Fast Spectral Clustering for Unsupervised Hyperspectral Image Classification
Abstract
:1. Introduction
- An novel multiplicative update optimization for eigenvalue decomposition is proposed for large-scale unsupervised HSIs classification. It is worth noting that the proposed method can be easily portable to the variants of spectral clustering methods with different regularization items only if the constraints are convex functions.
- Two affinity matrix approximation techniques, namely the anchor-based graph and the Nyström extension, are introduced to improve the affinity matrix by sampling limited samples (i.e., pixels or anchors).
- Comprehensive experiments on the HSI datasets illustrated that the proposed method achieved a good result in terms of efficiency and effectiveness, and the combination of multiplicative update method and affinity matrix approximation provided a better performance.
2. Overview
2.1. Notation
2.2. Normalized Cuts Revisit
3. Improved Spectral Clustering with Multiplicative Update Algorithm
3.1. Formulation and Motivation
3.2. Multiplicative Update Optimization
| Algorithm 1: Algorithm to solve the problem in Equation (6). |
 |
4. Approximated Affinity Matrix
4.1. Affinity Matrix with Nyström Extension
4.2. Affinity Matrix with Anchor-Based Graph
5. Experiments
5.1. Experimental Datasets
- Salinas and Salinas-A were acquired by the 224-band AVIRIS sensor over Salinas Valley, California, and characterized by high spatial resolution (3.7-m pixels). Salinas covers 512 lines by 217 samples at as scale of . Salinas ground truth contains 16 classes. Salinas-A is an small subscene of Salinas image and it comprises pixels located within the same scene at [samples, lines] = [591–676, 158–240] and includes six classes.
- Pavia University is the scene collected by the ROSIS sensor during a flight campaign over Pavia, northern Italy. The number of spectral bands is 103 for Pavia University. Pavia University is a pixels image, where some pixels contain no information and these samples are discarded. Both hyperspectral image ground truths differentiate nine classes.
- Kennedy Space Center was acquired by the NASA AVIRIS instrument over the Kennedy Space Center (KSC), Florida, on 23 March 1996. They acquired data in 224 bands of 10 nm width with center wavelengths from 400 to 2500 nm and 176 bands were used for the analysis. KSC hyperspectral image contains pixels. For classification purposes, 13 classes representing the various land cover types that occur in this environment were defined for the site.
- Samson dataset is an image with 95 × 95 pixels and each pixel was recorded at 156 channels covering the wavelengths from 401 nm to 889 nm. The spectral resolution is high up to 3.13 nm and it is not degraded by blank or noisy channels. There are three targets in this image: Soil, Tree and Water.
- Japser Ridge is a hyperspectral image with pixels. Each pixel was recorded at 224 channels ranging from 380 nm to 2500 nm. The spectral resolution is up to 9.46 nm. There are four end-members latent in these data: Road, Soil, Water and Tree.
- Urban has 210 wavelengths ranging from 400 nm to 2500 nm, resulting in a spectral resolution of 10 nm. There are pixels, each of which corresponding to a m area. There are three versions of the ground truth, which contain 4, 5 and 6 end-members respectively, and are introduced in the ground truth.
- Indian Pines was gathered by AVIRIS sensor in northwestern Indiana and consists of pixels and 224 spectral reflectance bands. The Indian Pines scene contains two-thirds agriculture, and one-third forest or other natural perennial vegetation. The ground truth available is designated into sixteen classes and we reduced the number of bands to 200 by removing bands covering the region of water absorption.
5.2. Evaluation Metrics
- P. is the most common metric for clustering results evaluation and it can be formulated aswhere is the clustering result set and is the ground truth. The worst clustering result is very close to 0 and the best clustering result has a purity value equal to 1.
- NMI is a normalization of the mutual information score to scale the results between 0 and 1 aswhere denotes the number of data contained in the cluster , is the number of data belonging to the , and denotes the number of data that are in the intersection between the cluster and the class . The larger is the NMI, the better is the clustering result.
5.3. Toy Example
5.4. HSI Clustering Analysis
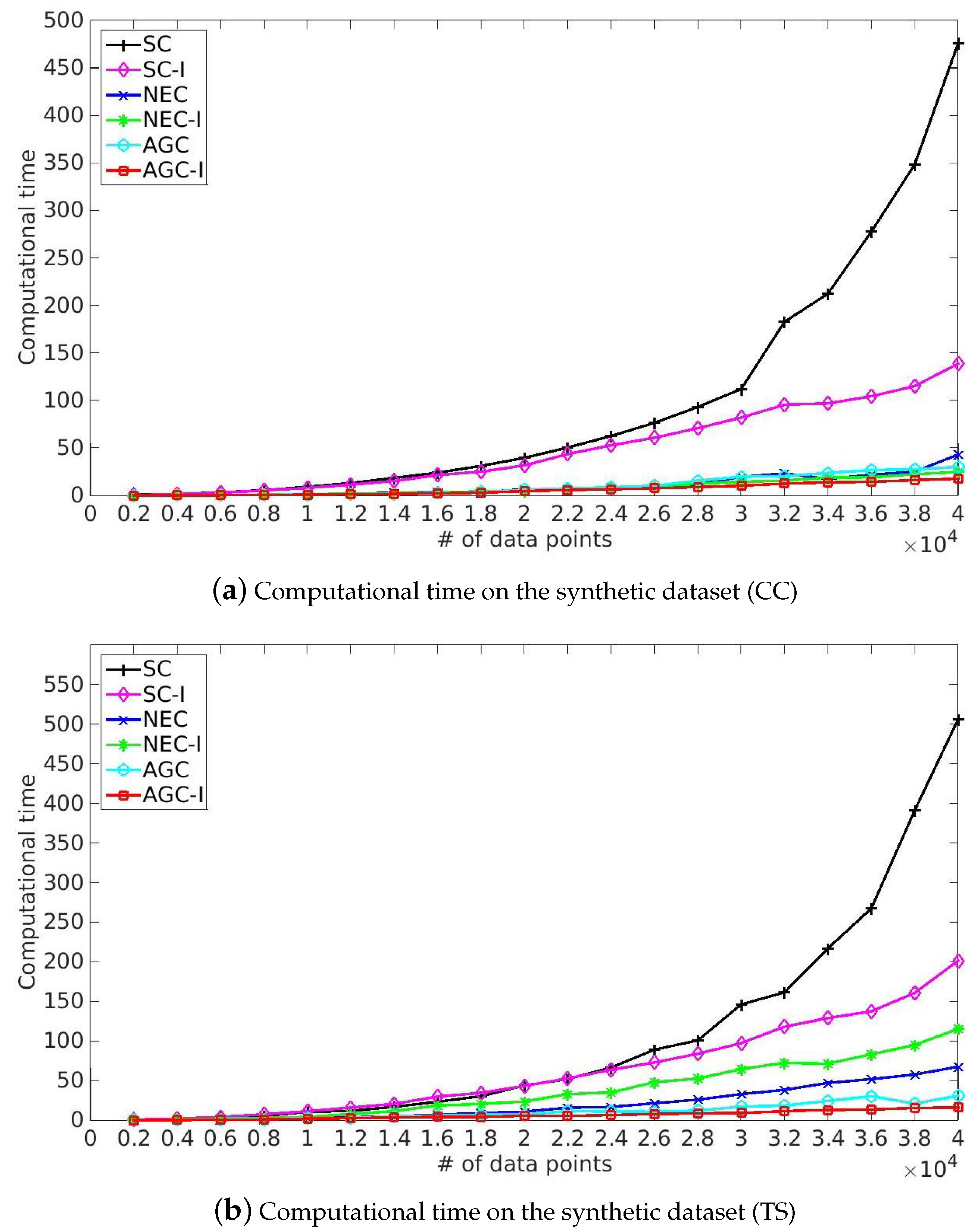
- SC and the corresponding improved algorithm SC-I achieved competitive performance in term of Purity and NMI. However, SC took more time solving eigenvalue decomposition of Laplacian matrix and our improved algorithm provided a more efficient solution because of the utilization of the multiplicative update optimization. Meanwhile, it took more time to process India Pines because of the rapid growth of time complexity of eigenvalue decomposition of Laplacian matrix caused by the increase of spatial resolution and classes. Note that SC-I, which is based on the multiplicative update algorithm, slightly outperformed SC in terms of Purity and NMI, illustrating that the nonnegative constraint and the orthonormal constraint provided a better indicator matrix. This made it easier to get a robust clustering result by the later processing, such as k-means.
- NEC and AGC are two efficient improved algorithms and they took only one-twentieth the time in our experiments. Moreover, NEC and AGC could be used on large-scale hyperspectral image datasets such as KSC and Urban, while SC ran out of memory in dealing with the above large-scale datasets because of the storage and time complexity of the affinity matrix. However, the experimental results also illustrate that NEC was not robust enough, which might be because the affinity matrix can be indefinite and the inverse matrix contains plural elements, making it difficult to get a robust clustering result by k-means. Besides NEC, the other methods did not struggle with this problem, and also provided a better performance than NEC.
- The proposed NEC-I and AGC-I outperformed the other methods in terms of effectiveness and efficiency. NEC-I and AGC-I firstly take the advantage of sample techniques including Nyström extension and anchor-based graph, which allow them to be used on large-scale hyperspectral image datasets. Furthermore, the proposed multiplicative update algorithm provided an efficient resolution for eigenvalue decomposition of Laplacian matrix. The results presented in Table 5 illustrate that NEC-I and AGC-I performed better than NEC and AGC in most cases. The proposed multiplicative update optimization is flexible and well-knit with the approximated affinity matrix such as Nyström extension and anchor-based graph.
5.5. Computational Time
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Wang, Q.; Wan, J.; Li, X. Robust Hierarchical Deep Learning for Vehicular Management. IEEE Trans. Veh. Technol. 2018. [Google Scholar] [CrossRef]
- Wang, Q.; Chen, M.; Nie, F.; Li, X. Detecting Coherent Groups in Crowd Scenes by Multiview Clustering. IEEE Trans. Pattern Anal. Mach. Intell. 2018. [Google Scholar] [CrossRef] [PubMed]
- Yan, Q.; Ding, Y.; Xia, Y.; Chong, Y.; Zheng, C. Class probability propagation of supervised information based on sparse subspace clustering for hyperspectral images. Remote Sens. 2017, 9, 1017. [Google Scholar] [CrossRef]
- Wang, Q.; Liu, S.; Chanussot, J.; Li, X. Scene classification with recurrent attention of vhr remote sensing images. IEEE Trans. Geosci. Remote Sens. 2018, 57, 1155–1167. [Google Scholar] [CrossRef]
- Wang, Q.; He, X.; Li, X. Locality and structure regularized low rank representation for hyperspectral image classification. IEEE Trans. Geosci. Remote Sens. 2018. [Google Scholar] [CrossRef]
- He, X.; Wang, Q.; Li, X. Spectral-spatial Hyperspectral image classification via locality and structure constrained low-rank representation. In Proceedings of the IEEE International Geoscience and Remote Sensing Symposium, Valencia, Spain, 23–27 July 2018. [Google Scholar]
- Wang, Q.; Meng, Z.; Li, X. Locality adaptive discriminant analysis for spectral-spatial classification of hyperspectral images. IEEE Geosci. Remote Sens. Lett. 2017, 14, 2077–2081. [Google Scholar] [CrossRef]
- Yuan, Y.; Lin, J.; Wang, Q. Hyperspectral image classification via multi-task joint sparse representation and stepwise mrf optimization. IEEE Trans. Cybern. 2016, 46, 2966–2977. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.; Yuan, Y.; Wang, Q. Hyperspectral anomaly detection via discriminative feature learning with multiple-dictionary sparse representation. Remote Sens. 2018, 10, 745. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, F.; Li, X. Optimal clustering framework for hyperspectral band selection. IEEE Trans. Geosci. Remote Sens. 2018, 56, 5910–5922. [Google Scholar] [CrossRef]
- Xie, H.; Zhao, A.; Huang, S.; Han, J.; Liu, S.; Xu, X.; Luo, X.; Pan, H.; Du, Q.; Tong, X. Unsupervised hyperspectral remote sensing image clustering based on adaptive density. IEEE Geosci. Remote Sens. Lett. 2018, 15, 632–636. [Google Scholar] [CrossRef]
- Chen, M.; Wang, Q.; Li, X. Discriminant analysis with graph learning for hyperspectral image classification. Remote Sens. 2018, 10, 836. [Google Scholar] [CrossRef]
- Fauvel, M.; Benediktsson, J.A.; Chanussot, J.; Sveinsson, J.R. Spectral and spatial classification of hyperspectral data using SVMs and morphological profiles. IEEE Trans. Geosci. Remote Sens. 2008, 46, 3804–3814. [Google Scholar] [CrossRef]
- Melgani, F.; Bruzzone, L. Classification of hyperspectral remote sensing images with support vector machines. IEEE Trans. Geosci. Remote Sens. 2004, 42, 1778–1790. [Google Scholar] [CrossRef]
- Rodarmel, C.; Shan, J. Principal component analysis for hyperspectral image classification. Surv. Land Inf. Sci. 2002, 62, 115–122. [Google Scholar]
- Wang, Q.; Lin, J.; Yuan, Y. Salient band selection for hyperspectral image classification via manifold ranking. IEEE Trans. Neural Netw. Learn. Syst. 2016, 27, 1279–1289. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Wan, J.; Nie, F.; Liu, B.; Yan, C.; Li, X. Hierarchical Feature Selection for Random Projection. IEEE Trans. Neural Netw. Learn. Syst. 2018. [Google Scholar] [CrossRef]
- Li, J.; Bioucas-Dias, J.M.; Plaza, A. Semisupervised hyperspectral image segmentation using multinomial logistic regression with active learning. IEEE Trans. Geosci. Remote Sens. 2010, 48, 4085–4098. [Google Scholar] [CrossRef]
- Chen, Y.; Nasrabadi, N.M.; Tran, T.D. Hyperspectral image classification using dictionary-based sparse representation. IEEE Trans. Geosci. Remote Sens. 2011, 49, 3973–3985. [Google Scholar] [CrossRef]
- Chen, Y.; Lin, Z.; Zhao, X.; Wang, G.; Gu, Y. Deep learning-based classification of hyperspectral data. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2014, 7, 2094–2107. [Google Scholar] [CrossRef]
- Zhang, H.; Zhai, H.; Zhang, L.; Li, P. Spectral-spatial sparse subspace clustering for hyperspectral remote sensing images. IEEE Trans. Geosci. Remote Sens. 2016, 54, 3672–3684. [Google Scholar] [CrossRef]
- Matasci, G.; Volpi, M.; Kanevski, M.; Bruzzone, L.; Tuia, D. Semisupervised transfer component analysis for domain adaptation in remote sensing image classification. IEEE Trans. Geosci. Remote Sens. 2015, 53, 3550–3564. [Google Scholar] [CrossRef]
- Crawford, M.M.; Tuia, D.; Yang, H.L. Active learning: Any value for classification of remotely sensed data? Proc. IEEE 2013, 101, 593–608. [Google Scholar] [CrossRef]
- Tuia, D.; Persello, C.; Bruzzone, L. Domain adaptation for the classification of remote sensing data: An overview of recent advances. IEEE Geosci. Remote Sens. Mag. 2016, 4, 41–57. [Google Scholar] [CrossRef]
- Guo, X.; Huang, X.; Zhang, L.; Zhang, L.; Plaza, A.; Benediktsson, J.A. Support tensor machines for classification of hyperspectral remote sensing imagery. IEEE Trans. Geosci. Remote Sens. 2016, 54, 3248–3264. [Google Scholar] [CrossRef]
- Wang, R.; Nie, F.; Yu, W. Fast spectral clustering with anchor graph for large hyperspectral images. IEEE Geosci. Remote Sens. Lett. 2017, 14, 2003–2007. [Google Scholar] [CrossRef]
- Nie, F.; Wang, X.; Jordan, M.I.; Huang, H. The constrained laplacian rank algorithm for graph-based clustering. In Proceedings of the 30th AAAI Conference on Artificial Intelligence, Phoenix, AZ, USA, 12–17 February 2016. [Google Scholar]
- Bezdek, J.C. Pattern recognition with fuzzy objective function algorithms. Adv. Appl. Pattern Recognit. 1981, 22, 203–239. [Google Scholar]
- Rodriguez, A.; Laio, A. Clustering by fast search and find of density peaks. Science 2014, 344, 1492–1496. [Google Scholar] [CrossRef]
- Buckley, J.J. Fuzzy hierarchical analysis. Fuzzy Sets Syst. 1985, 17, 233–247. [Google Scholar] [CrossRef]
- Vijendra, S. Efficient clustering for high dimensional data: Subspace based clustering and density based clustering. Inf. Technol. J. 2011, 10, 1092–1105. [Google Scholar] [CrossRef]
- Zhong, Y.; Zhang, L.; Gong, W. Unsupervised remote sensing image classification using an artificial immune network. Int. J. Remote Sens. 2011, 32, 5461–5483. [Google Scholar] [CrossRef]
- Zhong, Y.; Zhang, S.; Zhang, L. Automatic fuzzy clustering based on adaptive multi-objective differential evolution for remote sensing imagery. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2013, 6, 2290–2301. [Google Scholar] [CrossRef]
- Zhang, L.; You, J. A spectral clustering based method for hyperspectral urban image. In Proceedings of the 2017 Joint Urban Remote Sensing Event (JURSE), Dubai, UAE, 6–8 March 2017; pp. 1–3. [Google Scholar]
- Zhao, Y.; Yuan, Y.; Nie, F.; Wang, Q. Spectral clustering based on iterative optimization for large-scale and high-dimensional data. Neurocomputing 2018, 318, 227–235. [Google Scholar] [CrossRef]
- Bai, J.; Xiang, S.; Pan, C. A graph-based classification method for hyperspectral images. IEEE Trans. Geosci. Remote Sens. 2013, 51, 803–817. [Google Scholar] [CrossRef]
- Camps-Valls, G.; Marsheva, T.V.B.; Zhou, D. Semi-supervised graph-based hyperspectral image classification. IEEE Trans. Geosci. Remote Sens. 2007, 45, 3044–3054. [Google Scholar] [CrossRef]
- Wang, Q.; Qin, Z.; Nie, F.; Li, X. Spectral Embedded Adaptive Neighbors Clustering. IEEE Trans. Neural Netw. Learn. Syst. 2018. [Google Scholar] [CrossRef] [PubMed]
- Belongie, S.; Fowlkes, C.; Chung, F.; Malik, J. Spectral partitioning with indefinite kernels using the Nyström extension. In Proceedings of the European Conference on Computer Vision, Copenhagen, Denmark, 28–31 May 2002; pp. 531–542. [Google Scholar]
- Fowlkes, C.; Belongie, S.; Chung, F.; Malik, J. Spectral grouping using the Nystrom method. IEEE Trans. Pattern Anal. Mach. Intell. 2004, 26, 214–225. [Google Scholar] [CrossRef]
- Tang, X.; Jiao, L.; Emery, W.J.; Liu, F.; Zhang, D. Two-stage reranking for remote sensing image retrieval. IEEE Trans. Geosci. Remote Sens. 2017, 55, 5798–5817. [Google Scholar] [CrossRef]
- Zhang, X.; Jiao, L.; Liu, F.; Bo, L.; Gong, M. Spectral clustering ensemble applied to sar image segmentation. IEEE Trans. Geosci. Remote Sens. 2008, 46, 2126–2136. [Google Scholar] [CrossRef]
- Zhu, W.; Nie, F.; Li, X. Fast Spectral Clustering with efficient large graph construction. In Proceedings of the IEEE International Conference on Speech and Signal Processing, New Orleans, LA, USA, 5–9 March 2017; pp. 2492–2496. [Google Scholar]
- Nie, F.; Zhu, W.; Li, X. Unsupervised Large Graph Embedding. In Proceedings of the 31st AAAI Conference on Artificial Intelligence, San Francisco, CA, USA, 4–9 February 2017; pp. 2422–2428. [Google Scholar]
- Yokoya, N.; Yairi, T.; Iwasaki, A. Coupled nonnegative matrix factorization unmixing for hyperspectral and multispectral data fusion. IEEE Trans. Geosci. Remote Sens. 2012, 50, 528–537. [Google Scholar] [CrossRef]
- Jia, S.; Qian, Y. Constrained nonnegative matrix factorization for hyperspectral unmixing. IEEE Trans. Geosci. Remote Sens. 2009, 47, 161–173. [Google Scholar] [CrossRef]
- Shi, J.; Malik, J. Normalized cuts and image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2000, 22, 888–905. [Google Scholar]
- Nie, F.; Ding, C.; Luo, D.; Huang, H. Improved minmax cut graph clustering with nonnegative relaxation. In Proceedings of the Joint European Conference on Machine Learning and Knowledge Discovery in Databases, Barcelona, Spain, 20–24 September 2010; pp. 451–466. [Google Scholar]
- Türkmen, A.C. A review of nonnegative matrix factorization methods for clustering. Comput. Sci. 2015, 1, 405–408. [Google Scholar]
- Fowlkes, C.; Belongie, S.; Malik, J. Efficient spatiotemporal grouping using the Nyström method. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Kauai, HI, USA, 8–14 December 2001. [Google Scholar]
- Liu, W.; He, J.; Chang, S.F. Large Graph Construction for Scalable Semi-Supervised Learning. In Proceedings of the 27th International Conference on Machine Learning, Haifa, Israel, 22–24 June 2010. [Google Scholar]



| W | Affinity (or similarity) matrix |
| Diagonal matrix | |
| Laplacian matrix | |
| Cluster indicator matrix | |
| Cluster indicator | |
| Pixels (or data points) | |
| Identity matrix | |
| n | Number of pixels |
| m | Number of chosen pixels (or anchors) |
| d | Number of spectral bands |
| c | Number of classes |
| SC | SC-I | NEC | NEC-I | AGC | AGC-I | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | |
| Num. = 2000 | 1.00 | 1.00 | 1.02 | 1.00 | 1.00 | 0.35 | 0.68 | 0.25 | 0.05 | 1.00 | 1.00 | 0.08 | 1.00 | 1.00 | 0.05 | 1.00 | 1.00 | 0.06 |
| Num. = 4000 | 1.00 | 1.00 | 1.45 | 1.00 | 1.00 | 1.35 | 0.65 | 0.09 | 0.14 | 1.00 | 1.00 | 0.17 | 1.00 | 1.00 | 0.15 | 1.00 | 1.00 | 0.14 |
| Num. = 6000 | 1.00 | 1.00 | 3.12 | 1.00 | 1.00 | 2.94 | 0.69 | 0.26 | 0.36 | 1.00 | 1.00 | 0.48 | 1.00 | 1.00 | 0.26 | 1.00 | 1.00 | 0.29 |
| Num. = 8000 | 1.00 | 1.00 | 5.53 | 1.00 | 1.00 | 5.23 | 0.67 | 0.25 | 0.68 | 1.00 | 1.00 | 0.81 | 1.00 | 1.00 | 0.56 | 1.00 | 1.00 | 0.54 |
| Num. = 10,000 | 1.00 | 1.00 | 9.05 | 1.00 | 1.00 | 7.87 | 0.68 | 0.25 | 0.81 | 1.00 | 1.00 | 1.25 | 1.00 | 1.00 | 0.69 | 1.00 | 1.00 | 0.86 |
| Num. = 12,000 | 1.00 | 1.00 | 13.04 | 1.00 | 1.00 | 11.35 | 0.50 | 0.00 | 1.91 | 1.00 | 1.00 | 1.89 | 1.00 | 1.00 | 0.83 | 1.00 | 1.00 | 1.20 |
| Num. = 14,000 | 1.00 | 1.00 | 18.39 | 1.00 | 1.00 | 15.23 | 0.57 | 0.02 | 2.67 | 1.00 | 1.00 | 2.32 | 1.00 | 1.00 | 1.13 | 1.00 | 1.00 | 1.62 |
| Num. = 16,000 | 1.00 | 1.00 | 23.99 | 1.00 | 1.00 | 21.44 | 0.52 | 0.00 | 3.66 | 1.00 | 1.00 | 3.01 | 1.00 | 1.00 | 1.33 | 1.00 | 1.00 | 2.17 |
| Num. = 18,000 | 1.00 | 1.00 | 31.05 | 1.00 | 1.00 | 25.00 | 0.63 | 0.19 | 3.25 | 1.00 | 1.00 | 4.42 | 1.00 | 1.00 | 1.85 | 1.00 | 1.00 | 2.87 |
| Num. = 20,000 | 1.00 | 1.00 | 39.52 | 1.00 | 1.00 | 31.52 | 0.69 | 0.27 | 6.59 | 1.00 | 1.00 | 4.55 | 1.00 | 1.00 | 2.23 | 1.00 | 1.00 | 4.58 |
| Num. = 22,000 | 1.00 | 1.00 | 50.36 | 1.00 | 1.00 | 43.48 | 0.50 | 0.00 | 5.58 | 1.00 | 1.00 | 7.19 | 1.00 | 1.00 | 3.06 | 1.00 | 1.00 | 5.57 |
| Num. = 24,000 | 1.00 | 1.00 | 62.55 | 1.00 | 1.00 | 52.81 | 0.54 | 0.00 | 7.13 | 1.00 | 1.00 | 8.40 | 1.00 | 1.00 | 3.79 | 1.00 | 1.00 | 6.54 |
| Num. = 26,000 | 1.00 | 1.00 | 76.38 | 1.00 | 1.00 | 60.66 | 0.53 | 0.00 | 9.17 | 1.00 | 1.00 | 8.88 | 1.00 | 1.00 | 4.54 | 1.00 | 1.00 | 7.57 |
| Num. = 28,000 | 1.00 | 1.00 | 93.06 | 1.00 | 1.00 | 70.78 | 0.69 | 0.26 | 11.59 | 1.00 | 1.00 | 12.34 | 0.83 | 0.47 | 5.45 | 1.00 | 1.00 | 8.78 |
| Num. = 30,000 | 1.00 | 1.00 | 111.98 | 1.00 | 1.00 | 81.95 | 0.74 | 0.28 | 19.52 | 1.00 | 1.00 | 14.34 | 1.00 | 1.00 | 8.12 | 1.00 | 1.00 | 10.31 |
| Num. = 32,000 | 1.00 | 1.00 | 182.78 | 1.00 | 1.00 | 95.47 | 0.59 | 0.15 | 23.14 | 1.00 | 1.00 | 15.63 | 0.83 | 0.48 | 10.01 | 1.00 | 1.00 | 12.43 |
| Num. = 34,000 | 1.00 | 1.00 | 212.34 | 1.00 | 1.00 | 96.86 | 0.63 | 0.20 | 17.35 | 1.00 | 1.00 | 18.90 | 1.00 | 1.00 | 10.30 | 1.00 | 1.00 | 13.72 |
| Num. = 36,000 | 1.00 | 1.00 | 277.53 | 1.00 | 1.00 | 104.32 | 0.51 | 0.00 | 21.86 | 1.00 | 1.00 | 19.13 | 1.00 | 1.00 | 31.71 | 1.00 | 1.00 | 14.41 |
| Num. = 38,000 | 1.00 | 1.00 | 348.30 | 1.00 | 1.00 | 115.03 | 0.50 | 0.00 | 24.99 | 1.00 | 1.00 | 22.33 | 1.00 | 1.00 | 23.17 | 1.00 | 1.00 | 16.07 |
| Num. = 40,000 | 1.00 | 1.00 | 475.56 | 1.00 | 1.00 | 138.64 | 0.57 | 0.12 | 43.01 | 1.00 | 1.00 | 24.66 | 1.00 | 1.00 | 18.09 | 1.00 | 1.00 | 17.70 |
| Average | 1.00 | 1.00 | 101.85 | 1.00 | 1.00 | 49.11 | 0.60 | 0.13 | 10.17 | 1.00 | 1.00 | 8.54 | 0.98 | 0.95 | 6.37 | 1.00 | 1.00 | 6.37 |
| SC | SC-I | NEC | NEC-I | AGC | AGC-I | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | |
| Num. = 2000 | 0.97 | 0.83 | 1.01 | 1.00 | 0.98 | 0.50 | 0.50 | 0.01 | 0.08 | 0.99 | 0.93 | 0.17 | 1.00 | 1.00 | 0.82 | 0.95 | 0.71 | 0.14 |
| Num. = 4000 | 0.98 | 0.85 | 1.40 | 0.98 | 0.88 | 1.92 | 0.73 | 0.22 | 0.38 | 1.00 | 0.96 | 0.87 | 1.00 | 1.00 | 0.40 | 0.98 | 0.87 | 0.24 |
| Num. = 6000 | 0.97 | 0.83 | 2.95 | 0.97 | 0.82 | 3.98 | 0.50 | 0.00 | 0.68 | 1.00 | 0.98 | 1.76 | 1.00 | 1.00 | 0.87 | 0.99 | 0.93 | 0.43 |
| Num. = 8000 | 0.97 | 0.79 | 5.41 | 0.99 | 0.93 | 7.40 | 0.50 | 0.00 | 1.26 | 0.99 | 0.94 | 2.63 | 1.00 | 1.00 | 1.04 | 1.00 | 0.96 | 0.97 |
| Num. = 10,000 | 0.97 | 0.81 | 8.37 | 0.99 | 0.95 | 11.27 | 0.50 | 0.00 | 2.35 | 1.00 | 0.95 | 4.09 | 1.00 | 1.00 | 2.68 | 0.87 | 0.45 | 1.90 |
| Num. = 12,000 | 0.97 | 0.79 | 13.39 | 0.99 | 0.94 | 15.81 | 0.50 | 0.00 | 3.46 | 0.99 | 0.95 | 7.65 | 1.00 | 1.00 | 2.75 | 0.95 | 0.71 | 2.41 |
| Num. = 14,000 | 0.97 | 0.79 | 18.70 | 0.98 | 0.89 | 20.61 | 0.71 | 0.29 | 5.01 | 0.99 | 0.91 | 11.43 | 1.00 | 1.00 | 4.77 | 0.98 | 0.87 | 3.31 |
| Num. = 16,000 | 0.97 | 0.79 | 26.49 | 0.83 | 0.35 | 29.71 | 0.51 | 0.03 | 6.79 | 0.96 | 0.80 | 18.55 | 1.00 | 1.00 | 5.68 | 0.99 | 0.93 | 4.14 |
| Num. = 18,000 | 0.97 | 0.81 | 32.98 | 0.99 | 0.92 | 34.42 | 0.68 | 0.25 | 8.97 | 0.99 | 0.95 | 20.36 | 1.00 | 1.00 | 6.82 | 1.00 | 0.96 | 4.03 |
| Num. = 20,000 | 0.97 | 0.82 | 43.03 | 0.99 | 0.90 | 43.23 | 0.50 | 0.00 | 10.82 | 0.99 | 0.93 | 23.61 | 1.00 | 1.00 | 6.96 | 0.92 | 0.59 | 5.59 |
| Num. = 22,000 | 0.97 | 0.79 | 55.62 | 0.99 | 0.93 | 52.66 | 0.72 | 0.30 | 15.71 | 0.99 | 0.94 | 32.81 | 1.00 | 1.00 | 11.39 | 0.95 | 0.71 | 5.30 |
| Num. = 24,000 | 0.97 | 0.80 | 72.02 | 0.99 | 0.95 | 63.61 | 0.52 | 0.01 | 16.68 | 0.99 | 0.93 | 34.86 | 1.00 | 1.00 | 10.94 | 0.98 | 0.87 | 6.48 |
| Num. = 26,000 | 0.97 | 0.80 | 85.32 | 0.99 | 0.94 | 72.83 | 0.53 | 0.03 | 21.37 | 0.99 | 0.91 | 48.00 | 1.00 | 1.00 | 10.66 | 0.99 | 0.93 | 7.71 |
| Num. = 28,000 | 0.97 | 0.80 | 102.27 | 0.99 | 0.95 | 83.75 | 0.50 | 0.00 | 25.86 | 0.99 | 0.95 | 52.51 | 1.00 | 1.00 | 11.99 | 1.00 | 0.96 | 8.52 |
| Num. = 30,000 | 0.97 | 0.81 | 149.99 | 1.00 | 0.98 | 97.31 | 0.51 | 0.03 | 32.83 | 0.99 | 0.94 | 64.45 | 1.00 | 1.00 | 17.10 | 1.00 | 1.00 | 9.06 |
| Num. = 32,000 | 0.97 | 0.81 | 190.72 | 0.99 | 0.93 | 118.01 | 0.50 | 0.00 | 38.24 | 0.99 | 0.94 | 72.44 | 1.00 | 1.00 | 18.38 | 1.00 | 0.98 | 11.40 |
| Num. = 34,000 | 0.97 | 0.81 | 258.66 | 0.98 | 0.88 | 128.90 | 0.51 | 0.03 | 47.00 | 0.99 | 0.95 | 71.32 | 1.00 | 1.00 | 24.32 | 0.91 | 0.57 | 12.86 |
| Num. = 36,000 | 0.97 | 0.81 | 358.37 | 0.98 | 0.86 | 137.45 | 0.50 | 0.00 | 51.82 | 0.99 | 0.94 | 83.11 | 1.00 | 1.00 | 30.24 | 0.98 | 0.89 | 13.48 |
| Num. = 38,000 | 0.97 | 0.80 | 459.32 | 0.97 | 0.82 | 160.64 | 0.50 | 0.00 | 57.57 | 0.68 | 0.10 | 94.87 | 1.00 | 1.00 | 20.89 | 0.98 | 0.04 | 15.44 |
| Num. = 40,000 | 0.97 | 0.81 | 636.23 | 0.99 | 0.94 | 201.30 | 0.50 | 0.00 | 67.46 | 1.00 | 0.97 | 115.24 | 1.00 | 1.00 | 30.73 | 1.00 | 1.00 | 15.88 |
| Average | 0.97 | 0.81 | 126.31 | 0.98 | 0.89 | 64.27 | 0.55 | 0.06 | 20.72 | 0.98 | 0.89 | 38.04 | 1.00 | 1.00 | 10.97 | 0.97 | 0.80 | 6.46 |
| SC | SC-I | NEC | NEC-I | AGC | AGC-I | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | P. | NMI | CT | |
| Num. = 2000 | 1.00 | 1.00 | 0.38 | 1.00 | 0.98 | 0.70 | 0.50 | 0.00 | 0.09 | 1.00 | 1.00 | 0.17 | 0.56 | 0.19 | 0.86 | 1.00 | 1.00 | 0.08 |
| Num. = 4000 | 1.00 | 1.00 | 1.34 | 0.99 | 0.92 | 2.43 | 0.50 | 0.00 | 1.50 | 1.00 | 1.00 | 0.85 | 1.00 | 1.00 | 0.41 | 1.00 | 1.00 | 0.29 |
| Num. = 6000 | 1.00 | 1.00 | 2.70 | 0.99 | 0.90 | 5.22 | 0.50 | 0.00 | 1.08 | 1.00 | 1.00 | 1.63 | 1.00 | 1.00 | 1.14 | 1.00 | 1.00 | 0.81 |
| Num. = 8000 | 1.00 | 1.00 | 5.71 | 0.99 | 0.91 | 9.64 | 0.89 | 0.53 | 1.90 | 1.00 | 1.00 | 3.22 | 1.00 | 1.00 | 2.13 | 1.00 | 1.00 | 1.48 |
| Num. = 10,000 | 1.00 | 1.00 | 8.46 | 0.99 | 0.95 | 15.39 | 0.50 | 0.00 | 3.05 | 1.00 | 1.00 | 5.32 | 1.00 | 1.00 | 2.20 | 1.00 | 1.00 | 2.34 |
| Num. = 12,000 | 1.00 | 1.00 | 12.55 | 0.99 | 0.95 | 21.76 | 0.50 | 0.01 | 4.70 | 1.00 | 1.00 | 8.24 | 1.00 | 1.00 | 3.73 | 1.00 | 1.00 | 3.42 |
| Num. = 14,000 | 1.00 | 1.00 | 18.24 | 0.99 | 0.94 | 27.11 | 0.50 | 0.00 | 8.47 | 1.00 | 1.00 | 12.49 | 1.00 | 1.00 | 3.74 | 1.00 | 1.00 | 4.64 |
| Num. = 16,000 | 1.00 | 1.00 | 26.76 | 0.93 | 0.63 | 39.33 | 0.50 | 0.00 | 10.85 | 1.00 | 1.00 | 17.26 | 1.00 | 1.00 | 4.50 | 1.00 | 1.00 | 6.28 |
| Num. = 18,000 | 1.00 | 1.00 | 34.21 | 0.99 | 0.92 | 44.15 | 0.90 | 0.55 | 15.68 | 1.00 | 1.00 | 20.80 | 1.00 | 1.00 | 6.51 | 1.00 | 1.00 | 7.63 |
| Num. = 20,000 | 1.00 | 1.00 | 43.86 | 0.99 | 0.92 | 57.08 | 0.50 | 0.00 | 21.38 | 1.00 | 1.00 | 25.60 | 1.00 | 1.00 | 7.78 | 1.00 | 1.00 | 9.60 |
| Num. = 22,000 | 1.00 | 1.00 | 55.55 | 0.99 | 0.94 | 72.23 | 0.50 | 0.00 | 27.26 | 1.00 | 1.00 | 33.16 | 1.00 | 1.00 | 8.48 | 1.00 | 1.00 | 11.20 |
| Num. = 24,000 | 1.00 | 1.00 | 69.45 | 0.99 | 0.95 | 86.58 | 0.68 | 0.25 | 27.87 | 1.00 | 1.00 | 38.49 | 1.00 | 1.00 | 8.98 | 1.00 | 1.00 | 13.34 |
| Num. = 26,000 | 1.00 | 1.00 | 101.07 | 0.99 | 0.95 | 99.36 | 0.50 | 0.01 | 48.77 | 1.00 | 1.00 | 44.62 | 1.00 | 1.00 | 11.41 | 1.00 | 1.00 | 15.60 |
| Num. = 28,000 | 1.00 | 1.00 | 114.92 | 0.99 | 0.95 | 114.37 | 0.50 | 0.00 | 39.56 | 1.00 | 1.00 | 55.30 | 1.00 | 1.00 | 11.83 | 1.00 | 1.00 | 17.92 |
| Num. = 30,000 | 1.00 | 1.00 | 149.53 | 1.00 | 0.96 | 136.30 | 0.91 | 0.59 | 76.80 | 1.00 | 1.00 | 63.81 | 1.00 | 1.00 | 14.98 | 1.00 | 1.00 | 21.26 |
| Num. = 32,000 | 1.00 | 1.00 | 209.87 | 0.99 | 0.93 | 158.11 | 0.85 | 0.49 | 84.71 | 1.00 | 1.00 | 67.09 | 1.00 | 1.00 | 16.84 | 1.00 | 1.00 | 27.18 |
| Num. = 34,000 | 1.00 | 1.00 | 270.50 | 0.99 | 0.91 | 167.20 | 0.50 | 0.00 | 82.73 | 1.00 | 1.00 | 79.64 | 1.00 | 1.00 | 20.62 | 1.00 | 1.00 | 28.29 |
| Num. = 36,000 | 1.00 | 1.00 | 381.08 | 0.99 | 0.91 | 181.04 | 0.62 | 0.04 | 76.13 | 1.00 | 1.00 | 89.44 | 1.00 | 1.00 | 20.62 | 1.00 | 1.00 | 30.00 |
| Num. = 38,000 | 1.00 | 1.00 | 594.83 | 0.96 | 0.75 | 223.33 | 0.77 | 0.37 | 99.99 | 1.00 | 1.00 | 100.11 | 1.00 | 1.00 | 21.47 | 1.00 | 1.00 | 33.28 |
| Num. = 40,000 | 1.00 | 1.00 | 754.12 | 0.99 | 0.93 | 258.65 | 0.50 | 0.00 | 108.56 | 1.00 | 1.00 | 119.93 | 1.00 | 1.00 | 32.86 | 1.00 | 1.00 | 36.46 |
| Average | 1.00 | 1.00 | 142.76 | 0.99 | 0.91 | 86.00 | 0.61 | 0.14 | 37.05 | 1.00 | 1.00 | 39.36 | 0.98 | 0.96 | 10.05 | 1.00 | 1.00 | 13.55 |
| SC | SC-I | NEC | NEC-I | AGC | AGC-I | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PUI. | NMI | CT | PUI. | NMI | CT | PUI. | NMI | CT | PUI. | NMI | CT | PUI. | NMI | CT | PUI. | NMI | CT | |
| Samson | 0.85 | 0.61 | 6.57 | 0.85 | 0.60 | 5.77 | 0.73 | 0.53 | 0.10 | 0.85 | 0.60 | 0.17 | 0.88 | 0.73 | 0.19 | 0.91 | 0.75 | 0.19 |
| Jasper | 0.83 | 0.71 | 10.31 | 0.91 | 0.76 | 6.43 | 0.70 | 0.56 | 0.03 | 0.83 | 0.71 | 0.11 | 0.72 | 0.66 | 0.09 | 0.82 | 0.70 | 0.14 |
| SalinasA | 0.81 | 0.80 | 4.77 | 0.85 | 0.79 | 4.31 | 0.78 | 0.77 | 0.06 | 0.80 | 0.81 | 0.17 | 0.79 | 0.78 | 0.10 | 0.84 | 0.81 | 0.15 |
| India Pines | 0.36 | 0.44 | 66.21 | 0.46 | 0.46 | 45.37 | 0.43 | 0.45 | 0.53 | 0.43 | 0.49 | 1.29 | 0.35 | 0.43 | 0.58 | 0.42 | 0.46 | 1.46 |
| Salinas | - | - | - | - | - | - | 0.60 | 0.72 | 1.62 | 0.62 | 0.71 | 4.62 | 0.56 | 0.67 | 2.44 | 0.56 | 0.71 | 3.55 |
| Pavia Uni. | - | - | - | - | - | - | 0.47 | 0.34 | 1.34 | 0.61 | 0.57 | 3.34 | 0.46 | 0.51 | 3.40 | 0.54 | 0.57 | 3.67 |
| KSC | - | - | - | - | - | - | 0.46 | 0.57 | 1.16 | 0.51 | 0.52 | 5.97 | 0.47 | 0.52 | 6.10 | 0.51 | 0.53 | 6.48 |
| Urban | - | - | - | - | - | - | 0.40 | 0.12 | 0.41 | 0.45 | 0.21 | 3.01 | 0.51 | 0.33 | 1.14 | 0.50 | 0.29 | 3.12 |
| Average | 0.68 | 0.58 | 27.69 | 0.74 | 0.60 | 19.19 | 0.57 | 0.51 | 0.66 | 0.64 | 0.58 | 2.34 | 0.59 | 0.58 | 1.76 | 0.64 | 0.60 | 2.35 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, Y.; Yuan, Y.; Wang, Q. Fast Spectral Clustering for Unsupervised Hyperspectral Image Classification. Remote Sens. 2019, 11, 399. https://doi.org/10.3390/rs11040399
Zhao Y, Yuan Y, Wang Q. Fast Spectral Clustering for Unsupervised Hyperspectral Image Classification. Remote Sensing. 2019; 11(4):399. https://doi.org/10.3390/rs11040399
Chicago/Turabian StyleZhao, Yang, Yuan Yuan, and Qi Wang. 2019. "Fast Spectral Clustering for Unsupervised Hyperspectral Image Classification" Remote Sensing 11, no. 4: 399. https://doi.org/10.3390/rs11040399
APA StyleZhao, Y., Yuan, Y., & Wang, Q. (2019). Fast Spectral Clustering for Unsupervised Hyperspectral Image Classification. Remote Sensing, 11(4), 399. https://doi.org/10.3390/rs11040399






