Automatic Estimation of Crop Disease Severity Levels Based on Vegetation Index Normalization
Abstract
1. Introduction
2. Materials and Methods
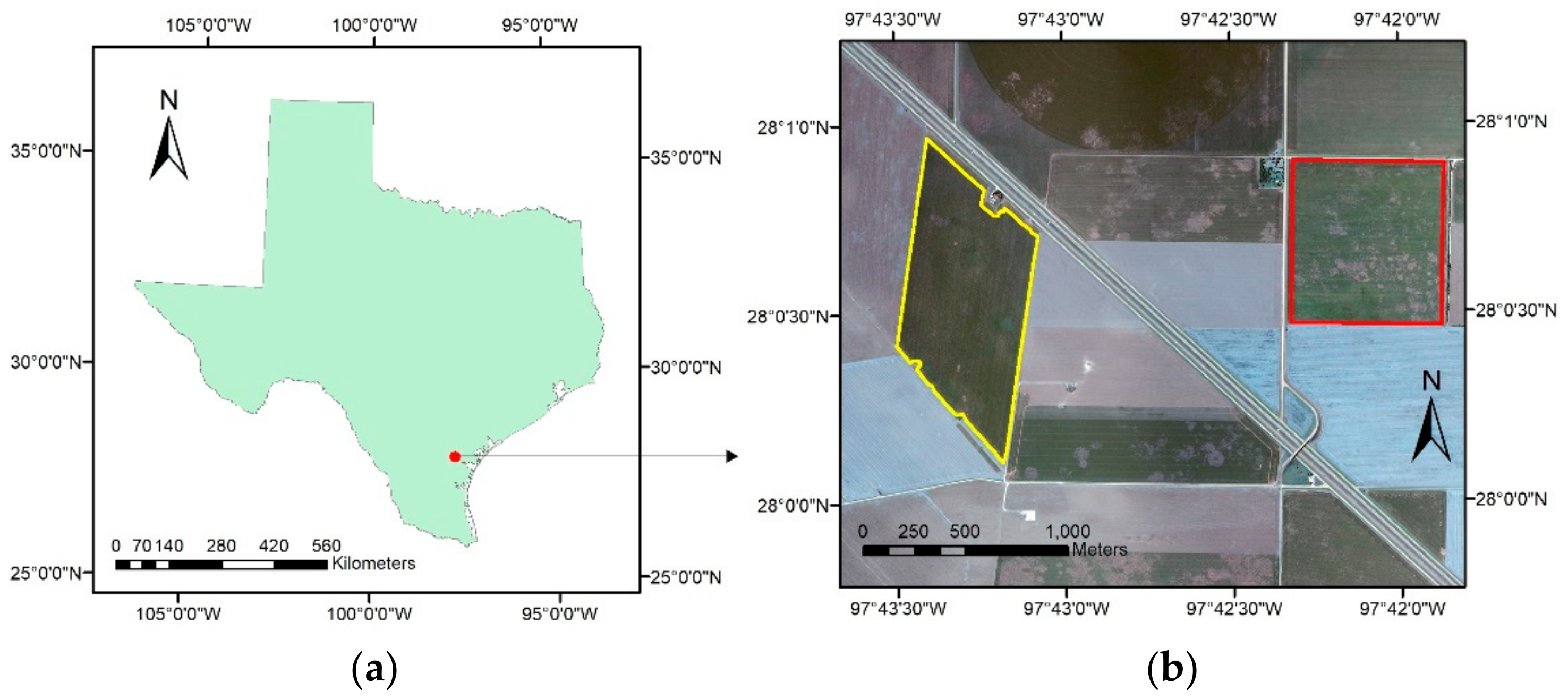
2.1. Study Area
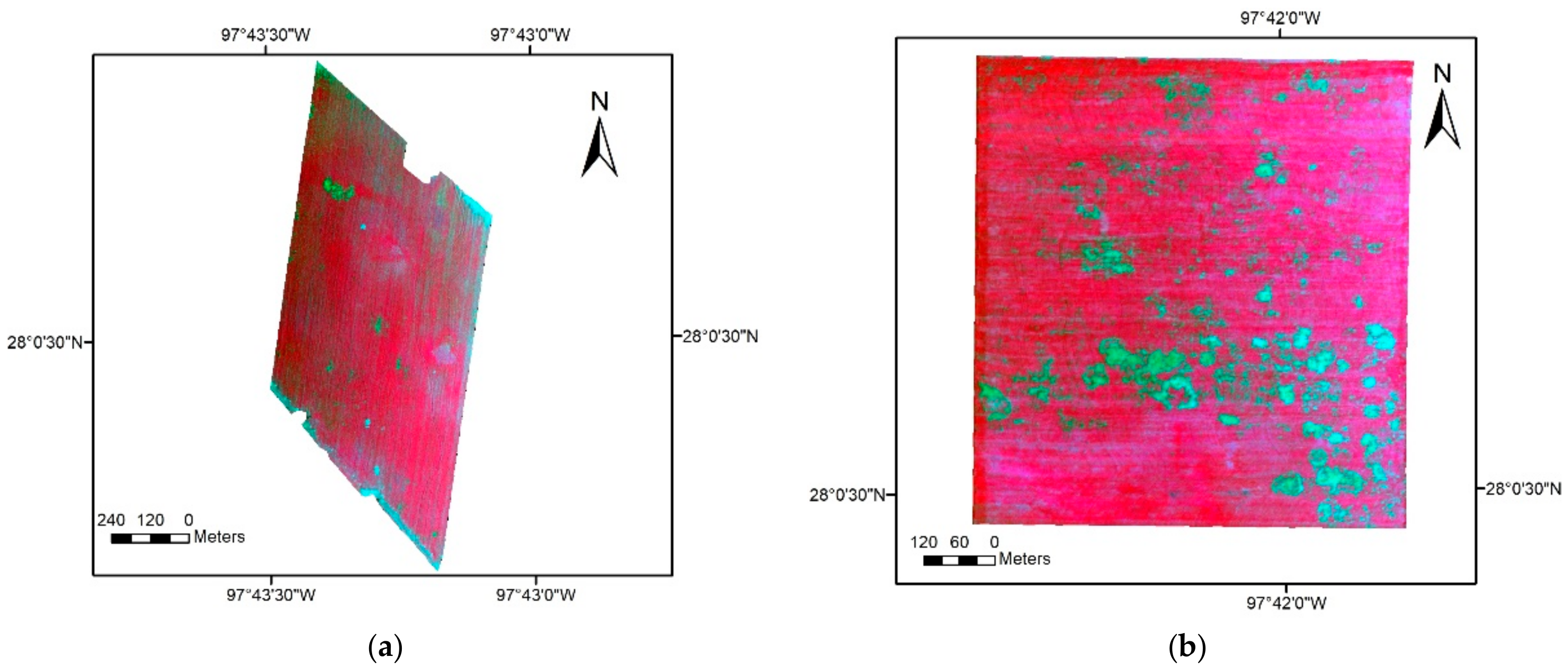
2.2. Image Acquisition and Processing
2.3. Data Analysis
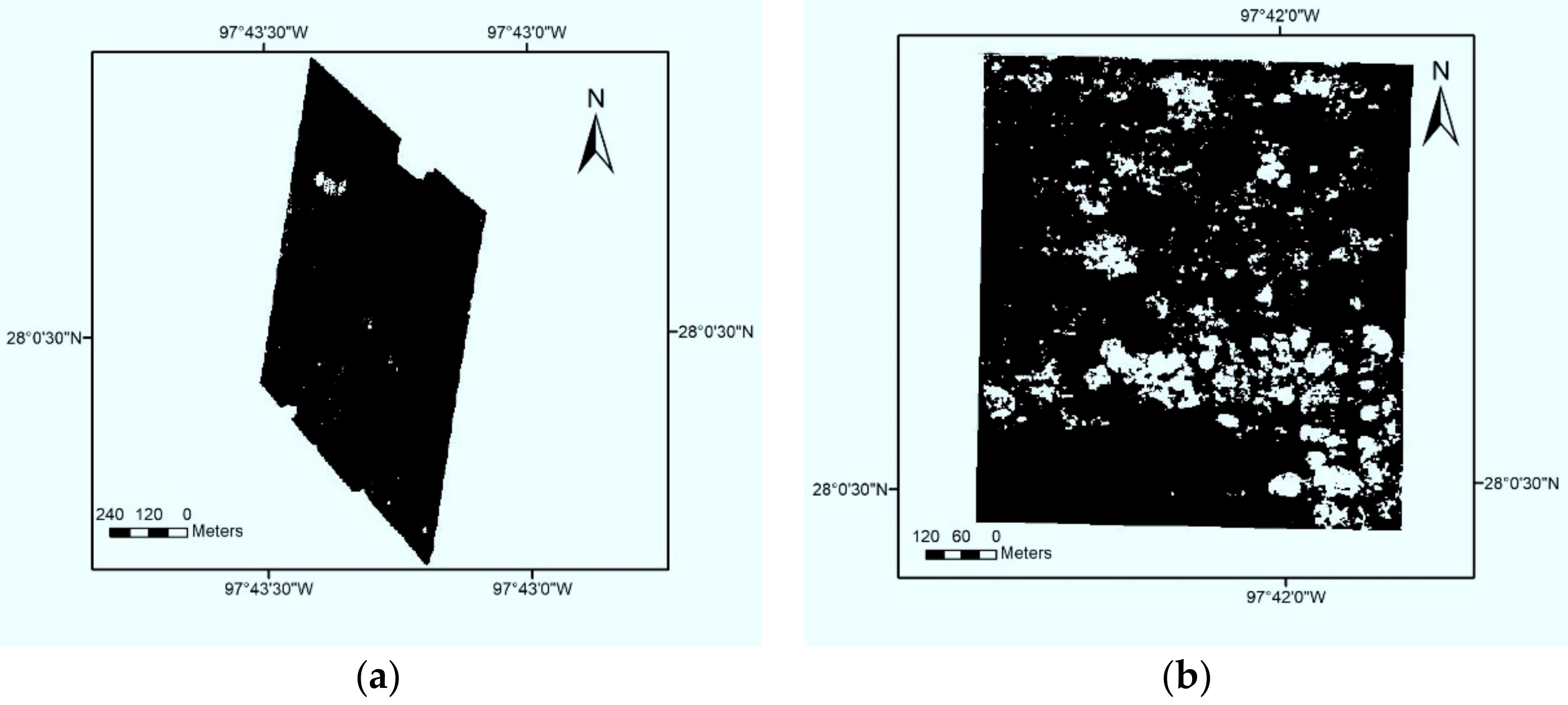
2.3.1. Cotton Root Rot Classification
2.3.2. Calculation and Normalization of Vegetation Indices
2.3.3. Disease Severity Assessment
2.3.4. Accuracy Evaluation of Disease Severity Levels
3. Results
3.1. Classification of Cotton Root Rot
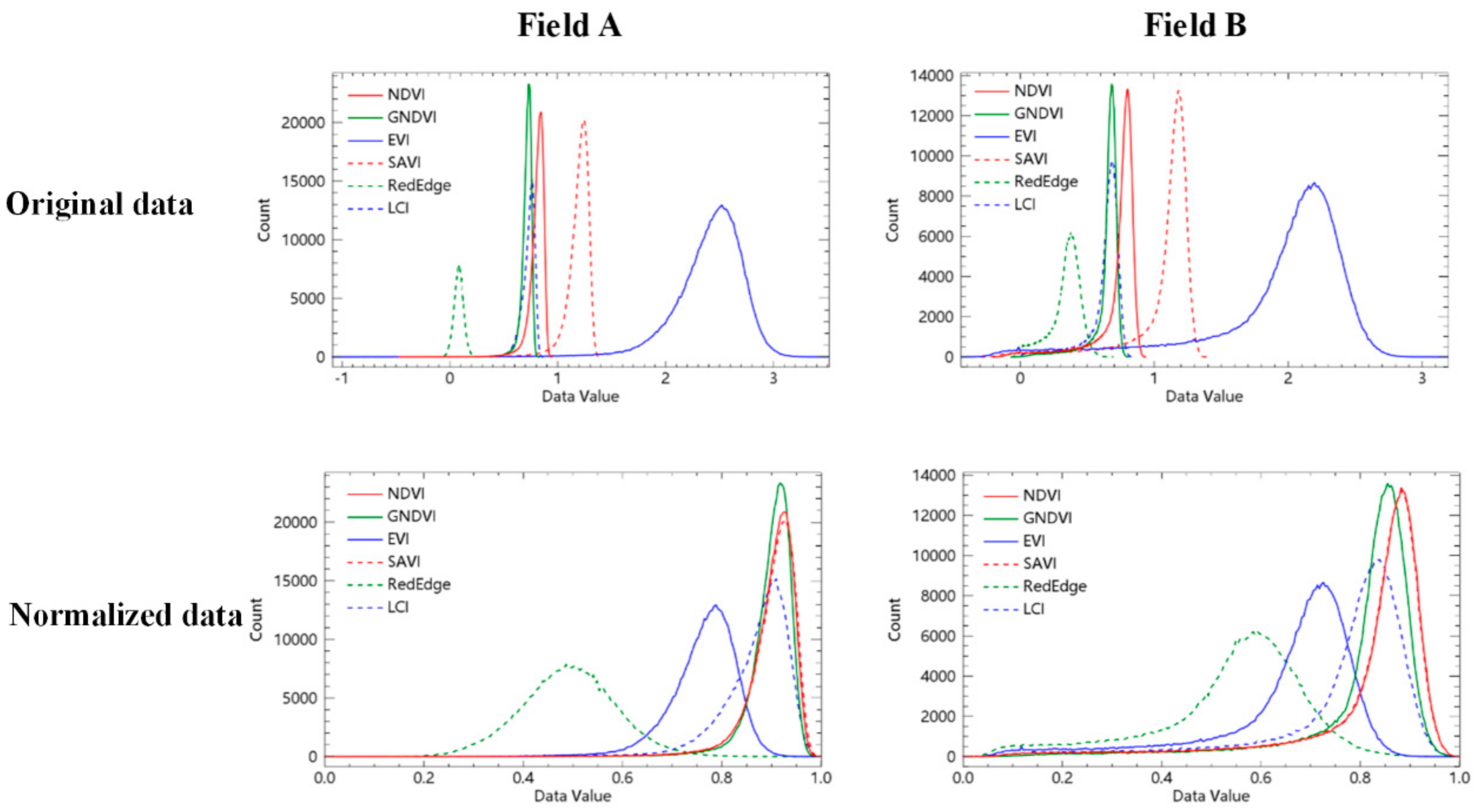
3.2. Calculation and Normalization of Vegetation Indices
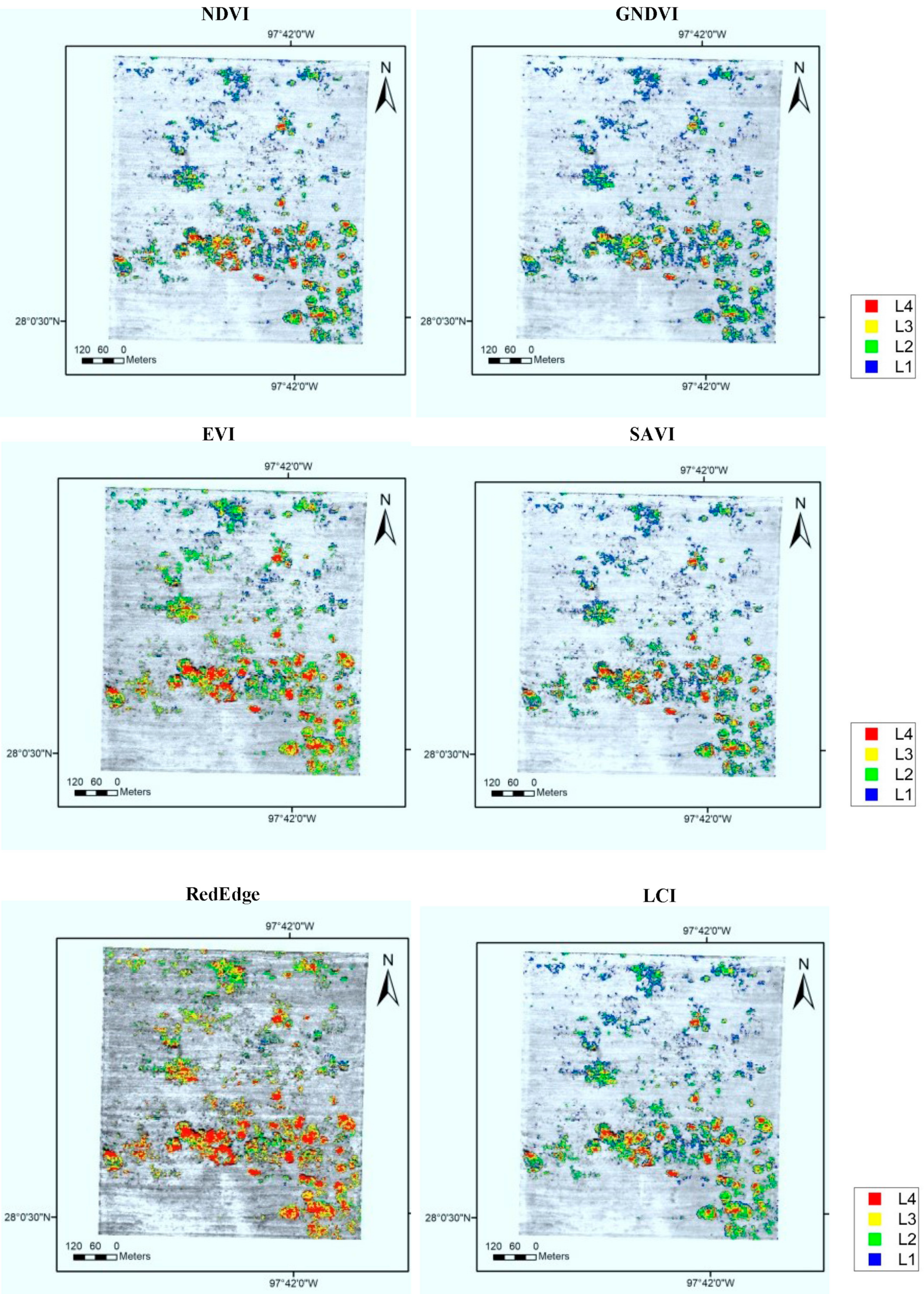
3.3. Root Rot Severity Assessment
3.4. Accuracy Assessment of Disease Severity Levels
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Liu, Z.; Huang, J.; Tao, R. Characterizing and estimating fungal disease severity of rice brown spot with hyperspectral reflectance data. Rice Sci. 2008, 15, 232–242. [Google Scholar] [CrossRef]
- Ashourloo, D.; Mobasheri, M.; Huete, A. Developing two spectral disease indices for detection of wheat leaf rust (Puccinia triticina). Remote Sens. 2014, 6, 4723–4740. [Google Scholar] [CrossRef]
- Jin, N.; Huang, W.; Ren, Y.; Luo, J.; Wu, Y.; Jing, Y.; Wang, D. Hyperspectral identification of cotton verticillium disease severity. Optik 2013, 124, 2569–2573. [Google Scholar] [CrossRef]
- Calderón, R.; Navas-Cortés, J.A.; Lucena, C.; Zarco-Tejada, P.J. High-resolution airborne hyperspectral and thermal imagery for early detection of Verticillium wilt of olive using fluorescence, temperature and narrow-band spectral indices. Remote Sens. Environ. 2013, 139, 231–245. [Google Scholar] [CrossRef]
- Zhang, J.; Pu, R.; Loraamm, R.W.; Yang, G.; Wang, J. Comparison between wavelet spectral features and conventional spectral features in detecting yellow rust for winter wheat. Comput. Electron. Agric. 2014, 100, 79–87. [Google Scholar] [CrossRef]
- Duveiller, G.; Weiss, M.; Baret, F.; Defourny, P. Retrieving wheat Green Area Index during the growing season from optical time series measurements based on neural network radiative transfer inversion. Remote Sens. Environ. 2011, 115, 887–896. [Google Scholar] [CrossRef]
- Zhang, J.; Pu, R.; Wang, J.; Huang, W.; Yuan, L.; Luo, J. Detecting powdery mildew of winter wheat using leaf level hyperspectral measurements. Comput. Electron. Agric. 2012, 85, 13–23. [Google Scholar] [CrossRef]
- Yang, C.; Everitt, J.H.; Fernandez, C.J. Comparison of airborne multispectral and hyperspectral imagery for mapping cotton root rot. Biosyst. Eng. 2010, 2, 131–139. [Google Scholar] [CrossRef]
- Yang, C.; Odvody, G.N.; Fernandez, C.J.; Landivar, J.A.; Minzenmayer, R.R.; Nichols, R.L. Evaluating unsupervised and supervised image classification methods for mapping cotton root rot. Precis. Agric. 2015, 16, 201–215. [Google Scholar] [CrossRef]
- Yang, C.H.; Odvody, G.N.; Fernandez, C.J.; Landivar, J.A.; Minzenmayer, R.R.; Nichols, R.L.; Thomasson, J.A. Monitoring cotton root rot progression within a growing season using airborne multispectral imagery. J. Cotton Sci. 2014, 18, 85–93. [Google Scholar]
- Yang, C.; Odvody, G.N.; Thomasson, J.A.; Isakeit, T.; Minzenmayer, R.R.; Drake, D.R.; Nichols, R.L. Site-Specific Management of Cotton Root Rot Using Airborne and High-Resolution Satellite Imagery and Variable-Rate Technology. Trans. ASABE. 2018, 61, 849–858. [Google Scholar] [CrossRef]
- Huang, W.; Guan, Q.; Luo, J.; Zhang, J.; Zhao, J.; Liang, D.; Huang, L.; Zhang, D. New optimized spectral indices for identifying and monitoring winter wheat diseases. IEEE J. Sel. Top. Appl. Earth Obs. Remote. Sens. 2014, 7, 2516–2524. [Google Scholar] [CrossRef]
- Kumar, N.; Niwas, R.; Khichar, M.L.; Saini, R.K.; Biswas, B. Effect of different growing environments on population dynamics of sucking pests in relation to various spectral indices in cotton. J. Indian Soc. Remote 2013, 41, 309–317. [Google Scholar] [CrossRef]
- Liu, L.; Song, X.; Li, C.; Qi, L.; Huang, W.; Wang, J. Monitoring and evaluation of the diseases of and yield winter wheat from multi-temporal remotely-sensed data. Trans. Chin. Soc. Agric. Eng. 2009, 25, 137–143. [Google Scholar]
- Prabhakar, M.; Prasad, Y.G.; Thirupathi, M.; Sreedevi, G.; Dharajothi, B.; Venkateswarlu, B. Use of ground based hyperspectral remote sensing for detection of stress in cotton caused by leafhopper (Hemiptera: Cicadellidae). Comput. Electron. Agric. 2011, 79, 189–198. [Google Scholar] [CrossRef]
- Chen, B.; Li, S.; Wang, K.; Zhou, G.; Bai, J. Evaluating the severity level of cotton Verticillium using spectral signature analysis. Int. J. Remote Sens. 2012, 33, 2706–2724. [Google Scholar] [CrossRef]
- Wu, M.; Yang, C.; Song, X.; Hoffmann, W.C.; Huang, W.; Niu, Z.; Wang, C.; Li, W.; Yu, B. Monitoring cotton root rot by synthetic Sentinel-2 NDVI time series using improved spatial and temporal data fusion. Sci. Rep. UK 2018, 8, 2016. [Google Scholar] [CrossRef]
- Ahamed, T.; Tian, L.; Zhang, Y.; Ting, K.C. A review of remote sensing methods for biomass feedstock production. Biomass Bioenergy 2011, 35, 2455–2469. [Google Scholar] [CrossRef]
- Yang, C.; Greenberg, S.M.; Everitt, J.H.; Fernandez, C.J. Assessing cotton defoliation, regrowth control and root rot infection using remote sensing technology. Int. J. Agric. Biol. Eng. 2011, 4, 1–11. [Google Scholar]
- Zhang, L.; Sun, X.; Wu, T.; Zhang, H. An analysis of shadow effects on spectral vegetation indexes using a ground-based imaging spectrometer. IEEE Geosci. Remote Sens. Lett. 2015, 12, 2188–2192. [Google Scholar] [CrossRef]
- Oumar, Z.; Mutanga, O.; Ismail, R. Predicting Thaumastocoris peregrinus damage using narrow band normalized indices and hyperspectral indices using field spectra resampled to the Hyperion sensor. Int. J. Appl. Earth Obs. 2013, 21, 113–121. [Google Scholar] [CrossRef]
- Tucker, C.J. Red and photographic infrared linear combinations for monitoring vegetation. Remote Sens. Environ. 1979, 8, 127–150. [Google Scholar] [CrossRef]
- Gitelson, A.A.; Merzlyak, M.N. Remote sensing of chlorophyll concentration in higher plant leaves. Adv. Space Res. 1998, 22, 689–692. [Google Scholar] [CrossRef]
- Haboudane, D.; Miller, J.R.; Tremblay, N.; Zarco-Tejada, P.J.; Dextraze, L. Integrated narrow-band vegetation indices for prediction of crop chlorophyll content for application to precision agriculture. Remote Sens. Environ. 2002, 81, 416–426. [Google Scholar] [CrossRef]
- Huete, A.R. A soil-adjusted vegetation index (SAVI). Remote Sens. Environ. 1988, 25, 295–309. [Google Scholar] [CrossRef]
- Cloutis, E.A.; Connery, D.R.; Major, D.J.; Dover, F.J. Airborne multi-spectral monitoring of agricultural crop status: Effect of time of year, crop type and crop condition parameter. Int. J. Remote Sens. 1996, 17, 2579–2601. [Google Scholar] [CrossRef]
- Datt, B. A new reflectance index for remote sensing of chlorophyll content in higher plants: Tests using Eucalyptus leaves. J. Plant. Physiol. 1999, 154, 30–36. [Google Scholar] [CrossRef]
- Kruse, F.A.; Lefkoff, A.B.; Boardman, J.W.; Heidebrecht, K.B.; Shapiro, A.T.; Barloon, P.J.; Goetz, A. The spectral image processing system (SIPS)—interactive visualization and analysis of imaging spectrometer data. Remote Sens. Environ. 1993, 44, 145–163. [Google Scholar] [CrossRef]
- Vishnu, S.; Nidamanuri, R.R.; Bremananth, R. Spectral material mapping using hyperspectral imagery: A review of spectral matching and library search methods. Geocarto Int. 2013, 28, 171–190. [Google Scholar] [CrossRef]
- Song, X.; Yang, C.; Wu, M.; Zhao, C.; Yang, G.; Hoffmann, W.; Huang, W. Evaluation of Sentinel-2A Satellite Imagery for Mapping Cotton Root Rot. Remote Sens. 2017, 9, 906. [Google Scholar] [CrossRef]
- Mirik, M.; Ansley, R.J.; Michels, G.J.; Elliott, N.C. Spectral vegetation indices selected for quantifying Russian wheat aphid (Diuraphis noxia) feeding damage in wheat (Triticum aestivum L.). Precis. Agric. 2012, 13, 501–516. [Google Scholar] [CrossRef]
- Izzuddin, M.A.; Nisfariza, M.N.; Ezzati, B.; Idris, A.S.; Steven, M.D.; Boyd, D. Analysis of airborne hyperspectral image using vegetation indices, red edge position and continuum removal for detection of ganoderma disease in oil palm. J. Oil Palm Res. 2018, 30, 416–428. [Google Scholar]
- Chen, B.; Li, S.K.; Wang, K.R.; Wang, F.Y.; Tan, H.Z.; Liu, G.Q.; Chen, J.L. Spectrum characteristics of cotton single leaf infected by verticillium wilt and estimation on severity level of disease. Sci. Agric. Sin. 2007, 40, 2709–2715. [Google Scholar]
- Clark, R.N.; Swayze, G.A.; Livo, K.E.; Kokaly, R.F.; Sutley, S.J.; Dalton, J.B.; McDougal, R.R.; Gent, C.A. Imaging spectroscopy: Earth and planetary remote sensing with the USGS Tetracorder and expert systems. J. Geophys. Res. 2003, 108, 5131. [Google Scholar] [CrossRef]
- Muhammed, H.H. Hyperspectral crop reflectance data for characterising and estimating fungal disease severity in wheat. Biosyst. Eng. 2005, 91, 9–20. [Google Scholar] [CrossRef]
- Lyon, J.G.; Yuan, D.; Lunetta, R.S.; Elvidge, C.D. A change detection experiment using vegetation indices. Photogramm. Eng. Remote Sens. 1998, 64, 143–150. [Google Scholar]
- Mutowo, G.; Mutanga, O.; Masocha, M. Mapping foliar N in miombo woodlands using sentinel-2 derived chlorophyll and structural indices. J. Appl. Remote Sens. 2018, 12, 46028. [Google Scholar] [CrossRef]
- Yang, C.; Cheng, C.; Chen, R. Changes in spectral characteristics of rice canopy infested with brown planthopper and leaffolder. Crop. Sci. 2007, 47, 329–335. [Google Scholar] [CrossRef]
- Adão, T.; Hruška, J.; Pádua, L.; Bessa, J.; Peres, E.; Morais, R.; Sousa, J.J. Hyperspectral imaging: A review on UAV-based sensors, data processing and applications for agriculture and forestry. Remote Sens. 2017, 9, 1110. [Google Scholar] [CrossRef]
- Raj, R.; Kar, S.; Nandan, R.; Jagarlapudi, A. Precision Agriculture and Unmanned Aerial Vehicles (UAVs). In Unmanned Aerial Vehicle: Applications in Agriculture and Environment; Springer: Berlin/Heidelberg, Germany, 2020; pp. 7–23. [Google Scholar]
- Qin, Z.; Zhang, M. Detection of rice sheath blight for in-season disease management using multispectral remote sensing. Int. J. Appl. Earth Obs. 2005, 7, 115–128. [Google Scholar] [CrossRef]
- Adams, M.L.; Norvell, W.A.; Philpot, W.D.; Peverly, J.H. Toward the discrimination of manganese, zinc, copper, and iron deficiency in ‘Bragg’soybean using spectral detection methods. Agron. J. 2000, 92, 268–274. [Google Scholar]
- Chen, P.; Zhang, J.; Li, M.; Lei, Y. Physiological change and hyperspectral character analysis of cotton leaves infested by Tetranychus turkestani. Chin. Bull. Entomol. 2007, 44, 61–65. [Google Scholar]



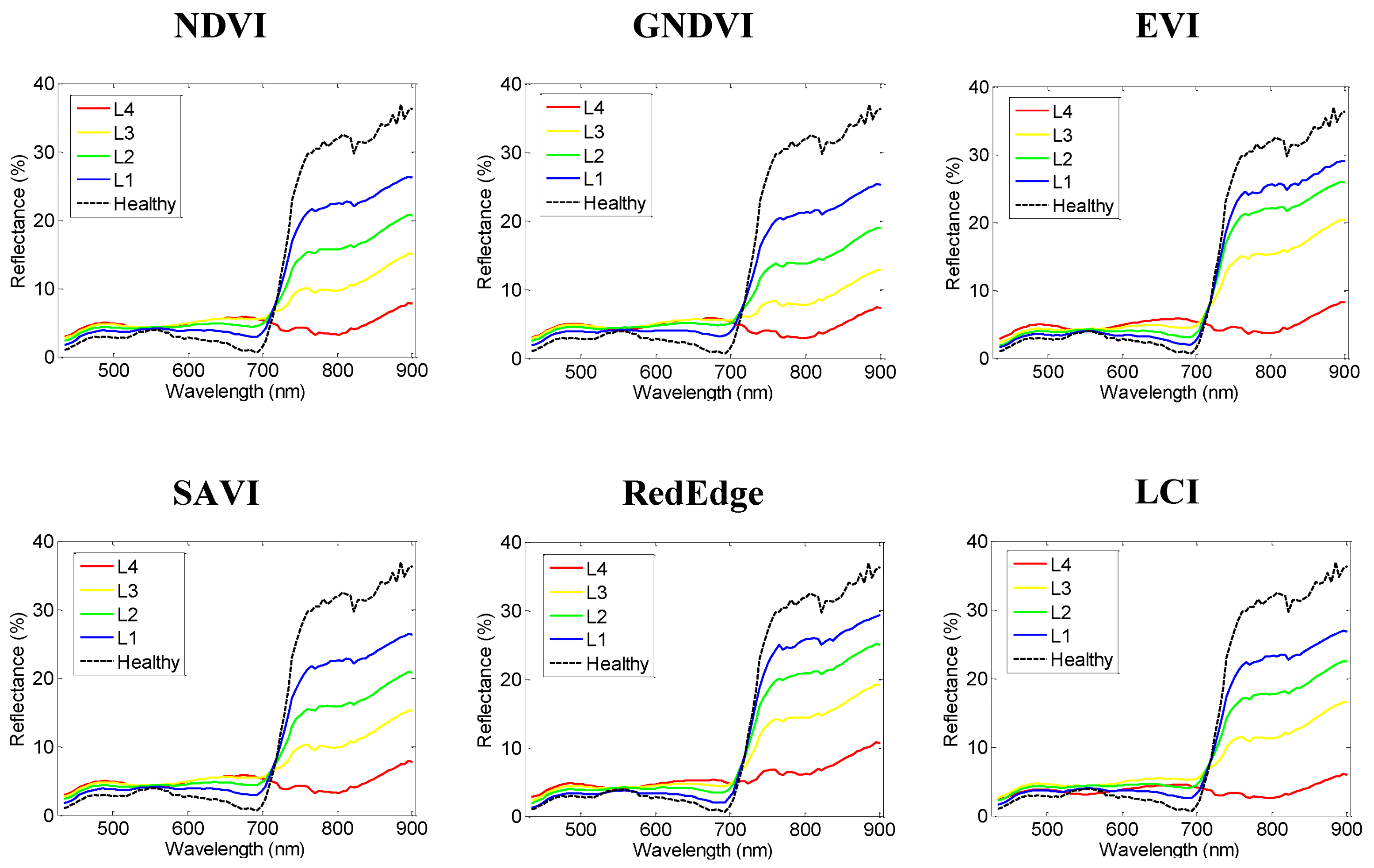
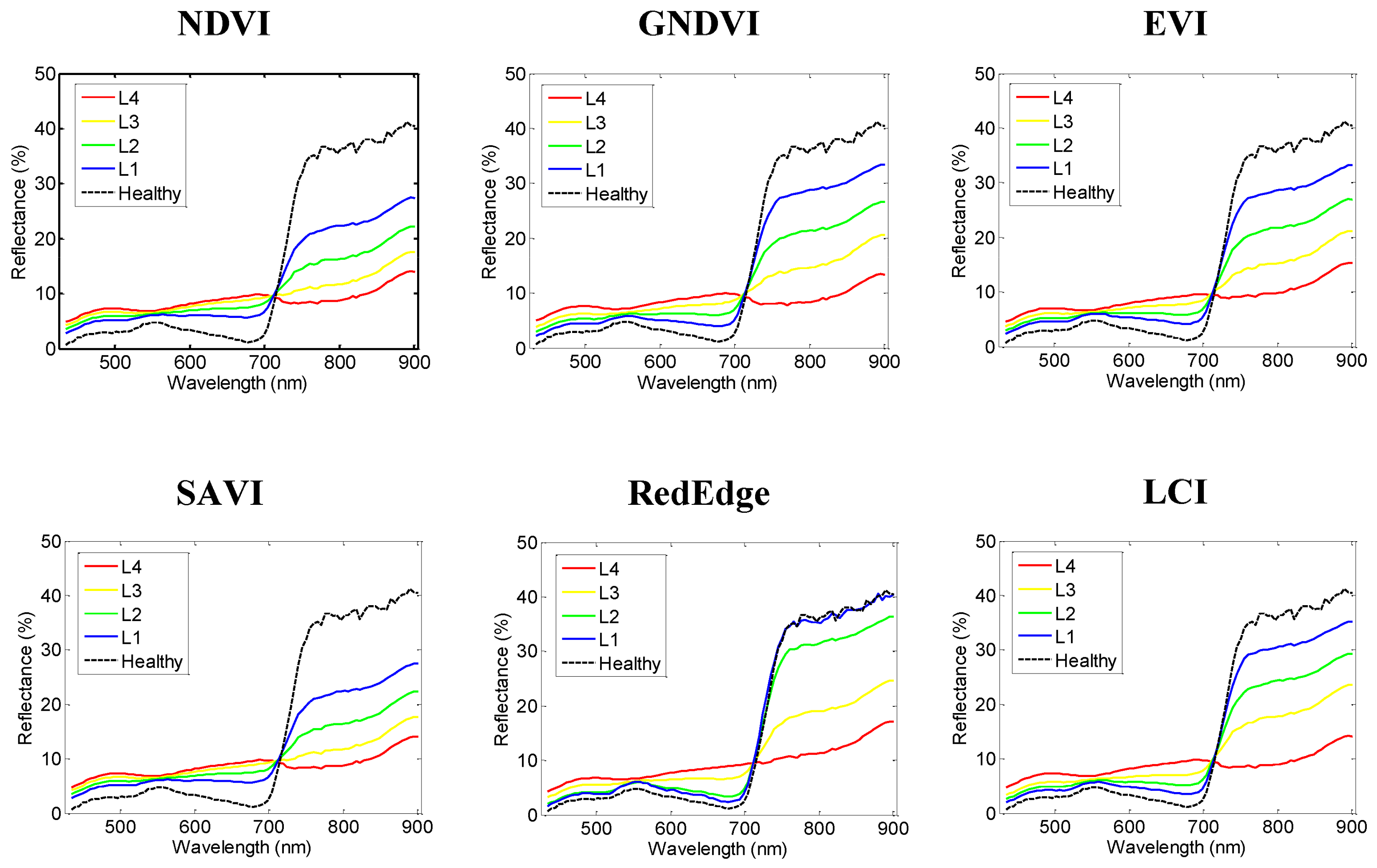
| Vegetation Index | Abbrev. | Formula | Application | Reference |
|---|---|---|---|---|
| Normalized difference vegetation index | NDVI | (R800 − R670)/(R800 + R670) | Biomass | [22] |
| Green normalized difference vegetation index | GNDVI | (R800 − R550)/(R800 + R550) | PAR, vegetation cover | [23] |
| Enhanced vegetation index | EVI | 2.5(R800 − R670)/(R800 + 6R670 − 7.5R450 + 1) | Biomass, LAI | [24] |
| Soil adjusted vegetation index | SAVI | (1 + L)(R800 − R670)/(R800 + R670 + L) | Biomass, soil background | [25] |
| Red edge 2 | RedEdge | (R710 − R680)/(R710 + R680) | Chlorophyll content | [26] |
| Leaf chlorophyll index | LCI | (R850 − R710)/(R850 + R680) | Chlorophyll content | [27] |
| Severity Level | Symptom Description | Normalized Vegetation Indices |
|---|---|---|
| Level 4 (L4) | Extremely severe | 0.8–1.0 |
| Level 3 (L3) | Severe | 0.6–0.8 |
| Level 2 (L2) | Medium | 0.4–0.6 |
| Level 1 (L1) | Slight | 0.2–0.4 |
| Level 0 (L0) | Latency period | 0.0–0.2 |
| Field | Vegetation Index | Min. | Max. | Mean | Std. Dev. |
|---|---|---|---|---|---|
| A | NDVI | −0.4697 | 0.9475 | 0.8079 | 0.0775 |
| GNDVI | −0.3225 | 0.8269 | 0.7130 | 0.0544 | |
| EVI | −1.0867 | 3.5126 | 2.4153 | 0.3185 | |
| SAVI | −0.6640 | 1.4000 | 1.1916 | 0.1154 | |
| RedEdge | −0.1445 | 0.3202 | 0.0840 | 0.0441 | |
| LCI | −0.1175 | 0.8571 | 0.7277 | 0.0678 | |
| B | NDVI | −0.2028 | 0.9356 | 0.7225 | 0.1724 |
| GNDVI | −0.0511 | 0.8066 | 0.6430 | 0.1093 | |
| EVI | −0.4417 | 3.1930 | 1.9429 | 0.5463 | |
| SAVI | −0.2942 | 1.3850 | 1.0673 | 0.2564 | |
| RedEdge | −0.0724 | 0.6904 | 0.3397 | 0.1159 | |
| LCI | −0.0633 | 0.8335 | 0.6265 | 0.1361 |
| Vegetation Index | Severity Level | Min. | Max. | Mean | Number of Pixels |
|---|---|---|---|---|---|
| NDVI | L4 | −0.4697 | −0.1873 | −0.2862 | 59 |
| L3 | −0.1750 | 0.0970 | −0.0156 | 176 | |
| L2 | 0.0979 | 0.3805 | 0.2667 | 484 | |
| L1 | 0.3808 | 0.6639 | 0.5460 | 1470 | |
| GNDVI | L4 | −0.3225 | −0.0935 | −0.1657 | 35 |
| L3 | −0.0922 | 0.1371 | 0.0389 | 116 | |
| L2 | 0.1417 | 0.3670 | 0.2748 | 288 | |
| L1 | 0.3681 | 0.5970 | 0.5124 | 1256 | |
| EVI | L4 | −1.0867 | −0.1726 | −0.4488 | 91 |
| L3 | −0.1699 | 0.7439 | 0.3749 | 537 | |
| L2 | 0.7473 | 1.6613 | 1.2551 | 1465 | |
| L1 | 1.6626 | 2.5721 | 2.0241 | 1433 | |
| SAVI | L4 | −0.6640 | −0.2678 | −0.4064 | 59 |
| L3 | −0.2501 | 0.1547 | −0.0113 | 188 | |
| L2 | 0.1626 | 0.5692 | 0.4029 | 500 | |
| L1 | 0.5698 | 0.9804 | 0.8079 | 1477 | |
| RedEdge | L4 | −0.0947 | −0.0521 | −0.0614 | 35 |
| L3 | −0.0518 | 0.0407 | 0.0087 | 2190 | |
| L2 | 0.0407 | 0.1329 | 0.0646 | 1317 | |
| L1 | 0.1341 | 0.1822 | 0.1522 | 23 | |
| LCI | L4 | −0.1175 | 0.0747 | 0.0065 | 75 |
| L3 | 0.0773 | 0.2708 | 0.1974 | 235 | |
| L2 | 0.2714 | 0.4649 | 0.3850 | 695 | |
| L1 | 0.4650 | 0.6589 | 0.5688 | 1687 |
| Vegetation Index | Severity Level | Min. | Max. | Mean | Number of Pixels |
|---|---|---|---|---|---|
| NDVI | L4 | −0.2028 | 0.0240 | −0.0436 | 5128 |
| L3 | 0.0240 | 0.2508 | 0.1411 | 10,249 | |
| L2 | 0.2508 | 0.4776 | 0.3716 | 13,750 | |
| L1 | 0.4776 | 0.7044 | 0.5901 | 17,668 | |
| GNDVI | L4 | −0.0511 | 0.1204 | 0.0792 | 2491 |
| L3 | 0.1205 | 0.2920 | 0.2119 | 8477 | |
| L2 | 0.2920 | 0.4635 | 0.3863 | 13,394 | |
| L1 | 0.4637 | 0.6350 | 0.5522 | 22,074 | |
| EVI | L4 | −0.2942 | 0.2853 | 0.0378 | 10,648 |
| L3 | 0.2853 | 1.0121 | 0.6572 | 17,483 | |
| L2 | 1.0122 | 1.7391 | 1.3579 | 17,366 | |
| L1 | 1.7392 | 2.4655 | 2.0048 | 8087 | |
| SAVI | L4 | −0.2942 | 0.0403 | −0.0607 | 5278 |
| L3 | 0.0404 | 0.3749 | 0.2126 | 10,328 | |
| L2 | 0.3749 | 0.7094 | 0.5528 | 13,884 | |
| L1 | 0.7094 | 1.0439 | 0.8750 | 17,521 | |
| RedEdge | L4 | −0.0724 | 0.0802 | 0.0258 | 17,931 |
| L3 | 0.0802 | 0.2327 | 0.1532 | 23,505 | |
| L2 | 0.2328 | 0.3853 | 0.2934 | 10,799 | |
| L1 | 0.3853 | 0.5356 | 0.4204 | 1538 | |
| LCI | L4 | −0.0633 | 0.1161 | 0.0620 | 5713 |
| L3 | 0.1161 | 0.2954 | 0.2088 | 12,194 | |
| L2 | 0.2955 | 0.4748 | 0.3897 | 16,599 | |
| L1 | 0.4748 | 0.6541 | 0.5580 | 16,063 |
| Field | Vegetation Index | L4 | L3 | L2 | L1 |
|---|---|---|---|---|---|
| A | NDVI | 0.8116 | 0.4159 | 0.2312 | 0.1086 |
| GNDVI | 0.8471 | 0.5020 | 0.2809 | 0.1261 | |
| EVI | 0.7764 | 0.2386 | 0.1172 | 0.0631 | |
| SAVI | 0.8116 | 0.4086 | 0.2277 | 0.1074 | |
| RedEdge | 0.5685 | 0.2534 | 0.1366 | 0.0590 | |
| LCI | 0.8076 | 0.3580 | 0.1910 | 0.0899 | |
| B | NDVI | 0.6825 | 0.5279 | 0.3563 | 0.2111 |
| GNDVI | 0.7140 | 0.4127 | 0.2307 | 0.1064 | |
| EVI | 0.6212 | 0.3899 | 0.2204 | 0.1157 | |
| SAVI | 0.6808 | 0.5246 | 0.3531 | 0.2088 | |
| RedEdge | 0.5459 | 0.2807 | 0.0815 | 0.0503 | |
| LCI | 0.6736 | 0.3142 | 0.1734 | 0.0838 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, H.; Yang, C.; Guo, W.; Zhang, L.; Zhang, D. Automatic Estimation of Crop Disease Severity Levels Based on Vegetation Index Normalization. Remote Sens. 2020, 12, 1930. https://doi.org/10.3390/rs12121930
Zhao H, Yang C, Guo W, Zhang L, Zhang D. Automatic Estimation of Crop Disease Severity Levels Based on Vegetation Index Normalization. Remote Sensing. 2020; 12(12):1930. https://doi.org/10.3390/rs12121930
Chicago/Turabian StyleZhao, Hengqian, Chenghai Yang, Wei Guo, Lifu Zhang, and Dongyan Zhang. 2020. "Automatic Estimation of Crop Disease Severity Levels Based on Vegetation Index Normalization" Remote Sensing 12, no. 12: 1930. https://doi.org/10.3390/rs12121930
APA StyleZhao, H., Yang, C., Guo, W., Zhang, L., & Zhang, D. (2020). Automatic Estimation of Crop Disease Severity Levels Based on Vegetation Index Normalization. Remote Sensing, 12(12), 1930. https://doi.org/10.3390/rs12121930






