Finite Difference Analysis and Bivariate Correlation of Hyperspectral Data for Detecting Laurel Wilt Disease and Nutritional Deficiency in Avocado
Abstract
1. Introduction
2. Materials and Methods
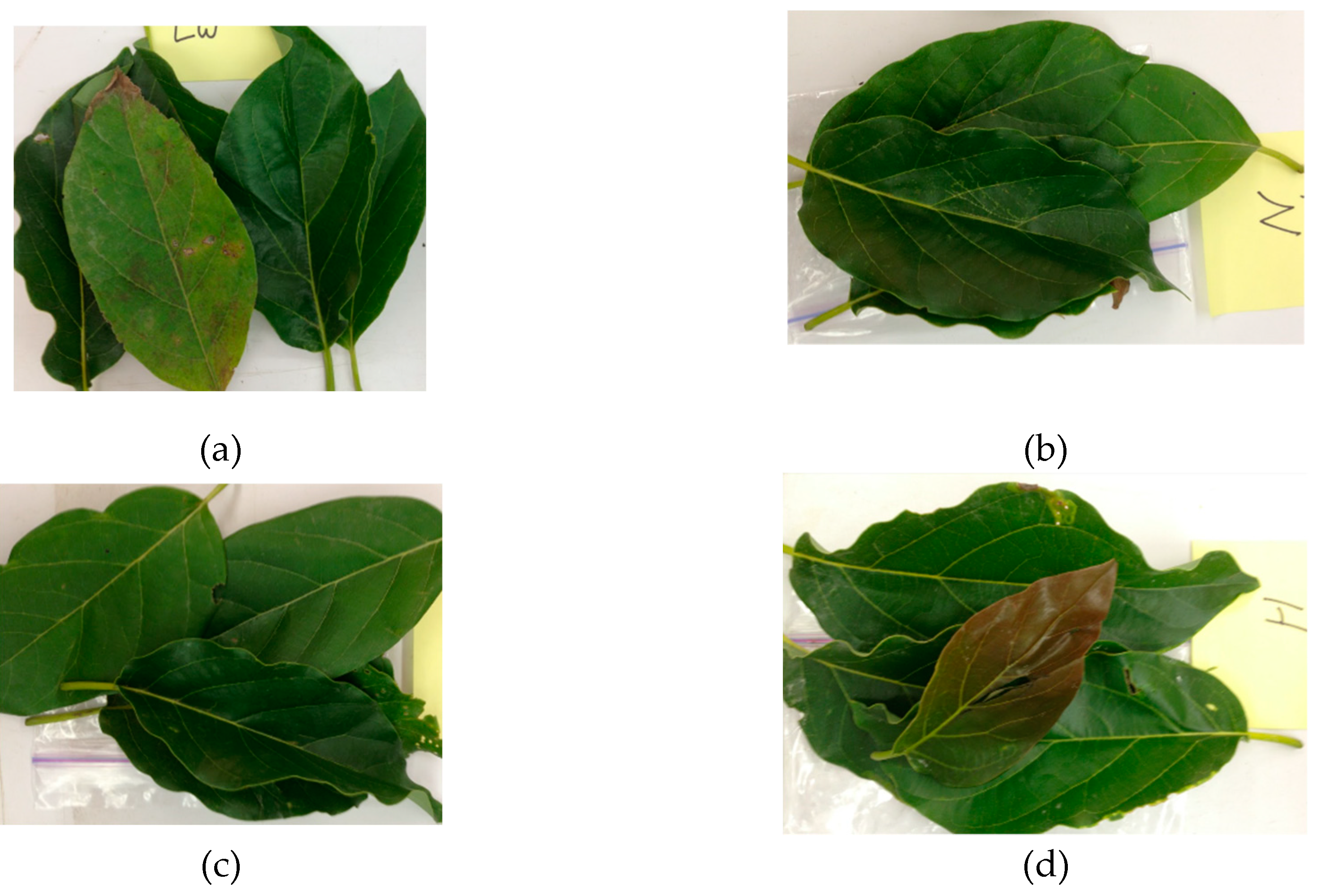
2.1. Plant and Sample Selection
2.2. Spectral Data Collection
2.3. Higher Order Spectra Enhancement Methodology
2.3.1. Higher-Order Spectral Analysis
2.3.2. Bivariate Correlation Process
3. Results
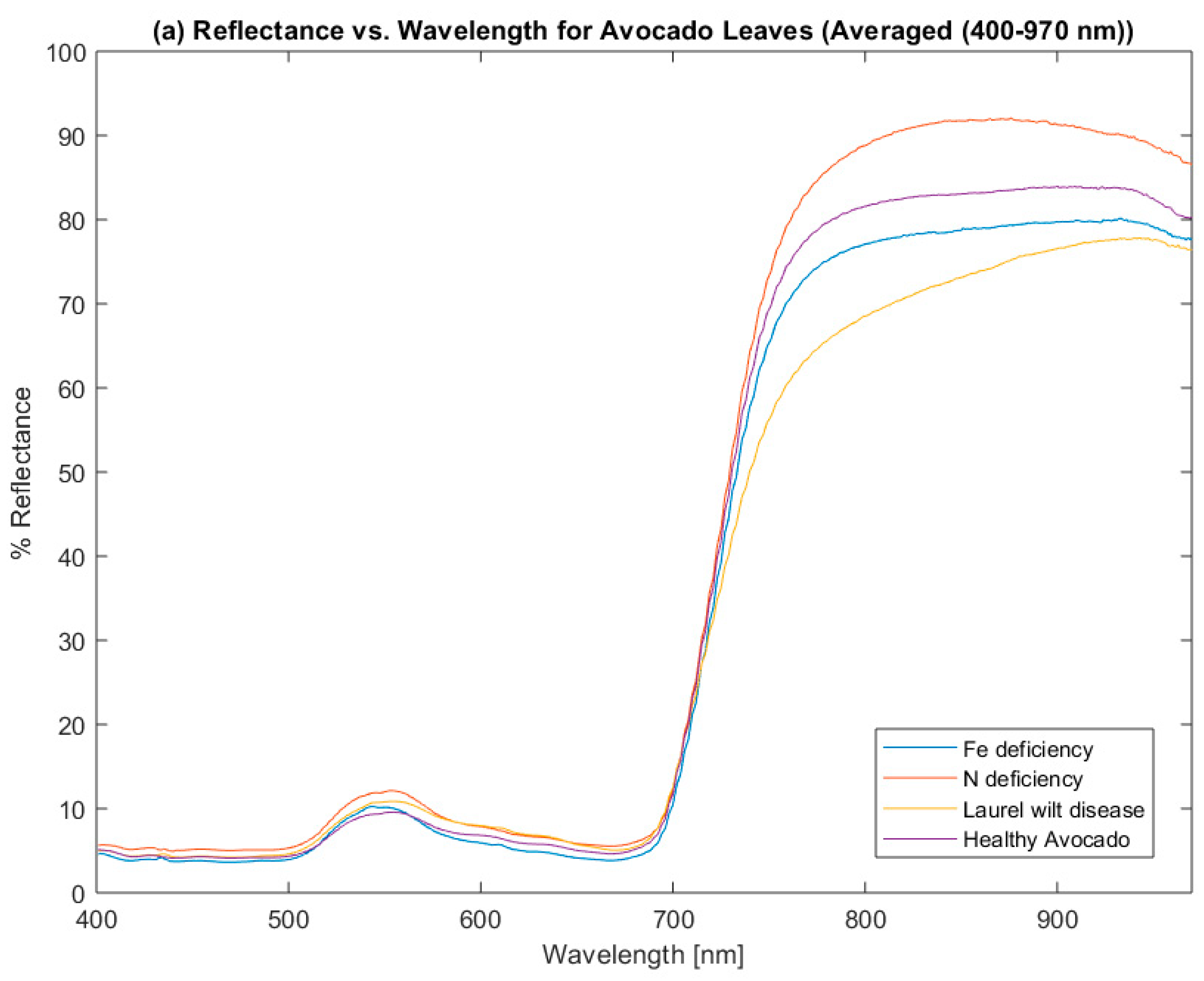
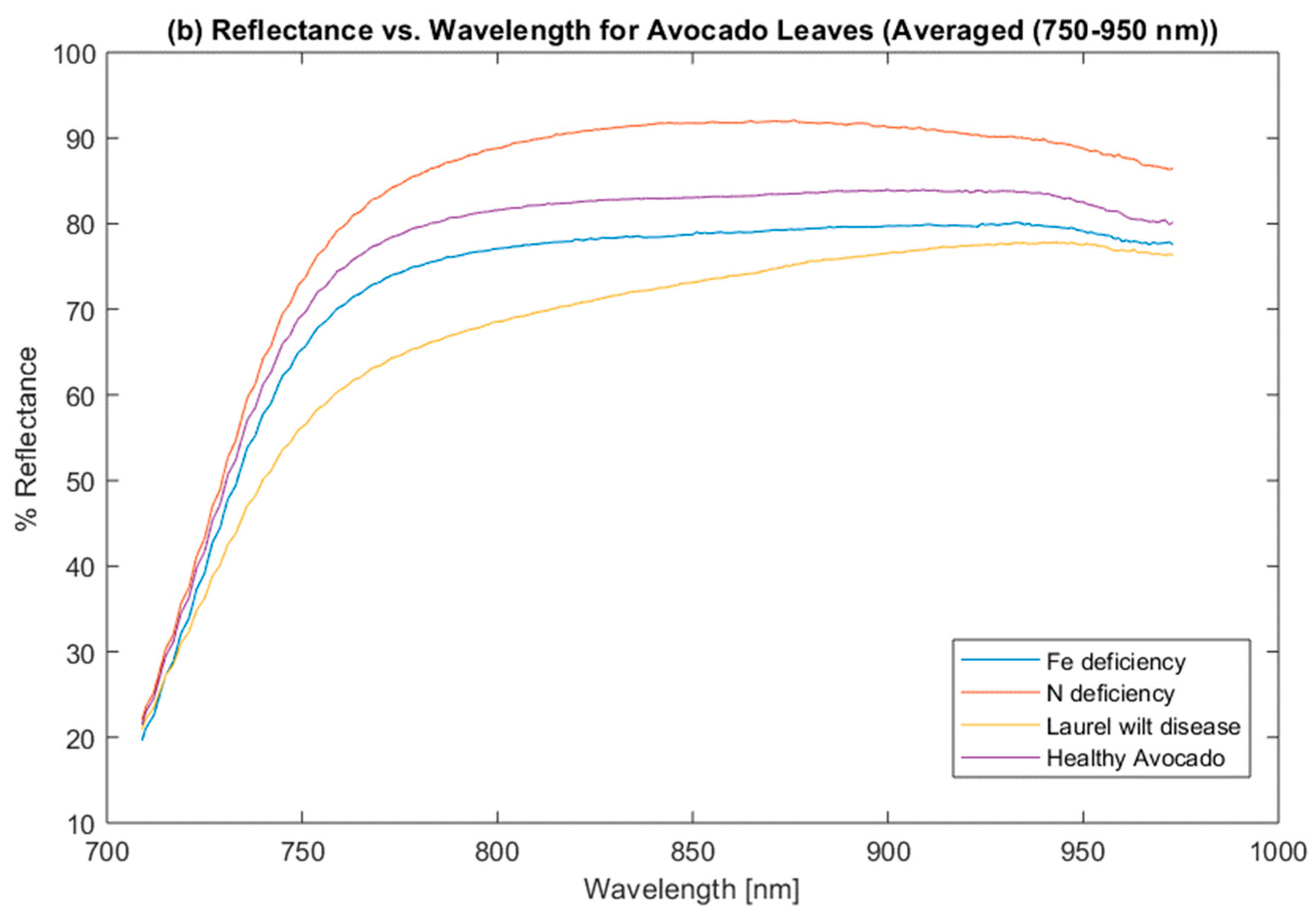
3.1. Data Analysis
3.2. Applied FDA and Bivariate Correlation Process
3.2.1. Fe-Deficient Classification Results
3.2.2. N-Deficient Classification Results
3.2.3. Lw-Disease Classification Results
3.2.4. Healthy Avocado Classification Results
3.3. Analysis of Classification Results
- (i)
- Normalization of the hyperspectral data.
- (ii)
- Polynomial fitting of data.
- (iii)
- Smoothing the data by moving median with absolute deviation.
- (iv)
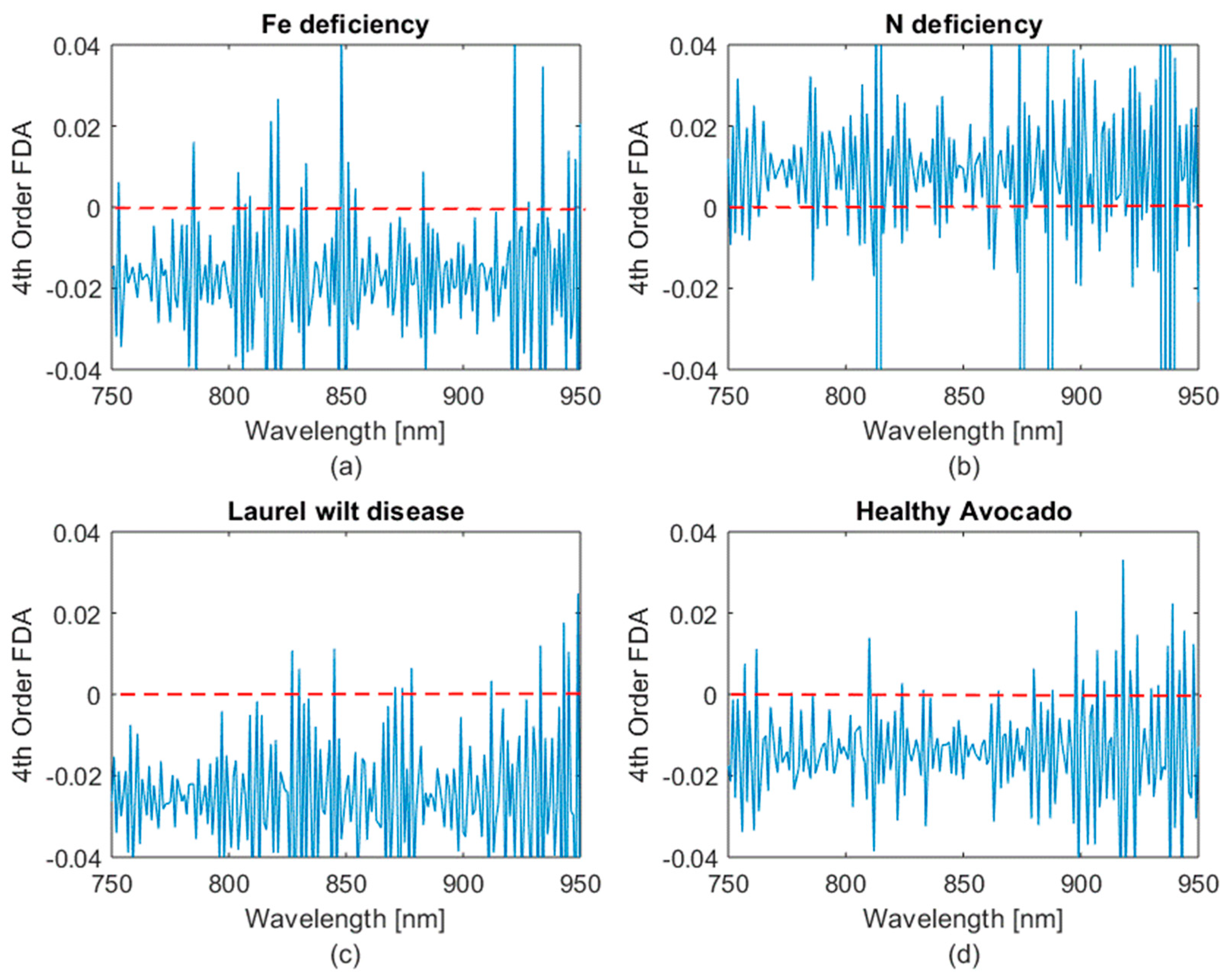
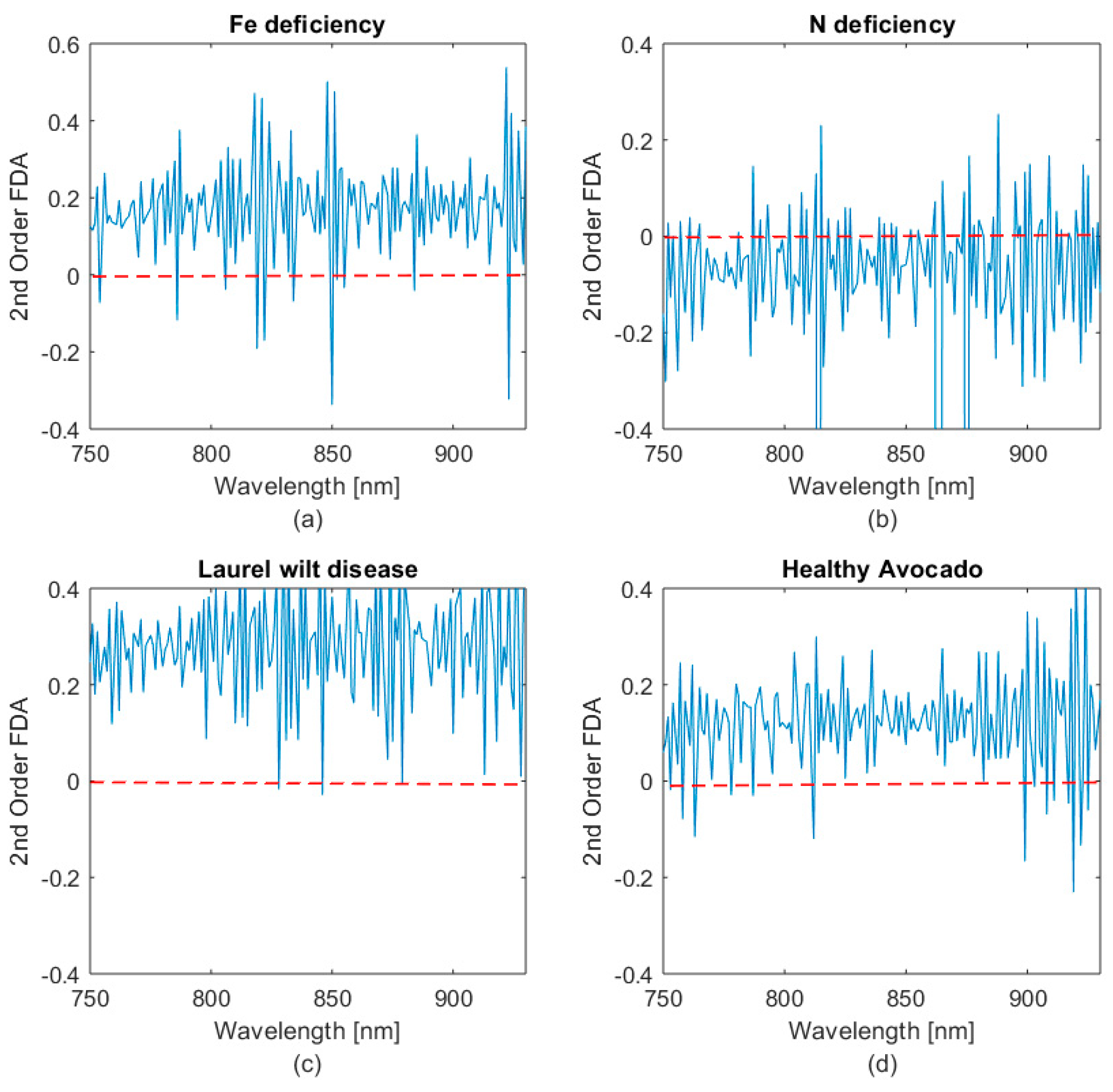
- Obtaining the second and fourth order finite difference approximation (FDA).
- (v)
- Establishing regions of interest (ROI) through FDA inflection point analysis of the spectra.
- (vi)
- Detecting and correctly categorizing the leaf sample into one of the healthy or diseased/deficient categories based on the correlation coefficient result.
- (vii)
- For correlations greater than 0.4, the leaf specimen was classified with the specimen group correlated against.
Confusion Matrix as Figure of Merit
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Evans, E.A.; Crane, J.; Hodges, A.; Osborne, J.L. Potential economic impact of laurel wilt disease on the Florida avocado industry. Horttechnology 2010, 20, 234–238. [Google Scholar] [CrossRef]
- Smith, J.A.; Dreaden, T.J.; Mayfield, A.E., III; Boone, A.; Fraedrich, S.W.; Bates, C. First report of laurel wilt disease caused by Raffaelea lauricola on sassafras in Florida and South Carolina. Plant Dis. 2009, 93, 1079. [Google Scholar] [CrossRef] [PubMed]
- Ploetz, R.C.; Pena, J.E.; Smith, J.A.; Dreaden, T.J.; Crane, J.H.; Schubert, T.; Dixon, W. Laurel wilt, caused by Raffaelea lauricola, is confirmed in Miami-Dade county, center of Florida’s commercial avocado production. Plant Dis. 2011, 95, 1589. [Google Scholar] [CrossRef] [PubMed]
- Carrillo, D.; Duncan, R.E.; Ploetz, J.N.; Campbell, A.F.; Ploetz, R.C.; Pena, J.E. Lateral transfer of a phytopathogenic symbiont among native and exotic ambrosia beetles. Plant Pathol. 2014, 63, 54–62. [Google Scholar] [CrossRef]
- Mayfield, A.E., III; Barnard, E.L.; Smith, J.A.; Bernick, S.C.; Eickwort, J.M.; Dreaden, T.J. Effect of propiconazole on laurel wilt disease development in redbay trees and on the pathogen in vitro. Arboric. Urban For. 2008, 34, 317–324. [Google Scholar]
- Abdulridha, J.; Ehsani, R.; Ampatzidis, Y.; de Castro, A. Evaluating the performance of spectral features and multivariate analysis tools to detect Laurel Wilt Disease and Nutritional Deficiency in Avocado. Comput. Electron. Agric. 2018, 155, 203–211. [Google Scholar] [CrossRef]
- Abdulridha, J.; Ehsani, R.; de Castro, A. Detection and Differentiation between Laurel Wilt Disease, Phytophthora Disease, and Salinity Damage Using a Hyperspectral Sensing Technique. Agriculture 2016, 6, 56. [Google Scholar] [CrossRef]
- Fraedrich, S.W.; Harrington, T.C.; Bates, C.A.; Johnson, J.; Reid, L.S.; Best, G.S.; Leininger, T.D.; Hawkins, T.S. Susceptibility to laurel wilt and disease incidence in two rare plant species, pond berry and pond spice. Plant Dis. 2011, 95, 1056–1062. [Google Scholar] [CrossRef]
- Luvisi, A.; Ampatzidis, Y.G.; De Bellis, L. Plant Pathology and Information Technology: Opportunity for Management of Disease Outbreak and Applications in Regulation Frameworks. Sustainability 2016, 8, 831. [Google Scholar] [CrossRef]
- Ampatzidis, Y.; De Bellis, L.; Luvisi, A. iPathology: Robotic Applications and Management of Plants and Plant Diseases. Sustainability 2017, 9, 1010. [Google Scholar] [CrossRef]
- Cruz, A.C.; Luvisi, A.; De Bellis, L.; Ampatzidis, Y. X-FIDO: An Effective Application for Detecting Olive Quick Decline Syndrome with Deep Learning and Data Fusion. Front. Plant Sci. 2017, 8, 1741. [Google Scholar] [CrossRef] [PubMed]
- Cruz, A.; Ampatzidis, Y.; Pierro, R.; Materazzi, A.; Panattoni, A.; De Bellis, L.; Luvisi, A. Detection of grapevine yellows symptoms in Vitis vinifera L. with artificial intelligence. Comput. Electron. Agric. 2019, 157, 63–76. [Google Scholar] [CrossRef]
- Ampatzidis, Y.; Partel, V. UAV-based High Throughput Phenotyping in Citrus Utilizing Multispectral Imaging and Artificial Intelligence. Remote Sens. 2019, 11, 410. [Google Scholar] [CrossRef]
- Ampatzidis, Y.; Kiner, J.; Abdolee, R.; Ferguson, L. Voice-Controlled and Wireless Solid Set Canopy Delivery (VCW-SSCD) System for Mist-Cooling. Sustainability 2018, 10, 421. [Google Scholar] [CrossRef]
- Partel, V.; Kakarla, C.; Ampatzidis, Y. Development and evaluation of a low-cost and smart technology for precision weed management utilizing artificial intelligence. Comput. Electron. Agric. 2019, 157, 339–350. [Google Scholar] [CrossRef]
- Partel, V.; Nunes, L.; Stansley, P.; Ampatzidis, Y. Automated Vision-based System for Monitoring Asian Citrus Psyllid in Orchards utilizing Artificial Intelligence. Comput. Electron. Agric. 2019, 162, 328–336. [Google Scholar] [CrossRef]
- Ampatzidis, Y.; Tan, L.; Haley, R.; Whiting, M.D. Cloud-based harvest management information system for hand-harvested specialty crops. Comput. Electron. Agric. 2016, 122, 161–167. [Google Scholar] [CrossRef]
- Ampatzidis, Y.G.; Vougioukas, S.G. Field experiments for evaluating the incorporation of RFID and barcode registration and digital weighing technologies in manual fruit harvesting. Comput. Electron. Agric. 2009, 66, 166–172. [Google Scholar] [CrossRef]
- Sankaran, S.; Ehsani, R.; Inch, S.A.; Ploetz, R.C. Evaluation of visible-near infrared reflectance spectra of avocado leaves as a non-destructive sensing tool for detection of laurel wilt. Plant Dis. 2012, 96, 1683–1689. [Google Scholar] [CrossRef]
- Moshou, D.; Bravo, C.; Oberti, R.; West, J.; Bodria, L.; McCartney, A.; Ramon, H. Plant disease detection based on data fusion of hyper-spectral and multi-spectral fluorescence imaging using Kohonen maps. Real Time Imaging 2005, 11, 75–83. [Google Scholar] [CrossRef]
- Varpe, A.B.; Surase, R.R.; Vibhute, A.D.; Gaikwad, S.V.; Rajendra, Y.D.; Kale, K.V.; Mehrotra, S.C. Synygium cumini Plant Photosynthetic Pigment Detection from Hyperspectral data sets using Spectral Indices. In Proceedings of the 2nd Intenational Conference on Man and Machine Interfacing (MAMI), Bhubaneswar, India, 21–23 December 2017. [Google Scholar]
- Ahmadi, P.; Muharam, F.M.; Ahmad, K.; Mansor, S.; Abu Seman, I. Early Detection of Ganoderma Basal Stem Rot of Oil Palms Using Artificial Neural Network Spectral Analysis. Plant Dis. 2017, 101, 1009–1016. [Google Scholar] [CrossRef] [PubMed]
- Corti, M.; Masseroni, D.; Marino Gallina, L. Use of Spectral and Thermal Imaging Sensors to Monitor Crop Water and Nitrogen Status. In First Conference on Proximal Sensing Supporting Precision Agriculture; EAGE: Houten, The Netherlands, 2015; Volume 3997/2214. [Google Scholar]
- Bravo, C.; Moshou, D.; West, J.; McCartney, A.; Ramon, H. Early disease detection in wheat fields using spectral reflectance. Biosyst. Eng. 2003, 84, 137–145. [Google Scholar] [CrossRef]
- Franke, J.; Menz, G. Multi-temporal wheat disease detection by multi-spectral remote sensing. Precis. Agric. 2007, 8, 161–172. [Google Scholar] [CrossRef]
- Perez-Bueno, M.L.; Pineda, M.; Vida, C.; Fernandez-Ortuno, D.; Tores, J.A.; de Vicente, A.; Cazorla, F.M.; Baron, M. Detection of White Root Rot in Avocado Trees by Remote Sensing. Plant Dis. 2019, 103, 1119–1125. [Google Scholar] [CrossRef] [PubMed]
- Calderon, R.; Navas-Cortes, J.A.; Lucena, C.; Zarco-Tejada, P.J. High-resolution airborne hyperspectral and thermal imagery for early, detection of Verticillium wilt of olive using fluorescence, temperature and narrow-band spectral indices. Remote Sens. Environ. 2013, 139, 231–245. [Google Scholar] [CrossRef]
- Ploetz, R.C.; Perez-Martinez, J.M.; Smith, J.A.; Hughes, M.; Dreaden, T.J.; Inch, S.A.; Fu, Y. Responses of avocado to laurel wilt, caused by Raffaelea lauricola. Plant Pathol. 2012, 61, 801–808. [Google Scholar] [CrossRef]
- Ye, D.D.; Sun, L.J.; Tan, W.Y.; Che, W.K.; Yang, M.C. Detecting and classifying minor bruised potato based on hyperspectral imaging. Chemom. Intell. Lab. Syst. 2018, 177, 129–139. [Google Scholar] [CrossRef]
- Susic, N.; Zibrat, U.; Sirca, S.; Strajnar, P.; Razinger, J.; Knapic, M.; Voncina, A.; Urek, G.; Stare, B.G. Discrimination between abiotic and biotic drought stress in tomatoes using hyperspectral imaging. Sens. Actuators B Chem. 2018, 273, 842–852. [Google Scholar] [CrossRef]

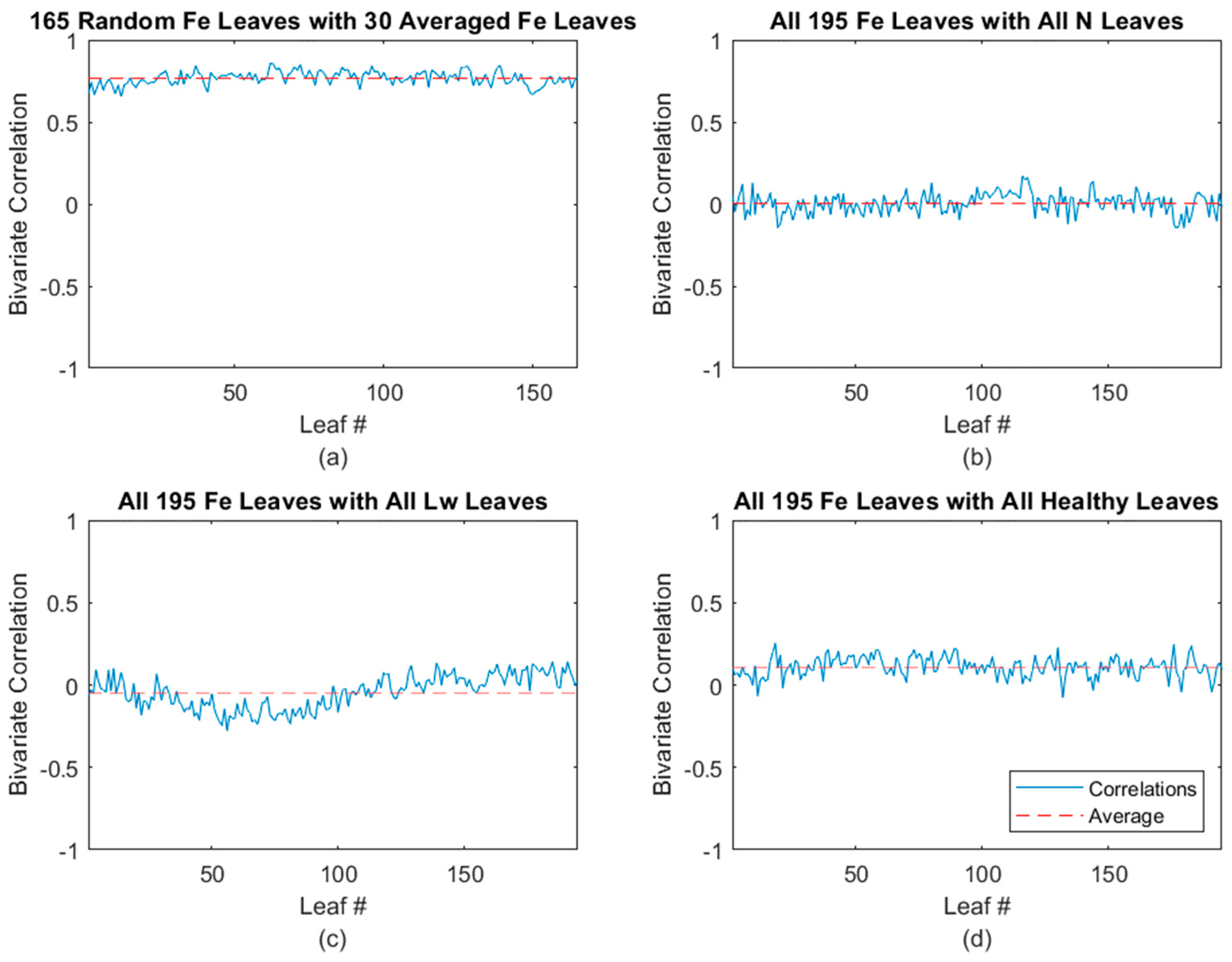
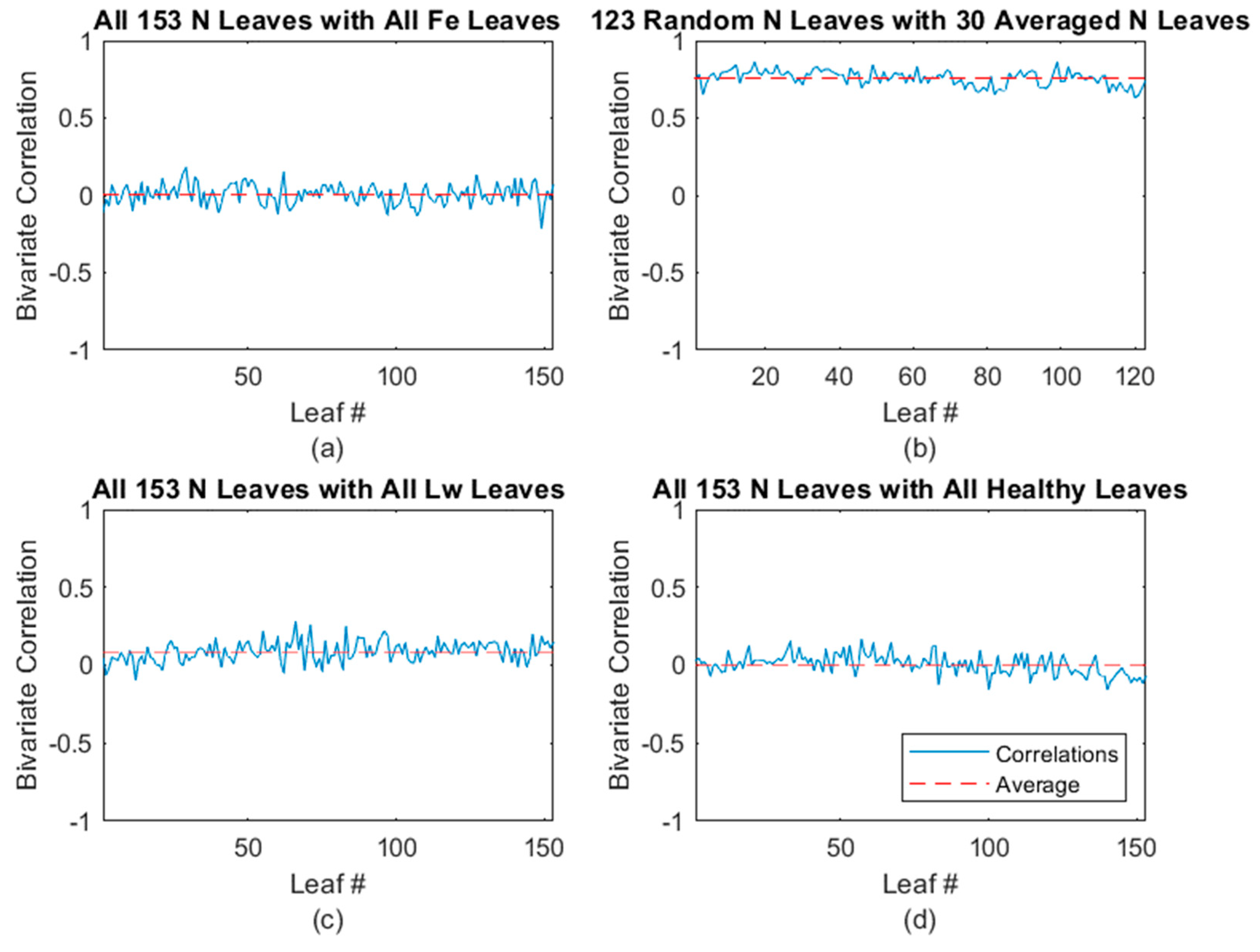
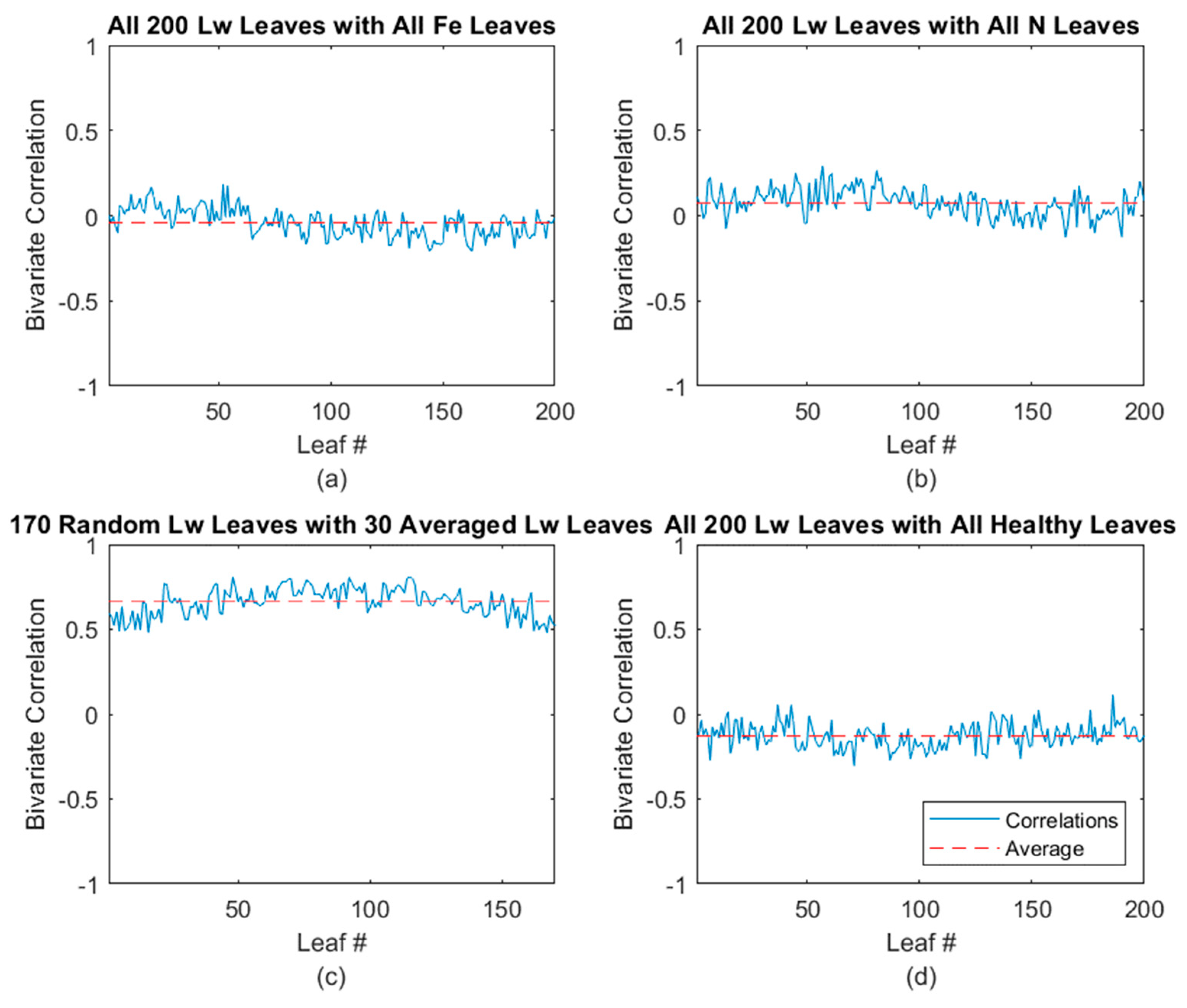
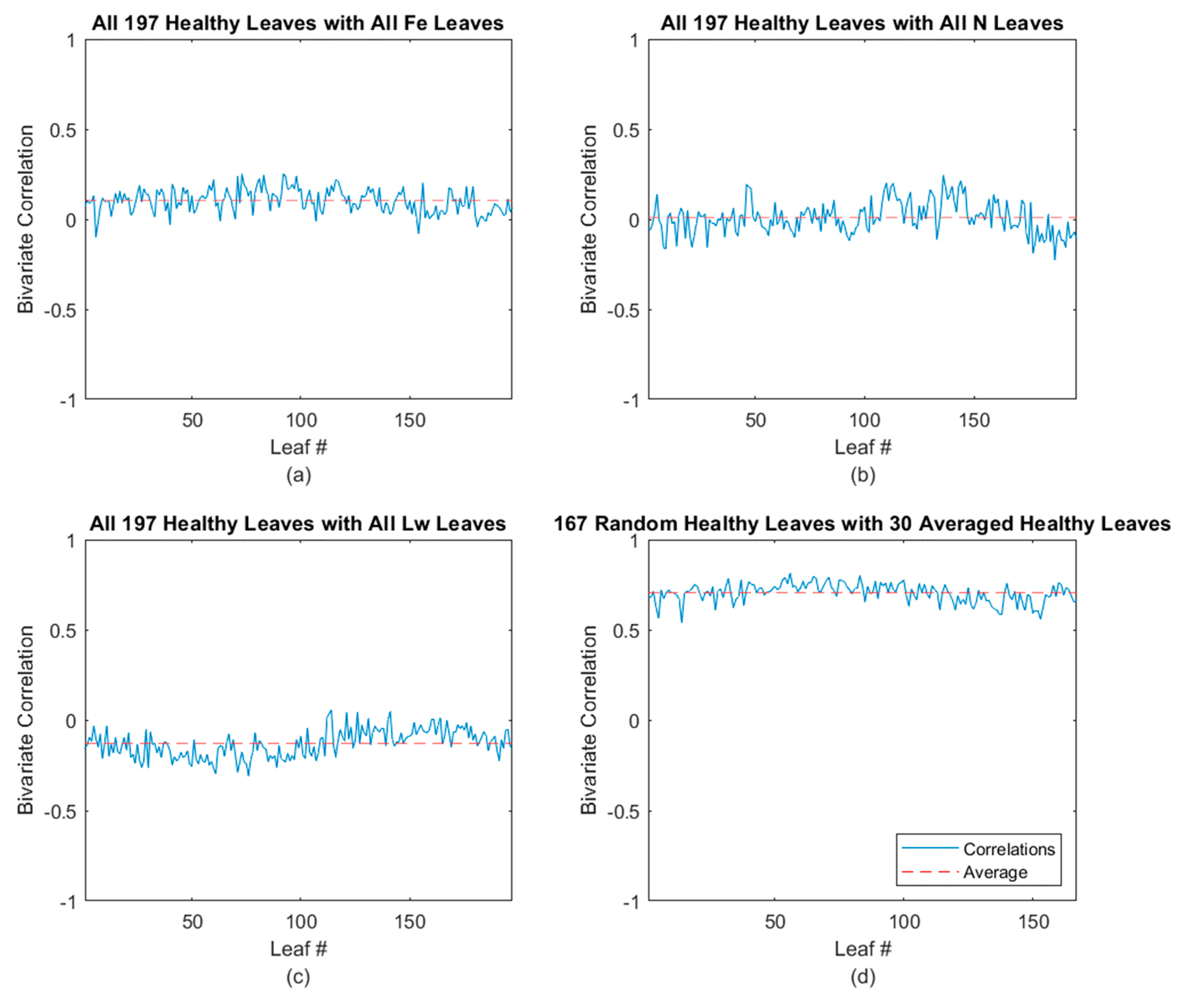
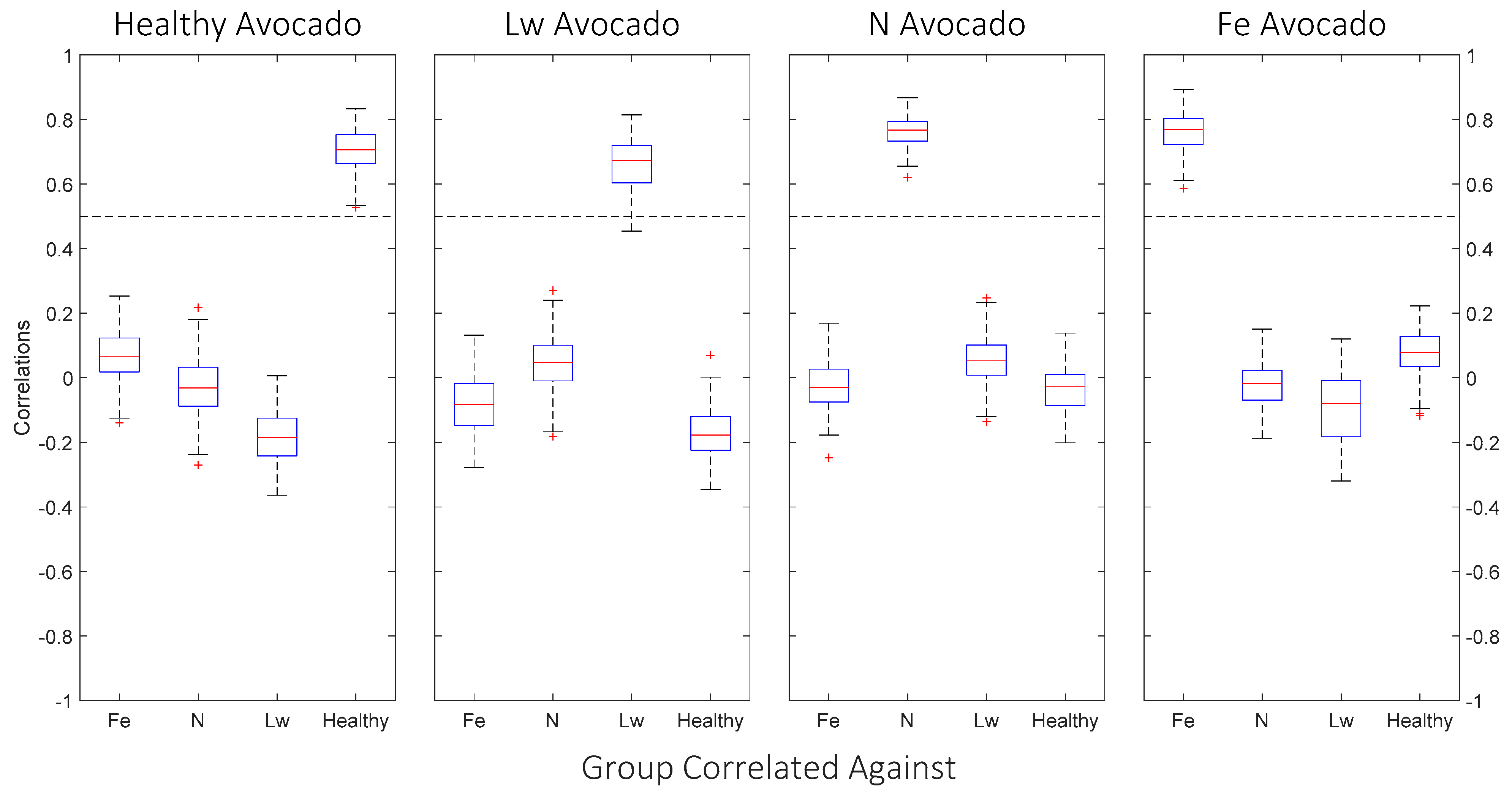
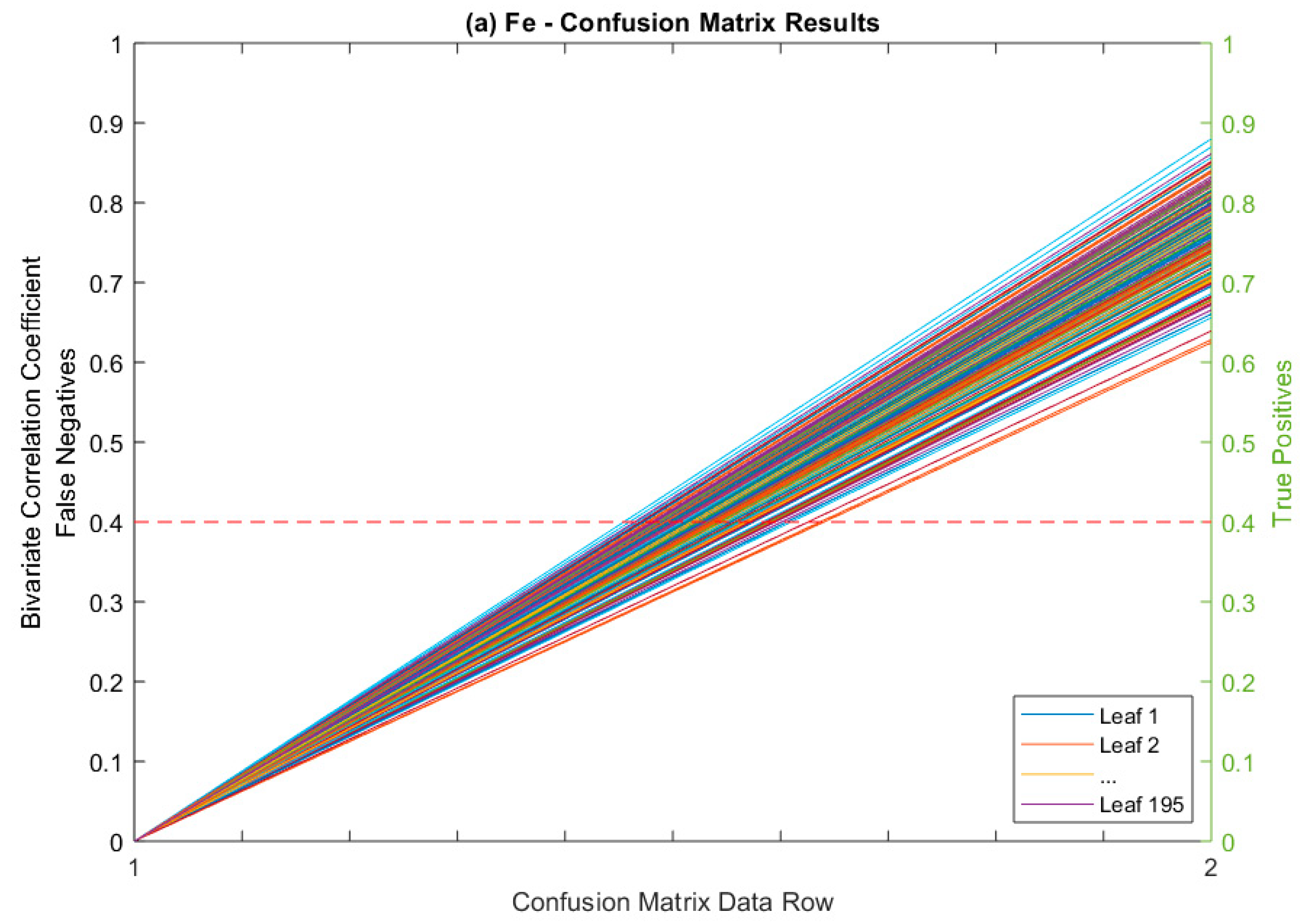
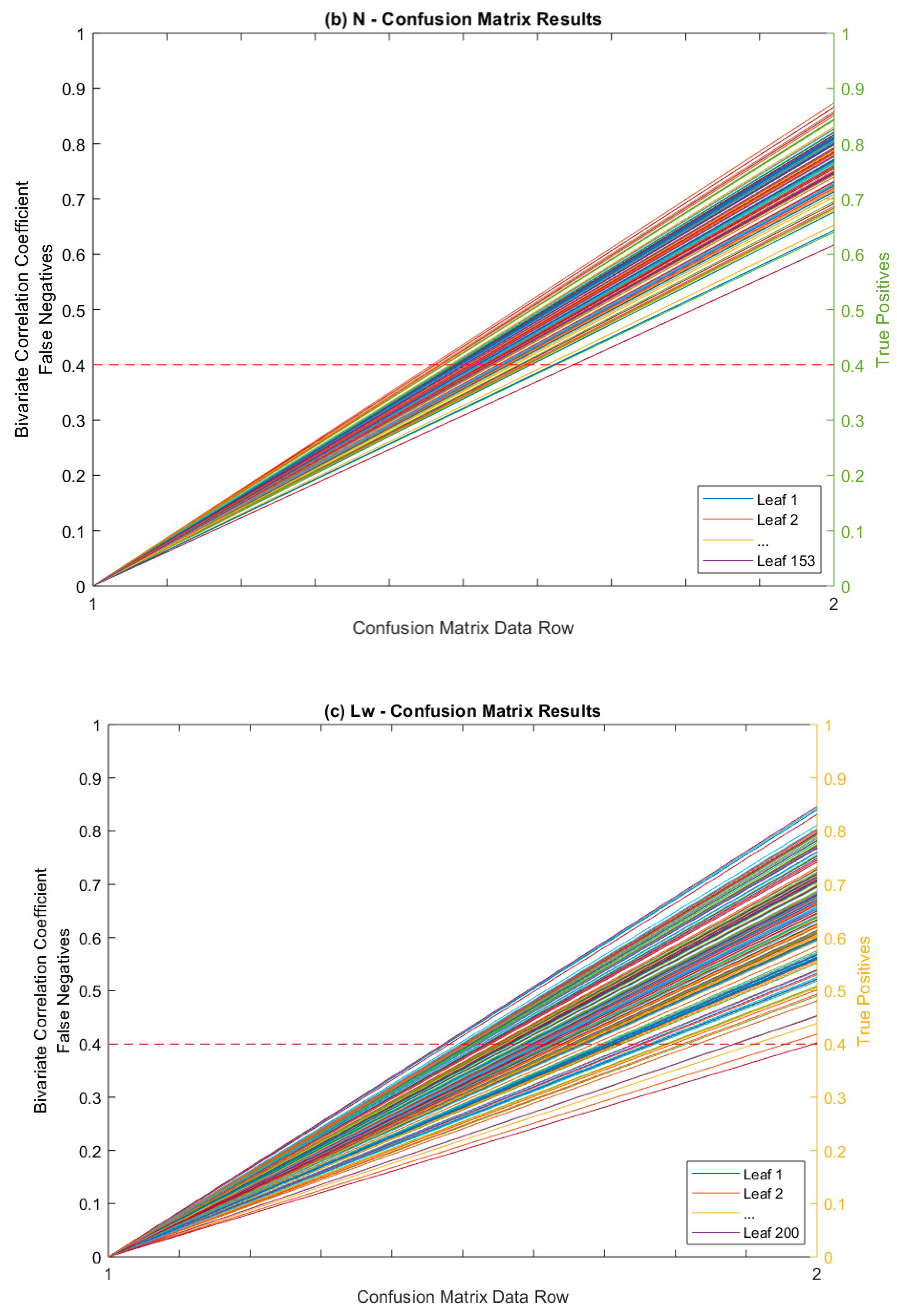
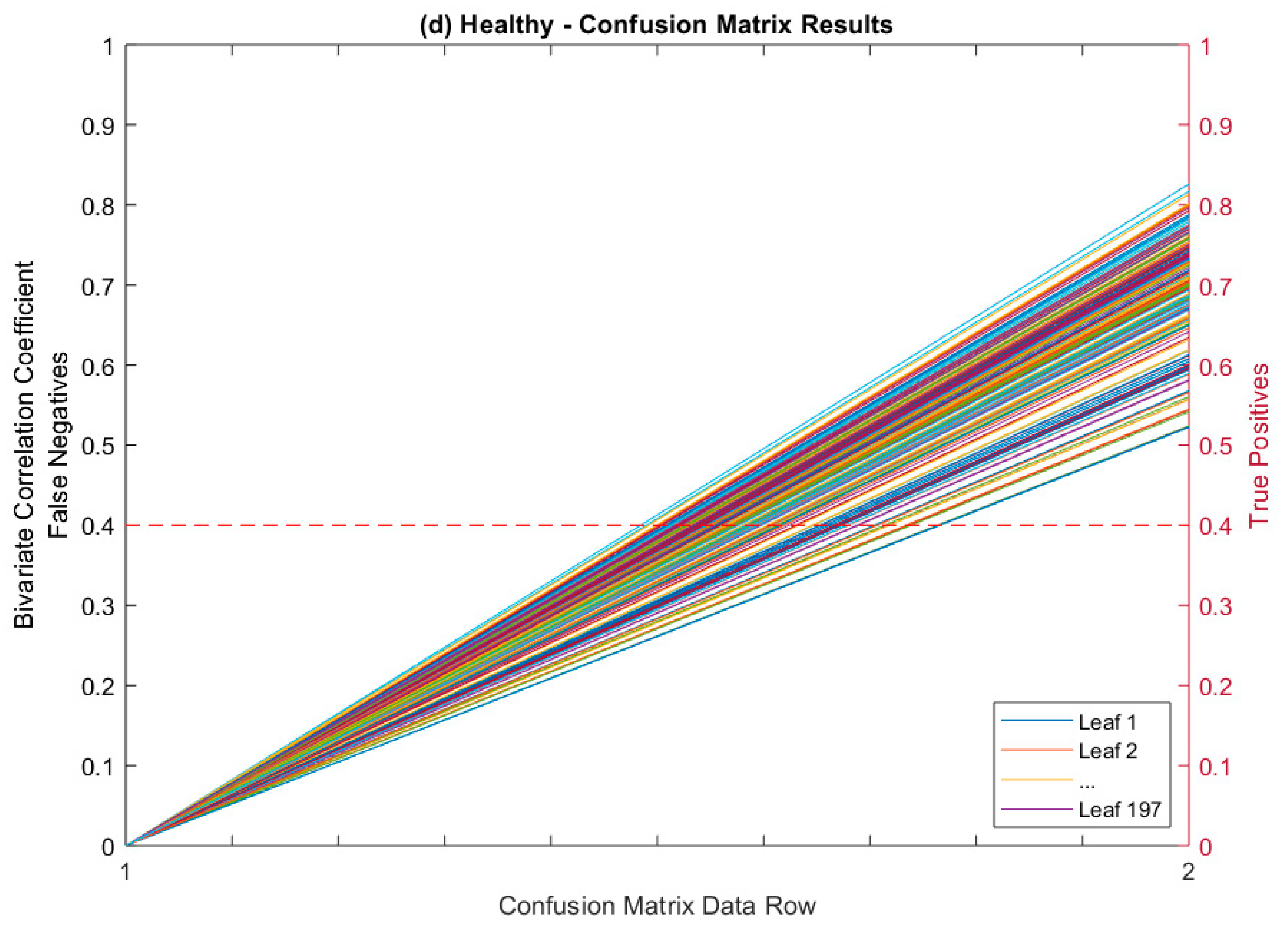
| Leaves Tested | Average Correlated With | Average ρ * | % (ρ) > 0.5 | % (ρ) > 0.4 |
|---|---|---|---|---|
| Healthy | Healthy | 0.698 | 100.00 | 100.00 |
| Fe-deficient | 0.071 | 0.00 | 0.00 | |
| N-deficient | −0.027 | 0.00 | 0.00 | |
| Laurel wilt | −0.181 | 0.00 | 0.00 | |
| Fe-deficient | Healthy | 0.076 | 0.00 | 0.00 |
| Fe-deficient | 0.778 | 100.00 | 100.00 | |
| N-deficient | −0.021 | 0.00 | 0.00 | |
| Laurel wilt | −0.090 | 0.00 | 0.00 | |
| N-deficient | Healthy | −0.032 | 0.00 | 0.00 |
| Fe-deficient | −0.021 | 0.00 | 0.00 | |
| N-deficient | 0.762 | 100.00 | 100.00 | |
| Laurel wilt | 0.027 | 0.00 | 0.00 | |
| Laurel wilt | Healthy | −0.186 | 0.00 | 0.00 |
| Fe-deficient | −0.078 | 0.00 | 0.00 | |
| N-deficient | 0.038 | 0.00 | 0.00 | |
| Laurel wilt | 0.645 | 100.00 | 100.00 |
| Confusion Matrix Results | ||||||
| Actual Class | ||||||
| Healthy | Fe-Deficient | N-deficient | Lw-Diseased | |||
| Predicted Class | Healthy | 197 | 0 | 0 | 0 | |
| Fe-Deficient | 0 | 195 | 0 | 0 | ||
| N-Deficient | 0 | 153 | 0 | |||
| Lw-Diseased | 0 | 0 | 200 | |||
| Totals | ||||||
| True Positives (TP) | 197 | 195 | 153 | 200 | 745 | |
| False Negatives (FN) | 0 | 0 | 0 | 0 | 0 | |
| True Negatives (TN) | 548 | 550 | 592 | 545 | 2235 | |
| False Positives (FP) | 0 | 0 | 0 | 0 | 0 | |
| ACC = (TP+TN)/(P+N) = 1.00 | ||||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hariharan, J.; Fuller, J.; Ampatzidis, Y.; Abdulridha, J.; Lerwill, A. Finite Difference Analysis and Bivariate Correlation of Hyperspectral Data for Detecting Laurel Wilt Disease and Nutritional Deficiency in Avocado. Remote Sens. 2019, 11, 1748. https://doi.org/10.3390/rs11151748
Hariharan J, Fuller J, Ampatzidis Y, Abdulridha J, Lerwill A. Finite Difference Analysis and Bivariate Correlation of Hyperspectral Data for Detecting Laurel Wilt Disease and Nutritional Deficiency in Avocado. Remote Sensing. 2019; 11(15):1748. https://doi.org/10.3390/rs11151748
Chicago/Turabian StyleHariharan, Jeanette, John Fuller, Yiannis Ampatzidis, Jaafar Abdulridha, and Andrew Lerwill. 2019. "Finite Difference Analysis and Bivariate Correlation of Hyperspectral Data for Detecting Laurel Wilt Disease and Nutritional Deficiency in Avocado" Remote Sensing 11, no. 15: 1748. https://doi.org/10.3390/rs11151748
APA StyleHariharan, J., Fuller, J., Ampatzidis, Y., Abdulridha, J., & Lerwill, A. (2019). Finite Difference Analysis and Bivariate Correlation of Hyperspectral Data for Detecting Laurel Wilt Disease and Nutritional Deficiency in Avocado. Remote Sensing, 11(15), 1748. https://doi.org/10.3390/rs11151748







