A Survey of Active Learning for Quantifying Vegetation Traits from Terrestrial Earth Observation Data
Abstract
1. Introduction
- Provide an up-to-date overview of the use of AL heuristics in the framework of biophysical and biochemical vegetation traits retrieval from terrestrial EO data;
- Identify optimal AL strategies to obtain efficient training datasets for kernel-based ML regression algorithms to be implemented in hybrid retrieval workflows;
- Give recommendations and inspirations for further research under the AL perspective and in the context of terrestrial vegetation monitoring from EO data.
2. Background: Active Learning Theory
- The sample must be (nearly) as informative as the full dataset, implying that a learning algorithm can extract the same essential information from the sample as it would from the full dataset [35];
- The sample should be as small as possible in order to reduce the computational load.
2.1. Active Learning for Classification
2.2. Active Learning for Emulation
- The maximization of the predictive variance of the emulation/regression model [62];
- A-optimality;
- D-optimality;
- E-optimality.
2.3. Active Learning for Regression
2.3.1. Uncertainty Criteria Methods
2.3.2. Diversity Criteria Methods
3. Literature Survey: Active Learning under the Earth Observation Perspective
3.1. Systematic Approach
3.2. Sensors and Estimated Variables
3.3. Applied ML Algorithms and AL Heuristics
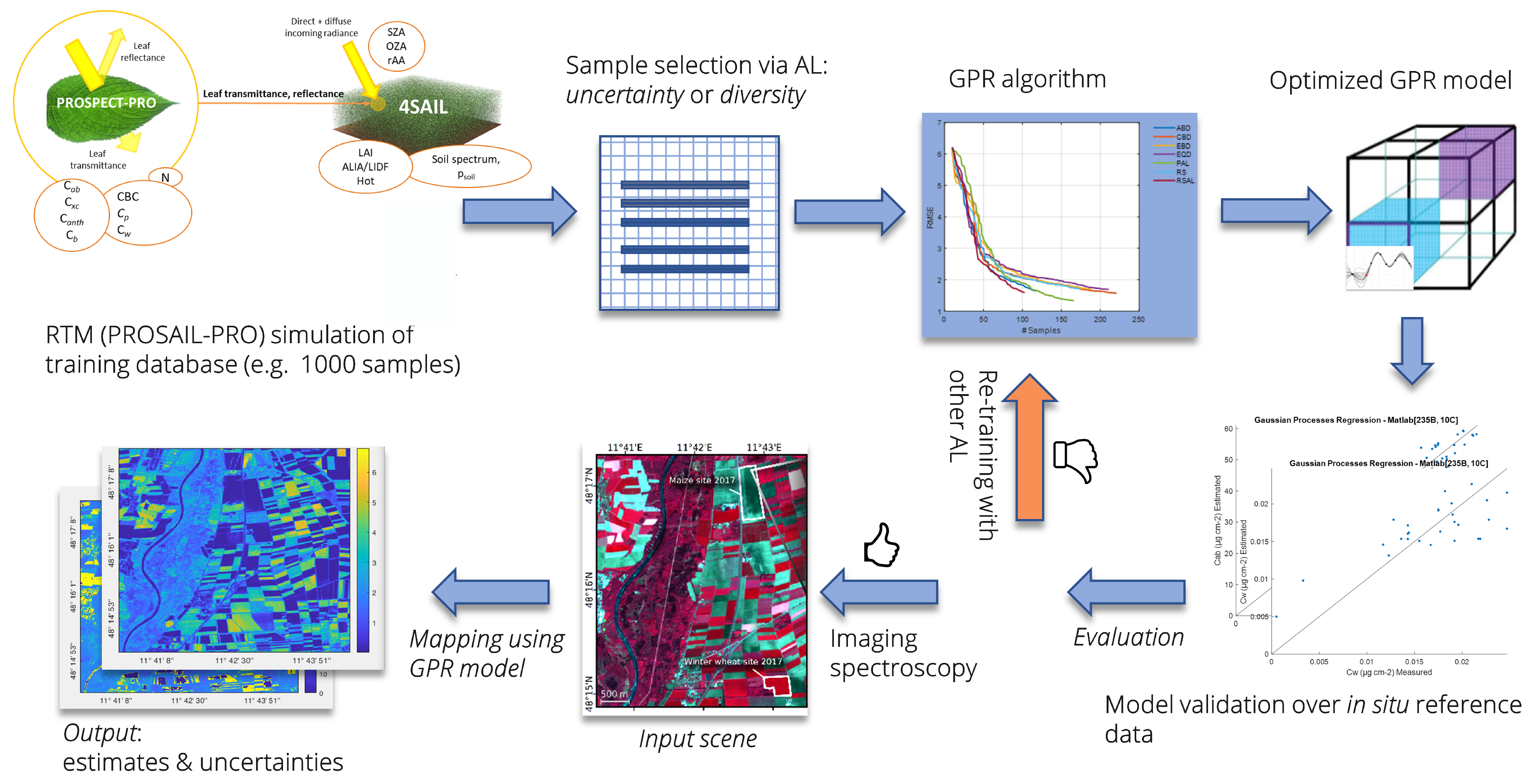
4. Experimental Case Study
4.1. Data Collection
4.2. Experimental Design
4.3. Evaluation
5. Towards Advanced Use of Active Learning for Vegetation Properties Retrieval
5.1. Discussion of Survey Results
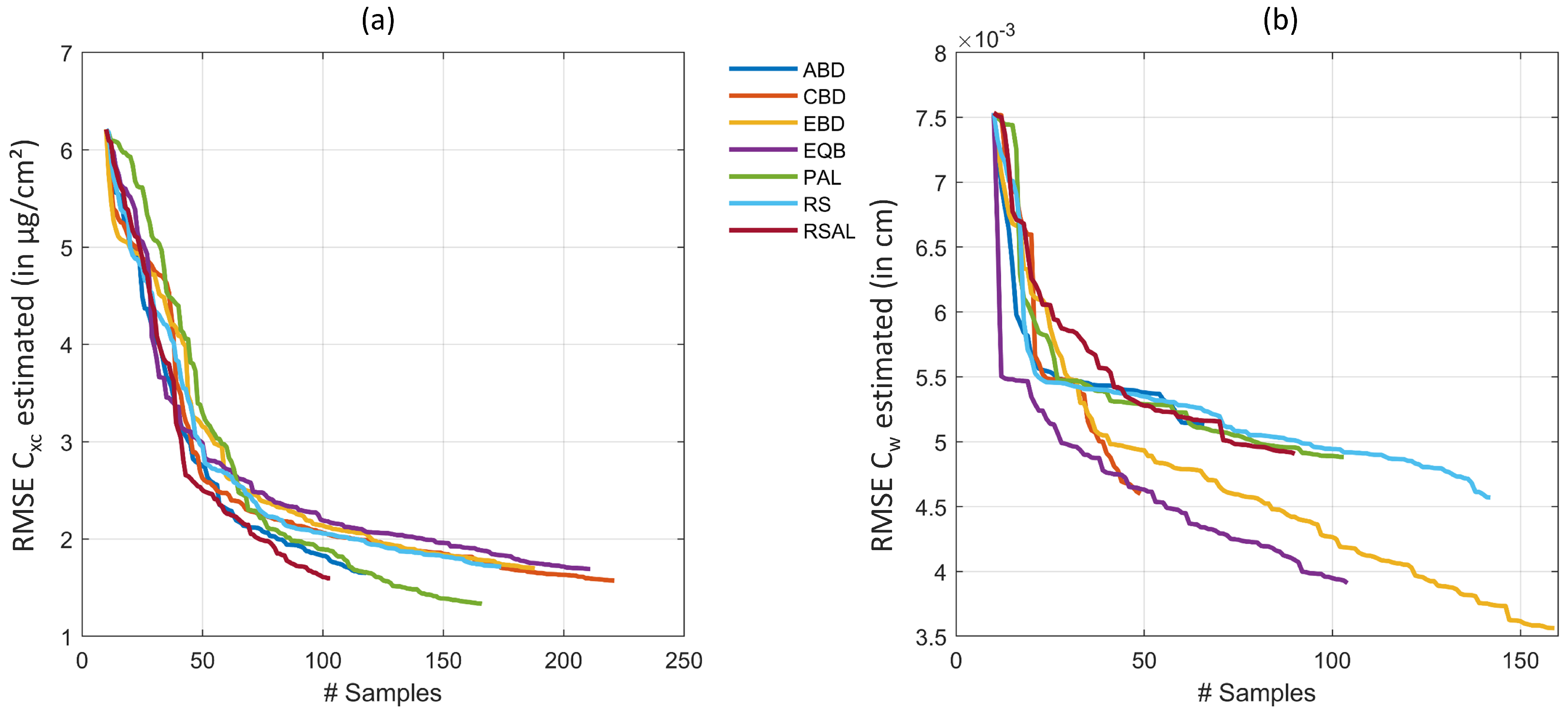
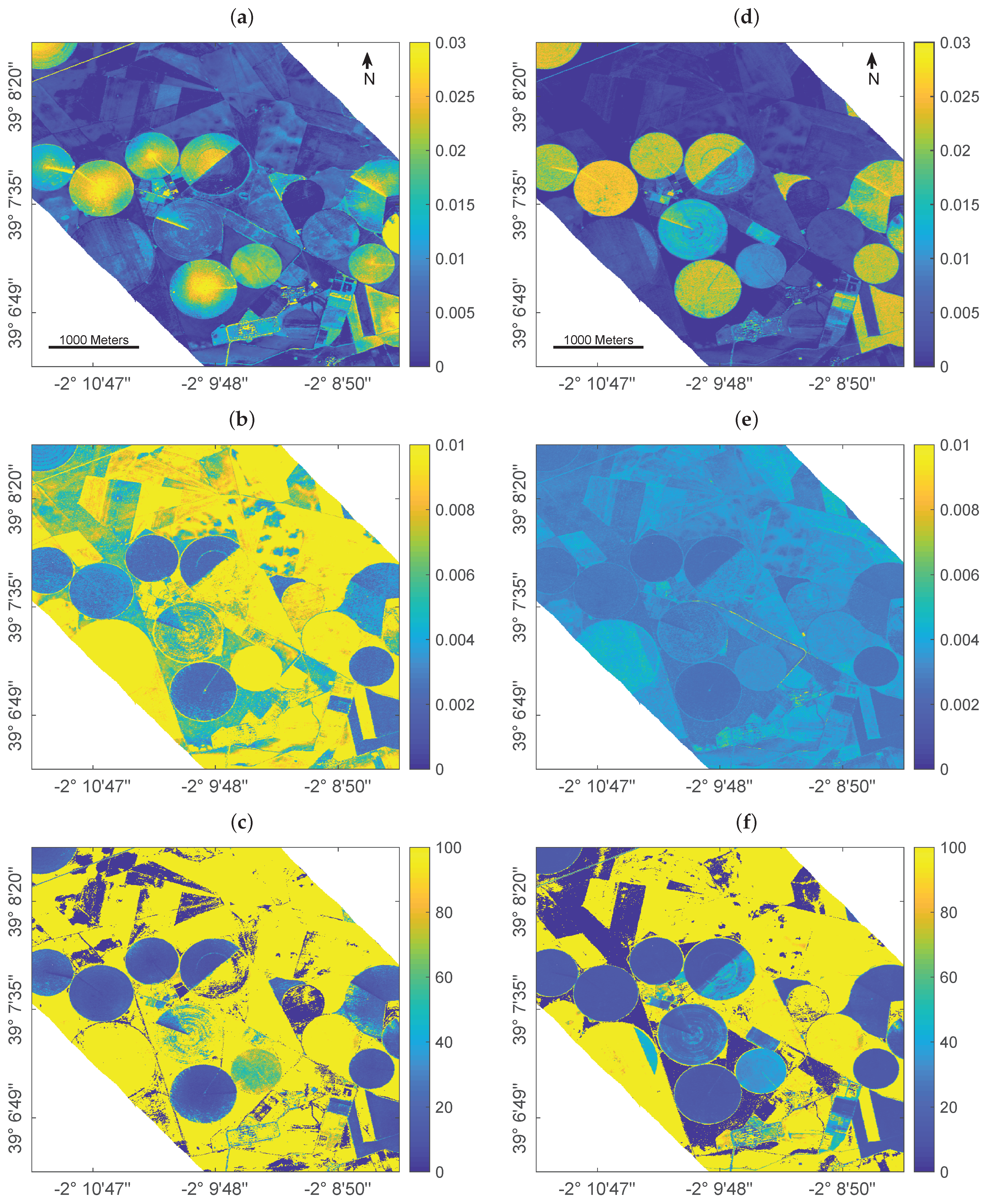
5.2. Discussion of Experimental Results
5.3. AL for Hybrid Retrieval Workflows: Ways Forward
6. Conclusions and Future Perspectives
- Use of full datasets may include redundant information, which potentially leads to decreasing retrieval accuracy compared to optimized training datasets;
- Efficient reduction of the training database (up to 80% when given 1k samples) results in decreased computational demand, hence increases processing speed for kernel-based algorithms;
- Lighter models established through AL-based sample selection facilitate their storage within software toolboxes;
- AL-based training datasets queried against in situ data are better adapted to real world situations due to the selective behaviour of the techniques;
- GPR trained with AL-reduced datasets resulted in lower retrieval uncertainties as opposed to training with a full data pool.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
Appendix A.1. Uncertainty Criteria Methods
- Entropy query-by-bagging
- Residual regression active learning
Appendix A.2. Diversity Criteria Methods
- Angle-based diversity
- Cluster-based diversity
References
- OECD. The Space Economy in Figures: How Space Contributes to the Global Economy; OECD: Paris, France, 2019. [Google Scholar]
- Nock, C.A.; Vogt, R.J.; Beisner, B.E. Functional Traits. In eLS; American Cancer Society: Atlanta, GA, USA, 2016; pp. 1–8. [Google Scholar]
- Guanter, L.; Kaufmann, H.; Segl, K.; Foerster, S.; Rogass, C.; Chabrillat, S.; Kuester, T.; Hollstein, A.; Rossner, G.; Chlebek, C.; et al. The EnMAP Spaceborne Imaging Spectroscopy Mission for Earth Observation. Remote Sens. 2015, 7, 8830. [Google Scholar] [CrossRef]
- Nieke, J.; Rast, M. Towards the Copernicus Hyperspectral Imaging Mission For The Environment (CHIME). In Proceedings of the IGARSS 2018—2018 IEEE International Geoscience and Remote Sensing Symposium, Valencia, Spain, 22–27 July 2018; pp. 157–159. [Google Scholar]
- Verrelst, J.; Camps-Valls, G.; Muñoz Marí, J.; Rivera, J.; Veroustraete, F.; Clevers, J.; Moreno, J. Optical remote sensing and the retrieval of terrestrial vegetation bio-geophysical properties—A review. ISPRS J. Photogramm. Remote. Sens. 2015, 108, 273–290. [Google Scholar] [CrossRef]
- Verrelst, J.; Malenovský, Z.; Van der Tol, C.; Camps-Valls, G.; Gastellu-Etchegorry, J.P.; Lewis, P.; North, P.; Moreno, J. Quantifying Vegetation Biophysical Variables from Imaging Spectroscopy Data: A Review on Retrieval Methods. Surv. Geophys. 2019, 40, 589–629. [Google Scholar] [CrossRef]
- Berger, K.; Verrelst, J.; Féret, J.B.; Wang, Z.; Wocher, M.; Strathmann, M.; Danner, M.; Mauser, W.; Hank, T. Crop nitrogen monitoring: Recent progress and principal developments in the context of imaging spectroscopy missions. Remote Sens. Environ. 2020, 242, 111758. [Google Scholar] [CrossRef]
- Estévez, J.; Vicent, J.; Rivera-Caicedo, J.P.; Morcillo-Pallarés, P.; Vuolo, F.; Sabater, N.; Camps-Valls, G.; Moreno, J.; Verrelst, J. Gaussian processes retrieval of LAI from Sentinel-2 top-of-atmosphere radiance data. ISPRS J. Photogramm. Remote Sens. 2020, 167, 289–304. [Google Scholar] [CrossRef]
- Brede, B.; Verrelst, J.; Gastellu-Etchegorry, J.P.; Clevers, J.G.; Goudzwaard, L.; den Ouden, J.; Verbesselt, J.; Herold, M. Assessment of workflow feature selection on forest LAI prediction with sentinel-2A MSI, landsat 7 ETM+ and Landsat 8 OLI. Remote Sens. 2020, 12, 915. [Google Scholar] [CrossRef]
- Berger, K.; Verrelst, J.; Feret, J.B. Retrieval of aboveground crop nitrogen content with a hybrid machine learning method. Int. J. Appl. Earth Obs. Geoinf. 2020, 92, 102174. [Google Scholar] [CrossRef]
- De Grave, C.; Verrelst, J.; Morcillo-Pallarés, P.; Pipia, L.; Rivera-Caicedo, J.P.; Amin, E.; Belda, S.; Moreno, J. Quantifying vegetation biophysical variables from the Sentinel-3/FLEX tandem mission: Evaluation of the synergy of OLCI and FLORIS data sources. Remote Sens. Environ. 2020, 251, 112101. [Google Scholar] [CrossRef]
- Danner, M.; Berger, K.; Wocher, M.; Mauser, W.; Hank, T. Efficient RTM-based training of machine learning regression algorithms to quantify biophysical & biochemical traits of agricultural crops. ISPRS J. Photogramm. Remote Sens. 2020. under review. [Google Scholar]
- Reichstein, M.; Camps-Valls, G.; Stevens, B.; Jung, M.; Denzler, J.; Carvalhais, N.; Prabhat. Deep learning and process understanding for data-driven Earth system science. Nature 2019, 566, 195–204. [Google Scholar] [CrossRef]
- Svendsen, D.; Martino, L.; Camps-Valls, G. Active emulation of computer codes with Gaussian processes - Application to remote sensing. Pattern Recognit. 2020, 100, 107103. [Google Scholar] [CrossRef]
- Féret, J.B.; Berger, K.; de Boissieu, F.; Malenovský, Z. PROSPECT-PRO for estimating content of nitrogen-containing leaf proteins and other carbon-based constituents. Remote Sens. Environ. 2021, 252, 112173. [Google Scholar] [CrossRef]
- Verhoef, W. Light scattering by leaf layers with application to canopy reflectance modeling: The SAIL model. Remote Sens. Environ. 1984, 16, 125–141. [Google Scholar] [CrossRef]
- Verhoef, W.; Bach, H. Coupled soil-leaf-canopy and atmosphere radiative transfer modeling to simulate hyperspectral multi-angular surface reflectance and TOA radiance data. Remote Sens. Environ. 2007, 109, 166–182. [Google Scholar] [CrossRef]
- Jacquemoud, S.; Verhoef, W.; Baret, F.; Bacour, C.; Zarco-Tejada, P.; Asner, G.; François, C.; Ustin, S. PROSPECT + SAIL models: A review of use for vegetation characterization. Remote Sens. Environ. 2009, 113, S56–S66. [Google Scholar] [CrossRef]
- Berger, K.; Atzberger, C.; Danner, M.; D’Urso, G.; Mauser, W.; Vuolo, F.; Hank, T. Evaluation of the PROSAIL model capabilities for future hyperspectral model environments: A review study. Remote Sens. 2018, 10, 85. [Google Scholar] [CrossRef]
- Weiss, M.; Baret, F.; Myneni, R.B.; Pragnère, A.; Knyazikhin, Y. Investigation of a model inversion technique to estimate canopy biophysical variables from spectral and directional reflectance data. Agronomie 2000, 20, 3–22. [Google Scholar] [CrossRef]
- Rasmussen, C.E.; Williams, C.K.I. Gaussian Processes for Machine Learning; The MIT Press: New York, NY, USA, 2006. [Google Scholar]
- Camps-Valls, G.; Verrelst, J.; Munoz-Mari, J.; Laparra, V.; Mateo-Jimenez, F.; Gomez-Dans, J. A survey on Gaussian processes for earth-observation data analysis: A comprehensive investigation. IEEE Geosci. Remote Sens. Mag. 2016, 4, 58–78. [Google Scholar] [CrossRef]
- Verrelst, J.; Rivera, J.; Moreno, J.; Camps-Valls, G. Gaussian processes uncertainty estimates in experimental Sentinel-2 LAI and leaf chlorophyll content retrieval. ISPRS J. Photogramm. Remote Sens. 2013, 86, 157–167. [Google Scholar] [CrossRef]
- Park, J.; Lechevalier, D.; Ak, R.; Ferguson, M.; Law, K.H.; Lee, Y.T.T.; Rachuri, S. Gaussian Process Regression (GPR) Representation in Predictive Model Markup Language (PMML). Smart Sustain. Manuf. Syst. 2017, 1, 121. [Google Scholar] [CrossRef]
- Estévez, J.; Berger, K.; Vicent, J.; Rivera-Caicedo, J.P.; Morcillo-Pallarés, P.; Wocher, M.; Verrelst, J. Top-of-atmosphere retrieval of multiple crop traits using variational heteroscedastic Gaussian processes within a hybrid workflow. Remote Sens. Environ. 2021. under review. [Google Scholar]
- Campos-Taberner, M.; Moreno-Martínez, Á.; García-Haro, F.J.; Camps-Valls, G.; Robinson, N.P.; Kattge, J.; Running, S.W. Global Estimation of Biophysical Variables from Google Earth Engine Platform. Remote Sens. 2018, 10, 1167. [Google Scholar] [CrossRef]
- Upreti, D.; Huang, W.; Kong, W.; Pascucci, S.; Pignatti, S.; Zhou, X.; Ye, H.; Casa, R. A comparison of hybrid machine learning algorithms for the retrieval of wheat biophysical variables from sentinel-2. Remote Sens. 2019, 11, 481. [Google Scholar] [CrossRef]
- Locherer, M.; Hank, T.; Danner, M.; Mauser, W. Retrieval of Seasonal Leaf Area Index from Simulated EnMAP Data through Optimized LUT-Based Inversion of the PROSAIL Model. Remote Sens. 2015, 7, 10321–10346. [Google Scholar] [CrossRef]
- Duan, S.B.; Li, Z.L.; Wu, H.; Tang, B.H.; Ma, L.; Zhao, E.; Li, C. Inversion of the PROSAIL model to estimate leaf area index of maize, potato, and sunflower fields from unmanned aerial vehicle hyperspectral data. Int. J. Appl. Earth Obs. Geoinf. 2014, 26, 12–20. [Google Scholar] [CrossRef]
- Darvishzadeh, R.; Matkan, A.A.; Ahangar, A.D. Inversion of a Radiative Transfer Model for Estimation of Rice Canopy Chlorophyll Content Using a Lookup-Table Approach. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2012, 5, 1222–1230. [Google Scholar] [CrossRef]
- Rivera-Caicedo, J.P.; Verrelst, J.; Muñoz-Marí, J.; Camps-Valls, G.; Moreno, J. Hyperspectral dimensionality reduction for biophysical variable statistical retrieval. ISPRS J. Photogramm. Remote. Sens. 2017, 132, 88–101. [Google Scholar] [CrossRef]
- Verrelst, J.; Rivera, J.P.; Gitelson, A.; Delegido, J.; Moreno, J.; Camps-Valls, G. Spectral band selection for vegetation properties retrieval using Gaussian processes regression. Int. J. Appl. Earth Obs. Geoinf. 2016, 52, 554–567. [Google Scholar] [CrossRef]
- Pasolli, E.; Melgani, F.; Alajlan, N.; Bazi, Y. Active Learning Methods for Biophysical Parameter Estimation. IEEE Trans. Geosci. Remote Sens. 2012, 50, 4071–4084. [Google Scholar] [CrossRef]
- Tuia, D.; Volpi, M.; Copa, L.; Kanevski, M.; Munoz-Mari, J. A Survey of Active Learning Algorithms for Supervised Remote Sensing Image Classification. IEEE J. Sel. Top. Signal Process. 2011, 5, 606–617. [Google Scholar] [CrossRef]
- Liu, H.; Motoda, H. Instance Selection and Construction for Data Mining; Springer: Springer Science+Business Media Dordrecht: Dordrecht, The Netherlands, 2001. [Google Scholar]
- Settles, B. Active Learning Literature Survey; Computer Sciences Technical Report 1648; University of Wisconsin–Madison: Madison, WI, USA, 2009. [Google Scholar]
- Settles, B. Active Learning; Morgan & Claypool Publishers: Williston, VT, USA, 2012. [Google Scholar]
- Meek, C.; Thiesson, B.; Heckerman, D. The learning-curve sampling method applied to model-based clustering. J. Mach. Learn. Res. 2002, 2, 397–418. [Google Scholar]
- Elrafey, A.; Wojtusiak, J. A Hybrid Active Learning and Progressive Sampling Algorithm. Int. J. Mach. Learn. Comput. 2018, 8. [Google Scholar] [CrossRef]
- Martino, L.; Svendsen, D.H.; Vicent, J.; Camps-Valls, G. Adaptive Sequential Interpolator Using Active Learning for Efficient Emulation of Complex Systems. In Proceedings of the ICASSP 2020—2020 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), Barcelona, Spain, 4–8 May 2020; pp. 3577–3581. [Google Scholar]
- Verrelst, J.; Dethier, S.; Rivera, J.P.; Munoz-Mari, J.; Camps-Valls, G.; Moreno, J. Active learning methods for efficient hybrid biophysical variable retrieval. IEEE Geosci. Remote. Sens. Lett. 2016, 13, 1012–1016. [Google Scholar] [CrossRef]
- Polewski, P.; Yao, W.; Heurich, M.; Krzystek, P.; Stilla, U. Combining Active and Semisupervised Learning of Remote Sensing Data Within a Renyi Entropy Regularization Framework. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2016, 9, 2910–2922. [Google Scholar] [CrossRef]
- Shi, Q.; Du, B.; Zhang, L. Spatial Coherence-Based Batch-Mode Active Learning for Remote Sensing Image Classification. IEEE Trans. Image Process. Publ. IEEE Signal Process. Soc. 2015, 24, 2037–2050. [Google Scholar]
- Pradhan, M.K.; Minz, S.; Shrivastava, V.K. Fast active learning for hyperspectral image classification using extreme learning machine. IET Image Proc. 2019, 13, 549–555. [Google Scholar] [CrossRef]
- Kumar, P.; Gupta, A. Active Learning Query Strategies for Classification, Regression, and Clustering: A Survey. J. Comput. Sci. Tech. 2020, 35, 913–945. [Google Scholar] [CrossRef]
- Wang, K.; Zhang, D.; Li, Y.; Zhang, R.; Lin, L. Cost-Effective Active Learning for Deep Image Classification. IEEE Trans. Circuits Syst. Video Technol. 2016, 27, 2591–2600. [Google Scholar] [CrossRef]
- Luo, Z.; Hauskrecht, M. Group-Based Active Learning of Classification Models. Proc. Int. Fla Res. Soc. Conf. Fla. Res. Symp. 2017, 2017, 92. [Google Scholar]
- Du, J.; Ling, C.X. Asking Generalized Queries to Domain Experts to Improve Learning. IEEE Trans. Knowl. Data Eng. 2010, 22, 812–825. [Google Scholar] [CrossRef]
- Nguyen, H.T.; Smeulders, A. Active Learning Using Pre-Clustering; Association for Computing Machinery: New York, NY, USA, 2004. [Google Scholar]
- Chakraborty, S.; Balasubramanian, V.; Panchanathan, S. Adaptive Batch Mode Active Learning. IEEE Trans. Neural Netw. Learn. Syst. 2014, 26, 1747–1760. [Google Scholar] [CrossRef] [PubMed]
- Bruzzone, L.; Persello, C. Active learning for classification of remote sensing images. In Proceedings of the 2009 IEEE International Geoscience and Remote Sensing Symposium, Cape Town, South Africa, 12–17 July 2009. [Google Scholar]
- Zhang, Z.; Pasolli, E.; Yang, H.L.; Crawford, M.M. Multimetric Active Learning for Classification of Remote Sensing Data. IEEE Geosci. Remote Sens. Lett. 2016, 13, 1007–1011. [Google Scholar] [CrossRef]
- Demir, B.; Persello, C.; Bruzzone, L. Batch-Mode Active-Learning Methods for the Interactive Classification of Remote Sensing Images. IEEE Trans. Geosci. Remote Sens. 2011, 49, 1014–1031. [Google Scholar] [CrossRef]
- Ienco, D.; Bifet, A.; Žliobaitė, I.; Pfahringer, B. Clustering Based Active Learning for Evolving Data Streams. In Discovery Science; Springer: Berlin, Germany, 2013; pp. 79–93. [Google Scholar]
- Rivera, J.P.; Verrelst, J.; Gómez-Dans, J.; Muñoz Marí, J.; Moreno, J.; Camps-Valls, G. An Emulator Toolbox to Approximate Radiative Transfer Models with Statistical Learning. Remote Sens. 2015, 7, 9347–9370. [Google Scholar] [CrossRef]
- Verrelst, J.; Sabater, N.; Rivera, J.P.; Muñoz Marí, J.; Vicent, J.; Camps-Valls, G.; Moreno, J. Emulation of Leaf, Canopy and Atmosphere Radiative Transfer Models for Fast Global Sensitivity Analysis. Remote Sens. 2016, 8, 673. [Google Scholar] [CrossRef]
- Verrelst, J.; Rivera-Caicedo, J.; Muñoz Marí, J.; Camps-Valls, G.; Moreno, J. SCOPE-Based Emulators for Fast Generation of Synthetic Canopy Reflectance and Sun-Induced Fluorescence Spectra. Remote Sens. 2017, 9, 927. [Google Scholar] [CrossRef]
- Vicent, J.; Verrelst, J.; Rivera Caicedo, J.; Sabater Medina, N.; Muñoz, J.; Camps-Valls, G.; Moreno, J. Emulation as an Accurate Alternative to Interpolation in Sampling Radiative Transfer Codes. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2018, 11, 1–14. [Google Scholar] [CrossRef]
- Smucker, B.; Krzywinski, M.; Altman, N. Optimal experimental design. Nat. Methods 2018, 15, 559–560. [Google Scholar] [CrossRef]
- Chaloner, K.; Verdinelli, I. Bayesian Experimental Design: A Review. Stat. Sci. 1995, 10, 273–304. [Google Scholar] [CrossRef]
- Ford, I.; Titterington, D.M.; Kitsos, C.P. Recent Advances in Nonlinear Experimental Design. Technometrics 1989, 31, 49–60. [Google Scholar] [CrossRef]
- Busby, D. Hierarchical adaptive experimental design for Gaussian process emulators. Reliab. Eng. Syst. Saf. 2009, 94, 1183–1193. [Google Scholar] [CrossRef]
- Krause, A.; Singh, A.; Guestrin, C. Near-Optimal Sensor Placements in Gaussian Processes: Theory, Efficient Algorithms and Empirical Studies. J. Mach. Learn. Res. 2008, 9, 235–284. [Google Scholar]
- Krause, A.; Guestrin, C.; Gupta, A.; Kleinberg, J. Near-optimal sensor placements: Maximizing information while minimizing communication cost. In Proceedings of the 2006 5th International Conference on Information Processing in Sensor Networks, Nashville, TN, USA, 19–21 April 2006; pp. 2–10. [Google Scholar]
- Niederreiter, H. Random Number Generation and Quasi-Monte Carlo Methods; Society for Industrial and Applied Mathematics: Philadelphia, PA, USA, 1992. [Google Scholar]
- Pronzato, L.; Müller, W. Design of computer experiments: space filling and beyond. Stat. Comput. 2012, 22, 681–701. [Google Scholar] [CrossRef]
- Borodin, A.; Olshanski, G. Distributions on partitions, point processes and the hypergeometric kernel. Commun. Math. Phys. 2000, 22, 335–358. [Google Scholar] [CrossRef]
- Borodin, A.; Diaconis, P.; Fulman, J. On adding a list of numbers (and other one-dependent determinantal processes). Bull. Am. Math. Soc. 2010, 47, 639–670. [Google Scholar] [CrossRef]
- Marvasti, F. Nonuniform Sampling: Theory and Practice; Springer: Berlin, Germany, 2012. [Google Scholar]
- Seleznjev, O.; Shykula, M. Uniform and non-uniform quantization of Gaussian processes. Math. Commun. 2012, 17, 447–460. [Google Scholar]
- Llorente, F.; Martino, L.V.; Elvira, D.D.; Lopez-Santiago, J. Adaptive quadrature schemes for Bayesian inference via active learning. arXiv 2020, arXiv:2006.00535. [Google Scholar]
- Kanagawa, M.; Hennig, P. Convergence guarantees for adaptive Bayesian quadrature methods. In Proceedings of the Advances in Neural Information Processing Systems, Vancouver, BC, Canada, 8–14 December 2019; pp. 6234–6245. [Google Scholar]
- Llorente, F.; Martino, L.V.; Elvira, D.D.; Lopez-Santiago, J. Deep Importance Sampling based on Regression for Model Inversion and Emulation. arXiv 2020, arXiv:2010.10346. [Google Scholar]
- Cleary, E.; Garbuno-Inigo, A.; Lan, S.; Schneider, T.; Stuart, A.M. Calibrate, emulate, sample. J. Comput. Phys. 2021, 424, 109716. [Google Scholar] [CrossRef]
- Servera, J.V.; Alonso, L.; Martino, L.; Sabater, N.; Verrelst, J.; Camps-Valls, G.; Moreno, J. Gradient-Based Automatic Lookup Table Generator for Radiative Transfer Models. IEEE Trans. Geosci. Remote Sens. 2018, 57, 1040–1048. [Google Scholar] [CrossRef]
- Vicent, J.; Verrelst, J.; Sabater, N.; Alonso, L.; Rivera-Caicedo, J.P.; Martino, L.; Muñoz Marí, J.; Moreno, J. Comparative analysis of atmospheric radiative transfer models using the Atmospheric Look-up table Generator (ALG) toolbox (version 2.0). Geosci. Model Dev. 2020, 13, 1945–1957. [Google Scholar] [CrossRef]
- Lewis, D.D.; Gale, W.A. A Sequential Algorithm for Training Text Classifiers. In SIGIR ’94; Springer: London, UK, 1994; pp. 3–12. [Google Scholar]
- Zhang, C.; Chen, T. An active learning framework for content-based information retrieval. IEEE Trans. Multimedia 2002, 4, 260–268. [Google Scholar] [CrossRef]
- He, T.; Zhang, S.; Xin, J.; Zhao, P.; Wu, J.; Xian, X.; Li, C.; Cui, Z. An active learning approach with uncertainty, representativeness, and diversity. Sci. World J. 2014, 2014, 827586. [Google Scholar] [CrossRef]
- Lu, X.; Zhang, J.; Li, T.; Zhang, Y. Incorporating diversity into self-learning for synergetic classification of hyperspectral and panchromatic images. Remote Sens. 2016, 8, 804. [Google Scholar] [CrossRef]
- Douak, F.; Melgani, F.; Benoudjit, N. Kernel ridge regression with active learning for wind speed prediction. Appl. Energy 2013, 103, 328–340. [Google Scholar] [CrossRef]
- Douak, F.; Benoudjit, N.; Melgani, F. A two-stage regression approach for spectroscopic quantitative analysis. Chemom. Intell. Lab. Syst. 2011, 109, 34–41. [Google Scholar] [CrossRef]
- Gu, Y.; Jin, Z.; Chiu, S. Active learning combining uncertainty and diversity for multi-class image classification. IET Comput. Vis. 2015, 9, 400–407. [Google Scholar] [CrossRef]
- Cronin, P.; Ryan, F.; Coughlan, M. Undertaking a literature review: A step-by-step approach. Br. J. Nurs. 2008, 17, 38–43. [Google Scholar] [CrossRef]
- Shahraiyni, T.H.; Schaale, M.; Fell, F.; Fischer, J.; Preusker, R.; Vatandoust, M.; Shouraki, B.S.; Tajrishy, M.; Khodaparast, H.; Tavakoli, A. Application of the Active Learning Method for the estimation of geophysical variables in the Caspian Sea from satellite ocean colour observations. Int. J. Remote Sens. 2007, 28, 4677–4683. [Google Scholar] [CrossRef]
- Douak, F.; Melgani, F.; Alajlan, N.; Pasolli, E.; Bazi, Y.; Benoudjit, N. Active learning for spectroscopic data regression. J. Chemom. 2012, 26, 374–383. [Google Scholar] [CrossRef]
- Djamai, N.; Fernandes, R. Active learning regularization increases clear sky retrieval rates for vegetation biophysical variables using Sentinel-2 data. Remote Sens. Environ. 2021, 254, 112241. [Google Scholar] [CrossRef]
- Baret, F.; Pavageau, K.; Béal, D.; Weiss, M.; Berthelot, B.; Regner, P. Algorithm Theoretical Basis Document for MERIS Top of Atmosphere Land Products (TOA VEG). Technical Report. March 2006. Available online: http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.565.5755&rep=rep1&type=pdf (accessed on 7 July 2020).
- Baret, F.; Buis, S. Estimating Canopy Characteristics from Remote Sensing Observations: Review of Methods and Associated Problems. In Advances in Land Remote Sensing: System, Modeling, Inversion and Application; Springer: Dordrecht, The Netherlands, 2008; pp. 173–201. [Google Scholar]
- Verrelst, J.; Berger, K.; Rivera-Caicedo, J.P. Intelligent sampling for vegetation nitrogen mapping based on hybrid machine learning algorithms. IEEE Geosci. Remote. Sens. Lett. 2020, in press. [Google Scholar] [CrossRef]
- Upreti, D.; Pignatti, S.; Pascucci, S.; Tolomio, M.; Huang, W.; Casa, R. Bayesian Calibration of the Aquacrop-OS Model for Durum Wheat by Assimilation of Canopy Cover Retrieved from VENμS Satellite Data. Remote Sens. 2020, 12, 2666. [Google Scholar] [CrossRef]
- Zhou, X.; Zhang, J.; Chen, D.; Huang, Y.; Kong, W.; Yuan, L.; Ye, H.; Huang, W. Assessment of Leaf Chlorophyll Content Models for Winter Wheat Using Landsat-8 Multispectral Remote Sensing Data. Remote Sens. 2020, 12, 2574. [Google Scholar] [CrossRef]
- Pipia, L.; Amin, E.; Belda, S.; Salinero Delgado, M.; Verrelst, J. LAI Green mapping and cloud gap-filling using Gaussian Process Regression in Google Earth Engine. Remote Sens. 2021. under review. [Google Scholar]
- Suykens, J.; Vandewalle, J. Least squares support vector machine classifiers. Neural Process. Lett. 1999, 9, 293–300. [Google Scholar] [CrossRef]
- Moore, C.J.; Chua, A.J.K.; Berry, C.P.L.; Gair, J.R. Fast methods for training Gaussian processes on large datasets. R. Soc. Open Sci. 2016, 3. [Google Scholar] [CrossRef] [PubMed]
- Gorelick, N.; Hancher, M.; Dixon, M.; Ilyushchenko, S.; Thau, D.; Moore, R. Google Earth Engine: Planetary-scale geospatial analysis for everyone. Remote. Sens. Environ. 2017, 202, 18–27. [Google Scholar] [CrossRef]
- Weiss, M.; Baret, F. S2ToolBox Level 2 products: LAI, FAPAR, FCOVER, Version 1.1. In ESA Contract nr 4000110612/14/I-BG (p. 52); INRA: Avignon, France, 2016. [Google Scholar]
- Lázaro-Gredilla, M.; Titsias, M.K.; Verrelst, J.; Camps-Valls, G. Retrieval of Biophysical Parameters With Heteroscedastic Gaussian Processes. IEEE Geosci. Remote Sens. Lett. 2013, 11, 838–842. [Google Scholar] [CrossRef]
- Wocher, M.; Berger, K.; Danner, M.; Mauser, W.; Hank, T. Physically-based retrieval of canopy equivalent water thickness using hyperspectral data. Remote Sens. 2018, 10, 1924. [Google Scholar] [CrossRef]
- Rocchi, L.; Rustioni, L.; Failla, O. Chlorophyll and carotenoid quantifications in white grape (Vitis vinifera L.) skins by reflectance spectroscopy. J. Grapevine Res. 2016, 55, 11–16. [Google Scholar]
- Chappelle, E.W.; Kim, M.S.; McMurtrey, J.E. Ratio Analysis of Reflectance Spectra (RARS)—An Algorithm for the Remote Estimation of the Concentrations of Chlorophyll a, Chlorophyll b, and Carotenoids in Soybean Leaves. 1992. Available online: https://www.sciencedirect.com/science/article/abs/pii/0034425792900893 (accessed on 7 July 2020).
- Danner, M.; Berger, K.; Wocher, M.; Mauser, W.; Hank, T. Fitted PROSAIL Parameterization of Leaf Inclinations, Water Content and Brown Pigment Content for Winter Wheat and Maize Canopies. Remote Sens. 2019, 11, 1150. [Google Scholar] [CrossRef]
- Baret, F.; Andrieu, B.; Guyot, G. A Simple Model for Leaf Optical Properties in Visible and Near-Infrared: Application to the Analysis of Spectral Shifts Determinism. In Applications of Chlorophyll Fluorescence in Photosynthesis Research, Stress Physiology, Hydrobiology and Remote Sensing: An Introduction to the Various Fields of Applications of the in vivo Chlorophyll Fluorescence also Including the Proceedings of the First International Chlorophyll Fluorescence Symposium Held in the Physikzentrum, Bad Honnef, F.R.G., 6–8 June 1998; Springer: Dordrecht, The Netherlands, 1988; pp. 345–351. [Google Scholar]
- Moreno, J.F.; Alonso, L.; Fernàndez, G.; Fortea, J.C.; Gandía, S. The SPECTRA Barrax Campaign (SPARC): Overview and First Results from CHRIS Data; European Space Agency, (Special Publication) ESA SP: Paris, France, 2004. [Google Scholar]
- Verrelst, J.; Rivera, J.; Veroustraete, F.; Muñoz Marí, J.; Clevers, J.; Camps-Valls, G.; Moreno, J. Experimental Sentinel-2 LAI estimation using parametric, non-parametric and physical retrieval methods—A comparison. ISPRS J. Photogramm. Remote Sens. 2015, 108, 260–272. [Google Scholar] [CrossRef]
- Wocher, M.; Berger, K.; Danner, M.; Mauser, W.; Hank, T. RTM-based dynamic absorption integrals for the retrieval of biochemical vegetation traits. Int. J. Appl. Earth Obs. Geoinf. 2020, 93, 102219. [Google Scholar] [CrossRef]
- Guanter, L.; Alonso, L.; Moreno, J. A method for the surface reflectance retrieval from PROBA/CHRIS data over land: Application to ESA SPARC campaigns. IEEE Trans. Geosci. Remote Sens. 2005, 43, 2908–2917. [Google Scholar] [CrossRef]
- Féret, J.B.; Gitelson, A.A.; Noble, S.D.; Jacquemoud, S. PROSPECT-D: Towards modeling leaf optical properties through a complete lifecycle. Remote Sens. Environ. 2017, 193, 204–215. [Google Scholar] [CrossRef]
- Verrelst, J.; Romijn, E.; Kooistra, L. Mapping vegetation density in a heterogeneous river floodplain ecosystem using pointable CHRIS/PROBA data. Remote Sens. 2012, 4, 2866–2889. [Google Scholar] [CrossRef]
- Caicedo, J.; Verrelst, J.; Munoz-Mari, J.; Moreno, J.; Camps-Valls, G. Toward a semiautomatic machine learning retrieval of biophysical parameters. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2014, 7, 1249–1259. [Google Scholar] [CrossRef]
- Demir, B.; Bruzzone, L. A multiple criteria active learning method for support vector regression. Pattern Recognit. 2014, 47, 2558–2567. [Google Scholar] [CrossRef]
- Berger, K.; Atzberger, C.; Danner, M.; Wocher, M.; Mauser, W.; Hank, T. Model-Based Optimization of Spectral Sampling for the Retrieval of Crop Variables with the PROSAIL Model. Remote Sens. 2018, 10, 2063. [Google Scholar] [CrossRef]
- Vicent, J.; Sabater, N.; Tenjo, C.; Acarreta, J.R.; Manzano, M.; Rivera, J.P.; Jurado, P.; Franco, R.; Alonso, L.; Verrelst, J.; et al. FLEX End-to-End Mission Performance Simulator. IEEE Trans. Geosci. Remote Sens. 2016, 54, 4215–4223. [Google Scholar] [CrossRef]
- Verrelst, J.; Rivera Caicedo, J.P.; Vicent, J.; Morcillo Pallarés, P.; Moreno, J. Approximating Empirical Surface Reflectance Data through Emulation: Opportunities for Synthetic Scene Generation. Remote Sens. 2019, 11, 157. [Google Scholar] [CrossRef]
- Aharon, S.; Peleg, Z.; Argaman, E.; Ben-David, R.; Lati, R.N. Image-Based High-Throughput Phenotyping of Cereals Early Vigor and Weed-Competitiveness Traits. Remote Sens. 2020, 12, 3877. [Google Scholar] [CrossRef]
- Weiss, M.; Jacob, F.; Duveiller, G. Remote sensing for agricultural applications: A meta-review. Remote. Sens. Environ. 2020, 236, 111402. [Google Scholar] [CrossRef]
- Jay, S.; Maupas, F.; Bendoula, R.; Gorretta, N. Retrieving LAI, chlorophyll and nitrogen contents in sugar beet crops from multi-angular optical remote sensing: Comparison of vegetation indices and PROSAIL inversion for field phenotyping. Field Crop. Res. 2017, 210, 33–46. [Google Scholar] [CrossRef]
- Patra, S.; Bruzzone, L. A cluster-assumption based batch mode active learning technique. Pattern Recognit. Lett. 2012, 33, 1042–1048. [Google Scholar] [CrossRef]
| References | Sensors | Estimated Traits | ML Algorithms | AL Methods |
|---|---|---|---|---|
| Verrelst et al. [41] | Sentinel-3 OLCI (simulated) | LAI, | KRR, GPR | PAL, EQB, RSAL, EBD, ABD, CBD |
| Upreti et al. [27] | Sentinel-2 | LAI, , Fcover, fAPAR | GPR | EBD, ABD, CBD |
| Verrelst et al. [90] | EnMAP (resampled) | KRR, VHGPR | PAL, EQB, RSAL, EBD, ABD, CBD | |
| Upreti et al. [91] | VENS | Fcover | GPR | EBD |
| Zhou et al. [92] | Landsat-8 OLI | GPR | PAL, EQB, RSAL, EBD, ABD, CBD | |
| Pipia et al. [93] | Sentinel-2 | green LAI | GPR | PAL, EQB, RSAL, EBD, ABD, CBD |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Berger, K.; Rivera Caicedo, J.P.; Martino, L.; Wocher, M.; Hank, T.; Verrelst, J. A Survey of Active Learning for Quantifying Vegetation Traits from Terrestrial Earth Observation Data. Remote Sens. 2021, 13, 287. https://doi.org/10.3390/rs13020287
Berger K, Rivera Caicedo JP, Martino L, Wocher M, Hank T, Verrelst J. A Survey of Active Learning for Quantifying Vegetation Traits from Terrestrial Earth Observation Data. Remote Sensing. 2021; 13(2):287. https://doi.org/10.3390/rs13020287
Chicago/Turabian StyleBerger, Katja, Juan Pablo Rivera Caicedo, Luca Martino, Matthias Wocher, Tobias Hank, and Jochem Verrelst. 2021. "A Survey of Active Learning for Quantifying Vegetation Traits from Terrestrial Earth Observation Data" Remote Sensing 13, no. 2: 287. https://doi.org/10.3390/rs13020287
APA StyleBerger, K., Rivera Caicedo, J. P., Martino, L., Wocher, M., Hank, T., & Verrelst, J. (2021). A Survey of Active Learning for Quantifying Vegetation Traits from Terrestrial Earth Observation Data. Remote Sensing, 13(2), 287. https://doi.org/10.3390/rs13020287