Advancing Forest Inventory and Fuel Monitoring with Multi-Sensor Hybrid Models: A Comparative Framework for Basal Area Estimation
Highlights
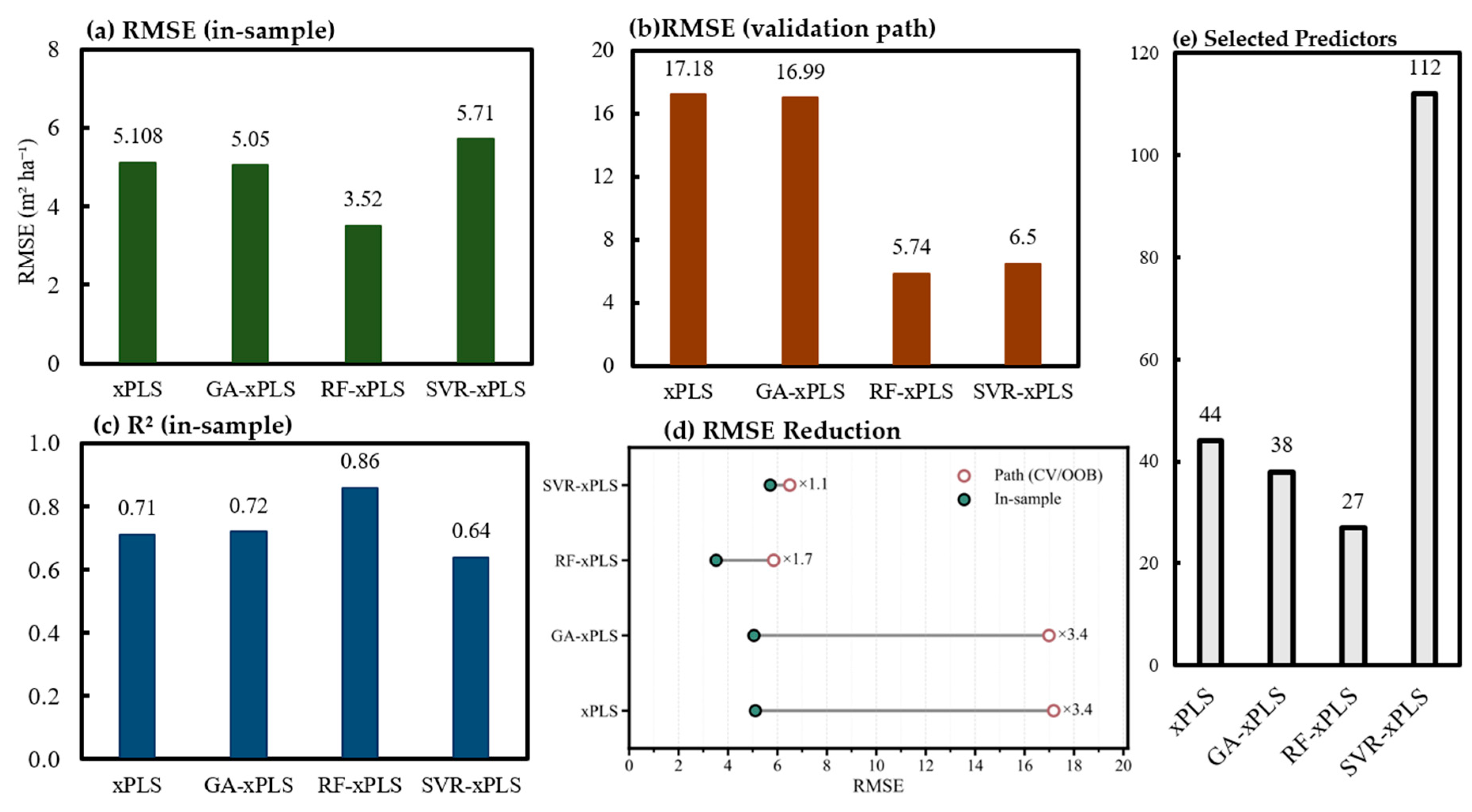
- RF-xPLS achieved the strongest pooled model performance for total and species-level basal area (BA), with well-behaved residuals and tight bootstrap confidence intervals.
- A parsimonious 27-predictor subset preserved most of the skill, dominated by SWIR/NDII (canopy water/dry-matter), red-edge/NIR greenness features, and a single LiDAR structure metric (HQUAD).
- Hybrid selection can reduce a high-dimensional, collinear multi-sensor feature space to a compact, interpretable set without sacrificing accuracy—supporting operational, wall-to-wall BA mapping.
- The selected predictors indicate which inputs are most informative and where additional field plots (especially in very high-BA stands) would most effectively improve generalization and uncertainty.
Abstract
1. Introduction
- Evaluate predictive performance of the four approaches (xPLS, GA-xPLS, RF-xPLS, and SVR-xPLS) in terms of RMSE and R2.
- Identify most influential predictors of total and species-specific conifer BA.
- Assess the complementarity of structural (LiDAR) and spectral (multispectral) predictors in explaining BA variation.
2. Materials and Methods
2.1. Study Area and Data Sources
2.1.1. Predictor Variables
2.1.2. Field Data and Response Variables
2.2. Modeling Framework
2.2.1. Iterative Exclusion PLS (xPLS)
2.2.2. GA-xPLS
2.2.3. RF-xPLS
2.2.4. SVR-xPLS
3. Results
3.1. Field Inventory and Predictor Overview
3.2. Model Selection and Cross-Validated Accuracy
3.3. Species-Level Performance
3.4. Model Skill Across Responses
3.5. Observation vs. Prediction
3.6. Selected Predictors by Recommended Model
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| (A) RF-xPLS: Iterative Exclusion with Random Forests, Selecting Removals by Minimum Pooled OOB RMSE | ||
| Item | Setting used | Implementation detail |
| RF algorithm | Random Forest regression | MATLAB TreeBagger (Statistics and ML Toolbox) |
| Number of trees | NumTrees = 500 | Fixed for all responses |
| Minimum leaf size | MinLeaf = 5 | Fixed for all responses |
| Predictors per split (mtry) | mtry = round(sqrt(p_init)), capped as p shrinks | NumPredictK0 = round(sqrt(size(Xmat,2))); mtry = min(size(Xok,2), NumPredictK0) |
| OOB prediction | On | ‘OOBPrediction’, ‘on’ |
| Selection criterion | Minimum pooled OOB RMSE across responses | At each step, drop variable yielding lowest pooled OOB RMSE |
| Permutation importance | OOB permuted importance (diagnostic) | oobPermutedPredictorImportance(tb) (try/catch for compatibility) |
| (B) GA-xPLS: Subset Search Under a Hard Size Cap, Using Repeated K-Fold CV Fitness | ||
| GA representation | Binary mask (0/1 predictors) | PopulationType = ‘doubleVector’ with IntCon = 1:nVars, rounded to 0/1 |
| Population size | 350 | ‘PopulationSize’, 350 |
| Max generations | 800 | ‘MaxGenerations’, 800 |
| Stall generation limit | Inf (disabled) | ‘StallGenLimit’, inf |
| Selection function | Tournament selection | ‘SelectionFcn’, {@selectiontournament, 6} |
| Tournament size | 6 | tournSize = 6 |
| Crossover function | Two-point crossover | ‘CrossoverFcn’, @crossovertwopoint |
| Crossover fraction | 0.7 | ‘CrossoverFraction’, 0.70 |
| Mutation function | Uniform mutation | ‘MutationFcn’, {@mutationuniform, 0.25} |
| Mutation rate | 0.25 | mutationRate = 0.25 |
| Size penalty weight | λ = 0.10 | penSize = λ × (s/p) |
| Band-violation penalty | 0.5 | bandPenalty = 0.5 |
| Fitness CV scheme | Repeated K-fold CV: K = 10, repeats = 3 | useKFoldFitness = true; cvK = 10; cvRepeats = 3 |
| (C) SVR-xPLS: Iterative Exclusion with SVR Using LOOCV, Fixed CCC, and Response-Specific ε | ||
| SVR algorithm | Support Vector Regression | MATLAB fitrsvm (Statistics and ML Toolbox) |
| Kernel | linear or RBF (user choice) | KernelFunction = ‘linear’ or ‘rbf’ |
| BoxConstraint (C) | C = 1.0 | globalC = 1.0 used for all responses |
| Epsilon (ε) | ε = 0.1 × std(y) (response-specific) | respEpsilon = std(y_current) × 0.1 per response model |
| CV scheme during exclusion | LOOCV | Inner loop over observations: for cvIdx = 1:numObs |
| Selection criterion | Minimum pooled RMSE across responses | pooled RMSE = sqrt(mean(rmseVec.^2)) |
Appendix B


Appendix C



Appendix D

Appendix E
| Scenario | nPred | nS2 | nL9 | nLiDAR | RMSE_Pooled_Train | R2_Pooled_Train |
|---|---|---|---|---|---|---|
| S2_only | 23 | 23 | 0 | 0 | 3.70 | 0.84 |
| L9_only | 3 | 0 | 3 | 0 | 6.13 | 0.59 |
| LiDAR_only | 1 | 0 | 0 | 1 | 7.27 | 0.42 |
| Optical_S2 + L9 | 26 | 23 | 3 | 0 | 3.55 | 0.86 |
| S2 + LiDAR | 24 | 23 | 0 | 1 | 3.56 | 0.86 |
| L9 + LiDAR | 4 | 0 | 3 | 1 | 5.59 | 0.66 |
| All_S2 + L9 + LiDAR | 27 | 23 | 3 | 1 | 3.48 | 0.88 |
References
- Heinselman, M.L. Fire in the virgin forests of the Boundary Waters Canoe Area, Minnesota. Quat. Res. 1973, 3, 329–382. [Google Scholar] [CrossRef]
- Ohmann, L.F.; Grigal, D.F. Early revegetation and nutrient dynamics following the 1971 Little Sioux Forest fire in northeastern Minnesota. For. Sci. 1979, 25, a0001–z0001. [Google Scholar] [CrossRef]
- Frelich, L.E.; Reich, P.B. Spatial patterns and succession in a Minnesota southern-boreal forest. Ecol. Monogr. 1995, 65, 325–346. [Google Scholar] [CrossRef]
- Ali, A.A.; Asselin, H.; Larouche, A.C.; Bergeron, Y.; Carcaillet, C.; Richard, P.J. Changes in fire regime explain the Holocene rise and fall of Abies balsamea in the coniferous forests of western Québec, Canada. Holocene 2008, 18, 693–703. [Google Scholar] [CrossRef]
- Scheller, R.M.; Mladenoff, D.J.; Crow, T.R.; Sickley, T.A. Simulating the effects of fire reintroduction versus continued fire absence on forest composition and landscape structure in the Boundary Waters Canoe Area, northern Minnesota, USA. Ecosystems 2005, 8, 396–411. [Google Scholar] [CrossRef]
- Brown, J.K.; Smith, J.K. Wildland Fire in Ecosystems: Effects of Fire on Flora; General Technical Report RMRS-GTR-42-vol. 2; US Department of Agriculture, Forest Service, Rocky Mountain Research Station: Ogden, UT, USA, 2000; 257p. [Google Scholar]
- de Lafontaine, G.; Payette, S. Long-term fire and forest history of subalpine balsam fir (Abies balsamea) and white spruce (Picea glauca) stands in eastern Canada inferred from soil charcoal analysis. Holocene 2011, 22, 191–201. [Google Scholar] [CrossRef]
- Dickinson, M.B.; Johnson, E.A.; Artiaga, R. Fire spread probabilities for experimental beds composed of mixedwood boreal forest fuels. Can. J. For. Res. 2013, 43, 321–330. [Google Scholar] [CrossRef]
- Kreider, M.R.; Higuera, P.E.; Parks, S.A.; Rice, W.L.; White, N.; Larson, A.J. Fire suppression makes wildfires more severe and accentuates impacts of climate change and fuel accumulation. Nat. Commun. 2024, 15, 2412. [Google Scholar] [CrossRef] [PubMed]
- Engelstad, P.S.; Falkowski, M.; Wolter, P.; Poznanovic, A.; Johnson, P. Estimating Canopy Fuel Attributes from Low-Density LiDAR. Fire 2019, 2, 38. [Google Scholar] [CrossRef]
- Szpakowski, D.M.; Jensen, J.L.R. A review of the applications of remote sensing in fire ecology. Remote Sens. 2019, 11, 2638. [Google Scholar] [CrossRef]
- Wolter, P.T.; Olbrich, J.J.; Johnson, P.J. Modeling sub-boreal forest canopy bulk density in Minnesota, USA, using synthetic aperture radar and optical satellite sensor data. Fire Ecol. 2021, 17, 26. [Google Scholar] [CrossRef]
- Wear, D.N.; Wibbenmeyer, M.; Joiner, E. Enhancing the economic feasibility of fuel treatments: Market and policy pathways for US Federal Lands. For. Policy Econ. 2024, 169, 103365. [Google Scholar] [CrossRef]
- Pierson, D.; Andika, R.; Brewen, J.; Clark, N.; Hardy, M.C.; McCollum, D.; McCormick, F.H.; Morisette, J.; Nicosia, T.; Page-Dumroese, D.; et al. Beyond the Basics: A Perspective on Barriers and Opportunities for Scaling Up Biochar Production from Forest Slash. Biochar 2024, 6, 1. [Google Scholar] [CrossRef]
- Puettmann, M.; Sahoo, K.; Wilson, K.; O’Neil, E. Life cycle assessment of biochar produced from forest residues using portable systems. J. Clean. Prod. 2020, 250, 119564. [Google Scholar] [CrossRef]
- Zhao, T.; Li, G.; Zhang, S.; Yang, H.; Ni, W.; Shao, A. Enhancement effect of biochar on the immobilization of heavy metals and anions in MSWI fly ash/bottom ash–coal fly ash-based cementitious materials: Risk assessment model and mechanisms. J. Hazard. Mater. 2025, 497, 139615. [Google Scholar] [CrossRef]
- Trapero, J.R.; Alcazar-Ruiz, A.; Dorado, F.; Sanchez-Silva, L. Biochar price forecasting: A novel methodology for enhancing market stability and economic viability. J. Environ. Manag. 2025, 377, 124681. [Google Scholar] [CrossRef]
- Wolter, P.T.; Townsend, P.A.; Sturtevant, B.R. Estimation of forest structural parameters using 5 and 10 meter SPOT-5 satellite data. Remote Sens. Environ. 2009, 113, 2019–2036. [Google Scholar] [CrossRef]
- Wolter, P.T.; Townsend, P.A. Multi-sensor data fusion for estimating forest species composition and abundance in northern Minnesota. Remote Sens. Environ. 2011, 115, 671–691. [Google Scholar] [CrossRef]
- Hovind, H.J.; Rieck, C.E. Basal Area and Point-Sampling: Interpretation and Application (No. 23); Department of Natural Resources: Madison, WI, USA, 1961. [Google Scholar]
- O’Brien, R. Comprehensive Inventory of Utah’s Forest Resources, 1993; US Department of Agriculture, Forest Service, Rocky Mountain Research Station: Ogden, UT, USA, 1999. [Google Scholar] [CrossRef]
- Vanclay, J.K.; Sands, P.J. Calibrating the self-thinning frontier. For. Ecol. Manag. 2009, 259, 81–85. [Google Scholar] [CrossRef]
- Bhattarai, R.; Rahimzadeh-Bajgiran, P.; Weiskittel, A.; Meneghini, A.; MacLean, D.A. Spruce budworm tree host species distribution and abundance mapping using multi-temporal Sentinel-1 and Sentinel-2 satellite imagery. ISPRS J. Photogramm. Remote Sens. 2021, 172, 28–40. [Google Scholar] [CrossRef]
- Balderas Torres, A.; Lovett, J.C. Using basal area to estimate aboveground carbon stocks in forests: La Primavera Biosphere’s Reserve, Mexico. Forestry 2013, 86, 267–281. [Google Scholar] [CrossRef]
- Fischer, S.M.; Wang, X.; Huth, A. Distinguishing mature and immature trees allows estimating forest carbon uptake from stand structure. Biogeosciences 2024, 21, 3305–3319. [Google Scholar] [CrossRef]
- Pretzsch, H. Forest Dynamics, Growth and Yield; Springer: Berlin/Heidelberg, Germany, 2009. [Google Scholar] [CrossRef]
- Yanai, R.D.; Young, A.; Campbell, J.L.; Westfall, J.A.; Barnett, C.J.; Dillon, G.A.; Green, M.B.; Woodall, C.W. Measurement uncertainty in a national forest inventory: Results from the northern region of the USA. Can. J. For. Res. 2023, 53, 163–177. [Google Scholar] [CrossRef]
- Duncanson, L.; Kellner, J.R.; Armston, J.; Dubayah, R.; Minor, D.M.; Hancock, S.; Healey, S.P.; Patterson, P.L.; Saarela, S.; Marselis, S.; et al. Aboveground biomass density models for NASA’s Global Ecosystem Dynamics Investigation (GEDI) lidar mission. Remote Sens. Environ. 2022, 270, 112845. [Google Scholar] [CrossRef]
- Lahssini, K.; Teste, F.; Dayal, K.R.; Durrieu, S.; Ienco, D.; Monnet, J.M. Combining LiDAR Metrics and Sentinel-2 Imagery to Estimate Basal Area and Wood Volume in Complex Forest Environment via Neural Networks. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2022, 15, 4337–4348. [Google Scholar] [CrossRef]
- Wulder, M.A.; White, J.C.; Nelson, R.F.; Næsset, E.; Ørka, H.O.; Coops, N.C.; Hilker, T.; Bater, C.W.; Gobakken, T. Lidar sampling for large-area forest characterization: A review. Remote Sens. Environ. 2012, 121, 196–209. [Google Scholar] [CrossRef]
- Féret, J.-B.; Asner, G.P. Mapping tropical forest canopy diversity using high-fidelity imaging spectroscopy. Ecol. Appl. 2014, 24, 1289–1296. [Google Scholar] [CrossRef] [PubMed]
- White, J.C.; Coops, N.C.; Wulder, M.A.; Vastaranta, M.; Hilker, T.; Tompalski, P. Remote Sensing Technologies for Enhancing Forest Inventories: A Review. Can. J. Remote Sens. 2016, 42, 619–641. [Google Scholar] [CrossRef]
- Allouis, T.; Durrieu, S.; Vega, C.; Couteron, P. Stem volume and above-ground biomass estimation of individual pine trees from lidar data: Contribution of full-waveform signals. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2013, 6, 924–934. [Google Scholar] [CrossRef]
- Luo, S.; Wang, C.; Xi, X.; Pan, F.; Peng, D.; Zou, J.; Nie, S.; Qin, H. Fusion of airborne LiDAR data and hyperspectral imagery for aboveground and belowground forest biomass estimation. Ecol. Indic. 2017, 73, 378–387. [Google Scholar] [CrossRef]
- Zhang, W.; Wan, P.; Wang, T.; Cai, S.; Chen, Y.; Jin, X.; Yan, G. A Novel Approach for the Detection of Standing Tree Stems from Plot-Level Terrestrial Laser Scanning Data. Remote Sens. 2019, 11, 211. [Google Scholar] [CrossRef]
- Berra, E.F.; Gaulton, R. Remote sensing of temperate and boreal forest phenology: A review of progress, challenges and opportunities in the intercomparison of in-situ and satellite phenological metrics. For. Ecol. Manag. 2021, 1480, 118663. [Google Scholar] [CrossRef]
- Gong, Z.; Ge, W.; Guo, J.; Liu, J. Satellite remote sensing of vegetation phenology: Progress, challenges, and opportunities. ISPRS J. Photogramm. Remote Sens. 2024, 217, 149–164. [Google Scholar] [CrossRef]
- Wolter, P.T.; Mladenoff, D.J.; Host, G.; Crow, T.R. Improved Forest classification in the northern Lake States using multi-temporal Landsat imagery. Photogramm. Eng. Remote Sens. 1995, 61, 1129–1143. [Google Scholar]
- Brown, S.; Narine, L.L.; Gilbert, J. Using airborne lidar, multispectral imagery, and field inventory data to estimate basal area, volume, and aboveground biomass in heterogeneous mixed species forests: A case study in southern Alabama. Remote Sens. 2022, 14, 2708. [Google Scholar] [CrossRef]
- Kuhn, M.; Johnson, K. Applied Predictive Modeling; Springer: Berlin/Heidelberg, Germany, 2013. [Google Scholar] [CrossRef]
- Maltamo, M.; Næsset, E.; Vauhkonen, J. Forestry Applications of Airborne Laser Scanning: Concepts and Case Studies; Springer Netherlands: Dordrecht, The Netherlands, 2014. [Google Scholar] [CrossRef]
- Wolter, P.; Townsend, P.A.; Sturtevant, B.R.; Kingdon, C.C. Remote sensing of the distribution and abundance of host species for spruce budworm in Northern Minnesota and Ontario. Remote Sens. Environ. 2008, 112, 3971–3982. [Google Scholar] [CrossRef]
- Chong, I.G.; Jun, C.H. Performance of some variable selection methods when multicollinearity is present. Chemom. Intell. Lab. Syst. 2005, 78, 103–112. [Google Scholar] [CrossRef]
- Spiegelman, C.H.; McShane, M.J.; Goetz, M.J.; Motamedi, M.; Yue, Q.L.; Coté, G.L. Theoretical justification of wavelength selection in PLS calibration: Development of a new algorithm. Anal. Chem. 1998, 70, 35–44. [Google Scholar] [CrossRef]
- Heinze, G.; Wallisch, C.; Dunkler, D. Variable selection—A review and recommendations for the practicing statistician. Biom. J. 2018, 60, 431–449. [Google Scholar] [CrossRef]
- Moser, P.; Vibrans, A.C.; McRoberts, R.E.; Næsset, E.; Gobakken, T.; Chirici, G.; Mura, M.; Marchetti, M. Methods for variable selection in LiDAR-assisted Forest inventories. Forestry 2017, 90, 112–124. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests. Mach. Learn. 2004, 5, 5–32. [Google Scholar] [CrossRef]
- Guyon, I.; Weston, J.; Barnhill, S.; Vapnik, V. Gene selection for cancer classification using Support Vector Machines. Mach. Learn. 2002, 46, 389–422. [Google Scholar] [CrossRef]
- Bhattarai, R.; Rahimzadeh-Bajgiran, P.; Weiskittel, A.; Homayouni, S.; Gara, T.W.; Hanavan, R.P. Estimating species-specific leaf area index and basal area using optical and SAR remote sensing data in Acadian mixed spruce-fir forests, USA. Int. J. Appl. Earth Obs. Geoinf. 2022, 108, 102727. [Google Scholar] [CrossRef]
- Casas-Gómez, P.; Torres, J.F.; Linares, J.C.; Troncoso, A.; Martínez-Álvarez, F. Forecasting basal area increment in forest ecosystems using deep learning: A multi-species analysis in the Himalayas. Ecol. Inform. 2025, 85, 102951. [Google Scholar] [CrossRef]
- McKearnan, S.B.; Vock, D.M.; Marai, G.E.; Canahuate, G.; Fuller, C.D.; Wolfson, J. Feature selection for support vector regression using a genetic algorithm. Biostatistics 2023, 24, 295–308. [Google Scholar] [CrossRef]
- Yang, P.; Li, C.; Qiu, Y.; Huang, S.; Zhou, J. Metaheuristic Optimization of Random Forest for Predicting Punch Shear Strength of FRP-Reinforced Concrete Beams. Materials 2023, 16, 4034. [Google Scholar] [CrossRef] [PubMed]
- Feng, G. Feature selection algorithm based on optimized genetic algorithm and the application in high-dimensional data processing. PLoS ONE 2024, 19, e0303088. [Google Scholar] [CrossRef] [PubMed]
- Leardi, R.; González, A.L. Genetic algorithms applied to feature selection in PLS regression: How and when to use them. Chemom. Intell. Lab. Syst. 1998, 41, 195–207. [Google Scholar] [CrossRef]
- Mehmood, T.; Liland, K.H.; Snipen, L.; Sæbø, S. A review of variable selection methods in partial least squares regression. Chemom. Intell. Lab. Syst. 2012, 118, 62–69. [Google Scholar] [CrossRef]
- Yun, Y.H.; Bin, J.; Liu, D.L.; Xu, L.; Yan, T.-L.; Cao, D.-S.; Xu, Q.-S. A hybrid variable selection strategy based on continuous shrinkage of variable space in multivariate calibration. Anal. Chim. Acta 2019, 1058, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Yoo, D.G.; Kim, J.H. Meta-heuristic algorithms as tools for hydrological science. Geosci. Lett. 2014, 1, 4. [Google Scholar] [CrossRef]
- Adnan, R.M.; Liang, Z.; Trajkovic, S.; Zounemat-Kermani, M.; Li, B.; Kisi, O. Daily streamflow prediction using optimally pruned extreme learning machine. J. Hydrol. 2021, 599, 126354. [Google Scholar] [CrossRef]
- Salehnia, N.; Ahn, J. Modelling and reconstructing tree ring growth index with climate variables through artificial intelligence and statistical methods. Ecol. Indic. 2022, 134, 108496. [Google Scholar] [CrossRef]
- Wolter, P.T.; Berkley, E.A.; Peckham, S.D.; Singh, A.; Townsend, P.A. Exploiting tree shadows on snow for estimating forest basal area using Landsat data. Remote Sens. Environ. 2012, 121, 69–79. [Google Scholar] [CrossRef][Green Version]
- Van Wagner, C.E. Conditions for the start and spread of crown fire. Can. J. For. Res. 1977, 7, 23–34. [Google Scholar] [CrossRef]
- Scott, J.H.; Reinhardt, E.D. Assessing Crown Fire Potential by Linking Models of Surface and Crown Fire Behavior (No. 29); US Department of Agriculture, Forest Service, Rocky Mountain Research Station: Fort Collins, CO, USA, 2001. [Google Scholar] [CrossRef]
- Brandt, J.P. The extent of the North American boreal zone. Environ. Rev. 2009, 17, 101–161. [Google Scholar] [CrossRef]
- Baker, W.L. Landscape ecology and nature reserve design in the Boundary Waters Canoe Area, Minnesota. Ecology 1989, 70, 23–35. [Google Scholar] [CrossRef]
- Wolter, P.T.; White, M.A. Recent forest cover type transitions and landscape structural changes in northeast Minnesota. Landsc. Ecol. 2002, 17, 133–155. [Google Scholar] [CrossRef]
- Pastor, J.; Sharp, A.; Wolter, P. An application of Markov models to the dynamics of Minnesota’s forests. Can. J. For. Res. 2005, 35, 3011–3019. [Google Scholar] [CrossRef]
- Friedman, S.K.; Reich, P.B. Regional legacies of logging: Departure from presettlement forest conditions in northern Minnesota. Ecol. Appl. 2005, 15, 726–744. [Google Scholar] [CrossRef]
- Grosenbaugh, L.R. Plotless timber estimates—New, fast, easy. J. For. 1952, 50, 322–337. [Google Scholar] [CrossRef]
- Joyce, M.; Moen, R. Accuracy of a Modular GPS/GLONASS Receiver; Report Number: NRRI/TR-2018/28, Release 1.0; University of Minnesota Duluth: Duluth, MN, USA, 2018; Available online: https://hdl.handle.net/11299/204331 (accessed on 20 December 2025).
- Wold, S.; Sjostrom, M.; Eriksson, L. PLS-regression: A basic tool of chemometrics. Chemom. Intell. Lab. Syst. 2001, 58, 109–130. [Google Scholar] [CrossRef]
- Tran, T.N.; Afanador, N.L.; Buydens, L.M.; Blanchet, L. Interpretation of variable importance in Partial Least Squares with Significance Multivariate Correlation (sMC). Chemom. Intell. Lab. Syst. 2014, 138, 153–160. [Google Scholar] [CrossRef]
- Hasegawa, K.; Miyashita, Y.; Funatsu, K. GA Strategy for Variable Selection in QSAR Studies: GA-Based PLS Analysis of Calcium Channel Antagonists. J. Chem. Inf. Comput. Sci. 1997, 37, 306–310. [Google Scholar] [CrossRef] [PubMed]
- Leardi, R. Application of genetic algorithm–PLS for feature selection in spectral data sets. J. Chemom. 2000, 14, 643–655. [Google Scholar] [CrossRef]
- Zareef, M.; Arslan, M.; Hassan Md, M.; Ahmad, W.; Chen, Q. Comparison of Si-GA-PLS and Si-CARS-PLS build algorithms for quantitation of total polyphenols in black tea using the spectral analytical system. J. Sci. Food Agric. 2023, 103, 7914–7920. [Google Scholar] [CrossRef]
- Altmann, A.; Tolosi, L.; Sander, O.; Lengauer, T. Permutation importance: A corrected feature importance measure. Bioinformatics 2010, 26, 1340–1347. [Google Scholar] [CrossRef] [PubMed]
- Genuer, R.; Poggi, J.M.; Tuleau-Malot, C. Variable selection using random forests. Pattern Recognit. Lett. 2010, 31, 2225–2236. [Google Scholar] [CrossRef]
- Smola, A.J.; Schölkopf, B. A tutorial on support vector regression. Stat. Comput. 2004, 14, 199–222. [Google Scholar] [CrossRef]
- Bland, J.M.; Altman, D.G. Statistics Notes: Bootstrap resampling methods. BMJ 2015, 350, h2622. [Google Scholar] [CrossRef]
- Keane, R.E.; Burgan, R.; van Wagtendonk, J. Mapping wildland fuels for fire management across multiple scales: Integrating remote sensing, GIS, and biophysical modeling. IJWF 2001, 10, 301–319. [Google Scholar] [CrossRef]
- Bell, D.M.; Gregory, M.J.; Churchill, D.J.; Smith, A.C. Mapping with height and spectral remote sensing implies that environment and forest structure jointly constrain tree community composition in temperate coniferous forests of eastern Washington, United States. Front. For. Glob. Change 2022, 5, 962816. [Google Scholar] [CrossRef]
- Richardson, J.J.; Moskal, L.M. Strengths and limitations of assessing forest density and spatial configuration with aerial LiDAR. Remote Sens. Environ. 2011, 115, 2640–2651. [Google Scholar] [CrossRef]
- Arellano-Pérez, S.; Castedo-Dorado, F.; López-Sánchez, C.A.; González-Ferreiro, E.; Yang, Z.; Díaz-Varela, R.A.; Álvarez-González, J.G.; Vega, J.A.; Ruiz-González, A.D. Potential of Sentinel-2A data to model surface and canopy fuel characteristics in relation to crown fire hazard. Remote Sens. 2018, 10, 1645. [Google Scholar] [CrossRef]
- Buckley, D.S.; Isebrands, J.G.; Sharik, T.L. Practical field methods of estimating canopy cover, PAR, and LAI in Michigan Oak and pine stands. North. J. Appl. For. 1999, 16, 25–32. [Google Scholar] [CrossRef]
- Féret, J.B.; Gitelson, A.A.; Noble, S.D.; Jacquemoud, S. PROSPECT-D: Towards modeling leaf optical properties through a complete lifecycle. Remote Sens. Environ. 2017, 193, 204–215. [Google Scholar] [CrossRef]
- Cavender-Bares, J.; Gamon, J.A.; Townsend, P.A. The Use of Remote Sensing to Enhance Biodiversity Monitoring and Detection: A Critical Challenge for the Twenty-First Century. In Remote Sensing of Plant Biodiversity; Cavender-Bares, J., Gamon, J.A., Townsend, P.A., Eds.; Springer: Cham, Switzerland, 2020. [Google Scholar] [CrossRef]
- Zhou, H.; Zhou, G.; Song, X.; He, Q. Dynamic Characteristics of Canopy and Vegetation Water Content during an Entire Maize Growing Season in Relation to Spectral-Based Indices. Remote Sens. 2022, 14, 584. [Google Scholar] [CrossRef]
- Eliades, F.; Sarris, D.; Bachofer, F.; Michaelides, S.; Hadjimitsis, D. Understanding Tree Mortality Patterns: A Comprehensive Review of Remote Sensing and Meteorological Ground-Based Studies. Forests 2024, 15, 1357. [Google Scholar] [CrossRef]
- Ma, H.; Weiss, M.; Malik, D.; Berthelot, B.; Yebra, M.; Nolan, R.H.; Mialon, A.; Zeng, J.; Quan, X.; Tagesson, H.T.; et al. Satellite canopy water content from Sentinel-2, Landsat-8 and MODIS: Principle, algorithm and assessment. Remote Sens. Environ. 2025, 326, 114801. [Google Scholar] [CrossRef]
- Belgiu, M.; Dragut, L. Random Forest in Remote Sensing: A Review of Applications and Future Directions. ISPRS J. Photogramm. Remote Sens. 2016, 114, 24–31. [Google Scholar] [CrossRef]
- Vidican, R.; Mălinaș, A.; Ranta, O.; Moldovan, C.; Marian, O.; Ghețe, A.; Ghișe, C.R.; Popovici, F.; Cătunescu, G.M. Using Remote Sensing Vegetation Indices for the Discrimination and Monitoring of Agricultural Crops: A Critical Review. Agronomy 2023, 13, 3040. [Google Scholar] [CrossRef]
- Liu, J.; Pattey, E.; Jego, G. Assessment of vegetation indices for regional crop green LAI estimation from Landsat images over multiple growing seasons. Remote Sens. Environ. 2012, 123, 347–358. [Google Scholar] [CrossRef]
- Vélez, S.; Martínez-Peña, R.; Castrillo, D. Beyond Vegetation: A Review Unveiling Additional Insights into Agriculture and Forestry through the Application of Vegetation Indices. J 2023, 6, 421–436. [Google Scholar] [CrossRef]
- Bhattarai, R.; Rahimzadeh-Bajgiran, P.; Mech, A. Estimating nutritive, non-nutritive and defense foliar traits in spruce-fir stands using remote sensing and site data. For. Ecol. Manag. 2023, 549, 121461. [Google Scholar] [CrossRef]
- Hyde, P.; Dubayah, R.; Walker, W.; Blair, J.B.; Hofton, M.; Hunsaker, C. Mapping Forest structure for wildlife habitat analysis using multi-sensor (LiDAR, SAR/InSAR, ETM+, Quickbird) synergy. Remote Sens. Environ. 2006, 102, 63–73. [Google Scholar] [CrossRef]
- Asner, G.P.; Mascaro, J.; Muller-Landau, H.C.; Vieilledent, G.; Vaudry, R.; Rasamoelina, M.; Hall, J.S.; van Breugel, M. A universal airborne LiDAR approach for tropical forest carbon mapping. Oecologia 2012, 168, 1147–1160. [Google Scholar] [CrossRef]
- Borsah, A.A.; Nazeer, M.; Wong, M.S. LIDAR-Based Forest biomass remote sensing: A review of metrics, methods, and assessment criteria for the selection of allometric equations. Forests 2023, 14, 2095. [Google Scholar] [CrossRef]
- Lefsky, M.A.; Cohen, W.B.; Parker, G.G.; Harding, D.J. Lidar remote sensing for ecosystem studies. BioScience 2002, 52, 19–30. [Google Scholar] [CrossRef]
- Chen, Q. LiDAR remote sensing of vegetation biomass. In Remote Sensing of Natural Resources, 1st ed.; Routledge: London, UK, 2013. [Google Scholar]
- Lu, D.; Chen, Q.; Wang, G.; Moran, E.; Batistella, M.; Zhang, M.; Vaglio Laurin, G.; Saah, D. Aboveground Forest biomass estimation with Landsat and LiDAR data and uncertainty analysis of the estimates. Int. J. For. Res. 2012, 2012, 436537. [Google Scholar] [CrossRef]
- Li, L.; Guo, Q.; Tao, S.; Kelly, M.; Xu, G. LiDAR with multi-temporal MODIS provides a means to upscale predictions of forest biomass. ISPRS J. Photogramm. Remote Sens. 2015, 102, 198–208. [Google Scholar] [CrossRef]
- Choate, M.J.; Rengarajan, R.; Storey, J.C.; Lubke, M. Landsat 9 Geometric Commissioning Calibration Updates and System Performance Assessment. Remote Sens. 2023, 15, 3524. [Google Scholar] [CrossRef]
- Lu, D.; Chen, Q.; Wang, G.; Liu, L.; Li, G.; Moran, E. A survey of remote sensing-based aboveground biomass estimation methods in forest ecosystems. Int. J. Digit. Earth. 2014, 9, 63–105. [Google Scholar] [CrossRef]









| Category | Variable |
|---|---|
| Sentinel-2 bands (10 and 20 m, multi-seasonal) | B2, B3, B4, B8 (Mar., Jun., Aug., Oct. [10 m]) |
| B2, B3, B4, B5, B6, B7, B8A, B11, B12 (Mar., Jun., Aug., Sep., Oct. [20 m]) | |
| Sentinel-2 derived vegetation indices | ARVI, EVI7, EVI8, MSR, SAVI, NDII11, TVI (Mar., May, Sep., Oct., Jun., Aug. [20 m]) |
| NDVI (Mar., May, Jun., Oct., Aug. [10 m and 20 m]) | |
| Landsat-9 (30-m) | B1, B2, B3, B4, B5, B6, B7, NDVI, NDAI, MSI, SVR (Sep.) |
| LiDAR-derived predictors (10 m spatial resolution via canopy metrics) | CHM, STRAT5_M, MEDMODE, MEDMAD, LSKEW, LMOM2, LMOM3, LMOM4, HVAR, HSTD, HSKEW, HQUAD, HCV, H10PCT, H50PCT, H60PCT, H70PCT, H75PCT |
| VIs | Equation |
|---|---|
| ARVI (Atmospherically Resistant Vegetation Index) | (B8A − 2B4 + B2)/(B8A + 2B4 + B2) |
| EVI7 (Enhanced Vegetation Index7) | 2.5 × (B7 − B4)/(1 + B7 + 6B4 − 7.5B2) |
| EVI8 (Enhanced Vegetation Index8) | 2.5 × (B8A − B4)/(1 + B8A + 6B4 − 7.5B2) |
| MSR (Modified Simple Ratio) | ((B7/B4) − 1)/sqrt ((B7/B4) + 1) |
| SAVI (Soil Adjusted Vegetation Index) | 1.5 × (B8A − B4)/(B8A + B4 + 0.5) |
| NDII11 (Normalized Difference Infrared Index11) | (B8A − B11)/(B8A + B11) |
| TVI (Triangular Vegetation Index) | 0.5 × {(120 × (B6 − B3)) − (200 × (B4 − B3))} |
| NDVI (Normalized Difference Vegetation Index) | (B8A − B4)/(B8A + B4) |
| VIs | Equation |
|---|---|
| NDVI (Normalized Difference Vegetation Index) | (B5 − B4)/(B5 + B4) |
| MSI (Moisture Stress Index) | (B6 − B5)/(B6 + B5) |
| NDAI (Normalized Difference Autumn Index) | (B4 − B2)/(B4 + B2) |
| SVR (Short Wave Infrared to Visible Ratio) | (B6 + B7)/(B2 + B3 + B4) |
| Predictors | Metric Description |
|---|---|
| CHM | Canopy Height Model |
| STRAT5_M | Mean height of vegetation > 5 m and ≤10 m |
| MEDMOD | Median absolute deviation from mode height |
| MEDMAD | Median absolute deviation from median height |
| LSKEW | L-moment skewness |
| LMOM4 | Fourth L-moment (kurtosis) |
| LMOM3 | Third L-moment (skewness) |
| LMOM2 | Second L-moment (scale/dispersion) |
| HVAR | Variance of heights |
| HSTD | Standard deviation of all return heights |
| HSKEW | Kurtosis of heights |
| HQUAD | Quadratic mean height |
| HCV | Coefficient of variation of heights |
| H75PCT | Average height 75th percentile |
| H70PCT | Average height 70th percentile |
| H60PCT | Average height 60th percentile |
| H50PCT | Average height 50th percentile (median) |
| H10PCT | Average height 10th percentile |
| Category | Predictors | Physical Signal |
|---|---|---|
| SWIR moisture and dry-matter chemistry (S2) | B11_Mar_20m, B12_Mar_20m, B11_Aug_20m, B12_Sep_20m, May_NDII11 | Liquid water content; cellulose/lignin; canopy/wood dryness (SWIR absorption; NDII11 water sensitivity). |
| Canopy density and red-edge/NIR structure (S2) | B6_Mar_20m, B8_May_10m, B5_May_20m, B8A_May_20m, B7_Oct_20m | Leaf/crown density; internal leaf structure; red-edge sensitivity to chlorophyll and canopy structure. |
| Greenness indices (soil/illumination-robust) (S2) | Mar_EVI7, May_SAVI, Sep_EVI8, OCT_TVI, AUG_TVI, NDVI_Jun_20m, NDVI_Oct_10m | Photosynthetic activity/pigment state; SAVI/TVI mitigate soil/illumination; NDVI seasonal amplitude. |
| SWIR-to-Visible Ratio (SVR; L9-only) | SVR_Mar_20m, SVR_Oct_20m, AUG_SVR_20m, Sep_SVR_20m | Broad VIS↔SWIR contrast: higher values track drier canopy/greater dry-matter (SWIR) relative to VIS brightness. |
| Phenology and VIS band proxies (incl. cross-sensor checks) | B3_Jun_20m, B3_AUG_10m, B2_Oct_10m (S2); B1_L9_Sep (coastal/aerosol), B4_L9_Sep (red) | Seasonal pigment dynamics (green-up/peak/senescence) and late-season VIS stability/contrast via cross-sensor bands |
| Vertical structure (LiDAR) | HQUAD | Height-distribution curvature (vertical layering) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Salehnia, N.; Wolter, P.; Sturtevant, B.R.; Abbas Iossifov, D. Advancing Forest Inventory and Fuel Monitoring with Multi-Sensor Hybrid Models: A Comparative Framework for Basal Area Estimation. Remote Sens. 2026, 18, 852. https://doi.org/10.3390/rs18060852
Salehnia N, Wolter P, Sturtevant BR, Abbas Iossifov D. Advancing Forest Inventory and Fuel Monitoring with Multi-Sensor Hybrid Models: A Comparative Framework for Basal Area Estimation. Remote Sensing. 2026; 18(6):852. https://doi.org/10.3390/rs18060852
Chicago/Turabian StyleSalehnia, Nasrin, Peter Wolter, Brian R. Sturtevant, and Dalia Abbas Iossifov. 2026. "Advancing Forest Inventory and Fuel Monitoring with Multi-Sensor Hybrid Models: A Comparative Framework for Basal Area Estimation" Remote Sensing 18, no. 6: 852. https://doi.org/10.3390/rs18060852
APA StyleSalehnia, N., Wolter, P., Sturtevant, B. R., & Abbas Iossifov, D. (2026). Advancing Forest Inventory and Fuel Monitoring with Multi-Sensor Hybrid Models: A Comparative Framework for Basal Area Estimation. Remote Sensing, 18(6), 852. https://doi.org/10.3390/rs18060852






