Interaction between an ADCY3 Genetic Variant and Two Weight-Lowering Diets Affecting Body Fatness and Body Composition Outcomes Depending on Macronutrient Distribution: A Randomized Trial
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Diet Intervention
2.3. General Measurements
2.4. Genotyping
2.5. Statistical Analyses
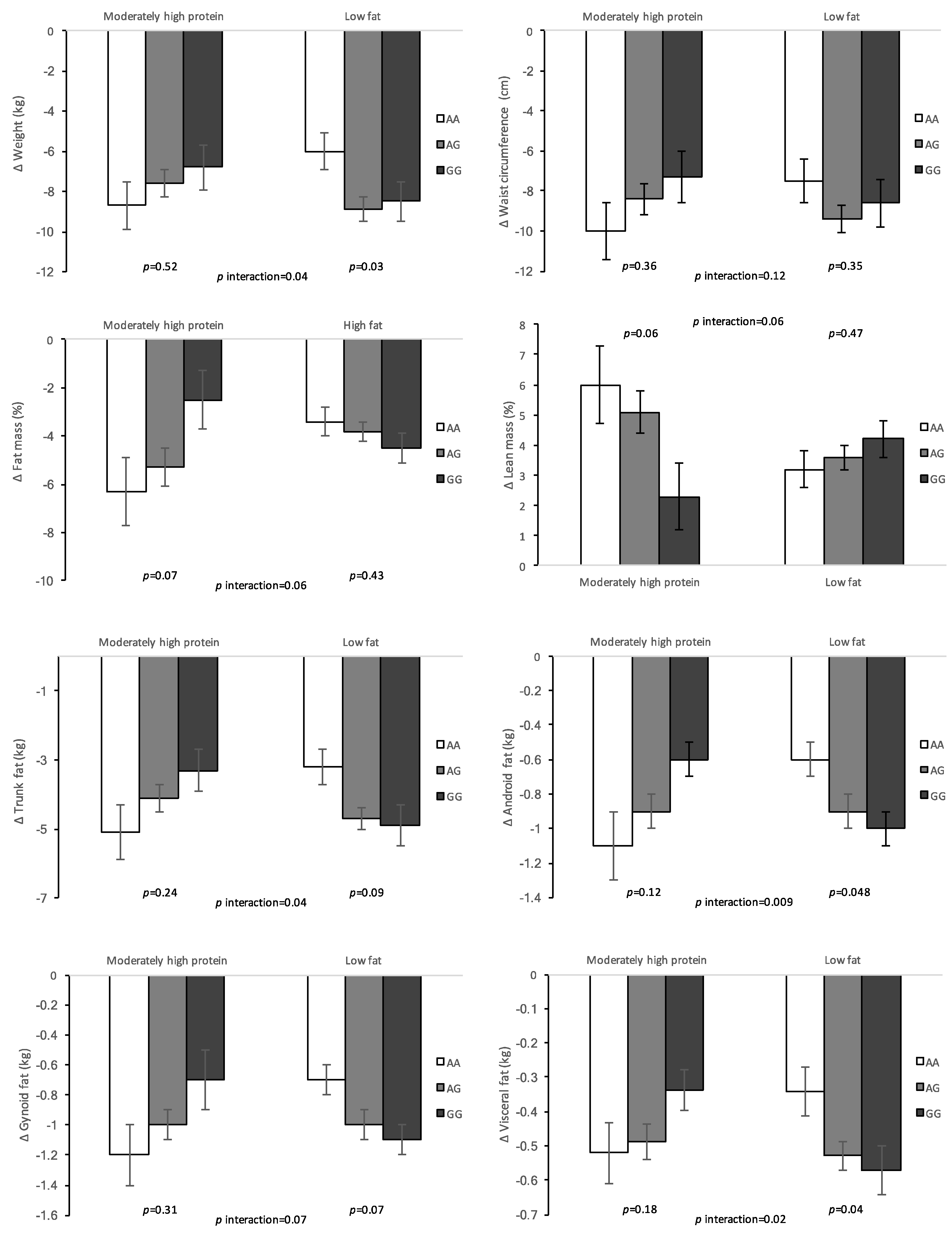
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Zhang, Y.; Liu, J.; Yao, J.; Ji, G.; Qian, L.; Wang, J.; Zhang, G.; Tian, J.; Nie, Y.; Zhang, Y.E.; et al. Obesity: Pathophysiology and intervention. Nutrients 2014, 6, 5153–5183. [Google Scholar] [CrossRef] [PubMed]
- Mariman, E.C.M.; Vink, R.G.; Roumans, N.J.T.; Bouwman, F.G.; Stumpel, C.T.R.M.; Aller, E.E.J.G.; van Baak, M.A.; Wang, P. The cilium: A cellular antenna with an influence on obesity risk. Br. J. Nutr. 2016, 116, 576–592. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Shen, C.; Seed Ahmed, M.; Östenson, C.-G.; Gu, H.F. Adenylate cyclase 3: A new target for anti-obesity drug development. Obes. Rev. 2016, 17, 907–914. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Halim, S.M.; Guenifi, A.; He, B.; Yang, B.; Mustafa, M.; Höjeberg, B.; Hillert, J.; Bakhiet, M.; Efendić, S. Mutations in the promoter of adenylyl cyclase (AC)-III gene, overexpression of AC-III mRNA, and enhanced cAMP generation in islets from the spontaneously diabetic GK rat model of type 2 diabetes. Diabetes 1998, 47, 498–504. [Google Scholar] [CrossRef] [PubMed]
- Seed Ahmed, M.; Kovoor, A.; Nordman, S.; Abu Seman, N.; Gu, T.; Efendic, S.; Brismar, K.; Östenson, C.-G.; Gu, H.F. Increased expression of adenylyl cyclase 3 in pancreatic islets and central nervous system of diabetic Goto-Kakizaki rats: A possible regulatory role in glucose homeostasis. Islets 2012, 4, 343–348. [Google Scholar] [CrossRef] [PubMed]
- Nordman, S.; Abulaiti, A.; Hilding, A.; Långberg, E.-C.; Humphreys, K.; Ostenson, C.-G.; Efendic, S.; Gu, H.F. Genetic variation of the adenylyl cyclase 3 (AC3) locus and its influence on type 2 diabetes and obesity susceptibility in Swedish men. Int. J. Obes. 2008, 32, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Wu, M.; Zhu, W.; Shen, J.; Shi, X.; Yang, J.; Zhao, Q.; Ni, C.; Xu, Y.; Shen, H.; et al. Evaluation of the association between the AC3 genetic polymorphisms and obesity in a Chinese Han population. PLoS ONE 2010, 5, e13851. [Google Scholar] [CrossRef] [PubMed]
- Speliotes, E.K.; Willer, C.J.; Berndt, S.I.; Monda, K.L.; Thorleifsson, G.; Jackson, A.U.; Lango Allen, H.; Lindgren, C.M.; Luan, J.; Mägi, R.; et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat. Genet. 2010, 42, 937–948. [Google Scholar] [CrossRef] [PubMed]
- Monda, K.L.; Chen, G.K.; Taylor, K.C.; Palmer, C.; Edwards, T.L.; Lange, L.A.; Ng, M.C.Y.; Adeyemo, A.A.; Allison, M.A.; Bielak, L.F.; et al. A meta-analysis identifies new loci associated with body mass index in individuals of African ancestry. Nat. Genet. 2013, 45, 690–696. [Google Scholar] [CrossRef] [PubMed]
- Berndt, S.I.; Gustafsson, S.; Magi, R.; Ganna, A.; Wheeler, E.; Feitosa, M.F.; Justice, A.E.; Monda, K.L.; Croteau-Chonka, D.C.; Day, F.R.; et al. Genome-wide meta-analysis identifies 11 new loci for anthropometric traits and provides insights into genetic architecture. Nat. Genet. 2013, 45, 501–512. [Google Scholar] [CrossRef] [PubMed]
- Graff, M.; Ngwa, J.S.; Workalemahu, T.; Homuth, G.; Schipf, S.; Teumer, A.; Vlzke, H.; Wallaschofski, H.; Abecasis, G.R.; Edward, L.; et al. Genome-wide analysis of BMI in adolescents and young adults reveals additional insight into the effects of genetic loci over the life course. Hum. Mol. Genet. 2013, 22, 3597–3607. [Google Scholar] [CrossRef] [PubMed]
- Locke, A.E.; Kahali, B.; Berndt, S.I.; Justice, A.E.; Pers, T.H.; Day, F.R.; Powell, C.; Vedantam, S.; Buchkovich, M.L.; Yang, J.; et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 2015, 518, 197–206. [Google Scholar] [CrossRef] [PubMed]
- World Medical Association. World Medical Association Declaration of Helsinki: Ethical principles for medical research involving human subjects. JAMA 2013, 310, 2191–2194. [Google Scholar] [CrossRef]
- Institute of Medicine. Dietary Reference Intakes for Energy, Carbohydrate, Fiber, Fat, Fatty Acids, Cholesterol, Protein, and Amino Acids (Macronutrients); The National Academies Press: Washington, DC, USA, 2005. [Google Scholar]
- Mifflin, M.D.; St Jeor, S.T.; Hill, L.A.; Scott, B.J.; Daugherty, S.A.; Koh, Y.O. A new predictive equation in healthy individuals for resting energy. Am. J. Clin. Nutr. 1990, 51, 241–247. [Google Scholar] [CrossRef] [PubMed]
- Seagle, H.M.; Strain, G.W.; Makris, A.; Reeves, R.S. Position of the American Dietetic Association: Weight management. J. Am. Diet. Assoc. 2009, 109, 330–346. [Google Scholar] [CrossRef] [PubMed]
- Mataix Verdú, J. Tabla de Compicion de Alimentos, 5th ed.; Universidad de Granada: Granada, Spain, 2009. [Google Scholar]
- Moreiras, O.; Carbajal, Á.; Cabrera, L.; Cuadrado, C. Tablas de Composición de Alimentos, 16th ed.; Piramide: Madrid, Spain, 2013. [Google Scholar]
- Farrán, A.; Zamora, R.; Cervera, P. Tablas de Composición de Alimentos del CESNID, 2nd ed.; McGraw Hill: Madrid, Spain, 2009. [Google Scholar]
- Perez, S.; Parra, M.D.; Martínez de Morentin, B.E.; Rodríguez, C.M.; Martínez, J.A. Evaluación de la variabilidad intraindividual de la medida de composición corporal mediante bioimpedancia en voluntarias sanas y su relación con el índice de masa corporal y el pliegue tricipital. Enfermería Clínica 2006, 15, 343–347. [Google Scholar] [CrossRef]
- Martin-Moreno, J.M.; Boyle, P.; Gorgojo, L.; Maisonneuve, P.; Fernandez-Rodriguez, J.C.; Salvini, S.; Willett, W.C. Development and validation of a food frequency questionnaire in Spain. Int. J. Epidemiol. 1993, 22, 512–519. [Google Scholar] [CrossRef] [PubMed]
- De la Fuente-Arrillaga, C.; Vázquez Ruiz, Z.; Bes-Rastrollo, M.; Sampson, L.; Martinez-González, M.A. Reproducibility of an FFQ validated in Spain. Public Health Nutr. 2010, 13, 1364–1372. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Ballart, J.D.; Piñol, J.L.; Zazpe, I.; Corella, D.; Carrasco, P.; Toledo, E.; Perez-Bauer, M.; Martínez-González, M.A.; Salas-Salvadó, J.; Martín-Moreno, J.M. Relative validity of a semi-quantitative food-frequency questionnaire in an elderly Mediterranean population of Spain. Br. J. Nutr. 2010, 103, 1808–1816. [Google Scholar] [CrossRef] [PubMed]
- Rogne, M.; Taskén, K. Compartmentalization of cAMP signaling in adipogenesis, lipogenesis, and lipolysis. Horm. Metab. Res. 2014, 46, 833–840. [Google Scholar] [CrossRef] [PubMed]
- Wurtz, P.; Wang, Q.; Kangas, A.J.; Richmond, R.C.; Skarp, J.; Tiainen, M.; Tynkkynen, T.; Soininen, P.; Havulinna, A.S.; Kaakinen, M.; et al. Metabolic signatures of adiposity in young adults: Mendelian randomization analysis and effects of weight change. PLoS Med. 2014, 11, e1001765. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Li, V.; Chan, G.C.K.; Phan, T.; Nudelman, A.S.; Xia, Z.; Storm, D.R. Adult type 3 adenylyl cyclase-deficient mice are obese. PLoS ONE 2009, 4, e6979. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.; Chen, X.; Yang, Y.; Storm, D.R. Disruption of type 3 adenylyl cyclase expression in the hypothalamus leads to obesity. Integr. Obes. Diabetes 2016, 28, 1304–1314. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Luo, J.; Leng, Y.; Yang, Y.; Zweifel, L.S.; Palmiter, R.D.; Storm, D.R. Ablation of type III adenylyl cyclase in mice causes reduced neuronal activity, altered sleep pattern, and depression-like phenotypes. Biol. Psychiatry 2016, 80, 836–848. [Google Scholar] [CrossRef] [PubMed]
- Qiu, L.; LeBel, R.P.; Storm, D.R.; Chen, X. Type 3 adenylyl cyclase: A key enzyme mediating the cAMP signaling in neuronal cilia. Int. J. Physiol. Pathophysiol. Pharmacol. 2016, 8, 95–108. [Google Scholar] [PubMed]
- You, P.; Hu, H.; Chen, Y.; Zhao, Y.; Yang, Y.; Wang, T.; Xing, R.; Shao, Y.; Zhang, W.; Li, D.; et al. Effects of melanocortin 3 and 4 receptor deficiency on energy homeostasis in rats. Sci. Rep. 2016, 6, 34938. [Google Scholar] [CrossRef] [PubMed]
- Tong, T.; Shen, Y.; Lee, H.-W.; Yu, R.; Park, T. Adenylyl cyclase 3 haploinsufficiency confers susceptibility to diet-induced obesity and insulin resistance in mice. Sci. Rep. 2016, 6, 34179. [Google Scholar] [CrossRef] [PubMed]
- Pitman, J.L.; Wheeler, M.C.; Lloyd, D.J.; Walker, J.R.; Glynne, R.J.; Gekakis, N. A gain-of-function mutation in adenylate cyclase 3 protects mice from diet-induced obesity. PLoS ONE 2014, 9, e110226. [Google Scholar] [CrossRef] [PubMed]
- Goni, L.; Cuervo, M.; Milagro, F.I.; Mart, J.A. Future Perspectives of Personalized Weight Loss Interventions Based on Nutrigenetic, Epigenetic, and Metagenomic Data. J. Nutr. 2016, 146, 905–912. [Google Scholar] [CrossRef] [PubMed]
| Moderately-High-Protein | Low-Fat | pa | pb | ||||
|---|---|---|---|---|---|---|---|
| Baseline (n = 72) | Change (n = 53) | Baseline (n = 75) | Change (n = 54) | ||||
| Age (years) | 45.6 (11.2) | - | 47.5 (8.6) | - | 0.28 | - | |
| Female sex | 48 (66.7) | - | 51 (68.0) | - | 0.87 | - | |
| Body weight (kg) c | 88.5 (13.2) | −7.6 (4.0) d | 88.8 (13.3) | −8.1 (4.1) d | 0.76 | 0.62 | |
| BMI (kg/m2) c | 31.7 (3.6) | −2.7 (1.4) d | 32.1 (3.8) | −2.9 (1.3) d | 0.56 | 0.71 | |
| WC (cm) c | 103.5 (11.2) | −8.4 (4.4) d | 103.8 (10.6) | −8.8 (4.5) d | 0.99 | 0.73 | |
| Body composition | |||||||
| Fat mass (%) c | 42.6 (9.1) | −4.8 (5.3) d | 41.9 (6.9) | −3.9 (2.5) d | 0.37 | 0.13 | |
| Lean mass (%) c | 54.2 (9.2) | 4.6 (5.4) d | 55.0 (6.6) | 3.6 (2.4) d | 0.34 | 0.12 | |
| Trunk fat (kg) c | 20.3 (5.0) | −4.1 (2.5) d | 20.6 (4.7) | −4.3 (2.4) d | 0.81 | 0.87 | |
| Android fat (kg) c | 3.6 (1.0) | −0.8 (0.5) d | 3.6 (0.9) | −0.9 (0.5) d | 0.78 | 0.98 | |
| Gynoid fat (kg) c | 5.9 (1.5) | −0.9 (0.7) d | 6.0 (1.7) | −1.0 (0.5) d | 0.74 | 0.84 | |
| Visceral fat (kg) c | 1.5 (0.9) | −0.5 (0.4) d | 1.7 (0.9) d | −0.5 (0.4) d | 0.37 | 0.92 | |
| AA (n = 35) | AG (n = 74) | GG (n = 38) | p | ||
|---|---|---|---|---|---|
| Age (years) | 50.9 (9.0) | 46.0 (9.5) | 43.8 (10.7) | 0.007 | |
| Sex | 0.16 | ||||
| Male | 11 (31.4) | 20 (27.0) | 17 (44.7) | ||
| Female | 24 (68.6) | 54 (73.0) | 21 (55.3) | ||
| Diet group | 0.84 | ||||
| Moderately-high-protein | 16 (45.7) | 36 (48.7) | 20 (52.6) | ||
| Low-fat | 19 (54.3) | 38 (51.3) | 18 (47.4) | ||
| Dietary intake per day | |||||
| Energy (kcal) | 2907 (849) | 3023 (855) | 3038 (1001) | 0.78 | |
| Protein (%) | 16.7 (3.4) | 16.5 (2.5) | 17.1 (3.1) | 0.55 | |
| Fat (%) | 38.8 (6.2) | 39.6 (6.0) | 40.0 (5.2) | 0.67 | |
| Carbohydrate (%) | 42.8 (8.5) | 41.9 (6.5) | 40.8 (7.0) | 0.52 | |
| Body weight (kg) | 88.2 (13.1) | 87.6 (12.6) | 91.0 (14.6) | 0.43 | |
| BMI (kg/m2) | 32.7 (3.9) | 31.6 (3.5) | 31.9 (3.6) | 0.34 | |
| WC (cm) | 105.9 (10.2) | 102.2 (10.5) | 104.5 (12.0) | 0.22 | |
| Body composition a | |||||
| Fat mass (%) | 42.1 (5.4) | 42.2 (8.9) | 42.4 (8.5) | 0.99 | |
| Lean mass (%) | 54.8 (5.1) | 54.7 (8.7) | 54.4 (8.5) | 0.98 | |
| Trunk fat (kg) | 20.7 (3.9) | 19.9 (4.8) | 21.2 (5.7) | 0.40 | |
| Android fat (kg) | 3.6 (0.8) | 3.5 (0.9) | 3.8 (1.2) | 0.30 | |
| Gynoid fat (kg) | 5.8 (1.5) | 6.0 (1.7) | 6.0 (1.6) | 0.87 | |
| Visceral fat (kg) | 1.7 (0.9) | 1.5 (0.9) | 1.7 (1.0) | 0.36 | |
| All Population | AA | AG | GG | p | ||
|---|---|---|---|---|---|---|
| Moderately-high protein diet a | ||||||
| Energy (kcal) | 1340 (286) | 1297 (180) | 1294 (238) | 1490 (414) | 0.11 | |
| Protein (%) | 28.9 (4.3) | 29.2 (3.5) | 29.7 (4.2) | 26.7 (4.4) | 0.12 | |
| Fat (%) | 30.8 (5.0) | 31.7 (5.4) | 30.1 (4.9) | 32.1 (5.1) | 0.43 | |
| Carbohydrate (%) | 42.5 (4.3) | 42.1 (3.7) | 41.9 (3.8) | 44.2 (5.7) | 0.27 | |
| Low-fat diet b | ||||||
| Energy (kcal) | 1324 (236) c | 1313 (274) | 1291 (231) | 1429 (185) | 0.29 | |
| Protein (%) | 22.3 (3.7) d | 22.0 (4.2) | 22.2 (3.2) | 22.8 (4.8) | 0.89 | |
| Fat (%) | 27.0 (5.7) d | 25.6 (8.3) | 27.4 (3.7) | 27.6 (6.6) | 0.61 | |
| Carbohydrate (%) | 53.3 (7.1) d | 54.5 (8.5) | 53.1 (6.5) | 52.1 (7.5) | 0.72 | |
| Moderately-High-Protein | Low-Fat | p Interaction | ||||
|---|---|---|---|---|---|---|
| β (SE) | p | β (SE) | p | |||
| Model 1 | ||||||
| Δ Weight (kg) | 0.95 (0.82) | 0.25 | −1.32 (0.69) | 0.06 | 0.02 | |
| Δ WC (cm) | 1.20 (0.95) | 0.21 | −0.88 (0.84) | 0.30 | 0.06 | |
| Δ Fat mass (kg) | 1.26 (0.76) | 0.10 | −0.95 (0.57) | 0.10 | 0.01 | |
| Δ Fat mass (%) | 1.65 (0.92) | 0.08 | −0.54 (0.44) | 0.22 | 0.03 | |
| Δ Lean mass (kg) | −0.54 (0.96) | 0.57 | −0.15 (0.30) | 0.60 | 0.58 | |
| Δ Lean mass (%) | −1.61 (0.90) | 0.08 | 0.48 (0.42) | 0.25 | 0.04 | |
| Δ Trunk fat (kg) | 0.89 (0.50) | 0.08 | −0.72 (0.41) | 0.08 | 0.008 | |
| Δ Android fat (kg) | 0.22 (0.11) | 0.049 | −0.18 (0.08) | 0.02 | 0.002 | |
| Δ Gynoid fat (kg) | 0.18 (0.14) | 0.20 | −0.19 (0.08) | 0.03 | 0.02 | |
| Δ Visceral fat (kg) | 0.08 (0.06) | 0.16 | −0.12 (0.05) | 0.02 | 0.005 | |
| Model 2 | ||||||
| Δ WC (cm) | 1.72 (0.94) | 0.07 | −0.61 (0.81) | 0.45 | 0.03 | |
| Δ Fat mass (kg) | 1.49 (0.78) | 0.06 | −1.08 (0.57) | 0.06 | 0.006 | |
| Δ Fat mass (%) | 2.21 (0.89) | 0.01 | −0.57 (0.43) | 0.20 | 0.01 | |
| Δ Lean mass (kg) | −0.41 (0.92) | 0.65 | −0.15 (0.30) | 0.63 | 0.52 | |
| Δ Lean mass (%) | −2.13 (0.84) | 0.01 | 0.50 (0.41) | 0.23 | 0.01 | |
| Δ Trunk fat (kg) | 1.15 (0.52) | 0.03 | −0.85 (0.40) | 0.04 | 0.006 | |
| Δ Android fat (kg) | 0.28 (0.11) | 0.01 | −0.19 (0.08) | 0.02 | 0.001 | |
| Δ Gynoid fat (kg) | 0.27 (0.14) | 0.05 | −0.20 (0.09) | 0.03 | 0.008 | |
| Δ Visceral fat (kg) | 0.10 (0.06) | 0.09 | −0.11 (0.05) | 0.02 | 0.005 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Goni, L.; Riezu-Boj, J.I.; Milagro, F.I.; Corrales, F.J.; Ortiz, L.; Cuervo, M.; Martínez, J.A. Interaction between an ADCY3 Genetic Variant and Two Weight-Lowering Diets Affecting Body Fatness and Body Composition Outcomes Depending on Macronutrient Distribution: A Randomized Trial. Nutrients 2018, 10, 789. https://doi.org/10.3390/nu10060789
Goni L, Riezu-Boj JI, Milagro FI, Corrales FJ, Ortiz L, Cuervo M, Martínez JA. Interaction between an ADCY3 Genetic Variant and Two Weight-Lowering Diets Affecting Body Fatness and Body Composition Outcomes Depending on Macronutrient Distribution: A Randomized Trial. Nutrients. 2018; 10(6):789. https://doi.org/10.3390/nu10060789
Chicago/Turabian StyleGoni, Leticia, Jose Ignacio Riezu-Boj, Fermín I. Milagro, Fernando J. Corrales, Lourdes Ortiz, Marta Cuervo, and J. Alfredo Martínez. 2018. "Interaction between an ADCY3 Genetic Variant and Two Weight-Lowering Diets Affecting Body Fatness and Body Composition Outcomes Depending on Macronutrient Distribution: A Randomized Trial" Nutrients 10, no. 6: 789. https://doi.org/10.3390/nu10060789
APA StyleGoni, L., Riezu-Boj, J. I., Milagro, F. I., Corrales, F. J., Ortiz, L., Cuervo, M., & Martínez, J. A. (2018). Interaction between an ADCY3 Genetic Variant and Two Weight-Lowering Diets Affecting Body Fatness and Body Composition Outcomes Depending on Macronutrient Distribution: A Randomized Trial. Nutrients, 10(6), 789. https://doi.org/10.3390/nu10060789






