, M.; Shuler, C.; Crowe, D.; Ohnuki, Y.; Dalrymple, K.; Saeki, Y. Expression of myogenic regulatory factors during the development of mouse tongue striated muscle. Arch. Oral Biol. 2000, 45, 71–78. [Google Scholar] [CrossRef]
- Nguyen, Q.G.; Buskin, J.N.; Himeda, C.L.; Fabre-Suver, C.; Hauschka, S.D. Transgenic and tissue culture analyses of the muscle creatine kinase enhancer Trex control element in skeletal and cardiac muscle indicate differences in gene expression between muscle types. Transgenic Res. 2003, 12, 337–349. [Google Scholar] [CrossRef]
- Gordon, P.V.; Keller, T.C., 3rd. Functional coupling to brush border creatine kinase imparts a selective energetic advantage to contractile ring myosin in intestinal epithelial cells. Cell Motil. Cytoskelet. 1992, 21, 38–44. [Google Scholar] [CrossRef]
- Sistermans, E.A.; de Kok, Y.J.; Peters, W.; Ginsel, L.A.; Jap, P.H.; Wieringa, B. Tissue- and cell-specific distribution of creatine kinase B: A new and highly specific monoclonal antibody for use in immunohistochemistry. Cell Tissue Res. 1995, 280, 435–446. [Google Scholar] [CrossRef]
- Mahajan, V.B.; Pai, K.S.; Lau, A.; Cunningham, D.D. Creatine kinase, an ATP-generating enzyme, is required for thrombin receptor signaling to the cytoskeleton. Proc. Natl. Acad. Sci. USA 2000, 97, 12062–12067. [Google Scholar] [CrossRef]
- Pollard, A.E.; Martins, L.; Muckett, P.J.; Khadayate, S.; Bornot, A.; Clausen, M.; Admyre, T.; Bjursell, M.; Fiadeiro, R.; Wilson, L.; et al. AMPK activation protects against diet-induced obesity through Ucp1-independent thermogenesis in subcutaneous white adipose tissue. Nat. Metab. 2019, 1, 340–349. [Google Scholar] [CrossRef]
- Rahbani, J.F.; Roesler, A.; Hussain, M.F.; Samborska, B.; Dykstra, C.B.; Tsai, L.; Jedrychowski, M.P.; Vergnes, L.; Reue, K.; Spiegelman, B.M.; et al. Creatine kinase B controls futile creatine cycling in thermogenic fat. Nature 2021, 590, 480–485. [Google Scholar] [CrossRef]
- Takahashi, M.; Kishimoto, H.; Shirasaka, Y.; Inoue, K. Functional characterization of monocarboxylate transporter 12 (SLC16A12/MCT12) as a facilitative creatine transporter. Drug Metab. Pharmacokinet. 2020, 35, 281–287. [Google Scholar] [CrossRef]
- Jomura, R.; Tanno, Y.; Akanuma, S.-i.; Kubo, Y.; Tachikawa, M.; Hosoya, K.-i. Monocarboxylate transporter 12 as a guanidinoacetate efflux transporter in renal proximal tubular epithelial cells. Biochim. Biophys. Acta BBA Biomembr. 2020, 1862. [Google Scholar] [CrossRef]
- Verouti, S.N.; Lambert, D.; Mathis, D.; Pathare, G.; Escher, G.; Vogt, B.; Fuster, D.G. Solute carrier SLC16A12 is critical for creatine and guanidinoacetate handling in the kidney. Am. J. Physiol. Ren. Physiol. 2021, 320, F351–F358. [Google Scholar] [CrossRef]
- Tachikawa, M.; Fujinawa, J.; Takahashi, M.; Kasai, Y.; Fukaya, M.; Sakai, K.; Yamazaki, M.; Tomi, M.; Watanabe, M.; Sakimura, K.; et al. Expression and possible role of creatine transporter in the brain and at the blood-cerebrospinal fluid barrier as a transporting protein of guanidinoacetate, an endogenous convulsant. J. Neurochem. 2008, 107, 768–778. [Google Scholar] [CrossRef]
- Colas, C.; Banci, G.; Martini, R.; Ecker, G.F. Studies of structural determinants of substrate binding in the Creatine Transporter (CreaT, SLC6A8) using molecular models. Sci. Rep. 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Saks, V.; Oudman, I.; Clark, J.F.; Brewster, L.M. The Effect of the Creatine Analogue Beta-guanidinopropionic Acid on Energy Metabolism: A Systematic Review. PLoS ONE 2013, 8. [Google Scholar] [CrossRef]
- Snow, R.J.; Murphy, R.M. Creatine and the creatine transporter: A review. Mol. Cell. Biochem. 2001, 224, 169–181. [Google Scholar] [CrossRef]
- Xu, W.; Liu, L.; Gorman, P.A.; Sheer, D.; Emson, P.C. Assignment of the human creatine transporter type 2 (SLC6A10) to chromosome band 16p11.2 by in situ hybridization. Cytogenet. Genome Res. 1997, 76, 19. [Google Scholar] [CrossRef]
- Eichler, E. Duplication of a gene-rich cluster between 16p11.1 and Xq28: A novel pericentromeric-directed mechanism for paralogous genome evolution. Hum. Mol. Genet. 1996, 5, 899–912. [Google Scholar] [CrossRef]
- Iyer, G.S.; Krahe, R.; Goodwin, L.A.; Doggett, N.A.; Siciliano, M.J.; Funanage, V.L.; Proujansky, R. Identification of a Testis-Expressed Creatine Transporter Gene at 16p11.2 and Confirmation of the X-Linked Locus to Xq28. Genomics 1996, 34, 143–146. [Google Scholar] [CrossRef]
- Bayou, N.; M’Rad, R.; Belhaj, A.; Daoud, H.; Zemni, R.; Briault, S.; Helayem, M.B.; Ben Jemaa, L.; Chaabouni, H. The Creatine Transporter Gene Paralogous at 16p11.2 Is Expressed in Human Brain. Comp. Funct. Genom. 2008, 2008, 1–5. [Google Scholar] [CrossRef]
- Kumar, R.A.; KaraMohamed, S.; Sudi, J.; Conrad, D.F.; Brune, C.; Badner, J.A.; Gilliam, T.C.; Nowak, N.J.; Cook, E.H., Jr.; Dobyns, W.B.; et al. Recurrent 16p11.2 microdeletions in autism. Hum. Mol. Genet. 2008, 17, 628–638. [Google Scholar] [CrossRef] [PubMed]
- Mayser, W.; Schloss, P.; Betz, H. Primary structure and functional expression of a choline transporter expressed in the rat nervous system. FEBS Lett. 1992, 305, 31–36. [Google Scholar] [CrossRef]
- Dodd, J.R.; Christie, D.L. Substituted Cysteine Accessibility of the Third Transmembrane Domain of the Creatine Transporter. J. Biol. Chem. 2005, 280, 32649–32654. [Google Scholar] [CrossRef]
- Barnwell, L.F.S.; Chaudhuri, G.; Townsel, J.G. Cloning and sequencing of a cDNA encoding a novel member of the human brain GABA/noradrenaline neurotransmitter transporter family. Gene 1995, 159, 287–288. [Google Scholar] [CrossRef]
- Gonzalez, A.M.; Uhl, G.R. ‘Choline/orphan V8-2-1/creatine transporter’ mRNA is expressed in nervous, renal and gastrointestinal systems. Mol. Brain Res. 1994, 23, 266–270. [Google Scholar] [CrossRef]
- Guerrero-Ontiveros, M.L.; Wallimann, T. Creatine supplementation in health and disease. Effects of chronic creatine ingestion in vivo: Down-regulation of the expression of creatine transporter isoforms in skeletal muscle. Mol. Cell. Biochem. 1998, 184, 427–437. [Google Scholar] [CrossRef]
- Martínez-Muñoz, C.; Rosenberg, E.H.; Jakobs, C.; Salomons, G.S. Identification, characterization and cloning of SLC6A8C, a novel splice variant of the creatine transporter gene. Gene 2008, 418, 53–59. [Google Scholar] [CrossRef]
- Ndika, J.D.T.; Martinez-Munoz, C.; Anand, N.; van Dooren, S.J.M.; Kanhai, W.; Smith, D.E.C.; Jakobs, C.; Salomons, G.S. Post-transcriptional regulation of the creatine transporter gene: Functional relevance of alternative splicing. Biochim. Biophys. Acta BBA Gen. Subj. 2014, 1840, 2070–2079. [Google Scholar] [CrossRef]
- Sitte, H.H.; Farhan, H.; Javitch, J.A. Sodium-dependent neurotransmitter transporters: Oligomerization as a determinant of transporter function and trafficking. Mol. Interv. 2004, 4, 38–47. [Google Scholar] [CrossRef] [PubMed]
- Peral, M.J.; García-Delgado, M.; Calonge, M.L.; Durán, J.M.; Horra, M.C.; Wallimann, T.; Speer, O.; Ilundáin, A.A. Human, rat and chicken small intestinal Na+-Cl−-creatine transporter: Functional, molecular characterization and localization. J. Physiol. 2002, 545, 133–144. [Google Scholar] [CrossRef]
- Odoom, J.; Kemp, G.; Radda, G. The regulation of total creatine content in a myoblast cell line. Mol. Cell. Biochem. 1996, 158. [Google Scholar] [CrossRef]
- Pramod, A.B.; Foster, J.; Carvelli, L.; Henry, L.K. SLC6 transporters: Structure, function, regulation, disease association and therapeutics. Mol. Aspects Med. 2013, 34, 197–219. [Google Scholar] [CrossRef]
- Rudnick, G.; Krämer, R.; Blakely, R.D.; Murphy, D.L.; Verrey, F. The SLC6 transporters: Perspectives on structure, functions, regulation, and models for transporter dysfunction. Pflügers Archiv Eur. J. Physiol. 2013, 466, 25–42. [Google Scholar] [CrossRef]
- Santacruz, L.; Darrabie, M.D.; Mishra, R.; Jacobs, D.O. Removal of Potential Phosphorylation Sites does not Alter Creatine Transporter Response to PKC or Substrate Availability. Cell. Physiol. Biochem. 2015, 37, 353–360. [Google Scholar] [CrossRef]
- Derave, W.; Straumann, N.; Olek, R.A.; Hespel, P. Electrolysis stimulates creatine transport and transporter cell surface expression in incubated mouse skeletal muscle: Potential role of ROS. Am. J. Physiol. Endocrinol. Metab. 2006, 291, E1250–E1257. [Google Scholar] [CrossRef] [PubMed]
- Shojaiefard, M.; Christie, D.L.; Lang, F. Stimulation of the creatine transporter SLC6A8 by the protein kinases SGK1 and SGK3. Biochem. Biophys. Res. Commun. 2005, 334, 742–746. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Cohen, P. Activation of serum- and glucocorticoid-regulated protein kinase by agonists that activate phosphatidylinositide 3-kinase is mediated by 3-phosphoinositide-dependent protein kinase-1 (PDK1) and PDK2. Biochem. J. 1999, 339, 319–328. [Google Scholar] [CrossRef] [PubMed]
- Fezai, M.; Warsi, J.; Lang, F. Regulation of the Na+,Cl- Coupled Creatine Transporter CreaT (SLC6A8) by the Janus Kinase JAK3. Neurosignals 2015, 23, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Almeida, L.S.; Salomons, G.S.; Hogenboom, F.; Jakobs, C.; Schoffelmeer, A.N.M. Exocytotic release of creatine in rat brain. Synapse 2006, 60, 118–123. [Google Scholar] [CrossRef] [PubMed]
- Brault, J.J.; Abraham, K.A.; Terjung, R.L. Muscle creatine uptake and creatine transporter expression in response to creatine supplementation and depletion. J. Appl. Physiol. 2003, 94, 2173–2180. [Google Scholar] [CrossRef]
- Tarnopolsky, M.; Parise, G.; Fu, M.H.; Brose, A.; Parshad, A.; Speer, O.; Wallimann, T. Acute and moderate-term creatine monohydrate supplementation does not affect creatine transporter mRNA or protein content in either young or elderly humans. Mol. Cell. Biochem. 2003, 244, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Jangid, N.; Surana, P.; Salmonos, G.; Jain, V. Creatine transporter deficiency, an underdiagnosed cause of male intellectual disability. BMJ Case Rep. 2020, 13. [Google Scholar] [CrossRef]
- Wang, Q.; Yang, J.; Liu, Y.; Li, X.; Luo, F.; Xie, J. A novel SLC6A8 mutation associated with intellectual disabilities in a Chinese family exhibiting creatine transporter deficiency: Case report. BMC Med. Genet. 2018, 19. [Google Scholar] [CrossRef]
- Dunbar, M.; Jaggumantri, S.; Sargent, M.; Stockler-Ipsiroglu, S.; van Karnebeek, C.D. Treatment of X-linked creatine transporter (SLC6A8) deficiency: Systematic review of the literature and three new cases. Mol. Genet. Metab. 2014, 112, 259–274. [Google Scholar] [CrossRef] [PubMed]
- Sharer, J.D.; Bodamer, O.; Longo, N.; Tortorelli, S.; Wamelink, M.M.; Young, S. Laboratory diagnosis of creatine deficiency syndromes: A technical standard and guideline of the American College of Medical Genetics and Genomics. Genet. Med. 2017, 19, 256–263. [Google Scholar] [CrossRef]
- Kaviani, M.; Izadi, A.; Heshmati, J. Would creatine supplementation augment exercise performance during a low carbohydrate high fat diet? Med. Hypotheses 2021, 146. [Google Scholar] [CrossRef]
- Ferretti, R.; Moura, E.G.; dos Santos, V.C.; Caldeira, E.J.; Conte, M.; Matsumura, C.Y.; Pertille, A.; Mosqueira, M. High-fat diet suppresses the positive effect of creatine supplementation on skeletal muscle function by reducing protein expression of IGF-PI3K-AKT-mTOR pathway. PLoS ONE 2018, 13. [Google Scholar] [CrossRef]
- Mine, M.; Mizuguchi, H.; Takayanagi, T. Kinetic analysis of the transphosphorylation with creatine kinase by pressure-assisted capillary electrophoresis/dynamic frontal analysis. Anal. Bioanal. Chem. 2021, 413, 1453–1460. [Google Scholar] [CrossRef] [PubMed]

| Enzyme Name and Commission Number | Isozyme | Gene Name | Ensembl ID † | Gene Location | UniprotKB | Subunit Structure and PDB Entry | Cellular Location | Tissue Location * |
|---|---|---|---|---|---|---|---|---|
| Creatine kinase EC: 2.7.3.2 | M-type | CKM | ENSG00000104879 | Chromosome 19: 45,306,414–45,322,977 Reverse strand. | P06732 | Dimer of identical or non-identical chains (1I0E) | Cytosol | Skeletal muscle & heart |
| B-type | CKB | ENSG00000166165 | Chromosome 14: 103,519,659–103,523,111 Reverse strand. | P12277 | Dimer of identical or non-identical chains (3B6R) | Cytosol, dendrite, extracellular exosome, extracellular space, mitochondrion, myelin sheath, neuronal cell body and nucleus | Mainly brain, but also in testes, retina, bone, among several others | |
| U-Type | CKMT1A | ENSG00000223572 | Chromosome 15: 43,692,886–43,699,222 Forward strand. | P12532 | Octamer of four CKMT dimers (1QK1) | Mitochondrial inner membrane and Extracellular exosome | Brain, heart, brown adipose tissue, among several others | |
| S-type | CKMT2 | ENSG00000131730 | Chromosome 5: 81,233,285–81,266,397 Forward strand. | P17540 | Octamer of four CKMT dimers (4Z9M) | Mitochondrial Inner Membrane | Mainly skeletal muscle |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bonilla, D.A.; Kreider, R.B.; Stout, J.R.; Forero, D.A.; Kerksick, C.M.; Roberts, M.D.; Rawson, E.S. Metabolic Basis of Creatine in Health and Disease: A Bioinformatics-Assisted Review. Nutrients 2021, 13, 1238. https://doi.org/10.3390/nu13041238
Bonilla DA, Kreider RB, Stout JR, Forero DA, Kerksick CM, Roberts MD, Rawson ES. Metabolic Basis of Creatine in Health and Disease: A Bioinformatics-Assisted Review. Nutrients. 2021; 13(4):1238. https://doi.org/10.3390/nu13041238
Chicago/Turabian StyleBonilla, Diego A., Richard B. Kreider, Jeffrey R. Stout, Diego A. Forero, Chad M. Kerksick, Michael D. Roberts, and Eric S. Rawson. 2021. "Metabolic Basis of Creatine in Health and Disease: A Bioinformatics-Assisted Review" Nutrients 13, no. 4: 1238. https://doi.org/10.3390/nu13041238
APA StyleBonilla, D. A., Kreider, R. B., Stout, J. R., Forero, D. A., Kerksick, C. M., Roberts, M. D., & Rawson, E. S. (2021). Metabolic Basis of Creatine in Health and Disease: A Bioinformatics-Assisted Review. Nutrients, 13(4), 1238. https://doi.org/10.3390/nu13041238