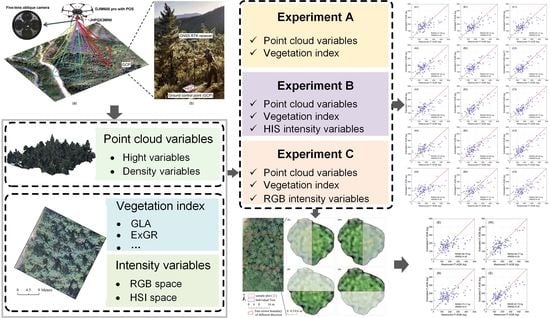
Estimating Individual Tree Above-Ground Biomass of Chinese Fir Plantation: Exploring the Combination of Multi-Dimensional Features from UAV Oblique Photos
Abstract
:1. Introduction
2. Materials and Methods
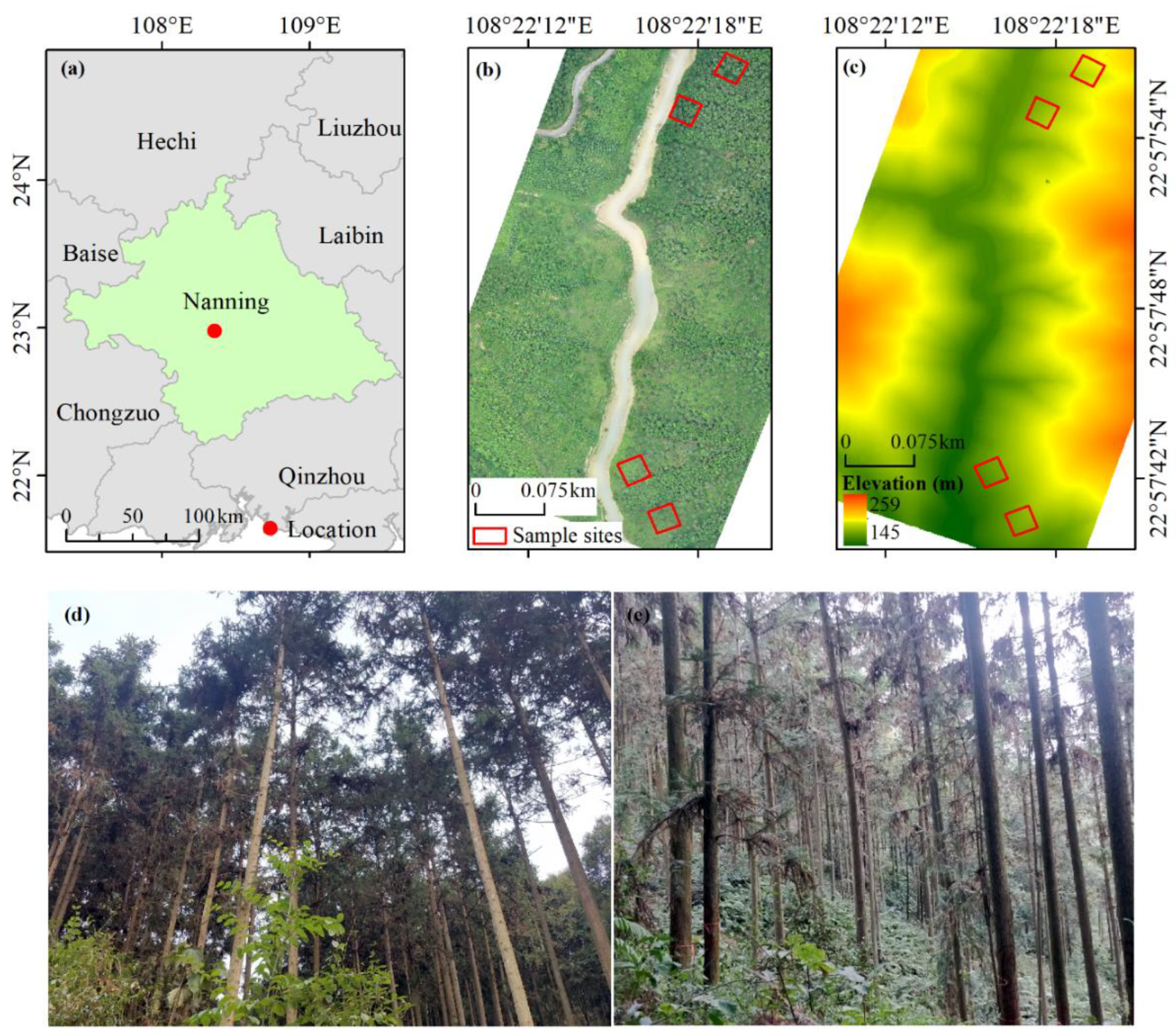
2.1. Study Area
2.2. Data
2.2.1. Field Data
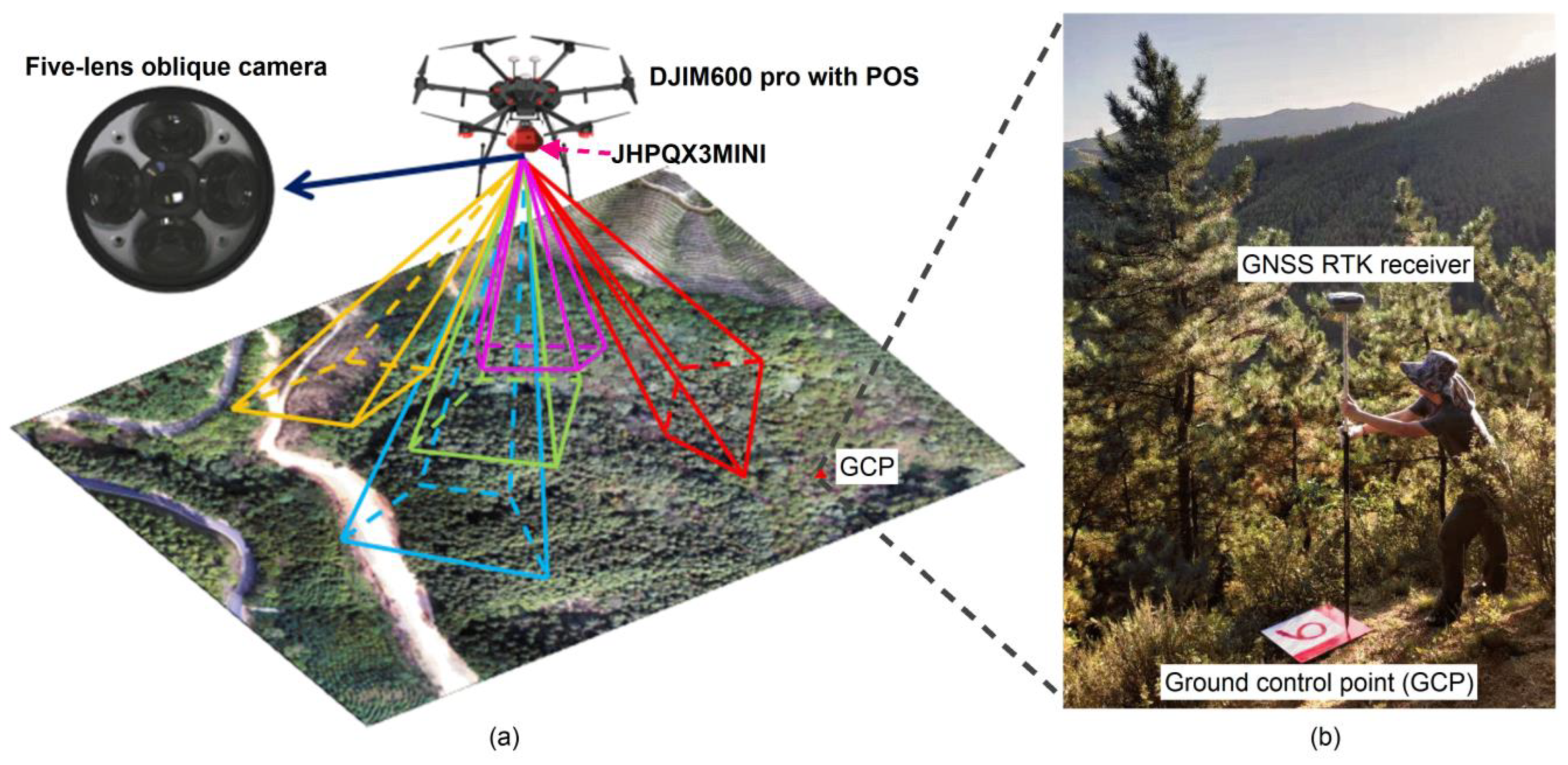
2.2.2. UAV Oblique Photography Data and Auxiliary Terrain Data
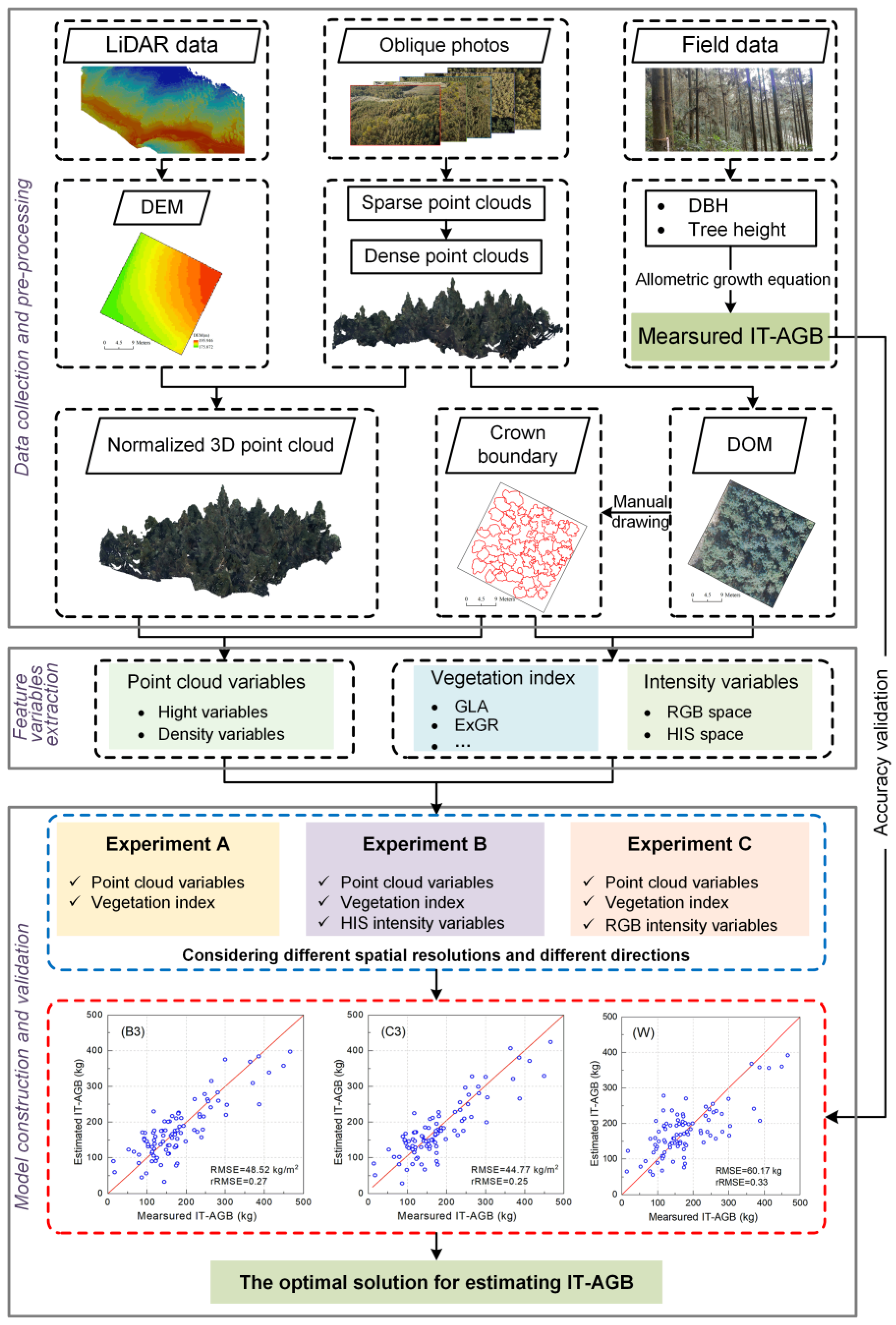
2.3. Data Processing
2.4. Methods
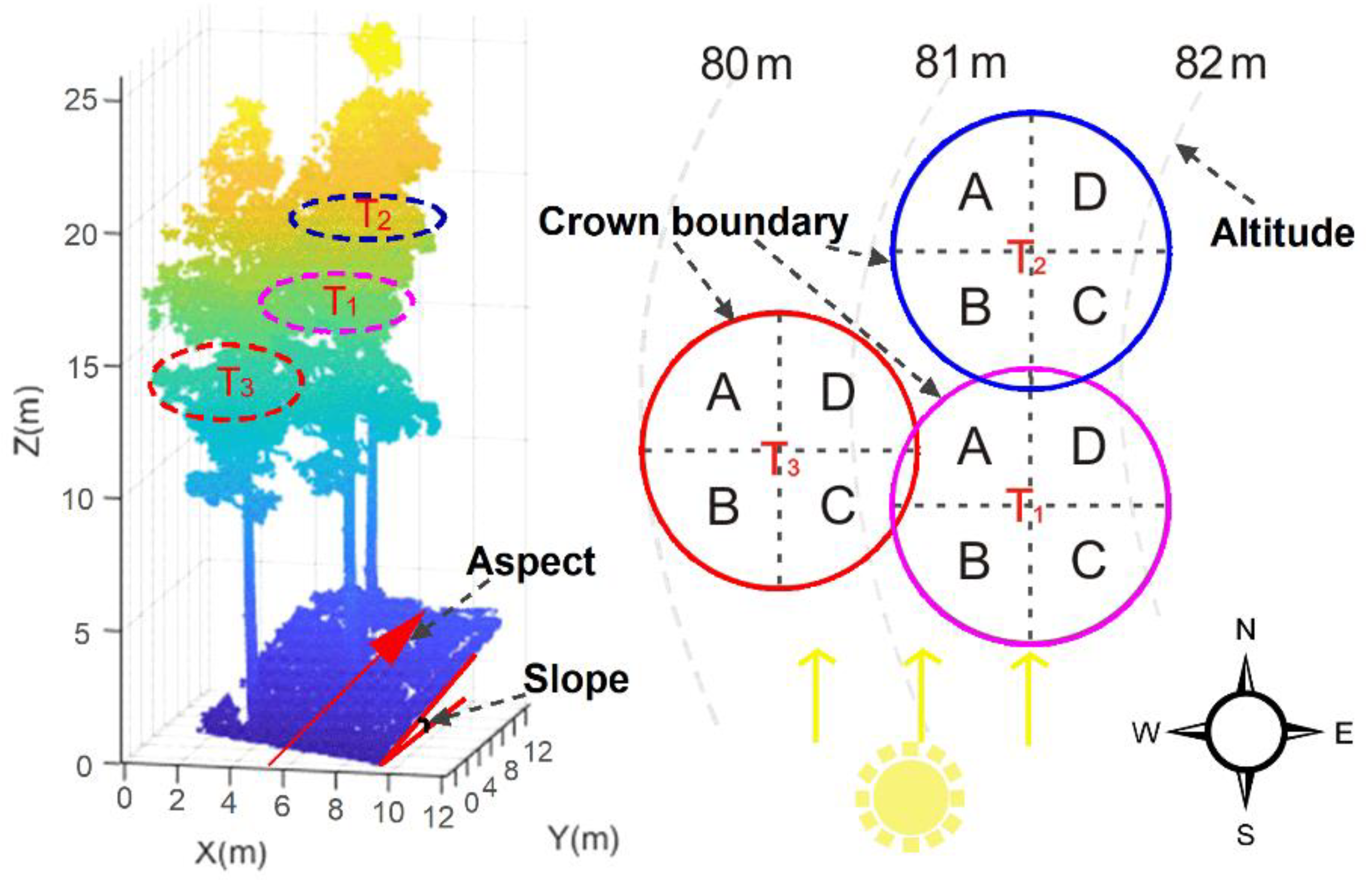
2.4.1. Individual Tree Segmentation
2.4.2. Feature Variables Extraction
2.4.3. Construction of Empirical Model
2.4.4. Accuracy Verification
3. Results
3.1. IT-AGB Distribution of Sample Plots
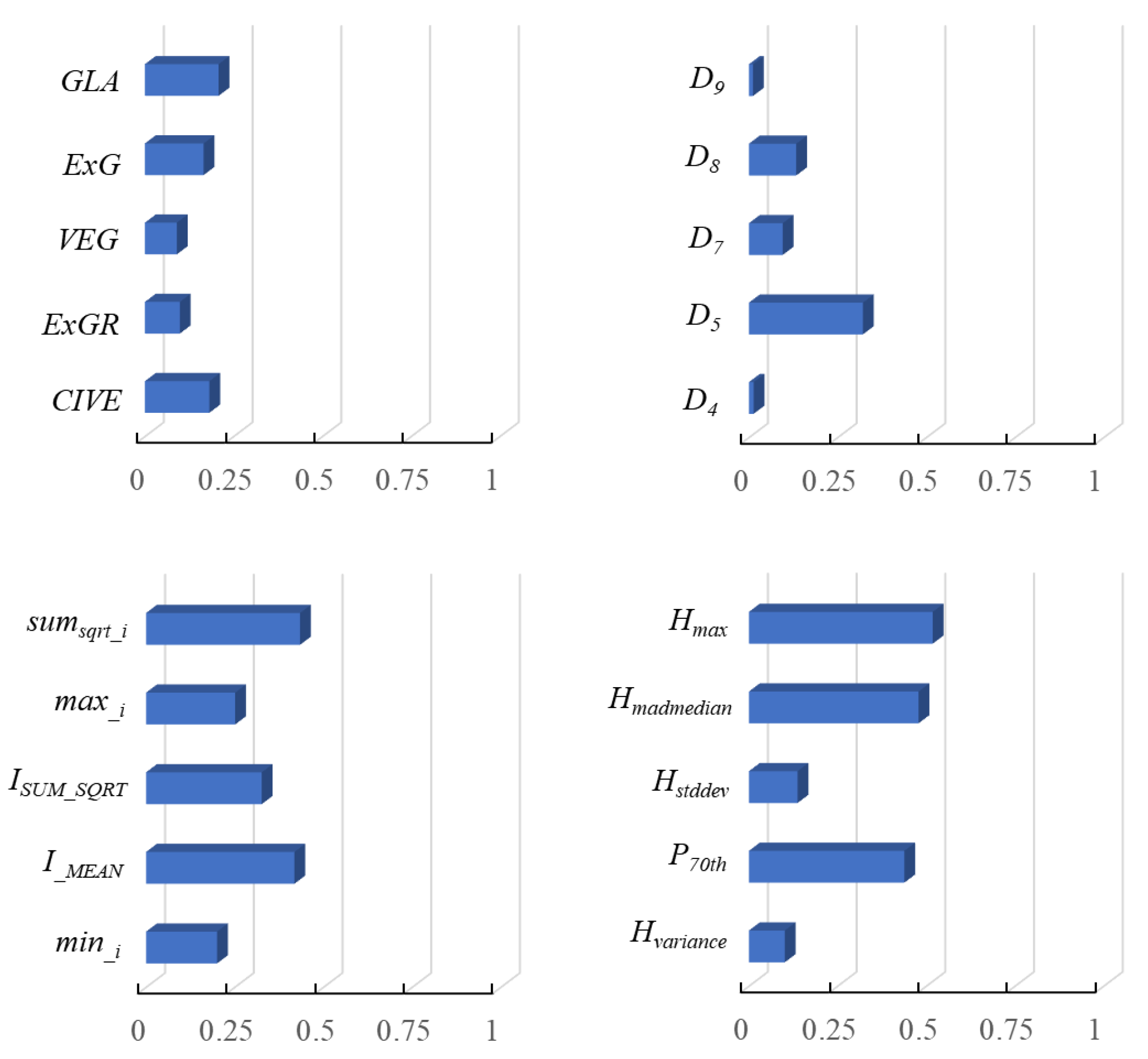
3.2. Features Selection
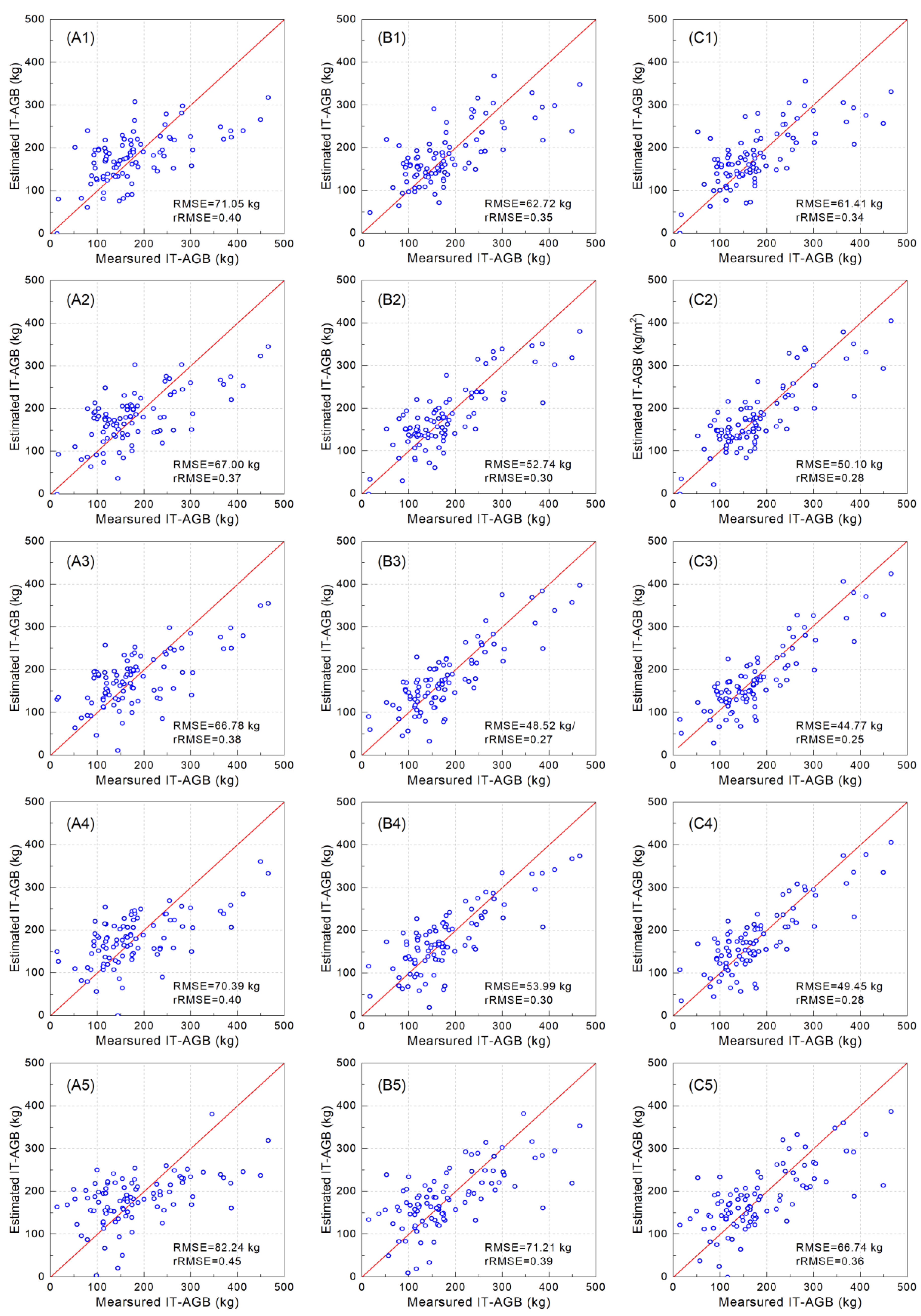
3.3. Contribution of Different Intensity Information and Spatial Resolution on Model
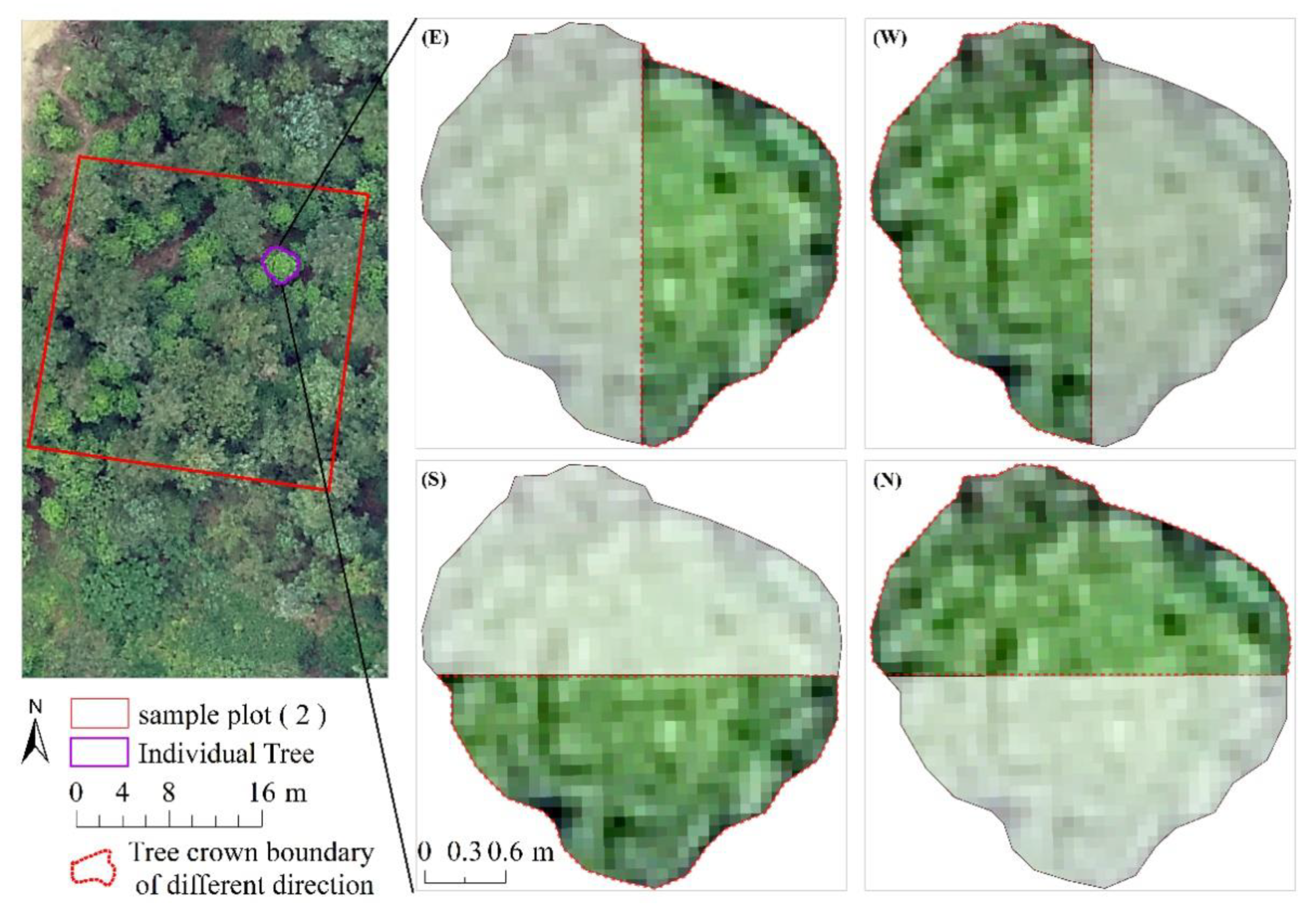
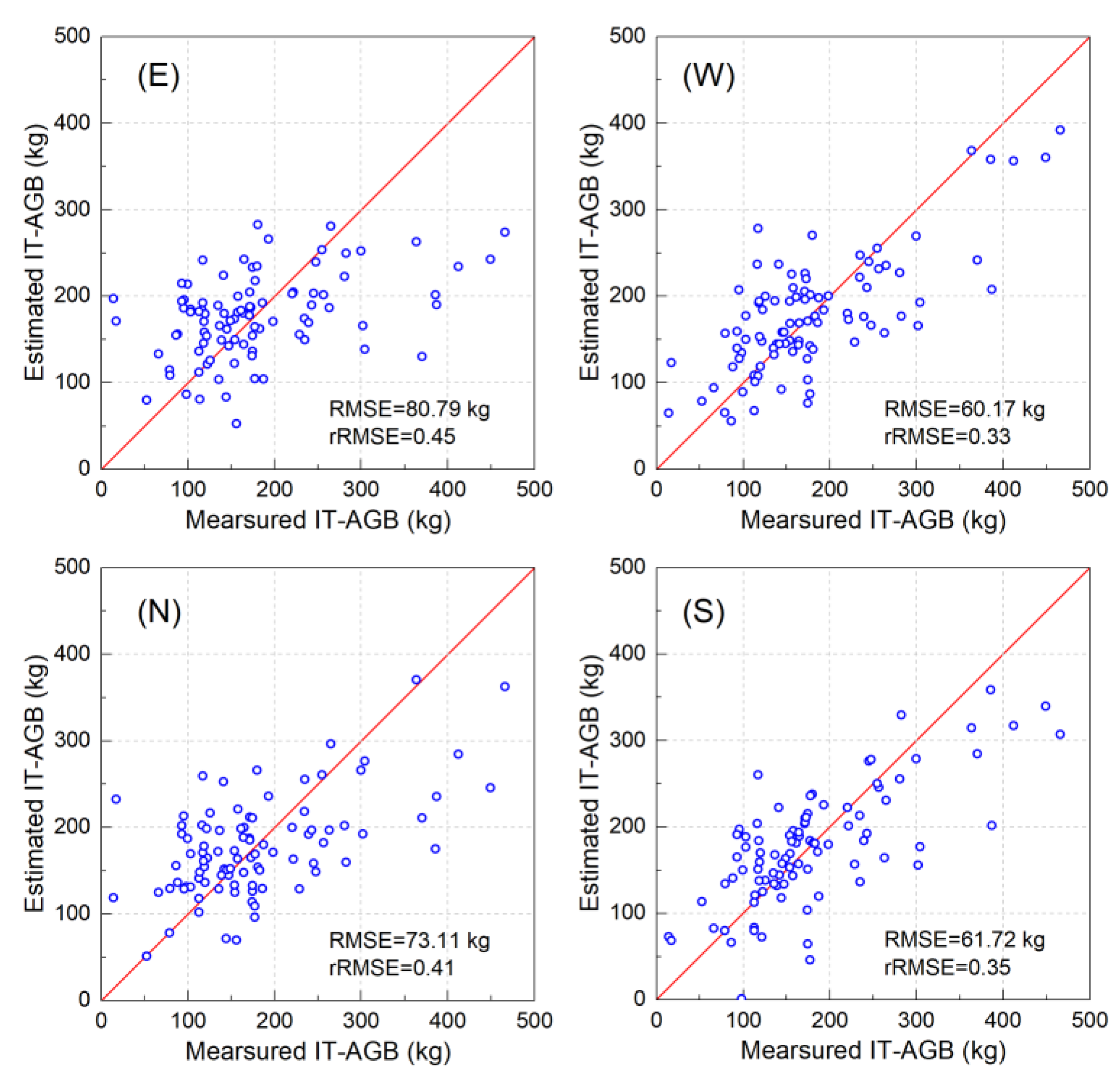
3.4. Comparisons of Models in Different Tree Canopy Directions
4. Discussion
4.1. Extraction of Feature Variables for Modeling
4.2. Effects of Different Spatial Resolution on Models
4.3. Effects of Intensity Information on Model
4.4. Effects of Different Direction on Model
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gómez, C.; White, J.C.; Wulder, M.A.; Alejandro, P. Historical forest biomass dynamics modelled with Landsat spectral trajectories. ISPRS J. Photogramm. Remote Sens. 2014, 93, 14–28. [Google Scholar] [CrossRef] [Green Version]
- Zhu, X.; Liu, D. Improving forest aboveground biomass estimation using seasonal Landsat NDVI time-series. ISPRS J. Photogramm. Remote Sens. 2015, 102, 222–231. [Google Scholar] [CrossRef]
- Li, L.; Guo, Q.; Tao, S.; Kelly, M.; Xu, G. Lidar with multi-temporal MODIS provide a means to upscale predictions of forest biomass. ISPRS J. Photogramm. Remote Sens. 2015, 102, 198–208. [Google Scholar] [CrossRef]
- Luo, S.; Wang, C.; Xi, X.; Nie, S.; Fan, X.; Chen, H.; Ma, D.; Liu, J.; Zou, J.; Lin, Y.; et al. Estimating forest aboveground biomass using small-footprint full-waveform airborne LiDAR data. Int. J. Appl. Earth Obs. Geoinf. 2019, 83, 101922. [Google Scholar] [CrossRef]
- Malhi, R.K.M.; Anand, A.; Srivastava, P.K.; Chaudhary, S.K.; Pandey, M.K.; Behera, M.D.; Kumar, A.; Singh, P.; Sandhya Kiran, G. Synergistic evaluation of Sentinel 1 and 2 for biomass estimation in a tropical forest of India. Adv. Space Res. 2021, in press. [Google Scholar] [CrossRef]
- Campbell, M.J.; Dennison, P.E.; Kerr, K.L.; Brewer, S.C.; Anderegg, W.R.L. Scaled biomass estimation in woodland ecosystems: Testing the individual and combined capacities of satellite multispectral and lidar data. Remote Sens. Environ. 2021, 262, 112511. [Google Scholar] [CrossRef]
- Wang, G.; Zhang, M.; Gertner, G.Z.; Oyana, T.; McRoberts, R.E.; Ge, H. Uncertainties of mapping aboveground forest carbon due to plot locations using national forest inventory plot and remotely sensed data. Scand. J. For. Res. 2011, 26, 360–373. [Google Scholar] [CrossRef]
- Wang, Y.; Ni, W.; Sun, G.; Chi, H.; Zhang, Z.; Guo, Z. Slope-adaptive waveform metrics of large footprint lidar for estimation of forest aboveground biomass. Remote Sens. Environ. 2019, 224, 386–400. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, X.; Guo, Z. Estimation of tree height and aboveground biomass of coniferous forests in North China using stereo ZY-3, multispectral Sentinel-2, and DEM data. Ecol. Indic. 2021, 126, 107645. [Google Scholar] [CrossRef]
- Liao, Z.; He, B.; Quan, X.; van Dijk, A.I.J.M.; Qiu, S.; Yin, C. Biomass estimation in dense tropical forest using multiple information from single-baseline P-band PolInSAR data. Remote Sens. Environ. 2019, 221, 489–507. [Google Scholar] [CrossRef]
- Ploton, P.; Barbier, N.; Couteron, P.; Antin, C.M.; Ayyappan, N.; Balachandran, N.; Barathan, N.; Bastin, J.F.; Chuyong, G.; Dauby, G.; et al. Toward a general tropical forest biomass prediction model from very high resolution optical satellite images. Remote Sens. Environ. 2017, 200, 140–153. [Google Scholar] [CrossRef]
- Askne, J.I.H.; Soja, M.J.; Ulander, L.M.H. Biomass estimation in a boreal forest from TanDEM-X data, lidar DTM, and the interferometric water cloud model. Remote Sens. Environ. 2017, 196, 265–278. [Google Scholar] [CrossRef]
- Yang, H.; Li, F.; Wang, W.; Yu, K. Estimating Above-Ground Biomass of Potato Using Random Forest and Optimized Hyperspectral Indices. Remote Sens. 2021, 13, 2339. [Google Scholar] [CrossRef]
- Proisy, C.; Couteron, P.; Fromard, F.O. Predicting and mapping mangrove biomass from canopy grain analysis using Fourier-based textural ordination (FOTO) of IKONOS images. Remote Sens. Environ. 2007, 109, 379–392. [Google Scholar] [CrossRef]
- Yadav, S.; Padalia, H.; Sinha, S.K.; Srinet, R.; Chauhan, P. Above-ground biomass estimation of Indian tropical forests using X band Pol-InSAR and Random Forest. Remote Sens. Appl. Soc. Environ. 2021, 21, 100462. [Google Scholar] [CrossRef]
- Li, G.; Xie, Z.; Jiang, X.; Lu, D.; Chen, E. Integration of ZiYuan-3 Multispectral and Stereo Data for Modeling Aboveground Biomass of Larch Plantations in North China. Remote Sens. 2019, 11, 2328. [Google Scholar] [CrossRef] [Green Version]
- Zhao, P.; Lu, D.; Wang, G.; Wu, C.; Huang, Y.; Yu, S. Examining Spectral Reflectance Saturation in Landsat Imagery and Corresponding Solutions to Improve Forest Aboveground Biomass Estimation. Remote Sens. 2016, 8, 469. [Google Scholar] [CrossRef] [Green Version]
- Hu, T.; Zhang, Y.; Su, Y.; Zheng, Y.; Lin, G.; Guo, Q. Mapping the Global Mangrove Forest Aboveground Biomass Using Multisource Remote Sensing Data. Remote Sens. 2020, 12, 1690. [Google Scholar] [CrossRef]
- Cao, L.; Coops, N.C.; Innes, J.L.; Sheppard, S.R.J.; Fu, L.; Ruan, H.; She, G. Estimation of forest biomass dynamics in subtropical forests using multi-temporal airborne LiDAR data. Remote Sens. Environ. 2016, 178, 158–171. [Google Scholar] [CrossRef]
- Bazezew, M.N.; Hussin, Y.A.; Kloosterman, E.H. Integrating Airborne LiDAR and Terrestrial Laser Scanner forest parameters for accurate above-ground biomass/carbon estimation in Ayer Hitam tropical forest, Malaysia. Int. J. Appl. Earth Obs. Geoinf. 2018, 73, 638–652. [Google Scholar] [CrossRef]
- Duncanson, L.I.; Dubayah, R.O.; Cook, B.D.; Rosette, J.; Parker, G. The importance of spatial detail: Assessing the utility of individual crown information and scaling approaches for lidar-based biomass density estimation. Remote Sens. Environ. 2015, 168, 102–112. [Google Scholar] [CrossRef]
- Lu, J.; Wang, H.; Qin, S.; Cao, L.; Pu, R.; Li, G.; Sun, J. Estimation of aboveground biomass of Robinia pseudoacacia forest in the Yellow River Delta based on UAV and Backpack LiDAR point clouds. Int. J. Appl. Earth Obs. Geoinf. 2020, 86, 102014. [Google Scholar] [CrossRef]
- Nord-Larsen, T.; Schumacher, J. Estimation of forest resources from a country wide laser scanning survey and national forest inventory data. Remote Sens. Environ. 2012, 119, 148–157. [Google Scholar] [CrossRef]
- Eitel, J.U.H.; Höfle, B.; Vierling, L.A.; Abellán, A.; Asner, G.P.; Deems, J.S.; Glennie, C.L.; Joerg, P.C.; LeWinter, A.L.; Magney, T.S.; et al. Beyond 3-D: The new spectrum of lidar applications for earth and ecological sciences. Remote Sens. Environ. 2016, 186, 372–392. [Google Scholar] [CrossRef] [Green Version]
- Su, Y.; Guo, Q.; Xue, B.; Hu, T.; Alvarez, O.; Tao, S.; Fang, J. Spatial distribution of forest aboveground biomass in China: Estimation through combination of spaceborne lidar, optical imagery, and forest inventory data. Remote Sens. Environ. 2016, 173, 187–199. [Google Scholar] [CrossRef] [Green Version]
- Liu, K.; Shen, X.; Cao, L.; Wang, G.; Cao, F. Estimating forest structural attributes using UAV-LiDAR data in Ginkgo plantations. ISPRS J. Photogramm. Remote Sens. 2018, 146, 465–482. [Google Scholar] [CrossRef]
- Jayathunga, S.; Owari, T.; Tsuyuki, S. The use of fixed–wing UAV photogrammetry with LiDAR DTM to estimate merchantable volume and carbon stock in living biomass over a mixed conifer–broadleaf forest. Int. J. Appl. Earth Obs. Geoinf. 2018, 73, 767–777. [Google Scholar] [CrossRef]
- Li, B.; Xu, X.; Zhang, L.; Han, J.; Bian, C.; Li, G.; Liu, J.; Jin, L. Above-ground biomass estimation and yield prediction in potato by using UAV-based RGB and hyperspectral imaging. ISPRS J. Photogramm. Remote Sens. 2020, 162, 161–172. [Google Scholar] [CrossRef]
- Fu, X.; Zhang, Z.; Cao, L.; Coops, N.C.; Goodbody, T.R.H.; Liu, H.; Shen, X.; Wu, X. Assessment of approaches for monitoring forest structure dynamics using bi-temporal digital aerial photogrammetry point clouds. Remote Sens. Environ. 2021, 255, 112300. [Google Scholar] [CrossRef]
- Dandois, J.P.; Ellis, E.C. Remote Sensing of Vegetation Structure Using Computer Vision. Remote Sens. 2010, 2, 1157–1176. [Google Scholar] [CrossRef] [Green Version]
- Goodbody, T.R.H.; Coops, N.C.; White, J.C. Digital Aerial Photogrammetry for Updating Area-Based Forest Inventories: A Review of Opportunities, Challenges, and Future Directions. Curr. For. Rep. 2019, 5, 55–75. [Google Scholar] [CrossRef] [Green Version]
- Pulkkinen, M.; Ginzler, C.; Traub, B.; Lanz, A. Stereo-imagery-based post-stratification by regression-tree modelling in Swiss National Forest Inventory. Remote Sens. Environ. 2018, 213, 182–194. [Google Scholar] [CrossRef]
- Watts, A.C.; Ambrosia, V.G.; Hinkley, E.A. Unmanned Aircraft Systems in Remote Sensing and Scientific Research: Classification and Considerations of Use. Remote Sens. 2012, 4, 1671–1692. [Google Scholar] [CrossRef] [Green Version]
- Puliti, S.; Ene, L.T.; Gobakken, T.; Næsset, E. Use of partial-coverage UAV data in sampling for large scale forest inventories. Remote Sens. Environ. 2017, 194, 115–126. [Google Scholar] [CrossRef]
- Dandois, J.P.; Ellis, E.C. High spatial resolution three-dimensional mapping of vegetation spectral dynamics using computer vision. Remote Sens. Environ. 2013, 136, 259–276. [Google Scholar] [CrossRef] [Green Version]
- Cunliffe, A.M.; Brazier, R.E.; Anderson, K. Ultra-fine grain landscape-scale quantification of dryland vegetation structure with drone-acquired structure-from-motion photogrammetry. Remote Sens. Environ. 2016, 183, 129–143. [Google Scholar] [CrossRef] [Green Version]
- Price, B.; Waser, L.T.; Wang, Z.; Marty, M.; Ginzler, C.; Zellweger, F. Predicting biomass dynamics at the national extent from digital aerial photogrammetry. Int. J. Appl. Earth Obs. Geoinf. 2020, 90, 102116. [Google Scholar] [CrossRef]
- Alonzo, M.; Dial, R.J.; Schulz, B.K.; Andersen, H.-E.; Lewis-Clark, E.; Cook, B.D.; Morton, D.C. Mapping tall shrub biomass in Alaska at landscape scale using structure-from-motion photogrammetry and lidar. Remote Sens. Environ. 2020, 245, 111841. [Google Scholar] [CrossRef]
- Luo, Y.; Lu, F. Comprehensive Database of Biomes Regressions for China’s Tree Species; China Forestry Publishing House: Beijing, China, 2015; p. 125. [Google Scholar]
- Michez, A.; Piegay, H.; Lisein, J.; Claessens, H.; Lejeune, P. Classification of riparian forest species and health condition using multi-temporal and hyperspatial imagery from unmanned aerial system. Environ. Monit. Assess 2016, 188, 146. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jing, R.; Gong, Z.; Zhao, W.; Pu, R.; Deng, L. Above-bottom biomass retrieval of aquatic plants with regression models and SfM data acquired by a UAV platform—A case study in Wild Duck Lake Wetland, Beijing, China. ISPRS J. Photogramm. Remote Sens. 2017, 134, 122–134. [Google Scholar] [CrossRef]
- Giannetti, F.; Chirici, G.; Gobakken, T.; Næsset, E.; Travaglini, D.; Puliti, S. A new approach with DTM-independent metrics for forest growing stock prediction using UAV photogrammetric data. Remote Sens. Environ. 2018, 213, 195–205. [Google Scholar] [CrossRef]
- Navarro, A.; Young, M.; Allan, B.; Carnell, P.; Macreadie, P.; Ierodiaconou, D. The application of Unmanned Aerial Vehicles (UAVs) to estimate above-ground biomass of mangrove ecosystems. Remote Sens. Environ. 2020, 242, 111747. [Google Scholar] [CrossRef]
- Puliti, S.; Ørka, H.; Gobakken, T.; Næsset, E. Inventory of Small Forest Areas Using an Unmanned Aerial System. Remote Sens. 2015, 7, 9632–9654. [Google Scholar] [CrossRef] [Green Version]
- Dalponte, M.; Ørka, H.O.; Ene, L.T.; Gobakken, T.; Næsset, E. Tree crown delineation and tree species classification in boreal forests using hyperspectral and ALS data. Remote Sens. Environ. 2014, 140, 306–317. [Google Scholar] [CrossRef]
- Hyyppa, J.; Kelle, M.; Lehikoinen, M.; Inkinen, M. A segmentation-based method to retrieve stem volume estimates from 3-D tree height models produced by laser scanners. IEEE Trans. Geosci. Remote Sens. 2001, 39, 969–975. [Google Scholar] [CrossRef]
- Lee, H.; Slatton, K.C.; Roth, B.E.; Cropper, W.P. Adaptive clustering of airborne LiDAR data to segment individual tree crowns in managed pine forests. Int. J. Remote Sens. 2010, 31, 117–139. [Google Scholar] [CrossRef]
- Vastaranta, M. Individual tree detection and area-based approach in retrieval of forest inventory characteristics from low-pulse airborne laser scanning data. Photogramm. J. Finl. 2011, 22, 1311–1317. [Google Scholar]
- Hu, B.; Li, J.; Jing, L.; Judah, A. Improving the efficiency and accuracy of individual tree crown delineation from high-density LiDAR data. Int. J. Appl. Earth Obs. Geoinf. 2014, 26, 145–155. [Google Scholar] [CrossRef]
- Mongus, D.; Žalik, B. An efficient approach to 3D single tree-crown delineation in LiDAR data. ISPRS J. Photogramm. Remote Sens. 2015, 108, 219–233. [Google Scholar] [CrossRef]
- Hu, X.; Chen, W.; Xu, W. Adaptive Mean Shift-Based Identification of Individual Trees Using Airborne LiDAR Data. Remote Sens. 2017, 9, 148. [Google Scholar] [CrossRef] [Green Version]
- Dai, W.; Yang, B.; Dong, Z.; Shaker, A. A new method for 3D individual tree extraction using multispectral airborne LiDAR point clouds. ISPRS J. Photogramm. Remote Sens. 2018, 144, 400–411. [Google Scholar] [CrossRef]
- Wu, Y.; Zhao, L.; Jiang, H.; Guo, X.; Huang, F. Image segmentation method for green crops using improved mean shift. Trans. Chin. Soc. Agric. Eng. 2014, 30, 161–167. [Google Scholar]
- Jing, R.; Deng, L.; Zhao, W.J.; Gong, Z.N. Object-oriented aquatic vegetation extracting approach based on visible vegetation indices. Chin. J. Appl. Ecol. 2016, 27, 1427–1436. [Google Scholar]
- Sun, G.; Wang, X.; Yan, T.; Li, X.; Chen, J. Inversion method of flora growth parameters based on machine vision. Trans. Chin. Soc. Agric. Eng. 2014, 30, 187–195. [Google Scholar]
- Gitelson, A.A.; Kaufman, Y.J.; Stark, R.; Rundquist, D. Novel algorithms for remote estimation of vegetation fraction. Remote Sens. Environ. 2002, 80, 76–87. [Google Scholar] [CrossRef] [Green Version]
- Hague, T.; Tillett, N.D.; Wheeler, H. Automated Crop and Weed Monitoring in Widely Spaced Cereals. Precis. Agric. 2006, 7, 21–32. [Google Scholar] [CrossRef]
- Montalvo, M.; Guerrero, J.M.; Romeo, J.; Emmi, L.; Guijarro, M.; Pajares, G. Automatic expert system for weeds/crops identification in images from maize fields. Expert Syst. Appl. 2013, 40, 75–82. [Google Scholar] [CrossRef]
- Wu, Y.; Zhang, H.; Lu, T.; Xu, H.; Ou, G. Aboveground biomass estimation of coniferous forests using Landsat 8 OLI and the determination of saturation points in the alpine and sub-alpine area of Northwest Yunnan. J. Yunnan Univ. Nat. Sci. Ed. 2021, 43, 818–830. [Google Scholar] [CrossRef]
- Knapp, N.; Fischer, R.; Huth, A. Linking lidar and forest modeling to assess biomass estimation across scales and disturbance states. Remote Sens. Environ. 2018, 205, 199–209. [Google Scholar] [CrossRef]
- Zhang, G.; Ganguly, S.; Nemani, R.R.; White, M.A.; Milesi, C.; Hashimoto, H.; Wang, W.; Saatchi, S.; Yu, Y.; Myneni, R.B. Estimation of forest aboveground biomass in California using canopy height and leaf area index estimated from satellite data. Remote Sens. Environ. 2014, 151, 44–56. [Google Scholar] [CrossRef]
- Gao, Y.; Liang, Z.; Wang, B.; Wu, Y.; Shiyu, L. UAV and satellite remote sensing images based aboveground biomass inversion in the meadows of Lake Shengjin. J. Lake Sci. 2019, 31, 517–528. [Google Scholar]
- Yan, E.; Lin, H.; Wang, G.; Chen, Z. Estimation of Hunan forest carbon density based on spectral mixture analysis of MODIS data. J. Appl. Ecol. 2015, 26, 3433–3442. [Google Scholar] [CrossRef]
- Kamiyama, M.; Taguchi, A. Color Conversion Formula with Saturation Correction from HSI Color Space to RGB Color Space. IEICE Trans. Fundam. Electron. Commun. Comput. Sci. 2021, E104, 1000–1005. [Google Scholar] [CrossRef]
- Hunt, R.W.G. The Reproduction of Colour; John Wiley & Sons: Hoboken, NJ, USA, 2004. [Google Scholar]



| Sensors and Flight Parameters | JHP QX3MINI |
|---|---|
| Sensor size (mm × mm) | 23.5 × 15.6 |
| Heading overlap (%) | 80 |
| Side overlap (%) | 70 |
| Horizontal speed (m·s−1) | 4~8 |
| Flight altitude (m) | 100 |
| Exposure interval (s) | 0.8~4.5 |
| Focal length (mm) | 35 |
| Scanning angle (°) | 45 |
| Single camera pixel numbers (million) | 42 |
| Sensors and Flight Parameters | Parameter Values |
|---|---|
| Wavelength (nm) | 1550 |
| Beam divergence angle (mrad) | 0.5 |
| Spot diameter (cm) | 45 |
| Pulse repetition rate (kHz) | 360 |
| Pulse emission frequency (Hz) | 112 |
| Flight altitude (m) | 900 |
| Flight speed (m/s) | 55 |
| Type | Variable | Formula | Describe |
|---|---|---|---|
| RGB space | —— | Maximum intensity | |
| —— | Minimum intensity | ||
| —— | Mean intensity | ||
| Total intensity | |||
| Square root of intensity value | |||
| HSI space | —— | Maximum intensity | |
| —— | Minimum intensity | ||
| —— | Mean intensity | ||
| —— | Total intensity | ||
| —— | Square root of intensity value | ||
| Hight variables | Haad | Mean absolute deviation, is the height of the ith point in each unit, is the average height of all points, n is the total number of points in each statistical unit. | |
| HAIQ | —— | Cumulative height percentile interquartile spacing | |
| Hkurtosis | Height kurtosis | ||
| Hcv | Variation coefficient of Z value of all points in a statistical unit, Zstd and Zmean are the standard deviation of the height values of all points and the average height of all points in each statistical unit. | ||
| Hredio | Canopy fluctuation rate, Zmin, Zmax, Zmean are the minimum height, maximum height and average height of all points in each statistical unit | ||
| Hstddev | Standard deviation of Z values of all points in a statistical unit | ||
| Hvariance | Variance of Z values of all points in a statistical unit | ||
| Hskewness | Height skewness | ||
| H1…99th | —— | 1…99% cumulative height percentile | |
| Hmax, min, mean, median | —— | The maximum, minimum, average and median of the point cloud after normalization | |
| P1st,…,99th | —— | 75%, 95% high percentile | |
| density variables | D0,…,9 | —— | The proportion of height points greater than 30%, 50%, 70%, and 90% to all points |
| Vegetation index | CIVE | Color index of vegetation [53] | |
| ExG | Excess green index [54] | ||
| ExGR | Excess green minus excess red index [55] | ||
| GLA | Green leaf algorithm [56] | ||
| NGRDI | Normalized green–red difference index [56] | ||
| VEG | Vegetation index [57] | ||
| COM | Combination index [58] |
| Sample Number | Number of Trees | Sample Characteristics | DBH (cm) | Tree Height (m) | AGB (kg) |
|---|---|---|---|---|---|
| 1 | 51 | Maximum | 36.90 | 24.50 | 448.85 |
| minimum | 10.10 | 9.20 | 21.85 | ||
| Mean | 20.90 | 18.53 | 153.23 | ||
| Standard deviation | 5.10 | 2.68 | 81.18 | ||
| 2 | 40 | Maximum | 31.50 | 23.60 | 376.41 |
| minimum | 12.50 | 15.80 | 51.88 | ||
| Mean | 21.72 | 20.16 | 175.35 | ||
| Standard deviation | 5.11 | 2.12 | 84.21 | ||
| 3 | 28 | Maximum | 33.90 | 25.80 | 386.73 |
| minimum | 18.00 | 14.10 | 98.07 | ||
| Mean | 25.75 | 19.84 | 227.85 | ||
| Standard deviation | 4.11 | 3.25 | 79.23 | ||
| 4 | 30 | Maximum | 34.80 | 26.10 | 465.611 |
| minimum | 5.90 | 4.50 | 19.94 | ||
| Mean | 18.55 | 14.19 | 153.89 | ||
| Standard deviation | 8.59 | 6.11 | 125.87 | ||
| Total | 149 | Maximum | 36.90 | 26.10 | 465.61 |
| minimum | 5.90 | 4.50 | 19.94 | ||
| Mean | 21.53 | 19.39 | 170.51 | ||
| Standard deviation | 6.04 | 3.97 | 92.73 |
| Model | Model Equation | R2 | RMSECV (kg) | rRMSECV | |
|---|---|---|---|---|---|
| 0.05 m | A | 0.44 | 71.05 | 0.40 | |
| B | 0.57 | 62.72 | 0.35 | ||
| C | 0.59 | 61.41 | 0.34 | ||
| 0.1 m | A | 0.49 | 67.00 | 0.37 | |
| B | 0.70 ** | 52.74 | 0.30 | ||
| C | 0.72 ** | 50.10 | 0.28 | ||
| 0.2 m | A | 0.50 | 66.78 | 0.38 | |
| B | 0.75 ** | 48.52 | 0.27 | ||
| C | 0.79 ** | 44.77 | 0.25 | ||
| 0.5 m | A | 0.44 | 70.39 | 0.40 | |
| B | 0.69 ** | 53.99 | 0.30 | ||
| C | 0.74 ** | 49.45 | 0.28 | ||
| 1 m | A | 0.34 | 82.24 | 0.45 | |
| B | 0.52 | 71.21 | 0.39 | ||
| C | 0.58 | 66.74 | 0.36 | ||
| Model | Model Equation | R2 | RMSECV (kg) | rRMSECV |
|---|---|---|---|---|
| East | 0.29 | 80.79 | 0.45 | |
| North | 0.42 | 73.11 | 0.41 | |
| South | 0.58 ** | 61.72 | 0.35 | |
| West | 0.60 ** | 60.17 | 0.33 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lei, L.; Chai, G.; Wang, Y.; Jia, X.; Yin, T.; Zhang, X. Estimating Individual Tree Above-Ground Biomass of Chinese Fir Plantation: Exploring the Combination of Multi-Dimensional Features from UAV Oblique Photos. Remote Sens. 2022, 14, 504. https://doi.org/10.3390/rs14030504
Lei L, Chai G, Wang Y, Jia X, Yin T, Zhang X. Estimating Individual Tree Above-Ground Biomass of Chinese Fir Plantation: Exploring the Combination of Multi-Dimensional Features from UAV Oblique Photos. Remote Sensing. 2022; 14(3):504. https://doi.org/10.3390/rs14030504
Chicago/Turabian StyleLei, Lingting, Guoqi Chai, Yueting Wang, Xiang Jia, Tian Yin, and Xiaoli Zhang. 2022. "Estimating Individual Tree Above-Ground Biomass of Chinese Fir Plantation: Exploring the Combination of Multi-Dimensional Features from UAV Oblique Photos" Remote Sensing 14, no. 3: 504. https://doi.org/10.3390/rs14030504
APA StyleLei, L., Chai, G., Wang, Y., Jia, X., Yin, T., & Zhang, X. (2022). Estimating Individual Tree Above-Ground Biomass of Chinese Fir Plantation: Exploring the Combination of Multi-Dimensional Features from UAV Oblique Photos. Remote Sensing, 14(3), 504. https://doi.org/10.3390/rs14030504