Antimicrobial Activity of EPA and DHA against Oral Pathogenic Bacteria Using an In Vitro Multi-Species Subgingival Biofilm Model
Abstract
:1. Introduction
2. Materials and Methods
2.1. Omega 3 Fatty-Acids
2.2. Bacterial Strains and Culture Conditions
2.3. Antibacterial Effect of EPA and DHA against Planktonic Bacteria
2.4. Biofilm Development
2.5. Antimicrobial Activity on Biofilms
2.6. Microbiological Outcomes: Quantitative Polymerase Chain Reaction (qPCR) Analysis
2.7. Confocal Laser Scanning Microscopy (CLSM) Analysis
2.8. Scanning Electron Microscopy (SEM) Analysis
2.9. Statistical Analyses
3. Results
3.1. Antibacterial Effect of DHA and EPA on Planktonic Bacteria
3.2. Antibacterial Effect of the DHA Extract on Biofilm
3.2.1. Analysis by qPCR
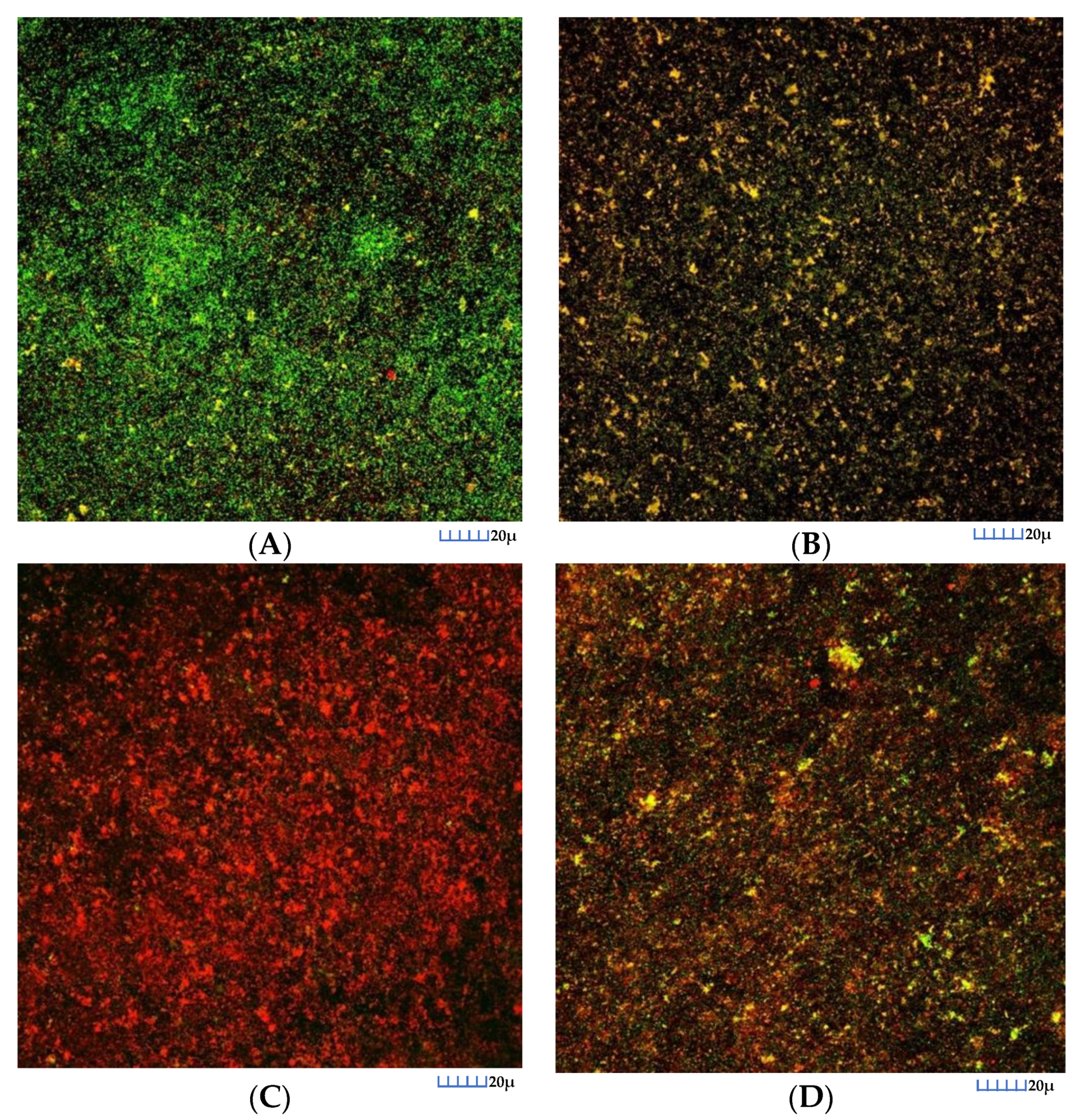
3.2.2. CLSM Analysis
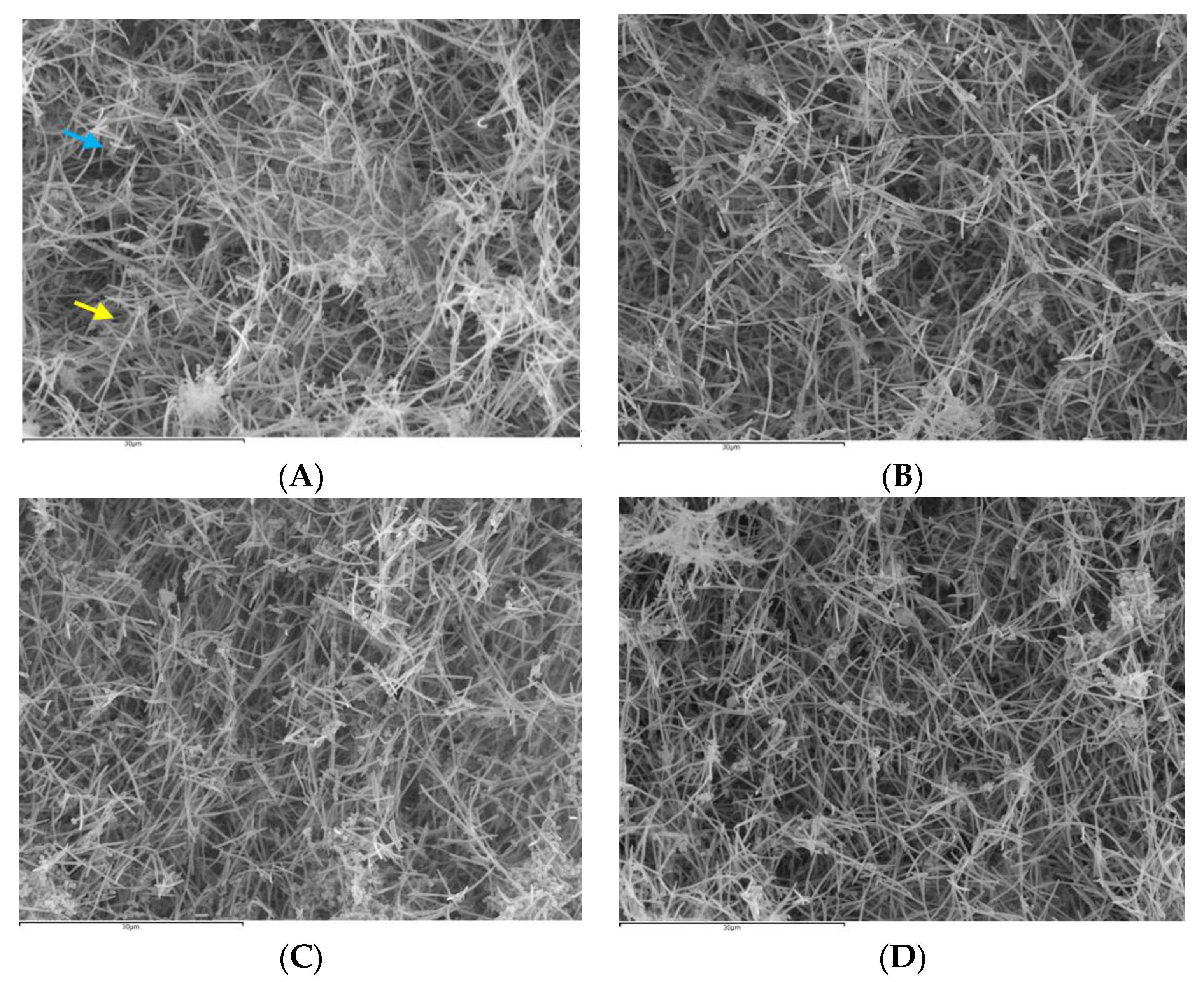
3.2.3. SEM Analysis
3.3. Antibacterial Effect of the EPA Extract
3.3.1. qPCR Analysis
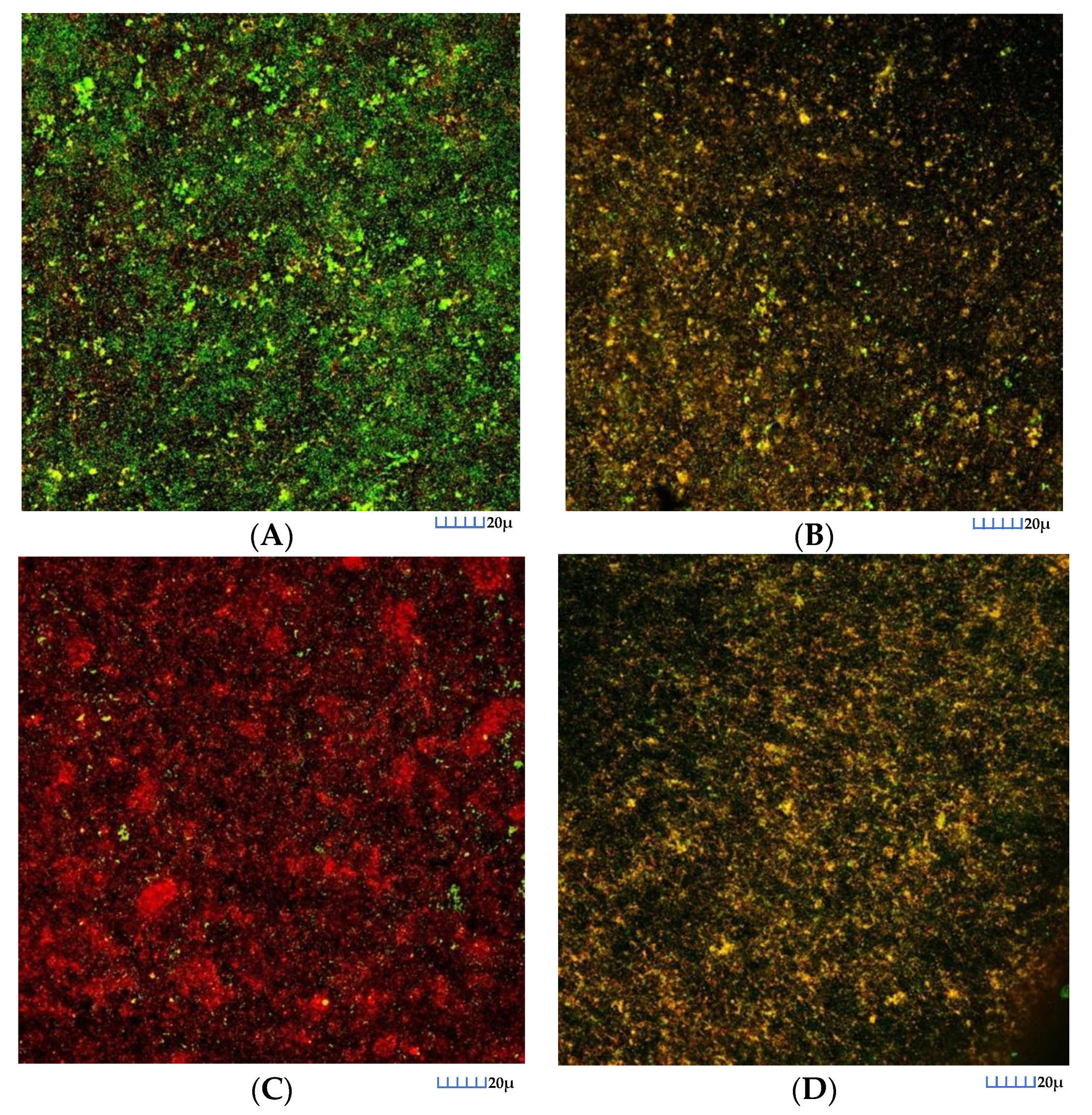
3.3.2. CLSM Analysis
3.3.3. SEM Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Papapanou, P.N.; Sanz, M.; Buduneli, N.; Dietrich, T.; Feres, M.; Fine, D.H.; Flemmig, T.F.; Garcia, R.; Giannobile, W.V.; Graziani, F.; et al. Periodontitis: Consensus report of workgroup 2 of the 2017 World Workshop on the Classification of Periodontal and Peri-Implant Diseases and Conditions. J. Clin. Periodontol. 2018, 45 (Suppl. S20), S162–S170. [Google Scholar] [CrossRef] [PubMed]
- Beikler, T.; Flemmig, T.F. Oral biofilm-associated diseases: Trends and implications for quality of life, systemic health and expenditures. Periodontology 2000 2011, 55, 87–103. [Google Scholar] [CrossRef] [PubMed]
- Paster, B.J.; Boches, S.K.; Galvin, J.L.; Ericson, R.E.; Lau, C.N.; Levanos, V.A.; Sahasrabudhe, A.; Dewhirst, F.E. Bacterial diversity in human subgingival plaque. J. Bacteriol. 2001, 183, 3770–3783. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Socransky, S.S.; Haffajee, A.D. Periodontal microbial ecology. Periodontology 2000 2005, 38, 135–187. [Google Scholar] [CrossRef]
- Romero-Lastra, P.; Sanchez, M.C.; Ribeiro-Vidal, H.; Llama-Palacios, A.; Figuero, E.; Herrera, D.; Sanz, M. Comparative gene expression analysis of Porphyromonas gingivalis ATCC 33277 in planktonic and biofilms states. PLoS ONE 2017, 12, e0174669. [Google Scholar] [CrossRef]
- Sanchez, M.C.; Romero-Lastra, P.; Ribeiro-Vidal, H.; Llama-Palacios, A.; Figuero, E.; Herrera, D.; Sanz, M. Comparative gene expression analysis of planktonic Porphyromonas gingivalis ATCC 33277 in the presence of a growing biofilm versus planktonic cells. BMC Microbiol. 2019, 19, 58. [Google Scholar] [CrossRef] [Green Version]
- Hajishengallis, G.; Lamont, R.J. Beyond the red complex and into more complexity: The polymicrobial synergy and dysbiosis (PSD) model of periodontal disease etiology. Mol. Oral Microbiol. 2012, 27, 409–419. [Google Scholar] [CrossRef] [Green Version]
- Sanz, M.; Beighton, D.; Curtis, M.A.; Cury, J.A.; Dige, I.; Dommisch, H.; Ellwood, R.; Giacaman, R.A.; Herrera, D.; Herzberg, M.C.; et al. Role of microbial biofilms in the maintenance of oral health and in the development of dental caries and periodontal diseases. Consensus report of group 1 of the Joint EFP/ORCA workshop on the boundaries between caries and periodontal disease. J. Clin. Periodontol. 2017, 44 (Suppl. S18), S5–S11. [Google Scholar] [CrossRef]
- Meyle, J.; Chapple, I. Molecular aspects of the pathogenesis of periodontitis. Periodontology 2000 2015, 69, 7–17. [Google Scholar] [CrossRef]
- Cobb, C.M. Clinical significance of non-surgical periodontal therapy: An evidence-based perspective of scaling and root planing. J. Clin. Periodontol. 2002, 29 (Suppl. S2), 6–16. [Google Scholar] [CrossRef]
- Heitz-Mayfield, L.J.; Trombelli, L.; Heitz, F.; Needleman, I.; Moles, D. A systematic review of the effect of surgical debridement vs non-surgical debridement for the treatment of chronic periodontitis. J. Clin. Periodontol. 2002, 29 (Suppl. S3), 92–102. [Google Scholar] [CrossRef] [PubMed]
- Herrera, D.; Alonso, B.; Leon, R.; Roldan, S.; Sanz, M. Antimicrobial therapy in periodontitis: The use of systemic antimicrobials against the subgingival biofilm. J. Clin. Periodontol. 2008, 35, 45–66. [Google Scholar] [CrossRef] [PubMed]
- Matesanz-Perez, P.; Garcia-Gargallo, M.; Figuero, E.; Bascones-Martinez, A.; Sanz, M.; Herrera, D. A systematic review on the effects of local antimicrobials as adjuncts to subgingival debridement, compared with subgingival debridement alone, in the treatment of chronic periodontitis. J. Clin. Periodontol. 2013, 40, 227–241. [Google Scholar] [CrossRef]
- Dabholkar, C.S.; Shah, M.; Kathariya, R.; Bajaj, M.; Doshi, Y. Comparative Evaluation of Antimicrobial Activity of Pomegranate-Containing Mouthwash Against Oral-Biofilm Forming Organisms: An Invitro Microbial Study. J. Clin. Diagn. Res. 2016, 10, Zc65–Zc69. [Google Scholar] [CrossRef]
- Muller, H.D.; Eick, S.; Moritz, A.; Lussi, A.; Gruber, R. Cytotoxicity and Antimicrobial Activity of Oral Rinses In Vitro. Biomed. Res. Int. 2017, 2017, 4019723. [Google Scholar] [CrossRef] [PubMed]
- van Winkelhoff, A.J.; Herrera, D.; Oteo, A.; Sanz, M. Antimicrobial profiles of periodontal pathogens isolated from periodontitis patients in The Netherlands and Spain. J. Clin. Periodontol. 2005, 32, 893–898. [Google Scholar] [CrossRef]
- Barbieri, R.; Coppo, E.; Marchese, A.; Daglia, M.; Sobarzo-Sanchez, E.; Nabavi, S.F.; Nabavi, S.M. Phytochemicals for human disease: An update on plant-derived compounds antibacterial activity. Microbiol. Res. 2017, 196, 44–68. [Google Scholar] [CrossRef] [PubMed]
- Serrano, J.; Escribano, M.; Roldan, S.; Martin, C.; Herrera, D. Efficacy of adjunctive anti-plaque chemical agents in managing gingivitis: A systematic review and meta-analysis. J. Clin. Periodontol. 2015, 42 (Suppl. S16), S106–S138. [Google Scholar] [CrossRef]
- Van Winkelhoff, A.J. Antibiotics in the treatment of peri-implantitis. Eur. J. Oral. Implantol. 2012, 5, S43–S50. [Google Scholar]
- Van Strydonck, D.A.; Slot, D.E.; Van der Velden, U.; Van der Weijden, F. Effect of a chlorhexidine mouthrinse on plaque, gingival inflammation and staining in gingivitis patients: A systematic review. J. Clin. Periodontol. 2012, 39, 1042–1055. [Google Scholar] [CrossRef]
- Rams, T.E.; Degener, J.E.; van Winkelhoff, A.J. Antibiotic resistance in human peri-implantitis microbiota. Clin. Oral Implant. Res. 2014, 25, 82–90. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Desbois, A.P.; Mearns-Spragg, A.; Smith, V.J. A fatty acid from the diatom Phaeodactylum tricornutum is antibacterial against diverse bacteria including multi-resistant Staphylococcus aureus (MRSA). Mar. Biotechnol. 2009, 11, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Desbois, A.P.; Smith, V.J. Antibacterial free fatty acids: Activities, mechanisms of action and biotechnological potential. Appl. Microbiol. Biotechnol. 2010, 85, 1629–1642. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Desbois, A.P. Potential applications of antimicrobial fatty acids in medicine, agriculture and other industries. Recent Pat. Antiinfect Drug Discov. 2012, 7, 111–122. [Google Scholar] [CrossRef] [PubMed]
- Correia, M.; Michel, V.; Matos, A.A.; Carvalho, P.; Oliveira, M.J.; Ferreira, R.M.; Dillies, M.A.; Huerre, M.; Seruca, R.; Figueiredo, C.; et al. Docosahexaenoic acid inhibits Helicobacter pylori growth in vitro and mice gastric mucosa colonization. PLoS ONE 2012, 7, e35072. [Google Scholar] [CrossRef]
- Correia, M.; Michel, V.; Osorio, H.; El Ghachi, M.; Bonis, M.; Boneca, I.G.; De Reuse, H.; Matos, A.A.; Lenormand, P.; Seruca, R.; et al. Crosstalk between Helicobacter pylori and gastric epithelial cells is impaired by docosahexaenoic acid. PLoS ONE 2013, 8, e60657. [Google Scholar] [CrossRef]
- Desbois, A.P.; Lawlor, K.C. Antibacterial activity of long-chain polyunsaturated fatty acids against Propionibacterium acnes and Staphylococcus aureus. Mar. Drugs 2013, 11, 4544–4557. [Google Scholar] [CrossRef] [Green Version]
- Mil-Homens, D.; Bernardes, N.; Fialho, A.M. The antibacterial properties of docosahexaenoic omega-3 fatty acid against the cystic fibrosis multiresistant pathogen Burkholderia cenocepacia. FEMS Microbiol. Lett. 2012, 328, 61–69. [Google Scholar] [CrossRef] [Green Version]
- Huang, C.B.; Ebersole, J.L. A novel bioactivity of omega-3 polyunsaturated fatty acids and their ester derivatives. Mol. Oral Microbiol. 2010, 25, 75–80. [Google Scholar] [CrossRef]
- Sheu, C.W.; Freese, E. Effects of fatty acids on growth and envelope proteins of Bacillus subtilis. J. Bacteriol. 1972, 111, 516–524. [Google Scholar] [CrossRef] [Green Version]
- Galbraith, H.; Miller, T.B. Effect of long chain fatty acids on bacterial respiration and amino acid uptake. J. Appl. Bacteriol. 1973, 36, 659–675. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.D.; Brown, K.E.; Morse, S.A. Inhibitory action of fatty acids on the growth of Neisseria gonorrhoeae. Infect. Immun. 1977, 17, 303–312. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boyaval, P.; Corre, C.; Dupuis, C.; Roussel, E. Effects of free fatty acids on propionic acid bacteria. Le Lait 1995, 75, 17–29. [Google Scholar] [CrossRef]
- Wieckowski, M.R.; Wojtczak, L. Fatty acid-induced uncoupling of oxidative phosphorylation is partly due to opening of the mitochondrial permeability transition pore. FEBS Lett. 1998, 423, 339–342. [Google Scholar] [CrossRef]
- Shaikh, S.R.; Jolly, C.A.; Chapkin, R.S. n-3 Polyunsaturated fatty acids exert immunomodulatory effects on lymphocytes by targeting plasma membrane molecular organization. Mol. Aspects Med. 2012, 33, 46–54. [Google Scholar] [CrossRef] [Green Version]
- Serhan, C.N. Resolution phase of inflammation: Novel endogenous anti-inflammatory and proresolving lipid mediators and pathways. Annu. Rev. Immunol. 2007, 25, 101–137. [Google Scholar] [CrossRef] [Green Version]
- Sun, M.; Zhou, Z.; Dong, J.; Zhang, J.; Xia, Y.; Shu, R. Antibacterial and antibiofilm activities of docosahexaenoic acid (DHA) and eicosapentaenoic acid (EPA) against periodontopathic bacteria. Microb. Pathog. 2016, 99, 196–203. [Google Scholar] [CrossRef]
- Sanchez, M.C.; Llama-Palacios, A.; Blanc, V.; Leon, R.; Herrera, D.; Sanz, M. Structure, viability and bacterial kinetics of an in vitro biofilm model using six bacteria from the subgingival microbiota. J. Periodontal Res. 2011, 46, 252–260. [Google Scholar] [CrossRef]
- Sanchez, M.C.; Ribeiro-Vidal, H.; Esteban-Fernandez, A.; Bartolome, B.; Figuero, E.; Moreno-Arribas, M.V.; Sanz, M.; Herrera, D. Antimicrobial activity of red wine and oenological extracts against periodontal pathogens in a validated oral biofilm model. BMC Complement. Altern. Med. 2019, 19, 145. [Google Scholar] [CrossRef]
- Sanchez, M.C.; Fernandez, E.; Llama-Palacios, A.; Figuero, E.; Herrera, D.; Sanz, M. Response to antiseptic agents of periodontal pathogens in in vitro biofilms on titanium and zirconium surfaces. Dent. Mater. 2017, 33, 446–453. [Google Scholar] [CrossRef]
- Sanchez, M.C.; Marin, M.J.; Figuero, E.; Llama-Palacios, A.; Leon, R.; Blanc, V.; Herrera, D.; Sanz, M. Quantitative real-time PCR combined with propidium monoazide for the selective quantification of viable periodontal pathogens in an in vitro subgingival biofilm model. J. Periodontal Res. 2014, 49, 20–28. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, M.C.; Llama-Palacios, A.; Fernandez, E.; Figuero, E.; Marin, M.J.; Leon, R.; Blanc, V.; Herrera, D.; Sanz, M. An in vitro biofilm model associated to dental implants: Structural and quantitative analysis of in vitro biofilm formation on different dental implant surfaces. Dent. Mater. 2014, 30, 1161–1171. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, M.C.; Ribeiro-Vidal, H.; Bartolome, B.; Figuero, E.; Moreno-Arribas, M.V.; Sanz, M.; Herrera, D. New Evidences of Antibacterial Effects of Cranberry Against Periodontal Pathogens. Foods 2020, 9, 246. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Amiri-Jami, M.; Lapointe, G.; Griffiths, M.W. Engineering of EPA/DHA omega-3 fatty acid production by Lactococcus lactis subsp. cremoris MG1363. Appl. Microbiol. Biotechnol. 2014, 98, 3071–3080. [Google Scholar] [CrossRef] [PubMed]
- Berge, J.P.; Barnathan, G. Fatty acids from lipids of marine organisms: Molecular biodiversity, roles as biomarkers, biologically active compounds, and economical aspects. Adv. Biochem. Eng. Biotechnol. 2005, 96, 49–125. [Google Scholar]
- Calder, P.C. Marine omega-3 fatty acids and inflammatory processes: Effects, mechanisms and clinical relevance. Biochim. Biophys. Acta 2015, 1851, 469–484. [Google Scholar] [CrossRef]
- Calder, P.C. n-3 fatty acids, inflammation and immunity: New mechanisms to explain old actions. Proc. Nutr. Soc. 2013, 72, 326–336. [Google Scholar] [CrossRef] [Green Version]
- Giordano, E.; Visioli, F. Long-chain omega 3 fatty acids: Molecular bases of potential antioxidant actions. Prostaglandins Leukot. Essent. Fatty Acids 2014, 90, 1–4. [Google Scholar] [CrossRef]
- Choi, J.S.; Park, N.H.; Hwang, S.Y.; Sohn, J.H.; Kwak, I.; Cho, K.K.; Choi, I.S. The antibacterial activity of various saturated and unsaturated fatty acids against several oral pathogens. J. Environ. Biol. 2013, 34, 673–676. [Google Scholar]
- Gilbert, P.; Das, J.; Foley, I. Biofilm susceptibility to antimicrobials. Adv. Dent. Res. 1997, 11, 160–167. [Google Scholar] [CrossRef]
- Costerton, J.W.; Stewart, P.S.; Greenberg, E.P. Bacterial biofilms: A common cause of persistent infections. Science 1999, 284, 1318–1322. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stewart, P.S.; Costerton, J.W. Antibiotic resistance of bacteria in biofilms. Lancet 2001, 358, 135–138. [Google Scholar] [CrossRef]
- Gilbert, P.; Maira-Litran, T.; McBain, A.J.; Rickard, A.H.; Whyte, F.W. The physiology and collective recalcitrance of microbial biofilm communities. Adv. Microb. Physiol. 2002, 46, 202–256. [Google Scholar] [PubMed]
- Socransky, S.S.; Haffajee, A.D. Dental biofilms: Difficult therapeutic targets. Periodontology 2000 2002, 28, 12–55. [Google Scholar] [CrossRef]
- Marsh, P.D.; Moter, A.; Devine, D.A. Dental plaque biofilms: Communities, conflict and control. Periodontology 2000 2011, 55, 16–35. [Google Scholar] [CrossRef] [PubMed]
- Sedlacek, M.J.; Walker, C. Antibiotic resistance in an in vitro subgingival biofilm model. Oral Microbiol. Immunol. 2007, 22, 333–339. [Google Scholar] [CrossRef] [Green Version]
- Hojo, K.; Nagaoka, S.; Ohshima, T.; Maeda, N. Bacterial interactions in dental biofilm development. J. Dent. Res. 2009, 88, 982–990. [Google Scholar] [CrossRef]
- Peng, Y.; Zheng, Y.; Zhang, Y.; Zhao, J.; Chang, F.; Lu, T.; Zhang, R.; Li, Q.; Hu, X.; Li, N. Different effects of omega-3 fatty acids on the cell cycle in C2C12 myoblast proliferation. Mol. Cell. Biochem. 2012, 367, 165–173. [Google Scholar] [CrossRef]
- Al-Radha, A.S.; Younes, C.; Diab, B.S.; Jenkinson, H.F. Essential oils and zirconia dental implant materials. Int. J. Oral Maxillofac. Implant. 2013, 28, 1497–1505. [Google Scholar] [CrossRef]
- Bonesvoll, P.; Lokken, P.; Rolla, G. Influence of concentration, time, temperature and pH on the retention of chlorhexidine in the human oral cavity after mouth rinses. Arch. Oral Biol. 1974, 19, 1025–1029. [Google Scholar] [CrossRef]
- Bonesvoll, P.; Lokken, P.; Rolla, G.; Paus, P.N. Retention of chlorhexidine in the human oral cavity after mouth rinses. Arch. Oral Biol. 1974, 19, 209–212. [Google Scholar] [CrossRef]
- Schiott, C.R.; Loe, H.; Jensen, S.B.; Kilian, M.; Davies, R.M.; Glavind, K. The effect of chlorhexidine mouthrinses on the human oral flora. J. Periodontal Res. 1970, 5, 84–89. [Google Scholar] [CrossRef] [PubMed]


| Treatments | Mean (SD) | Global p | % of Reduction of Viable CFUs mL−1 as Compared with PBS | |
|---|---|---|---|---|
| S. oralis | PBS | 7.80 × 107 (3.46 × 107) | <0.001 | |
| DHA | 3.49 × 104 (4.53 × 103) | 99.96% | ||
| EtOH | 1.99 × 107 (1.23 × 107) | 74.49% | ||
| CHX | 1.39 × 106 (3.81 × 105) | 98.22% | ||
| A. naeslundii | PBS | 3.22 × 106 (1.04 × 106) | <0.001 | |
| DHA | 5.88 × 103 (1.51 × 103) | 99.82% | ||
| EtOH | 1.58 × 106 (9.42 × 105) | 51.04% | ||
| CHX | 3.23 × 105 (1.88 × 105) | 90.07% | ||
| V. parvula | PBS | 4.56 × 107 (9.86 × 106) | <0.001 | |
| DHA | 8.83 × 104 (2.60 × 104) | 99.80% | ||
| EtOH | 1.68 × 107 (5.40 × 106) | 63.16% | ||
| CHX | 8.18 × 106 (9.59 × 106) | 82.07% | ||
| F. nucleatum | PBS | 2.24 × 106 (1.04 × 106) | <0.001 | |
| DHA | 2.01 × 103 (8.68 × 102) | 99.91% | ||
| EtOH | 7.36 × 105 (6.70 × 105) | 67.15% | ||
| CHX | 1.62 × 106 (1.15 × 106) | 27.68% | ||
| P. gingivalis | PBS | 2.32 × 107 (8.31 × 106) | <0.001 | |
| DHA | 1.77 × 104 (2.71 × 103) | 99.92% | ||
| EtOH | 2.37 × 106 (1.20 × 106) | 89.79% | ||
| CHX | 1.82 × 106 (3.13 × 106) | 92.16% | ||
| A. actinomycetemcomitans | PBS | 1.14 × 107 (4.84 × 106) | <0.001 | |
| DHA | 1.12 × 104 (4.71 × 103) | 99.90% | ||
| EtOH | 4.62 × 106 (2.25 × 106) | 59.48% | ||
| CHX | 2.10 × 106 (5.64 × 105) | 81.58% |
| Comparisons | Mean Difference | 95% Confidence Interval for Difference | Post Hoc p | ||
|---|---|---|---|---|---|
| Lower Bound | Upper Bound | ||||
| S. oralis | PBS-DHA | 7.80 × 107 | 5.36 × 107 | 1.02 × 108 | <0.001 |
| PBS-EtOH | 5.81 × 107 | 3.37 × 107 | 8.25 × 107 | <0.001 | |
| PBS-CHX | 7.66 × 107 | 5.23 × 107 | 1.01 × 108 | <0.001 | |
| EtOH-DHA | 1.99 × 107 | −4.47 × 106 | 4.43 × 107 | 0.170 | |
| CHX-DHA | 1.35 × 106 | −2.30 × 107 | 2.57 × 107 | 1.000 | |
| EtOH-CHX | 1.86 × 107 | −5.82 × 106 | 4.29 × 107 | 0.240 | |
| A. naeslundii | PBS-DHA | 3.22 × 106 | 2.28 × 106 | 4.15 × 106 | <0.001 |
| PBS-EtOH | 1.64 × 106 | 7.05 × 105 | 2.58 × 106 | <0.001 | |
| PBS-CHX | 2.90 × 106 | 1.96 × 106 | 3.84 × 106 | <0.001 | |
| EtOH-DHA | 1.57 × 106 | 6.31 × 105 | 2.51 × 106 | <0.001 | |
| CHX-DHA | 3.17 × 105 | −6.23 × 105 | 1.26 × 106 | 1.000 | |
| EtOH-CHX | 1.25 × 106 | 3.14 × 105 | 2.19 × 106 | 0.004 | |
| V. parvula | PBS-DHA | 4.55 × 107 | 3.57 × 107 | 5.53 × 107 | <0.001 |
| PBS-EtOH | 2.88 × 107 | 1.90 × 107 | 3.86 × 107 | <0.001 | |
| PBS-CHX | 3.74 × 107 | 2.76 × 107 | 4.72 × 107 | <0.001 | |
| EtOH-DHA | 1.67 × 107 | 6.89 × 106 | 2.65 × 107 | <0.001 | |
| CHX-DHA | 8.09 × 106 | −1.70 × 106 | 1.79 × 107 | 0.160 | |
| EtOH-CHX | 8.59 × 106 | −1.21 × 106 | 1.84 × 107 | 0.115 | |
| F. nucleatum | PBS-DHA | 2.24 × 106 | 1.12 × 106 | 3.36 × 106 | <0.001 |
| PBS-EtOH | 1.50 × 106 | 3.86 × 105 | 2.62 × 106 | 0.004 | |
| PBS-CHX | 6.15 × 105 | −5.03 × 105 | 1.73 × 106 | 0.789 | |
| EtOH-DHA | 7.34 × 105 | −3.84 × 105 | 1.85 × 106 | 0.445 | |
| CHX-DHA | 1.62 × 106 | 5.05 × 105 | 2.74 × 106 | 0.002 | |
| EtOH-CHX | −8.89 × 105 | −3.84 × 105 | 1.85 × 106 | 0.194 | |
| P. gingivalis | PBS-DHA | 2.32 × 107 | 1.73 × 107 | 2.92 × 107 | <0.001 |
| PBS-EtOH | 2.09 × 107 | 1.49 × 107 | 2.68 × 107 | <0.001 | |
| PBS-CHX | 2.14 × 107 | 1.55 × 107 | 2.73 × 107 | <0.001 | |
| EtOH-DHA | 2.36 × 106 | −3.58 × 106 | 8.29 × 106 | 1.000 | |
| CHX-DHA | 1.81 × 106 | −4.13 × 106 | 7.74 × 106 | 1.000 | |
| EtOH-CHX | 5.50 × 105 | −5.39 × 106 | 6.49 × 106 | 1.000 | |
| A. actinomycetemcomitans | PBS-DHA | 1.14 × 107 | 7.81 × 106 | 1.49 × 107 | <0.001 |
| PBS-EtOH | 6.76 × 106 | 3.20 × 106 | 1.03 × 107 | <0.001 | |
| PBS-CHX | 9.28 × 106 | 5.73 × 106 | 1.28 × 107 | <0.001 | |
| EtOH-DHA | 4.61 × 106 | 1.06 × 106 | 8.17 × 106 | 0.006 | |
| CHX-DHA | 2.08 × 106 | −1.47 × 106 | 5.64 × 106 | 0.654 | |
| EtOH-CHX | 2.53 × 106 | −1.03 × 106 | 6.08 × 106 | 0.325 | |
| Treatments | Mean (SD) | Global p | % of Reduction in Viable CFUs mL−1 as Compared with PBS | ||
|---|---|---|---|---|---|
| S. oralis | PBS | 4.71 × 107 (1.38 × 107) | <0.001 | ||
| EPA | 1.34 × 106 (5.12 × 105) | 97.16% | |||
| EtOH | 9.17 × 106 (2.46 × 106) | 80.54% | |||
| CHX | 1.64 × 106 (7.58 × 105) | 96.52% | |||
| A. naeslundii | PBS | 3.63 × 106 (1.47 × 106) | <0.001 | ||
| EPA | 5.98 × 104 (1.82 × 104) | 98.36% | |||
| EtOH | 6.08 × 105 (1.91 × 105) | 83.25% | |||
| CHX | 3.66 × 105 (1.30 × 105) | 89.92% | |||
| V. parvula | PBS | 6.43 × 107 (1.66 × 107) | <0.001 | ||
| EPA | 2.94 × 106 (8.30 × 105) | 95.43% | |||
| EtOH | 2.10 × 107 (3.14 × 106) | 67.34% | |||
| CHX | 1.80 × 107 (1.08 × 107) | 72.01% | |||
| F. nucleatum | PBS | 2.16 × 106 (9.67 × 105) | <0.001 | ||
| EPA | 2.88 × 104 (1.45 × 104) | 98.67% | |||
| EtOH | 6.66 × 105 (5.88 × 105) | 69.17% | |||
| CHX | 1.24 × 106 (4.58 × 105) | 42.60% | |||
| P. gingivalis | PBS | 1.27 × 107 (1.60 × 106) | <0.001 | ||
| EPA | 3.16 × 105 (1.55 × 105) | 97.51% | |||
| EtOH | 1.92 × 106 (5.04 × 105) | 84.88% | |||
| CHX | 1.63 × 106 (2.18 × 105) | 87.17% | |||
| A. actinomycetemcomitans | PBS | 6.84 × 106 (3.40 × 106) | <0.001 | ||
| EPA | 5.91 × 105 (3.18 × 105) | 91.36% | |||
| EtOH | 2.07 × 106 (7.91 × 105) | 69.74% | |||
| CHX | 2.29 × 106 (1.62 × 106) | 66.52% |
| Comparisons | Mean Difference | 95% Confidence Interval for Difference | Post Hoc p | ||
|---|---|---|---|---|---|
| Lower Bound | Upper Bound | ||||
| S. oralis | PBS-EPA | 4.57 × 107 | 3.64 × 107 | 5.50 × 107 | <0.001 |
| PBS-EtOH | 3.79 × 107 | 2.86 × 107 | 4.72 × 107 | <0.001 | |
| PBS-CHX | 4.54 × 107 | 3.61 × 107 | 5.47 × 107 | <0.001 | |
| EtOH-EPA | 7.83 × 106 | −1.47 × 106 | 1.71 × 107 | 0.144 | |
| CHX-EPA | 2.96 × 105 | −9.00 × 106 | 9.59 × 106 | 1.000 | |
| EtOH-CHX | 7.53 × 106 | −1.76 × 106 | 1.68 × 107 | 0.177 | |
| A. naeslundii | PBS-EPA | 3.57 × 106 | 2.59 × 106 | 4.56 × 106 | <0.001 |
| PBS-EtOH | 3.02 × 106 | 2.04 × 106 | 4.01 × 106 | <0.001 | |
| PBS-CHX | 3.27 × 106 | 2.28 × 106 | 4.25 × 106 | <0.001 | |
| EtOH-EPA | 5.48 × 105 | −4.39 × 105 | 1.54 × 106 | 0.768 | |
| CHX-EPA | 3.06 × 105 | −6.81 × 105 | 1.29 × 106 | 1.000 | |
| EtOH-CHX | 2.42 × 105 | −7.45 × 105 | 1.23 × 106 | 1.000 | |
| V. parvula | PBS-EPA | 6.14 × 107 | 4.81 × 107 | 7.47 × 107 | <0.001 |
| PBS-EtOH | 4.33 × 107 | 3.00 × 107 | 5.66 × 107 | <0.001 | |
| PBS-CHX | 4.63 × 107 | 3.30 × 107 | 5.96 × 107 | <0.001 | |
| EtOH-EPA | 1.81 × 107 | 4.82 × 106 | 3.14 × 107 | 0.003 | |
| CHX-EPA | 1.51 × 107 | 1.81 × 106 | 2.84 × 107 | 0.019 | |
| EtOH-CHX | 3.00 × 106 | −1.03 × 107 | 1.63 × 107 | 1.000 | |
| F. nucleatum | PBS-EPA | 2.13 × 106 | 1.32 × 106 | 2.94 × 106 | <0.001 |
| PBS-EtOH | 1.50 × 106 | 6.88 × 105 | 2.31 × 106 | <0.001 | |
| PBS-CHX | 9.27 × 105 | 1.18 × 105 | 1.74 × 106 | 0.018 | |
| EtOH-EPA | 6.37 × 105 | −1.72 × 105 | 1.45 × 106 | 0.205 | |
| CHX-EPA | 1.21 × 106 | 3.98 × 105 | 2.02 × 106 | 0.001 | |
| EtOH-CHX | −5.70 × 105 | −1.38 × 106 | 2.39 × 105 | 0.336 | |
| P. gingivalis | PBS-EPA | 1.23 × 107 | 1.12 × 107 | 1.35 × 107 | <0.001 |
| PBS-EtOH | 1.07 × 107 | 9.61 × 106 | 1.19 × 107 | <0.001 | |
| PBS-CHX | 1.10 × 107 | 9.90 × 106 | 1.21 × 107 | <0.001 | |
| EtOH-EPA | 1.60 × 106 | 4.77 × 105 | 2.73 × 106 | 0.002 | |
| CHX-EPA | 1.32 × 106 | 1.93 × 105 | 2.44 × 106 | 0.014 | |
| EtOH-CHX | 2.84 × 105 | −8.41 × 105 | 1.41 × 106 | 1.000 | |
| A. actinomycetemcomitans | PBS-EPA | 6.25 × 106 | 3.70 × 106 | 8.81 × 106 | <0.001 |
| PBS-EtOH | 4.77 × 106 | 2.21 × 106 | 7.33 × 106 | <0.001 | |
| PBS-CHX | 4.55 × 106 | 2.00 × 106 | 7.11 × 106 | <0.001 | |
| EtOH-EPA | 1.48 × 106 | −1.07 × 106 | 4.04 × 106 | 0.676 | |
| CHX-EPA | 1.70 × 106 | −8.55 × 105 | 4.26 × 106 | 0.423 | |
| EtOH-CHX | −2.18 × 105 | −2.77 × 106 | 2.34 × 106 | 1.000 | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ribeiro-Vidal, H.; Sánchez, M.C.; Alonso-Español, A.; Figuero, E.; Ciudad, M.J.; Collado, L.; Herrera, D.; Sanz, M. Antimicrobial Activity of EPA and DHA against Oral Pathogenic Bacteria Using an In Vitro Multi-Species Subgingival Biofilm Model. Nutrients 2020, 12, 2812. https://doi.org/10.3390/nu12092812
Ribeiro-Vidal H, Sánchez MC, Alonso-Español A, Figuero E, Ciudad MJ, Collado L, Herrera D, Sanz M. Antimicrobial Activity of EPA and DHA against Oral Pathogenic Bacteria Using an In Vitro Multi-Species Subgingival Biofilm Model. Nutrients. 2020; 12(9):2812. https://doi.org/10.3390/nu12092812
Chicago/Turabian StyleRibeiro-Vidal, Honorato, María Carmen Sánchez, Andrea Alonso-Español, Elena Figuero, Maria José Ciudad, Luís Collado, David Herrera, and Mariano Sanz. 2020. "Antimicrobial Activity of EPA and DHA against Oral Pathogenic Bacteria Using an In Vitro Multi-Species Subgingival Biofilm Model" Nutrients 12, no. 9: 2812. https://doi.org/10.3390/nu12092812
APA StyleRibeiro-Vidal, H., Sánchez, M. C., Alonso-Español, A., Figuero, E., Ciudad, M. J., Collado, L., Herrera, D., & Sanz, M. (2020). Antimicrobial Activity of EPA and DHA against Oral Pathogenic Bacteria Using an In Vitro Multi-Species Subgingival Biofilm Model. Nutrients, 12(9), 2812. https://doi.org/10.3390/nu12092812








