Hierarchical Segmentation Framework for Identifying Natural Vegetation: A Case Study of the Tehachapi Mountains, California
Abstract
: Two critical limitations of very high resolution imagery interpretations for time-series analysis are higher imagery variances and large data sizes. Although object-based analyses with a multi-scale framework for diverse object sizes are one potential solution, more data requirements and large amounts of testing at high costs are required. In this study, I applied a three-level hierarchical vegetation framework for reducing those costs, and a three-step procedure was used to evaluate its effects on a digital orthophoto quadrangles with 1 m spatial resolution. Step one and step two were for image segmentation optimized for delineation of tree density, which involved global Otsu’s method followed by the random walker algorithm. Step three was for detailed species delineations, which were derived from multiresolution segmentation, in two test areas. Step one and step two were able to delineating tree density segments and label species association robustly, compared to previous hierarchical frameworks. However, step three was limited by less image information to produce detailed, reasonable image objects with optimal scale parameters for species labeling. This hierarchical vegetation framework has potential to develop baseline data for evaluating climate change impacts on vegetation at lower cost using widely available data and a personal laptop.1. Introduction
1.1. Motivation
Vegetation mapping is required for biological conservation and forest inventory; especially time series mapping is commonly used to detect transformations of species or suitable habitats (e.g., [1–3]) and evaluate impacts of climate change on species (e.g., [4,5]). Species distribution models (SDMs, also called habitat suitability models), which correlate environment variables with species sampling data to map species occurrences [6], have been used for vegetation mapping. However, the field-based approach is cost and labor-consuming, while pure remote sensing imagery interpretations are lacking in ability to detect individual species [7]. SDMs are viewed as static and equilibrium models capturing species-environment relations at large scales, but ignoring dynamic biological interactions, such as dispersal, migration, facilitation, competition, mutualism, and predation at local scales [8]. Mapping species directly using very high resolution imagery (1 to 2 m), and tracking vegetation transformations over time through repeated mapping provided the potential for building more dynamic vegetation mapping.
With advances of sensors, spatial resolutions of remotely sensed data have been improved to centimeter levels and provide more spatial information on species distributions. For example, WorldView-1, which was launched on 18 September 2007, collects panchromatic imagery at 0.5 m, and WorldView-2, which was launched on 8 October 2007, collects panchromatic imagery at 0.46 m and multispectral imagery at 1.84 m. GeoEye-1, which was launched on 6 September 2008, collects panchromatic imagery at 0.41 m and multispectral imagery at 1.65 m. As Nagendra and Rocchini [9] indicated, very high resolution imagery, which spatial resolution is fewer than 5 m, but only with four to eight bands or less bands, provides more accurate locations of tree canopies. The ability is better than hyperspectral imagery, which has a series of continuous spectral bands (over 100 bands), but coarser spatial resolution (20–30 at best). However, higher spatial resolutions may not have higher classification accuracies, due to higher variances within classes [10] (also called H-resolution problem, which occurs when land cover elements are larger than pixel sizes [11]). In addition, more pixels in very high resolution imagery result in large data sizes, and increase loadings of computer hardware requirements (i.e., segmentation procedures have very high memory and CPU requirements), so most studies only test in small areas, because of memory limitations of personal computers and long processing times. Object-based analysis can improve salt and pepper effects and increase classification accuracies over pixel-based image classifications, which ignore similarity of near pixels [12–18]. In addition, imagery can be stratified into smaller subsets [19] within the processing capabilities of personal computers.
Object-based image classification includes a two-step procedure, image segmentation and image classification. Image segmentation gathers several similar neighbor pixels together as objects, and image classification categorizes or labels objects as land cover types. In theory, image objects have equal internal variances at a common scale [20], and very high resolution imagery is helpful to derive correct locations. However, each class with varying sizes needs different scales to define appropriate objects [14,20,21], and multispectral bands are helpful to labeling or classification procedures.
The appropriate scale for identifying objects can be found by an iterative workflow [22] using hierarchical semantic models or knowledge [23]. In other words, there is no single optimal scale parameter, but a spatially-nested (multi-scale) structure [20–24] can be used to identify objects with different sizes and describe the object traits, especially in imagery without many spectral bands. For example, Yu et al., [18] pointed out vegetation alliances, which are more general than species types, are also critical to define tree species segmentation, and the similarity of near objects may decrease classification accuracies. Kim et al., [25] indicated that multiscale image classification involving both spectral and texture traits can increase classification accuracies. Nevertheless, those hierarchical-segmentation applications in very high resolution imagery are still limited to classify primary land cover categories [25] or estimate forest parameters [26–28] in small test areas, and there is lacking in a general framework for specific species in large areas.
Previous vegetation mapping frameworks have taken two approaches, the data-based orientation and vegetation-type orientation. In the data-based approach, integrating multiple remote sensing data layers can increase interpretations of vegetation traits [25,26,28,29]. In one example, Xie et al., [29] used this approach to identify exotic Australian pine with three-level segmentations: NDVI to distinguish vegetation from non-vegetation, tree heights, derived from LIDAR, to distinguish trees from short trees (lower than 3 m), shrubs and grasses, which did not belong to Australian Pine, and image traits using shape/color and smooth/compactness weighting parameters for target objects. However, higher data requirements raise costs, and may not be extensively adapted to other regions. In the vegetation-type approach, vegetation nested structures from tree, stands, forest types (e.g., pine, oak and red fir) and vegetation types (e.g., wetland, forest and grassland) are another solution [20]. This framework, incorporating object-based image segmentation [20] and labeling procedures [30], can be used to identify more general vegetation life form or land-cover types (e.g., conifer forest, hardwood forest, chaparral, soft chaparral) and within them, more specific vegetation types. As an example, the level of Associations (e.g., Jeffrey pine, black oak and coast live oak) is nested under the US National Vegetation Classification Systems ( http://usnvc.org). However, the labeling procedures for each individual species type, based on multispectral classification and image interpretation, may not all reach equally high accuracies (e.g., accuracies for conifer types ranged from 23% to 100% in Franklin et al., [30], but ecological studies may need to focus on certain species with lower accuracies).
1.2. Problem Statement and Literature Review
The goal of this study was to implement a hierarchical framework for segmenting specific tree species using three bands derived from 1 m digital orthophoto quadrangles (DOQ) and an evaluative framework to assess the segment results. For the purpose of time-series image processing, imagery, which governments started to conduct aerial surveys for, with only one single black/white band is the most widely available historical (long-term) sources. Thus, this study focused on the approach, which better used spatial information, rather than spectral analyses, as DOQ, which has very high spatial resolution, but the digital number may not be real. A further goal was to evaluate methods that can be implemented on personal computers (with conventional amounts of memory). Although identifying individual species can be demonstrated by machine learning methods (e.g., [31]), the hyperspectral and/or very high resolution imagery (less than 0.5 m) that these methods relied on is not always available. Thus, the hierarchical framework, which was built on widely available data and computing capabilities, and incorporated ecological knowledge of the study area, provided the ability to label segments within specific species association or species types, even if imagery data were not able to support species identifications.
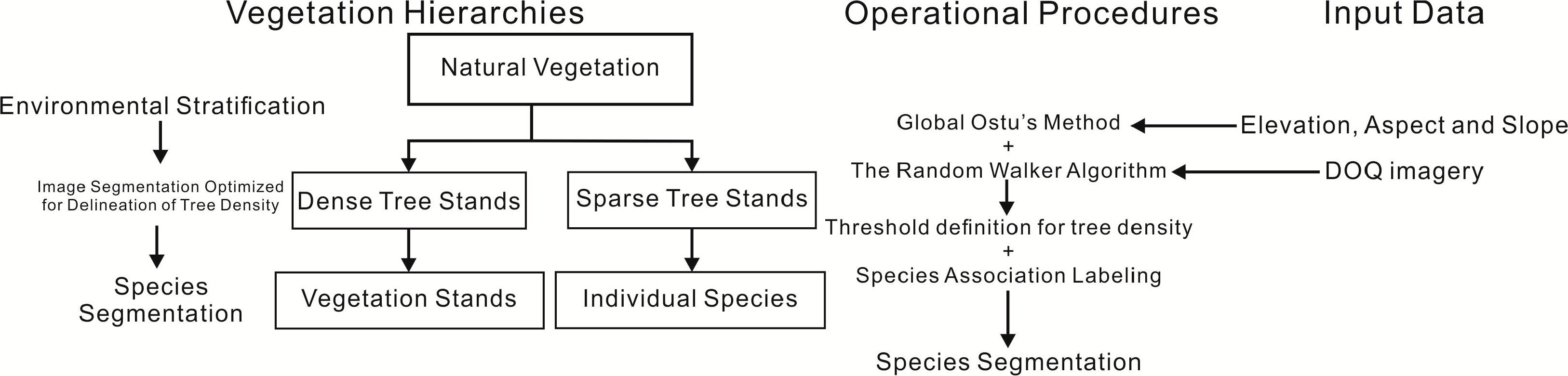
Partitioning the image based on tree density was a key part of the natural vegetation framework used in this study (Figure 1). Previous studies emphasized intensive data requirements [25,26,28–30] and repetitive testing on the applications of vegetation mapping, but those approaches may not form robust procedures toward long-term image processing with uncertain data sources (i.e., validation points) and the lack of multiple attributes. Moreover, segmentation studies (reviewed by [32–37]) have not been extensively applied to natural vegetation using a robust approach. Among those segmentation methods, which were classified by Fu and Mui [32], edge-based algorithms were the most vulnerable to noise (i.e., heterogeneous pixels with higher variances, especially in sparse tree stands) [32], and threshold-based algorithms cannot deal with imagery complexities, even using local threshold approaches [38,39]. In contrast, region-based algorithms were widely incorporated with other algorithms, including the Woodcock and Harward [20] region growing algorithm, eCognition’s imagery merging and fractal net evolution approach [21,23,24], and multiscale object-specific segmentation (MOSS) using size constrained region merging [21,40]. Those algorithms could better deal with noise and avoid over-segmentation on vegetation mapping, compared to other algorithms, such as the watershed algorithm and the region growing algorithm [34,37,41]. However, two-step labeling procedures [30] or repetitive testing on parameters (e.g., scale, compact and shape parameters for multiresolution segmentation in eCognition) to construct hierarchical frameworks [21–24] were still time-consuming and data-intensive. Therefore, a robust alternative using image segmentation optimized for delineation of tree density, which distinguishes a specific threshold for separating meaningful species association and selecting segmentation algorithm or parameter, was necessary for simplifying those procedures and data requirements in time-series image analysis.
Furthermore, tree density was the main trait of the hierarchical vegetation framework for two reasons (Figure 1). Tree density distributions, which reflected specific ecological site factors, were more appropriate for occurrence of specific species association (e.g., shade-intolerant species and shade-tolerant species) [42–45]. In theory, shade-intolerant species tend to occur at sparse tree stands, while shade-tolerant species tend to occur at dense tree stands. Additionally, different tree density patterns were required to identify two kinds of image objects, tree species and vegetation stands, using two processing procedures. For identifying individual tree species, tree crown delineation algorithms and labeling classifiers using multispectral bands were widely used, especially for interpreting species counts and types, which can substitute for field-based surveys [46–50] to some extents. Those algorithms included valley-following, region growing and watershed segmentation [51–55]. Nevertheless, the fundamental assumptions were that tree crowns should be clearly separated in space (not overlap) and should be in regular shapes and similar sizes, according to algorithm functions [52,53]. Thus, no general rule in the literature to select algorithms is available [52,53]. Furthermore, this approach was less effective in heterogeneous and denser hardwood stands than conifer landscapes [51,56]. For vegetation stands, decomposing landscapes into smaller objects (vegetation stands, a group of trees) and labeling species by the majority classified pixels within segmentations [15], vegetation gradient model adding spectral mixture analysis [30] or non-parametric classifiers (e.g., [18,23]) were usually applied (e.g., [18,57–60]). Nevertheless, segmentations in sparse tree stands may not be partitioned well, due to effects of non-vegetation areas or shadows. In addition, precise vegetation boundaries were still challenging in denser tree stands, because region-based segmentation partitioned imagery, based on its variances, rather than true objects. To sum up, preferable algorithms in the literature for the two kinds of objects had their own problems, and no general solution was available. As a result, one of goals in this study was to test a flexible guideline for algorithm or parameter selections.
Specifically, the hypotheses tested in this study were:
Environmental stratification, which environmental variables were separated by the global Otsu’s method for strata, will reduce the image variation of digital number (DN) and texture. It is assumed that reducing variance in image DN and texture indices simplified species composition based on studies showing that texture indices (local variances and second order textures) have high correlations with forest structural parameters (e.g., standard deviation of diameters and basal area) [61,62] and can clearly distinguish different forest parameters (e.g., stand ages) [63].
Identifying tree-density patterns will be used to simplify species association in a given area where there was no recent disturbance. This hypothesis was based on assumptions that shade tolerance was related to variations in tree architectural parameters, such as stem and crown dimensions [42–45], and species life history traits (e.g., different seedling time between oaks and pines) and environmental conditions (e.g., topographic and climatic conditions), which constrained species regenerations, resulting in different species dominances (pine dominance and oak-pine codominance) in stands with different tree densities [64,65].
Different tree density distributions will have different optimal scale parameters for image segmentation where the optimal scale parameter was that with the lowest segmentation variances and spatial autocorrelations [66–68]. The hypothesis was based on studies indicating that different image objects, such as sizes of tree crowns or clumps are a function of different spatial scales [10].
2. Methodology
Step one and step two, which involved the global Otsu’s method using environmental variables and the random walker algorithm on the imagery, was used to identify two tree density patterns. Step three was the multiresolution segmentation for extracting species objects (Figure 1). Environmental variables, based on elevation, slope or aspect, were used to partition the whole image into smaller subsets. Large images needed to be subset in order to reduce processing time and variation partitioning can help to statistically characterize certain components by increased stationarity or homogeneity within strata [69,70]. A non-parametric test was used to see which terrain variable was most effective at reducing variation in local DN and texture measures within strata. The reasons for using one environmental variable, rather than all three, were that one of goals in this study was to build a parsimonious procedure for reducing computations, and stratification based on a single variable was simple to implement. Then, the random walker algorithm was applied within the environmental strata to partition imagery into smaller subsets based on tree density patterns. The segmentation was evaluated by independent vegetation maps (the Timber survey and CALVEG) for segmentation accuracies. The goal was to evaluate whether the segmentation distinguishes image regions associated with particular species or species associates (Figure 1). Finally, the segmentation of tree densities was further partitioned by multiresolution segmentation with different scale parameters for species or stand level segmentation. The segmentation efficiency was evaluated by whether the detailed species segmentation related to specific guidelines of scale parameters on identifying species objects (Figure 1). So as to minimize costs, this study mainly used open source python libraries, scikit-image and other free software, and free DOQ imagery, and reduced processing time in commercial software to achieve the above goals.
2.1. Study Targets and Data Sources
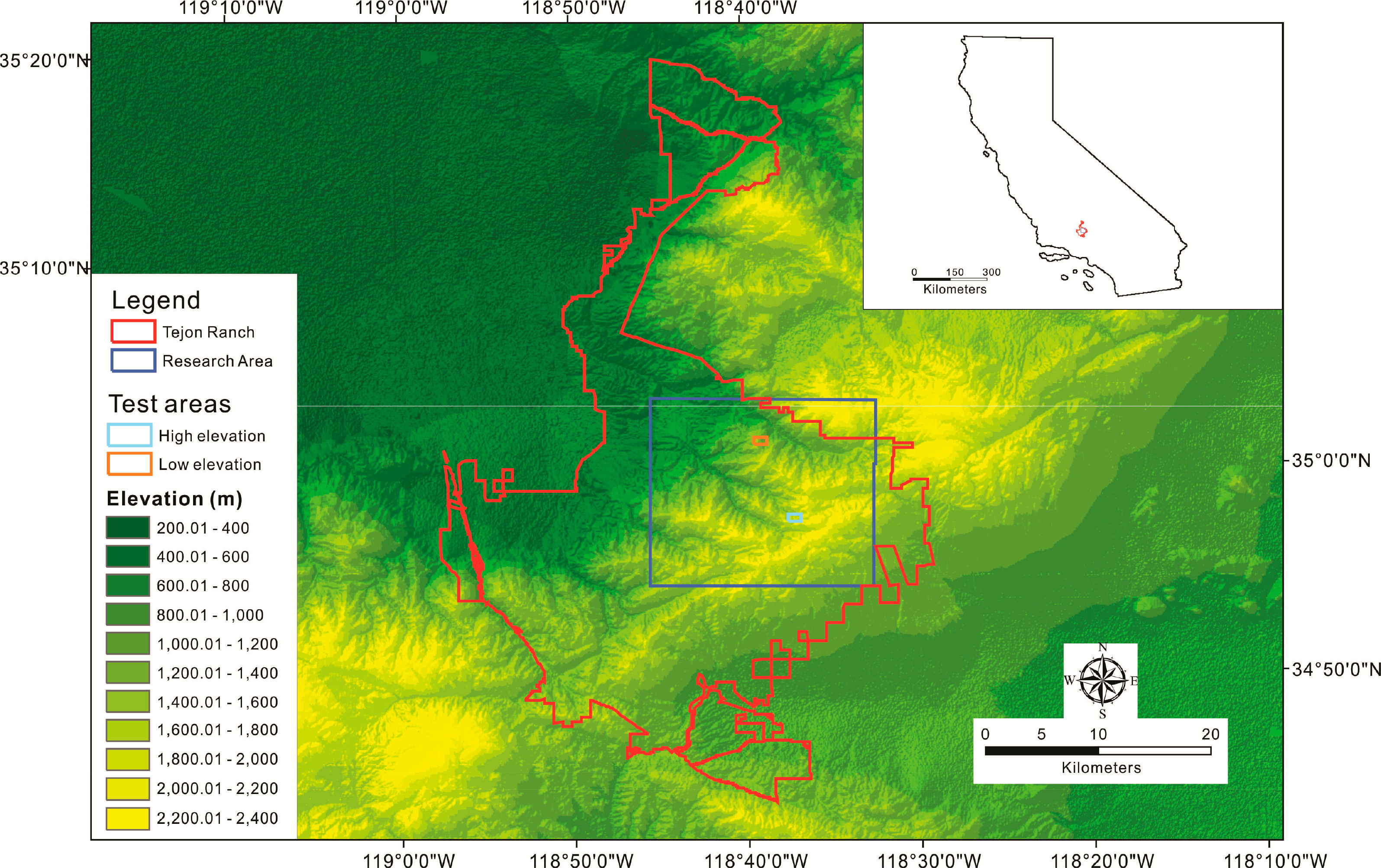
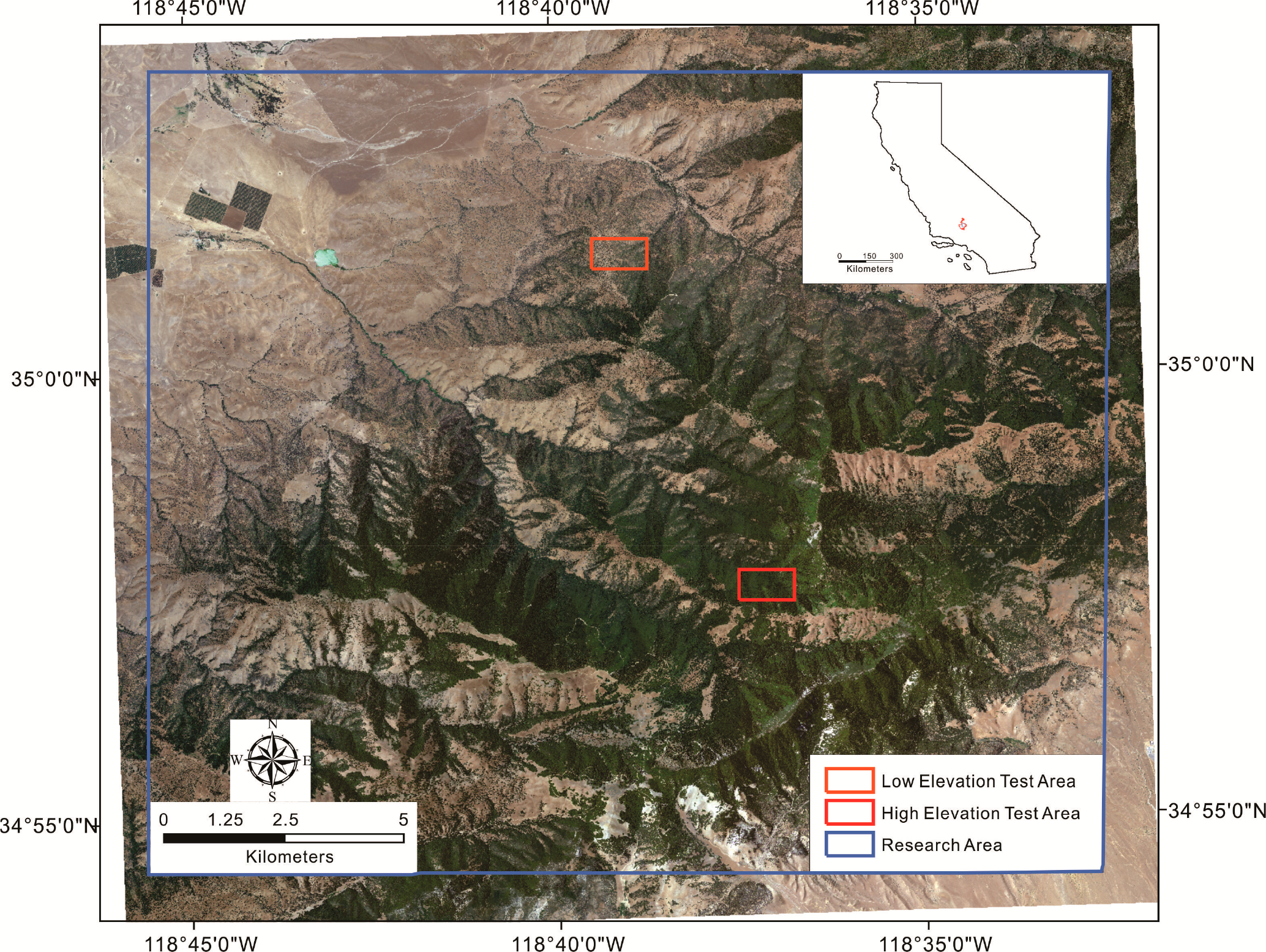
Tejon Ranch, which belongs to Tejon Ranch Company, is located in the convergence of four eco-regions: the Mojave Desert, the Central Valley, the Sierra Nevada, and the Transverse Ranges [71]. The research area is located in the Tehachapi Mountains, elevation ranging from 368 to 2360 m [72] (Figure 2). Based on the only climate station within Tejon Ranch, 434.3 m elevation, the average yearly temperature is 59.61 °C (1895–2011) and yearly rainfall is 11.29 inches (1899–2011) [73]. This area has a typical Mediterranean climate. The main rainfall season concentrates in the winter from October to March of the next year, while the dry season is in the summer from July to September. The main vegetation type is oak woodlands, including canyon live oak, interior live oak, blue oak, California black oak, scrub oak, and others along with ponderosa pine and gray pine.
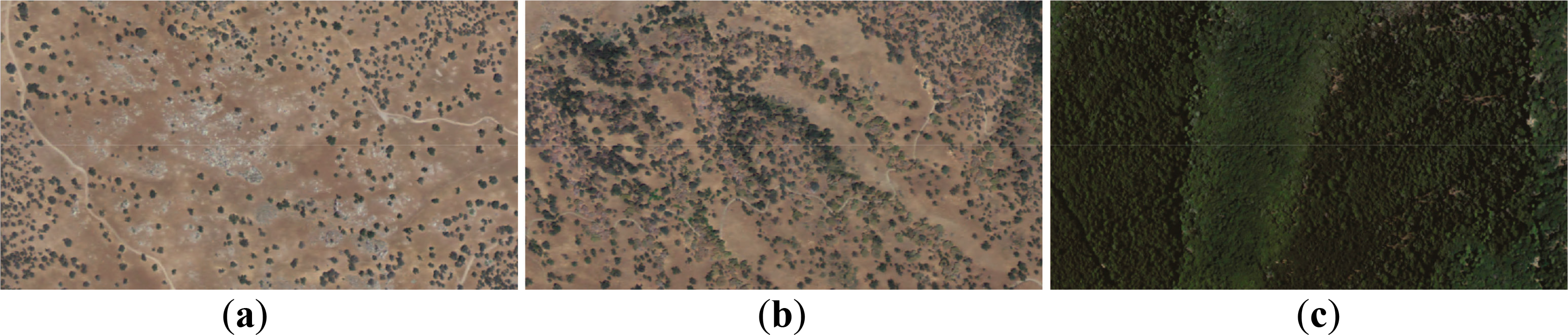
This study focused on two hardwood species, blue oak and California black oak, and two conifer species, ponderosa pine and grey pine. The four species are shade-intolerant [74], and tend to occur in sparse tree stands. Nevertheless, the shade tolerance of California black oak varies with age (higher shade-tolerant in saplings and less shade tolerance up maturity), so it can sustain in denser tree stands [74]. More importantly, owing to the long-term fire disturbance history, tree density does not always achieve its maximum possible value, and local adaptations to microenvironments and regeneration played more important roles in determining tree density. For example, ponderosa pines in southwestern United States were often found in dense stands changing from a range of 49–124 trees ha-1 at the time of Euro-American settlement to a range of 1235–2470 trees ha-1 now [75–77], due to high density of seedlings and saplings and fast growth near burned areas [65]. To sum up, species successions were not clearly identified in previous literatures and records and any rules about determinants of tree densities were not extensively applied to regions [74], so it is assumed that species have similar tree density patterns within a given area.
Data sources included a one-meter DOQ, SRTM 30 m Digital Elevation Data v.4 and two archived vegetation type maps, referred to in this study as Timber survey and the CALVEG. The DOQ was imagery with three 8-bit bands, which was rectified by digital terrain models and ground position points to remove terrain relief and camera tilt [78] so every land element was in corrected ground position. The horizon geometric accuracy, which was evaluate by a root-mean-square-error (RMSE), is 0.82 m [79]. The image was collected in 9 December 2009. This imagery is freely available from United States Department of Agriculture: Natural Resources Conservation Service Geospatial Data Gateway ( http://datagateway.nrcs.usda.gov/GDGOrder.aspx) or United States Geological Survey: The National Map Viewer and Download Platform ( http://viewer.nationalmap.gov/viewer/). However, because the DOQ was produced by aerial photos, the radiometric ranges cannot reflect the real spectral digital numbers. The SRTM 30m Digital Elevation Data, which were produced by National Aeronautics and Space Administration (NASA), were created by collecting elevation points from SRTM3 and a series of auxiliary digital terrain models for the purpose of interpolating voids to create seamless topography [72]. The data provided not only elevation information in 30 m spatial resolution but also slope and aspect gradients by calculating the rates of maximum change and their directions. The Timber survey was a field-surveyed map generated in 1980 for distributions of oak woodlands on Tejon Ranch [80]. The CALVEG (Classification and Assessment with Landsat of Visible Ecological Groupings) was produced by the USDA Forest Service using Landsat Thematic Mapper to construct a vegetation database across California in 1990s, and redeveloped in 2009 through 2010 using Landsat Thematic Mapper and Spot 5 panchromatic images [30]. Although the time of existing vegetation maps cannot correspond to the DOQ’s, this study assumed that the vegetation did not change much since 1980. The assumption was based on the fact of poor regeneration of California oak specie [81].
2.2. Environmental Stratification
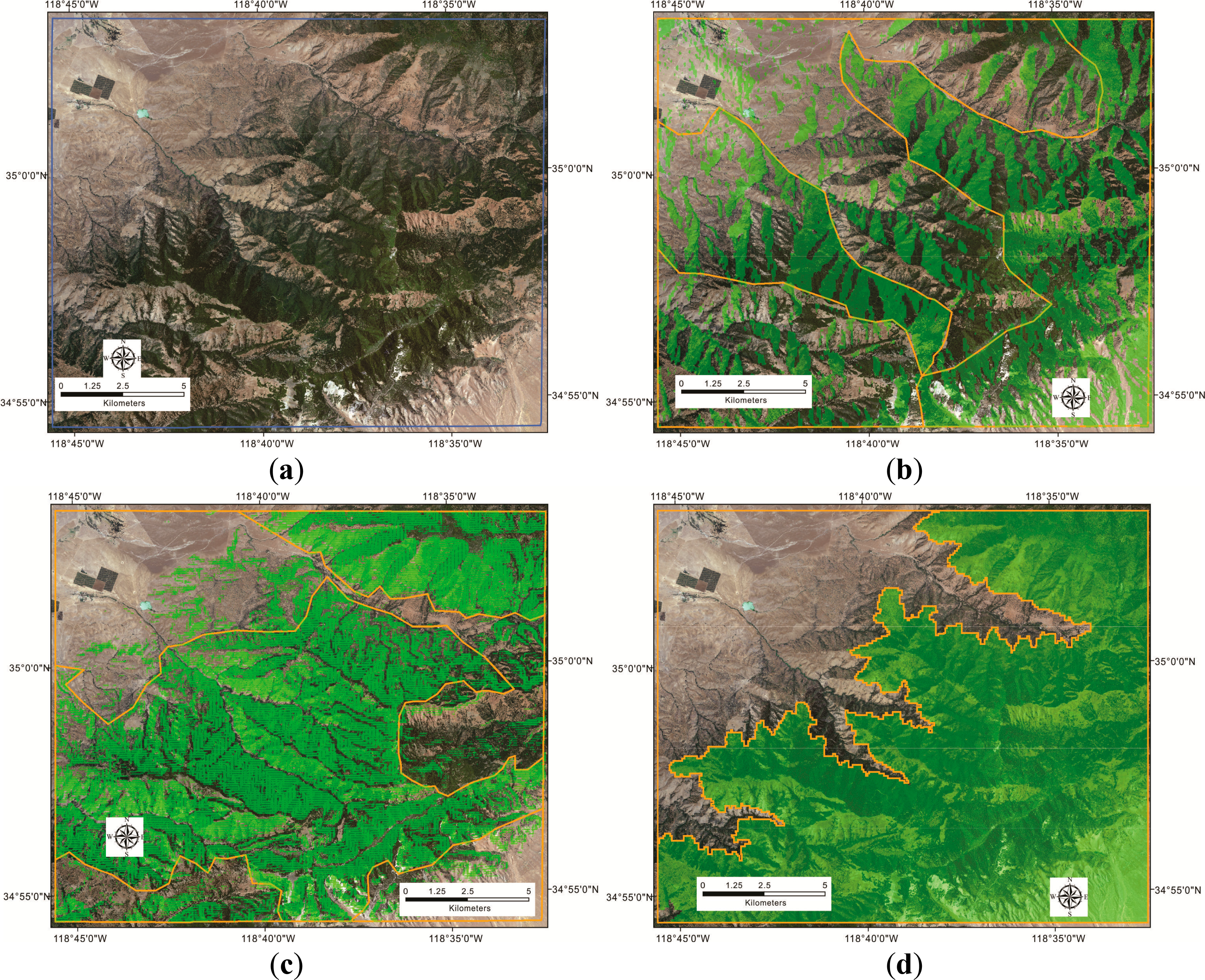
To create environmental stratifications, a global Otsu’s method was used to separate elevation, slope and aspect images into smaller areas using scikit-image. The scikit-image ( http://scikit-image.org/), written by a community of volunteers, is one of python open-source peer-reviewed code libraries. The library is a collection of image processing algorithms in previous studies. The goal applying global Otsu’s method was to find an optimal global threshold in image histogram (one-dimension intensity measurements) to partition imagery into two groups. The function was to maximize the intra-group variances and minimize within-group variances [82] in order to categorize objects from backgrounds, such as tree crowns from barren lands. Preliminary analyses determined optimal thresholds; the three stratifications were partitioned by thresholds of 1169 m for elevation, 41° for slope and 149° for aspect (Figure 3).
The global Otsu’s method has been shown to be more effective than other binarization methods [35,39] with lower computational time. However, those binarization methods were not spatially separated [32,35]. The strata boundaries needed to be delineated by hand, depending on the spatial distributions of environmental variables beyond or below the thresholds. The manual boundaries of strata did show diverse patterns to reflect tree covers. Then, the random walker algorithm was implemented within each stratification for further partitions.
2.3. Image Segmentation Optimized for Delineation of Tree Density
The random walker algorithm was used for image segmentation. Segmentation was based on single first band. The goal of using a single band, instead of three bands, was to save computation time, if the single band can reach the goal to identify tree density. Additionally, DOQ may not represent the true spectral digital numbers. The random walker algorithm, which was proposed by Grady [83], was used to partition imagery into smaller subsets more robustly using scikit-image. The random walker algorithm originated from the graph theory to view the whole imagery as the combinations of vertexes (nodes) and edges (arcs), and the random walkers, which represented each individual pixel, were trying to formulate a path to their neighbors, according to certain probability distributions. The algorithm started from defining the markers, a group of seeds as the sampling of desired imagery objects. Then, this algorithm would assign each unseeded pixel a probability and a weight. The probability was assigned by absolute distances from seeds to each individual image pixels (unseeded pixels), while the weight was converted by the image intensity. Finally, the algorithm would assign a seed class to those unseeded pixels for cuts, mainly according to the probability, and the cuts might be adjusted by the weights to avoid crossing sharp image intensities. For example, if three out of four neighbors belong to one class, then the focal pixel is assigned to this class. Thus, the random walker algorithm can keep locally consistent boundaries, regardless of spatial extents and attribute ranges.
However, how to define the markers was not described in details in the original paper, and few studies have used the random walker algorithm to identify vegetation types, because the random walker algorithm was under the assumptions (also called supervised segmentation algorithms) that a series of pixels for desired objects and backgrounds were known and nearby pixels between desired objects and backgrounds can evolve to desired boundaries. To handle this issue, the grey-level image DN, which was assumed to be a function of tree densities, was used in the marker designs. The image DN was higher, and the tree densities were sparser. Using natural break optimizing, the map classification was to reduce within-group variances, in three classes to derive the thresholds for desired objects and backgrounds. As a result, barren lands or individual trees over the highest threshold were viewed as the desired objects, and denser tree stands below the lowest threshold were treated as the background (Figure 4). Then, the random walker algorithm could separate desired objects from backgrounds. In the second segmentation, the same procedures were applied to separate lower tree density stands from higher tree density stands (Figure 4). Finally, barren lands or individual trees and lower tree density stands were combined as lower tree density stands for species association labeling (Figure 4).
2.4. Species Segmentation
Detailed vegetation objects were further partitioned following the image segmentation optimized for delineation of tree density using multiresolution in eCognition 8, which has been extensively used in studies of object-base image classification for vegetation inventories (e.g., [18,58,59]). The algorithm of multiresolution segmentation was demonstrated by optimization procedures to minimize heterogeneity for each individual merging [24]. Although many algorithms were developed to delineate tree crowns and open for free, those algorithms are scene-dependent and time-consuming to test them all in various operational system and computer languages. Thus, eCognition can be the simplest way to see the effects of the hierarchical framework on species segmentation. Segmentation was produced by 17 attributes or features: RGB; first-order textures: mean, standard deviation, kurtosis, mean Euclidean distance, skewness, variance; second-order textures: mean, variance, homogeneity, contrast, dissimilar, entropy, second moment, and correlation [84,85]. For the purpose of understanding which attributes can form better segmentation, several attribute combinations were tested, especially for the second-order textures, which literature pointed out high correlations with vegetation characteristics [61,62]. Nevertheless, multiresolution segmentation counted on classifiers to label after segmentation procedures. Therefore, as a prior framework, the multiresolution segmentation cannot be implemented with appropriate parameters or ruleset beforehand, and applied the same setting to other images directly. Optimal scale parameters were needed to be identified. The tested scale parameters (without units, depending on image heterogeneity), which determined the object merges and object sizes for the purpose of reducing heterogeneity [24], included 0.5, 1, 1.5, 2, 2.5, 3, 3.5, 4, 4.5, 5, and 5.5.
2.5. Segmentation Evaluations
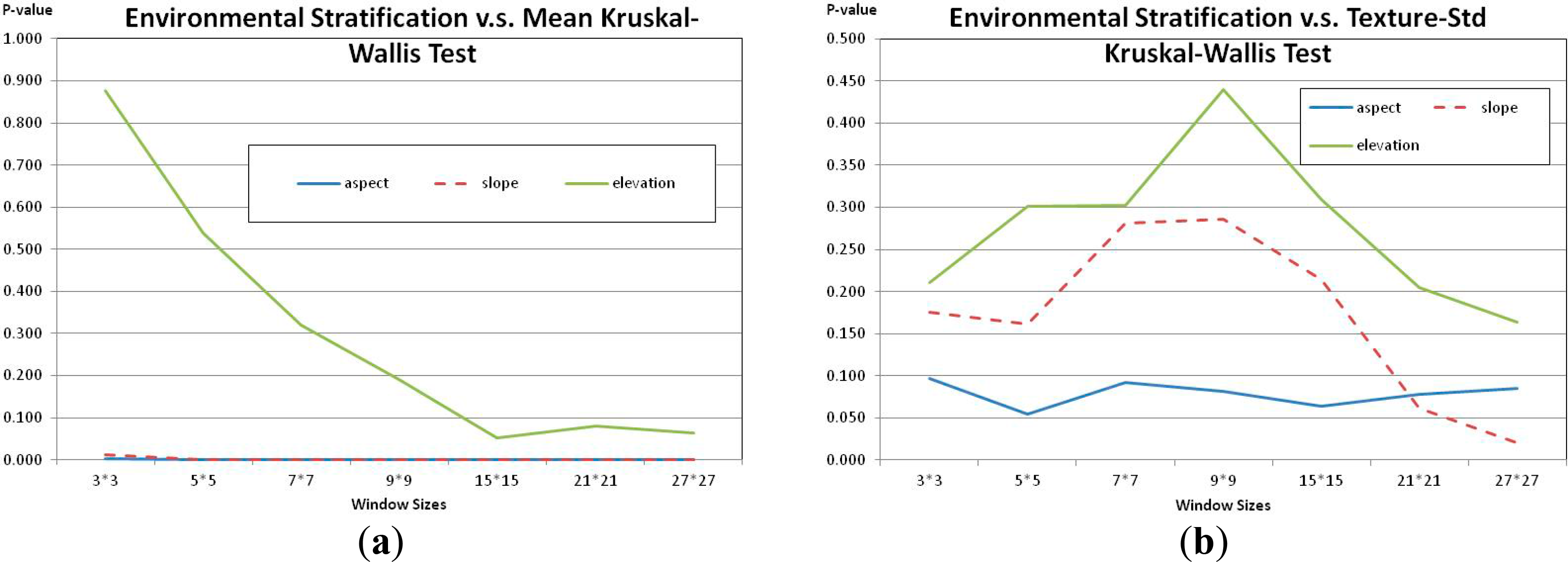
To test the hierarchical vegetation framework and the three hypotheses, a series of evaluations were used. First, image local DN and texture, mean and standard deviation in the first band, were used. The local mean and standard deviation were calculated based on window sizes, 3 by 3, 5 by 5, 7 by 7, 9 by 9, 15 by 15, 21 by 21 and 27 by 27. The goal was to see which terrain variable could be used to stratify the image most effectively, because image DN and texture correlates with forest structure parameters. A non-parametric test, Kruskal-Wallis test was applied using SPSS Statistics 20.0 to test whether variances (the local mean and standard deviation) were equal among groups (different environmental stratification).
Second, four focal species were chose to examine the effectiveness of the random walker algorithm by determining whether the two tree density patterns, which were combined from three tree density types (Figure 4), reflected different species association distributions as determined by the archival vegetation maps (reference data). In other words, the CALVEG and the timber survey were used as reference data for sampling four focal species types. The species samples were combined into species association and used to compare with tree density distributions within the study area. Indices of map agreement, including the kappa value, overall accuracies, user’s accuracies and producer’s accuracies, are based on the values in a confusion matrix [30,86]. Sampling the maps in order to calculate those indices were based on a stratified random approach accomplished using a “plug-in” sampling design tool in ArcGIS 10.0 to allocate 100 points proportionally by class areas. The strata were based on the three image segmentation optimized for delineation of tree density (Figure 4), where larger segmentation areas had more sampling points.
In order to evaluate the scale parameters used in the multiresolution segmentation, this study applied an objective (unsupervised) evaluation approach by using Moran’s I [67,68] as a measure of spatial autocorrelation to measure the similarity of segment-averaged attributes as a function of the distance between segments. There are several reasons to select the unsupervised evaluation approach. First, the approach could be applied in areas without abundant field data for supervised evaluations. The CALVEG was classified from SPOT (20 m) and Landsat TM (30 m), while the Timber survey was in coarser segmentation than the CALVEG. Also, the two reference maps were created at different scales [80], so it would not be possible to distinguish whether inaccuracies of vegetation segmentations were from boundary errors in maps made at other scales. In brief, it was questionable to sample points for validation of species distributions in 1-meter DOQ. Second, it is still challenging to evaluate results of object-based image analysis, compared to accuracy assessments in pixel-based image classifications, which applied sampling points to evaluate agreements [57,87,88]. Pixels within segmentations were not spectrally homogeneous [89], so the sampling points may not represent the whole objects. In addition, some of segmentation tools are proprietary software, such as Berkeley Image Segmentation (BIS) ( http://www.berkenviro.com/berkeleyimgseg/) and eCognition ( http://www.ecognition.com/). Although they have been described in the literature (e.g., [22–24,90]), there was no way to evaluate the segmentation accuracies by algorithms.
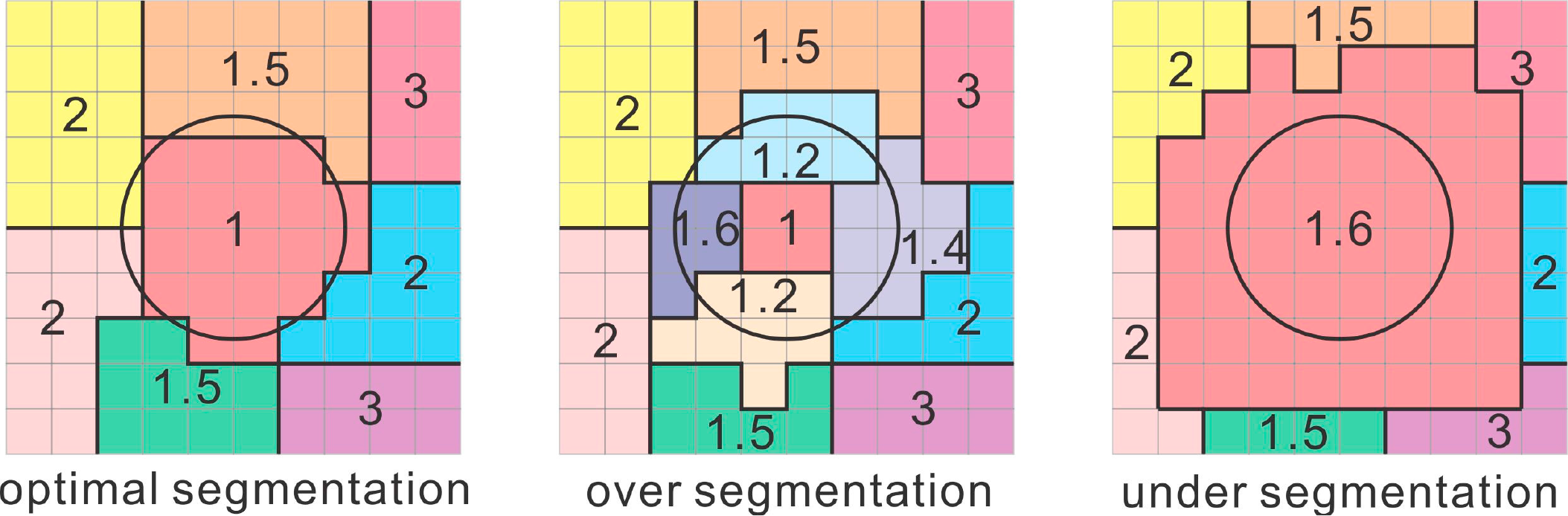
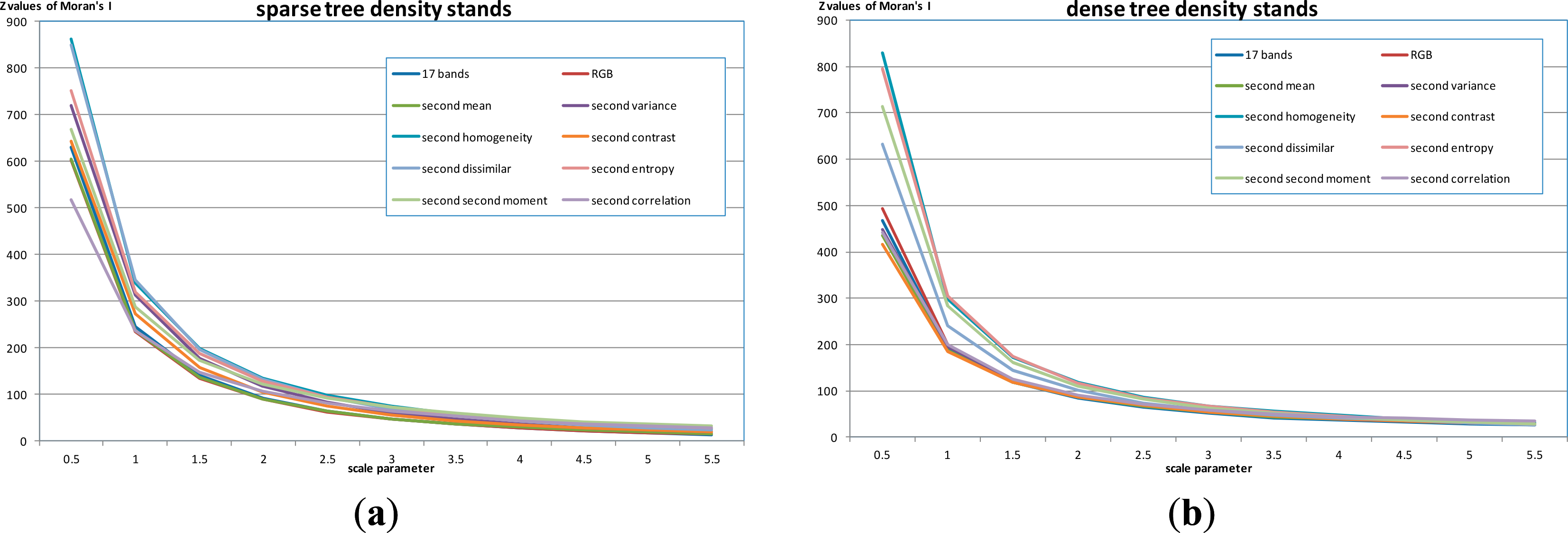
The basic idea of unsupervised evaluation was that attributes of optimal segmentations with clear boundaries (such as average spectral or texture values) should have low between-segment spatial autocorrelation, whereas over-segmentation (many small polygons), and under-segmentation (where the polygons are larger than the optimal segmentations), yielded segments whose attributes had higher between-segment spatial autocorrelation (Figure 5). Kim et al., [68] further pointed out that the segmentations in optimal scale parameters can contribute to better classification accuracies. Therefore, the segmentations based on different scale parameters were evaluated based on the Z values of Moran’s I in two test areas, sparse tree density stands and dense tree density stands (Figure 6). The goal was to see whether there were different optimal scale parameters for segmentation of two tree density patterns, because each has different sizes of individual trees and tree stands. This reason to use the Z values of Moran’s I, instead of Moran’s I in Kim et al. (2008) and Kim et al. (2009) [67,68] was that image segmentations with different scale parameters resulted in different weight matrices among images. For comparisons for an appropriate scale parameter, Moran’s I is required to standardized as the Z values of Moran’s I.
3. Results
3.1. Environmental Stratification
The aspect stratification had the best performance in terms of increasing stationarity within strata. Both DN and texture, mean and standard deviation of first-band digital number (DN) for different window sizes, showed significantly reduced variance within strata based on aspect, and the slope strata had the second best performance (Figure 7). Visually, the aspect stratification was most effective in separating tree crowns from barren lands, and the slope stratification again had the second performance (Figure 3). The slope stratification could be used to identify parts of barren lands, which had less steep slopes, but the elevation stratification was not useful for distinguishing woodland cover strata in this study area. Therefore, the aspect and slope variables produced strata that reduced variance in local measures of DN and texture, supporting the first hypothesis.
3.2. Image Segmentation on Tree Density
The two image segmentation optimized for delineation of tree density based on the random walker algorithm effectively separated the two species association. Ponderosa pines and California black oaks tended to occur in dense tree stands, while grey pines and blue oaks tended to occur in sparse tree stands within the study area. Agreement of species association with image segmentation optimized for delineation of tree density reached about 80% overall accuracies and 0.6 kappa values. In particular, using the aspect stratification for the thresholds of desired objects (firstly, for barren lands or individual tree stands and secondly for lower tree density stands) by natural break optimizing for the random walker algorithm did perform the best to produce image segmentation optimized for delineation of tree density (Table 1). Visually, step one and step two were effective to separate tree density patterns into three tree density categories, barren lands or individual trees, lower tree density stands and higher tree density stands, and the two selected regions of two tree densities had similar performances on separations of tree densities (Figure 8).
3.3. Species Segmentation
The evaluation of scale parameters suggested that multiresolution segmentation may not delineate appropriate detailed vegetation segmentation based on this imagery. Neither lower tree density stands nor higher tree density stands had an optimal scale parameter. The optimal scale parameter indicated that certain scale parameter would reach a minimum spatial autocorrelation, compared to its larger and smaller scale parameters. The Z values of Moran’s I continuously declined as the scale parameters increase, showing no local minimum (Figure 9). The difference between results for sparse tree density stands and dense tree density stands was the magnitude of Z values of Moran’s I in most attribute combinations. The sparse tree density stands had higher values, while the dense tree density stands had lower values (Figure 9). High variances of sparse tree stands, especially in large areas of barren lands, dominated the patterns of segmentations, while mixed species and overlapping tree crowns in dense tree stands resulted in similarity among segmentations. However, several segmentations, which were based on single second-order texture, especially homogeneity, dissimilarity and entropy, in both sparse tree density stands and dense tree density stands did not perform well (Figure 9).
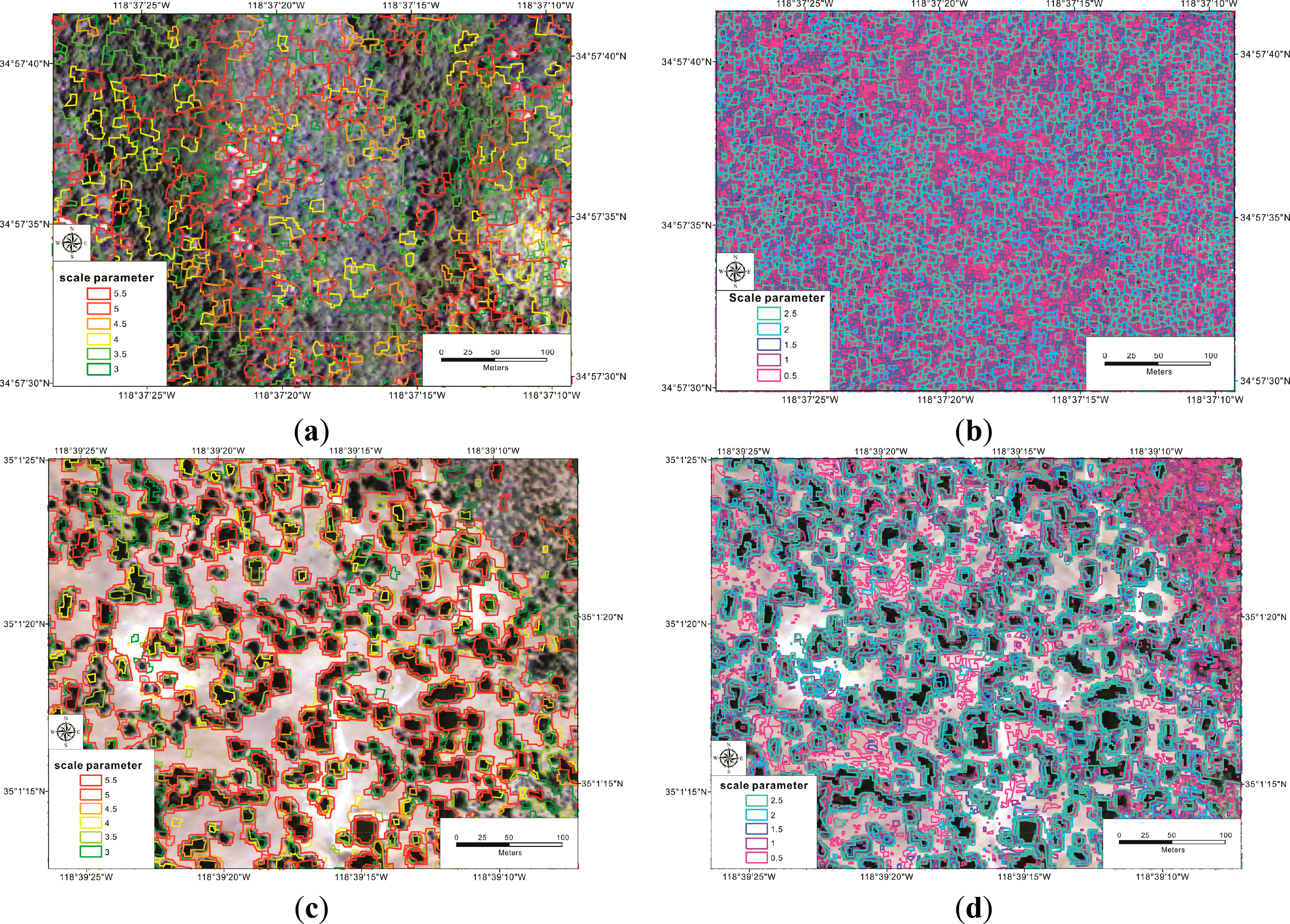
Visually, similar neighbor segmentation was the main problem in either denser tree density stands or sparse tree density stands, regardless of scale parameters. In the sparse tree density stands, although the main tree crowns could be identified, the boundaries of tree crowns were not precise. Concretely, near segmentations of barren lands confused with parts of tree crowns using smaller scale parameter (over segmentation), while tree species segmentations included parts of barren lands using larger scale parameter (under segmentation). Under the tested scale parameters, there was no optimal points to balance the over segmentation and under segmentation. In the dense tree stands, the similarity among segmentation was still high. Segmentations using smaller scale parameters resulted in very small, complex segmentation. Through spectral difference segmentation, original segmentations with larger scale parameters merged into very large segmentations (Figure 10).
4. Discussion
4.1. Implications of Image Segmentation Optimized for Delineation of Tree Density
Under the limitations of long-term available data and computation abilities of personal computers, image segmentation optimized for delineation of tree density demonstrated in this study provided an alternative framework for vegetation mapping, instead of data intensive analyses. The approach involved both the global Otsu’s method and the random walker algorithm using terrain variables and 1 m very high resolution imagery with only three bands as limited inputs. Those procedures can be carried out in Window 7 64-bit operating system using Intel(R) Core(TM) i7-2860QM CPU with 12 GB memory. The concrete goals were to add environmental variables, which were correlated with tree covers, and reduce variances within environmental strata and image segmentations. Specifically, those segmentation results reflected the applications of ecological understandings, instead of intensive data requirements and repetitive tests on segmentation and labeling. For one thing, water availability, which west-side slopes, confronting the ocean, had more precipitations than east-side slopes, might be the reason for the best performances of the aspect stratification, although no high-density climate stations or climate mapping at high spatial resolution can be used to validate. For another thing, because species associations are correlated with forest density in the study area, image segmentation optimized for delineation of tree density could be labeled by specific species associations, even though the third-step procedure of detailed species segmentations did not work well at identifying individual species segmentation, due to coarse spatial resolutions and less attributes.
In this study area, although four target species are shade-intolerant, they do not all occur in early-successional or low density stands as expected for shade-intolerant species, because of their varying shade tolerance at different ages (i.e., California black oak) and the human management history (i.e., ponderosa pine). Ponderosa pines occur in dense tree stands as do California black oaks, while blue oaks and grey pines occur in sparse tree stands. The agreements between image segmentation optimized for delineation of tree density and species association reached about 80% (overall accuracy). Further testing of alternative species labeling strategies in different ecosystems is needed for general applications.
Indices summarizing the confusion matrix were used to examine hypothesis 2, species association labeling. However, those common indices, especially the kappa value, have confronted harsh challenges in recent decades. The main criticism was that the kappa value was constructed by comparisons between the reference map and classification map, but the two maps may not be meaningfully comparable. Overall, the kappa value requires that both maps followed some assumptions, such as fitting normality and not being affected by other covariates, so adjustments on each map were the key, such as the weighted kappa coefficient [91]. In remote sensing cases, the kappa value was only a summary statistics through randomness sampling as a baseline, not a meaningful index to indicate quantity disagreement and location disagreement [92]. This study applied the kappa values and the overall accuracy as indices, because the stratified random approach using environmental variables, instead of wholly random sampling was effective to emphasize species locations (e.g., [93]), and tree density focused on whether species association labels were in specific image segmentation optimized for delineation of tree density, rather than improving detailed land use mapping or species mapping. Thus, simple indices were adequate for comparing the effects of environmental stratifications and assessing the correctness of species association locations.
4.2. Implications of Species Segmentation
The implications of species segmentation were to test the applications of the vegetation hierarchical framework, and whether the results related to specific guidelines of algorithm or parameter selections. The results employed an unsupervised evaluation (the Z values of Moran’s I under 999 permutations to evaluate whether the attributes of segmentations (average within segmentation) were significantly different from the neighbor segmentation. Although all showing decreases of the Z values of Moran’s I with increases of the scale parameters using different attribute combinations for segmentation (Figure 9), and image objects in two test areas both violated the object assumption with equal internal variances [20], the two tree density patterns did have fairly different reactions in most of attribute combinations on defining detailed species objects.
The unsuccessful segmentation at the third level resulted from two aspects in object definitions. In dense tree density stands, individual tree crowns could not be delineated, since the 1 m spatial resolution imagery was still too coarse. The spatial resolution is larger than the tree crowns. The result was similar to the outcomes of Woodcock and Strahler [10], where the coarse spatial resolutions did not have an optimal peak to reflect tree crown sizes (30 m imagery presented an asymptote, which local variances decreased as spatial resolutions increased, while 0.75 m imagery showed a local peak). The Z values of Moran’s I also showed an asymptote, which the Z values of Moran’s I decreased as scale parameters increased. In addition, homogeneity throughout the whole image resulted in very complex segmentations, especially using smaller scale parameters. As a result, adding higher spatial resolution image data or ancillary data, especially for oak woodlands, which often have lower classification accuracies (e.g., [56]), and using larger scale parameters for homogeneous areas may be a better alternative.
In the sparse tree density stands, the segmentations using multiresolution segmentation also could not derive correct tree crown objects, although individual tree crowns were clearly separated visually. The reason came from not enough information to delineate tree crown objects from the neighbor segmentation. Even if involving multiple texture features, segmentations were still limited to characterizing individual tree crowns. The segmentation may improve performances by implementing other segmentation algorithms, such as the watershed algorithm, as the example in Baatz et al., [22] or the valley-following algorithms and adding attributes, which can reflect vegetation characteristics (e.g., near-infrared band).
Although the results showed that multiresolution segmentation cannot deal with detailed species segmentations based on the DOQ imagery, the third evaluation still provided guidelines. As Fu and Mui [32] pointed out, region-based algorithms may differ in segmentation results, according to the order of region-merging, even if region-based algorithms were widely used and had higher classification accuracies than pixel-based imagery classifications. As a result, applying global information as nested models within certain classes is necessary. In other words, a single algorithm may not apply to whole hierarchical frameworks, even with the multi-scale concept.
4.3. Applications of Hierarchical Vegetation Framework
The hierarchical vegetation framework, developed in this study, is more time- and cost-saving for species association labeling. For one thing, data can be collected and automatically processed at lower costs. Very high resolution imagery in the form of digital photography is widely available, and environmental variables help to improve imagery processing for broad-area investigations. For another thing, the approach can incorporate current imagery processing methods, and does not require repetitive testing, like procedures in eCognition, and was effective to reduce time on using eCognition.
Concretely, aerial photos with single imagery DN are more available data sources for vegetation mapping, and sometimes, aerial photos with very high spatial resolution are in archives, either national-level imagery datasets (e.g., USDA: Natural Resources Conservation Service Geospatial Data Gateway or ASO Taiwan Image supplier and services System) or international-level imagery datasets (e.g., EarthExplorer). For example, Corona Lanyard (1963) and Corona KH-4B (from 1967 to 1972) both provided 1.8 m imagery worldwide, and Corona imagery can be bought through EarthExplorer website ( http://earthexplorer.usgs.gov/) with very small costs. Meanwhile, SRTM, a digital terrain dataset has worldwide coverage in 90 m spatial resolution, and can be used to assist image processing.
In contrast, Landsat imagery with coarser spatial resolutions has been widely used in long-term land use/land cover mapping and monitoring [94], because of its long-term consistent platforms, since 1972 and multiple bands, which reflect vegetation distributions (near-infrared) and temperature (thermal-infrared). Nevertheless, except in the United States, Landsat satellites may not offer enough imagery with continuous or regular time intervals and good imagery qualities, such as low cloud covers in wetter tropical areas. Furthermore, other detailed ancillary data, which were helpful to assist imagery interpretations, may be available at national-level datasets, such as U.S. National Elevation Dataset 10 Meter ( http://datagateway.nrcs.usda.gov/GDGOrder.aspx), U.S. General Soil Map (STATSGO) ( http://websoilsurvey.nrcs.usda.gov/app/WebSoilSurvey.aspx), or LiDAR Topography Data ( http://opentopo.sdsc.edu/gridsphere/gridsphere?cid=datasets). However, those datasets cannot reflect temporal changes of species distributions and are limited in survey areas.
This research provided a robust way for time-series image processing. The approach can be achieved by a single-band, very high resolution image at lower time, labor and facility costs. However, this approach still suffers several limitations. First, the global Otsu’s method applying to the aspect factor did not consider areas where the terrains mostly face north, since the situation (when 360 deg. = 0 deg) was not included. As the aspect factor performed the best as the first step to reduce variances with ecological meaning (tree covers), the global Otsu’s method is also the best way of fast computation. Second, species association labeling may only be effective in certain areas, as the kappa values only reached 0.6. Both disturbance frequencies and intensities may violate the second hypothesis. Especially, the approach cannot apply to areas suffering large, infrequent disturbances, which may result in more heterogeneous species distributions [95]. The goal of this study was not to provide an ultimate solution for general ecosystem labeling, but to propose an examined framework for an efficient approach on time-series images. Third, compared to commercial software, open-source solution cannot always provide equal functions, regardless of personal expertise. As open-source tools may be developed for usage in other fields (e.g. medical image processing, instead of vegetation mapping), and code quality is not certain among different libraries, it takes much time and effort for researchers to write their own codes or test the workable functions for research questions. For these limitations, there are several reasons to use the open-source tools as the solution. For one thing, this research selected a peer-reviewed image processing library, scikit-image, which reduced code uncertainty. For another thing, this research was not trying to fill the gap between commercial software and open-source tool, but provided the best way of using current available data and tools, especially for long-term image processing. To be brief, regardless of limited image information, this research intended to offer a starting point of using open-source solutions on time-series image processing.
Future studies should focus on two aspects. One is finding a new method for species association labeling in different ecosystems. Ecological understanding and local knowledge should be applied to vegetation mapping. The other focus of future research should be the techniques themselves. Algorithms of tree crown delineations can be tested at sparse tree density stands and appropriate object definitions should be established for dense tree density stands. In particular, individual tree density patterns reflect different biological interactions over times. For example, at the species spreading front, reduced intra-specific competition in dense populations may increase population growth rates and migrations, but sparse populations may suffer decreased migrations [96]. However, highly overlapping tree crowns make it difficult to detect changes in density of individual species (e.g., [97]), and tree crowns in sparse tree stands cannot be identified using multiresolution segmentation. Therefore, based on the hierarchical framework, two separate algorithms with more precise object definition are required.
5. Conclusions
Object-based analysis with a multiple scale framework is one of solutions toward time-series image processing. However, the data-based orientation approach required more data sources and the vegetation-type orientation approach was limited to improve classification accuracies of specific species types for ecological modeling. In particular, long-term image sources, which have less image information and are lacking in detailed validation data are still challenging to deal with. Also, large amounts of tests on labeling procedures and obtaining the best multiresolution framework are still required. As a result, a robust segmentation approach mainly using open-source tools with less data requirements (i.e., SRTM, digital orthophoto quadrangles with 1 m spatial resolution and existing vegetation maps as validation data) was proposed.
The author applied a three-step procedure for the above issues. At the first level-segmentation, the global Otsu’s method was applied to characterize vegetation traits and reduce computation loads by gathering similar components together into strata on three environmental variables (i.e., slope, aspect and elevation). With the evaluation by the Kruskal-Wallis test, this step, especially the aspect factor was proved effective to reduce image variances in local DN and texture and more importantly, characterize similar components of tree covers within strata. At the second step, image segmentation optimized for delineation of tree density was used with the least computation requirements. Species association labeling for four target species, which were grouped into two species associations was tested for effectiveness, and indicated about 80% in overall accuracy and 0.6 in kappa values between image segmentation optimized for delineation of tree density and species associations, which were derived from two existing vegetation maps. In the third step, the image segmentation, optimized for delineation of tree density, was further tested for its applications on partitioning species segmentation using the Z values of Moran’s I. However, this step using multi-resolution segmentation did not bring an optimal scale parameter, due to limited information on a single-band imagery. More importantly, the results, which showed two reflections on the Z values of Moran’s I, indicated the suggestions for segmentation algorithm selections and developments.
Although the image information is limited to identifying individual species, and the species association labeling for the evaluation of image segmentation optimized for delineation of tree density is area-based, the study still provides the following contributions to vegetation mapping. First, the hierarchical vegetation framework for image segmentation provided the possibility of long-term change detections, at least to compare certain ecological indices over time. For example, worldview-3 or Geo-eye provide more opportunities to delineate species segmentation, while Landsat MSS, TM, ETM+ or Corona are unable to segment species, as compared to recent imagery. As more and more spatial-temporal modeling approaches are developed, long-term imagery change detections in object (entity) forms provide the potentials to extend current models to the recent past (over 70 years)[98]. Second, the hierarchical vegetation framework with ecological meanings is effective to reduce computation loads and indicate the processing procedures. In the former two steps, the whole image dataset (22393 × 19458) was processed, while the third step was limited by the commercial software, which required extra facility, such as the eCognition server ( http://www.ecognition.com/products/ecognition-server) for big data processing. As more historical images are available on the Internet, this approach can provide solutions to process large amounts of image sources with diverse quality. Third, based on the current object definition, the limited image information was proven unable to produce detailed, reasonable image segmentation with an optimal scale parameter. As the Z values of Moran’s I are more proper to be used as the evaluation index for appropriate segmentations, the index was also used to evaluate the optimal segmentation level. In this study, the second level segmentation is optimal, since the near segmentations at the third level are similar. Although multiresolution segmentation can produce segmentations, those segmentations may not be meaningful, as spectral difference segmentation further aggregated all segmentations together. At this point, the open-source solution using threshold-based segmentation is helpful to deal with those kinds of images.
Acknowledgments
The author would like to thank Janet Franklin for advising and encouragement to provide me the writing guideline, comments and edit for completion of this paper. Also thank the data support by Collaborative NSF award EF-1065826 to J. Franklin, ASU (PI: F.W. Davis, UCSB). The author appreciates Sergio Rey for teaching and discussions on python programming and spatial analyses that formed the main technical basis of this paper. The author is also grateful Billie Turner and Soe Myint to read through the paper and offer some suggestions, and in particular, the author appreciates Soe Myint for allowing me access to ENVI and eCognition. Additionally, the author thanks to Kuo-Chen Chang for the access of eCognition in the revision. The author is grateful to four anonymous reviewers and the guest editor for detailed and constructive criticisms and further edit.
Conflicts of Interest
The author declares no conflict of interest.
References
- Egbert, S.L.; Lawrence, K.S.; Martinez-Meyer, E.; Ortega-Huerta, M.; Townsend Peterson, A. Use of datasets derived from time-series AVHRR imagery as surrogates for land cover maps in predicting species’ distributions. Proceedings of 2002 IEEE International Geoscience and Remote Sensing Symposium, IGARSS’02, Toronto, ON, Canada, 24–28 June 2002; pp. 2337–2339.
- Hill, R.A.; Wilson, A.K.; George, M.; Hinsley, S.A. Mapping tree species in temperate deciduous woodland using time-series multi-spectral data. Appl. Veg. Sci 2010, 13, 86–99. [Google Scholar]
- Loh, J.; Green, R.E.; Ricketts, T.; Lamoreux, J.; Jenkins, M.; Kapos, V.; Randers, J. The living planet index: Using species population time series to track trends in biodiversity. Philos. Trans. R. Soc. B Biol. Sci 2005, 360, 289–295. [Google Scholar]
- Donohue, R.J.; Mcvicar, T.R.; Roderick, M.L. Climate-Related trends in australian vegetation cover as inferred from satellite observations, 1981–2006. Glob. Change Biol 2009, 15, 1025–1039. [Google Scholar]
- Pettorelli, N.; Vik, J.O.; Mysterud, A.; Gaillard, J.-M.; Tucker, C.J.; Stenseth, N.C. Using the satellite-derived NDVI to assess ecological responses to environmental change. Trends Ecol. Evol 2005, 20, 503–510. [Google Scholar]
- Franklin, J. Mapping Species Distributions: Spatial Inference and Prediction; Cambridge University Press: New York, NY, USA, 2010. [Google Scholar]
- Franklin, J. Predictive vegetation mapping: Geographic modelling of biospatial patterns in relation to environmental gradients. Progr. Phys. Geogr 1995, 19, 474–499. [Google Scholar]
- Pearson, R.G.; Dawson, T.P. Predicting the impacts of climate change on the distribution of species: Are bioclimate envelope models useful? Glob. Ecol. Biogeogr 2003, 12, 361–371. [Google Scholar]
- Nagendra, H.; Rocchini, D. High resolution satellite imagery for tropical biodiversity studies: The devil is in the detail. Biodiver. Conserv 2008, 17, 3431–3442. [Google Scholar]
- Woodcock, C.E.; Strahler, A.H. The factor of scale in remote sensing. Remote Sens. Environ 1987, 21, 311–332. [Google Scholar]
- Strahler, A.H.; Woodcock, C.E.; Smith, J.A. On the nature of models in remote sensing. Remote Sens. Environ 1986, 20, 121–139. [Google Scholar]
- Blaschke, T. Object based image analysis for remote sensing. ISPRS J. Photogramm. Remote Sens 2010, 65, 2–16. [Google Scholar]
- Budreski, K.A.; Wynne, R.H.; Browder, J.O.; Campbell, J.B. Non-Parametric land cover classification in the brazilian amazon using multitemporal landsat TM/ETM imagery. Photogramm. Eng. Remote Sens 2007, 73, 813–827. [Google Scholar]
- Huang, H.; Wu, B.; Fan, J. Analysis to the relationship of classification accuracy, segmentation scale, image resolution. Proceedings of the 2003 IEEE International Geoscience and Remote Sensing Symposium, Toulouse, France, 21–25 July 2003; pp. 3671–3673.
- Shandley, J.; Franklin, J.; White, T. Testing the woodcock-harward image segmentation algorithm in an area of southern California chaparral and woodland vegetation. Int. J. Remote Sens 1996, 17, 983–1004. [Google Scholar]
- Varela, R.A.D.; Rego, P.R.; Iglesias, S.C.; Sobrino, C.M. Automatic habitat classification methods based on satellite images: A practical assessment in the NW iberia coastal mountains. Environ. Monit. Assess 2008, 144, 229–250. [Google Scholar]
- Yan, G.; Mas, J.-F.; Maathuis, B.H.P.; Xiangmin, Z.; Dijk, P.M.V. Comparison of pixel-based and object-oriented image classification approaches—A case study in a coal fire area, wuda, inner Mongolia, China. Int. J. Remote Sens 2006, 27, 4039–4055. [Google Scholar]
- Yu, Q.; Gong, P.; Clinton, N.; Biging, G.; Kelly, M.; Schirokauer, D. Object-Based detailed vegetation classification with airborne high spatial resolution remote sensing imagery. Photogramm. Eng. Remote Sens 2006, 72, 799–811. [Google Scholar]
- Strahler, A.H. Stratification of natural vegetation for forest and rangeland inventory using landsat digital imagery and collateral data. Int. J. Remote Sens 1981, 2, 15–41. [Google Scholar]
- Woodcock, C.E.; Harward, V.J. Nested-hierarchical scene models and image segmentation. Int. J. Remote Sens 1992, 13, 3167–3187. [Google Scholar]
- Hay, G.J.; Blaschke, T.; Marceau, D.J.; Bouchard, A. A comparison of three image-object methods for the multiscale analysis of landscape structure. ISPRS J. Photogramm. Remote Sens 2003, 57, 327–345. [Google Scholar]
- Baatz, M.; Hoffmann, C.; Willhauck, G. Progressing from object-based to object-oriented image analysis. In Object-Based Image Analysis: Spatial Concepts for Knowledge-Driven Remote Sensing Applications; Blaschke, T., Lang, S., Hay, G.J., Eds.; Springer-Verlag: Berlin Heidelberg, Germany, 2008; pp. 29–42. [Google Scholar]
- Benz, U.C.; Hofmann, P.; Willhauck, G.; Lingenfelder, I.; Heynen, M. Multi-resolution, object-oriented fuzzy analysis of remote sensing data for gis-ready information. ISPRS J. Photogramm. Remote Sens 2004, 58, 239–258. [Google Scholar]
- Baatz, M.; Schäpe, A. Multiresolution segmentation—An optimization approach for high quality multi-scale image segmentation. In Angewandte Geographische Informationsverarbeitung XII; Strobl, J., Blaschke, T., Griesebner, G., Eds.; Wichmann-Verlag: Heidelberg, Germany, 2000; pp. 12–23. [Google Scholar]
- Kim, M.; Warner, T.A.; Madden, M.; Atkinson, D.S. Multi-scale geobia with very high spatial resolution digital aerial imagery: Scale, texture and image objects. Int. J. Remote Sens 2011, 32, 2825–2850. [Google Scholar]
- Chen, G.; Hay, G.J. An airborne lidar sampling strategy to model forest canopy height from quickbird imagery and geobia. Remote Sens. Environ 2011, 115, 1532–1542. [Google Scholar]
- Chen, G.; Hay, G.J.; St-Onge, B. A geobia framework to estimate forest parameters from lidar transects, quickbird imagery and machine learning: A case study in Quebec, Canada. Int. J. Appl. Earth Observ. Geoinf 2012, 15, 28–37. [Google Scholar]
- Chen, Q.; Laurin, G.V.; Battles, J.J.; Saah, D. Integration of airborne lidar and vegetation types derived from aerial photography for mapping aboveground live biomass. Remote Sens. Environ 2012, 121, 108–117. [Google Scholar]
- Xie, Z.; Roberts, C.; Johnson, B. Object-based target search using remotely sensed data: A case study in detecting invasive exotic australian pine in south Florida. ISPRS J. Photogramm. Remote Sens 2008, 63, 647–660. [Google Scholar]
- Franklin, J.; Woodcock, C.E.; Warbington, R. Multi-attribute vegetation maps of forest service lands in California supporting resource management decisions. Photogramm. Eng. Remote Sens 2000, 66, 1209–1217. [Google Scholar]
- Foody, G.M.; Atkinson, P.M.; Gething, P.W.; Ravenhill, N.A.; Kelly, C.K. Identification of specific tree species in ancient semi-natural woodland from digital aerial sensor imagery. Ecol. Appl 2005, 15, 1233–1244. [Google Scholar]
- Fu, K.S.; Mui, J.K. A survey on image segmentation. Pattern Recognit 1981, 13, 3–16. [Google Scholar]
- Jain, A.K.; Murty, M.N.; Flynn, P.J. Data clustering: A review. ACM Comput. Surv. (CSUR) 1999, 31, 264–323. [Google Scholar]
- Ma, Z.; Tavares, J.M.R.S.; Jorge, R.N.; Mascarenhas, T. A review of algorithms for medical image segmentation and their applications to the female pelvic cavity. Comput. Methods Biomech. Biomed. Eng 2010, 13, 235–246. [Google Scholar]
- Pal, N.R.; Pal, S.K. A review on image segmentation techniques. Pattern Recognit 1993, 26, 1277–1294. [Google Scholar]
- Pham, D.L.; Xu, C.; Prince, J.L. Current methods in medical image segmentation. Ann. Rev. Biomed. Eng 2000, 2, 315–337. [Google Scholar]
- Sonka, M.; Hlavac, V.; Boyle, R. Image Processing, Analysis, and Machine Vision; Thomson: Toronto, ON, Canada, 2008. [Google Scholar]
- Sezgin, M.; Sankur, B. Survey over image thresholding techniques and quantitative performance evaluation. J. Electr. Imaging 2004, 13, 146–165. [Google Scholar]
- Trier, Ø.D.; Taxt, T. Evaluation of binarization methods for document images. IEEE Trans. Pattern Anal. Mach. Intell 1995, 17, 312–315. [Google Scholar]
- Hay, G.J.; Castilla, G.; Wulder, M.A.; Ruiz, J.R. An automated object-based approach for the multiscale image segmentation of forest scenes. Int. J. Appl. Earth Observ. Geoinf 2005, 7, 339–359. [Google Scholar]
- Wang, D. A multiscale gradient algorithm for image segmentation using watershelds. Pattern Recognit 1997, 30, 2043–2052. [Google Scholar]
- King, D.A. Allometry and life history of tropical trees. J. Trop. Ecol 1996, 12, 25–44. [Google Scholar]
- Lines, E.R.; Zavala, M.A.; Purves, D.W.; Coomes, D.A. Predictable changes in aboveground allometry of trees along gradients of temperature, aridity and competition. Glob. Ecol. Biogeogr 2012, 21, 1017–1028. [Google Scholar]
- Poorter, L.; Lianes, E.; de las Heras, M.; Zavala, M.A. Architecture of iberian canopy tree species in relation to wood density, shade tolerance and climate. Plant Ecol 2012, 213, 707–722. [Google Scholar]
- Van Gelder, H.A.; Poorter, L.; Sterck, F.J. Wood mechanics, allometry, and life-history variation in a tropical rain forest tree community. New Phytol 2006, 171, 367–378. [Google Scholar]
- Gougeon, F.A.; Leckie, D.G. The individual tree crown approach applied to ikonos images of a coniferous plantation area. Photogramm. Eng. Remote Sens 2006, 72, 1287–1297. [Google Scholar]
- Haara, A.; Haarala, M. Tree species classifcation using semi-automatic delineation of trees on aerial images. Scand. J. For. Res 2002, 17, 556–565. [Google Scholar]
- Katoh, M.; Gougeon, F.A.; Leckie, D.G. Application of high-resolution airborne data using individual tree crowns in Japanese conifer plantations. J. For. Res 2009, 14, 10–19. [Google Scholar]
- Leckie, D.G.; Gougeon, F.A.; Tinis, S.; Nelson, T.; Burnett, C.N.; Paradine, D. Automated tree recognition in old growth conifer stands with high resolution digital imagery. Remote Sens. Environ 2005, 94, 311–326. [Google Scholar]
- Leckie, D.G.; Gougeon, F.A.; Walsworth, N.; Paradine, D. Stand delineation and composition estimation using semi-automated individual tree crown analysis. Remote Sens. Environ 2003, 85, 355–369. [Google Scholar]
- Culvenor, D.S. Extracting individual tree information a survey of techniques for high spatial resolution imagery. In Remote Sensing of Forest Environments: Concepts and Case Studies; Wulder, M., Franklin, S., Eds.; Springer: Berlin, Germany, 2003; pp. 255–277. [Google Scholar]
- Ke, Y.; Quackenbush, L.J. A comparison of three methods for automatic tree crown detection and delineation from high spatial resolution imagery. Int. J. Remote Sens 2011, 32, 3625–3647. [Google Scholar]
- Ke, Y.; Quackenbush, L.J. A review of methods for automatic individual tree-crown detection and delineation from passive remote sensing. Int. J. Remote Sens 2011, 32, 4725–4747. [Google Scholar]
- Larsen, M.; Eriksson, M.; Oescombes, X.; Perrin, G.; Brandtberg, T.; Gougeon, F.A. Comparison of six individual tree crown detection algorithms evaluated under varying forest conditions. Int. J. Remote Sens 2011, 32, 5827–5852. [Google Scholar]
- Li, Z.; Hayward, R.F.; Zhang, J.; Liu, Y. Individual tree crown delineation techniques for vegetation management in power line corridor. Proceedings of the 10th International Conference on Digital Image Computing: Techniques and Applications, Canberra, Australia, 1–3 December 2008.
- Katoh, M. Classifying tree species in a northern mixed forest using high-resolution ikonos data. J. For. Res 2004, 9, 7–14. [Google Scholar]
- Heumann, B.W. An object-based classification of mangroves using a hybrid decision tree—support vector machine approach. Remote Sens 2011, 3, 2440–2460. [Google Scholar]
- Laliberte, A.S.; Fredrickson, E.L.; Rango, A. Combining decision trees with hierarchical object-oriented image analysis for mapping arid rangelands. Photogramm. Eng. Remote Sens 2007, 73, 197–207. [Google Scholar]
- Mallinis, G.; Koutsias, N.; Tsakiri-Strati, M.; Karteris, M. Object-based classification using quickbird imagery for delineating forest vegetation polygons in a mediterranean test site. ISPRS J. Photogramm. Remote Sens 2008, 63, 237–250. [Google Scholar]
- Myint, S.W.; Giri, C.P.; Wang, L.; Zhu, Z.; Gillette, S.C. Identifying mangrove species and their surrounding land use and land cover classes using an object-oriented approach with a lacunarity spatial measure. GISci. Remote Sens 2008, 45, 188–208. [Google Scholar]
- Klobučar, D.; Subašić, M.; Pernar, R. Estimation of stands parameters from IKONOS satellite images using textural features. Proceedings of the 7th International Symposium on Image and Signal Processing and Analysis (ISPA 2011), Dubrovnik, Croatia, 4–6 September 2011.
- Ozdemir, I.; Karnieli, A. Predicting forest structural parameters using the image texture derived from worldview-2 multispectral imagery in a dryland forest, Israel. Int. J. Appl. Earth Observ. Geoinf 2011, 13, 701–710. [Google Scholar]
- Franklin, S.E.; Wulder, M.A.; Gerylo, G.R. Texture analysis of IKONOS panchromatic data for Douglas-fir forest age class separability in British Columbia. Int. J. Remote Sens 2001, 22, 2627–2632. [Google Scholar]
- Gracia, M.; Retana, J.; Roig, P. Mid-term successional patterns after fire of mixed pine-oak forests in NE Spain. Acta Oecol 2002, 23, 405–411. [Google Scholar]
- Zavala, M.A.; Zea, E. Mechanisms maintaining biodiversity in mediterranean pine-oak forests: Insights from a spatial simulation model. Plant Ecol 2004, 171, 197–207. [Google Scholar]
- Johnson, B.; Xie, Z. Unsupervised image segmentation evaluation and refinement using a multi-scale approach. ISPRS J. Photogramm. Remote Sens 2011, 66, 473–483. [Google Scholar]
- Kim, M.; Madden, M.; Warner, T. Estimation of optimal image object size for the segmentation of forest stands with multispectral IKONOS imagery. In Object-Based Image Analysis: Spatial Concepts for Knowledge-Driven Remote Sensing Applications; Blaschke, T., Lang, S., Hay, G.J., Eds.; Springer-Verlag: Berlin Heidelberg, Germany, 2008; pp. 91–110. [Google Scholar]
- Kim, M.; Madden, M.; Warner, T.A. Forest type mapping using object-specific texture measures from multispectral ikonos imagery: Segmentation quality and image classification issues. Photogramm. Eng. Remote Sens 2009, 75, 819–829. [Google Scholar]
- Osborne, P.E.; Suárez-Seoane, S. Should data be partitioned spatially before building large-scale distribution models? Ecol. Model 2002, 157, 249–259. [Google Scholar]
- Peres-Neto, P.R.; Legendre, P.; Dray, S.; Borcard, D. Variation partitioning of species data matrices: Estimation and comparison of fractions. Ecology 2006, 87, 2614–2625. [Google Scholar]
- Bailey, R.G. Description of the Ecoregions of the United States; United States Forest Service: Ogden, UT, USA, 1995. [Google Scholar]
- Jarvis, A.; Reuter, H.I.; Nelson, A.; Guevara, E. Hole-Filled Seamless Srtm Data v4, International Centre for Tropical Agriculture (CIAT). Available online: http://srtm.csi.cgiar.org/ (accessed on 20 June 2012).
- Menne, M.J.; Williams, J.; Vose, R.S. Long-Term Daily and Monthly Climate Records from Stations Across the Contiguous United States. Available online: http://cdiac.ornl.gov/epubs/ndp/ushcn/ushcn.html (accessed on 30 April 2014).
- Burns, R.M.; Honkala, B.H. Silvics of North America: 1. Conifers; 2. Hardwoods; United States Department of Agriculture Forest Service: Washington, DC, USA, 1990; Volume 2. [Google Scholar]
- DeGomez, T. Determining ponderosa pine tree density on small lots. Backyards Beyond 2008, 8, 8–10. [Google Scholar]
- Fitzgerald, S.A. Fire Ecology of Ponderosa Pine and the Rebuilding of Fire-Resilient Ponderosa Pine Ecosystems; U.S. Department of Agriculture, Forest Service: Albany, CA, USA, 2005. [Google Scholar]
- Habeck, R.J. Pinus Ponderosa var. Ponderosa, 1992, Available online: http://egsc.usgs.gov/isb/pubs/factsheets/fs05701.html (accessed on 10 Decebmer 2012).
- Survey, U.S.G. Digital Orthophoto Quadrangles Fact Sheet 057–01; United State Geological Survey: Reston, VA, USA, 2001. [Google Scholar]
- Part. 2 Specifications Standards for Digital Orthophotos; U.S. Geological Survey National Mapping Division, 1996. Available online: http://nationalmap.gov/standards/doqstds.html (accessed on 10 March 2014).
- Hoagland, S.; Krieger, A.; Moy, S.; Shepard, A. Ecology and Management of Oak Woodlands on Tejon Ranch: Recommendations for Conserving a Valuable California Ecosystem; 2011 Group Project Final Report; University of California: Santa Barbara, CA, USA, 2011. [Google Scholar]
- McCreary, D.D. Demography and Recruitment Limitations of Three Oak Species in California; University of California: Oakland, CA, USA, 2001. [Google Scholar]
- Otsu, N. A tlreshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern 1979, 9, 62–66. [Google Scholar]
- Grady, L. Random walks for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell 2006, 28, 1768–1783. [Google Scholar]
- Haralick, R.M. Statistical and structural approach to texture. Proc. IEEE 1979, 67, 786–804. [Google Scholar]
- Haralick, R.M.; Shanmugam, K.; Dinstein, I. Textural features for image classification. IEEE Trans. Syst 1973, 3, 610–621. [Google Scholar]
- Lillesand, T.M.; Kiefer, R.W.; Chipman, J.W. Remote Sensing and Image Interpretation; Wiley: Hoboken, NJ, USA, 2004. [Google Scholar]
- Liu, D.; Xia, F. Asscssing object-based classification: Advantages and limitations. Remote Sens. Lett 2010, 1, 187–194. [Google Scholar]
- Morgan, J.L.; Gergel, S.E.; Coops, N.C. Aerial photography: A rapidly evolving tool for ecological management. BioScience 2010, 60, 47–59. [Google Scholar]
- Ryherd, S.; Woodcock, C. Combining spectral and texture data in the segmentation of remotely sensed images. Photogramm. Eng. Remote Sens 1996, 62, 181–194. [Google Scholar]
- Clinton, N.; Holt, A.; Scarborough, J.; Yan, L.; Gong, P. Accuracy assessment measures for object-based image segmentation goodness. Photogramm. Eng. Remote Sens 2010, 76, 289–299. [Google Scholar]
- Banerjee, M.; Capozzoli, M.; McSweeney, L.; Sinha, D. Beyond kappa: A review of interrater agreement measures. Can. J. Stat 1999, 27, 3–23. [Google Scholar]
- Pontius, R.G., Jr.; Millones, M. Death to kappa birth of quantity disagreement and allocation disagreement for accuracy assessment. Int. J. Remote Sens 2011, 32, 4407–4429. [Google Scholar]
- Franklin, J.; Keeler-Wolf, T.; Thomas, K.A.; Shaari, D.A.; Stine, P.A.; Michaelsen, J.; Miller, J. Stratified sampling for field survey of environmental gradients in the mojave desert ecoregion. In GIS and Remote Sensing Application in Biogeography and Ecology; Millington, A.C., Walsh, S.J., Osborne, P.E., Eds.; Kluwer Academic Publishers: Massachusetts, MA, USA, 1999; pp. 229–253. [Google Scholar]
- Cohen, W.B.; Goward, S.N. Landsat’s role in ecological applications of remote sensing. BioScience 2004, 54, 535–545. [Google Scholar]
- Turner, M.G.; Dale, V.H. Comparing large, infrequent disturbances: What have we learned? Ecosystems 1998, 1, 493–496. [Google Scholar]
- Thuiller, W.; Albert, C.; Araújo, M.B.; Berry, P.M.; Cabeza, M.; Guisan, A.; Hickler, T.; Midgley, G.F.; Paterson, J.; Schurr, F.M.; et al. Predicting global change impacts on plant species’ distributions: Future challenges. Perspect. Plant Ecol. Evol. Syst 2008, 9, 137–152. [Google Scholar]
- Tyler, C.M.; Kuhn, B.; Davis, F.W. Demography and recruitment limitations of three oak species in California. Quart. Rev. Biol 2006, 81, 127–152. [Google Scholar]
- Brown, D.G.; Band, L.E.; Green, K.O.; Irwin, E.G.; Jain, A.; Lambin, E.F.; Pontius, R.G.J.; Seto, K.C.; Turner, B.L.I.; Verburg, P.H. Advancing Land Change Modeling: Opportunities and Research Requirements; The National Academies Press: Washington, DC, USA, 2013. [Google Scholar]


| (a) Timber Survey | |||
| Elevation Stratification | Slope Stratification | Aspect Stratification | |
| Kappa value | 0.52 | 0.56 | 0.65 |
| Overall accuracies | 0.76 | 0.78 | 0.83 |
| Users’ accuracies | |||
| Barren land | 0.72 | 0.8 | 0.88 |
| High density | 0.81 | 0.76 | 0.76 |
| Producers’ accuracies | |||
| Blue-oak and grey pine | 0.82 | 0.8 | 0.81 |
| Black oak and ponderosa pine | 0.71 | 0.76 | 0.84 |
| (b) CALVEG | |||
| Elevation Stratification | Slope Stratification | Aspect Stratification | |
| Kappa value | 0.61 | 0.64 | 0.65 |
| Overall accuracies | 0.80 | 0.82 | 0.82 |
| Users’ accuracies | |||
| Barren land | 0.64 | 0.76 | 0.72 |
| High density | 0.96 | 0.88 | 0.92 |
| Producers’ accuracies | |||
| Blue-oak and grey pine | 0.94 | 0.86 | 0.9 |
| Black oak and ponderosa pine | 0.74 | 0.79 | 0.77 |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Liau, Y.-T. Hierarchical Segmentation Framework for Identifying Natural Vegetation: A Case Study of the Tehachapi Mountains, California. Remote Sens. 2014, 6, 7276-7302. https://doi.org/10.3390/rs6087276
Liau Y-T. Hierarchical Segmentation Framework for Identifying Natural Vegetation: A Case Study of the Tehachapi Mountains, California. Remote Sensing. 2014; 6(8):7276-7302. https://doi.org/10.3390/rs6087276
Chicago/Turabian StyleLiau, Yan-Ting. 2014. "Hierarchical Segmentation Framework for Identifying Natural Vegetation: A Case Study of the Tehachapi Mountains, California" Remote Sensing 6, no. 8: 7276-7302. https://doi.org/10.3390/rs6087276
APA StyleLiau, Y.-T. (2014). Hierarchical Segmentation Framework for Identifying Natural Vegetation: A Case Study of the Tehachapi Mountains, California. Remote Sensing, 6(8), 7276-7302. https://doi.org/10.3390/rs6087276




