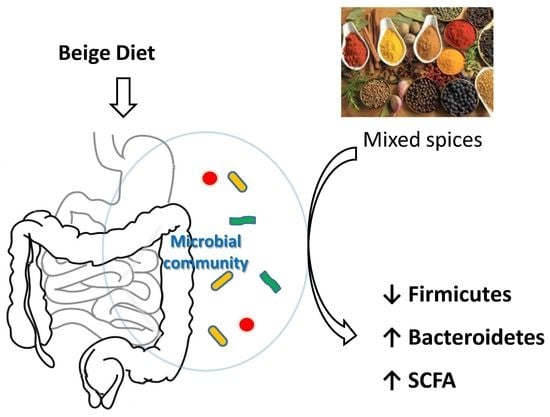
Mixed Spices at Culinary Doses Have Prebiotic Effects in Healthy Adults: A Pilot Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design and Spice Intervention
2.2. Sample Collection
2.3. Determination of Fecal SCFA and Urinary Rosmarinic Acid
2.4. DNA Extraction, 16S rRNA Sequencing, and Taxonomic Assignment
2.5. Identification of Changes in OTU Abundance Associated with Intervention
2.6. Statistical Analysis
3. Results
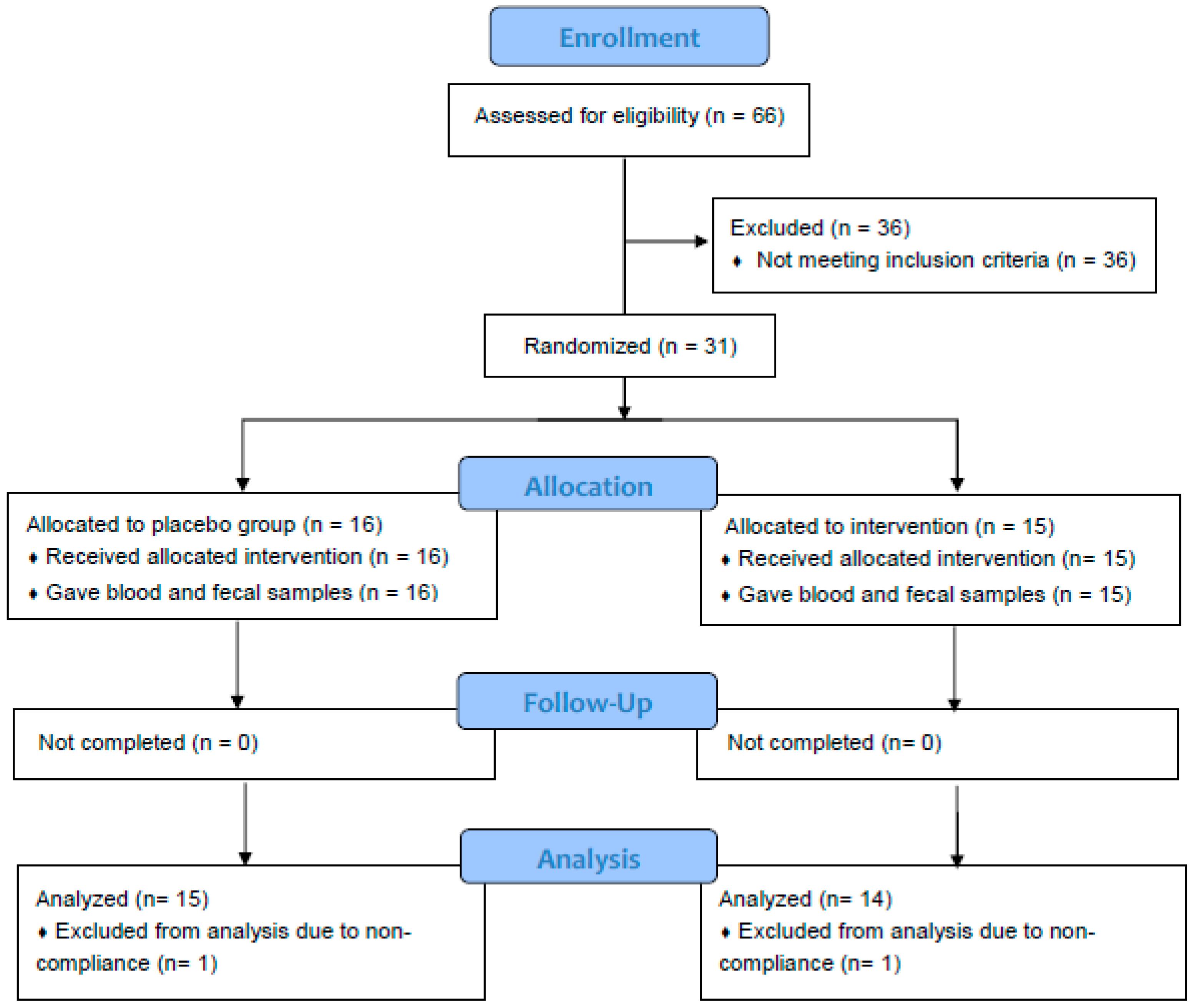
3.1. Subjects
3.2. Body Mass Index, Body Composition, Wellness Score, and Bristol Stool Scores
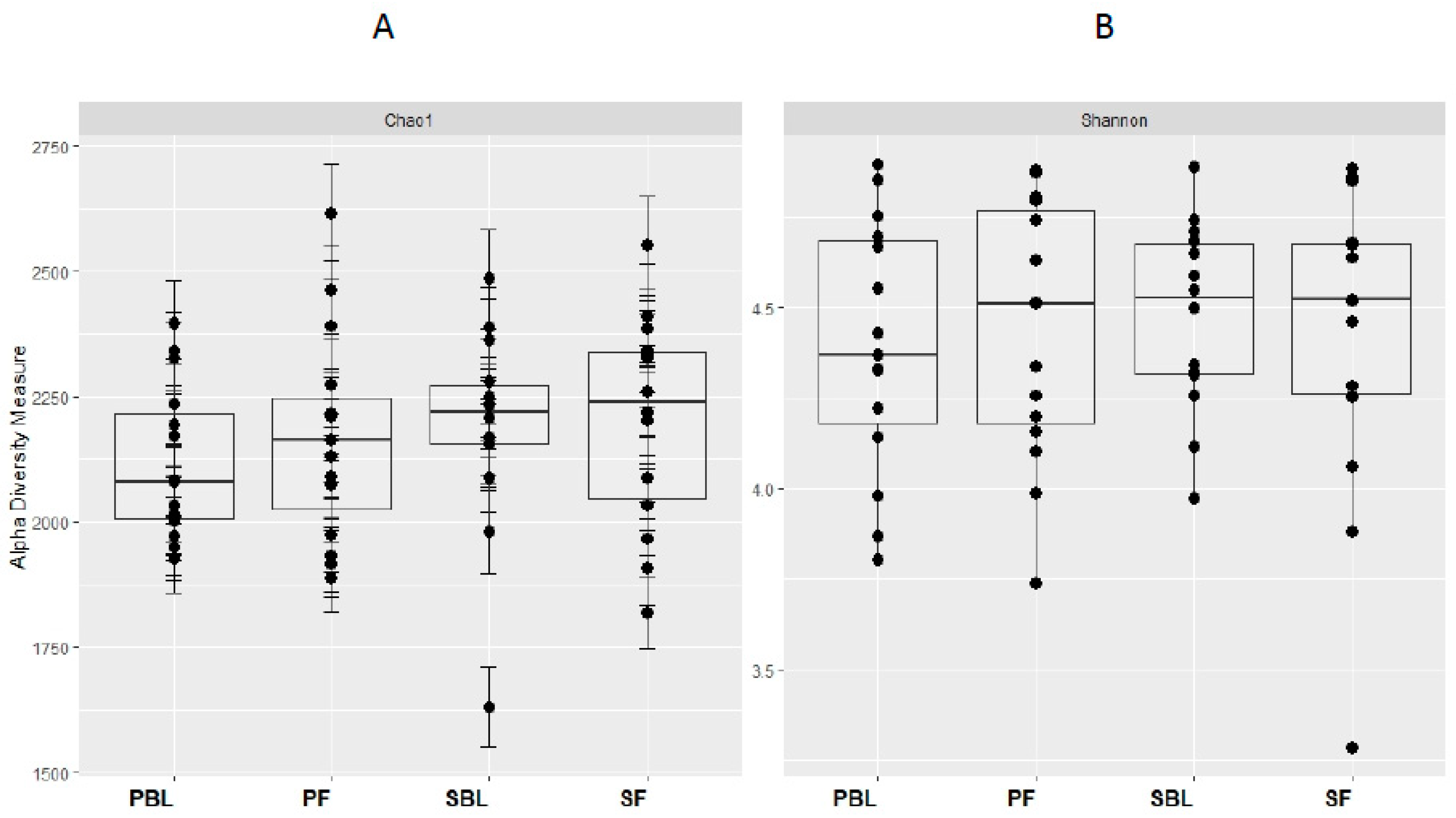
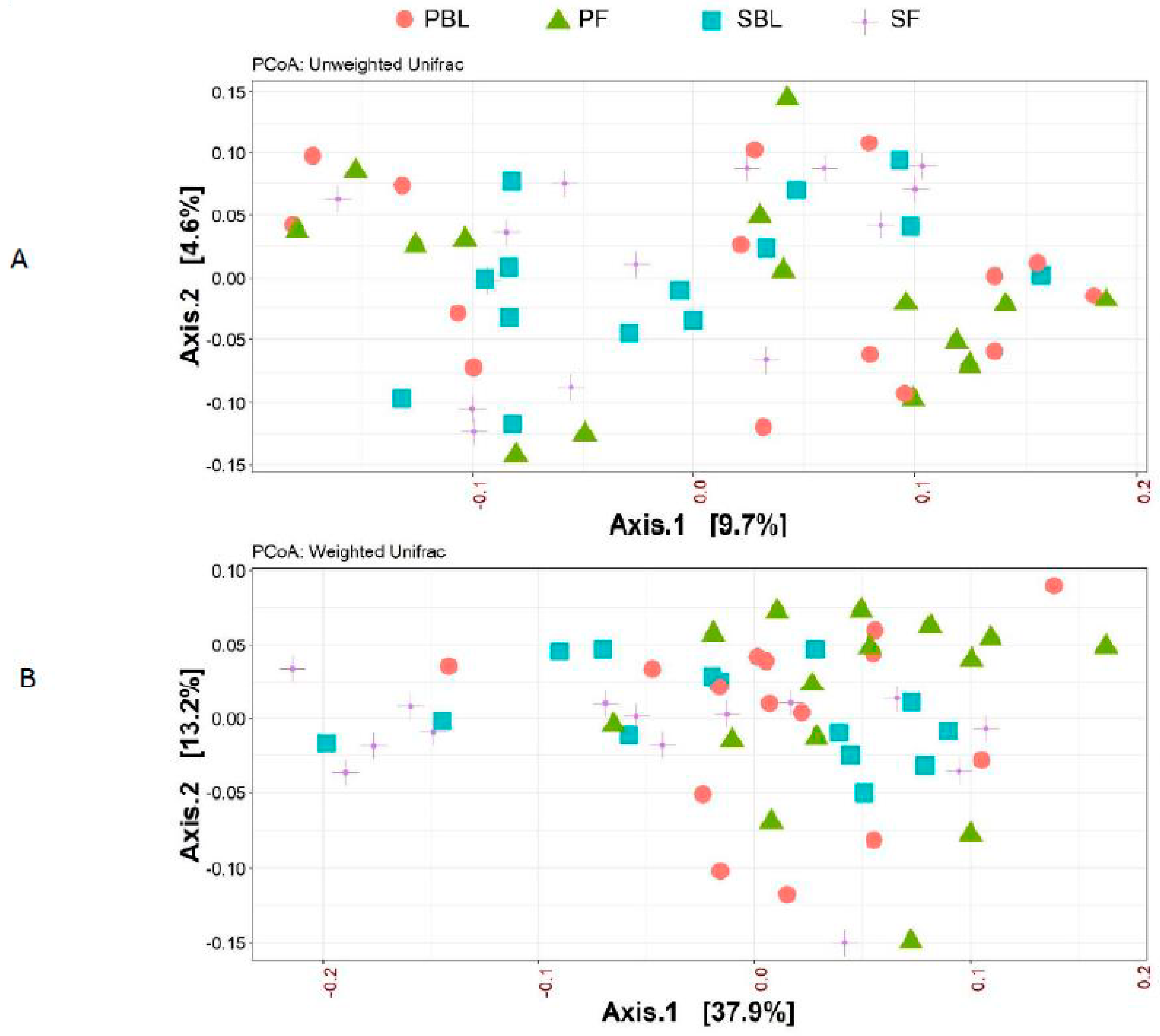
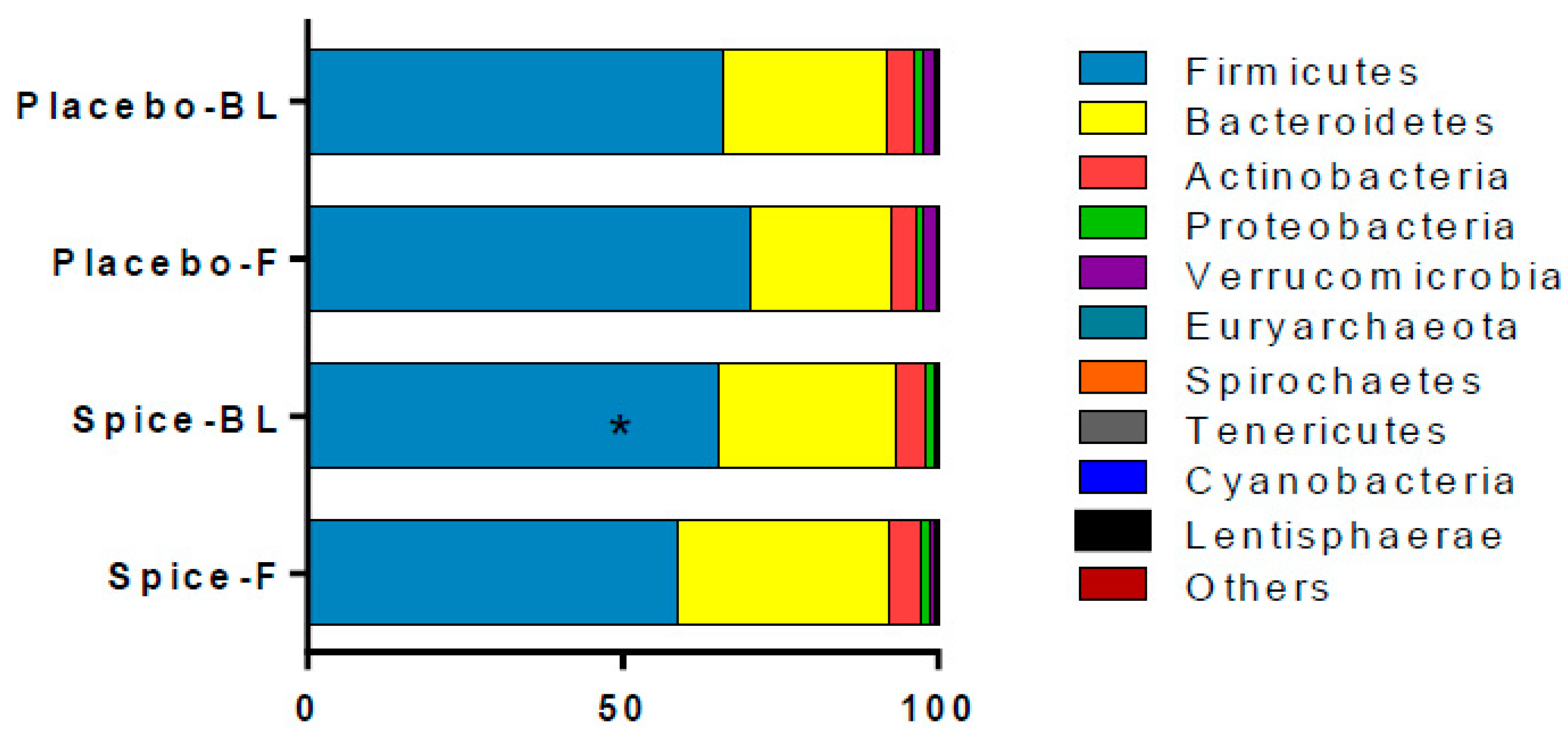
3.3. Microbial Composition
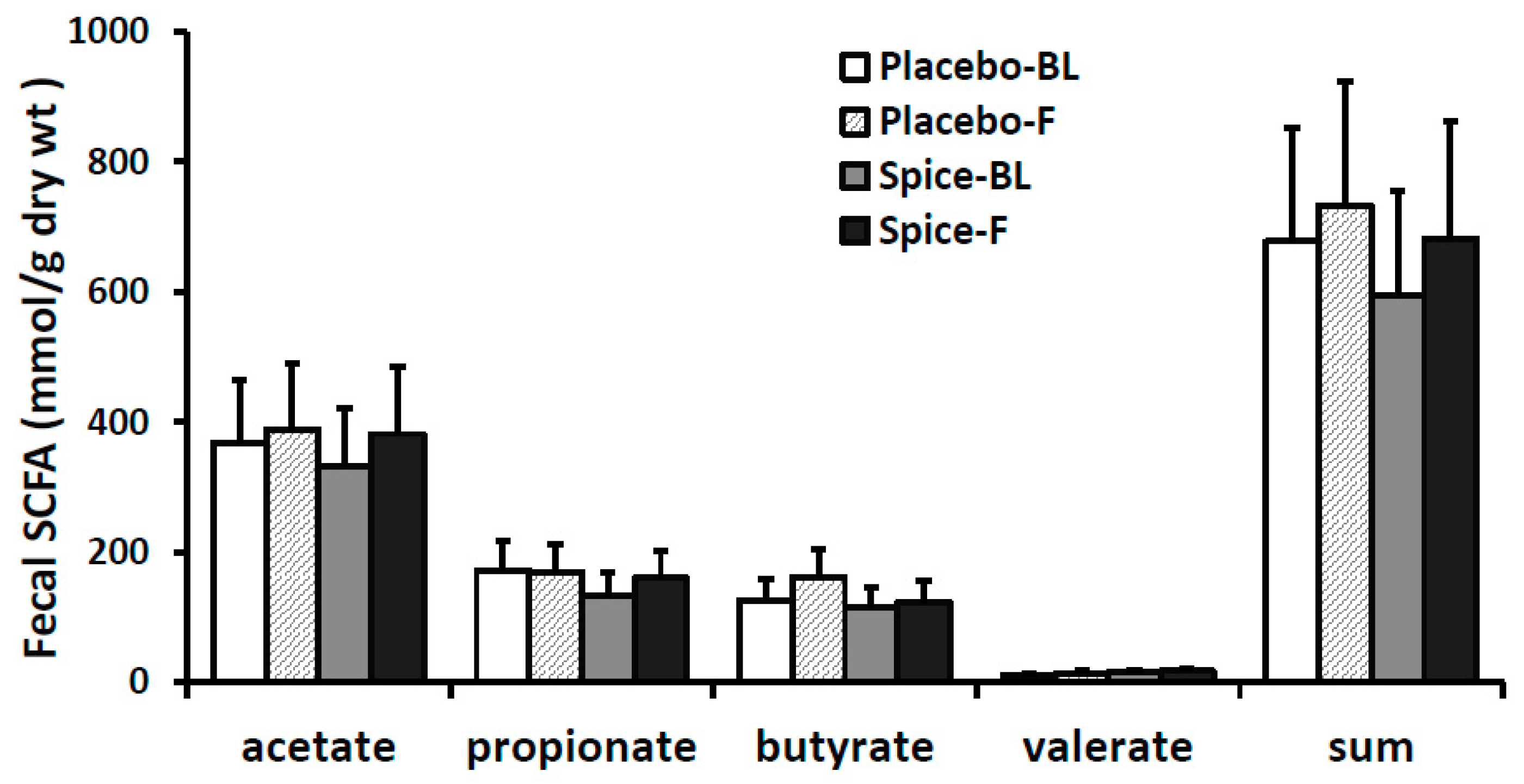
3.4. Fecal Short Chain Fatty Acids and Correlation Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Wu, G.D.; Chen, J.; Hoffmann, C.; Bittinger, K.; Chen, Y.Y.; Keilbaugh, S.A.; Bewtra, M.; Knights, D.; Walters, W.A.; Knight, R.; et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 2011, 334, 105–108. [Google Scholar] [CrossRef]
- David, L.A.; Materna, A.C.; Friedman, J.; Campos-Baptista, M.I.; Blackburn, M.C.; Perrotta, A.; Erdman, S.E.; Alm, E.J. Host lifestyle affects human microbiota on daily timescales. Genome Biol. 2014, 15, R89. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.K.; Chang, H.W.; Yan, D.; Lee, K.M.; Ucmak, D.; Wong, K.; Abrouk, M.; Farahnik, B.; Nakamura, M.; Zhu, T.H.; et al. Influence of diet on the gut microbiome and implications for human health. J. Transl. Med. 2017, 15, 73. [Google Scholar] [CrossRef]
- Hsiao, E.Y.; McBride, S.W.; Hsien, S.; Sharon, G.; Hyde, E.R.; McCue, T.; Codelli, J.A.; Chow, J.; Reisman, S.E.; Petrosino, J.F.; et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 2013, 155, 1451–1463. [Google Scholar] [CrossRef] [PubMed]
- Mangiola, F.; Ianiro, G.; Franceschi, F.; Fagiuoli, S.; Gasbarrini, G.; Gasbarrini, A. Gut microbiota in autism and mood disorders. World J. Gastroenterol. 2016, 22, 361–368. [Google Scholar] [CrossRef] [PubMed]
- Ley, R.E.; Turnbaugh, P.J.; Klein, S.; Gordon, J.I. Microbial ecology—Human gut microbes associated with obesity. Nature 2006, 444, 1022–1023. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Klipfell, E.; Bennett, B.J.; Koeth, R.; Levison, B.S.; Dugar, B.; Feldstein, A.E.; Britt, E.B.; Fu, X.; Chung, Y.M.; et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 2011, 472, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Qin, J.; Li, Y.; Cai, Z.; Li, S.; Zhu, J.; Zhang, F.; Liang, S.; Zhang, W.; Guan, Y.; Shen, D.; et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 2012, 490, 55–60. [Google Scholar] [CrossRef]
- Tzounis, X.; Rodriguez-Mateos, A.; Vulevic, J.; Gibson, G.R.; Kwik-Uribe, C.; Spencer, J.P. Prebiotic evaluation of cocoa-derived flavanols in healthy humans by using a randomized, controlled, double-blind, crossover intervention study. Am. J. Clin. Nutr. 2011, 93, 62–72. [Google Scholar] [CrossRef]
- Guglielmetti, S.; Fracassetti, D.; Taverniti, V.; Del Bo, C.; Vendrame, S.; Klimis-Zacas, D.; Arioli, S.; Riso, P.; Porrini, M. Differential modulation of human intestinal bifidobacterium populations after consumption of a wild blueberry (Vaccinium angustifolium) drink. J. Agric. Food Chem. 2013, 61, 8134–8140. [Google Scholar] [CrossRef]
- Lu, Q.Y.; Summanen, P.H.; Lee, R.P.; Huang, J.; Henning, S.M.; Heber, D.; Finegold, S.M.; Li, Z. Prebiotic Potential and Chemical Composition of Seven Culinary Spice Extracts. J. Food Sci. 2017, 82, 1807–1813. [Google Scholar] [CrossRef] [PubMed]
- London, S.J.; Daly, A.K.; Cooper, J.; Navidi, W.C.; Carpenter, C.L.; Idle, J.R. Polymorphism of glutathione S-transferase M1 and lung cancer risk among African-Americans and Caucasians in Los Angeles County, California. J. Natl. Cancer Inst. 1995, 87, 1246–1253. [Google Scholar] [CrossRef] [PubMed]
- Henning, S.M.; Yang, J.P.; Grojean, E.M.; Lee, R.P.; Hsu, M.; Heber, D.; Li, Z.L. Effect of decaffeinated green and black tea extracts on energy metabolism in mice fed a high fat/high sucrose/western diet. FASEB J. 2016, 30 (Suppl. 1), 428.4. [Google Scholar]
- Baba, S.; Osakabe, N.; Natsume, M.; Yasuda, A.; Muto, Y.; Hiyoshi, K.; Takano, H.; Yoshikawa, T.; Terao, J. Absorption, metabolism, degradation and urinary excretion of rosmarinic acid after intake of Perilla frutescens extract in humans. Eur. J. Nutr. 2005, 44, 1–9. [Google Scholar] [CrossRef] [PubMed]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- McMurdie, P.J.; Holmes, S. Phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE 2013, 8, 61217. [Google Scholar] [CrossRef]
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. R Package. Version 2.5-4. Available online: https://CRAN.R-project.org/package=vegan (accessed on 25 June 2019).
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Storey, J.D.; Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 2003, 100, 9440–9445. [Google Scholar] [CrossRef]
- Remely, M.; Hippe, B.; Zanner, J.; Aumueller, E.; Brath, H.; Haslberger, A.G. Gut Microbiota of Obese, Type 2 Diabetic Individuals is Enriched in Faecalibacterium prausnitzii, Akkermansia muciniphila and Peptostreptococcus anaerobius after Weight Loss. Endocr. Metab. Immune 2016, 16, 99–106. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef]
- Cani, P.D.; Bibiloni, R.; Knauf, C.; Neyrinck, A.M.; Neyrinck, A.M.; Delzenne, N.M.; Burcelin, R. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes 2008, 57, 1470–1481. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Rottier, O.; Goiot, Y.; Neyrinck, A.; Geurts, L.; Delzenne, N. Changes in gut microbiota control intestinal permeability-induced inflammation in obese and diabetic mice through unexpected dependent mechanisms. Diabetologia 2008, 51, S34–S35. [Google Scholar]
- Arboleya, S.; Watkins, C.; Stanton, C.; Ross, R.P. Gut Bifidobacteria Populations in Human Health and Aging. Front. Microbiol. 2016, 7, 1204. [Google Scholar] [CrossRef] [PubMed]
- Million, M.; Angelakis, E.; Maraninchi, M.; Henry, M.; Giorgi, R.; Valero, R.; Vialettes, B.; Raoult, D. Correlation between body mass index and gut concentrations of Lactobacillus reuteri, Bifidobacterium animalis, Methanobrevibacter smithii and Escherichia coli. Int. J. Obes. (Lond.) 2013, 37, 1460–1466. [Google Scholar] [CrossRef] [PubMed]
- Cuervo, A.; Valdes, L.; Salazar, N.; de los Reyes-Gavilan, C.G.; Ruas-Madiedo, P.; Gueimonde, M.; Gonzalez, S. Pilot study of diet and microbiota: Interactive associations of fibers and polyphenols with human intestinal bacteria. J. Agric. Food Chem. 2014, 62, 5330–5336. [Google Scholar] [CrossRef] [PubMed]
- Vendrame, S.; Guglielmetti, S.; Riso, P.; Arioli, S.; Klimis-Zacas, D.; Porrini, M. Six-week consumption of a wild blueberry powder drink increases bifidobacteria in the human gut. J. Agric. Food Chem. 2011, 59, 12815–12820. [Google Scholar] [CrossRef]
- He, X.; Slupsky, C.M.; Dekker, J.W.; Haggarty, N.W.; Lonnerdal, B. Integrated Role of Bifidobacterium animalis subsp. lactis Supplementation in Gut Microbiota, Immunity, and Metabolism of Infant Rhesus Monkeys. Msystems 2016, 1, e00128-16. [Google Scholar] [PubMed]
- Wescombe, P.A.; Hale, J.D.F.; Heng, N.C.K.; Tagg, J.R. Developing oral probiotics from Streptococcus salivarius. Future Microbiol. 2012, 7, 1355–1371. [Google Scholar] [CrossRef]
- Qin, J.; Li, R.; Raes, J.; Arumugam, M.; Burgdorf, K.S.; Manichanh, C.; Nielsen, T.; Pons, N.; Levenez, F.; Yamada, T.; et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 2010, 464, 59–65. [Google Scholar] [CrossRef]
- Flint, H.J.; Bayer, E.A.; Rincon, M.T.; Lamed, R.; White, B.A. Polysaccharide utilization by gut bacteria: Potential for new insights from genomic analysis. Nat. Rev. Microbiol. 2008, 6, 121–131. [Google Scholar] [CrossRef]
- Ze, X.; Duncan, S.H.; Louis, P.; Flint, H.J. Ruminococcus bromii is a keystone species for the degradation of resistant starch in the human colon. ISME J. 2012, 6, 1535–1543. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.K.; Mehrabian, P.; Boyajian, S.; Wu, W.L.; Selicha, J.; Vonderfecht, S.; Mazmanian, S.K. The Protective Role of Bacteroides fragilis in a Murine Model of Colitis-Associated Colorectal Cancer. MSphere 2018, 3, e00587-18. [Google Scholar] [CrossRef] [PubMed]
- Salyers, A.A.; Vercellotti, J.R.; West, S.E.H.; Wilkins, T.D. Fermentation of Mucin and Plant Polysaccharides by Strains of Bacteroides from Human Colon. Appl. Environ. Microb. 1977, 33, 319–322. [Google Scholar]
- Wexler, A.G.; Goodman, A.L. An insider’s perspective: Bacteroides as a window into the microbiome. Nat. Microbiol. 2017, 2, 17026. [Google Scholar] [CrossRef] [PubMed]
- Mazmanian, S.K.; Round, J.L.; Kasper, D.L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 2008, 453, 620–625. [Google Scholar] [CrossRef]
- McNabney, S.M.; Henagan, T.M. Short Chain Fatty Acids in the Colon and Peripheral Tissues: A Focus on Butyrate, Colon Cancer, Obesity and Insulin Resistance. Nutrients 2017, 9, 1348. [Google Scholar] [CrossRef] [PubMed]
- Canfora, E.E.; Jocken, J.W.; Blaak, E.E. Short-chain fatty acids in control of body weight and insulin sensitivity. Nat. Rev. Endocrinol. 2015, 11, 577–591. [Google Scholar] [CrossRef]
- Louis, P.; Flint, H.J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 2017, 19, 29–41. [Google Scholar] [CrossRef]
- Wolever, T.M.; Spadafora, P.; Eshuis, H. Interaction between colonic acetate and propionate in humans. Am. J. Clin. Nutr. 1991, 53, 681–687. [Google Scholar] [CrossRef]
- Louis, P.; Flint, H.J. Diversity, metabolism and microbial ecology of butyrate-producing bacteria from the human large intestine. FEMS Microbiol. Lett. 2009, 294, 1–8. [Google Scholar] [CrossRef]
- Morrison, D.J.; Preston, T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 2016, 7, 189–200. [Google Scholar] [CrossRef] [PubMed]
- McDonald, J.A.K.; Mullish, B.H.; Pechlivanis, A.; Liu, Z.G.; Brignardello, J.; Kao, D.; Holmes, E.; Li, J.V.; Clarke, T.B.; Thursz, M.R.; et al. Inhibiting Growth of Clostridioides difficile by Restoring Valerate, Produced by the Intestinal Microbiota. Gastroenterology 2018, 155, 1495–1507. [Google Scholar] [CrossRef] [PubMed]
- Kang, C.; Zhang, Y.; Zhu, X.H.; Liu, K.; Wang, X.L.; Chen, M.T.; Wang, J.; Chen, H.; Hui, S.C.; Huang, L.; et al. Healthy Subjects Differentially Respond to Dietary Capsaicin Correlating with Specific Gut Enterotypes. J. Clin. Endocr. Metab. 2016, 101, 4681–4689. [Google Scholar] [CrossRef] [PubMed]
- Van Hul, M.; Geurts, L.; Plovier, H.; Druart, C.; Everard, A.; Stahlman, M.; Rhimi, M.; Chira, K.; Teissedre, P.L.; Delzenne, N.M.; et al. Reduced obesity, diabetes, and steatosis upon cinnamon and grape pomace are associated with changes in gut microbiota and markers of gut barrier. Am. J. Physiol. Endocrinol. Metab. 2018, 314, E334–E352. [Google Scholar] [CrossRef] [PubMed]
- Khare, P.; Jagtap, S.; Jain, Y.; Baboota, R.K.; Mangal, P.; Boparai, R.K.; Bhutani, K.K.; Sharma, S.S.; Premkumar, L.S.; Kondepudi, K.K.; et al. Cinnamaldehyde supplementation prevents fasting-induced hyperphagia, lipid accumulation, and inflammation in high-fat diet-fed mice. Biofactors 2016, 42, 201–211. [Google Scholar] [PubMed]
- Ezzat, S.M.; Ezzat, M.I.; Okba, M.M.; Menze, E.T.; Abdel-Naim, A.B. The hidden mechanism beyond ginger (Zingiber officinale Rosc.) potent in vivo and in vitro anti-inflammatory activity. J. Ethnopharmacol. 2018, 214, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Zhou, Y.; Li, Y.; Xu, D.P.; Li, S.; Li, H.B. Spices for Prevention and Treatment of Cancers. Nutrients 2016, 8, 495. [Google Scholar] [CrossRef]
- Rastogi, S.; Pandey, M.M.; Rawat, A.K.S. Spices: Therapeutic Potential in Cardiovascular Health. Curr. Pharm. Des. 2017, 23, 989–998. [Google Scholar] [CrossRef]
- Finegold, S.M.; Li, Z.; Summanen, P.H.; Downes, J.; Thames, G.; Corbett, K.; Dowd, S.; Krak, M.; Heber, D. Xylooligosaccharide increases bifidobacteria but not lactobacilli in human gut microbiota. Food Funct. 2014, 5, 436–445. [Google Scholar] [CrossRef]
| Placebo Group | Spice Group | p-Value | |
|---|---|---|---|
| Enrolled/randomized | 15 | 14 | |
| Age, years | 36.7 ± 13.3 | 34.4 ± 12.5 | 0.63 a |
| Sex | |||
| Male (N) | 4 | 4 | |
| Female (N) | 11 | 10 | 0.91 b |
| Race | |||
| African American (N) | 1 | 3 | |
| White (N) | 9 | 6 | |
| Asian (N) | 3 | 3 | |
| Multi-Racial (N) | 2 | 2 | |
| Other | 0 | 1 | 0.57 |
| Weight, kg | 75.6 ± 16.9 | 76.7 ± 16.4 | 0.95 |
| BMI, kg/m2 | 26.9 ± 4.5 | 28.2 ± 7.0 | 0.54 |
| Body fat c (%) | 30.0 ± 7.9 | 32.0 ± 12.5 | 0.63 |
| Phylum/OTU No. | Class, Order, Family, Genus, and Species | log2 FC | p | padj |
|---|---|---|---|---|
| Actinobacteria | ||||
| 4426298 | Actinobacteria; Bifidobacteriales; Bifidobacteriaceae; Bifidobacterium; animalis | 4.578 | 0.004 | 0.496 |
| 681370 | Actinobacteria; Bifidobacteriales; Bifidobacteriaceae; Bifidobacterium; pseudolongum | 1.890 | 0.002 | 0.442 |
| Bacteroidetes | ||||
| 583656 | Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; | 3.675 | 0.000 | 0.037 |
| 175535 | Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; | 3.138 | 0.014 | 0.839 |
| 344525 | Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; eggerthii | 3.045 | 0.013 | 0.839 |
| 351231 | Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; fragilis | 2.949 | 0.004 | 0.498 |
| Firmicutes | ||||
| 679245 | Bacilli; Lactobacillales; Lactobacillaceae; Lactobacillus; | 6.297 | 0.002 | 0.442 |
| 1033473 | Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; | 3.138 | 0.002 | 0.442 |
| 579608 | Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; | 2.899 | 0.002 | 0.442 |
| 828043 | Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; | 2.865 | 0.003 | 0.496 |
| 780650 | Clostridia; Clostridiales; Clostridiaceae; | 2.175 | 0.021 | 0.988 |
| 558420 | Clostridia; Clostridiales; Clostridiaceae; Clostridium; | −4.226 | 0.005 | 0.530 |
| 323778 | Clostridia; Clostridiales; Lachnospiraceae; | −3.353 | 0.014 | 0.839 |
| 176450 | Clostridia; Clostridiales; Lachnospiraceae; Blautia; | 2.720 | 0.007 | 0.655 |
| 589313 | Clostridia; Clostridiales; Lachnospiraceae; Blautia; producta | −4.178 | 0.005 | 0.568 |
| 297182 | Clostridia; Clostridiales; Lachnospiraceae; Dorea; | 3.264 | 0.020 | 0.988 |
| 647215 | Clostridia; Clostridiales; Ruminococcaceae; Oscillospira; | −3.902 | 0.012 | 0.839 |
| 359950 | Clostridia; Clostridiales; Ruminococcaceae; Ruminococcus; | 7.302 | 0.000 | 0.002 |
| 193336 | Clostridia; Clostridiales; Ruminococcaceae; Ruminococcus; | 3.631 | 0.009 | 0.777 |
| 2979308 | Clostridia; Clostridiales; Ruminococcaceae; Ruminococcus; | 2.520 | 0.009 | 0.777 |
| 173744 | Clostridia; Clostridiales; Veillonellaceae; Megasphaera; | 2.129 | 0.014 | 0.839 |
| 587530 | Erysipelotrichi; Erysipelotrichales; Erysipelotrichaceae; Eubacterium; dolichum | 2.819 | 0.015 | 0.839 |
| Proteobacteria | ||||
| 4435655 | Alphaproteobacteria;RF32; | 3.483 | 0.010 | 0.778 |
| 211720 | Alphaproteobacteria;RF32; | 6.491 | 0.016 | 0.839 |
| 636296 | Betaproteobacteria; Burkholderiales; Alcaligenaceae; Sutterella; | −3.958 | 0.016 | 0.839 |
| 865469 | Gammaproteobacteria; Pasteurellales; Pasteurellaceae; Haemophilus; parainfluenzae | 4.238 | 0.021 | 0.988 |
| Acetate | Propionate | Butyrate | Sum | |||||
|---|---|---|---|---|---|---|---|---|
| R | p | R | p | R | p | R | p | |
| Firmicutes | −0.254 | 0.27 | −0.391 | 0.04 | −0.210 | 0.28 | −0.290 | 0.13 |
| Bacteroidetes | 0.287 | 0.14 | 0.435 | 0.07 | 0.216 | 0.27 | −0.083 | 0.27 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, Q.-Y.; Rasmussen, A.M.; Yang, J.; Lee, R.-P.; Huang, J.; Shao, P.; Carpenter, C.L.; Gilbuena, I.; Thames, G.; Henning, S.M.; et al. Mixed Spices at Culinary Doses Have Prebiotic Effects in Healthy Adults: A Pilot Study. Nutrients 2019, 11, 1425. https://doi.org/10.3390/nu11061425
Lu Q-Y, Rasmussen AM, Yang J, Lee R-P, Huang J, Shao P, Carpenter CL, Gilbuena I, Thames G, Henning SM, et al. Mixed Spices at Culinary Doses Have Prebiotic Effects in Healthy Adults: A Pilot Study. Nutrients. 2019; 11(6):1425. https://doi.org/10.3390/nu11061425
Chicago/Turabian StyleLu, Qing-Yi, Anna M. Rasmussen, Jieping Yang, Ru-Po Lee, Jianjun Huang, Paul Shao, Catherine L. Carpenter, Irene Gilbuena, Gail Thames, Susanne M. Henning, and et al. 2019. "Mixed Spices at Culinary Doses Have Prebiotic Effects in Healthy Adults: A Pilot Study" Nutrients 11, no. 6: 1425. https://doi.org/10.3390/nu11061425
APA StyleLu, Q.-Y., Rasmussen, A. M., Yang, J., Lee, R.-P., Huang, J., Shao, P., Carpenter, C. L., Gilbuena, I., Thames, G., Henning, S. M., Heber, D., & Li, Z. (2019). Mixed Spices at Culinary Doses Have Prebiotic Effects in Healthy Adults: A Pilot Study. Nutrients, 11(6), 1425. https://doi.org/10.3390/nu11061425