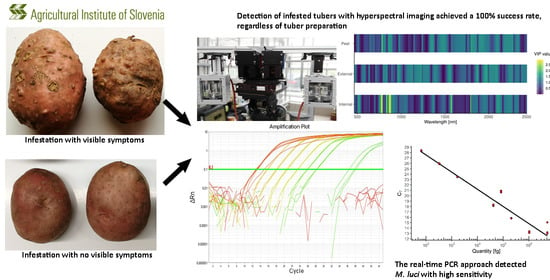
Detection of Root-Knot Nematode Meloidogyne luci Infestation of Potato Tubers Using Hyperspectral Remote Sensing and Real-Time PCR Molecular Methods
Abstract
1. Introduction
2. Materials and Methods
2.1. Microplot Potato Cultivation
2.2. Tuber Preparation
2.3. Remote Detection
2.3.1. Image Acquisition
2.3.2. Image Segmentation and Data Pre-Processing
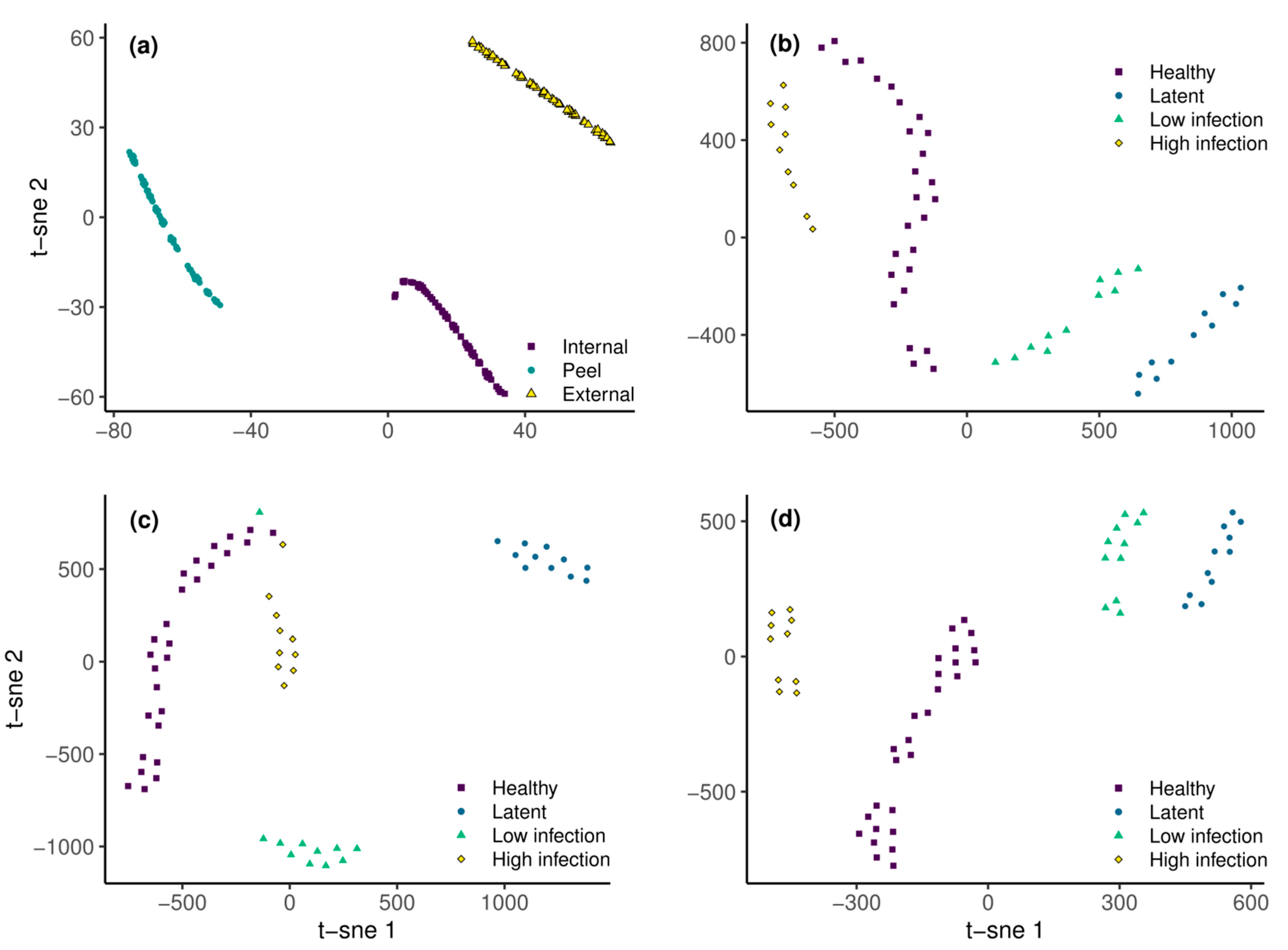
2.3.3. Data Dimensionality Reduction and Feature Extraction
2.3.4. Supervised Classification
2.4. Molecular Detection
2.4.1. Sampling
2.4.2. DNA Isolation
2.4.3. Real-Time PCR
3. Results
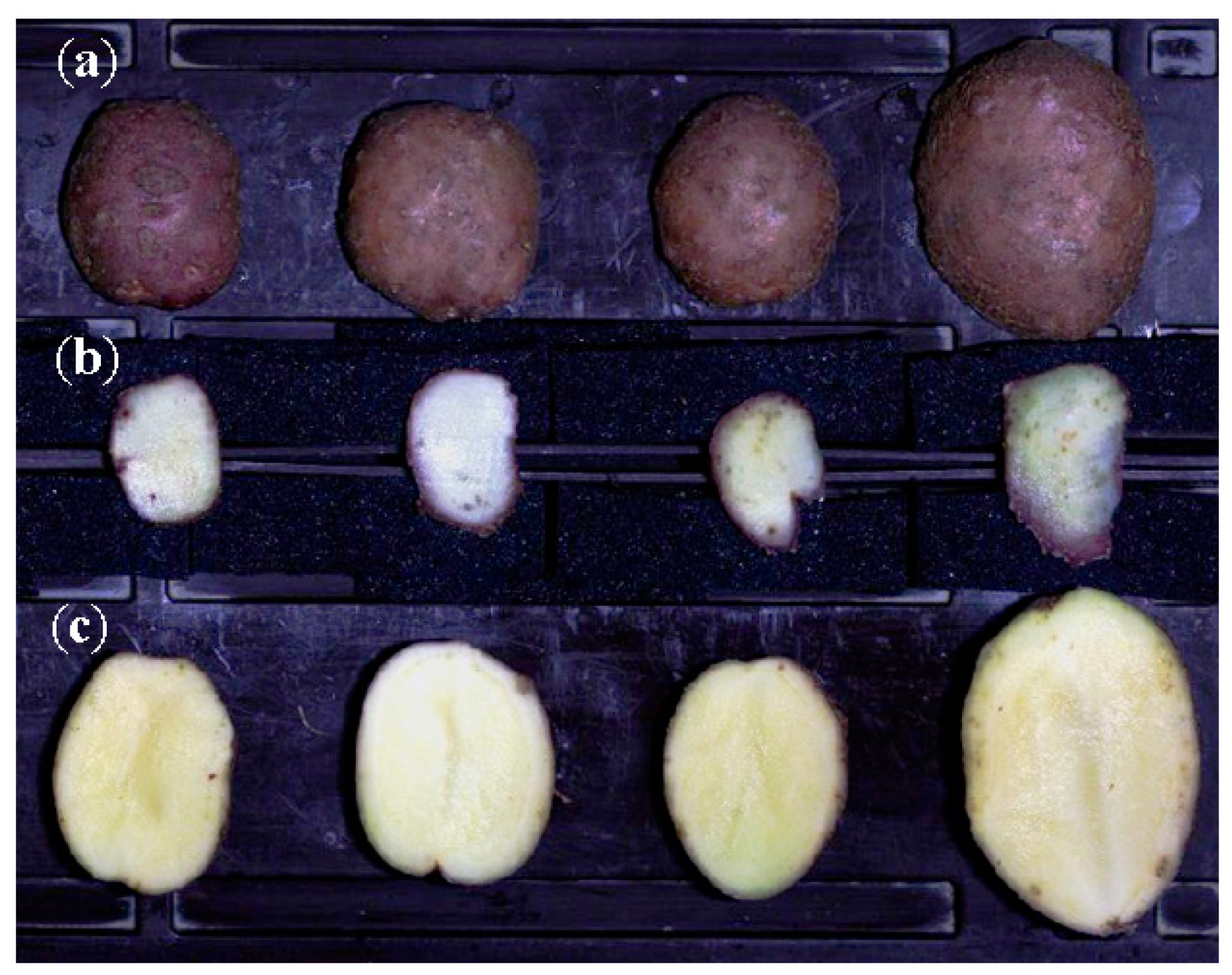
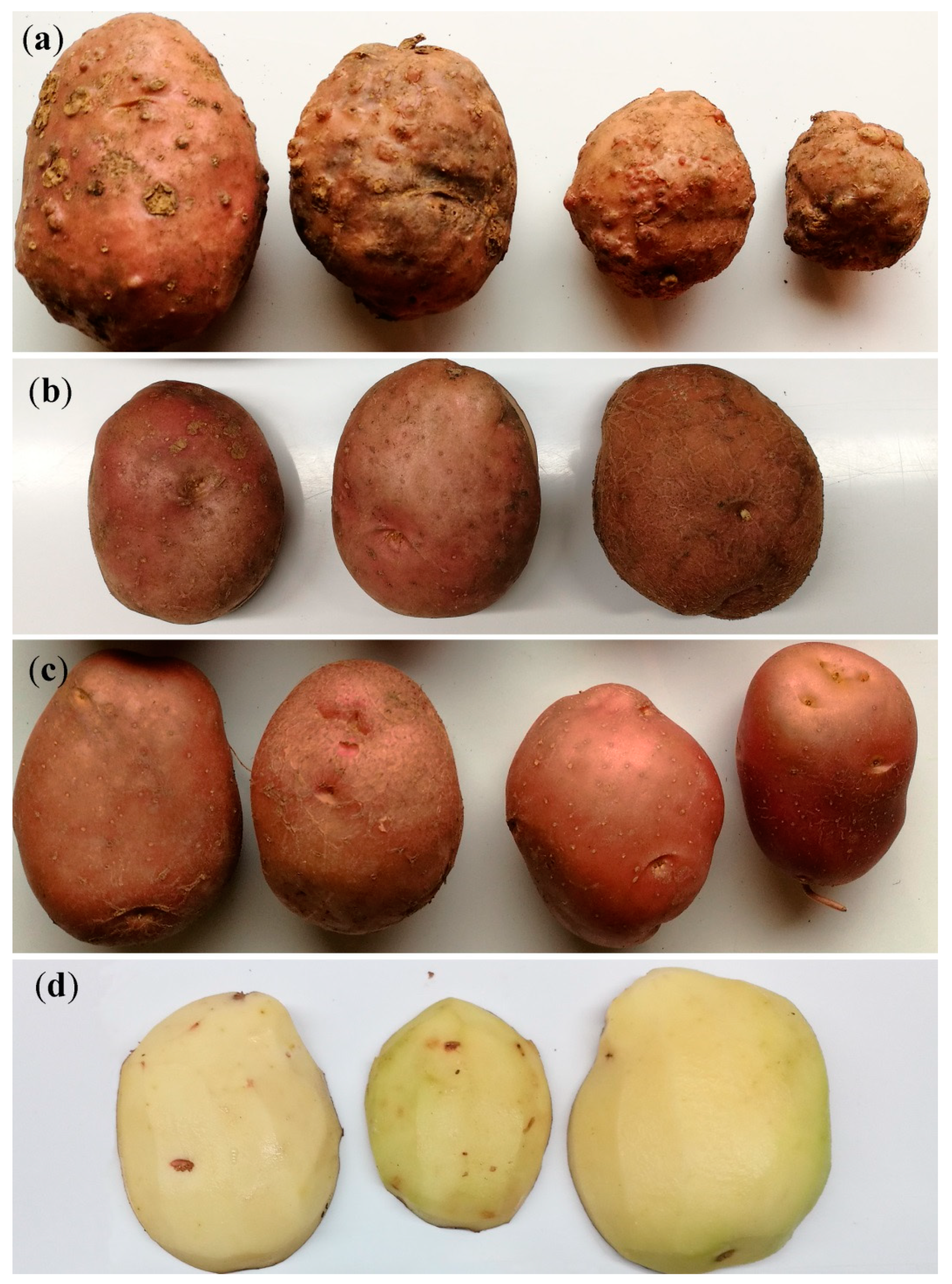
3.1. Symptoms of M. luci Infestation
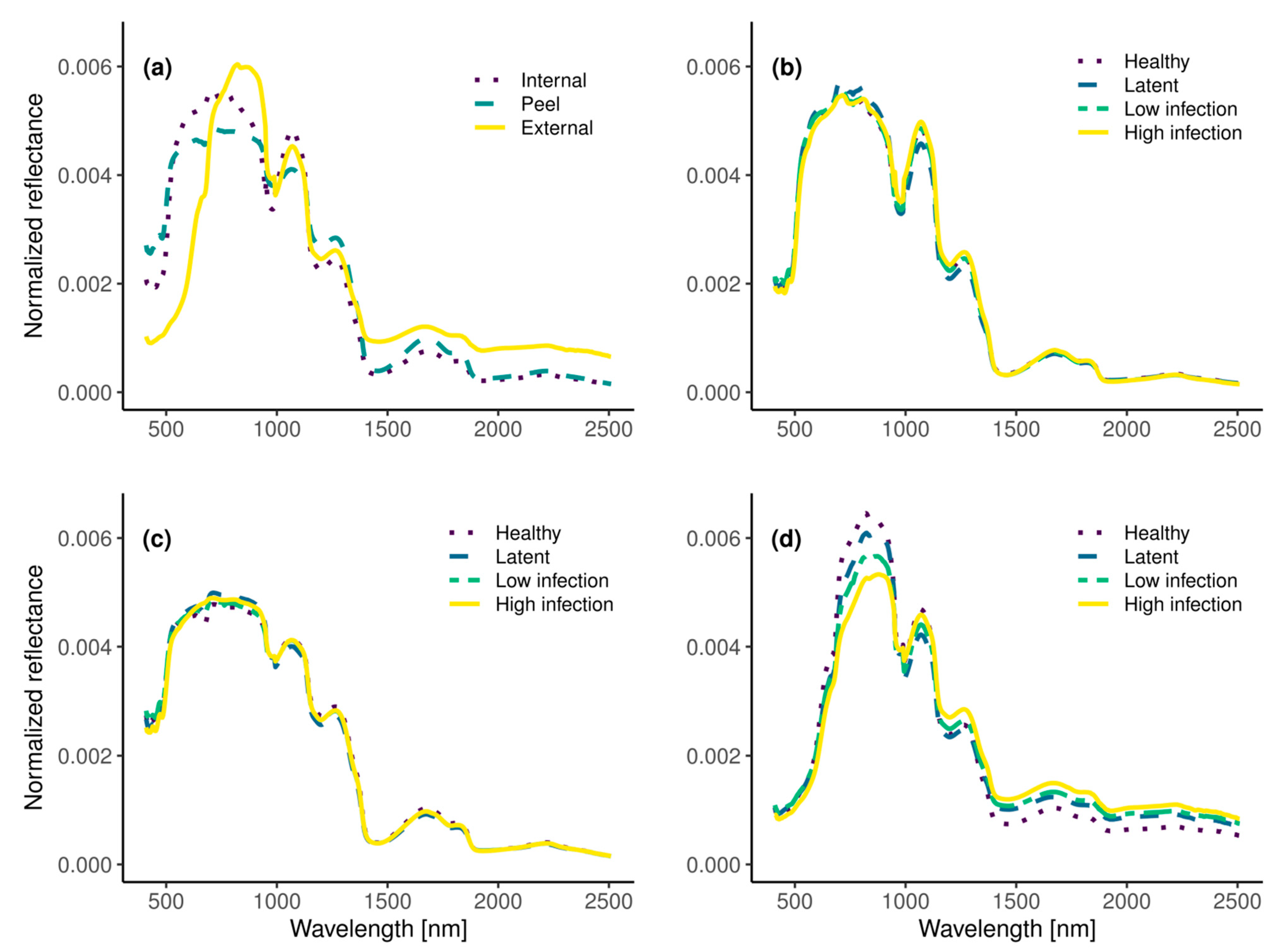
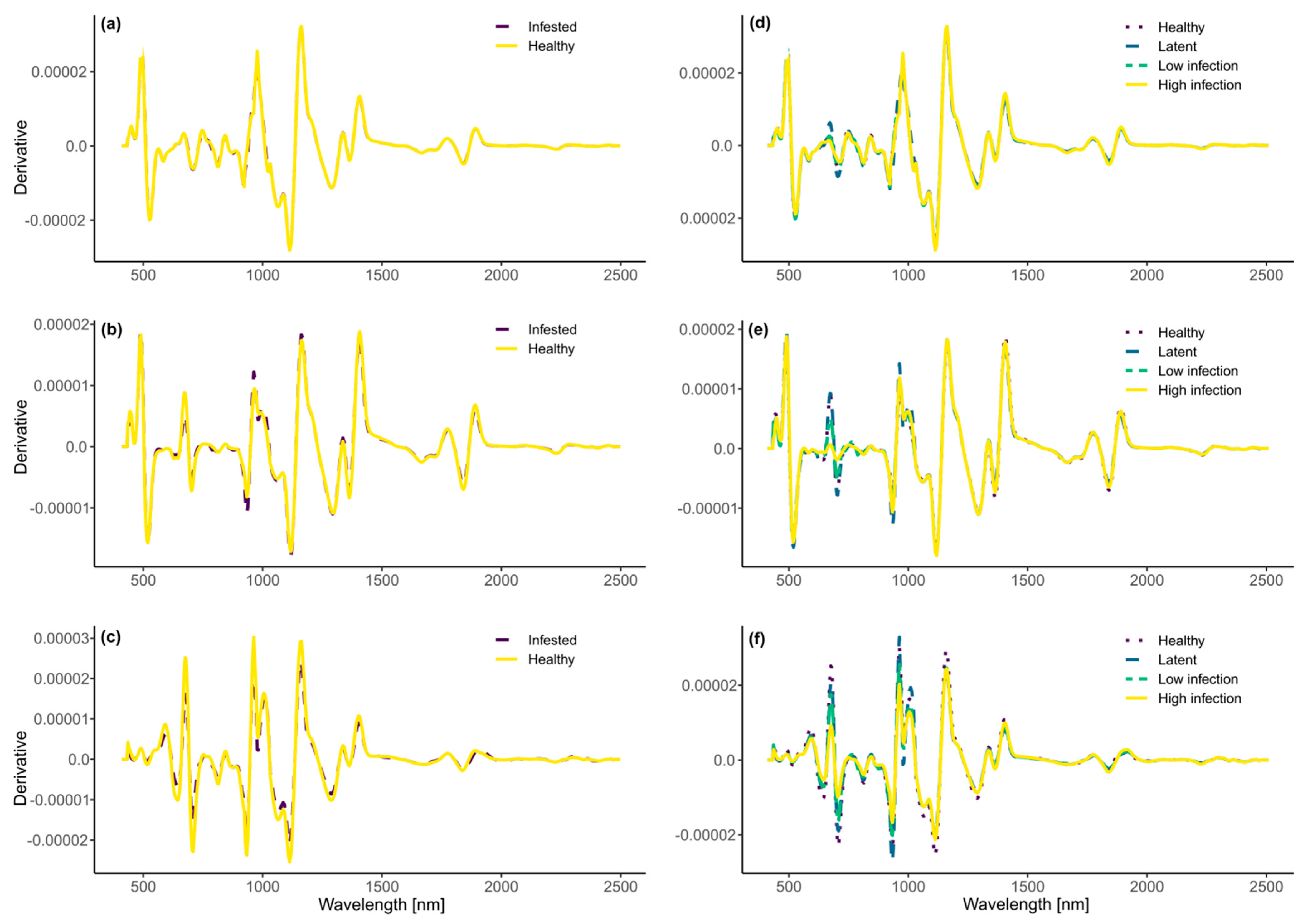
3.2. Remote Detection
3.3. Molecular Detection
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- EPPO Global Database: Meloidogyne chitwoodi. EPPO Datasheets on Pests Recommended for Regulation. Available online: https://gd.eppo.int (accessed on 9 April 2021).
- Maleita, C.; Esteves, I.; Cardoso, J.M.S.; Cunha, M.J.; Carneiro, R.M.D.G.; Abrantes, I. Meloidogyne luci, a new root-knot nematode parasitizing potato in Portugal. Plant Pathol. 2018, 67, 366–376. [Google Scholar] [CrossRef]
- Carneiro, R.M.D.G.; Correa, V.R.; Almeida, M.R.A.; Gomes, A.C.M.M.; Deimi, A.M.; Castagnone-Sereno, P.; Karssen, G. Meloidogyne luci n. sp. (Nematoda: Meloidogynidae), a root-knot nematode parasitising different crops in Brazil, Chile and Iran. Nematology 2014, 16, 289–301. [Google Scholar] [CrossRef]
- Gerič Stare, B.; Strajnar, P.; Susič, N.; Urek, G.; Širca, S. Reported populations of Meloidogyne ethiopica in Europe identified as Meloidogyne luci. Plant Dis. 2017, 101, 1627–1632. [Google Scholar] [CrossRef] [PubMed]
- Širca, S.; Urek, G.; Karssen, G. First report of the root-knot nematode Meloidogyne ethiopica on tomato in Slovenia. Plant Dis. 2004, 88, 680. [Google Scholar] [CrossRef]
- Gerič, S.B.; Strajnar, P.; Širca, S.; Susič, N.; Urek, G. Record of a new location for tropical root knot nematode Meloidogyne luci in Slovenia. Bull. OEPP/EPPO Bull. 2018, 48, 135–137. [Google Scholar] [CrossRef]
- Maleita, C.M.; José, S.M.; Egas, C.; Curtis, R.H.C.; Abrantes, I.M.O. Biometrical, biochemical, and molecular diagnosis of Portuguese Meloidogyne hispanica isolates. Plant Dis. 2012, 96, 865–874. [Google Scholar] [CrossRef]
- Conceição, I.L.; Tzortzakakis, E.A.; Gomes, P.; Abrantes, I.; Cunha, M.J. Detection of the root-knot nematode Meloidogyne ethiopica in Greece. Eur. J. Plant Pathol. 2012, 134, 451–457. [Google Scholar] [CrossRef]
- Santos, D.; Correia, A.; Abrantes, I.; Maleita, C. New hosts and records in Portugal for the root-knot nematode Meloidogyne luci. J. Nematol. 2019, 51, 1–4. [Google Scholar] [CrossRef]
- Rusinque, L.; Nóbrega, F.; Cordeiro, L.; Serra, C.; Inácio, M.L. First detection of Meloidogyne luci (Nematoda: Meloidogynidae) parasitizing potato in the Azores, Portugal. Plants 2021, 10, 99. [Google Scholar] [CrossRef]
- Strajnar, P.; Širca, S.; Knapič, M.; Urek, G. Effect of Slovenian climatic conditions on the development and survival of the root-knot nematode Meloidogyne Ethiopica. Eur. J. Plant Pathol. 2011, 129, 81–88. [Google Scholar] [CrossRef]
- EPPO Alert List: Addition of Meloidogyne luci together with M. ethiopica; EPPO Reporting Service No. 11-2017, Num. Article: 2017/218. Available online: https://gd.eppo.int/reporting/article-6186 (accessed on 2 March 2021).
- Cunha, G.T.; Visôtto, L.I.; Lopes, E.A.; Oliveira, C.M.G.; God, P.I.V.G. Diagnostic methods for identification of root-knot nematodes species from Brazil. Crop Prot. 2018, 48, 2. [Google Scholar] [CrossRef]
- Elmasry, G.; Kamruzzaman, M.; Sun, D.W.; Allen, P. Principles and applications of hyperspectral imaging in quality evaluation of agro-food products: A review. Crit. Rev. Food Sci. Nutr. 2012, 52, 999–1023. [Google Scholar] [CrossRef]
- Brugger, A.; Behmann, J.; Paulus, S.; Luigs, H.G.; Kuska, M.T.; Schramowski, P.; Kersting, K.; Steiner, U.; Mahlein, A.K. Extending hyperspectral imaging for plant phenotyping to the UV-range. Remote Sens. 2019, 11, 1401. [Google Scholar] [CrossRef]
- Voigt, C.A. Callose-mediated resistance to pathogenic intruders in plant defence-related papillae. Front. Plant Sci. 2014, 5, 168. [Google Scholar] [CrossRef] [PubMed]
- Susič, N.; Žibrat, U.; Širca, S.; Strajnar, P.; Razinger, J.; Knapič, M.; Vončina, A.; Urek, G.; Gerič Stare, B. Discrimination between abiotic and biotic drought stress in tomatoes using hyperspectral imaging. Sens. Actuators B Chem. 2018, 273, 842–852. [Google Scholar] [CrossRef]
- Susič, N.; Žibrat, U.; Sinkovič, L.; Vončina, A.; Razinger, J.; Knapič, M.; Sedlar, A.; Širca, S.; Gerič, S.B. From genome to field—Observation of the multimodal nematicidal and plant growth-promoting effects of Bacillus firmus I-1582 on tomatoes using hyperspectral remote sensing. Plants 2020, 9, 592. [Google Scholar] [CrossRef] [PubMed]
- Jensen, J.R. Remote Sensing of the Environment: An Earth Resource Perspective, 2nd ed.; Pearson Prentice Hall: Upper Saddle River, NJ, USA, 2007; p. 592. [Google Scholar]
- Sankaran, S.; Mishra, A.; Ehsani, R.; Davis, C. A review of advanced techniques for detecting plant diseases. Comput. Electron. Agric. 2010, 72, 1–13. [Google Scholar] [CrossRef]
- Mahlein, A.K.; Oerke, E.C.; Steiner, U.; Dehne, H.W. Recent advances in sensing plant diseases for precision crop protection. Eur. J. Plant Pathol. 2012, 133, 197–209. [Google Scholar] [CrossRef]
- Ji, Y.; Sun, L.; Li, Y.; Li, J.; Liu, S.; Xie, X.X.Y. Non-destructive classification of defective potatoes based on hyperspectral imaging and support vector machine. Infrared Phys. Technol. 2019, 99, 71–79. [Google Scholar] [CrossRef]
- Lopez, M.A.; Keresztes, J.C.; Goodarzi, M.; Arazuri, A.; Jaren, C.; Saeys, W. Non-destructive detection of blackspot in potatoes by Vis-NIR and SWIR hyperspectralimaging. Food Control 2016, 70, 229–241. [Google Scholar] [CrossRef]
- Amjad, W.; Crichton, S.O.J.; Munir, A.; Hensel, O.; Sturm, B. Hyperspectral imaging for the determination of potato slice moisture content and chromaticity during the convective hot air drying process. Biosyst. Eng. 2018, 166, 170–182. [Google Scholar] [CrossRef]
- Ji, Y.; Sun, L.; Li, Y.; Ye, D. Detection of bruised potatoes using hyperspectral imaging technique based on discrete wavelet transform. Infrared Phys. Technol. 2019, 103, 103054. [Google Scholar] [CrossRef]
- Su, W.H.; Bakalis, S.; Sun, D.W. Fourier transform mid-infrared-attenuated total reflectance (FTMIR-ATR) microspectroscopy for determining textural property ofmicrowave baked tuber. J. Food Eng. 2018, 218, 1–13. [Google Scholar] [CrossRef]
- Su, W.H.; Sun, D.W. Chemical imaging for measuring the time series variations of tuber dry matter and starch concentration. Comput. Electron. Agric. 2017, 140, 361–373. [Google Scholar] [CrossRef]
- Ayvaz, H.; Santos, A.M.; Moyseenko, J.; Kleinhenz, M.; Rodriguez-Saona, L.E. Application of a portable infrared instrument for simultaneous analysis of sugars, asparagine and glutamine levels in raw potato tubers. Plant Foods Hum. Nutr. 2015, 70, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Dacal-Nieto, A.; Formella, A.; Carrión, P.; Vazquez-Fernandez, E.; Fernández-Delgado, M. Non–Destructive Detection of Hollow Heart in Potatoes Using Hyperspectral Imaging. In Proceedings of the Computer Analysis of Images and Patterns; Real, P., Diaz-Pernil, D., Molina-Abril, H., Berciano, A., Kropatsch, W., Eds.; Springer: Berlin/Heidelberg, Germany, 2011; pp. 180–187. [Google Scholar] [CrossRef]
- Huang, T.; Li, X.Y.; Xu, M.L.; Jin, R.; Ku, J.; Xu, S.M. Non-destructive detection research for hollow heart of potato based on semi-transmission hyperspectral imaging and SVM. Spectrosc. Spect. Anal. 2015, 35, 198–202. [Google Scholar] [CrossRef]
- Riza, D.F.A.; Suzuki, T.; Ogawa, Y.; Kondo, N. Diffuse reflectance characteristic of potato surface for external defects discrimination. Postharvest Biol. Technol. 2017, 133, 12–19. [Google Scholar] [CrossRef]
- Santos, D.; Martins da Silva, P.; Abrantes, I.; Maleita, C. Tomato Mi-1.2 gene confers resistance to Meloidogyne luci and M. ethiopica. Eur. J. Plant Pathol. 2020, 156, 571–580. [Google Scholar] [CrossRef]
- Žibrat, U.; Širca, S.; Susič, N.; Knapič, M.; Gerič Stare, B.; Urek, G. Noninvasive detection of plant parasitic nematodes using hyperspectral and other remote sensing systems. In Hyperspectral Remote Sensing: Theory and Applications; Earth Observation, Series; Pandey, P.C., Srivastava, P.K., Balzter, H., Bhattacharya, B., Petropoulos, G.P., Eds.; Elsevier: Amsterdam, The Netherlands, 2020; pp. 357–375. [Google Scholar] [CrossRef]
- Janssen, T.; Karssen, G.; Verhaeven, M.; Coyne, D.; Bert, W. Mitochondrial coding genome analysis of tropical root-knot nematodes (Meloidogyne) supports haplotype based diagnostics and reveals evidence of recent reticulate evolution. Sci. Rep. 2016, 6, 1–13. [Google Scholar] [CrossRef]
- Correa, V.R.; Mattos, V.S.; Almeida, M.R.A.; Santos, M.F.A.; Tigano, M.S.; Castagnone-Sereno, P.; Carneiro, R.M.D.G. Genetic diversity of the root-knot nematode Meloidogyne ethiopica and development of a species-specific SCAR marker for its diagnosis. Plant Pathol. 2014, 63, 476–483. [Google Scholar] [CrossRef]
- Santos, M.F.A.; Mattos, V.S.; Monteiro, J.M.S.; Almeida, M.R.A.; Jorge, A.S.; Cares, J.E.; Castagnone-Sereno, P.; Coyne, D.; Carneiro, R.M.D.G. Diversity of Meloidogyne spp. from peri-urban areas of sub-Saharan Africa and their genetic similarity with populations from the Latin America. Physiol. Mol. Plant Pathol. 2019, 105, 110–118. [Google Scholar] [CrossRef]
- Gerič Stare, B.; Aydınlı, G.; Devran, Z.; Mennan, S.; Strajnar, P.; Urek, G.; Širca, S. Recognition of species belonging to Meloidogyne ethiopica group and development of a diagnostic method for its detection. Eur. J. Plant Pathol. 2019, 154, 621–633. [Google Scholar] [CrossRef]
- Hussey, R.S.; Barker, K.R. Comparison of methods of collecting inocula for Meloidogyne spp., including a new technique. Plant Dis. Rep. 1973, 57, 1025–1028. [Google Scholar]
- Piqueras, S.; Burger, J.; Tauler, R.; Juan, A. Relevant aspects of quantification and sample heterogeneity in hyperspectral image resolution. Chemometr. Intell. Lab. Syst. 2012, 117, 169–182. [Google Scholar] [CrossRef]
- Žibrat, U.; Susič, N.; Knapič, M.; Širca, S.; Strajnar, P.; Razinger, J.; Vončina, A.; Urek, G.; Gerič Stare, B. Pipeline for imaging, extraction, pre-processing, and processing of time-series hyperspectral data for discriminating drought stress origin in tomatoes. MethodsX 2019, 6, 300–408. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.I. An information-theoretic approach to spectral variability, similarity, and discrimination for hyperspectral image analysis. IEEE Trans. Inf. Theory 2000, 46, 1927–1932. [Google Scholar] [CrossRef]
- Maaten, L.; Hinton, G.E. Visualizing data using t-SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
- Ballabio, D.; Consonni, V. Classification tools in chemistry. Part 1: Linear models. Anal. Methods 2013, 5, 3790–3798. [Google Scholar] [CrossRef]
- Chong, I.G.; Jun, C.H. Performance of some variable selection methods when multicollinearity is present. Chemom. Intell. Lab. Syst. 2005, 78, 103–112. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2015. [Google Scholar]
- Rohart, F.; Gautier, B.; Singh, A.; Le Cao, K.A. mixOmics: An R package for ’omics feature selection and multiple data integration. PLoS Comput. Biol. 2017, 13. [Google Scholar] [CrossRef]
- Kuhn, M. Caret: Classification and Regression Training; (Version 6.0.86); R. Astrophysics Source Code Library. CRAN: 2015. Available online: https://CRAN.R-project.org/package=caret (accessed on 30 March 2021).
- Donaldson, J. tsne: T-Distributed Stochastic Neighbor Embedding for R (t-SNE); (Version 0.1.3); R. CRAN: 2016. Available online: https://CRAN.R-project.org/package=tsne (accessed on 30 March 2021).
- Susič, N.; Koutsovoulos, G.D.; Riccio, C.; Danchin, E.G.J.; Blaxter, M.L.; Lunt, D.H.; Strajnar, P.; Širca, S.; Urek, G.; Stare, B.G. Genome sequence of the root-knot nematode Meloidogyne luci. J. Nematol. 2020, 52, 1–5. [Google Scholar] [CrossRef]
- Jones, J.T.; Haegeman, A.; Danchin, E.G.J.; Gaur, H.S.; Helder, J.; Jones, M.G.K.; Kikuchi, T.; Manzanilla-López, R.; Palomares-Rius, J.E.; Wesemael, W.M.L.; et al. Top 10 plant-parasitic nematodes in molecular plant pathology. Mol. Plant Pathol. 2013, 14, 946–961. [Google Scholar] [CrossRef]
- Lima, F.S.O.; Mattos, S.V.; Silva, S.E.; Carvalho, M.A.S.; Teixeira, R.A.; Silva, J.C.; Correa, V.R. Nematodes Affecting Potato and Sustainable Practices for Their Management. In Potato—From Incas to All over the World, 1st ed.; Yildiz, M., Ed.; IntechOpen: London, UK, 2018; Volume 1, pp. 107–121. [Google Scholar]
- EPPO. PM 3/69 (2) Meloidogyne chitwoodi and M. fallax sampling potato tubers for detection. Bull. OEPP/EPPO Bull. 2019, 49, 486–487. [Google Scholar] [CrossRef]
- De Haan, E.G.; Dekker, C.C.E.M.; Tameling, W.I.L.; den Nijs, L.J.M.F.; van den Bovenkamp, G.W.; Kooman-Gersmann, M. The MeloTuber test: A real-time TaqMan® PCR-based assay to detect the root-knot nematodes Meloidogyne chitwoodi and M. fallax directly in potato tubers. Bull. OEPP/EPPO Bull. 2014, 44, 166–175. [Google Scholar] [CrossRef]
- Viaene, N.; Mahieu, T.; Peña, E. Distribution of Meloidogyne chitwoodi in potato tubers and comparison of extraction methods. Nematology 2007, 9, 143–150. [Google Scholar] [CrossRef]
- Finley, A.M. Histopathology of Meloidogyne chitwoodi (Golden et al.) on Russet Burbank potato. J. Nematol. 1981, 13, 483. [Google Scholar]
- Workman, J., Jr.; Weyer, L. Practical Guide and Spectral Atlas for Interpretative Near-Infrared Spectroscopy, 2nd ed.; CRC Press: New York, NY, USA, 2012; pp. 231–249. ISBN 9781439875254. [Google Scholar]
- Abd-Elgawad, M.M.M.; Kerlan, M.-C.; Molinari, S.; Abd-El-Kareem, F.; Kabeil, S.S.A.; Mohamad, M.M.; El-Nagdi, W.A. Histopathological changes and enzymatic activities induced by Meloidogyne incognita on resistant and susceptible potato. J. Plant Pathol. 2012, 1, 62–72. [Google Scholar] [CrossRef]
- Colović, M.B.; Krstić, D.Z.; Lazarević-Pašti, T.D.; Bondžić, A.M.; Vasić, V.M. Acetylcholinesterase inhibitors: Pharmacology and toxicology. Curr. Neuropharmacol. 2013, 11, 315–335. [Google Scholar] [CrossRef]
- Mandel, F. Tomato glycoalkaloids: Role in the plant and in the diet. J. Agric. Food Chem. 2002, 50, 5751–5780. [Google Scholar] [CrossRef]


| Sample Type | Biological Replicates | Technical Replicates DNA Isolation | Technical Replicates Real-Time PCR | CT |
|---|---|---|---|---|
| 50 peels from the RKN treatment—tubers with visible symptoms and 50 peels from the uninfested (NC) treatment | 1 | 1 | 1 | 23.0 |
| 1 | 2 a | 1 | 26.1 | |
| 1 | 3 b | 1 | 25.0 | |
| 100 peels from the RKN treatment—tubers without visible symptoms (latent infestation) | 1 | 1 | 1 | 24.5 |
| 1 | 2 a | 1 | 29.2 | |
| 1 | 3 b | 1 | 26.7 | |
| 2 | 1 | 1 | 25.4 | |
| 2 | 2 | 1 | 22.5 | |
| 2 | 3 | 1 | 24.0 | |
| 99 peels from the uninfested (NC) treatment and one peel from the RKN treatment, one tuber with visible symptoms | 1 | 1 | 1 | 28.6 |
| 1 | 2 a | 1 | 31.1 | |
| 1 | 3 b | 1 | 30.9 | |
| 2 | 1 | 1 | 28.4 | |
| 2 | 1 | 2 | 32.4 | |
| 2 | 1 | 3 | 32.5 | |
| 2 | 2 | 1 | 31.0 | |
| 2 | 3 | 1 | 28.1 | |
| 3 | 1 | 1 | 28.6 | |
| 3 | 2 | 1 | 28.3 | |
| 3 | 3 | 1 | 28.5 | |
| 100 peels from the uninfested (NC) treatment with addition of three RKN females | 1 | 1 | 1 | 29.6 |
| 1 | 2 a | 1 | 32.0 | |
| 1 | 3 b | 1 | 32.3 | |
| 100 peels from the uninfested (NC) treatment with addition of one RKN female | 1 | 1 | 1 | und. |
| 1 | 2 a | 1 | 34.2 | |
| 1 | 3 b | 1 | und. | |
| 2 | 1 | 1 | und. | |
| 2 | 2 a | 1 | und. | |
| 2 | 3 b | 1 | und. | |
| 100 peels from the uninfested (NC) treatment | 1 | 1 | 1 | und. |
| 1 | 2 a | 1 | und. | |
| 1 | 3 b | 1 | und. | |
| 2 | 1 | 1 | und. | |
| 2 | 2 | 1 | und. | |
| 2 | 3 | 1 | und. | |
| 3 | 1 | 1 | und. | |
| 3 | 2 | 1 | und. | |
| 3 | 3 | 1 | und. |
| Group | Treatment | N | PLS-DA | PLS-SVM | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| LV | Var (%) | RMSECV | c | Gamma | Accuracy | CL | Kappa | |||
| M. luci | External | 59 | 10 | 75.4 | 0.71 | 0.77 | 0.1 | 0.983 | 0.909–0.999 | 0.975 |
| Internal | 59 | 10 | 81.9 | 0.75 | 0.527 | 0.046 | 0.983 | 0.909–0.999 | 0.975 | |
| Peeled | 59 | 10 | 68.0 | 0.61 | 1 | 0.1 | 0.983 | 0.909–0.999 | 0.975 | |
| M. fallax | External | 65 | 5 | 80.5 | 0.8 | 1 | 0.032 | 1 | 0.945–1 | 1 |
| Internal | 65 | 6 | 64.8 | 0.81 | 10 | 0.01 | 1 | 0.945–1 | 1 | |
| Peeled | 65 | 9 | 78.3 | 0.76 | 0.599 | 0.017 | 0.985 | 0.917–0.999 | 0.973 | |
| Predicted | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Group | Lt | Lo | Hi | He | Sensitivity | Specificity | Precision | Recall | F1 | Balanced Accuracy | |
| Lt | 10 | 1 | 1 | 0.98 | 0.91 | 1 | 0.952 | 0.99 | |||
| Lo | 11 | 0.917 | 1 | 1 | 0.917 | 0.957 | 0.958 | ||||
| Ex | Hi | 10 | 1 | 1 | 1 | 1 | 1 | 1 | |||
| He | 27 | 1 | 1 | 1 | 1 | 1 | 1 | ||||
| Lt | 11 | 1 | 1 | 1 | 1 | 1 | 1 | ||||
| Lo | 10 | 1 | 1 | 0.98 | 0.909 | 1 | 0.952 | 0.99 | |||
| In | Hi | 10 | 0.909 | 1 | 1 | 0.909 | 0.952 | 0.95 | |||
| He | 27 | 1 | 1 | 1 | 1 | 1 | 1 | ||||
| Lt | 11 | 1 | 1 | 1 | 1 | 1 | 1 | ||||
| Pe | Lo | 11 | 0.917 | 1 | 1 | 0.917 | 0.957 | 0.958 | |||
| Hi | 1 | 9 | 1 | 0.98 | 0.9 | 1 | 0.947 | 0.99 | |||
| He | 27 | 1 | 1 | 1 | 1 | 1 | 1 | ||||
| Mf | Ml | He | Sensitivity | Specificity | Precision | Recall | F1 | Balanced Accuracy | |||
| Mf | 6 | 1 | 1 | 1 | 1 | 1 | 1 | ||||
| Ex | Ml | 32 | 1 | 1 | 1 | 1 | 1 | 1 | |||
| He | 27 | 1 | 1 | 1 | 1 | 1 | 1 | ||||
| Mf | 6 | 1 | 1 | 1 | 1 | 1 | 1 | ||||
| In | Ml | 32 | 1 | 1 | 1 | 1 | 1 | 1 | |||
| He | 27 | 1 | 1 | 1 | 1 | 1 | 1 | ||||
| Mf | 6 | 1 | 1 | 1 | 1 | 1 | 1 | ||||
| Pe | Ml | 31 | 1 | 1 | 0.971 | 0.969 | 1 | 0.984 | 0.985 | ||
| He | 27 | 0.964 | 1 | 1 | 0.964 | 0.982 | 0.982 | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Žibrat, U.; Gerič Stare, B.; Knapič, M.; Susič, N.; Lapajne, J.; Širca, S. Detection of Root-Knot Nematode Meloidogyne luci Infestation of Potato Tubers Using Hyperspectral Remote Sensing and Real-Time PCR Molecular Methods. Remote Sens. 2021, 13, 1996. https://doi.org/10.3390/rs13101996
Žibrat U, Gerič Stare B, Knapič M, Susič N, Lapajne J, Širca S. Detection of Root-Knot Nematode Meloidogyne luci Infestation of Potato Tubers Using Hyperspectral Remote Sensing and Real-Time PCR Molecular Methods. Remote Sensing. 2021; 13(10):1996. https://doi.org/10.3390/rs13101996
Chicago/Turabian StyleŽibrat, Uroš, Barbara Gerič Stare, Matej Knapič, Nik Susič, Janez Lapajne, and Saša Širca. 2021. "Detection of Root-Knot Nematode Meloidogyne luci Infestation of Potato Tubers Using Hyperspectral Remote Sensing and Real-Time PCR Molecular Methods" Remote Sensing 13, no. 10: 1996. https://doi.org/10.3390/rs13101996
APA StyleŽibrat, U., Gerič Stare, B., Knapič, M., Susič, N., Lapajne, J., & Širca, S. (2021). Detection of Root-Knot Nematode Meloidogyne luci Infestation of Potato Tubers Using Hyperspectral Remote Sensing and Real-Time PCR Molecular Methods. Remote Sensing, 13(10), 1996. https://doi.org/10.3390/rs13101996