Combining Airborne Laser Scanning and Aerial Imagery Enhances Echo Classification for Invasive Conifer Detection
Abstract
:1. Introduction
2. Materials and Methods
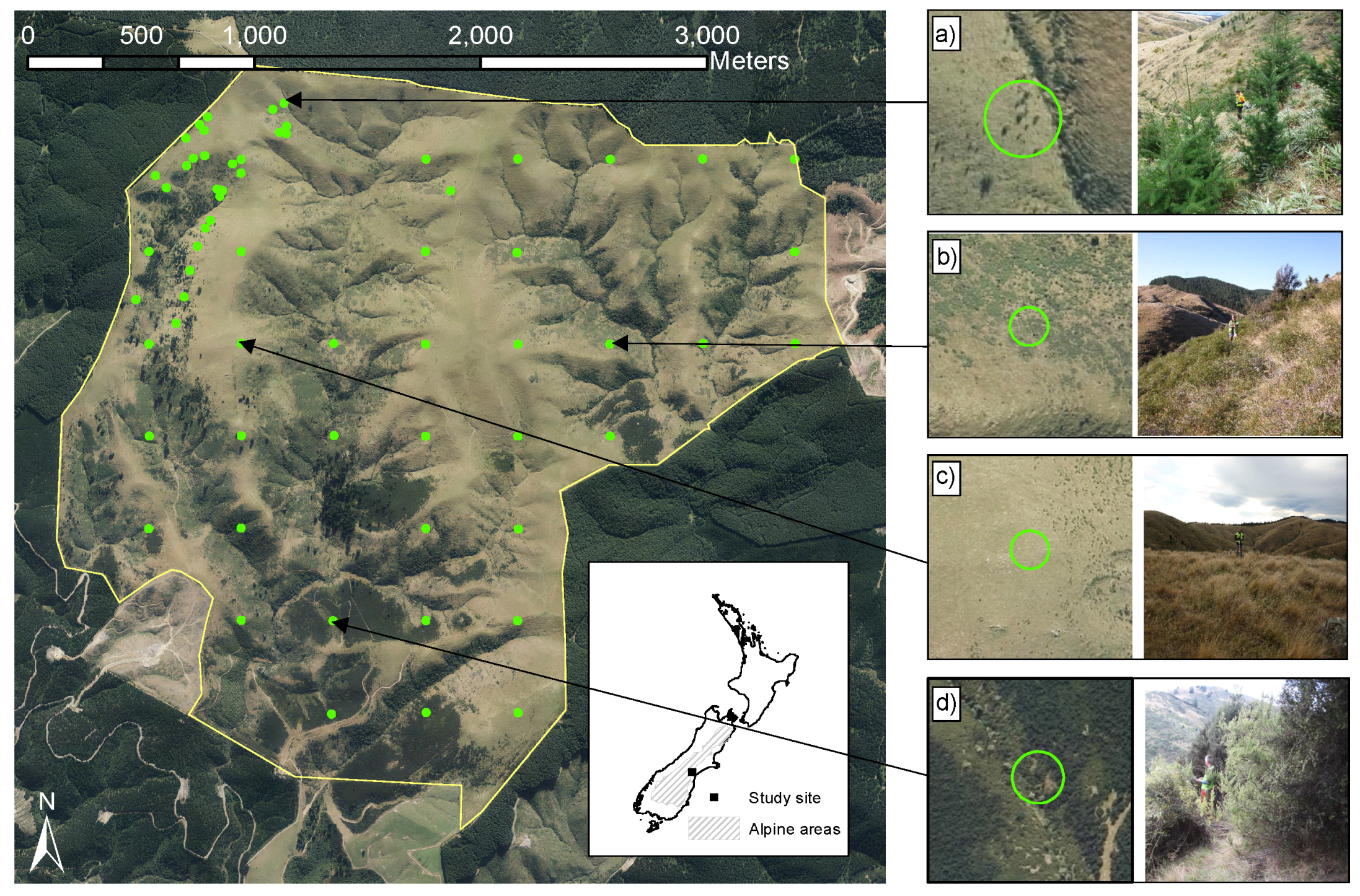
2.1. Study Site
2.2. Field Data
2.3. ALS Data
2.4. ALS Data Thinning
2.5. Aerial Imagery
2.6. Echo Classification
2.7. Random Forest
2.8. Accuracy Assessment
3. Results
3.1. Spectral and Structural Properties
3.2. Classification Accuracy
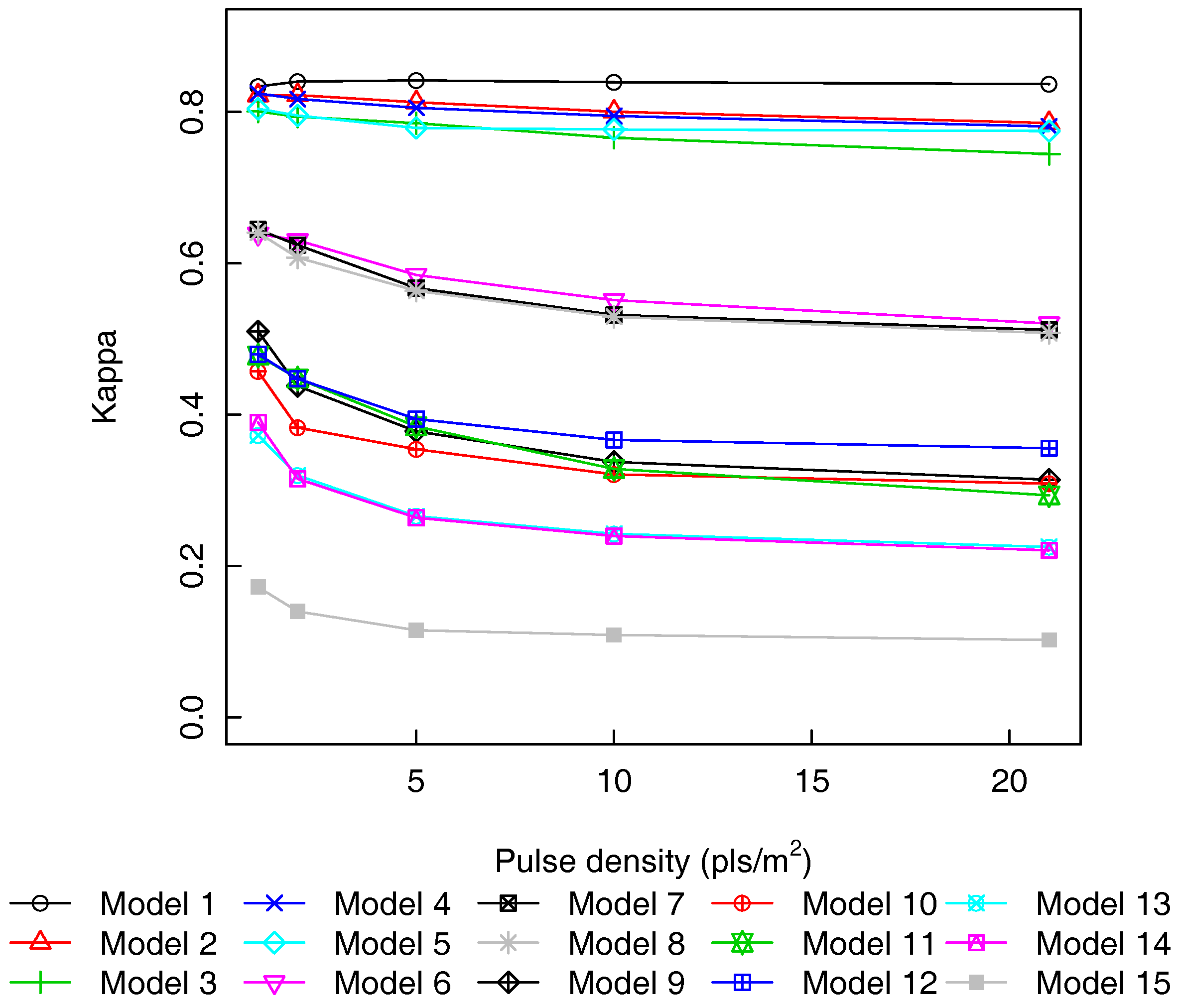
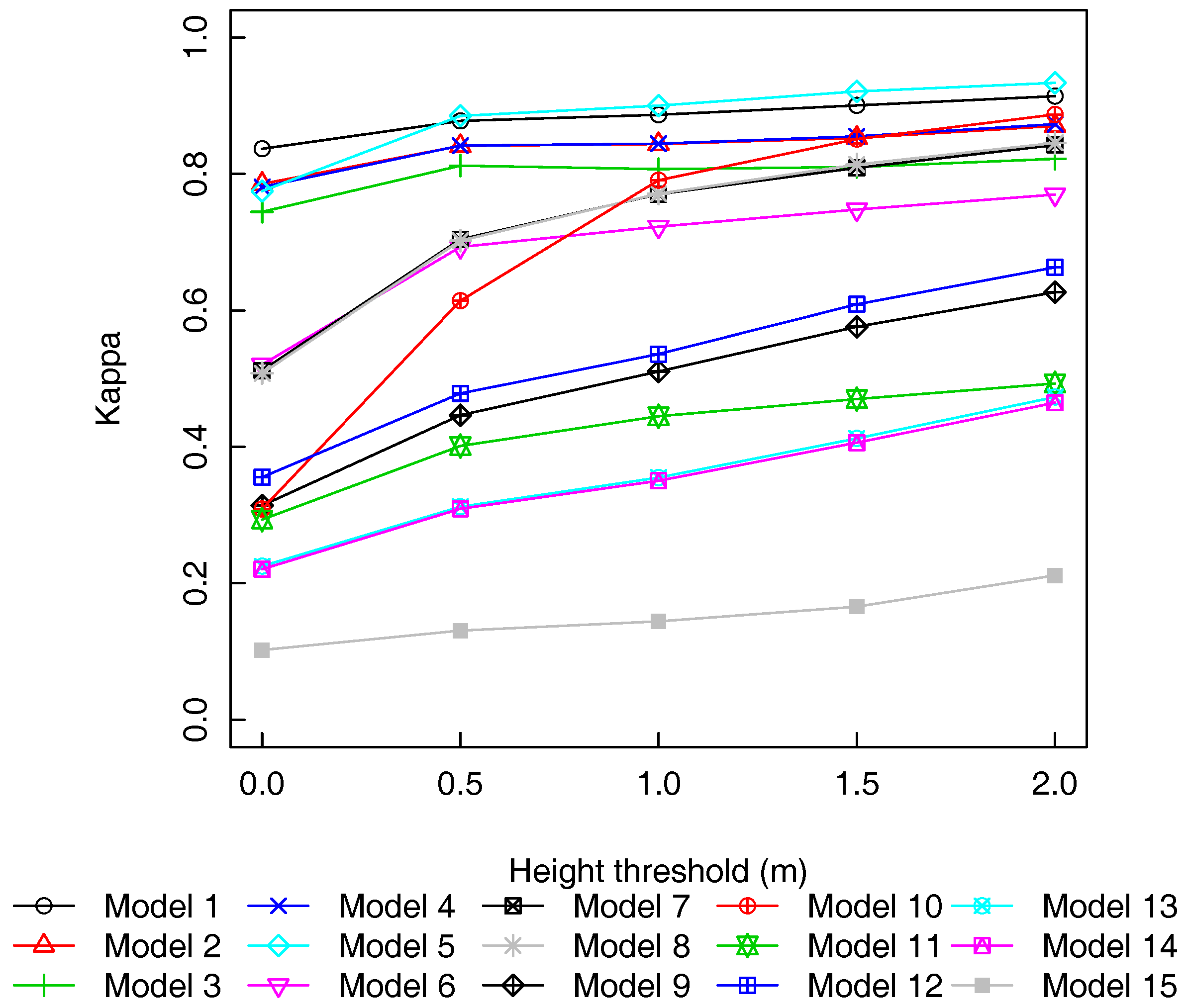
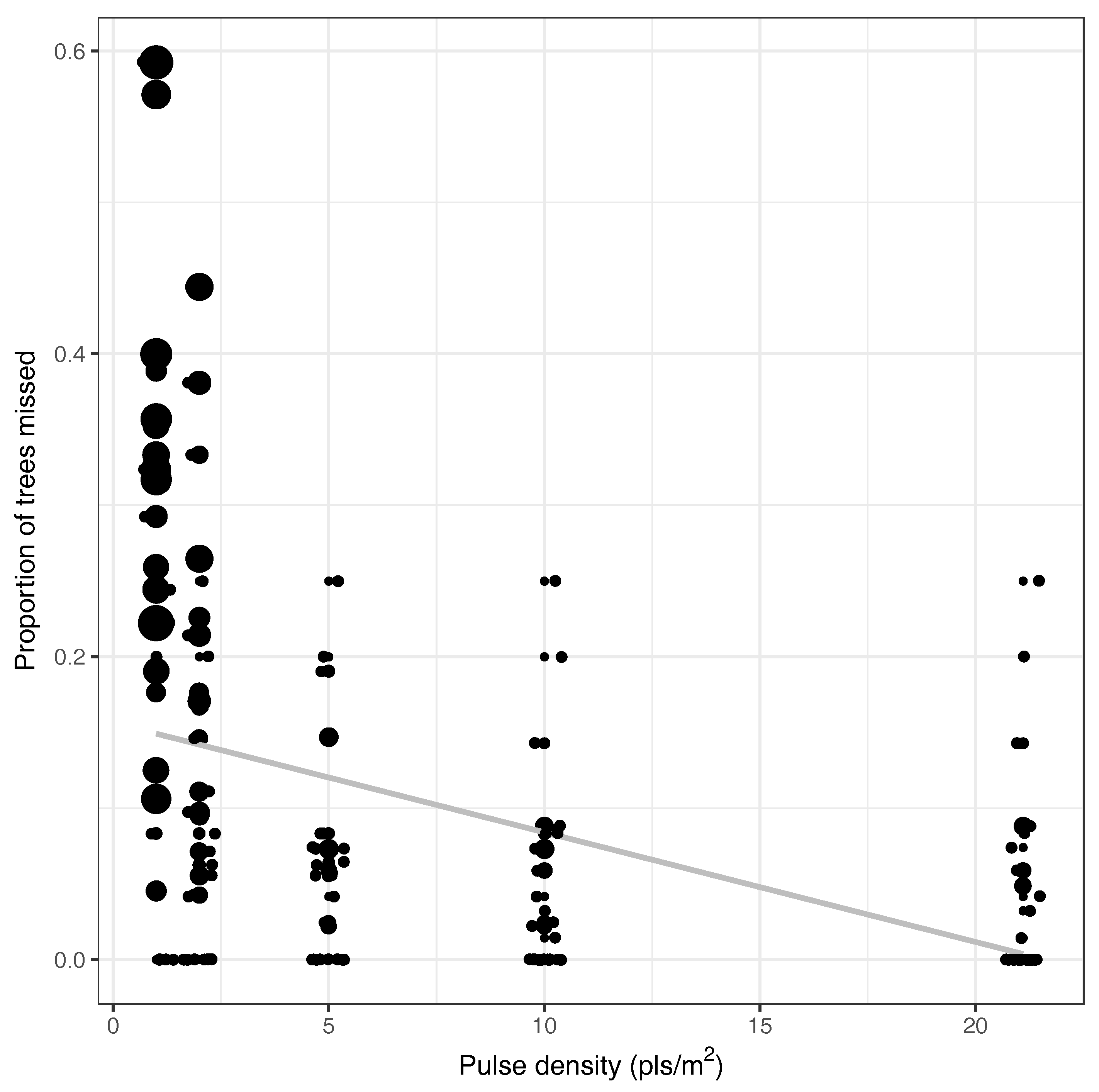
3.3. Pulse Density and Height Threshold
3.4. Leave One Plot Out Cross-Validation
3.5. Out-Of-Sample Errors
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| ALS | Airborne laser scanning |
| AUC | Area under the curve |
| AUCC | 95% confidence interval associated with AUC |
| CHM | Canopy height model |
| DTM | Digital terrain model |
| GSD | Ground surface distance |
| Kappa | Cohen’s kappa coefficient |
| KappaCI | 95% confidence interval associated with a kappa value |
| LiDAR | Light detection and ranging |
| LINZ | Land Information New Zealand |
| LOESS | Locally-weighted scatter plot smoothing |
| LOOCV | Leave one out cross-validation |
| LOPOCV | Leave one plot out cross-validation |
| MDA | Mean decrease in accuracy |
| NIR | Near-infrared |
| OOB | Out-of-bag |
| pls/m2 | Laser pulses emitted from the airborne laser scanner per m2 |
| pts/m2 | Points (or echoes) returned to the airborne sensor per m2 |
| RF | Random forest |
| ROC | Receiver operator characteristic |
| UCE | Ultra cam Eagle |
References
- Richardson, D.M.; Rejmánek, M. Conifers as invasive aliens: A global survey and predictive framework. Divers. Distrib. 2004, 10, 321–331. [Google Scholar] [CrossRef]
- Richardson, D.M.; Higgins, S.I. Pines as invaders of the southern hemisphere. In Ecology and Biogeography of Pinus; Richardson, D.M., Ed.; Cambridge University Press: Cambridge, UK, 1998; pp. 450–470. [Google Scholar]
- Howell, C.J.; McAlpine, K.G. Native plant species richness in non-native Pinus contorta forest. N. Z. J. Ecol. 2016, 40, 1. [Google Scholar]
- Ledgard, N.J. Wilding Control. Guidelines for the Control of Wilding Conifers; Scion: Rotorua, New Zealand, 2009. [Google Scholar]
- Anonymous. The Right Tree in the Right Place—New Zealand Wilding Conifer Management Strategy 2015–2030; Ministry for Primary Industries: Wellington, New Zealand, 2011.
- Velarde, S.J.; Paul, T.; Monge, J.; Yao, R. Cost Benefit Analysis of Wilding Conifer Management in New Zealand. Part I—Important Impacts Under Current Management; Report S0013; Scion: Bloxham, UK, 2015. [Google Scholar]
- Ledgard, N. Wilding control guidelines for farmers and land managers. N. Z. Plant Prot. 2009, 62, 380–386. [Google Scholar]
- Froude, V.A. Wilding Conifers in New Zealand: Beyond the Status Report; Report Prepared for the Ministry of Agriculture and Forestry, Pacific Eco-Logic, Bay of Islands, 44; Ministry of Agriculture and Forestry: Wellington, New Zealand, 2011.
- Clifford, V.; Paul, T.; Pearce, G. Quantifying the Change in High Country Fire Hazard From Wilding Trees; Report Prepared for Rural Fire New Zealand; New Zealand Fire Service Commission: Wellington, New Zealand, 2013. [Google Scholar]
- Ledgard, N.; Paul, T. Vegetation successions over 30 years of high country grassland invasion by Pinus contorta. N. Z. Plant Prot. 2008, 61, 98–104. [Google Scholar]
- Buckley, Y.M.; Brockerhoff, E.; Langer, L.; Ledgard, N.; North, H.; Rees, M. Slowing down a pine invasion despite uncertainty in demography and dispersal. J. Appl. Ecol. 2005, 42, 1020–1030. [Google Scholar] [CrossRef]
- Woods, D. The highs and lows of wilding conifer control operations: The good, the bad and the ugly! In Managing Wilding Conifers in New Zealand—Present and Future, Proceedings of the NZ Plant Protection Society Workshop, Christchurch, New Zealand, 11 August 2003; Hill, R.L., Zydenbos, S.M., Bezar, C.M., Eds.; New Zealand Plant Protection Society: Christchurch, New Zealand, 2003; pp. 55–63. [Google Scholar]
- Cochrane, P.; Grove, P. Exotic Wilding Conifer Spread Within Defined Areas of Canterbury High Counrty; Environment Canterbury: Prebbleton, New Zealand, 2013. [Google Scholar]
- Mureriwa, N.; Adam, E.; Sahu, A.; Tesfamichael, S. Spectral Discrimination of Prosopis Glandulosa (Mesquite) in Arid Environment of South Africa: Testing the Utility of in Situ Hyperspectral Data and Guided Regularized Random Forest Algorithm; Asian Association on Remote Sensing: Manila, Philippines, 2015. [Google Scholar]
- Hestir, E.L.; Khanna, S.; Andrew, M.E.; Santos, M.J.; Viers, J.H.; Greenberg, J.A.; Rajapakse, S.S.; Ustin, S.L. Identification of invasive vegetation using hyperspectral remote sensing in the California Delta ecosystem. Remote Sens. Environ. 2008, 112, 4034–4047. [Google Scholar] [CrossRef]
- HALL, S.J.; ASNER, G.P. Biological invasion alters regional nitrogen-oxide emissions from tropical rainforests. Glob. Chang. Biol. 2007, 13, 2143–2160. [Google Scholar] [CrossRef]
- Underwood, E.; Ustin, S.; DiPietro, D. Mapping nonnative plants using hyperspectral imagery. Remote Sens. Environ. 2003, 86, 150–161. [Google Scholar] [CrossRef]
- Niphadkar, M.; Nagendra, H. Remote sensing of invasive plants: incorporating functional traits into the picture. Int. J. Remote Sens. 2016, 37, 3074–3085. [Google Scholar] [CrossRef]
- Næsset, E.; Nelson, R. Using airborne laser scanning to monitor tree migration in the boreal–alpine transition zone. Remote Sens. Environ. 2007, 110, 357–369. [Google Scholar] [CrossRef]
- Stumberg, N.; Ørka, H.O.; Bollandsås, O.M.; Gobakken, T.; Næsset, E. Classifying tree and nontree echoes from airborne laser scanning in the forest–tundra ecotone. Can. J. Remote Sens. 2013, 38, 655–666. [Google Scholar] [CrossRef]
- Hantson, W.; Kooistra, L.; Slim, P.A. Mapping invasive woody species in coastal dunes in the Netherlands: A remote sensing approach using LIDAR and high-resolution aerial photographs. Appl. Veg. Sci. 2012, 15, 536–547. [Google Scholar] [CrossRef]
- Bork, E.W.; Su, J.G. Integrating LIDAR data and multispectral imagery for enhanced classification of rangeland vegetation: A meta analysis. Remote Sens. Environ. 2007, 111, 11–24. [Google Scholar] [CrossRef]
- Rees, W.G. Characterisation of Arctic treelines by LiDAR and multispectral imagery. Polar Rec. 2007, 43, 345–352. [Google Scholar] [CrossRef]
- Thieme, N.; Martin Bollandsås, O.; Gobakken, T.; Næsset, E. Detection of small single trees in the forest–tundra ecotone using height values from airborne laser scanning. Can. J. Remote Sens. 2011, 37, 264–274. [Google Scholar] [CrossRef]
- Stumberg, N.; Bollandsås, O.M.; Gobakken, T.; Næsset, E. Automatic detection of small single trees in the Forest-Tundra Ecotone using airborne laser scanning. Remote Sens. 2014, 6, 10152–10170. [Google Scholar] [CrossRef]
- Næsset, E. Discrimination between Ground vegetation and small pioneer trees in the Boreal-Alpine Ecotone using intensity metrics derived from airborne laser scanner data. Remote Sens. 2016, 8, 548. [Google Scholar] [CrossRef]
- Thomas, V.; Treitz, P.; McCaughey, J.H.; Morrison, I. Mapping stand-level forest biophysical variables for a mixedwood boreal forest using LiDAR: An examination of scanning density. Can. J. For. Res. 2006, 36, 34–47. [Google Scholar] [CrossRef]
- Morsdorf, F.; Frey, O.; Meier, E.; Itten, K.I.; Allgöwer, B. Assessment of the influence of flying altitude and scan angle on biophysical vegetation products derived from airborne laser scanning. Int. J. Remote Sens. 2008, 29, 1387–1406. [Google Scholar] [CrossRef]
- Wilkes, P.; Jones, S.; Suarez, L.; Haywood, A.; Woodgate, W.; Soto-Berelov, M.; Mellor, A.; Skidmore, A. Understanding the effects of als pulse density for metric retrieval across diverse forest types. Photogramm. Eng. Remote Sens. 2015, 81, 625–635. [Google Scholar] [CrossRef]
- Keranen, J.; Maltamo, M.; Packalen, P. Effect of flying altitude, scanning angle and scanning mode on the accuracy of ALS based forest inventory. Int. J. Appl. Earth Obs. Geoinf. 2016, 52, 349–360. [Google Scholar] [CrossRef]
- Maltamo, M.; Eerikäinen, K.; Packalén, P.; Hyyppä, J. Estimation of stem volume using laser scanning-based canopy height metrics. Forestry 2006, 79, 217–229. [Google Scholar] [CrossRef]
- Gobakken, T.; Næsset, E. Assessing effects of laser point density, ground sampling intensity, and field sample plot size on biophysical stand properties derived from airborne laser scanner data. Can. J. For. Res. 2008, 38, 1095–1109. [Google Scholar] [CrossRef]
- Watt, M.S.; Adams, T.; Gonzalez Aracil, S.; Marshall, H.; Watt, P. The influence of LiDAR pulse density and plot size on the accuracy of New Zealand plantation stand volume equations. N. Z. J. For. Sci. 2013, 43, 1–10. [Google Scholar] [CrossRef]
- Khosravipour, A.; Skidmore, A.K.; Isenburg, M.; Wang, T.; Hussin, Y.A. Generating pit-free canopy height models from airborne LiDAR. Photogramm. Eng. Remote Sens. 2014, 80, 863–872. [Google Scholar] [CrossRef]
- Hauglin, M.; Næsset, E. Detection and segmentation of small trees in the Forest-Tundra Ecotone using airborne laser scanning. Remote Sens. 2016, 8, 407. [Google Scholar] [CrossRef]
- Taylor, K.T.; Maxwell, B.D.; Pauchard, A.; Nuñez, M.A.; Peltzer, D.A.; Terwei, A.; Rew, L.J. Drivers of plant invasion vary globally: Evidence from pine invasions within six ecoregions. Glob. Ecol. Biogeogr. 2015, 25, 96–106. [Google Scholar] [CrossRef]
- Reese, H.; Nyström, M.; Nordkvist, K.; Olsson, H. Combining airborne laser scanning data and optical satellite data for classification of alpine vegetation. Int. J. Appl. Earth Obs. Geoinf. 2014, 27, 81–90. [Google Scholar] [CrossRef]
- Hauglin, M.; Ørka, H.O. Discriminating between Native Norway Spruce and Invasive Sitka Spruce—A comparison of multitemporal Landsat 8 imagery, aerial images and airborne laser scanner data. Remote Sens. 2016, 8, 363. [Google Scholar] [CrossRef]
- Liu, X. Airborne LiDAR for DEM generation: Some critical issues. Prog. Phys. Geogr. 2008, 32, 31–49. [Google Scholar]
- Baltsavias, E. Airborne laser scanning: basic relations and formulas. ISPRS J. Photogramm. Remote Sens. 1999, 54, 199–214. [Google Scholar] [CrossRef]
- Jakubowski, M.K.; Guo, Q.; Kelly, M. Tradeoffs between LiDAR pulse density and forest measurement accuracy. Remote Sens. Environ. 2013, 130, 245–253. [Google Scholar] [CrossRef]
- Millman, K.J.; Aivazis, M. Python for Scientists and Engineers. Comput. Sci. Eng. 2011, 13, 9–12. [Google Scholar] [CrossRef]
- Silva, C.A.; Crookston, N.L.; Hudak, A.T.; Vierling, L.A. rLiDAR: LiDAR Data Processing and Visualization, R package version 0.1. 2015.
- Wing, B.M.; Ritchie, M.W.; Boston, K.; Cohen, W.B.; Gitelman, A.; Olsen, M.J. Prediction of understory vegetation cover with airborne LiDAR in an interior ponderosa pine forest. Remote Sens. Environ. 2012, 124, 730–741. [Google Scholar] [CrossRef]
- Kaufman, Y.J.; Tanre, D. Atmospherically resistant vegetation index (ARVI) for EOS-MODIS. IEEE Trans. Geosci. Remote Sens. 1992, 30, 261–270. [Google Scholar] [CrossRef]
- Packalen, P.; Suvanto, A.; Maltamo, M. A Two Stage Method to estimate species-specific growing stock. Photogramm. Eng. Remote Sens. 2009, 75, 1451–1460. [Google Scholar] [CrossRef]
- Packalen, P.; Maltamo, M. Predicting the plot volume by tree species using airborne laser scanning and aerial photographs. For. Sci. 2006, 52, 611–622. [Google Scholar]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Mellor, A.; Haywood, A.; Stone, C.; Jones, S. The performance of random forests in an gperational settingfor large area sclerophyll forest classification. Remote Sens. 2013, 5, 2838–2856. [Google Scholar] [CrossRef]
- Dash, J.P.; Marshall, H.M.; Rawley, B. Methods for estimating multivariate stand yields and errors using k-NN and aerial laser scanning. Forestry 2015, 88, 237–247. [Google Scholar] [CrossRef]
- Dash, J.P.; Watt, M.S.; Bhandari, S.; Watt, P. Characterising forest structure using combinations of airborne laser scanning data, RapidEye satellite imagery and environmental variables. Forestry 2016, 89, 159–169. [Google Scholar] [CrossRef]
- Watt, M.S.; Dash, J.P.; Bhandari, S.; Watt, P. Comparing parametric and non-parametric methods of predicting Site Index for radiata pine using combinations of data derived from environmental surfaces, satellite imagery and airborne laser scanning. For. Ecol. Manag. 2015, 357, 1–9. [Google Scholar] [CrossRef]
- Watt, M.S.; Dash, J.P.; Watt, P.; Bhandari, S. Multi-sensor modelling of a forest productivity index for radiata pine plantations. N. Z. J. For. Sci. 2016, 46, 1–14. [Google Scholar] [CrossRef]
- Cutler, D.R.; Edwards, T.C.; Beard, K.H.; Cutler, A.; Hess, K.T.; Gibson, J.; Lawler, J.J. Random forests for classification in ecology. Ecology 2007, 88, 2783–2792. [Google Scholar] [CrossRef] [PubMed]
- Criminisi, A.; Konukoglu, E.; Shotton, J. Decision Forests: A Unified Framework for Classification, Regression, Density Estimation, Manifold Learning and Semi-Supervised Learning; NOW Publishers: New York, NY, USA, 2012. [Google Scholar]
- Wright, M.N. Ranger: A Fast Implementation of Random Forests, R package version 0.5.0. 2016.
- Cohen, J. A Coefficient of Agreement for Nominal Scales. Educ. Psychol. Meas. 1960, 20, 37–46. [Google Scholar] [CrossRef]
- Revelle, W. Psych: Procedures for Psychological, Psychometric, and Personality Research, R package version 1.6.12; Northwestern University: Evanston, IL, USA, 2016. [Google Scholar]
- Richard, J.; Landis, G.G.K. The measurement of observer agreement for categorical data. Biometrics 1977, 33, 159–174. [Google Scholar]
- Fleiss, J.; Cohen, J.; Everitt, B. Large sample standard errors of kappa and weighted kappa. Psychol. Bull. 1969, 72, 323–327. [Google Scholar] [CrossRef]
- Robin, X.; Turck, N.; Hainard, A.; Tiberti, N.; Lisacek, F.; Sanchez, J.C.; Müller, M. pROC: An open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform. 2011, 12, 77. [Google Scholar] [CrossRef] [PubMed]
- Holmgren, J.; Persson, A.; Soderman, U. Species identification of individual trees by combining high resolution LiDAR data with multi-spectral images. Int. J. Remote Sens. 2008, 29, 1537–1552. [Google Scholar] [CrossRef]
- Dalponte, M.; Bruzzone, L.; Gianelle, D. Tree species classification in the Southern Alps based on the fusion of very high geometrical resolution multispectral/hyperspectral images and LiDAR data. Remote Sens. Environ. 2012, 123, 258–270. [Google Scholar] [CrossRef]
- Ghosh, A.; Fassnacht, F.E.; Joshi, P.; Koch, B. A framework for mapping tree species combining hyperspectral and LiDAR data: Role of selected classifiers and sensor across three spatial scales. Int. J. Appl. Earth Obs. Geoinf. 2014, 26, 49–63. [Google Scholar] [CrossRef]
- Goodwin, N.R.; Coops, N.C.; Culvenor, D.S. Assessment of forest structure with airborne LiDAR and the effects of platform altitude. Remote Sens. Environ. 2006, 103, 140–152. [Google Scholar] [CrossRef]
- Lovell, J.L.; Jupp, D.L.; Culvenor, D.S.; Coops, N.C. Using airborne and ground-based ranging LiDAR to measure canopy structure in Australian forests. Can. J. Remote Sens. 2003, 29, 607–622. [Google Scholar] [CrossRef]
- Disney, M.; Kalogirou, V.; Lewis, P.; Prieto-Blanco, A.; Hancock, S.; Pfeifer, M. Simulating the impact of discrete-return LiDAR system and survey characteristics over young conifer and broadleaf forests. Remote Sens. Environ. 2010, 114, 1546–1560. [Google Scholar] [CrossRef]
- Gates, D.M.; Keegan, H.J.; Schleter, J.C.; Weidner, V.R. Spectral Properties of Plants. Appl. Opt. 1965, 4, 11–20. [Google Scholar] [CrossRef]
- Slaton, M.R.; Raymond Hunt, E.; Smith, W.K. Estimating near-infrared leaf reflectance from leaf structural characteristics. Am. J. Bot. 2001, 88, 278–284. [Google Scholar] [CrossRef] [PubMed]
- Williams, D.L. A comparison of spectral reflectance properties at the needle, branch, and canopy level for selected Conifer species. Remote Sens. Environ. 1991, 35, 79–93. [Google Scholar] [CrossRef]
- Nyström, M.; Holmgren, J.; Olsson, H. Change detection of mountain birch using multi-temporal ALS point clouds. Remote Sens. Lett. 2013, 4, 190–199. [Google Scholar] [CrossRef]
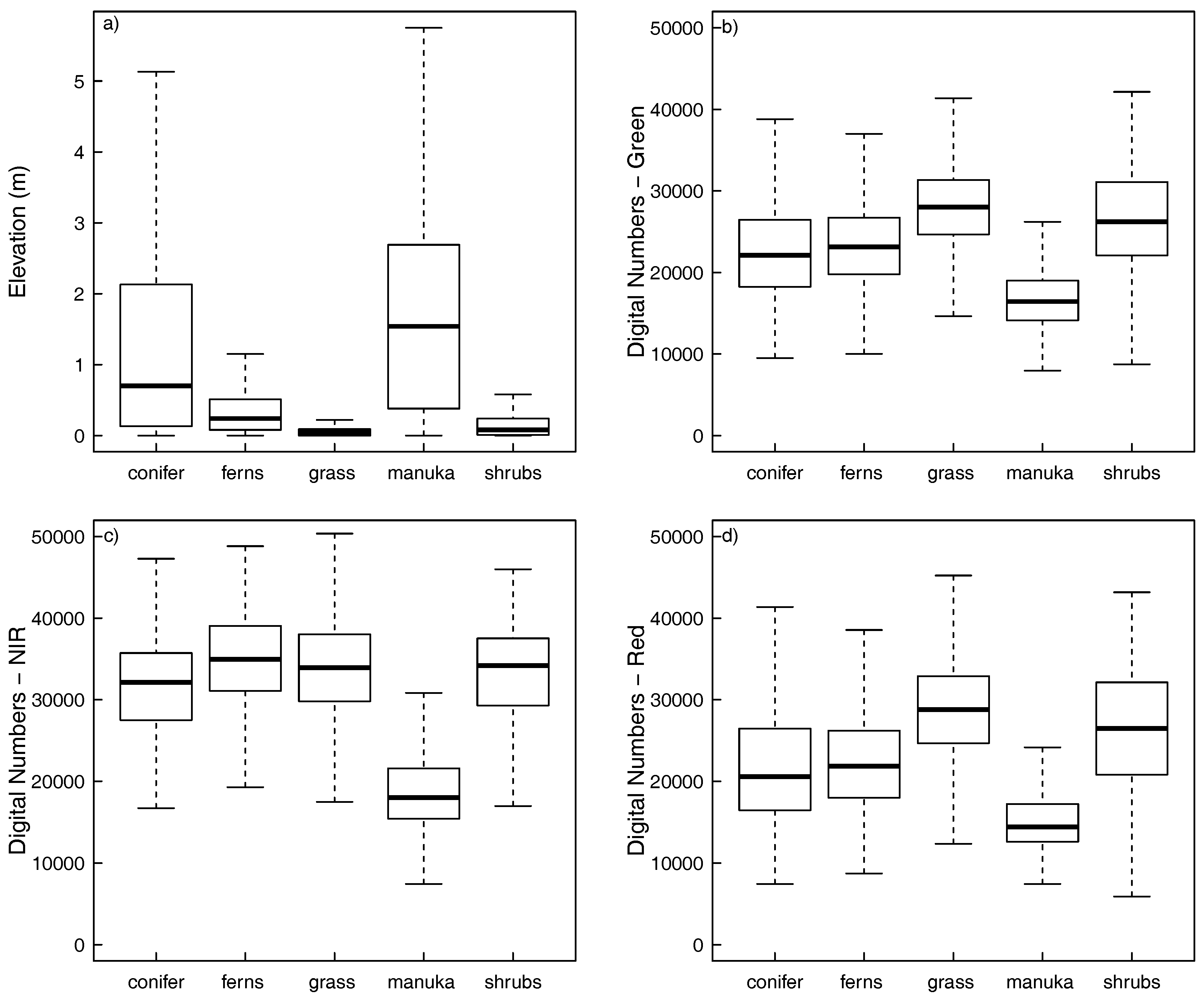


| Species | Count | Mean Height (m) | Mean Diameter (mm) |
|---|---|---|---|
| Ps. menz. | 813 | 1.72 (0.05–12.90) | 28.18 (4–250) |
| P. muricata | 8 | 1.99 (1.02–3.94) | 22.16 (7–40) |
| P. radiata | 4 | 2.00 (1.28–2.70) | 20.50 (5–41) |
| Variable | Value |
|---|---|
| Scanner | Riegl Q1560, 2 channel |
| Laser pulse rate | 330 kHz |
| Max scan angle | 14° |
| Echo types | 1st, 2nd, 3rd, …, 7th and last |
| File format | LAS 1.4 |
| Map projection | NZTM2000 |
| Horizontal datum | NZGD2000 |
| Vertical datum | NZVD2009 |
| Mean pulse density | 21.1 pls/m2 |
| Mean echo density | 36.5 pts/m2 |
| Pulse spacing | 0.22 m |
| Target Pulse Density (pls/m2) | Realised Point Density (pts/m2) | Realised Point Spacing (m) | Realised Pulse Density (pls/m2) | Realised Pulse Spacing (m) |
|---|---|---|---|---|
| 1 | 1.8 | 0.8 | 1.1 | 1 |
| 2 | 3.7 | 0.5 | 2.1 | 0.7 |
| 5 | 9.1 | 0.3 | 5.3 | 0.4 |
| 10 | 18.2 | 0.2 | 10.6 | 0.3 |
| unthinned | 36.5 | 0.2 | 21.1 | 0.2 |
| Variable | Value |
|---|---|
| Radiometric resolution | 32-bit colour (4 × 8 bits per band) |
| Spectral resolution | Red, green, blue, near-infrared |
| Pixel resolution | 0.3 m GSD |
| Spatial accuracy | ±4 m at the 95% confidence interval in the clear open space (2 sigma) over the area of interest |
| Data format | GeoTiff with associated world file (TFW) |
| Forward overlap | 60% (min 54%) |
| Side overlap | 30% (min 15%) |
| Model | Predictor Variables | Statistics | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifier | Elevation | Intensity | NIR | Red | Green | Kappa | KappaCI | AUC | AUCCI |
| 1 | * | * | * | * | * | 0.837 | 0.835–0.839 | 0.885 | 0.884–0.887 |
| 2 | * | * | * | * | 0.785 | 0.782–0.788 | 0.856 | 0.854–0.858 | |
| 3 | * | * | * | * | 0.744 | 0.741–0.747 | 0.828 | 0.826–0.829 | |
| 4 | * | * | * | * | 0.781 | 0.777–0.783 | 0.854 | 0.851–0.855 | |
| 5 | * | * | * | * | 0.773 | 0.771–0.776 | 0.849 | 0.847–0.850 | |
| 6 | * | * | * | 0.521 | 0.517–0.524 | 0.698 | 0.695–0.699 | ||
| 7 | * | * | * | 0.513 | 0.508–0.515 | 0.693 | 0.691–0.694 | ||
| 8 | * | * | * | 0.508 | 0.504–0.511 | 0.691 | 0.689–0.693 | ||
| 9 | * | * | * | 0.316 | 0.313–0.321 | 0.602 | 0.601–0.604 | ||
| 10 | * | * | * | 0.308 | 0.306–0.313 | 0.599 | 0.597–0.600 | ||
| 11 | * | * | 0.292 | 0.288–0.295 | 0.597 | 0.595–0.598 | |||
| 12 | * | * | 0.355 | 0.352–0.359 | 0.625 | 0.623–0.626 | |||
| 13 | * | * | 0.224 | 0.220–0.227 | 0.571 | 0.569–0.572 | |||
| 14 | * | * | 0.221 | 0.218–0.225 | 0.569 | 0.568–0.571 | |||
| 15 | * | 0.101 | 0.099–0.104 | 0.529 | 0.528–0.531 | ||||
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dash, J.P.; Pearse, G.D.; Watt, M.S.; Paul, T. Combining Airborne Laser Scanning and Aerial Imagery Enhances Echo Classification for Invasive Conifer Detection. Remote Sens. 2017, 9, 156. https://doi.org/10.3390/rs9020156
Dash JP, Pearse GD, Watt MS, Paul T. Combining Airborne Laser Scanning and Aerial Imagery Enhances Echo Classification for Invasive Conifer Detection. Remote Sensing. 2017; 9(2):156. https://doi.org/10.3390/rs9020156
Chicago/Turabian StyleDash, Jonathan P., Grant D. Pearse, Michael S. Watt, and Thomas Paul. 2017. "Combining Airborne Laser Scanning and Aerial Imagery Enhances Echo Classification for Invasive Conifer Detection" Remote Sensing 9, no. 2: 156. https://doi.org/10.3390/rs9020156
APA StyleDash, J. P., Pearse, G. D., Watt, M. S., & Paul, T. (2017). Combining Airborne Laser Scanning and Aerial Imagery Enhances Echo Classification for Invasive Conifer Detection. Remote Sensing, 9(2), 156. https://doi.org/10.3390/rs9020156





