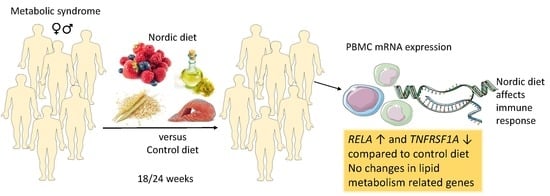
An Isocaloric Nordic Diet Modulates RELA and TNFRSF1A Gene Expression in Peripheral Blood Mononuclear Cells in Individuals with Metabolic Syndrome—A SYSDIET Sub-Study
Abstract
1. Introduction
2. Materials and Methods
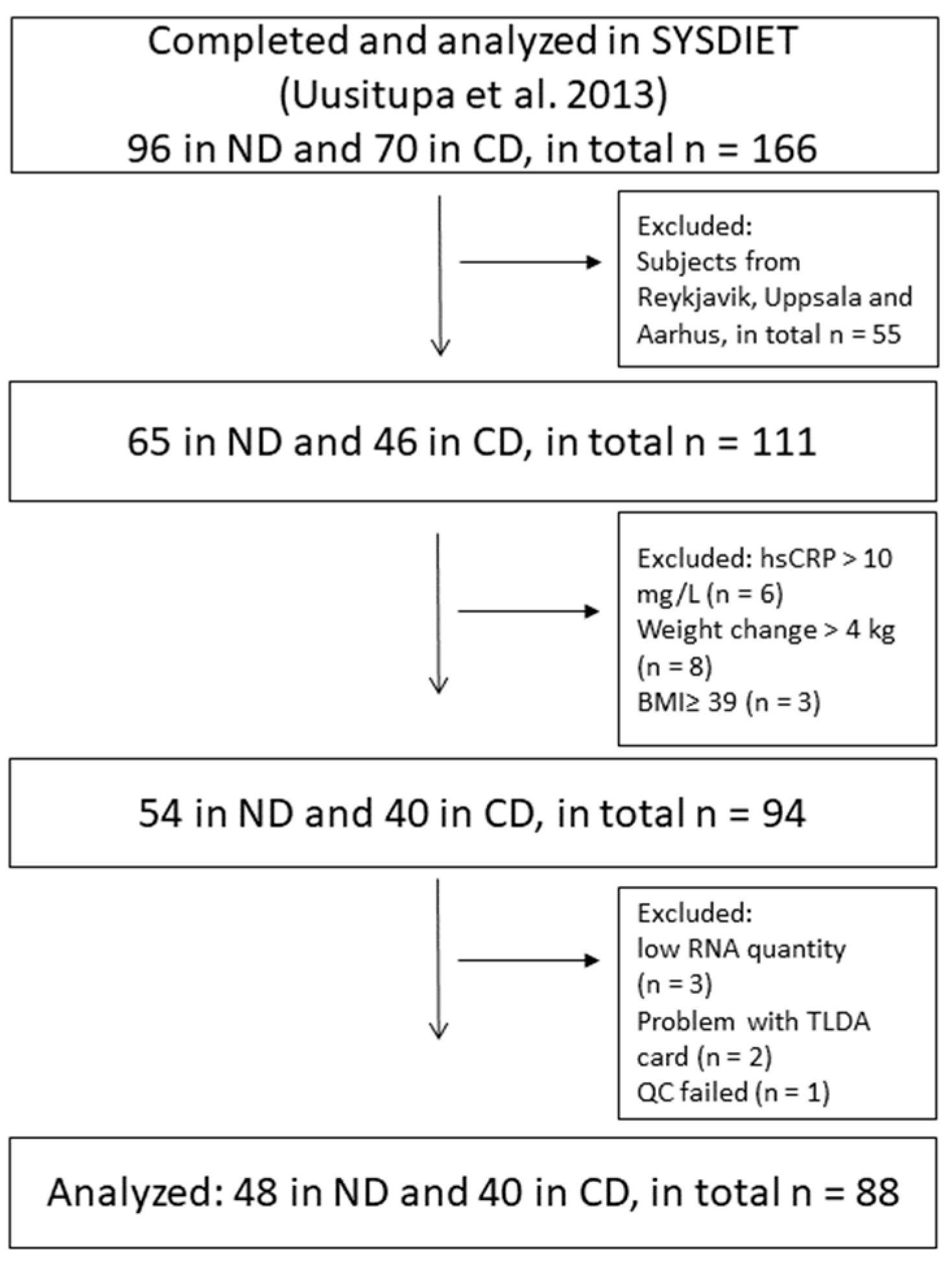
2.1. The SYSDIET Study
2.2. Diet
2.3. Biochemical Measurements
2.4. Sampling of PBMCs and RNA Extraction
2.5. Real-Time Polymerase Chain Reaction RT-qPCR
2.6. Statistical Analysis
3. Results
3.1. Baseline Characteristics
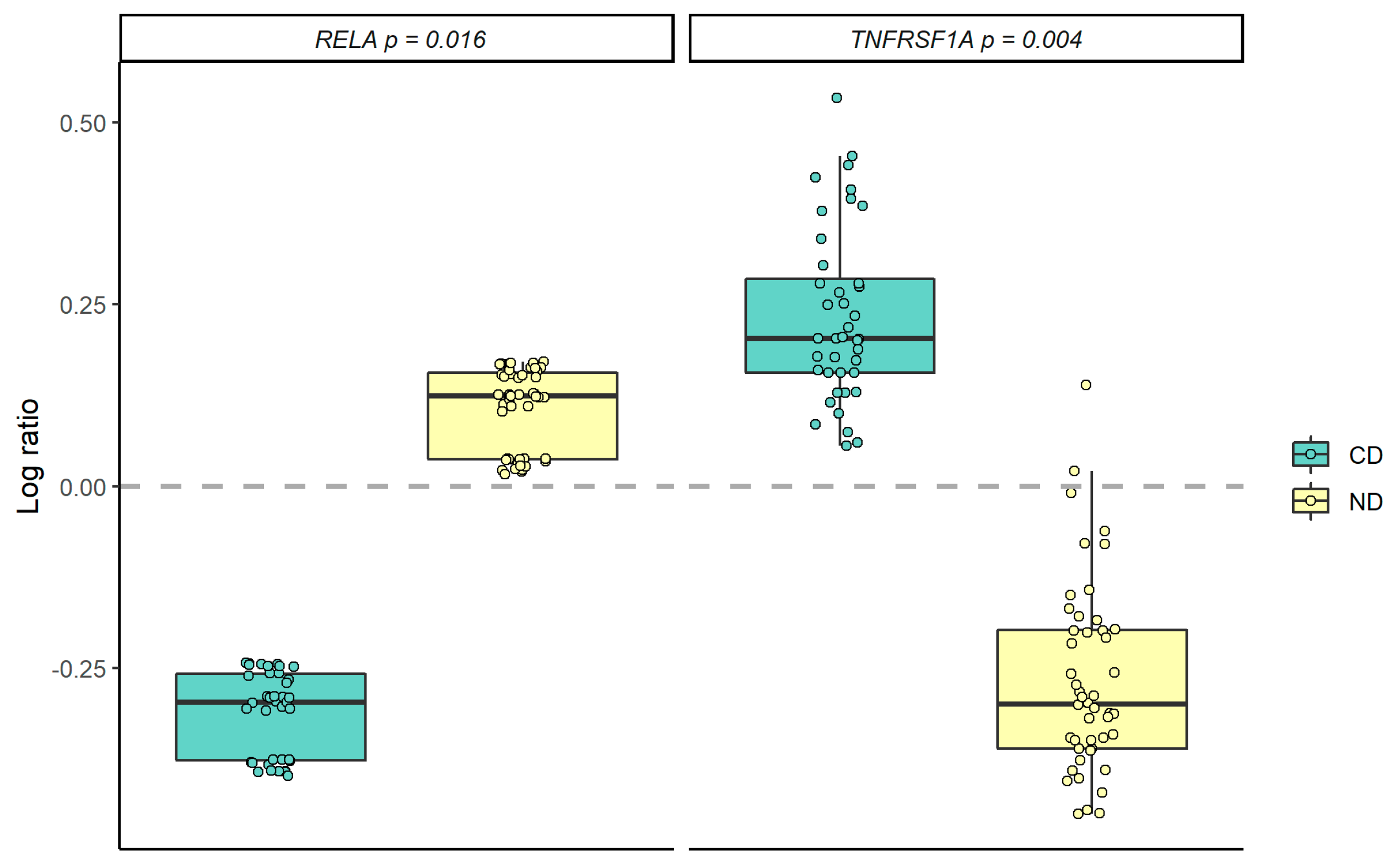
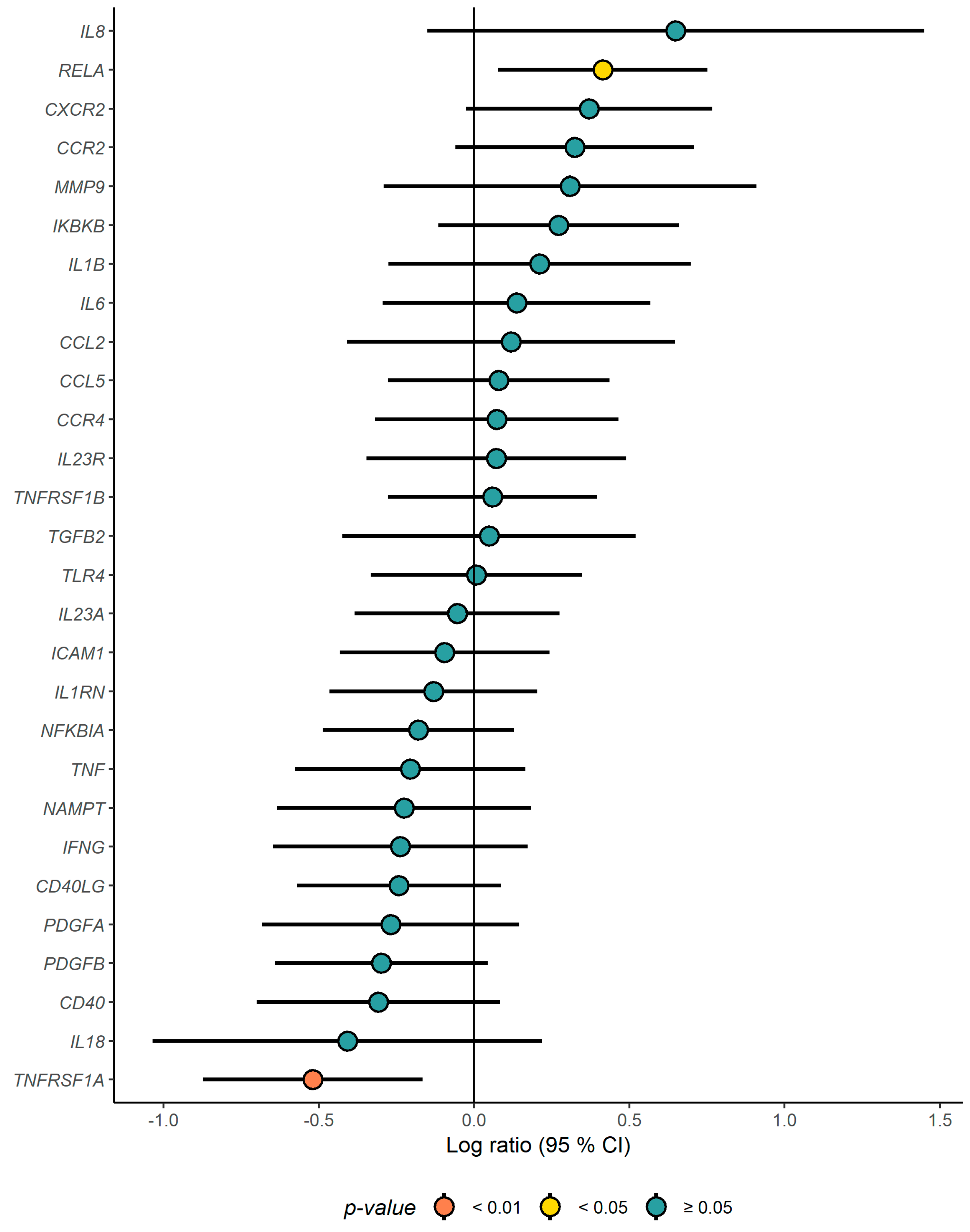
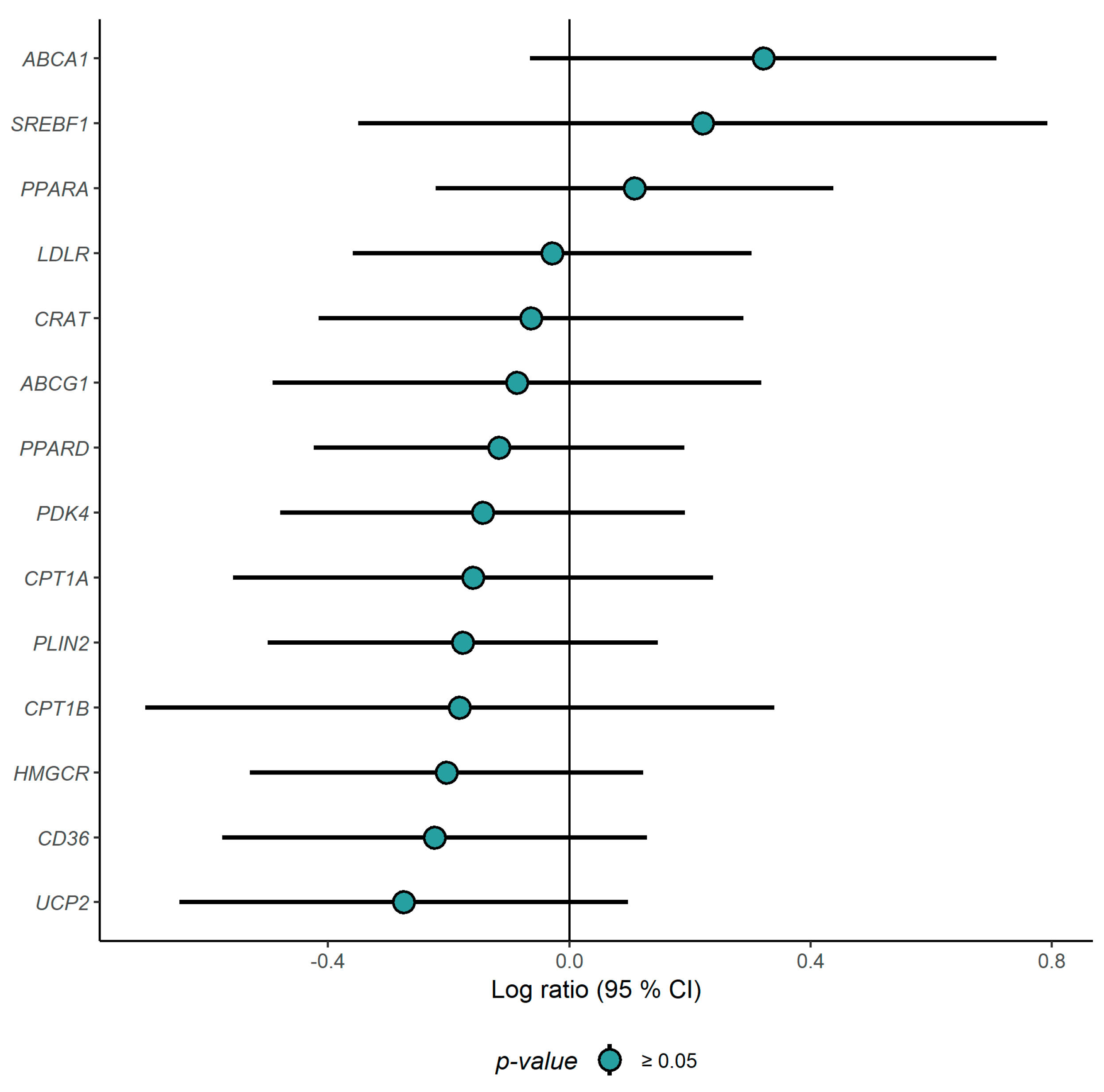
3.2. Gene Expression Profiling in PBMCs
3.3. Correlation Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wellen, K.E.; Hotamisligil, G.S. Inflammation, stress, and diabetes. J. Clin. Investig. 2005, 115, 1111–1119. [Google Scholar] [CrossRef] [PubMed]
- Wellen, K.E.; Hotamisligil, G.S. Obesity-induced inflammatory changes in adipose tissue. J. Clin. Investig. 2003, 112, 1785–1788. [Google Scholar] [CrossRef] [PubMed]
- Bastard, J.P.; Maachi, M.; Lagathu, C.; Kim, M.J.; Caron, M.; Vidal, H.; Capeau, J.; Feve, B. Recent advances in the relationship between obesity, inflammation, and insulin resistance. Eur. Cytokine Netw. 2006, 17, 4–12. [Google Scholar] [PubMed]
- Panagiotakos, D.B.; Pitsavos, C.; Chrysohoou, C.; Skoumas, J.; Tousoulis, D.; Toutouza, M.; Toutouzas, P.; Stefanadis, C. Impact of lifestyle habits on the prevalence of the metabolic syndrome among Greek adults from the ATTICA study. Am. Heart J. 2004, 147, 106–112. [Google Scholar] [CrossRef]
- Esposito, K.; Marfella, R.; Ciotola, M.; Di Palo, C.; Giugliano, F.; Giugliano, G.; D’Armiento, M.; D’Andrea, F.; Giugliano, D. Effect of a mediterranean-style diet on endothelial dysfunction and markers of vascular inflammation in the metabolic syndrome: A randomized trial. JAMA 2004, 292, 1440–1446. [Google Scholar] [CrossRef]
- De Mello, V.D.; Erkkilä, A.T.; Schwab, U.S.; Pulkkinen, L.; Kolehmainen, M.; Atalay, M.; Mussalo, H.; Lankinen, M.; Orešič, M.; Lehto, S.; et al. The effect of fatty or lean fish intake on inflammatory gene expression in peripheral blood mononuclear cells of patients with coronary heart disease. Eur. J. Nutr. 2009, 48, 447–455. [Google Scholar] [CrossRef]
- van Dijk, S.J.; Feskens, E.J.; Bos, M.B.; de Groot, L.C.; de Vries, J.H.; Müller, M.; Afman, L.A. Consumption of a high monounsaturated fat diet reduces oxidative phosphorylation gene expression in peripheral blood mononuclear cells of abdominally overweight men and women. J. Nutr. 2012, 142, 1219–1225. [Google Scholar] [CrossRef]
- Myhrstad, M.C.W.; Ulven, S.M.; Günther, C.C.; Ottestad, I.; Holden, M.; Ryeng, E.; Borge, G.I.; Kohler, A.; Brønner, K.W.; Thoresen, M.; et al. Fish oil supplementation induces expression of genes related to cell cycle, endoplasmic reticulum stress and apoptosis in peripheral blood mononuclear cells: A transcriptomic approach. J. Intern. Med. 2014, 276, 498–511. [Google Scholar] [CrossRef]
- Radler, U.; Stangl, H.; Lechner, S.; Lienbacher, G.; Krepp, R.; Zeller, E.; Brachinger, M.; Eller-Berndl, D.; Fischer, A.; Anzur, C.; et al. A combination of (omega-3) polyunsaturated fatty acids, polyphenols and L-carnitine reduces the plasma lipid levels and increases the expression of genes involved in fatty acid oxidation in human peripheral blood mononuclear cells and HepG2 cells. Ann. Nutr. Metab. 2011, 58, 133–140. [Google Scholar] [CrossRef]
- Afman, L.; Milenkovic, D.; Roche, H.M. Nutritional aspects of metabolic inflammation in relation to health-insights from transcriptomic biomarkers in PBMC of fatty acids and polyphenols. Mol. Nutr. food Res. 2014, 58, 1708–1720. [Google Scholar] [CrossRef]
- Kolehmainen, M.; Mykkänen, O.; Kirjavainen, P.V.; Leppänen, T.; Moilanen, E.; Adriaens, M.; Laaksonen, D.E.; Hallikainen, M.; Pimiä, P.R.; Pulkkinen, L.; et al. Bilberries reduce low-grade inflammation in individuals with features of metabolic syndrome. Mol. Nutr. Food Res. 2012, 56, 1501–1510. [Google Scholar] [CrossRef] [PubMed]
- Kolehmainen, M.; Ulven, S.M.; Paananen, J.; de Mello, V.; Schwab, U.; Carlberg, C.; Myhrstad, M.; Pihlajamäki, J.; Dungner, E.; Sjölin, E.; et al. Healthy Nordic diet downregulates the expression of genes involved in inflammation in subcutaneous adipose tissue in individuals with features of the metabolic syndrome. Am. J. Clin. Nutr. 2015, 101, 228–239. [Google Scholar] [CrossRef] [PubMed]
- Leder, L.; Kolehmainen, M.; Narverud, I.; Dahlman, I.; Myhrstad, M.C.; De Mello, V.D.; Paananen, J.; Carlberg, C.; Schwab, U.; Herzig, K.-H.; et al. Effects of a healthy Nordic diet on gene expression changes in peripheral blood mononuclear cells in response to an oral glucose tolerance test in subjects with metabolic syndrome: A SYSDIET sub-study. Genes Nutr. 2016, 11, 3. [Google Scholar] [CrossRef] [PubMed]
- Myhrstad, M.C.; de Mello, V.D.; Dahlman, I.; Kolehmainen, M.; Paananen, J.; Rundblad, A.; Carlberg, C.; Olstad, O.K.; Pihlajamäki, J.; Holven, K.B.; et al. Healthy Nordic Diet Modulates the Expression of Genes Related to Mitochondrial Function and Immune Response in Peripheral Blood Mononuclear Cells from Subjects with Metabolic Syndrome-A SYSDIET Sub-Study. Mol. Nutr. Food Res. 2019, 68, 1801405. [Google Scholar] [CrossRef]
- de Mello, V.D.; Kolehmanien, M.; Schwab, U.; Pulkkinen, L.; Uusitupa, M. Gene expression of peripheral blood mononuclear cells as a tool in dietary intervention studies: What do we know so far? Mol. Nutr. Food Res. 2012, 56, 1160–1172. [Google Scholar] [CrossRef]
- Uusitupa, M.; Hermansen, K.; Savolainen, M.J.; Schwab, U.; Kolehmainen, M.; Brader, L.; Mortensen, L.S.; Cloetens, L.; Persson, A.J.; Önning, G.; et al. Effects of an isocaloric healthy Nordic diet on insulin sensitivity, lipid profile and inflammation markers in metabolic syndrome—A randomized study (SYSDIET). J. Intern. Med. 2013, 274, 52–66. [Google Scholar] [CrossRef]
- Alberti, K.G.; Eckel, R.H.; Grundy, S.M.; Zimmet, P.Z.; Cleeman, J.I.; Donato, K.A.; Fruchart, J.C.; James, W.P.T.; Loria, C.M.; Smith, S.C., Jr. Harmonizing the metabolic syndrome: A joint interim statement of the International Diabetes Federation Task Force on Epidemiology and Prevention; National Heart, Lung, and Blood Institute; American Heart Association; World Heart Federation; International Atherosclerosis Society; and International Association for the Study of Obesity. Circulation 2009, 120, 1640–1645. [Google Scholar]
- Nordic Council of Ministers. Nordic Nutrition Recommendations 2012: Intergrating Nutrition and Physical Activity; Nordic Council of Ministers: Copenhagen, Denmark, 2014. [Google Scholar]
- De Mello, V.D.F.; Kolehmainen, M.; Pulkkinen, L.; Schwab, U.; Mager, U.; Laaksonen, D.E.; Gylling, H.; Atalay, M.; Rauramaa, R.; Uusitupa, M. Downregulation of genes involved in NFkappaB activation in peripheral blood mononuclear cells after weight loss is associated with the improvement of insulin sensitivity in individuals with the metabolic syndrome: The GENOBIN study. Diabetologia 2008, 51, 2060–2067. [Google Scholar] [CrossRef]
- Rosqvist, F.; Iggman, D.; Kullberg, J.; Cedernaes, J.; Johansson, H.E.; Larsson, A. Overfeeding polyunsaturated and saturated fat causes distinct effects on liver and visceral fat accumulation in humans. Diabetes 2014, 63, 2356–2368. [Google Scholar] [CrossRef]
- Rosqvist, F.; Kullberg, J.; Ståhlman, M.; Cedernaes, J.; Heurling, K.; Johansson, H.E.; Iggman, D.; Wilking, H.; Larsson, A.; Eriksson, O.; et al. Overeating saturated fat promotes fatty liver and ceramides compared to polyunsaturated fat: A randomized trial. J. Clin. Endocrinol. Metab. 2019, 104, 6207–6219. [Google Scholar] [CrossRef]
- Bjermo, H.; Iggman, D.; Kullberg, J.; Dahlman, I.; Johansson, L.; Persson, L.; Berglund, J.; Pulkki, K.; Basu, S.; Uusitupa, M.; et al. Effects of n-6 PUFAs compared with SFAs on liver fat, lipoproteins, and inflammation in abdominal obesity: A randomized controlled trial. Am. J. Clin. Nutr. 2012, 95, 1003–1012. [Google Scholar] [CrossRef] [PubMed]
- Preuss, H.G.; Kaats, G.R.; Mrvichin, N.; Swaroop, A.; Bagchi, D.; Clouatre, D. Examining the Relationship Between Nonalcoholic Fatty Liver Disease and the Metabolic Syndrome in Nondiabetic Subjects. J. Am. Coll. Nutr. 2018, 37, 457–465. [Google Scholar] [CrossRef] [PubMed]
- Aparicio-Vergara, M.; Hommelberg, P.P.; Schreurs, M.; Gruben, N.; Stienstra, R.; Shiri-Sverdlov, R.; Kloosterhuis, N.J.; de Bruin, A.; van de Sluis, B.; Koonen, D.P.Y.; et al. Tumor necrosis factor receptor 1 gain-of-function mutation aggravates nonalcoholic fatty liver disease but does not cause insulin resistance in a murine model. Hepatology 2013, 57, 566–576. [Google Scholar] [CrossRef] [PubMed]
- O’Grada, C.M.; Morine, M.J.; Morris, C.; Ryan, M.; Dillon, E.T.; Walsh, M.; Gibney, R.R.; Brennan, L.; Gibney, M.J.; Roche, H.M. PBMCs reflect the immune component of the WAT transcriptome—Implications as biomarkers of metabolic health in the postprandial state. Mol. Nutr. Food Res. 2014, 58, 808–820. [Google Scholar] [CrossRef]
- Bouwens, M.; Afman, L.A.; Müller, M. Fasting induces changes in peripheral blood mononuclear cell gene expression profiles related to increases in fatty acid beta-oxidation: Functional role of peroxisome proliferator activated receptor alpha in human peripheral blood mononuclear cells. Am. J. Clin. Nutr. 2007, 86, 1515–1523. [Google Scholar] [CrossRef]
- Li, J.; Sapper, T.N.; Mah, E.; Moller, M.V.; Kim, J.B.; Chitchumroonchokchai, C. Green tea extract treatment reduces NFkappaB activation in mice with diet-induced nonalcoholic steatohepatitis by lowering TNFR1 and TLR4 expression and ligand availability. J. Nutr. Biochem. 2017, 41, 34–41. [Google Scholar] [CrossRef]
- Ye, X.; Jiang, X.; Guo, W.; Clark, K.; Gao, Z. Overexpression of NF-kappaB p65 in macrophages ameliorates atherosclerosis in apoE-knockout mice. Am. J. Physiol. Endocrinol. Metab. 2013, 305, E1375–E1383. [Google Scholar] [CrossRef][Green Version]
- Ulven, S.M.; Leder, L.; Elind, E.; Ottestad, I.; Christensen, J.J.; Telle-Hansen, V.H. Exchanging a few commercial, regularly consumed food items with improved fat quality reduces total cholesterol and LDL-cholesterol: A double-blind, randomised controlled trial. Br. J. Nutr. 2016, 116, 1383–1393. [Google Scholar] [CrossRef]
- Ulven, S.M.; Christensen, J.J.; Nygård, O.; Svardal, A.; Leder, L.; Ottestad, I. Using metabolic profiling and gene expression analyses to explore molecular effects of replacing saturated fat with polyunsaturated fat-a randomized controlled dietary intervention study. Am. J. Clin. Nutr. 2019, 109, 1239–1250. [Google Scholar] [CrossRef]
| CD (n = 40) | ND (n = 48) | |||
|---|---|---|---|---|
| Male (n,%) | 15 | (37.5%) | 15 | (31.3%) |
| Age (years) | 55.8 | (7.8) | 54.2 | (8.3) |
| BMI (kg/m2) | 31.9 | (2.7) | 31.7 | (3.1) |
| Waist circumference (cm) | 105.4 | (9.3) | 102.6 | (9.0) |
| BP systolic (mmHg) | 131 | (17) | 127 | (14) |
| BP diastolic (mmHg) | 82 | (12) | 83 | (10) |
| Glucose (mmol/L) | 5.8 | (0.6) | 5.9 | (0.6) |
| Insulin (pmol/L) | 59.5 | (47–80.8) | 56.0 | (41.8–75.3) |
| Triglycerides (mmol/L) | 1.5 | (0.5) | 1.5 | (0.7) |
| Total cholesterol (mmol/L) | 5.3 | (1) | 5.3 | (1) |
| HDL cholesterol (mmol/L) | 1.3 | (0.5) | 1.4 | (0.3) |
| LDL cholesterol (mmol/L) | 3.3 | (0.9) | 3.2 | (0.9) |
| hsCRP (mg/L) | 1.5 | (0.9–3.1) | 1.5 | (0.8–2.8) |
| sTNFRII (ng/L) | 1900 | (415) | 1953 | (466) |
| IL-6 (ng/L) | 1.3 | (1.1–1.7) | 1.3 | (1–1.8) |
| IL-10 (ng/L) | 0.9 | (0.8–1.5) | 0.8 | (0.8–1.5) |
| IL-1β (ng/L) | 0.12 | (0.12–0.17) | 0.12 | (0.12–0.13) |
| IL1 Ra (ng/L) | 309 | (238–463) | 301 | (220–466) |
| HMW adiponectin (µg/L) | 3.6 | (2.2–6.7) | 3.9 | (2.8–6) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ulven, S.M.; Holven, K.B.; Rundblad, A.; Myhrstad, M.C.W.; Leder, L.; Dahlman, I.; Mello, V.D.d.; Schwab, U.; Carlberg, C.; Pihlajamäki, J.; et al. An Isocaloric Nordic Diet Modulates RELA and TNFRSF1A Gene Expression in Peripheral Blood Mononuclear Cells in Individuals with Metabolic Syndrome—A SYSDIET Sub-Study. Nutrients 2019, 11, 2932. https://doi.org/10.3390/nu11122932
Ulven SM, Holven KB, Rundblad A, Myhrstad MCW, Leder L, Dahlman I, Mello VDd, Schwab U, Carlberg C, Pihlajamäki J, et al. An Isocaloric Nordic Diet Modulates RELA and TNFRSF1A Gene Expression in Peripheral Blood Mononuclear Cells in Individuals with Metabolic Syndrome—A SYSDIET Sub-Study. Nutrients. 2019; 11(12):2932. https://doi.org/10.3390/nu11122932
Chicago/Turabian StyleUlven, Stine M., Kirsten B. Holven, Amanda Rundblad, Mari C. W. Myhrstad, Lena Leder, Ingrid Dahlman, Vanessa D. de Mello, Ursula Schwab, Carsten Carlberg, Jussi Pihlajamäki, and et al. 2019. "An Isocaloric Nordic Diet Modulates RELA and TNFRSF1A Gene Expression in Peripheral Blood Mononuclear Cells in Individuals with Metabolic Syndrome—A SYSDIET Sub-Study" Nutrients 11, no. 12: 2932. https://doi.org/10.3390/nu11122932
APA StyleUlven, S. M., Holven, K. B., Rundblad, A., Myhrstad, M. C. W., Leder, L., Dahlman, I., Mello, V. D. d., Schwab, U., Carlberg, C., Pihlajamäki, J., Hermansen, K., Dragsted, L. O., Gunnarsdottir, I., Cloetens, L., Åkesson, B., Rosqvist, F., Hukkanen, J., Herzig, K.-H., Savolainen, M. J., ... Kolehmainen, M. (2019). An Isocaloric Nordic Diet Modulates RELA and TNFRSF1A Gene Expression in Peripheral Blood Mononuclear Cells in Individuals with Metabolic Syndrome—A SYSDIET Sub-Study. Nutrients, 11(12), 2932. https://doi.org/10.3390/nu11122932