Urolithin Metabotypes Can Determine the Modulation of Gut Microbiota in Healthy Individuals by Tracking Walnuts Consumption over Three Days
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design and Participants
2.2. Dietary Estimation
2.3. Biological Samples
2.4. Urolithin Quantification in Urine
2.5. Gut Microbiota Composition Analysis
2.6. Statistical Analysis
2.7. Gut Microbiota Activity Analysis
3. Results
3.1. Characteristics of Subjects
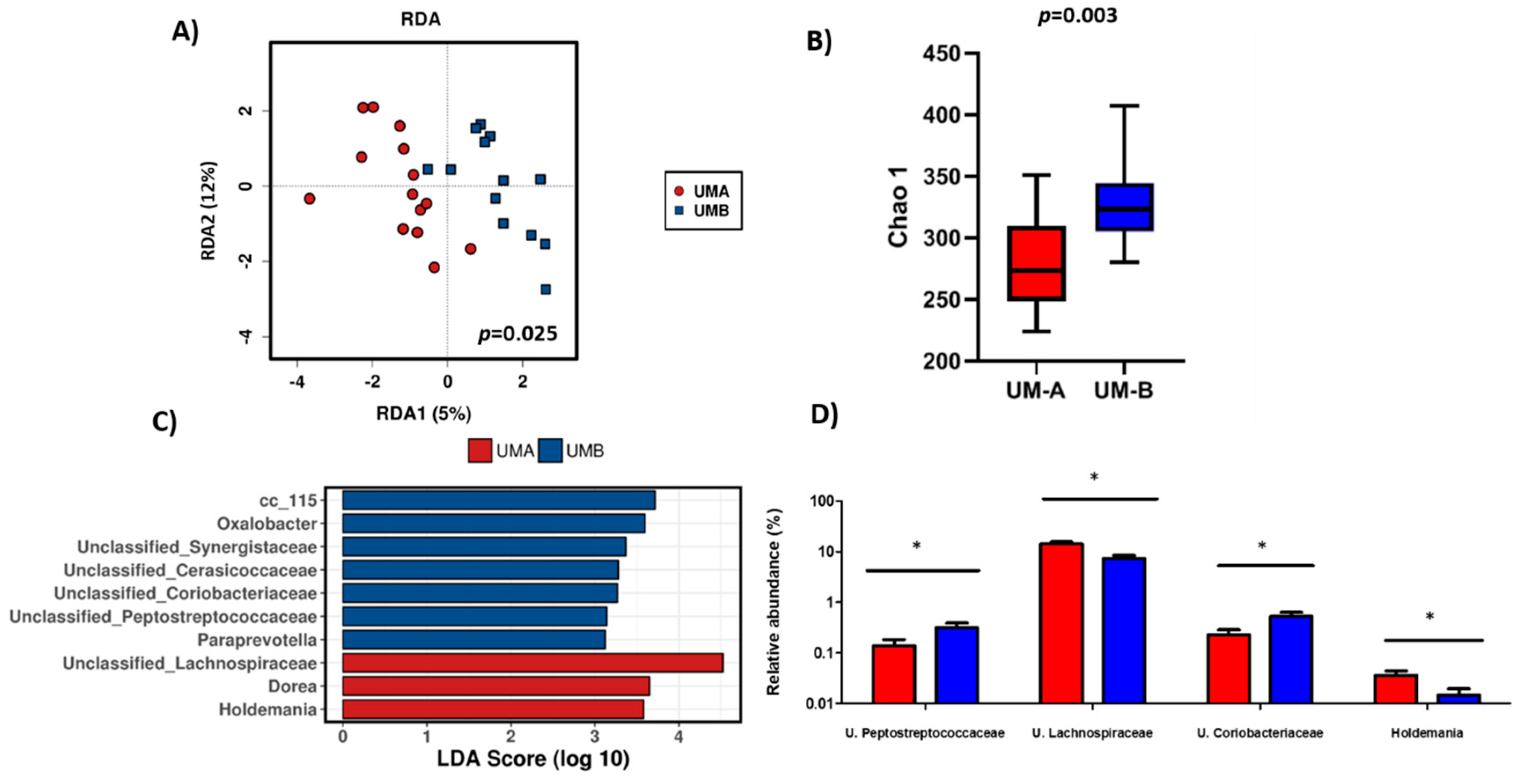
3.2. Baseline Microbiota Composition is Urolithin-Metabotype Dependent
3.3. Microbiota Composition of UM-B is Sensitive to Walnuts Consumption
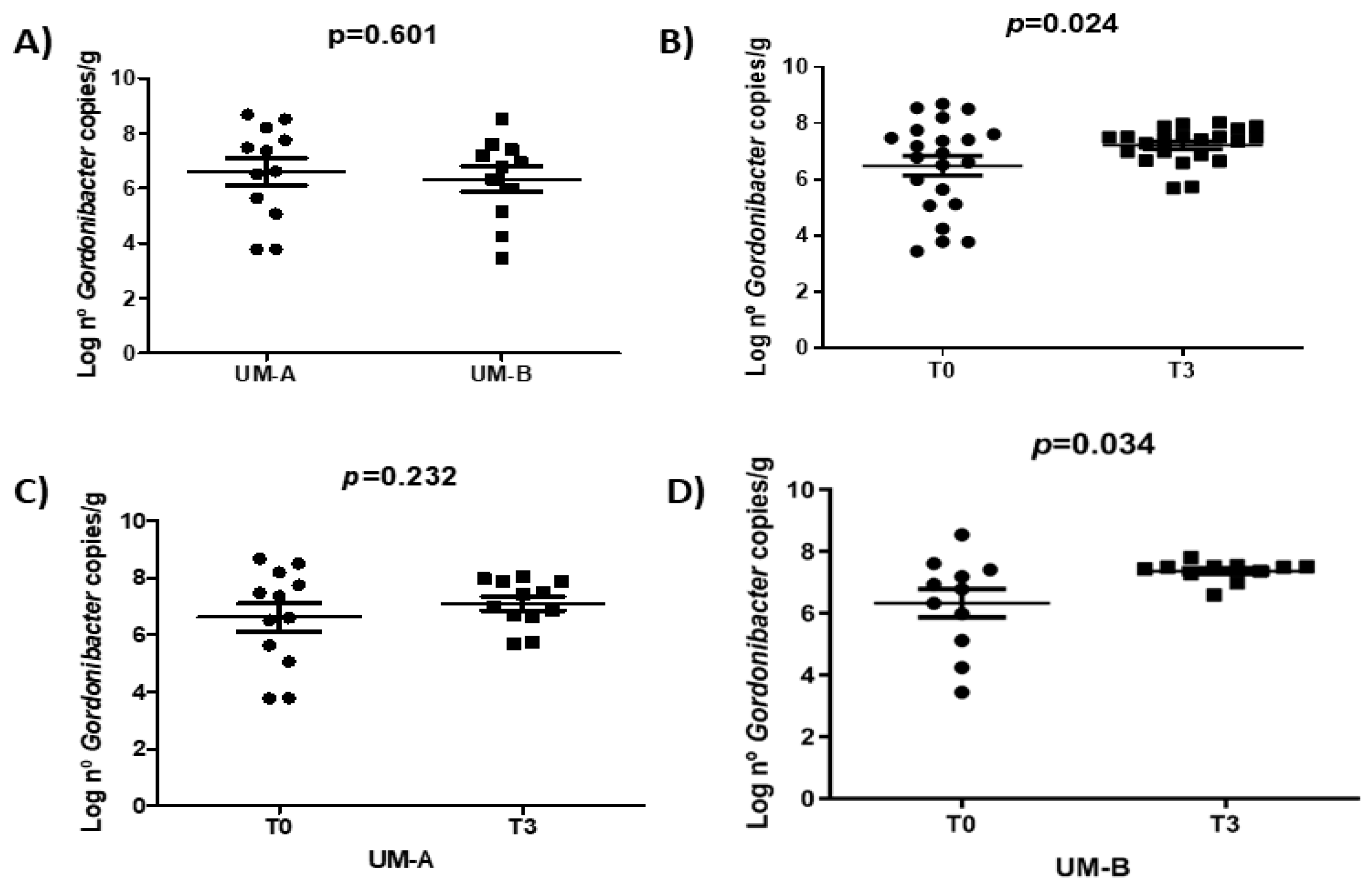
3.4. Walnut Consumption Increased Faecal Gordonibacter Levels in UM-B
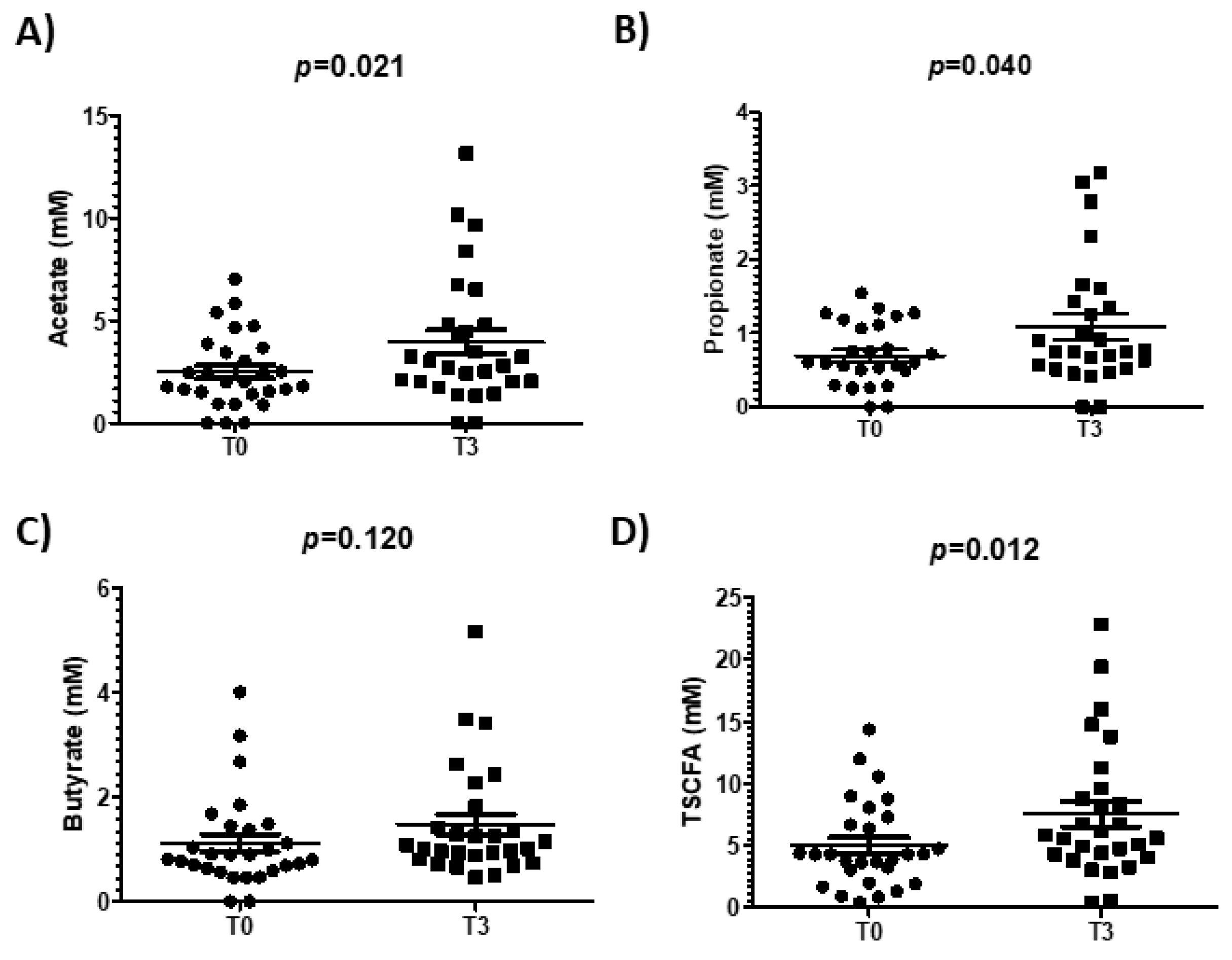
3.5. Effect of Walnut Consumption on Gut Microbiota Activity
3.6. Associations between Gut Microbiota Composition/Activity and Walnut Intervention
4. Discussion and Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| GM | gut microbiota |
| UM | urolithin metabotype |
| qPCR | real-time quantitative polymerase chain reaction |
| UHPLC/Q-TOF-MS | ultra-high performance liquid chromatography-quadrupole time-of-flight mass spectrometry |
| ETs | ellagitannins |
| EA | ellagic acid |
| Uro | urolithin |
| IsoUro | isourolithin |
| CVD | cardiovascular disease |
| SCFA | short chain fatty acids |
| BMI | body mass index |
| FFQ | food frequency questionnaire |
| CESNID | Centro de Enseñanza Superior de Nutrición Humana y Dietética |
| PREDIMED | PREvención con DIeta MEDiterránea |
| LEfSe | linear discriminant analysis effect size |
| RDA | redundancy analysis |
References
- Ros, E. Health benefits of nut consumption. Nutrients 2010, 2, 652–682. [Google Scholar] [CrossRef] [PubMed]
- Larrosa, M.; González-Sarrías, A.; Yáñez-Gascón, M.J.; Selma, M.V.; Azorín-Ortuño, M.; Toti, S.; Tomás-Barberán, F.; Dolara, P.; Espín, J.C. Anti-inflammatory properties of a pomegranate extract and its metabolite urolithin-A in a colitis rat model and the effect of colon inflammation on phenolic metabolism. J. Nutr. Biochem. 2010, 21, 717–725. [Google Scholar] [CrossRef] [PubMed]
- Selma, M.V.; Espín, J.C.; Tomás-Barberán, F.A. Interaction between Phenolics and Gut Microbiota: Role in Human Health. J. Agric. Food Chem. 2009, 57, 6485–6501. [Google Scholar] [CrossRef] [PubMed]
- Salas-Salvadó, J.; Bulló, M.; Pérez-Heras, A.; Ros, E. Dietary fibre, nuts and cardiovascular diseases. Br. J. Nutr. 2006, 96, S45–S51. [Google Scholar] [CrossRef]
- Cardona, F.; Andrés-Lacueva, C.; Tulipani, S.; Tinahones, F.J.; Queipo-Ortuño, M.I. Benefits of polyphenols on gut microbiota and implications in human health. J. Nutr. Biochem. 2013, 24, 1415–1422. [Google Scholar] [CrossRef]
- González-Sarrías, A.; García-Villalba, R.; Romo-Vaquero, M.; Alasalvar, C.; Örem, A.; Zafrilla, P.; Tomás-Barberán, F.A.; Selma, M.V.; Espín, J.C. Clustering according to urolithin metabotype explains the interindividual variability in the improvement of cardiovascular risk biomarkers in overweight-obese individuals consuming pomegranate: A randomized clinical trial. Mol. Nutr. Food Res. 2017, 61, 1600830. [Google Scholar] [CrossRef]
- Byerley, L.O.; Samuelson, D.; Blanchard, E.; Luo, M.; Lorenzen, B.N.; Banks, S.; Ponder, M.A.; Welsh, D.A.; Taylor, C.M. Changes in the gut microbial communities following addition of walnuts to the diet. J. Nutr. Biochem. 2017, 48, 94–102. [Google Scholar] [CrossRef]
- Holscher, H.D.; Guetterman, H.M.; Swanson, K.S.; An, R.; Matthan, N.R.; Lichtenstein, A.H.; Novotny, J.A.; Baer, D.J. Walnut Consumption Alters the Gastrointestinal Microbiota, Microbially Derived Secondary Bile Acids, and Health Markers in Healthy Adults: A Randomized Controlled Trial. J. Nutr. 2018, 148, 861–867. [Google Scholar] [CrossRef]
- Holscher, H.D.; Taylor, A.M.; Swanson, K.S.; Novotny, J.A.; Baer, D.J. Almond Consumption and Processing Affects the Composition of the Gastrointestinal Microbiota of Healthy Adult Men and Women: A Randomized Controlled Trial. Nutrients 2018, 10, 126. [Google Scholar] [CrossRef]
- Han, Y.; Song, M.; Gu, M.; Ren, D.; Zhu, X.; Cao, X.; Li, F.; Wang, W.; Cai, X.; Yuan, B.; et al. Dietary Intake of Whole Strawberry Inhibited Colonic Inflammation in Dextran-Sulfate-Sodium-Treated Mice via Restoring Immune Homeostasis and Alleviating Gut Microbiota Dysbiosis. J. Agric. Food Chem. 2019. [Google Scholar] [CrossRef]
- Tomás-Barberán, F.A.; García-Villalba, R.; González-Sarrías, A.; Selma, M.V.; Espín, J.C. Ellagic Acid Metabolism by Human Gut Microbiota: Consistent Observation of Three Urolithin Phenotypes in Intervention Trials, Independent of Food Source, Age, and Health Status. J. Agric. Food Chem. 2014, 62, 6535–6538. [Google Scholar] [CrossRef] [PubMed]
- Tomás-Barberán, F.A.; González-Sarrías, A.; García-Villalba, R.; Núñez-Sánchez, M.A.; Selma, M.V.; García-Conesa, M.T.; Espín, J.C. Urolithins, the rescue of “old” metabolites to understand a “new” concept: Metabotypes as a nexus among phenolic metabolism, microbiota dysbiosis, and host health status. Mol. Nutr. Food Res. 2017, 61, 1500901. [Google Scholar] [CrossRef] [PubMed]
- Beltrán, D.; Romo-Vaquero, M.; Espín, J.C.; Tomás-Barberán, F.A.; Selma, M.V. Ellagibacter isourolithinifaciens gen. nov., sp. nov., a new member of the family Eggerthellaceae, isolated from human gut. Int. J. Syst. Evol. Microbiol. 2018, 68, 1707–1712. [Google Scholar] [CrossRef] [PubMed]
- Selma, M.V.; Beltrán, D.; García-Villalba, R.; Espín, J.C.; Tomás-Barberán, F.A. Description of urolithin production capacity from ellagic acid of two human intestinal Gordonibacter species. Food Funct. 2014, 5, 1779–1784. [Google Scholar] [CrossRef]
- Selma, M.V.; Tomas-Barberan, F.A.; Beltran, D.; Garcia-Villalba, R.; Espin, J.C. Gordonibacter urolithinfaciens sp. nov., a urolithin-producing bacterium isolated from the human gut. Int. J. Syst. Evol. Microbiol. 2014, 64, 2346–2352. [Google Scholar] [CrossRef]
- Selma, M.V.; Beltrán, D.; Luna, M.C.; Romo-Vaquero, M.; García-Villalba, R.; Mira, A.; Espín, J.C.; Tomás-Barberán, F.A. Isolation of Human Intestinal Bacteria Capable of Producing the Bioactive Metabolite Isourolithin A from Ellagic Acid. Front. Microbiol. 2017, 8, 1521. [Google Scholar] [CrossRef]
- Selma, M.V.; González-Sarrías, A.; Salas-Salvadó, J.; Andrés-Lacueva, C.; Alasalvar, C.; Örem, A.; Tomás-Barberán, F.A.; Espín, J.C. The gut microbiota metabolism of pomegranate or walnut ellagitannins yields two urolithin-metabotypes that correlate with cardiometabolic risk biomarkers: Comparison between normoweight, overweight-obesity and metabolic syndrome. Clin. Nutr. 2018, 37, 897–905. [Google Scholar] [CrossRef]
- Cortés-Martín, A.; Selma, M.V.; Espín, J.C.; García-Villalba, R. The Human Metabolism of Nuts Proanthocyanidins does not Reveal Urinary Metabolites Consistent with Distinctive Gut Microbiota Metabotypes. Mol. Nutr. Food Res. 2019, 63, 1800819. [Google Scholar] [CrossRef]
- García-Villalba, R.; Espín, J.C.; Aaby, K.; Alasalvar, C.; Heinonen, M.; Jacobs, G.; Voorspoels, S.; Koivumäki, T.; Kroon, P.A.; Pelvan, E.; et al. Validated Method for the Characterization and Quantification of Extractable and Nonextractable Ellagitannins after Acid Hydrolysis in Pomegranate Fruits, Juices, and Extracts. J. Agric. Food Chem. 2015, 63, 6555–6566. [Google Scholar] [CrossRef]
- Salas-Salvadó, J.; Rubio, M.A.; Barbany, M.; Moreno, B. Consenso SEEDO 2007 para la evaluación del sobrepeso y la obesidad y el establecimiento de criterios de intervención terapéutica. Conferencia de consenso. Med. Clin. 2007, 128, 184–196. [Google Scholar]
- Fernández-Ballart, J.D.; Piñol, J.L.; Zazpe, I.; Corella, D.; Carrasco, P.; Toledo, E.; Perez-Bauer, M.; Martínez-González, M.Á.; Salas-Salvadó, J.; Martín-Moreno, J.M. Relative validity of a semi-quantitative food-frequency questionnaire in an elderly Mediterranean population of Spain. Br. J. Nutr. 2010, 103, 1808–1816. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Gonzalez, M.A.; Corella, D.; Salas-Salvado, J.; Ros, E.; Covas, M.I.; Fiol, M.; Warnberg, J.; Aros, F.; Ruiz-Gutierrez, V.; Lamuela-Raventos, R.M.; et al. Cohort Profile: Design and methods of the PREDIMED study. Int. J. Epidemiol. 2012, 41, 377–385. [Google Scholar] [CrossRef] [PubMed]
- García-Villalba, R.; Espín, J.C.; Tomás-Barberán, F.A. Chromatographic and spectroscopic characterization of urolithins for their determination in biological samples after the intake of foods containing ellagitannins and ellagic acid. J. Chromatogr. A 2016, 1428, 162–175. [Google Scholar] [CrossRef] [PubMed]
- Cortés-Martín, A.; García-Villalba, R.; González-Sarrías, A.; Romo-Vaquero, M.; Loria-Kohen, V.; Ramírez-de-Molina, A.; Tomás-Barberán, F.A.; Selma, M.V.; Espín, J.C. The gut microbiota urolithin metabotypes revisited: The human metabolism of ellagic acid is mainly determined by aging. Food Funct. 2018, 9, 4100–4106. [Google Scholar] [CrossRef]
- Garcia-Mantrana, I.; Selma-Royo, M.; Alcantara, C.; Collado, M.C. Shifts on Gut Microbiota Associated to Mediterranean Diet Adherence and Specific Dietary Intakes on General Adult Population. Front. Microbiol. 2018, 9, 890. [Google Scholar] [CrossRef]
- Romo-Vaquero, M.; García-Villalba, R.; González-Sarrías, A.; Beltrán, D.; Tomás-Barberán, F.A.; Espín, J.C.; Selma, M.V. Interindividual variability in the human metabolism of ellagic acid: Contribution of Gordonibacter to urolithin production. J. Funct. Foods 2015, 17, 785–791. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef]
- Andersen, S.J.; Hennebel, T.; Gildemyn, S.; Coma, M.; Desloover, J.; Berton, J.; Tsukamoto, J.; Stevens, C.; Rabaey, K. Electrolytic Membrane Extraction Enables Production of Fine Chemicals from Biorefinery Sidestreams. Environ. Sci. Technol. 2014, 48, 7135–7142. [Google Scholar] [CrossRef]
- Espín, J.C.; González-Sarrías, A.; Tomás-Barberán, F.A. The gut microbiota: A key factor in the therapeutic effects of (poly)phenols. Biochem. Pharmacol. 2017, 139, 82–93. [Google Scholar] [CrossRef]
- Healey, G.R.; Murphy, R.; Brough, L.; Butts, C.A.; Coad, J. Interindividual variability in gut microbiota and host response to dietary interventions. Nutr. Rev. 2017, 75, 1059–1080. [Google Scholar] [CrossRef] [PubMed]
- Romo-Vaquero, M.; Cortés-Martín, A.; Loria-Kohen, V.; Ramírez-de-Molina, A.; García-Mantrana, I.; Collado, M.C.; Espín, J.C.; Selma, M.V. Deciphering the Human Gut Microbiome of Urolithin Metabotypes: Association with Enterotypes and Potential Cardiometabolic Health Implications. Mol. Nutr. Food Res. 2019, 63, 1800958. [Google Scholar] [CrossRef] [PubMed]
- Tulipani, S.; Urpi-Sarda, M.; Garcı́a-Villalba, R.; Rabassa, M.; López-Uriarte, P.; Bulló, M.; Jáuregui, O.; Tomás-Barberán, F.; Salas-Salvadó, J.; Espı́n, J.C.; et al. Urolithins Are the Main Urinary Microbial-Derived Phenolic Metabolites Discriminating a Moderate Consumption of Nuts in Free-Living Subjects with Diagnosed Metabolic Syndrome. J. Agric. Food Chem. 2012, 60, 8930–8940. [Google Scholar] [CrossRef] [PubMed]
- Ukhanova, M.; Wang, X.; Baer, D.J.; Novotny, J.A.; Fredborg, M.; Mai, V. Effects of almond and pistachio consumption on gut microbiota composition in a randomised cross-over human feeding study. Br. J. Nutr. 2014, 111, 2146–2152. [Google Scholar] [CrossRef] [PubMed]
- Bamberger, C.; Rossmeier, A.; Lechner, K.; Wu, L.; Waldmann, E.; Fischer, S.; Stark, R.; Altenhofer, J.; Henze, K.; Parhofer, K. A Walnut-Enriched Diet Affects Gut Microbiome in Healthy Caucasian Subjects: A Randomized, Controlled Trial. Nutrients 2018, 10, 244. [Google Scholar] [CrossRef] [PubMed]
- Ros, E.; Mataix, J. Fatty acid composition of nuts-implications for cardiovascular health. Br. J. Nutr. 2006, 96, S29–S35. [Google Scholar] [CrossRef]
- Lamuel-Raventos, R.M.; Onge, M.-P.S. Prebiotic nut compounds and human microbiota. Crit. Rev. Food Sci. Nutr. 2017, 57, 3154–3163. [Google Scholar] [CrossRef]
- Costantini, L.; Molinari, R.; Farinon, B.; Merendino, N. Impact of Omega-3 Fatty Acids on the Gut Microbiota. Int. J. Mol. Sci. 2017, 18, 2645. [Google Scholar] [CrossRef]
- Burns, A.M.; Zitt, M.A.; Rowe, C.C.; Langkamp-Henken, B.; Mai, V.; Nieves, C.; Ukhanova, M.; Christman, M.C.; Dahl, W.J. Diet quality improves for parents and children when almonds are incorporated into their daily diet: A randomized, crossover study. Nutr. Res. 2016, 36, 80–89. [Google Scholar] [CrossRef]
- Rivière, A.; Selak, M.; Lantin, D.; Leroy, F.; De Vuyst, L. Bifidobacteria and Butyrate-Producing Colon Bacteria: Importance and Strategies for Their Stimulation in the Human Gut. Front. Microbiol. 2016, 7, 979. [Google Scholar] [CrossRef]
- Chambers, E.S.; Byrne, C.S.; Morrison, D.J.; Murphy, K.G.; Preston, T.; Tedford, C.; Garcia-Perez, I.; Fountana, S.; Serrano-Contreras, J.I.; Holmes, E.; et al. Dietary supplementation with inulin-propionate ester or inulin improves insulin sensitivity in adults with overweight and obesity with distinct effects on the gut microbiota, plasma metabolome and systemic inflammatory responses: A randomised cross-over trial. Gut 2019, 68, 1430–1438. [Google Scholar] [PubMed]
- Banel, D.K.; Hu, F.B. Effects of walnut consumption on blood lipids and other cardiovascular risk factors: A meta-analysis and systematic review. Am. J. Clin. Nutr. 2009, 90, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Lapiscina, E.H.; Clavero, P.; Toledo, E.; San Julian, B.; Sanchez-Tainta, A.; Corella, D.; Lamuela-Raventos, R.M.; Martinez, J.A.; Martinez-Gonzalez, M.Á. Virgin olive oil supplementation and long-term cognition: The Predimed-Navarra randomized, trial. J. Nutr. Health Aging 2013, 17, 544–552. [Google Scholar] [CrossRef] [PubMed]
- Estruch, R.; Ros, E.; Salas-Salvadó, J.; Covas, M.-I.; Corella, D.; Arós, F.; Gómez-Gracia, E.; Ruiz-Gutiérrez, V.; Fiol, M.; Lapetra, J.; et al. Primary Prevention of Cardiovascular Disease with a Mediterranean Diet. N. Engl. J. Med. 2013, 368, 1279–1290. [Google Scholar] [CrossRef] [PubMed]
- Louis, P.; Flint, H.J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 2017, 19, 29–41. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Chandrashekharappa, S.; Bodduluri, S.R.; Baby, B.V.; Hegde, B.; Kotla, N.G.; Hiwale, A.A.; Saiyed, T.; Patel, P.; Vijay-Kumar, M.; et al. Enhancement of the gut barrier integrity by a microbial metabolite through the Nrf2 pathway. Nat. Commun. 2019, 10, 89. [Google Scholar] [CrossRef]
- Sakata, T. Pitfalls in short-chain fatty acid research: A methodological review. Anim. Sci. J. 2019, 90, 3–13. [Google Scholar] [CrossRef]

| Urolithin Metabotypes | ||||
|---|---|---|---|---|
| Total (n = 27) | UM-A (n = 14) | UM-B (n = 13) | p Value | |
| Age (years) | 39.5 ± 7.3 | 36.1 ± 7.2 | 43.1 ± 5.6 | 0.521 |
| BMI (kg/cm2) | 23.3 ± 3.2 | 23.9 ± 3.5 | 22.7 ± 2.8 | 0.625 |
| Normal Weight (%) | 70.4 | 64.3 | 76.9 | 0.961 |
| Overweight (%) | 29.6 | 35.7 | 23.1 | 0.393 |
| Sex | ||||
| Female (%) | 55.5 | 57.1 | 61.5 | 0.471 |
| Male (%) | 40.7 | 42.8 | 38.5 | 0.565 |
| MD (%) | 55.5 | 57.1 | 53.8 | 0.669 |
| Total Protein (g/day) | 93.9 ± 15.5 | 95.5 ± 13.5 | 92.2 ± 17.9 | 0.597 |
| Animal-Derived Protein (g/day) | 47.7 ± 13.9 | 50.8 ± 12.4 | 44.4 ± 15.1 | 0.237 |
| Plant-Derived Protein (g/day) | 44.1 ± 7.7 | 42.4 ± 7.4 | 45.9 ± 7.9 | 0.232 |
| Lipids (g/day) | 78.8 ± 9.5 | 80.9 ± 8.6 | 76.5 ± 10.3 | 0.246 |
| SFA | 16.5 ± 3.2 | 16.1 ± 3.5 | 16.9 ± 3.0 | 0.527 |
| MUFA | 33.5 ± 4.9 | 33.7 ± 5.4 | 33.3 ± 4.5 | 0.844 |
| PUFA | 12.8 ± 2.5 | 12.6 ± 2.1 | 13.0 ± 2.9 | 0.684 |
| Total Carbohydrates (g/day) | 215.4 ± 29.7 | 207.8 ± 24.3 | 223.6 ± 33.6 | 0.171 |
| Dietary Fiber (g/day) | 26.1 ± 5.9 | 26.7 ± 6.4 | 25.4 ± 5.7 | 0.593 |
| Insoluble Dietary Fiber (g/day) | 16.8 ± 4.8 | 17.2 ± 4.9 | 16.3 ± 4.8 | 0.636 |
| Soluble Dietary Fiber (g/day) | 3.1 ± 0.8 | 3.0 ± 0.9 | 3.2 ± 0.8 | 0.553 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
García-Mantrana, I.; Calatayud, M.; Romo-Vaquero, M.; Espín, J.C.; Selma, M.V.; Collado, M.C. Urolithin Metabotypes Can Determine the Modulation of Gut Microbiota in Healthy Individuals by Tracking Walnuts Consumption over Three Days. Nutrients 2019, 11, 2483. https://doi.org/10.3390/nu11102483
García-Mantrana I, Calatayud M, Romo-Vaquero M, Espín JC, Selma MV, Collado MC. Urolithin Metabotypes Can Determine the Modulation of Gut Microbiota in Healthy Individuals by Tracking Walnuts Consumption over Three Days. Nutrients. 2019; 11(10):2483. https://doi.org/10.3390/nu11102483
Chicago/Turabian StyleGarcía-Mantrana, Izaskun, Marta Calatayud, María Romo-Vaquero, Juan Carlos Espín, María V. Selma, and María Carmen Collado. 2019. "Urolithin Metabotypes Can Determine the Modulation of Gut Microbiota in Healthy Individuals by Tracking Walnuts Consumption over Three Days" Nutrients 11, no. 10: 2483. https://doi.org/10.3390/nu11102483
APA StyleGarcía-Mantrana, I., Calatayud, M., Romo-Vaquero, M., Espín, J. C., Selma, M. V., & Collado, M. C. (2019). Urolithin Metabotypes Can Determine the Modulation of Gut Microbiota in Healthy Individuals by Tracking Walnuts Consumption over Three Days. Nutrients, 11(10), 2483. https://doi.org/10.3390/nu11102483






