Common Genetic Variations Involved in the Inter-Individual Variability of Circulating Cholesterol Concentrations in Response to Diets: A Narrative Review of Recent Evidence
Abstract
1. Introduction
2. Search Process and Criteria
3. Studies Investigating the Effect of SNPs Taken Individually
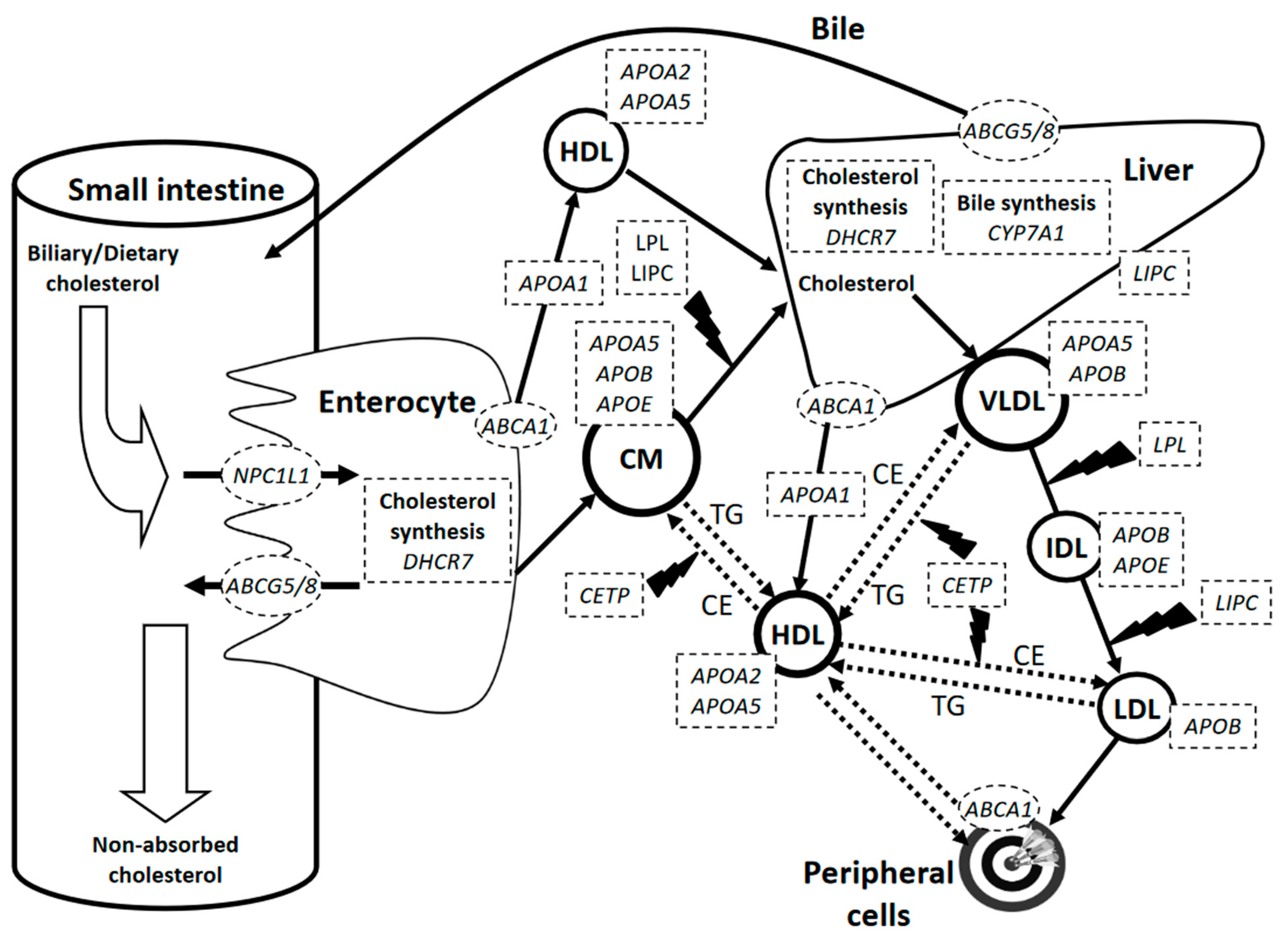
3.1. SNPs in Genes Encoding Transporters Involved in Cholesterol Absorption
3.2. SNPs in Genes Encoding Apolipoproteins
3.3. Genes Encoding Additional Proteins Involved in the Cholesterol Metabolic Pathway
4. Combinatory Patterns of SNPs Influencing Changes in Blood Cholesterol Concentrations in Response to Dietary Interventions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Floris, M.; Cano, A.; Porru, L.; Addis, R.; Cambedda, A.; Idda, M.L.; Steri, M.; Ventura, C.; Maioli, M. Direct-to-Consumer Nutrigenetics Testing: An Overview. Nutrients 2020, 12, 566. [Google Scholar] [CrossRef]
- Guasch-Ferre, M.; Dashti, H.S.; Merino, J. Nutritional Genomics and Direct-to-Consumer Genetic Testing: An Overview. Adv. Nutr. 2018, 9, 128–135. [Google Scholar] [CrossRef]
- Price, H.C.; Nicholls, A. Primary prevention of CVD: Diet. BMJ Clin. Evid. 2014, 2014, 0219. [Google Scholar]
- Tognon, G.; Lissner, L.; Saebye, D.; Walker, K.Z.; Heitmann, B.L. The Mediterranean diet in relation to mortality and CVD: A Danish cohort study. Br. J. Nutr. 2014, 111, 151–159. [Google Scholar] [CrossRef]
- Adams, J.N.; Raffield, L.M.; Freedman, B.I.; Langefeld, C.D.; Ng, M.C.; Carr, J.J.; Cox, A.J.; Bowden, D.W. Analysis of common and coding variants with cardiovascular disease in the Diabetes Heart Study. Cardiovasc. Diabetol. 2014, 13, 77. [Google Scholar] [CrossRef]
- Teslovich, T.M.; Musunuru, K.; Smith, A.V.; Edmondson, A.C.; Stylianou, I.M.; Koseki, M.; Pirruccello, J.P.; Ripatti, S.; Chasman, D.I.; Willer, C.J.; et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 2010, 466, 707–713. [Google Scholar] [CrossRef] [PubMed]
- Willer, C.J.; Sanna, S.; Jackson, A.U.; Scuteri, A.; Bonnycastle, L.L.; Clarke, R.; Heath, S.C.; Timpson, N.J.; Najjar, S.S.; Stringham, H.M.; et al. Newly identified loci that influence lipid concentrations and risk of coronary artery disease. Nat. Genet. 2008, 40, 161–169. [Google Scholar] [CrossRef]
- Abdullah, M.M.; Jones, P.J.; Eck, P.K. Nutrigenetics of cholesterol metabolism: Observational and dietary intervention studies in the postgenomic era. Nutr. Rev. 2015, 73, 523–543. [Google Scholar] [CrossRef] [PubMed]
- Vazquez-Vidal, I.; Desmarchelier, C.; Jones, P.J.H. Nutrigenetics of Blood Cholesterol Concentrations: Towards Personalized Nutrition. Curr. Cardiol. Rep. 2019, 21, 38. [Google Scholar] [CrossRef]
- Hill, W.G.; Goddard, M.E.; Visscher, P.M. Data and theory point to mainly additive genetic variance for complex traits. PLoS Genet. 2008, 4, e1000008. [Google Scholar] [CrossRef] [PubMed]
- Maki-Tanila, A.; Hill, W.G. Influence of gene interaction on complex trait variation with multilocus models. Genetics 2014, 198, 355–367. [Google Scholar] [CrossRef] [PubMed]
- Bruford, E.A.; Braschi, B.; Denny, P.; Jones, T.E.M.; Seal, R.L.; Tweedie, S. Guidelines for human gene nomenclature. Nat. Genet. 2020, 52, 754–758. [Google Scholar] [CrossRef] [PubMed]
- Jacobo-Albavera, L.; Posadas-Romero, C.; Vargas-Alarcon, G.; Romero-Hidalgo, S.; Posadas-Sanchez, R.; Gonzalez-Salazar Mdel, C.; Carnevale, A.; Canizales-Quinteros, S.; Medina-Urrutia, A.; Antunez-Arguelles, E.; et al. Dietary fat and carbohydrate modulate the effect of the ATP-binding cassette A1 (ABCA1) R230C variant on metabolic risk parameters in premenopausal women from the Genetics of Atherosclerotic Disease (GEA) Study. Nutr. Metab. 2015, 12, 45. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Lin, J.; Zhu, X.; Li, Y.; Fan, M.; Zhang, R.; Fang, D. Effects of R219K polymorphism of ATP-binding cassette transporter 1 gene on serum lipids ratios induced by a high-carbohydrate and low-fat diet in healthy youth. Biol. Res. 2014, 47, 4. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Abdullah, M.M.; Cyr, A.; Lepine, M.C.; Eck, P.K.; Couture, P.; Lamarche, B.; Jones, P.J. Common Variants in Cholesterol Synthesis- and Transport-Related Genes Associate with Circulating Cholesterol Responses to Intakes of Conventional Dairy Products in Healthy Individuals. J. Nutr. 2016, 146, 1008–1016. [Google Scholar] [CrossRef] [PubMed]
- Granado-Lorencio, F.; de Las Heras, L.; Millan, C.S.; Garcia-Lopez, F.J.; Blanco-Navarro, I.; Perez-Sacristan, B.; Dominguez, G. β-Cryptoxanthin modulates the response to phytosterols in post-menopausal women carrying NPC1L1 L272L and ABCG8 A632 V polymorphisms: An exploratory study. Genes Nutr. 2014, 9, 428. [Google Scholar] [CrossRef] [PubMed]
- de Luis, D.A.; Izaola, O.; Primo, D.; Aller, R. Role of rs670 variant of APOA1 gene on lipid profile, insulin resistance and adipokine levels in obese subjects after weight loss with a dietary intervention. Diabetes Res. Clin. Pract. 2018, 142, 139–145. [Google Scholar] [CrossRef]
- de Luis, D.; Izaola, O.; Primo, D.; Aller, R. Role of rs670 variant of APOA1 gene on metabolic response after a high fat vs. a low fat hypocaloric diets in obese human subjects. J. Diabetes Complicat. 2019, 33, 249–254. [Google Scholar] [CrossRef]
- Noorshahi, N.; Sotoudeh, G.; Djalali, M.; Eshraghian, M.R.; Keramatipour, M.; Basiri, M.G.; Doostan, F.; Koohdani, F. APOA II genotypes frequency and their interaction with saturated fatty acids consumption on lipid profile of patients with type 2 diabetes. Clin. Nutr. 2016, 35, 907–911. [Google Scholar] [CrossRef]
- Dominguez-Reyes, T.; Astudillo-Lopez, C.C.; Salgado-Goytia, L.; Munoz-Valle, J.F.; Salgado-Bernabe, A.B.; Guzman-Guzman, I.P.; Castro-Alarcon, N.; Moreno-Godinez, M.E.; Parra-Rojas, I. Interaction of dietary fat intake with APOA2, APOA5 and LEPR polymorphisms and its relationship with obesity and dyslipidemia in young subjects. Lipids Health Dis. 2015, 14, 106. [Google Scholar] [CrossRef]
- Lim, H.H.; Choi, M.; Kim, J.Y.; Lee, J.H.; Kim, O.Y. Increased risk of obesity related to total energy intake with the APOA5-1131T > C polymorphism in Korean premenopausal women. Nutr. Res. 2014, 34, 827–836. [Google Scholar] [CrossRef]
- Doo, M.; Won, S.; Kim, Y. Association between the APOB rs1469513 polymorphism and obesity is modified by dietary fat intake in Koreans. Nutrition 2015, 31, 653–658. [Google Scholar] [CrossRef]
- Shatwan, I.M.; Weech, M.; Jackson, K.G.; Lovegrove, J.A.; Vimaleswaran, K.S. Apolipoprotein E gene polymorphism modifies fasting total cholesterol concentrations in response to replacement of dietary saturated with monounsaturated fatty acids in adults at moderate cardiovascular disease risk. Lipids Health Dis. 2017, 16, 222. [Google Scholar] [CrossRef]
- Weber, K.S.; Knebel, B.; Strassburger, K.; Kotzka, J.; Stehle, P.; Szendroedi, J.; Mussig, K.; Buyken, A.E.; Roden, M. Associations between explorative dietary patterns and serum lipid levels and their interactions with ApoA5 and ApoE haplotype in patients with recently diagnosed type 2 diabetes. Cardiovasc. Diabetol. 2016, 15, 138. [Google Scholar] [CrossRef]
- MacKay, D.S.; Eck, P.K. CYP7A1-rs3808607 and APOE isoform associate with LDL cholesterol lowering after plant sterol consumption in a randomized clinical trial. Am. J. Clin. Nutr. 2015, 102, 951–957. [Google Scholar] [CrossRef]
- Griffin, B.A.; Walker, C.G.; Jebb, S.A.; Moore, C.; Frost, G.S.; Goff, L.; Sanders, T.A.B.; Lewis, F.; Griffin, M.; Gitau, R.; et al. APOE4 Genotype Exerts Greater Benefit in Lowering Plasma Cholesterol and Apolipoprotein B than Wild Type (E3/E3), after Replacement of Dietary Saturated Fats with Low Glycaemic Index Carbohydrates. Nutrients 2018, 10, 1524. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Rios, A.; Alcala-Diaz, J.F.; Gomez-Delgado, F.; Delgado-Lista, J.; Marin, C.; Leon-Acuna, A.; Camargo, A.; Rodriguez-Cantalejo, F.; Blanco-Rojo, R.; Quintana-Navarro, G.; et al. Beneficial effect of CETP gene polymorphism in combination with a Mediterranean diet influencing lipid metabolism in metabolic syndrome patients: CORDIOPREV study. Clin. Nutr. 2018, 37, 229–234. [Google Scholar] [CrossRef] [PubMed]
- Hosseini-Esfahani, F.; Esfandiar, Z.; Mirmiran, P.; Daneshpour, M.S.; Ghanbarian, A.; Azizi, F. The interaction of cholesteryl ester transfer protein gene variations and diet on changes in serum lipid profiles. Eur. J. Clin. Nutr. 2019, 73, 1291–1298. [Google Scholar] [CrossRef]
- Wang, Y.; Harding, S.V.; Eck, P.; Thandapilly, S.J.; Gamel, T.H.; Abdel-Aal el, S.M.; Crow, G.H.; Tosh, S.M.; Jones, P.J.; Ames, N.P. High-Molecular-Weight β-Glucan Decreases Serum Cholesterol Differentially Based on the CYP7A1 rs3808607 Polymorphism in Mildly Hypercholesterolemic Adults. J. Nutr. 2016, 146, 720–727. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.C.; Lin, J.; Wang, Q.; Liu, H.; Qiu, L.; Fang, D.Z. Associations of lipoprotein lipase gene rs326 with changes of lipid profiles after a high-carbohydrate and low-fat diet in healthy Chinese Han youth. Int. J. Environ. Res. Public Health 2014, 11, 4544–4554. [Google Scholar] [CrossRef]
- Ayyappa, K.A.; Shatwan, I.; Bodhini, D.; Bramwell, L.R.; Ramya, K.; Sudha, V.; Anjana, R.M.; Lovegrove, J.A.; Mohan, V.; Radha, V.; et al. High fat diet modifies the association of lipoprotein lipase gene polymorphism with high density lipoprotein cholesterol in an Asian Indian population. Nutr. Metab. 2017, 14, 8. [Google Scholar] [CrossRef]
- Smith, C.E.; Van Rompay, M.I.; Mattei, J.; Garcia, J.F.; Garcia-Bailo, B.; Lichtenstein, A.H.; Tucker, K.L.; Ordovas, J.M. Dietary fat modulation of hepatic lipase variant -514 C/T for lipids: A crossover randomized dietary intervention in Caribbean Hispanics. Physiol. Genom. 2017. [Google Scholar] [CrossRef]
- Xu, M.; Ng, S.S.; Bray, G.A.; Ryan, D.H.; Sacks, F.M.; Ning, G.; Qi, L. Dietary Fat Intake Modifies the Effect of a Common Variant in the LIPC Gene on Changes in Serum Lipid Concentrations during a Long-Term Weight-Loss Intervention Trial. J. Nutr. 2015, 145, 1289–1294. [Google Scholar] [CrossRef]
- Davis, H.R., Jr.; Altmann, S.W. Niemann-Pick C1 Like 1 (NPC1L1) an intestinal sterol transporter. Biochim. Biophys. Acta 2009, 1791, 679–683. [Google Scholar] [CrossRef]
- Davis, H.R., Jr.; Basso, F.; Hoos, L.M.; Tetzloff, G.; Lally, S.M.; Altmann, S.W. Cholesterol homeostasis by the intestine: Lessons from Niemann-Pick C1 Like 1 (NPC1L1). Atheroscler. Suppl. 2008, 9, 77–81. [Google Scholar] [CrossRef]
- Temel, R.E.; Tang, W.; Ma, Y.; Rudel, L.L.; Willingham, M.C.; Ioannou, Y.A.; Davies, J.P.; Nilsson, L.M.; Yu, L. Hepatic Niemann-Pick C1-like 1 regulates biliary cholesterol concentration and is a target of ezetimibe. J. Clin. Investig. 2007, 117, 1968–1978. [Google Scholar] [CrossRef] [PubMed]
- De Smet, E.; Mensink, R.P.; Plat, J. Effects of plant sterols and stanols on intestinal cholesterol metabolism: Suggested mechanisms from past to present. Mol. Nutr. Food Res. 2012, 56, 1058–1072. [Google Scholar] [CrossRef]
- Lee, J.Y.; Parks, J.S. ATP-binding cassette transporter AI and its role in HDL formation. Curr. Opin. Lipidol. 2005, 16, 19–25. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.; Gillard, B.K.; Gotto, A.M., Jr.; Rosales, C.; Pownall, H.J. ABCA1-Derived Nascent High-Density Lipoprotein-Apolipoprotein AI and Lipids Metabolically Segregate. Arterioscler. Thromb. Vasc. Biol. 2017, 37, 2260–2270. [Google Scholar] [CrossRef] [PubMed]
- Acuna-Alonzo, V.; Flores-Dorantes, T.; Kruit, J.K.; Villarreal-Molina, T.; Arellano-Campos, O.; Hunemeier, T.; Moreno-Estrada, A.; Ortiz-Lopez, M.G.; Villamil-Ramirez, H.; Leon-Mimila, P.; et al. A functional ABCA1 gene variant is associated with low HDL-cholesterol levels and shows evidence of positive selection in Native Americans. Hum. Mol. Genet. 2010, 19, 2877–2885. [Google Scholar] [CrossRef]
- Patel, S.B.; Graf, G.A.; Temel, R.E. ABCG5 and ABCG8: More than a defense against xenosterols. J. Lipid Res. 2018, 59, 1103–1113. [Google Scholar] [CrossRef]
- Ajagbe, B.O.; Othman, R.A.; Myrie, S.B. Plant Sterols, Stanols, and Sitosterolemia. J. AOAC Int. 2015, 98, 716–723. [Google Scholar] [CrossRef]
- Myrie, S.B.; Mymin, D.; Triggs-Raine, B.; Jones, P.J. Serum lipids, plant sterols, and cholesterol kinetic responses to plant sterol supplementation in phytosterolemia heterozygotes and control individuals. Am. J. Clin. Nutr. 2012, 95, 837–844. [Google Scholar] [CrossRef]
- Plat, J.; Bragt, M.C.; Mensink, R.P. Common sequence variations in ABCG8 are related to plant sterol metabolism in healthy volunteers. J. Lipid Res. 2005, 46, 68–75. [Google Scholar] [CrossRef] [PubMed]
- Viturro, E.; de Oya, M.; Lasuncion, M.A.; Gorgojo, L.; Moreno, J.M.; Benavente, M.; Cano, B.; Garces, C. Cholesterol and saturated fat intake determine the effect of polymorphisms at ABCG5/ABCG8 genes on lipid levels in children. Genet. Med. Off. J. Am. Coll. Med. Genet. 2006, 8, 594–599. [Google Scholar] [CrossRef]
- Bandarian, F.; Hedayati, M.; Daneshpour, M.S.; Naseri, M.; Azizi, F. Genetic polymorphisms in the APOA1 gene and their relationship with serum HDL cholesterol levels. Lipids 2013, 48, 1207–1216. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Smith, J.D. ABCA1 and nascent HDL biogenesis. BioFactors 2014, 40, 547–554. [Google Scholar] [CrossRef] [PubMed]
- Bandarian, F.; Daneshpour, M.S.; Hedayati, M.; Naseri, M.; Azizi, F. Identification of Sequence Variation in the Apolipoprotein A2 Gene and Their Relationship with Serum High-Density Lipoprotein Cholesterol Levels. Iran. Biomed. J. 2016, 20, 84–90. [Google Scholar]
- Beckstead, J.A.; Oda, M.N.; Martin, D.D.; Forte, T.M.; Bielicki, J.K.; Berger, T.; Luty, R.; Kay, C.M.; Ryan, R.O. Structure-function studies of human apolipoprotein A-V: A regulator of plasma lipid homeostasis. Biochemistry 2003, 42, 9416–9423. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lin, Y.C.; Nunez, V.; Johns, R.; Shiao, S.P. APOA5 Gene Polymorphisms and Cardiovascular Diseases: Metaprediction in Global Populations. Nurs. Res. 2017, 66, 164–174. [Google Scholar] [CrossRef]
- Zhao, T.; Zhao, J. Association of the apolipoprotein A5 gene −1131 T > C polymorphism with fasting blood lipids: A meta-analysis in 37,859 subjects. BMC Med. Genet. 2010, 11, 120. [Google Scholar] [CrossRef]
- Agoston-Coldea, L.; Zdrenghea, D.; Pop, D.; Craciun, A.; Rusu, M.L.; Mocan, T. Apolipoproteins A-I and B-markers in coronary risk evaluation. Rom. J. Intern. Med. Rev. Roum. Med. Interne 2007, 45, 251–258. [Google Scholar]
- Bennet, A.M.; Di Angelantonio, E.; Ye, Z.; Wensley, F.; Dahlin, A.; Ahlbom, A.; Keavney, B.; Collins, R.; Wiman, B.; de Faire, U.; et al. Association of apolipoprotein E genotypes with lipid levels and coronary risk. JAMA 2007, 298, 1300–1311. [Google Scholar] [CrossRef] [PubMed]
- Boekholdt, S.M.; Kuivenhoven, J.A.; Hovingh, G.K.; Jukema, J.W.; Kastelein, J.J.; van Tol, A. CETP gene variation: Relation to lipid parameters and cardiovascular risk. Curr. Opin. Lipidol. 2004, 15, 393–398. [Google Scholar] [CrossRef]
- Justesen, J.M.; Allin, K.H.; Sandholt, C.H.; Borglykke, A.; Krarup, N.T.; Grarup, N.; Linneberg, A.; Jørgensen, T.; Hansen, T.; Pedersen, O. Interactions of Lipid Genetic Risk Scores with Estimates of Metabolic Health in a Danish Population. Circ. Cardiovasc. Genet. 2015, 8, 465–472. [Google Scholar] [CrossRef][Green Version]
- San-Cristobal, R.; Navas-Carretero, S.; Livingstone, K.M.; Celis-Morales, C.; Macready, A.L.; Fallaize, R.; O’Donovan, C.B.; Lambrinou, C.P.; Moschonis, G.; Marsaux, C.F.M.; et al. Mediterranean Diet Adherence and Genetic Background Roles within a Web-Based Nutritional Intervention: The Food4Me Study. Nutrients 2017, 9, 1107. [Google Scholar] [CrossRef] [PubMed]
- Abdullah, M.M.H.; Eck, P.K.; Couture, P.; Lamarche, B.; Jones, P.J.H. The combination of single nucleotide polymorphisms rs6720173 (ABCG5), rs3808607 (CYP7A1), and rs760241 (DHCR7) is associated with differing serum cholesterol responses to dairy consumption. Appl. Physiol. Nutr. Metab. Physiol. Appl. Nutr. Metab. 2018, 43, 1090–1093. [Google Scholar] [CrossRef]
- Guevara-Cruz, M.; Medina-Vera, I.; Flores-López, A.; Aguilar-López, M.; Smith, C.E.; Parnell, L.D.; Lee, Y.C.; Lai, C.Q.; Tovar, A.R.; Ordovás, J.M.; et al. Development of a Genetic Score to Predict an Increase in HDL Cholesterol Concentration After a Dietary Intervention in Adults with Metabolic Syndrome. J. Nutr. 2019, 149, 1116–1121. [Google Scholar] [CrossRef] [PubMed]
- Mihaylova, B.; Emberson, J.; Blackwell, L.; Keech, A.; Simes, J.; Barnes, E.H.; Voysey, M.; Gray, A.; Collins, R.; Baigent, C. The effects of lowering LDL cholesterol with statin therapy in people at low risk of vascular disease: Meta-analysis of individual data from 27 randomised trials. Lancet 2012, 380, 581–590. [Google Scholar] [CrossRef]
- Robinson, J.G.; Smith, B.; Maheshwari, N.; Schrott, H. Pleiotropic effects of statins: Benefit beyond cholesterol reduction? A meta-regression analysis. J. Am. Coll. Cardiol. 2005, 46, 1855–1862. [Google Scholar] [CrossRef] [PubMed]
- Shameer, K.; Johnson, K.W.; Glicksberg, B.S.; Dudley, J.T.; Sengupta, P.P. Machine learning in cardiovascular medicine: Are we there yet? Heart 2018, 104, 1156–1164. [Google Scholar] [CrossRef] [PubMed]
| Author | Gene | SNP | Study Design | Diet | Population and Ethnicity | Major Cholesterol Outcomes |
|---|---|---|---|---|---|---|
| Jacobo-Albavera et al. (2015) [13] | ABCA1 | rs9282541 | Cross-sectional | Dietary CHO and fat intakes | 1598 premenopausal females, Mexicans | HDL-C concentrations negatively correlated with CHO intake and positively correlated with fat intake in T allele carriers but not CC homozygotes |
| Liu et al. (2014) [14] | ABCA1 | rs2230806 | Dietary intervention | 7-d of washout diet followed by 6-d of high-CHO/low-fat diet | 56 healthy adults, Chinese | Lower LDL-C/HDL-C concentration ratios in A allele male carriers and GG female homozygotes after vs. before high-CHO/low-fat diet |
| Abdullah et al. (2016) [15] | ABCG5 | rs6720173 | Crossover | Dairy vs. dairy-free diets | 101 healthy adults, Canadians | Higher TC and LDL-C concentrations in GG homozygotes vs. C allele carriers after 3 servings/d of dairy vs. dairy-free diets |
| Granado-Lorencio et al. (2014) [16] | ABCG8 | rs6544718 | Crossover | 750 μg β-cryptoxanthin and 1.5 g/d PS, single and combined | 19 postmenopausal females, Spanish | Lower TC concentrations in CC homozygotes vs. T allele carriers after β-cryptoxanthin + PS combined intake |
| de Luis et al. (2018) [17] | APOA1 | rs670 | Dietary intervention | Hypocaloric diet of one arm | 82 obese adults, Spanish | Lower HDL-C concentrations in GG homozygotes vs. A allele carriers at baseline and after hypocaloric diet |
| de Luis et al. (2019) [18] | APOA1 | rs670 | Dietary intervention | Hypocaloric high-fat vs. low-fat diets | 282 obese adults, Spanish | Higher HDL-C concentrations in A allele carriers vs. GG homozygotes at baseline and after both high-fat and low-fat diets |
| Noorshahi et al. (2016) [19] | APOA2 | rs5082 | Cross-sectional | Dietary SFA intake (>28.5 g/d), assessed by FFQ | 697 type 2 diabetic adults, Iranians | Higher LDL-C/HDL-C concentration ratio in CC homozygotes vs. T allele carriers with higher SFA intake |
| Dominguez-Reyes et al. (2015) [20] | APOA5 | rs662799 | Cross-sectional | Dietary fat intake, assessed by FFQ | 200 young normal-weight and obese adults, Mexicans | Lower HDL-C concentrations and higher PUFA intake in C allele carriers vs. TT homozygotes |
| Lim et al. (2014) [21] | APOA5 | rs662799 | Cross-sectional | Dietary intake, assessed by 24-h recall and FFQ | 1128 premenopausal females, Koreans | Lower HDL-C concentrations in CC homozygotes with higher total energy intake (≥2001 kcal/d) |
| Doo et al. (2015) [22] | APOB | rs1469513 | Cross-sectional (‘KoGES’ Study) | Total energy and macronutrient intake, assessed by FFQ | 6470 adults, Koreans | Higher TC and LDL-C concentrations in G allele carriers with higher energy or fat intake, and in AA homozygotes with higher CHO intake |
| Shatwan et al. (2017) [23] | APOE | rs1064725 | 16-wk parallel dietary intervention (‘DIVAS’ Study) | Isoenergetic diets rich in SFA, MUFA, or n-6 PUFA | 120 adults with moderate cardiovascular risk, Caucasians | Lower TC concentrations only in TT homozygotes after MUFA-rich diet vs. SFA-rich or n-6 PUFA-rich diets |
| Weber et al. (2016) [24] | APOE | rs429258 rs7412 | Cross-sectional (German Diabetes Study) | Dietary fat intake, assessed by FFQ | 348 diabetics, Germans | Lower LDL-C concentrations in APOE ε2 carriers with lower vs. higher intake frequencies of butter, cream cake, French fries, or alcoholic beverages |
| Mackay et al. (2015) [25] | APOE | rs429258 rs7412 | Crossover | 2 g/d of PS | 63 mildly hypercholesterolemic adults, Canadians | Greater reduction in LDL-C concentrations after PS intake in APOE ε4 vs. APOE ε3 carriers |
| Griffin et al. (2018) [26] | APOE | rs429258 rs7412 | 24-wk five-arm parallel dietary intervention (‘RISCK’ Study) | Replacing SFA with either MUFA or carbohydrate of high or low GI | 389 adults at increased risk of developing MetS, white Caucasians | Greater decreases in TC concentrations in carriers of E4 vs. E3/E3 when SFA was replaced with low GI carbohydrate on a lower fat diet, and an increase in TC concentrations when SFA was replaced with MUFA and high GI carbohydrates |
| Garcia-Rios et al. (2016) [27] | CETP | rs3764261 | 1-y dietary intervention (‘CORDIOPREV’ Study) | Mediterranean diet (35% fat, 22% MUFA) vs. low-fat diet (28% fat, 12% MUFA) | 424 MetS subjects, Spanish | Higher HDL-C concentrations after Mediterranean diet in T allele carriers vs. GG homozygotes |
| Hosseini-Esfahani et al (2019) [28] | CETP | rs3764261 | Population-based prospective design (Tehran Lipid and Glucose Study) | Usual dietary intake, assessed by FFQ | 4700 adults, Iranians | Lower TC concentrations and higher fish intakes in T allele carriers vs. GG homozygotes after 3.6 years of follow-up |
| Abdullah et al. (2016) [15] | CYP7A1 | rs3808607 | Crossover | Dairy vs. dairy-free diets | 101 healthy adults, Canadians | Higher TC concentrations in G allele carriers vs. TT homozygotes after 3 servings/d of dairy vs. dairy-free diets |
| Mackay et al. (2015) [25] | CYP7A1 | rs3808607 | Crossover | 2 g/d of PS | 63 mildly hypercholesterolemic adults, Canadians | Greater reduction in LDL-C concentrations after PS intake in GG vs. TT homozygotes |
| Wang et al. (2016) [29] | CYP7A1 | rs3808607 | Crossover | Barley β-glucan vs. control diet | 30 mildly hypercholesterolemic adults, Canadians | Lower TC concentrations after 3 g/d of high molecular weight barley β-glucan in G allele carriers vs. TT homozygotes |
| Abdullah et al. (2016) [15] | DHCR7 | rs760241 | Crossover | Dairy vs. dairy-free diets | 101 healthy adults, Canadians | Higher LDL-C concentrations in A allele carriers vs. GG homozygotes after 3 servings/d of dairy vs. dairy-free diets |
| Zhu et al. (2014) [30] | LPL | rs326 | Dietary intervention | 7-d of washout diet followed by 6-d of high-CHO (70% energy)/low-fat (~14% energy) diet | 56 healthy Chinese Han youth | Higher HDL-C concentrations in G allele male carriers after vs. before high-CHO/low-fat diet |
| Ayyappa et al. (2017) [31] | LPL | rs1121923 | Cross-sectional (Chennai Urban Rural Epidemiological Study) | Dietary intakes, assessed by FFQ | 788 type 2 diabetes cases and 1057 controls, Asian Indians | Higher HDL-C concentrations in T allele carriers with high fat diet vs. CC homozygotes |
| Smith et al. (2017) [32] | LIPC | rs1800588 | Crossover | Comparing a high-fat Western diet and a low-fat traditional Hispanic diet | 42 adults, Caribbean Hispanic descent | Higher HDL-C concentrations in CC homozygotes after high-fat Western diet (39% energy) vs. low-fat traditional Hispanic diet (20% energy) |
| Xu et al. (2015) [33] | LIPC | rs2070895 | 2-y randomized weight-loss dietary intervention (‘POUNDS LOST’ Study) | Dietary intakes | 743 overweight or obese adults, multiethnic groups | Higher TC concentrations after high fat intake, and higher HDL-C concentrations after low fat intake, in A allele carriers vs. GG homozygotes |
| Granado-Lorencio et al. (2014) [16] | NPC1L1 | rs2072183 | Crossover | 750 μg β-cryptoxanthin and 1.5 g/d PS, single and combined | 19 postmenopausal females, Spanish | Higher TC concentrations in CC homozygotes vs. G allele carriers after PS intake only |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abdullah, M.M.H.; Vazquez-Vidal, I.; Baer, D.J.; House, J.D.; Jones, P.J.H.; Desmarchelier, C. Common Genetic Variations Involved in the Inter-Individual Variability of Circulating Cholesterol Concentrations in Response to Diets: A Narrative Review of Recent Evidence. Nutrients 2021, 13, 695. https://doi.org/10.3390/nu13020695
Abdullah MMH, Vazquez-Vidal I, Baer DJ, House JD, Jones PJH, Desmarchelier C. Common Genetic Variations Involved in the Inter-Individual Variability of Circulating Cholesterol Concentrations in Response to Diets: A Narrative Review of Recent Evidence. Nutrients. 2021; 13(2):695. https://doi.org/10.3390/nu13020695
Chicago/Turabian StyleAbdullah, Mohammad M. H., Itzel Vazquez-Vidal, David J. Baer, James D. House, Peter J. H. Jones, and Charles Desmarchelier. 2021. "Common Genetic Variations Involved in the Inter-Individual Variability of Circulating Cholesterol Concentrations in Response to Diets: A Narrative Review of Recent Evidence" Nutrients 13, no. 2: 695. https://doi.org/10.3390/nu13020695
APA StyleAbdullah, M. M. H., Vazquez-Vidal, I., Baer, D. J., House, J. D., Jones, P. J. H., & Desmarchelier, C. (2021). Common Genetic Variations Involved in the Inter-Individual Variability of Circulating Cholesterol Concentrations in Response to Diets: A Narrative Review of Recent Evidence. Nutrients, 13(2), 695. https://doi.org/10.3390/nu13020695






