Gut Microbiota Bacterial Species Associated with Mediterranean Diet-Related Food Groups in a Northern Spanish Population
Abstract
1. Introduction
2. Material and Methods
2.1. Participants
2.2. Anthropometric and Biochemical Measurements
2.3. Dietary Estimation
2.4. Faecal Sample Collection and DNA Extraction
2.5. Metagenomic Data: Library Preparation
2.6. Metagenomics Data: Analysis and Processing
2.7. Richness and Evenness
2.8. Statistical Analysis
2.9. Prediction of Functional Potential of Gut Microbiota
3. Results
3.1. Participant Characteristics
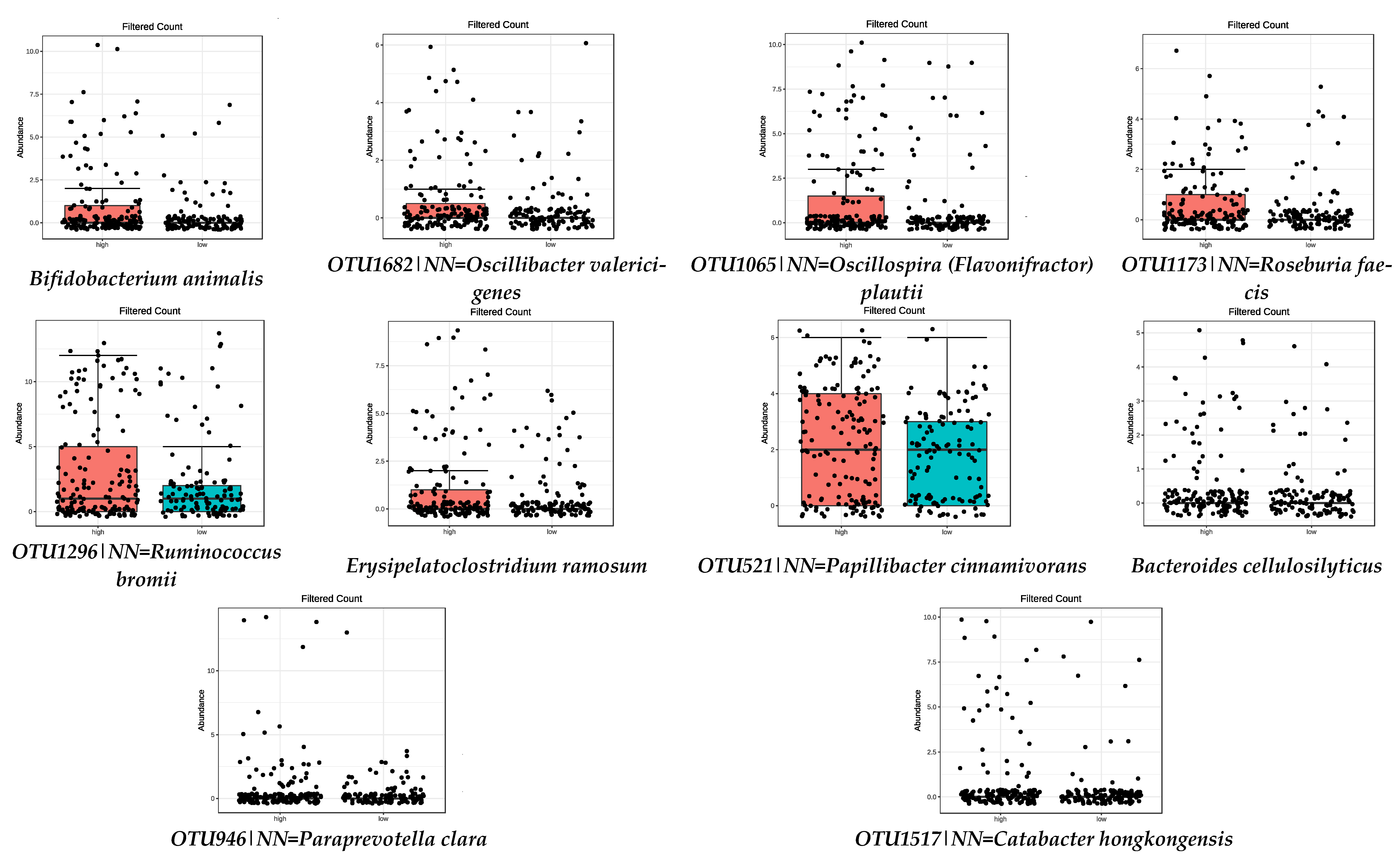
3.2. Microbiota Composition: MD Adherence
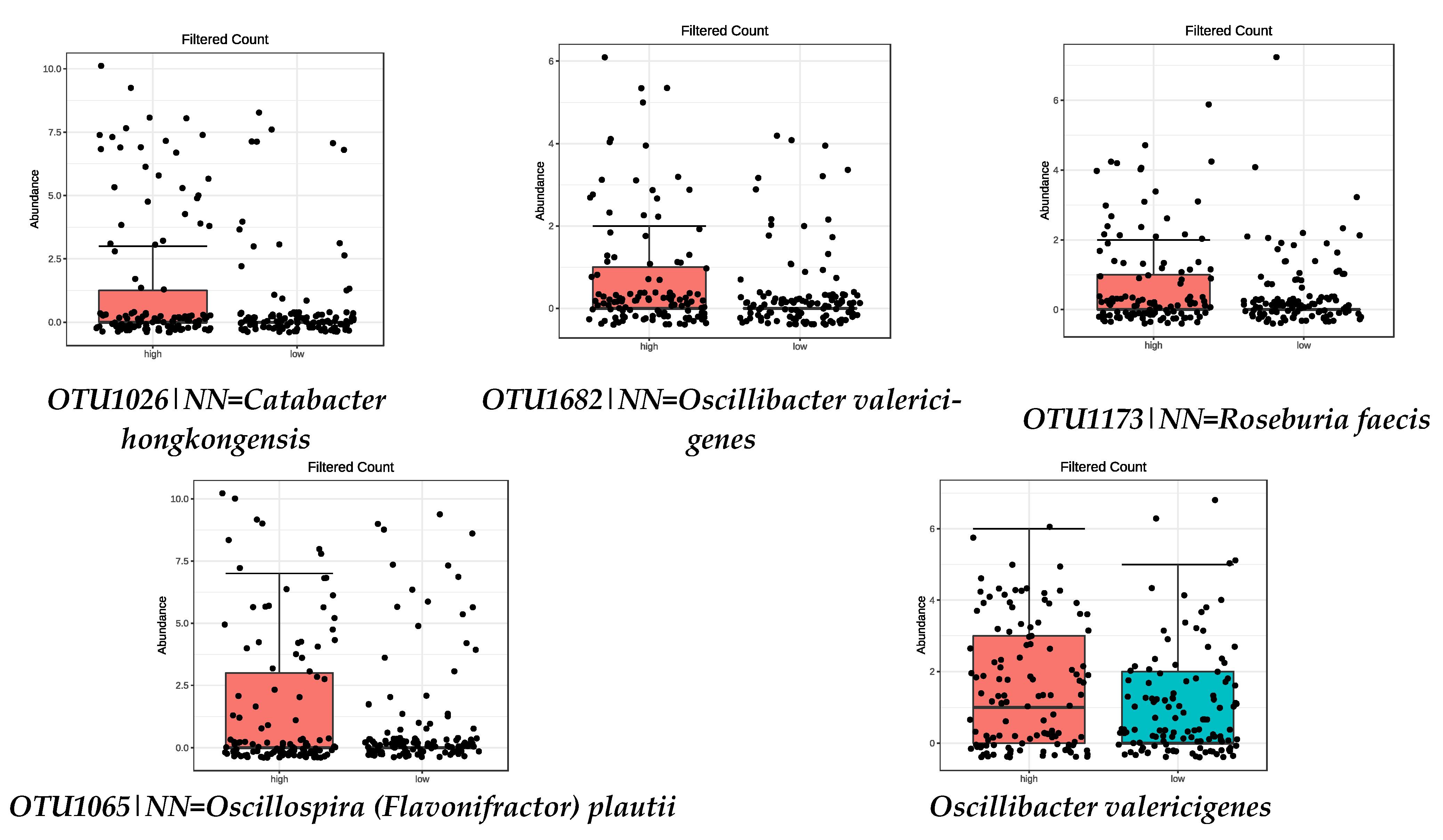
3.3. Microbiota Composition: Fibre Intake
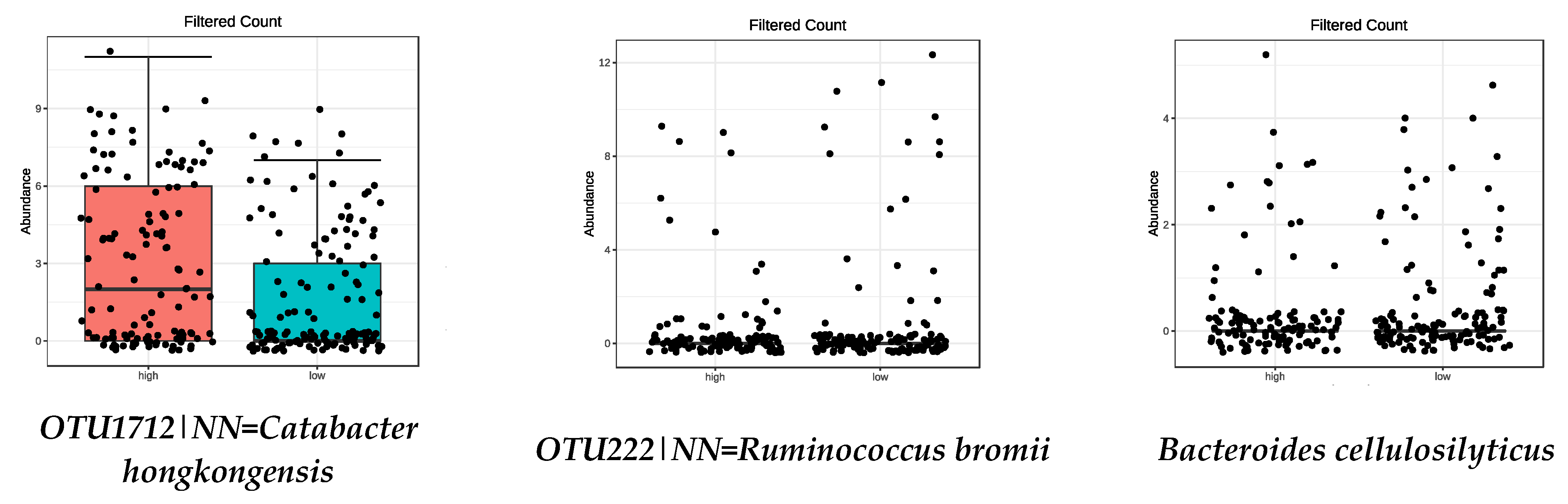
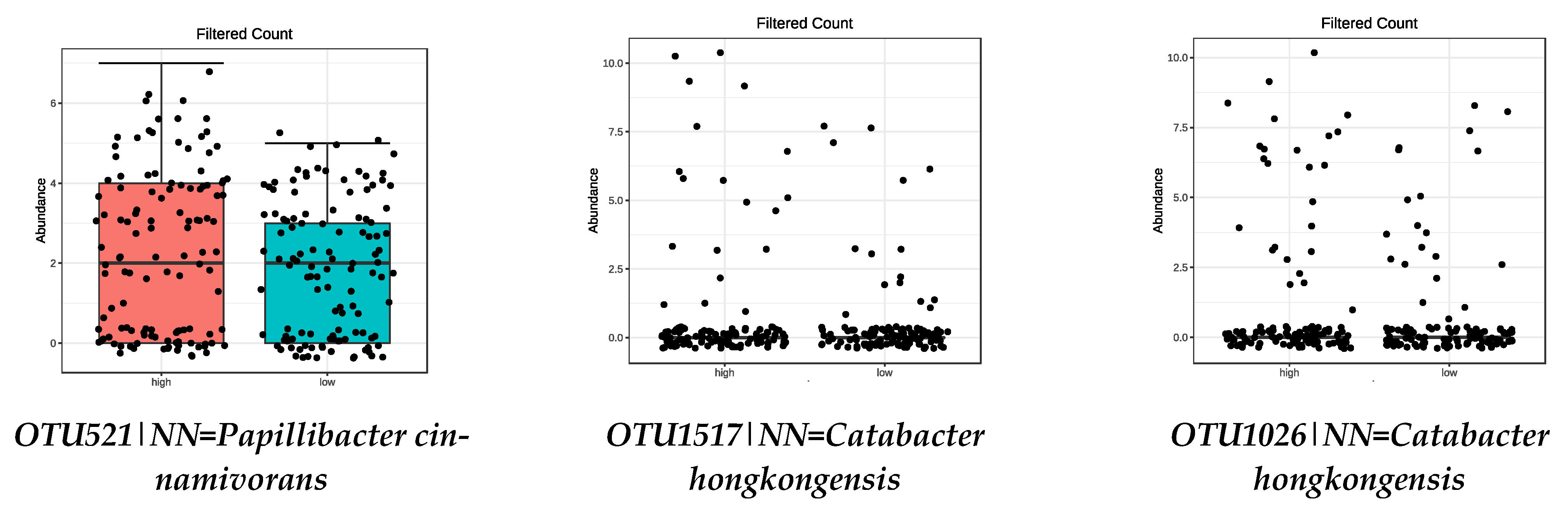
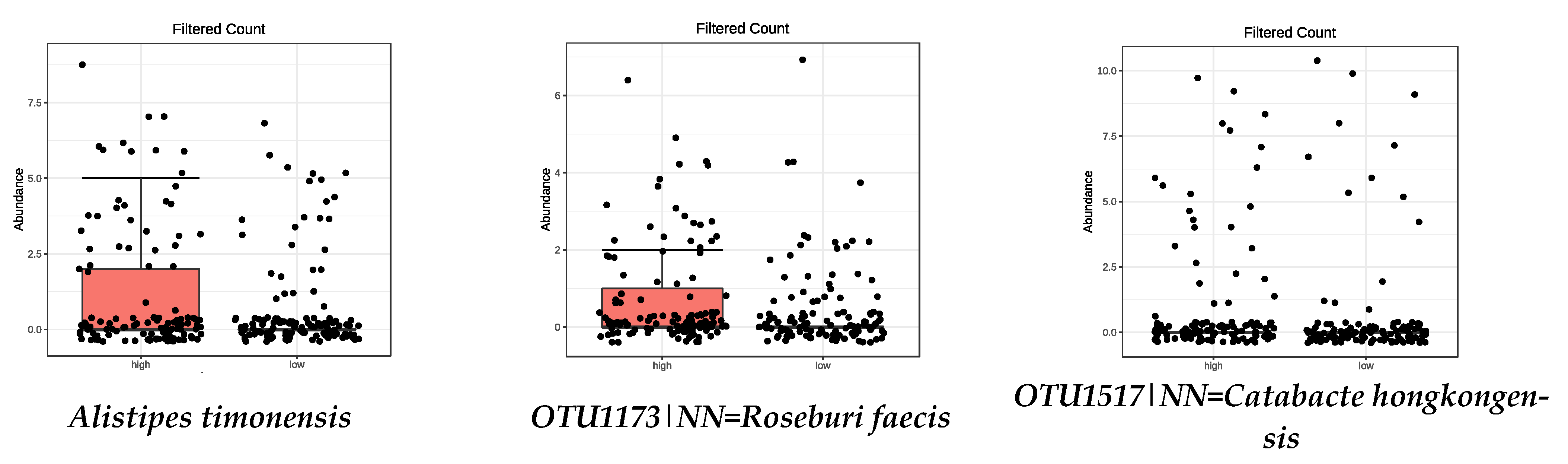
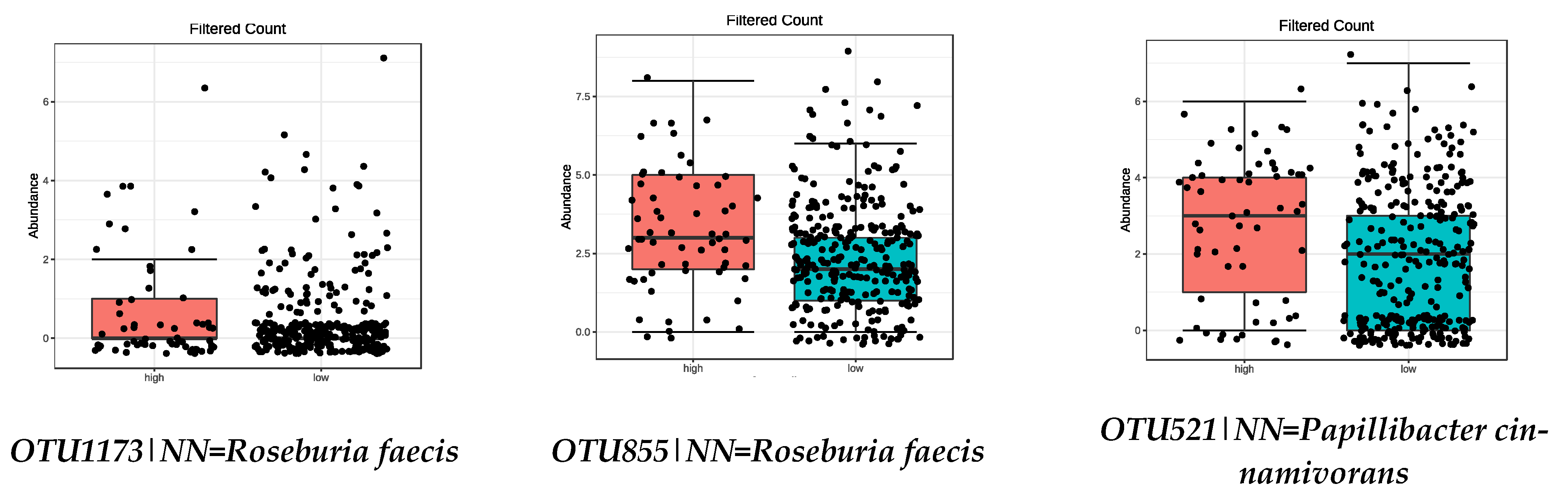
3.4. Microbiota Composition: Food Groups
3.5. Species Richness, Evenness, and Diversity
3.6. Comparison of Functional Potential of the Gut Microbiota
4. Discussion
5. MD High-Adherence Species
6. Study Limitations
7. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Maslowski, K.M.; Mackay, C.R. Diet, gut microbiota and immune responses. Nat. Immunol. 2011, 12, 5–9. [Google Scholar] [CrossRef]
- Myles, I.A. Fast food fever: Reviewing the impacts of the Western diet on immunity. Nutr. J. 2014, 13, 61. [Google Scholar] [CrossRef] [PubMed]
- Biro, F.M.; Wien, M. Childhood obesity and adult morbidities. Am. J. Clin. Nutr. 2010, 91, 1499–1505. [Google Scholar] [CrossRef]
- Milner, J.J.; Beck, M.A. The impact of obesity on the immune response to infection. Proc. Nutr. Soc. 2012, 71, 298–306. [Google Scholar] [CrossRef]
- Rook, G.A.W. 99th Dahlem Conference on Infection, Inflammation and Chronic Inflammatory Disorders: Darwinian medicine and the ‘hygiene’ or ‘old friends’ hypothesis. Clin. Exp. Immunol. 2010, 160, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Sofi, F.; Macchi, C.; Abbate, R.; Gensini, G.F.; Casini, A. Mediterranean diet and health status: An updated meta-analysis and a proposal for a literature-based adherence score. Public Health Nutr. 2014, 17, 2769–2782. [Google Scholar] [CrossRef]
- Le Chatelier, E.; Nielsen, T.; Qin, J.; Prifti, E.; Hildebrand, F.; Falony, G.; Almeida, M.; Arumugam, M.; Batto, J.-M.; Kennedy, S.; et al. Richness of human gut microbiome correlates with metabolic markers. Nature 2013, 500, 541–546. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Van Hul, M. Novel opportunities for next-generation probiotics targeting metabolic syndrome. Curr. Opin. Biotechnol. 2015, 32, 21–27. [Google Scholar] [CrossRef] [PubMed]
- Ogce, F.; Ceber, E.; Ekti, R.; Oran, N.T. Comparison of mediterranean, Western and Japanese diets and some recommendations. Asian Pac. J. Cancer Prev. 2008, 9, 351–356. [Google Scholar]
- Garcia-Mantrana, I.; Selma-Royo, M.; Alcantara, C.; Collado, M.C. Shifts on Gut Microbiota Associated to Mediterranean Diet Adherence and Specific Dietary Intakes on General Adult Population. Front. Microbiol. 2018, 9, 890. [Google Scholar] [CrossRef]
- Meslier, V.; Laiola, M.; Roager, H.M.; De Filippis, F.; Roume, H.; Quinquis, B.; Giacco, R.; Mennella, I.; Ferracane, R.; Pons, N.; et al. Mediterranean diet intervention in overweight and obese subjects lowers plasma cholesterol and causes changes in the gut microbiome and metabolome independently of energy intake. Gut 2020, 69, 1258–1268. [Google Scholar] [CrossRef]
- Ghosh, T.S.; Rampelli, S.; Jeffery, I.B.; Santoro, A.; Neto, M.; Capri, M.; Giampieri, E.; Jennings, A.; Candela, M.; Turroni, S.; et al. Mediterranean diet intervention alters the gut microbiome in older people reducing frailty and improving health status: The NU-AGE 1-year dietary intervention across five European countries. Gut 2020, 69, 1218–1228. [Google Scholar] [CrossRef]
- Rivière, A.; Selak, M.; Lantin, D.; Leroy, F.; De Vuyst, L. Bifidobacteria and Butyrate-Producing Colon Bacteria: Importance and Strategies for Their Stimulation in the Human Gut. Front. Microbiol. 2016, 7, 979. [Google Scholar] [CrossRef]
- Gutiérrez-Díaz, I.; Fernández-Navarro, T.; Salazar, N.; Bartolomé, B.; Moreno-Arribas, M.V.; De Andres-Galiana, E.J.; Fernández-Martínez, J.L.; De Los Reyes-Gavilán, C.G.; Gueimonde, M.; González, S. Adherence to a mediterranean diet influences the fecal metabolic profile of microbial-derived phenolics in a Spanish cohort of middle-age and older people. J. Agric. Food Chem. 2017, 65, 586–595. [Google Scholar] [CrossRef]
- De Filippis, F.; Pellegrini, N.; Vannini, L.; Jeffery, I.B.; La Storia, A.; Laghi, L.; Serrazanetti, D.I.; Di Cagno, R.; Ferrocino, I.; Lazzi, C.; et al. High-level adherence to a Mediterranean diet beneficially impacts the gut microbiota and associated metabolome. Gut 2016, 65, 1812–1821. [Google Scholar] [CrossRef] [PubMed]
- Mitsou, E.K.; Kakali, A.; Antonopoulou, S.; Mountzouris, K.C.; Yannakoulia, M.; Panagiotakos, D.B.; Kyriacou, A. Adherence to the Mediterranean diet is associated with the gut microbiota pattern and gastrointestinal characteristics in an adult population. Br. J. Nutr. 2017, 117, 1645–1655. [Google Scholar] [CrossRef]
- World Medical Association. World Medical Association Declaration of Helsinki: Ethical principles for medical research involving human subjects. J. Am. Med. Assoc. 2013, 310, 2191–2194. [Google Scholar] [CrossRef] [PubMed]
- Ramos-Lopez, O.; Riezu-Boj, J.I.; Milagro, F.I.; Goni, L.; Cuervo, M.; Martinez, J.A. Differential lipid metabolism outcomes associated with ADRB2 gene polymorphisms in response to two dietary interventions in overweight/obese subjects. Nutr. Metab. Cardiovasc. Dis. 2018, 28, 165–172. [Google Scholar] [CrossRef] [PubMed]
- Zulet, M.A.; Bondia-Pons, I.; Abete, I.; de la Iglesia, R.; López-Legarrea, P.; Forga, L.; Navas-Carretero, S.; Martínez, J.A. The reduction of the metabolyc syndrome in Navarra-Spain (RESMENA-S) study: A multidisciplinary strategy based on chrononutrition and nutritional education, together with dietetic and psychological control. Nutr. Hosp. 2011, 26, 16–26. [Google Scholar] [CrossRef] [PubMed]
- Shatenstein, B.; Nadon, S.; Godin, C.; Ferland, G. Development and Validation of a Food Frequency Questionnaire. Can. J. Diet. Pract. Res. 2005, 66, 67–75. [Google Scholar] [CrossRef]
- Moreiras, O.; Carbajal, Á.; Cabrera, L.; Cuadrado, C. Tablas de Composición de Alimentos. Guía de Prácticas, 16th ed.; Pirámide: Madrid, Spain, 2013. [Google Scholar]
- Martínez-González, M.A.; García-Arellano, A.; Toledo, E.; Salas-Salvadó, J.; Buil-Cosiales, P.; Corella, D.; Covas, M.I.; Schröder, H.; Arós, F.; Gómez-Gracia, E.; et al. A 14-Item Mediterranean Diet Assessment Tool and Obesity Indexes among High-Risk Subjects: The PREDIMED Trial. PLoS ONE 2012, 7, e43134. [Google Scholar] [CrossRef]
- Illumina 16S Metagenomic Sequencing Library Preparation. Available online: https://emea.illumina.com/search.html?q=16s metagenomic sequencing library preparation&filter=all&p=1&langsel=/es/ (accessed on 4 November 2019).
- Hildebrand, F.; Tito, R.Y.; Voigt, A.; Bork, P.; Raes, J. Correction: LotuS: An efficient and user-friendly OTU processing pipeline. Microbiome 2014, 2, 37. [Google Scholar] [CrossRef]
- Edgar, R.C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996–998. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. UCHIME2: Improved chimera prediction for amplicon sequencing. bioRxiv 2016, 074252. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Ritari, J.; Salojärvi, J.; Lahti, L.; de Vos, W.M. Improved taxonomic assignment of human intestinal 16S rRNA sequences by a dedicated reference database. BMC Genom. 2015, 16, 1056. [Google Scholar] [CrossRef]
- Gentleman, R.C.; Carey, V.J.; Bates, D.M.; Bolstad, B.; Dettling, M.; Dudoit, S.; Ellis, B.; Gautier, L.; Ge, Y.; Gentry, J.; et al. Bioconductor: Open software development for computational biology and bioinformatics. Genome Biol. 2004, 5, 80. [Google Scholar] [CrossRef]
- Dhariwal, A.; Chong, J.; Habib, S.; King, I.L.; Agellon, L.B.; Xia, J. MicrobiomeAnalyst: A Web-Based Tool for Comprehensive Statistical, Visual and Meta-Analysis of Microbiome Data. Available online: https://academic.oup.com/nar/article-abstract/45/W1/W180/3760191 (accessed on 21 April 2020).
- OMS|10 Datos Sobre la Obesidad. Available online: https://www.who.int/features/factfiles/obesity/facts/es/ (accessed on 8 June 2020).
- Lobionda, S.; Sittipo, P.; Kwon, H.Y.; Lee, Y.K. The role of gut microbiota in intestinal inflammation with respect to diet and extrinsic stressors. Microorganisms 2019, 7, 271. [Google Scholar] [CrossRef]
- Graf, D.; Di Cagno, R.; Fåk, F.; Flint, H.J.; Nyman, M.; Saarela, M.; Watzl, B. Contribution of diet to the composition of the human gut microbiota. Microb. Ecol. Health Dis. 2015, 26. [Google Scholar] [CrossRef]
- Mendez, M.A.; Popkin, B.M.; Jakszyn, P.; Berenguer, A.; Tormo, M.J.; Sanchéz, M.J.; Quirós, J.R.; Pera, G.; Navarro, C.; Martinez, C.; et al. Adherence to a Mediterranean Diet Is Associated with Reduced 3-Year Incidence of Obesity. J. Nutr. 2006, 136, 2934–2938. [Google Scholar] [CrossRef]
- Lopez-Legarrea, P.; Fuller, N.R.; Zulet, M.A.; Martinez, J.A.; Caterson, I.D. The influence of Mediterranean, carbohydrate and high protein diets on gut microbiota composition in the treatment of obesity and associated inflammatory state. Asia Pac. J. Clin. Nutr. 2014, 23, 360–368. [Google Scholar] [CrossRef] [PubMed]
- AlEssa, H.B.; Malik, V.S.; Yuan, C.; Willett, W.C.; Huang, T.; Hu, F.B.; Tobias, D.K. Dietary patterns and cardiometabolic and endocrine plasma biomarkers in US women. Am. J. Clin. Nutr. 2017, 105, 432–441. [Google Scholar] [CrossRef]
- Million, M.; Maraninchi, M.; Henry, M.; Armougom, F.; Richet, H.; Carrieri, P.; Valero, R.; Raccah, D.; Vialettes, B.; Raoult, D. Obesity-associated gut microbiota is enriched in Lactobacillus reuteri and depleted in Bifidobacterium animalis and Methanobrevibacter smithii. Int. J. Obes. 2012, 36, 817–825. [Google Scholar] [CrossRef]
- Horiuchi, H.; Kamikado, K.; Aoki, R.; Suganuma, N.; Nishijima, T.; Nakatani, A.; Kimura, I. Bifidobacterium animalis subsp. lactis GCL2505 modulates host energy metabolism via the short-chain fatty acid receptor GPR43. Sci. Rep. 2020, 10, 4158. [Google Scholar] [CrossRef] [PubMed]
- Den Besten, G.; Van Eunen, K.; Groen, A.K.; Venema, K.; Reijngoud, D.J.; Bakker, B.M. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid. Res. 2013, 54, 2325–2340. [Google Scholar] [CrossRef]
- Robert, C.; Chassard, C.; Lawson, P.A.; Bernalier-Donadille, A. Bacteroides cellulosilyticus sp. nov., a cellulolytic bacterium from the human gut microbial community. Int. J. Syst. Evol. Microbiol. 2007, 57, 1516–1520. [Google Scholar] [CrossRef] [PubMed]
- McNulty, N.P.; Wu, M.; Erickson, A.R.; Pan, C.; Erickson, B.K.; Martens, E.C.; Pudlo, N.A.; Muegge, B.D.; Henrissat, B.; Hettich, R.L.; et al. Effects of Diet on Resource Utilization by a Model Human Gut Microbiota Containing Bacteroides cellulosilyticus WH2, a Symbiont with an Extensive Glycobiome. PLoS Biol. 2013, 11, e1001637. [Google Scholar] [CrossRef]
- Morotomi, M.; Nagai, F.; Sakon, H.; Tanaka, R. Paraprevotella clara gen. nov., sp. nov. and Paraprevotella xylaniphila sp. nov., members of the family “Prevotellaceae” isolated from human faeces. Int. J. Syst. Evol. Microbiol. 2009, 59, 1895–1900. [Google Scholar] [CrossRef]
- Li, J.; Hou, Q.; Zhang, J.; Xu, H.; Sun, Z.; Menghe, B.; Zhang, H. Carbohydrate Staple Food Modulates Gut Microbiota of Mongolians in China. Front. Microbiol. 2017, 8, 484. [Google Scholar] [CrossRef]
- Flint, H.J.; Bayer, E.A.; Rincon, M.T.; Lamed, R.; White, B.A. Polysaccharide utilization by gut bacteria: Potential for new insights from genomic analysis. Nat. Rev. Microbiol. 2008, 6, 121–131. [Google Scholar] [CrossRef]
- Iino, T.; Mori, K.; Tanaka, K.; Suzuki, K.I.; Harayama, S. Oscillibacter valericigenes gen. nov., sp. nov., a valerate-producing anaerobic bacterium isolated from the alimentary canal of a Japanese corbicula clam. Int. J. Syst. Evol. Microbiol. 2007, 57, 1840–1845. [Google Scholar] [CrossRef]
- Yuille, S.; Reichardt, N.; Panda, S.; Dunbar, H.; Mulder, I.E. Human gut bacteria as potent class I histone deacetylase inhibitors in vitro through production of butyric acid and valeric acid. PLoS ONE 2018, 13. [Google Scholar] [CrossRef]
- Walker, A.W.; Ince, J.; Duncan, S.H.; Webster, L.M.; Holtrop, G.; Ze, X.; Brown, D.; Stares, M.D.; Scott, P.; Bergerat, A.; et al. Dominant and diet-responsive groups of bacteria within the human colonic microbiota. ISME J. 2011, 5, 220–230. [Google Scholar] [CrossRef] [PubMed]
- Borrelli, L.; Coretti, L.; Dipineto, L.; Bovera, F.; Menna, F.; Chiariotti, L.; Nizza, A.; Lembo, F.; Fioretti, A. Insect-based diet, a promising nutritional source, modulates gut microbiota composition and SCFAs production in laying hens. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Kasai, C.; Sugimoto, K.; Moritani, I.; Tanaka, J.; Oya, Y.; Inoue, H.; Tameda, M.; Shiraki, K.; Ito, M.; Takei, Y.; et al. Comparison of the gut microbiota composition between obese and non-obese individuals in a Japanese population, as analyzed by terminal restriction fragment length polymorphism and next-generation sequencing. BMC Gastroenterol. 2015, 15, 100. [Google Scholar] [CrossRef]
- Konikoff, T.; Gophna, U. Oscillospira: A Central, Enigmatic Component of the Human Gut Microbiota. Trends Microbiol. 2016, 24, 523–524. [Google Scholar] [CrossRef]
- Ryan, K.K.; Tremaroli, V.; Clemmensen, C.; Kovatcheva-Datchary, P.; Myronovych, A.; Karns, R.; Wilson-Pérez, H.E.; Sandoval, D.A.; Kohli, R.; Bäckhed, F.; et al. FXR is a molecular target for the effects of vertical sleeve gastrectomy. Nature 2014, 509, 183–188. [Google Scholar] [CrossRef]
- Lau, S.K.P.; Mcnabb, A.; Woo, G.K.S.; Hoang, L.; Fung, A.M.Y.; Chung, L.M.W.; Woo, P.C.Y.; Yuen, K.-Y.; Hospital, M.; Kong, H. Catabacter hongkongensis gen. nov., sp. nov., Isolated from Blood Cultures of Patients from Hong Kong and Canada. J. Clin. Microbiol. 2007, 45, 395–401. [Google Scholar] [CrossRef] [PubMed]
- Venkataraman, A.; Sieber, J.R.; Schmidt, A.W.; Waldron, C.; Theis, K.R.; Schmidt, T.M. Variable responses of human microbiomes to dietary supplementation with resistant starch. Microbiome 2016, 4, 33. [Google Scholar] [CrossRef]
- Peng, L.; Li, Z.-R.; Green, R.S.; Holzman, I.R.; Lin, J. Butyrate Enhances the Intestinal Barrier by Facilitating Tight Junction Assembly via Activation of AMP-Activated Protein Kinase in Caco-2 Cell Monolayers. J. Nutr. 2009, 139, 1619–1625. [Google Scholar] [CrossRef]
- Ruemmele, F.M.; Schwartz, S.; Seidman, E.G.; Dionne, S.; Levy, E.; Lentze, M.J. Butyrate induced Caco-2 cell apoptosis is mediated via the mitochondrial pathway. Gut 2003, 52, 94–100. [Google Scholar] [CrossRef]
- Mikkelsen, K.H.; Allin, K.H.; Knop, F.K. Effect of antibiotics on gut microbiota, glucose metabolism and body weight regulation: A review of the literature. Diabetes Obes. Metab. 2016, 18, 444–453. [Google Scholar] [CrossRef]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef]
- Midtvedt, T.; Gustafsson, B.E. Microbial conversation of bilirubin to urobilins in vitro and in vivo. Acta Pathol. Microbiol. Scand. Sect. B Microbiol. 2009, 89, 57–60. [Google Scholar] [CrossRef]
- Becker, N.; Kunath, J.; Loh, G.; Blaut, M. Human intestinal microbiota: Characterization of a simplified and stable gnotobiotic rat model. Gut Microbes 2011, 25, 25–33. [Google Scholar] [CrossRef]
- Mandić, A.D.; Woting, A.; Jaenicke, T.; Sander, A.; Sabrowski, W.; Rolle-Kampcyk, U.; von Bergen, M.; Blaut, M. Clostridium ramosum regulates enterochromaffin cell development and serotonin release. Sci. Rep. 2019, 9, 1177. [Google Scholar] [CrossRef] [PubMed]
- Munch Roager, H.; Vogt, J.K.; Kristensen, M.; Hansen, L.B.S.; Ibrügger, S.; Maerkedahl, R.B.; Bahl, M.I.; Lind, M.V.; Nielsen, R.L.; Frøkiaer, H.; et al. Whole grain-rich diet reduces body weight and systemic low-grade inflammation without inducing major changes of the gut microbiome: A randomised cross-over trial. Gut 2019, 68, 83–93. [Google Scholar] [CrossRef]
- Mabrok, H.B.; Klopfleisch, R.; Ghanem, K.Z.; Clavel, T.; Blaut, M.; Loh, G. Lignan transformation by gut bacteria lowers tumor burden in a gnotobiotic rat model of breast cancer. Carcinogenesis 2012, 33, 203–208. [Google Scholar] [CrossRef] [PubMed]
- Elmassry, M.M.; Chung, E.; Cao, J.J.; Hamood, A.N.; Shen, C.-L. Osteoprotective effect of green tea polyphenols and annatto-extracted tocotrienol in obese mice is associated with enhanced microbiome vitamin K2 biosynthetic pathways. J. Nutr. Biochem. 2020, 86, 108492. [Google Scholar] [CrossRef] [PubMed]
- Boureau, H.; Decré, D.; Carlier, J.P.; Guichet, C.; Bourlioux, P. Identification of a Clostridium cocleatum strain involved in an anti-Clostridium difficile barrier effect and determination of its mucin-degrading enzymes. Res. Microbiol. 1993, 144, 405–410. [Google Scholar] [CrossRef]
- Biagi, E.; Nylund, L.; Candela, M.; Ostan, R.; Bucci, L.; Pini, E.; Nikkïla, J.; Monti, D.; Satokari, R.; Franceschi, C.; et al. Through ageing, and beyond: Gut microbiota and inflammatory status in seniors and centenarians. PLoS ONE 2010, 5, e10667. [Google Scholar] [CrossRef]
- Landberg, R.; Hanhineva, K. Biomarkers of a healthy nordic diet—From dietary exposure biomarkers to microbiota signatures in the metabolome. Nutrients 2019, 12, 27. [Google Scholar] [CrossRef]
- Kushida, M.; Sugawara, S.; Asano, M.; Yamamoto, K.; Fukuda, S.; Tsuduki, T. Effects of the 1975 Japanese diet on the gut microbiota in younger adults. J. Nutr. Biochem. 2019, 64, 121–127. [Google Scholar] [CrossRef] [PubMed]
| Variables | High Adherence (3rd Tertile) (n = 94) | Low Adherence (1st Tertile) (n = 128) | p Value |
|---|---|---|---|
| Age (y) | 47.3 ± 1.2 | 41.7 ± 0.9 | 0.020 * |
| BMI | 27.7 ± 0.6 | 29.9 ± 0.6 | 0.002 * |
| Glucose (mg/dL) | 95 ± 2 | 93 ± 1 | 0.394 |
| Total cholesterol (mg/dL) | 209 ± 4 | 208 ± 4 | 0.822 |
| HDL (mg/dL) | 59 ± 1 | 57 ± 1 | 0.261 |
| LDL (mg/dL) | 133 ± 4 | 132 ± 3 | 0.904 |
| Triglycerides (mg/dL) | 87 ± 5 | 93 ± 4 | 0.336 |
| HOMA-IR | 1.4 ± 0.15 | 1.6 ± 0.1 | 0.011 * |
| Carbohydrate intake (%) | 54.5 ± 0.7 | 54.7 ± 0.6 | 0.135 |
| Protein intake (%) | 21.8 ± 0.4 | 22.2 ± 0.4 | 0.286 |
| Fat intake (%) | 23.6 ± 0.5 | 23.0 ± 0.4 | 0.085 |
| Fiber intake (g/day) | 35.9 ± 1.6 | 24.5 ± 0.7 | 0.000 * |
| Energy intake (kcal/day) | 2932 ± 106 | 2872 ± 84 | 0.680 |
| High Adherence (3rd Tertile) | Low Adherence (1st Tertile) | ||
|---|---|---|---|
| SPECIES | FDR | SPECIES | FDR |
| Bifidobacterium animalis | 1.21 × 10−7 | OTU100|NN = Eubacterium saphenum GU427005|D = 91 | 4.44 × 10−5 |
| Bacteroides cellulosilyticus | 4.47 × 10−7 | OTU375|NN = Succinivibrio dextrinosolvens Y17600|D = 97 | 0.0001 |
| OTU946|NN = Paraprevotella clara AB331896|D = 86.8 | 1.72 × 10−5 | OTU759|NN = Gordonibacter pamelaeae AB566419|D = 87.6 | 0.0005 |
| OTU1682|NN = Oscillibacter valericigenes AB238598|D = 91.1 | 3.42 × 10−5 | OTU11|NN = Butyricicoccus pullicaecorum EU410376|D = 89 | 0.0002 |
| OTU1065|NN = Oscillospira (Flavonifractor) plautii Y18187|D = 86.6 | 3.42 × 10−5 | Christensenella minuta | 0.0020 |
| OTU1173|NN = Roseburia faecis AY804149|D = 94.9 | 0.0008 | Parabacteroides goldsteinii | 0.0073 |
| OTU1517|NN = Catabacter hongkongensis AB671763|D = 87 | 0.0008 | OTU1625|NN = Anaerotruncus colihominis DQ002932|D = 89 | 0.0120 |
| OTU1296|NN = Ruminococcus bromii DQ882649|D = 92.3 | 0.0120 | Alistipes timonensis | 0.0155 |
| Erysipelatoclostridium ramosum | 0.0176 | Prevotella corporis | 0.0192 |
| OTU521|NN = Papillibacter cinnamivorans AF167711|D = 89 | 0.0463 | ||
| SPECIES | FDR |
|---|---|
| High intake of FIBRE | |
| OTU1026|NN = Catabacter hongkongensis AB671763|D = 82.4 | 8.48 × 10−11 |
| OTU1682|NN = Oscillibacter valericigenes AB238598|D = 91.1 | 6.73 × 10−5 |
| OTU1173|NN = Roseburia faecis AY804149|D = 94.9 | 8.38 × 10−5 |
| OTU1065|NN = Oscillospira (Flavonifractor) plautii Y18187|D = 86.6 | 0.0018 |
| Oscillibacter valericigenes | 0.0045 |
| High intake of LEGUMES | |
| OTU222|NN = Ruminococcus bromii DQ882649|D = 89.9 | 7.45 × 10−5 |
| OTU1712|NN = Catabacter hongkongensis AB671763|D = 84 | 0.0011 |
| Bacteroides cellulosilyticus | 0.0001 |
| High intake of VEGETABLES | |
| OTU1517|NN = Catabacter hongkongensis AB671763|D = 87 | 1.19 × 10−11 |
| OTU1026|NN = Catabacter hongkongensis AB671763|D = 82.4 | 1.43 × 10−6 |
| OTU521|NN = Papillibacter cinnamivorans AF167711|D = 89 | 0.0019 |
| High intake of FRUIT | |
| OTU1517|NN = Catabacter hongkongensis AB671763|D = 87 | 0.0008 |
| OTU1173|NN = Roseburia faecis AY804149|D = 94.9 | 0.0012 |
| High intake of NUTS | |
| OTU1173|NN = Roseburia faecis AY804149|D = 94.9 | 8.11 × 10−5 |
| OTU855|NN = Roseburia faecis AY804149|D = 95.5 | 0.0001 |
| OTU521|NN = Papillibacter cinnamivorans AF167711|D = 89 | 0.0024 |
| K0 Name | Definition | Log2FC | p Value | FDR |
|---|---|---|---|---|
| K00720 | Ceramide glucosyltransferase | 1.803 | 2.93 × 10−7 | 0.001 |
| K01884 | Cysteinyl-tRNA synthetase | 1.792 | 8.67 × 10−7 | 0.002 |
| K03653 | N-glycosylase/DNA lyase | 1.6569 | 1.57 × 10−6 | 0.002 |
| K02106 | Short-chain fatty acids transporter | 1.0933 | 2.26 × 10−5 | 0.03 |
| K07486 | Transposase | 0.9605 | 4.51 × 10−5 | 0.04 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rosés, C.; Cuevas-Sierra, A.; Quintana, S.; Riezu-Boj, J.I.; Martínez, J.A.; Milagro, F.I.; Barceló, A. Gut Microbiota Bacterial Species Associated with Mediterranean Diet-Related Food Groups in a Northern Spanish Population. Nutrients 2021, 13, 636. https://doi.org/10.3390/nu13020636
Rosés C, Cuevas-Sierra A, Quintana S, Riezu-Boj JI, Martínez JA, Milagro FI, Barceló A. Gut Microbiota Bacterial Species Associated with Mediterranean Diet-Related Food Groups in a Northern Spanish Population. Nutrients. 2021; 13(2):636. https://doi.org/10.3390/nu13020636
Chicago/Turabian StyleRosés, Carles, Amanda Cuevas-Sierra, Salvador Quintana, José I. Riezu-Boj, J. Alfredo Martínez, Fermín I. Milagro, and Anna Barceló. 2021. "Gut Microbiota Bacterial Species Associated with Mediterranean Diet-Related Food Groups in a Northern Spanish Population" Nutrients 13, no. 2: 636. https://doi.org/10.3390/nu13020636
APA StyleRosés, C., Cuevas-Sierra, A., Quintana, S., Riezu-Boj, J. I., Martínez, J. A., Milagro, F. I., & Barceló, A. (2021). Gut Microbiota Bacterial Species Associated with Mediterranean Diet-Related Food Groups in a Northern Spanish Population. Nutrients, 13(2), 636. https://doi.org/10.3390/nu13020636








