1. Introduction
Plant disease epidemics cause substantial economic losses in agricultural settings worldwide [
1]. Monitoring plant health and detecting plant diseases at early growth stages are essential to control disease spread and facilitate sustainable, environment-friendly, and cost-effective management practices [
2] in growers’ fields. This paper focuses on the potential of using satellite imagery to detect soybean sudden death syndrome (SDS), a disease of significant economic importance in North and South America [
3,
4,
5]. The disease is caused by
Fusarium virguliforme (
Fv), a soilborne fungal pathogen [
6], and is widely distributed across 23 U.S. states, including those accounting for most U.S. soybean production. During the years from 2006 to 2014, annual yield losses in the U.S. due to SDS were estimated at between 0.6 and 1.9 million metric tons [
7], representing 200 to 750 million dollars in monetary losses [
8].
The pathogen starts infecting roots during early soybean growth stages [
9,
10] and causes root rot and poor root development [
3]. Root infections are favored by cool, wet soil environments [
11,
12]. Foliar symptoms of interveinal chlorosis (yellowing between leaf veins) and necrosis (tissue browning following cell death) typically appear during reproductive stages [
13] and cause premature defoliation and senescence under severe disease pressure [
14]. The initial foliar symptoms show only as yellow traces on lower leaves, which makes the disease difficult to detect at early stages. Abundant soil moisture favors SDS foliar symptom expression [
12,
15], whereas infected plants may not develop foliar symptoms under dry field conditions. Disease distribution within a field is limited by the spatial distribution of the pathogen at the beginning of the growing season.
Scouting for SDS foliar symptoms in the field is made difficult by the relatively late onset of visible foliar symptom expression, which often occurs after the soybean canopy has closed, and by the patchy distribution of SDS in soybean fields. Scouting for symptomatic plants is time-consuming, and confirmation of
Fv infection requires destructive sampling [
16]. Therefore, a more effective method for monitoring and quantifying the distribution of SDS in the field is needed.
Early detection of plant diseases through remote sensing can be difficult when foliar symptoms are mild [
16], because multiple factors can contribute to the biophysical and chemical changes that are associated with plant disease [
17]. Plants with severe foliar symptoms differ significantly in canopy color from healthy or slightly infected plants [
18]. Thus, plants with highly visible symptoms can be more accurately detected than plants with mild symptoms [
19].
SDS causes foliar symptoms of chlorosis and necrosis. Canopy damage is the result of disease-induced changes in physiological functions, leaf pigments [
19], chlorophyll content, and biomass [
16]. Zwiggelaar [
20] stated that alteration in leaf pigment content results in changes in optical spectra, such as a decrease in canopy reflectance in the near-infrared band and an increase of reflectance in the red band [
21]. Both the red and NIR bands in the light spectrum respond to changes in chlorophyll pigment. These two bands are also highly sensitive to changes in leaf water stress [
22] and the total green leaf area [
23]. Reduced green leaf area may be present due to reduced plant growth during mid-season or due to early defoliation at the end of the season.
Leaf chlorophyll level is inversely related to spectral reflectance in the blue and red wavebands of visible light, and is dependent on the interactions between plant tissues and absorbed light [
24]. SDS-infected plants tend to show early chlorophyll deterioration [
16], resulting in spectral responses that increase reflectance in the blue and red bands while decreasing reflectance in the green and NIR regions. When plant diseases such as SDS produce necrotic or chlorotic symptoms on leaves, diseased plants show an overall increase in reflectance in the visible region, especially in the red band [
16,
25], and simultaneously decreased reflectance in the green and NIR regions. Reflectance in the NIR regions is affected primarily by leaf structure and canopy biomass, which are influenced by plant water and health status [
22,
25,
26,
27].
Recently, successful early detection of SDS in soybean leaves [
16,
25] and canopies [
25,
28] has been achieved using handheld, tractor-mounted, and unmanned aerial vehicle (UAV)-mounted remote sensing tools. Although these tools have proven to be promising for early detection of SDS, they still require considerable time, capital, and equipment investments for data collection. Also, these cutting-edge tools can be expensive and cumbersome for growers to operate and maintain in a commercial system. Data generated from these tools may lack spatial information, which must be obtained via additional geographic information system (GIS) and global positioning system (GPS) technologies before targeted and site-specific management applications can be implemented.
Satellite imagery, in comparison with ground-based and aerial remote sensing platforms, covers a wide ground swath and can provide high temporal resolution due to frequent revisit times. Consequently, there has been a growing interest in using high-resolution multispectral satellite images to monitor crop diseases [
29,
30,
31]. Image acquisition costs vary considerably depending on the provider and product specifications, but more low-cost or free image products are available now than in the past. For example, PlanetScope is a 4-band multispectral satellite that collects imagery at 3-m spatial resolution with 1- to 10-day temporal resolution (Planet Labs
https://www.planet.com/). PlanetScope ortho scenes are geo-rectified (i.e., processed to remove distortions caused by tilt and terrain), radiometrically and atmospherically corrected, and projected.
Different machine learning classification methods, such as support vector machines (SVM), artificial neural networks (ANNs), linear discriminant analysis (LDA), and random forest (RF) [
16,
25,
29,
32,
33,
34,
35], have been used for early detection of plant diseases based on remote sensing data. Random forest is a flexible and powerful machine learning classifier [
36] that has been utilized in the classification of remote-sensing-based information [
29,
37,
38,
39,
40,
41]. The random forest classifier can handle huge, multidimensional datasets and performs both classification and regression functions without over-fitting the model [
36,
42]. The random forest algorithm also evaluates the predictive importance of input features, such as reflectance wavebands in our study, hence supporting feature selection for subsequent analysis [
36].
Greater access to low-cost, multispectral, high-resolution satellite imagery provides opportunities for scientists to combine these large datasets with machine learning technologies [
27] to devise reliable automated systems for detecting and mapping plant diseases. These systems could support grower decisions for site-specific management applications, such as planting resistant soybean varieties and using diverse crop rotations [
43] in infested areas. They have the potential to significantly reduce harmful chemical seed treatment applications and the resulting economic expense and ecological impact on soybean production systems.
The objective of this study was to detect SDS using high-resolution satellite imagery collected over different vegetative and reproductive growth stages of the soybean crop in a long-term crop rotation field trial, with a history of rotation-influenced differences in SDS symptom intensity [
44,
45]. We hypothesized that spectral data based on blue, green, red, near-infrared (NIR) spectral bands, and the calculated normalized difference vegetation index (NDVI), combined with ground-based crop management information, would allow detection of SDS in larger field-scale plots. Specifically, we wished to investigate the potential of using:
- (1)
high-resolution satellite images to assess soybean crop health at the canopy scale, which would permit inspection of larger field areas.
- (2)
RF algorithms to classify disease status using image data from single dates collected over a two-month period, to see if and when subtle changes in canopy reflectance properties not associated with typical foliar symptoms would permit classification of crop disease status.
- (3)
knowledge of field areas likely to exhibit differences in disease intensity.
2. Materials and Methods
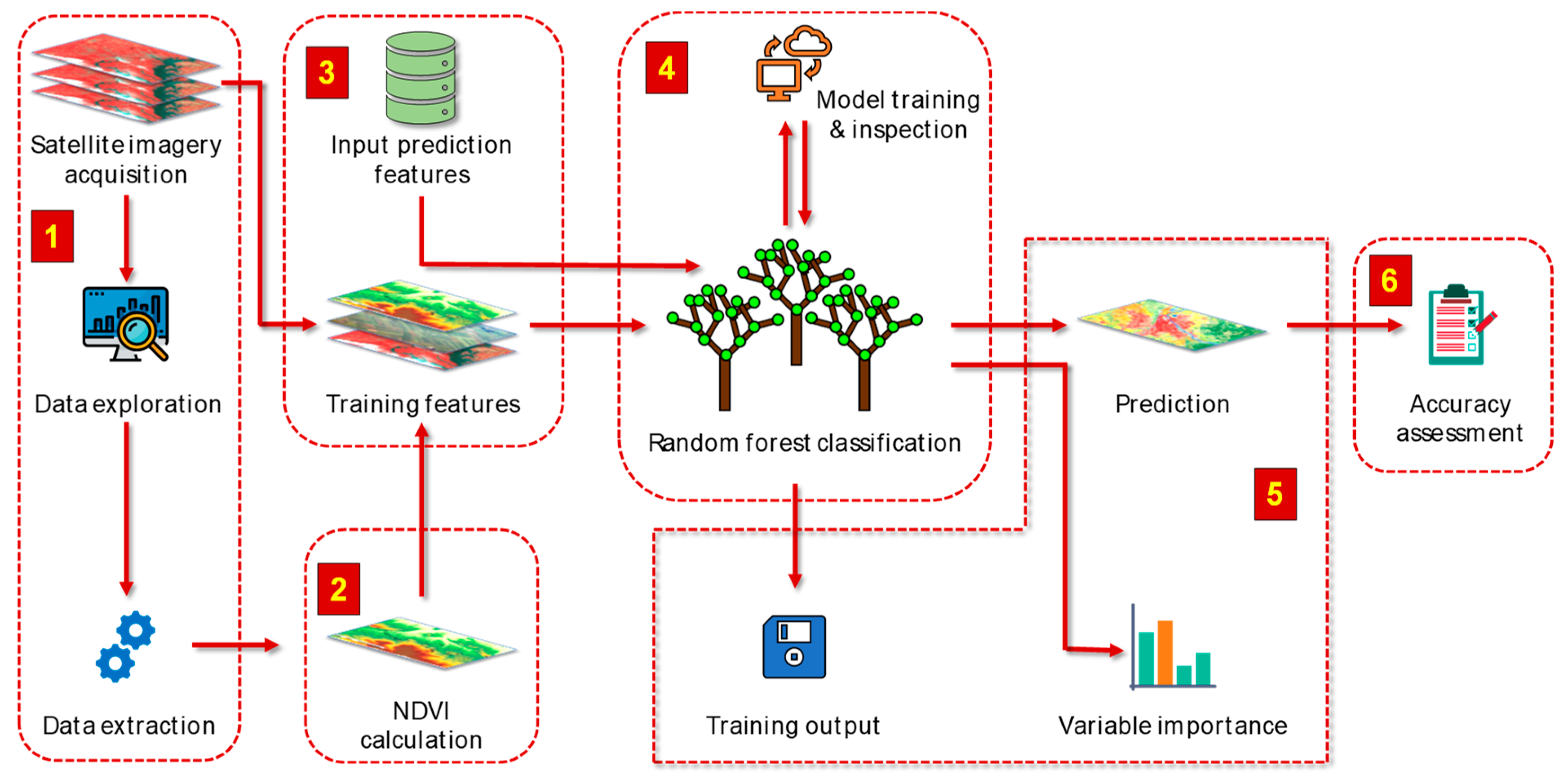
We used 4-band PlanetScope multispectral imagery for the 2016 to 2018 soybean growing seasons, along with crop rotation information, and analyzed the data using the random forest algorithm for SDS detection. Our method for data processing and analysis for the detection SDS is illustrated in
Figure 1. The method encompasses six key components: (1) image acquisition and data extraction, (2) normalized difference vegetation index (NDVI) calculation, (3) exploration and preparation of input datasets, (4) training and inspection of the random forest model, (5) model classification of diseased or healthy status, and (6) assessment of classification accuracy.
2.1. Study Site
Data for this study were collected from an ongoing soybean field experiment located at Iowa State University’s Marsden Farm in Boone County, Iowa [
44,
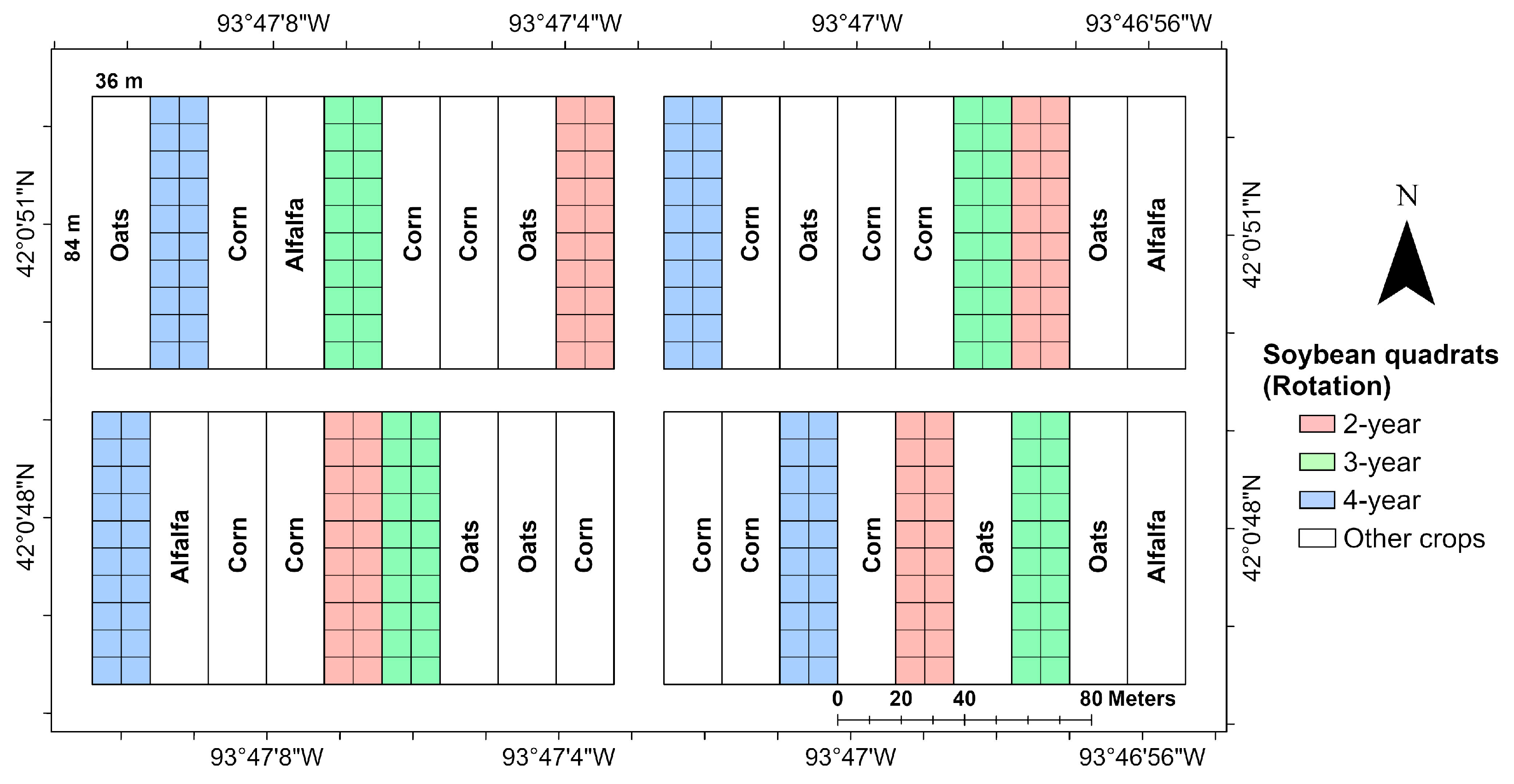
45]. This experiment was established in 2001 as a randomized complete block design (RCBD) with four blocks. Each block consists of nine main plots (18 m × 84 m), with two subplots (9 m × 84 m) within each main plot (
Figure 2).
Three crop rotation systems (2-year, 3-year, and 4-year) were applied to the main plots [
43]. The 2-year cropping system was planted with corn and soybean in alternate years and managed with extension-recommended practices and applications of synthetic fertilizers. In the 3-year cropping system, an oat-red clover forage mix was planted in the year after soybean and before corn. In the 4-year cropping system, oat and alfalfa were seeded in the 3rd year and followed by alfalfa in the 4th year. Crop nutrients for the 3-year and 4-year cropping systems were provided via applications of composted cattle manure and reduced rates of synthetic fertilizers. All crop phases of the three rotation systems were planted each year.
No artificial
Fv inoculum was added to the field plots in this experiment. However, in every year since 2010, the crop rotation treatments have been associated with differences in SDS intensity at this site [
15]. Therefore, this site was chosen to test using satellite imagery for SDS detection.
The soybean variety Latham L2758 R2, a maturity group 2.7 variety rated as resistant to SDS, was planted on 21 May in 2016, 15 May in 2017, and 17 May in 2018. Each soybean plot consisted of 24 rows, with row spacing of 0.76 m. Plots were subdivided into 20 quadrats, each roughly 8 × 9 m (
Figure 2). For visual disease assessment, rating areas (3 m wide × 1.5 m long) were flagged inside each quadrat area, 8.4 m apart in the outer six rows along both sides of the plot, resulting in 20 rating areas in each soybean main plot. Center coordinates for each quadrat were collected using a handheld Trimble GeoXT GPS Device (Trimble Inc., Sunnyvale, CA, USA). Each visual rating area spanned four soybean rows.
2.2. Disease Assessment
SDS foliar incidence, i.e., the percentage of plants in a quadrat with foliar symptoms [
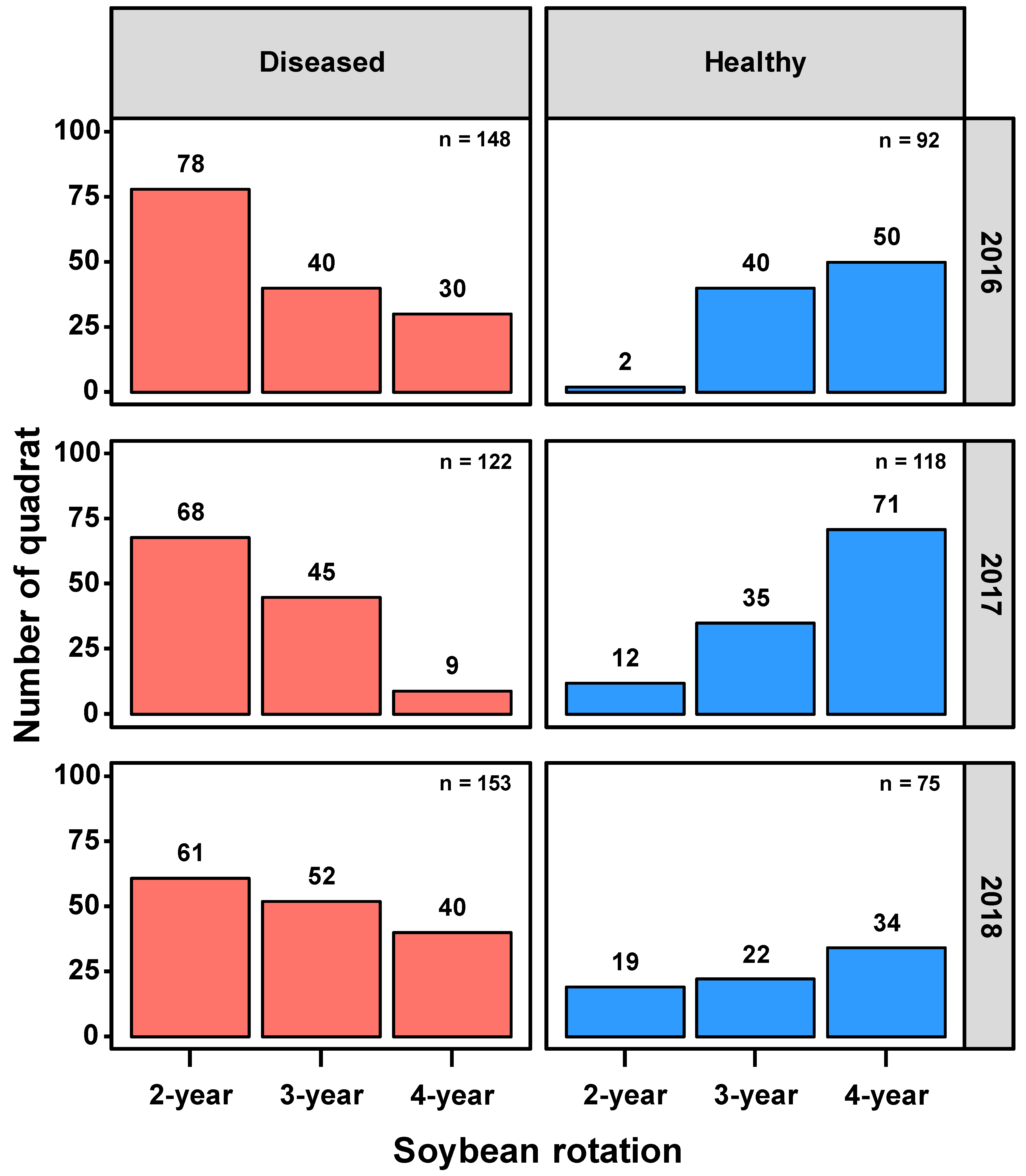
46], was assessed weekly in all soybean quadrats, from the time of initial foliar symptom onset in the trial until crop senescence (
Figure 3). For analysis, soybean quadrats were assigned to two main classes, i.e., healthy or diseased, based on foliar symptom incidence. Soybean quadrats with no foliar symptoms or less than 5% foliar disease incidence were classified as healthy quadrats, while quadrats showing more than 5% foliar incidence at the end of the season were classified as diseased quadrats. Disease was rated by a team of plant pathologists, including researchers who had been studying SDS at this site since 2010. All plants within the visual rating areas (1.5 m L × 3 m W) were examined. Disease was confirmed in the field by examining a few plants for internal stem discoloration (which distinguishes foliar symptoms caused by SDS from symptoms that may be caused by the brown stem rot pathogen,
Phialophora gregata) [
3], and by molecular detection of the pathogen in soybean roots that were collected for concurrent studies [
42] (L. Leandro, unpublished data). Plants were dug using hand trowels to minimize soil disturbance and loss of adjacent plants. Foliar symptoms were confirmed when disease symptoms in quadrats were noted for two or more consecutive rating dates. Plant stand data for the same quadrats were used to convert counts of diseased plants to percentages.
2.3. Imagery Acquisition
PlanetScope (PS) satellite imagery was acquired from Planet Labs Inc. (San Francisco, CA, USA), a private imaging company. PS satellites orbit at 400 km (51.6° inclination) altitude and can provide images with red, green, and blue (RGB) and near-infrared (NIR) data (
Table 1). The image product is delivered as a continuous, split-frame strip, with half-frames containing RGB and NIR imagery.
We utilized PS ortho scenes (Planet Labs image product Level 3B), which are orthorectified (i.e., processed to remove geometric distortions caused by tilt and terrain) and radiometrically and sensor corrected. The geometric correction was based on digital elevation models (DEMs) with post-spacing between 30 and 90 m, with an error specification of < 10 Root Mean Square Error (RMSE). Ortho scenes were projected to the universal transverse mercator (UTM) WGS84 cartographic projection prior to product delivery. The PS ortho tile product included a single strip of multiple orthorectified scenes that were merged and divided according to a defined grid. The 3-m resolution ortho scenes were obtained for the months of July and August in all years. We obtained 4, 15, and 18 cloud-free ortho scenes in 2016, 2017, and 2018, respectively. All 2016 images were created by generation 0e (A) satellites and all 2018 images by generation 0f or 10 (B) satellites. A and B have different spectral response curves. Beginning with 20-Jul-2017, the 2017 images switched from A to B, with the sole exception of 23-Aug-2017, which again used A.
Image alignment was checked visually in ArcGIS and compared to ground-based GIS reference points. Differences in image brightness and color—due to differences in satellite sensor hardware, acquisition time, sun angle, cloud conditions, and dates—are likely to be present, and were noted in the images we used. Because our intent was to evaluate a method for analyzing data from single dates, we did not standardize image brightness during pre-processing.
2.4. Data Extraction
Soybean plot locations varied from year to year and followed crop rotation sequences assigned to main plots. Quadrat locations for each year were described by shapefiles, which included crop rotation information (2-year, 3-year, or 4-year) and quadrat health status at the end of the season (SDS incidence reached 5% or higher, or plants remained healthy). For data extraction from satellite images and subsequent analysis, soybean quadrats were generalized to large quadrats (8.6 m W × 9.1 m L) (
Figure 2). Via the ArcPy Python Application Programming Interface (API) [
47], the ArcGIS Zonal statistics tool was used to calculate the mean values of all image pixels that were covered by each quadrat; mean digital number (DN) values were calculated for each of the four bands (R, G, B, and NIR). This process was repeated for each of the images obtained for a given year, so that each quadrat polygon feature contained the crop rotation type, the mean R, G, B, and NIR values of the pixels covered by the polygon, a calculated NDVI value (explained below), and a binary value for SDS health status.
Preliminary inspection of the image data indicated that band 4 (NIR) mean DN values for the two disease classes were significantly different in 24 of 37 images, as determined by Wilcoxon’s rank test (two-tailed test, p ≤ 0.05) performed via PROC NPAR1WAY i.e., the procedure to perform Wilcoxon’s rank test in Statistical analysis system (SAS Institute, Cary, NC, USA). Similar tests for bands 1, 2, and 3 indicated 13, 10, and 17 images showed class differences when quadrat mean DN values were tested. Exploratory analyses detected class differences in more of the images when single pixel data were examined (data not shown). Crop rotation, R, G, B, NIR, and NDVI were then used as exploratory variables (corresponding to “features” in the random forest methodology), and SDS status was used as the “ground truth” response variable.
A practical example of the data extraction process (zonal statistics) via Python is given in the Jupyter Notebook “Prepare data for random forest classification.ipynb”, which can be found in the data repository:
https://doi.org/10.25380/iastate.11356430 [
48]. The repository also contains gis project data in a ESRI (Environmental Systems Research Institute) file geodatabase (.gdb); specifically a set of preprocessed quadrat polygons and images (cropped to cover just the area under investigation) for a selection of dates in 2016, 2017, and 2018. Finally, the repository contains an ArcGIS Pro project file (.aprx) with which the data can be visualized. See
supplementary materials for more information.
2.5. NDVI Calculation
Although several conventional and newly-modified vegetation indices based on the red-edge bands have been explored to evaluate crop health [
16,
25,
29,
32], NDVI [
49] is one of the most commonly used vegetation indices. To evaluate its suitability for detecting and discriminating healthy and diseased soybean quadrats, the NDVI was calculated for each quadrat using Equation (1):
2.6. Data Analysis
Random forest classification [
36] was performed using R and the Scikit-learn [
50] library in Python 3.7.3 [
51]. Random forest is a supervised machine learning algorithm that applies bootstrap aggregation (bagging) [
52,
53] and random feature selection [
54,
55]. Random forest is a robust, nonparametric method that builds a forest of many decision trees from a training dataset and tests the accuracy of the forest on a test dataset. Although individual decision trees may not be accurate, the combination of many trees significantly increases the robustness of the model and leads to more accurate predictions. When the test datasets are passed through the trained decision trees, each decision tree votes on an outcome. The forest model then aggregates the votes from all decision trees and predicts the final class based on the largest number of votes. In our case, the outcome is in the form of a binary variable (healthy, SDS = 0, or diseased, SDS = 1); thus, we performed a random forest classification [
36] rather than a regression.
2.6.1. Random Forest Model Training and Prediction
Data from each ortho scene were analyzed separately, beginning by splitting the data into training and test datasets (70% and 30% of the quadrats, respectively), followed by model training and classification of the test data. Model parameters included the number of trees in the forest (n_estimators), number of features to consider when looking for the best split (max_features), the maximum tree depth allowed (max_depth), and minimum number of samples required to be a leaf node (min_sample_leaf).
These parameters were tuned using the 5-fold cross-validation grid search method. Grid search is a simple strategy used to evaluate all possible combinations of given, discrete parameter spaces and derive their optimal combination for a tuned model. Using these tuned models, SDS predictions were made using the test set. This procedure was repeated for each ortho scene in 2016, 2017, and 2018.
The data repository
https://doi.org/10.25380/iastate.11356430 contains a Jupyter Notebook “Using Random Forest Models for SDS.ipynb”, which demonstrates the entire process of splitting the data into training and test datasets, tuning the random forest model on the training dataset, performing the predictions of the test dataset using the tuned model, and judging the prediction quality of the model on the quadrat data discussed earlier.
2.6.2. Variable Importance
We measured the importance of predictor variables via the permutation variable importance measure procedure [
56], which is based on random selection and index reordering. First, the random forest model calculates prediction accuracy in the out-of-bag (OOB) observations (observations from the training set that are left out of the bootstrap samples and not used to construct the decision trees), and then it randomly shuffles values of a predictor variable to break the association between response and predictor values and recalculate the accuracy in OOB observations. Then, it calculates the difference in model accuracy before and after shuffling [
57]. The average of this number over all trees in the forest represents the raw importance score for the variable. If the predictor never had any meaningful relationship with the response, shuffling its values will produce very little change in the model accuracy. However, if a predictor was strongly associated with the response, permutations should create a significant decrease in accuracy. Permutation of variable importance evaluates the individual impact, including interactions, of each tree predictor in the random forest model. The procedure was used to evaluate all variables.
2.6.3. Accuracy Assessment
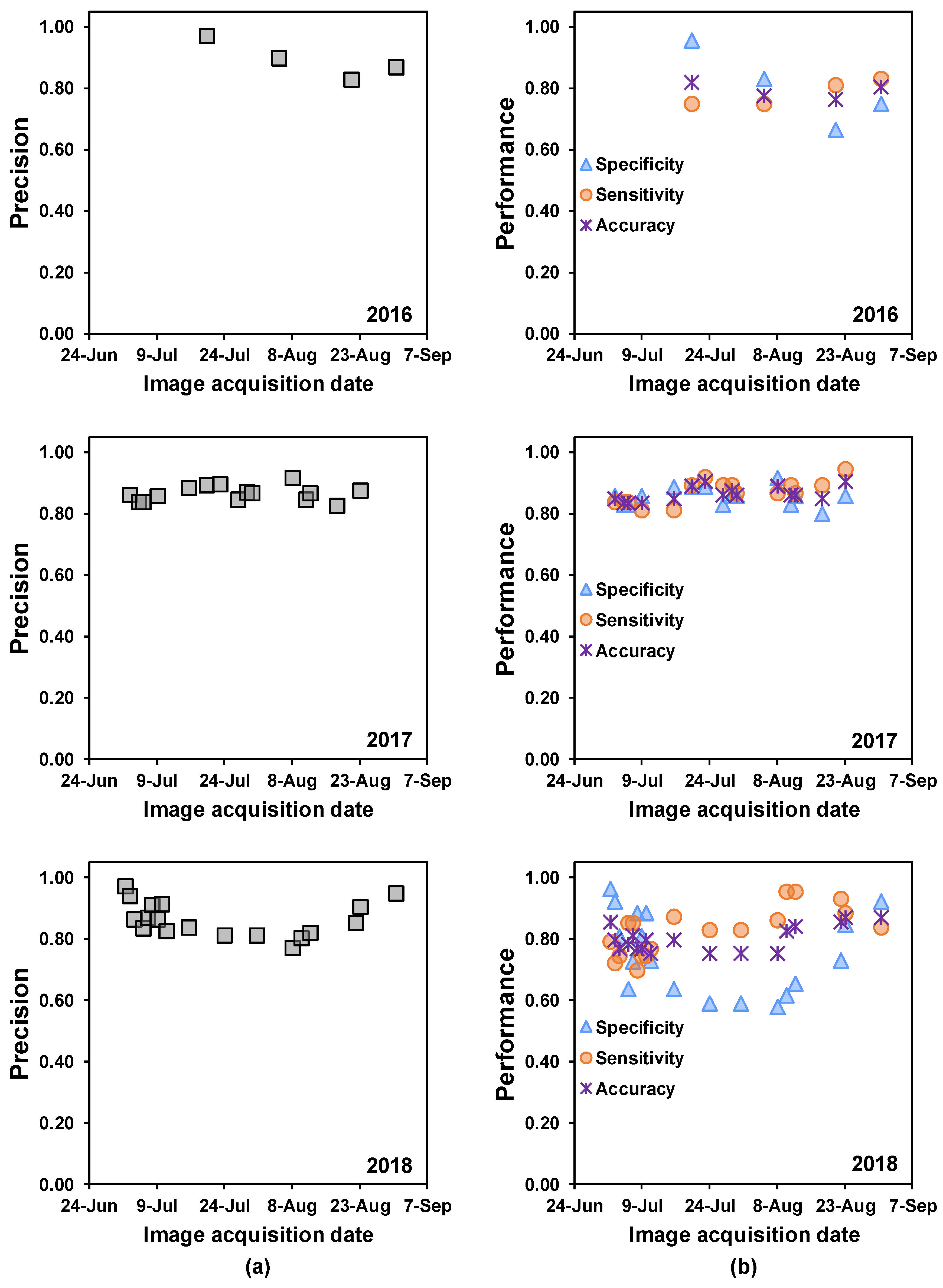
Confusion matrices (
Table 2) were constructed to assess the precision, specificity, sensitivity, and overall accuracy [
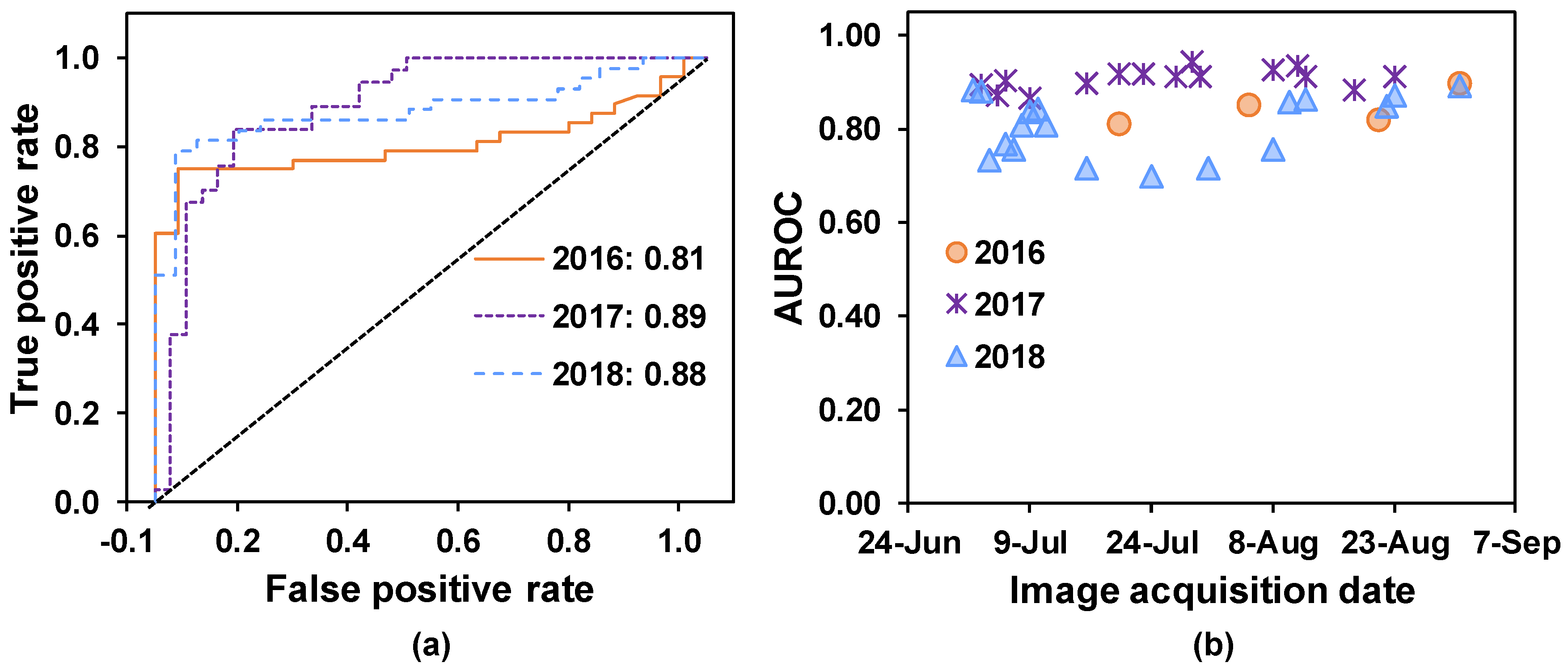
58] of SDS predictions made based on data from each PS ortho scene. Precision describes the percentage of “diseased” classifications that were assigned correctly. Specificity explains the percentage of correctly classified healthy quadrats, and sensitivity describes the percentage of correctly classified diseased quadrats. The overall accuracy is given by the proportion of correctly classified quadrats (of either status), which explains the ability of random forest trained models to classify healthy and diseased quadrats correctly. In addition, the area under the receiver operating characteristic (AUROC) curve was computed to evaluate the overall model performance for predicting SDS in the unknown test set [
59]. The receiver operating curve is a graphical plot that summarizes the performance of a classifier (i.e., a model) over all possible thresholds, which indicates the diagnostic ability of the model. This process was repeated for all ortho scenes collected from the three soybean growing seasons.
4. Discussion
Remote sensing data have the potential to detect SDS-stress-triggered changes in the pattern of light emission from plant canopies [
60]. Our findings suggest that canopy changes due to soybean SDS can be detected using high-resolution satellite imagery, even before the development of foliar symptoms in soybean fields (
Figure 7), and that prediction accuracy improves when known site-specific factors are included in the model.
Both healthy and diseased soybean quadrats were classified with >77% precision based on PlanetScope 3-m-resolution images collected from 2016 through 2018 i.e., quadrats were correctly classified as diseased or healthy more than 77% of the time. Overall, SDS classification sensitivity was higher than specificity, which indicates that classification accuracy was higher for diseased soybean quadrats than for healthy quadrats.
In exploratory data analyses, we saw that diseased soybean quadrats had high reflectance in the visible light bands (RGB) and low reflectance in the NIR band. However, healthy soybean quadrats showed lower reflectance in the visible light bands and higher reflectance in the NIR band. These patterns became more evident for disease ratings near the end of the season, when SDS foliar symptom onset and incidence increased and loss of leaf biomass and green pigment were readily observed. Despite the low level of incidence and foliar symptoms in the 2017 and 2018 growing seasons, trained random forest models detected diseased quadrats with high accuracy, even when plants were at vegetative growth stages.
In all three years, SDS-symptomatic soybean quadrats were classified correctly with 75% or better classification accuracy (
Table 3). Moreover, in 2016 and 2017, diseased and healthy soybean quadrats were detected with more than 81% AUROC, indicating that random forest trained models detected SDS with good discriminative ability, even before foliar symptom development. An AUROC value of 1.0 represents perfect classification, whereas an AUROC value of 0.5 indicates pure chance.
ROC curves are useful tools for measuring classification accuracy in binary-class problems such as ours [
61]. In 2018, AUROC values decreased for satellite images obtained between July 16 and August 8 and improved in subsequent images. Foliar damage caused by application of post-emergent herbicides (glyphosate as potassium salt (1.54 kg active ingredient/ha) plus lactofen (0.14 kg active ingredient/ha)) on July 11, 2018, at soybean growth stage R1-R2, likely altered soybean canopy reflectance during this period. Plant stress and foliar tissue damage were observed after the spray, and we postulate that this could have interfered with SDS detection in images collected during that time period. Despite these facts, the overall AUROC values ranged from 0.70 to 0.89, demonstrating the random forest models’ ability to detect SDS-associated changes with reasonable accuracy.
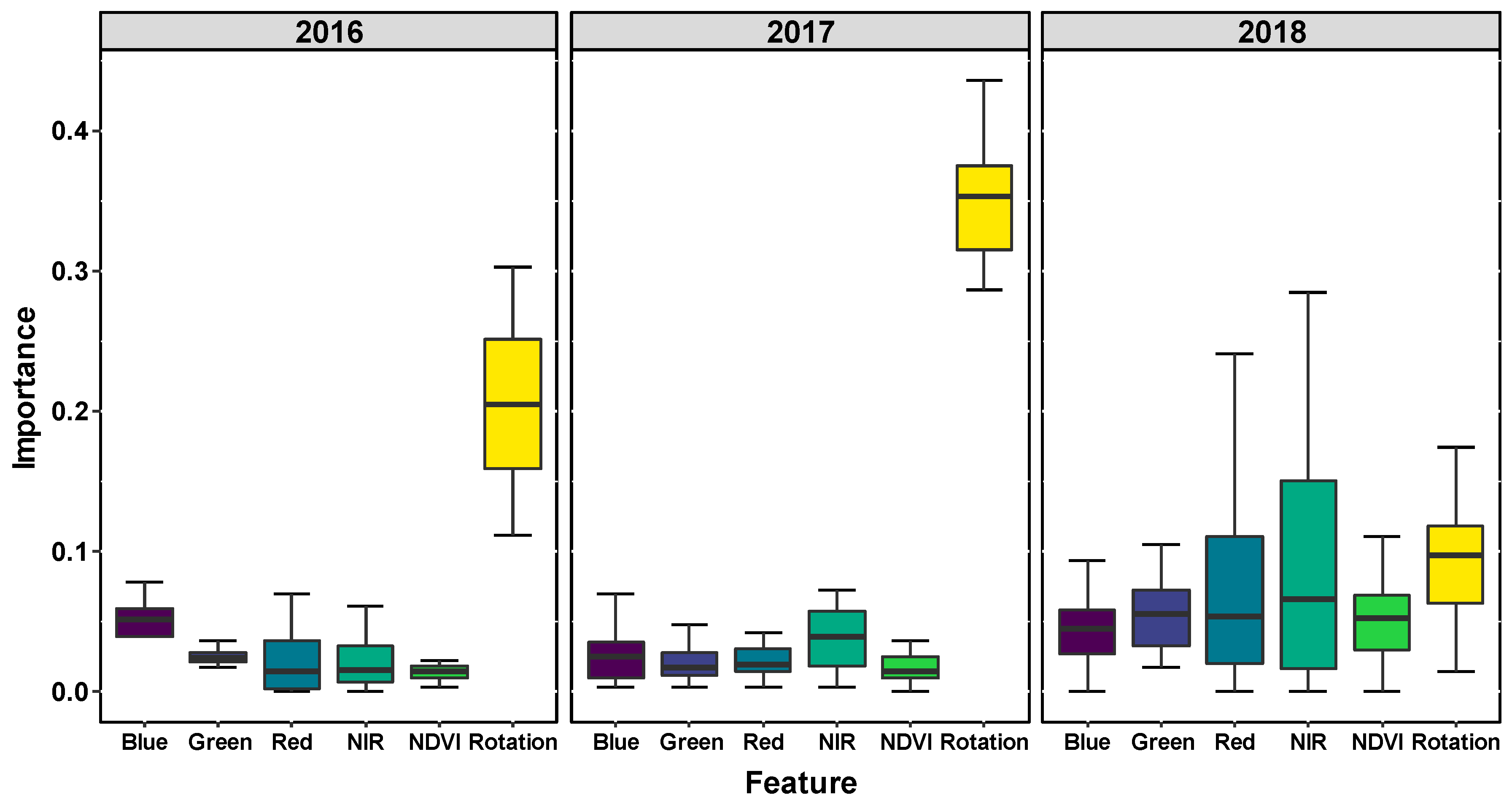
The random forest algorithm allows the evaluation of the predictive importance of the different input features (exploratory variables). By testing different feature combinations for SDS detection, we found high and consistent accuracy for AUROC values when using all features to train the models. The importance of features (variables) varied. In many cases, crop rotation information was more important than any of the spectral features. However, this alone was never sufficient for good results. For the best results, a set of all features (i.e., spectral and known environmental or treatment factors) should be used. It should be left to the RF model to prioritize certain features from this set to arrive at an optimal outcome.
A recent study conducted at the same site suggests that the long-term, diversified crop rotation systems have a significant effect on SDS foliar severity and incidence [
43]. Because rotation-specific differences in SDS incidence have been documented since 2010, we included rotation information in the SDS detection model along with remote sensing variables. The importance of crop rotation information for SDS detection in our study suggests that features specific to a study site (e.g., prior knowledge of disease occurrence and intensity, landscape features, environmental inputs, or cultivar susceptibility) may be as important as spectral features. Yang et al. [
62] used a combination of geographic features and satellite image data to predict SDS risk in commercial soybean fields throughout the state of Iowa. Their field survey data confirmed that information from early-season NDVI values and geographical features (e.g., field slope and distance from water flow lines) were correlated with late-season SDS intensity.
In another study, Herrmann et al. [
25] detected SDS before visual symptom onset with a classification accuracy of 88% and 91% from canopy and leaf-scale reflectance spectra, respectively. These classification accuracies were achieved for observations made after mid-July. In comparison, we observed similar or higher levels of SDS detection accuracy for the first week of July in the 2017 and 2018 cropping seasons. Our study differed from that of Herrmann et al. [
25] in that they discriminated between artificially inoculated and non-inoculated plots, with unknown levels of naturally occurring inoculum, whereas we assessed naturally infested plots and classified “true” disease status at the end of the growing season.
In another microplot study, Hatton et al. [
28] used thermal infrared sensors mounted on an unmanned aerial vehicle to assess SDS development in soybean plots. They conducted flights once foliar symptom development started and found a strong correlation between canopy temperatures and SDS symptom development at the end of the growing season. However, they did not address the early detection of SDS before foliar symptom onset.
5. Limitations and Future Work
Although we obtained high classification accuracies, our study has limitations. Our site already had a multiyear disease history documented by very detailed and diligent ground-truthing. Rotation turned out to be an important predictor at our site, as we have multiple years of data to document differences in SDS disease intensity. However, in other field situations, no scouting may have been performed to establish SDS at specific locations or no history of SDS occurrence may be available.
As we only used the test subset for our prediction, it is not known how useful a model trained on data from prior years would be for predicting disease in fields with unknown or poorly described prior disease histories. Even with a well-documented dataset like ours, additional studies are needed to evaluate if a model trained with data from past years could be used to predict SDS in future years.
Another limitation is that we tested only one sampling design for our study, and it is highly likely that sampling strategies could be optimized. Because of the high degree of manual work involved, we ground-truthed only the center (1.5 × 3 m) of each 9 × 9 m quadrat. However, in the random forest model, we used the average of the entire 9 × 9 m quadrat as spectral values for the quadrat. Estimation of disease status in large areas using scouting data may be improved by estimating disease at multiple points within an area of interest, or by utilizing data from areas with a range of disease intensities. Evaluation of sampling designs should be conducted using multiple sites and years, with data collected at both small and large spatial scales.
The current work showed that a consistent separation between the two classes (healthy vs. SDS) in all bands cannot realistically be expected when using satellite images with moderately-large pixels collected on fairly uniform, green plant canopies. The random forest models were able to incorporate small differences in just a few bands (or even one band), but these typically were less important than the crop rotation. However, a greater degree of consistent separation between multiple bands would certainly be desirable. Focusing only on the pixels covering the ground-truthed 1.5 × 3 m area within each quadrat may offer better results in the future. The use of multiband imagery acquired by drones (UAVs), which are becoming more common and provide a far higher resolution, is an alternative we are currently exploring.
Random forest models are known to cope well with a high number of variables (features). Similarly to previous work in this field, our study used NDVI as a variable for the model. However, it is unclear if this is permissible. One might argue that as NDVI is a ratio involving only the red and NIR bands, and as the mean of each of these bands is already used, no new information is added to the model when using NDVI. Instead, it might be more advantageous to explore the inclusion of other variables that describe each of the spectral variables in more detail, such as the minimum, maximum, standard deviation, median, or other quantiles. Our study also did not encode the location of the quadrats relative to each other for the random forest model, and this spatial (distance) information could also be used. Depending on their scale, future studies might potentially include soil types, distance to flowlines [
62], and climatic and environmental trends [
15].
A minor limitation arises from the satellite generations being switched (with different spectral response curves) in late July 2017. This means that the graphs showing our results along timelines may exhibit a break at that date. However, the impact of this change in response curves on the random forest models is unclear. While it is true that a model from 2016 is, therefore, not comparable to a 2018 model, we never compared models and their results directly, other than via the afore-mentioned graphs.
In the current work, each RF model was trained on a single date. In the future, we plan to explore the use of models that are based on multiple dates (e.g., all late growing season dates), which we expect should increase the predictive power of RF models. In this scenario, the generation of the satellite will be used as an independent variable when generating the random forest model.
6. Conclusions
In this study, we explored the potential of using multispectral PlanetScope satellite imagery for the classification of soybean quadrats as either healthy or SDS-infected using random forest models with spectral data (blue, green, red, and NIR bands), calculated NDVI, and crop rotation information obtained for three years (2016–2018). We provided example data and Python code to enable others to test, extend, and further develop this technique. Splitting the data into training (70%) and test (30%) subsets, healthy and diseased soybean quadrats were detected with more than 75% accuracy and 70% AUROC in all years, suggesting that high-resolution satellite imagery has the potential for early and accurate detection of SDS in soybean fields.
High-resolution multispectral imagery of cropland represents a relatively untapped, vast resource to explore. Early detection of SDS may be used by the industry to develop novel, sustainable, and environment-friendly means to mitigate this disease at early stages, and may meet practical demands for detection of SDS at a regional scale to inform field management applications and crop insurance decisions. Future studies could evaluate high-resolution multispectral imagery at the regional scale for early detection and monitoring of other economically significant diseases, which may facilitate effective policymaking, support decision making, and guide recommendations for sustainable and efficient allocation of management resources in precision agriculture systems.