Constrained Spectral Clustering of Individual Trees in Dense Forest Using Terrestrial Laser Scanning Data
Abstract
1. Introduction
2. Study Area and Data
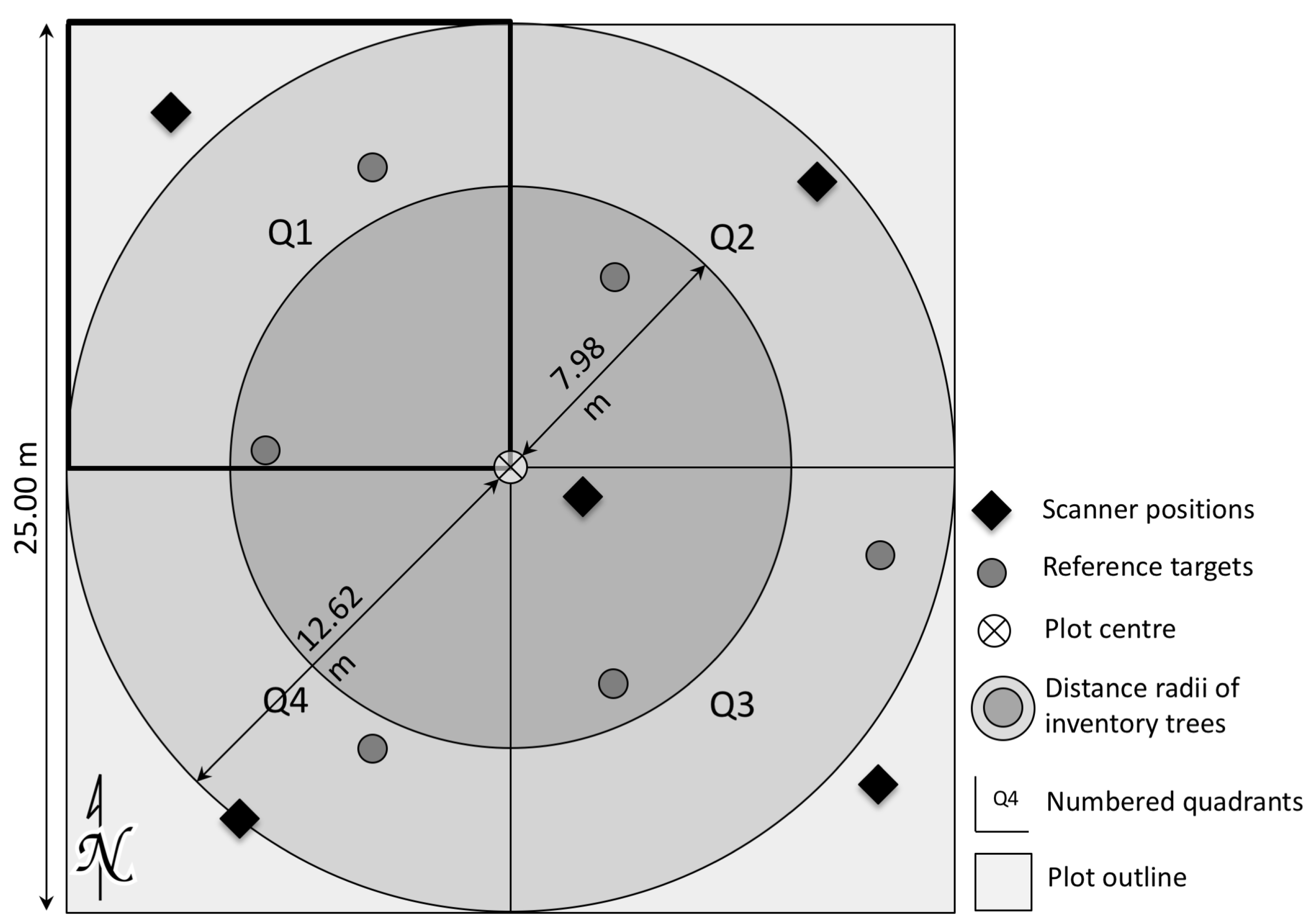
2.1. Study Area
2.2. LiDAR Data
2.3. Reference Data
3. Method
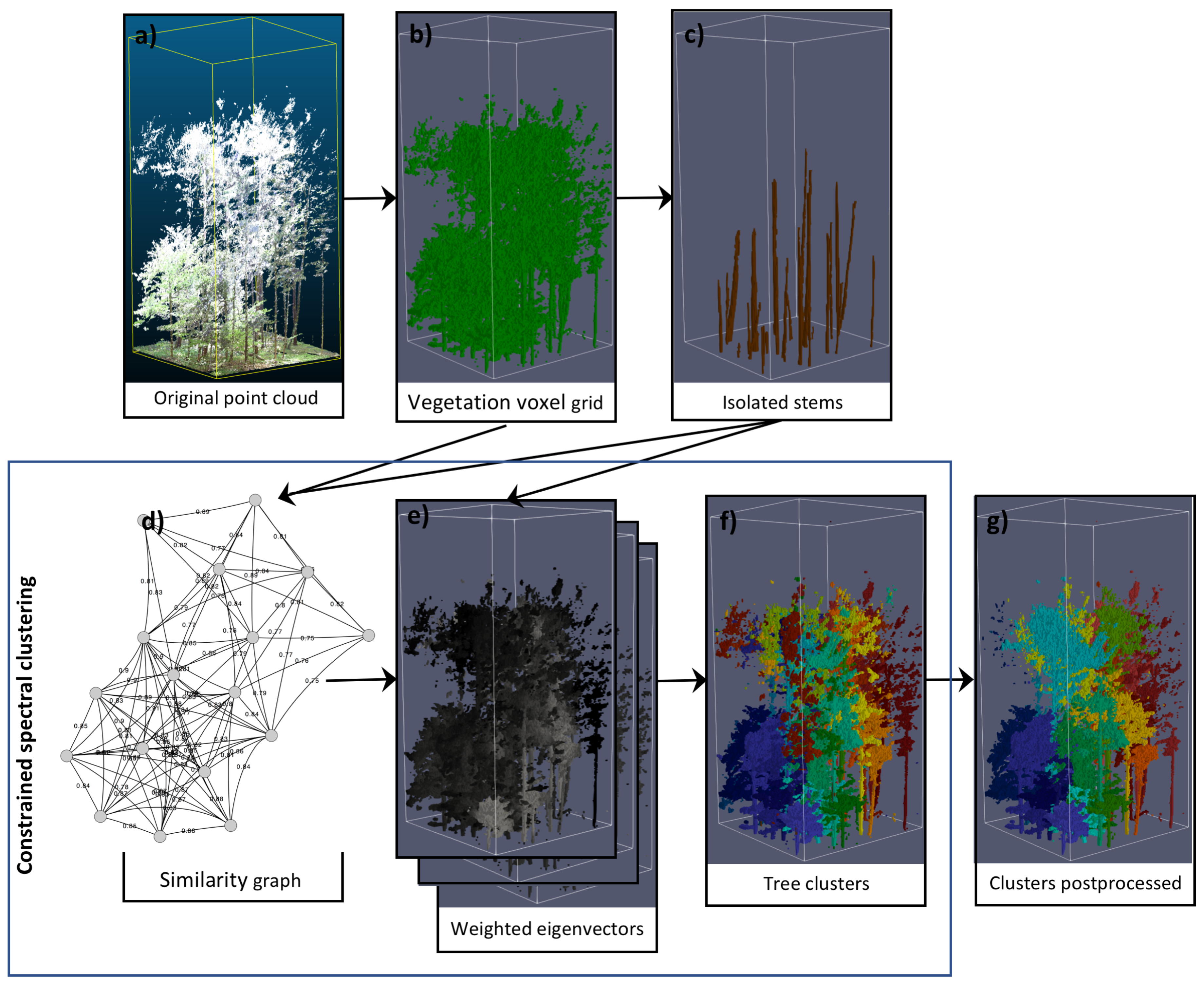
3.1. Outline
3.2. Tree Stem Detection
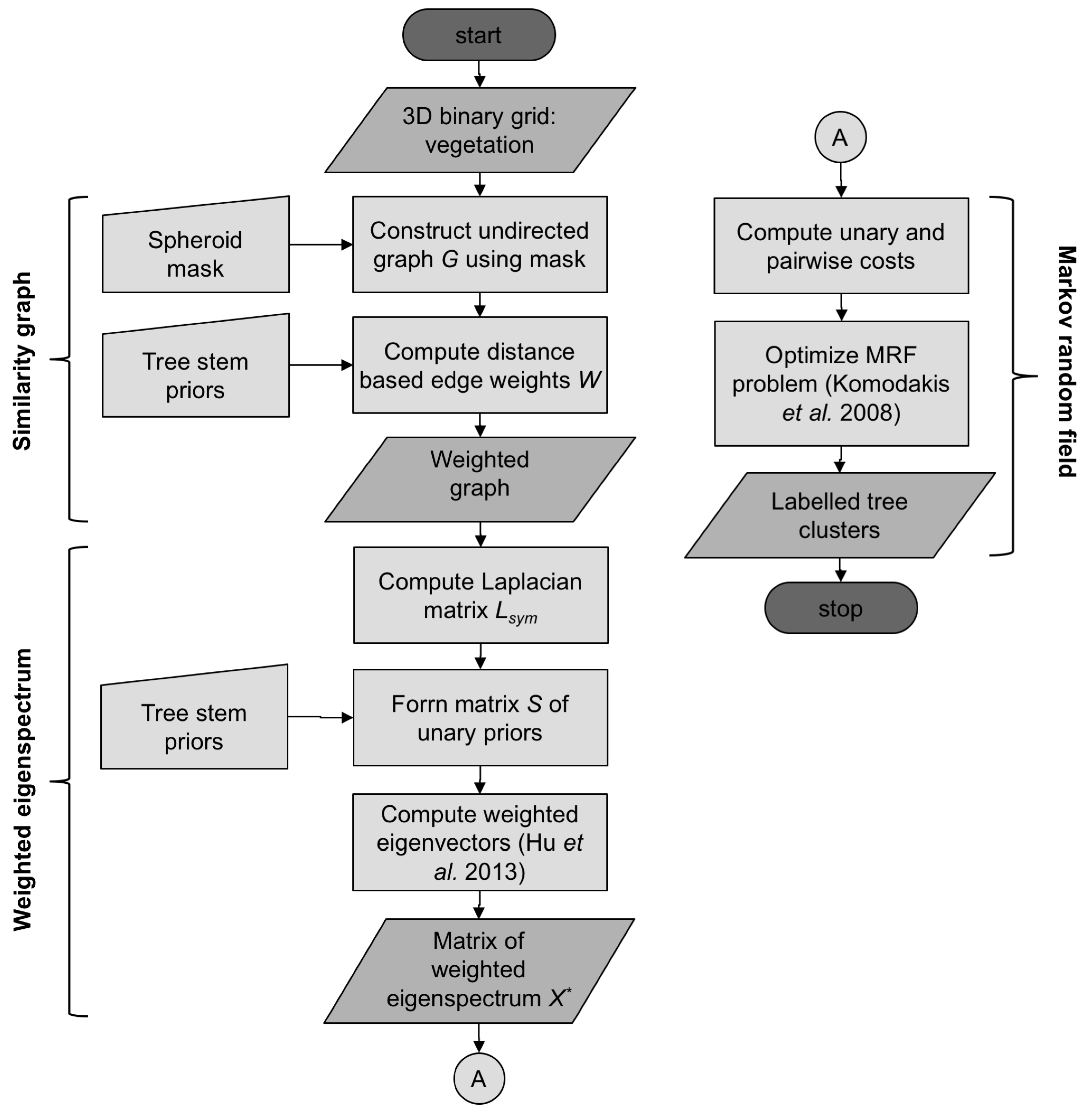
3.3. Constrained Spectral Clustering
3.3.1. Similarity Graph
3.3.2. Weighted Eigenspectrum
3.3.3. Markov Random Fields
3.4. Postprocessing
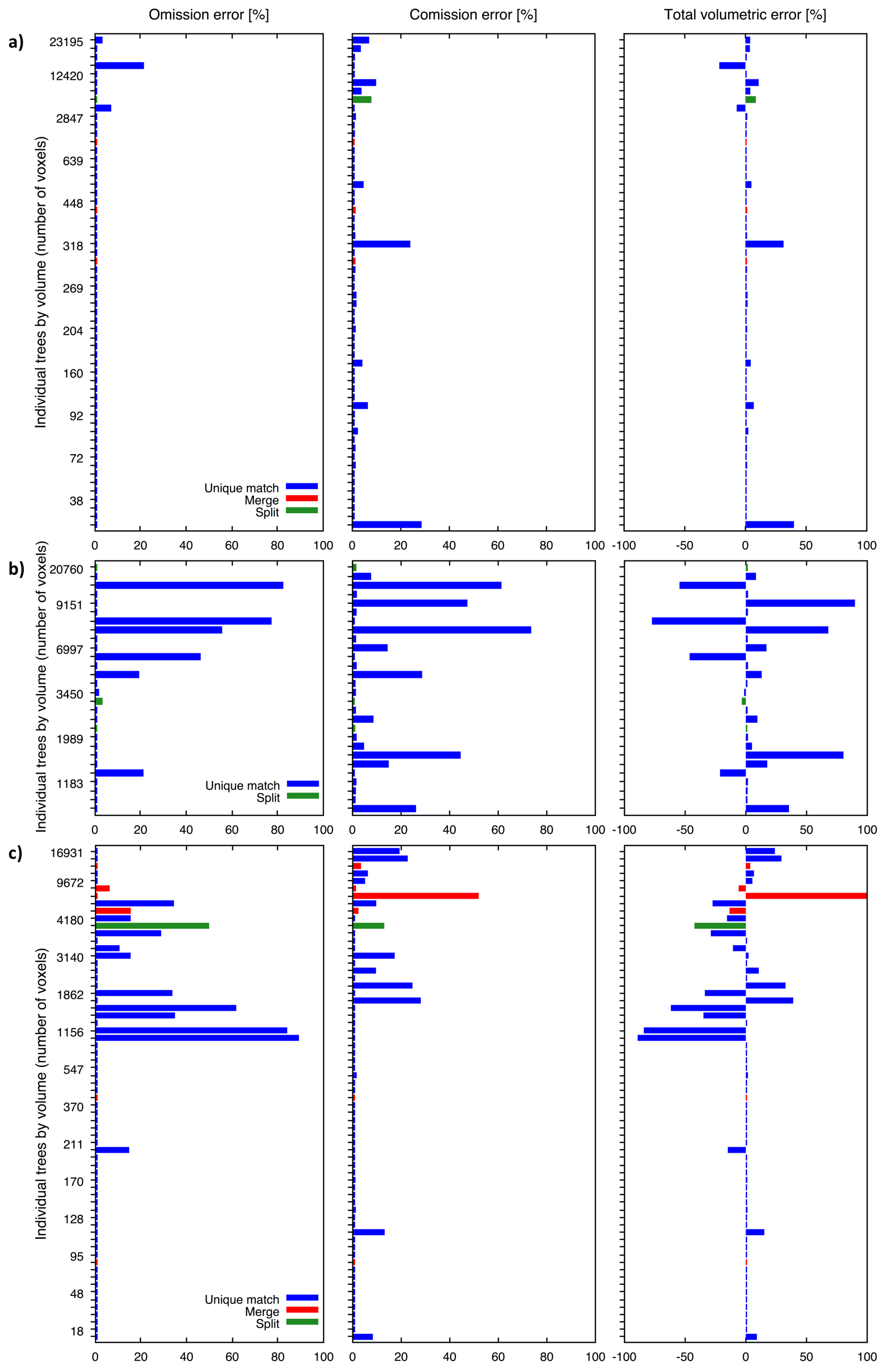
3.5. Accuracy Assessment
4. Results
5. Discussion
5.1. Methodical Approach
5.2. Interpretation of the Results
5.3. Comparison to Other Studies
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pinz, A.J. Final results of the vision expert system VES: Finding trees in aerial photographs. In Wissensbasierte Mustererkennung; OCG Schriftenreihe; Institut für Elektrische Messtechnik Und Messsignalverarbeitung: Graz, Austria, 1989; Volume 49, pp. 90–111. [Google Scholar]
- Reitberger, J.; Schnörr, C.; Krzystek, P.; Stilla, U. 3D segmentation of full waveform LIDAR data for single tree detection using normalized cut. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2008, 37, 77–83. [Google Scholar]
- Strîmbu, V.F.; Strîmbu, B.M. A graph-based segmentation algorithm for tree crown extraction using airborne LiDAR data. ISPRS J. Photogramm. Remote Sens. 2015, 104, 30–43. [Google Scholar] [CrossRef]
- Mongus, D.; Žalik, B. An efficient approach to 3D single tree-crown delineation in LiDAR data. ISPRS J. Photogramm. Remote Sens. 2015, 108, 219–233. [Google Scholar] [CrossRef]
- Heinzel, J.N.; Weinacker, H.; Koch, B. Prior-knowledge-based single-tree extraction. Int. J. Remote Sens. 2011, 32, 4999–5020. [Google Scholar] [CrossRef]
- Leckie, D.; Gougeon, F.; Hill, D.; Quinn, R.; Armstrong, L.; Shreenan, R. Combined high-density lidar and multispectral imagery for individual tree crown analysis. Can. J. Remote Sens. 2003, 29, 633–649. [Google Scholar] [CrossRef]
- Hackenberg, J.; Morhart, C.; Sheppard, J.; Spiecker, H.; Disney, M. Highly Accurate Tree Models Derived from Terrestrial Laser Scan Data: A Method Description. Forests 2014, 5, 1069–1105. [Google Scholar] [CrossRef]
- Raumonen, P.; Kaasalainen, M.; Åkerblom, M.; Kaasalainen, S.; Kaartinen, H.; Vastaranta, M.; Holopainen, M.; Disney, M.; Lewis, P. Fast Automatic Precision Tree Models from Terrestrial Laser Scanner Data. Remote Sens. 2013, 5, 491–520. [Google Scholar] [CrossRef]
- Hosoi, F.; Nakai, Y.; Omasa, K. 3-D voxel-based solid modeling of a broad-leaved tree for accurate volume estimation using portable scanning lidar. ISPRS J. Photogramm. Remote Sens. 2013, 82, 41–48. [Google Scholar] [CrossRef]
- Raumonen, P.; Casella, E.; Calders, K.; Murphy, S.; Åkerbloma, M.; Kaasalainen, M. Massive-scale tree modelling from TLS data. ISPRS Ann. Photogramm. Remote Sens. Spat. Inf. Sci. 2015, II-3/W4, 189–196. [Google Scholar] [CrossRef]
- Hu, S.; Li, Z.; Zhang, Z.; He, D.; Wimmer, M. Efficient tree modeling from airborne LiDAR point clouds. Comput. Graph. 2017, 67, 1–13. [Google Scholar] [CrossRef]
- Calders, K.; Newnham, G.; Burt, A.; Murphy, S.; Raumonen, P.; Herold, M.; Culvenor, D.; Avitabile, V.; Disney, M.; Armston, J.; et al. Nondestructive estimates of above-ground biomass using terrestrial laser scanning. Methods Ecol. Evol. 2014, 6, 198–208. [Google Scholar] [CrossRef]
- Kaartinen, H.; Hyyppä, J.; Yu, X.; Vastaranta, M.; Hyyppä, H.; Kukko, A.; Holopainen, M.; Heipke, C.; Hirschmugl, M.; Morsdorf, F.; et al. An International Comparison of Individual Tree Detection and Extraction Using Airborne Laser Scanning. Remote Sens. 2012, 4, 950–974. [Google Scholar] [CrossRef]
- Menk, J.; Dorren, L.; Heinzel, J.; Marty, M.; Huber, M. Evaluation automatischer Einzelbaumerkennung aus luftgestützten Laserscanning-Daten. Schweizerische Zeitschrift fur Forstwesen 2017, 168, 151–159. [Google Scholar] [CrossRef]
- Kwak, D.A.; Lee, W.K.; Lee, J.H.; Biging, G.S.; Gong, P. Detection of individual trees and estimation of tree height using LiDAR data. J. For. Res. 2007, 12, 425–434. [Google Scholar] [CrossRef]
- Baumann, A.B.G.; Baltsavias, E. Einzelbaumdetektion in bewaldetem Gebiet auf Basis von luftgestützten LiDAR-Daten. In Proceedings of the Dreiländertagung der DGPF, der OVG und der SGPF, Bern, Schweiz, 6–9 June 2016; Kersten, T.P., Ed.; Volume 25, pp. 506–515. [Google Scholar]
- Chmielewski, L.J.; Bator, M.; Zasada, M.; Stereńczak, K.; Strzeliński, P. Fuzzy Hough Transform-Based Methods for Extraction and Measurements of Single Trees in Large-Volume 3D Terrestrial LIDAR Data. In Computer Vision and Graphics; Springer: Berlin/Heidelberg, Germany, 2010; pp. 265–274. [Google Scholar]
- Bienert, A.; Scheller, S.; Keane, E.; Mohan, F.; Nugent, C. Tree detection and diameter estimations by analysis of forest terrestrial laserscanner point clouds. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2007, XXXVI-3/W52, 50–55. [Google Scholar]
- Lee, H.; Slatton, K.C.; Roth, B.E.; Cropper, W.P. Adaptive clustering of airborne LiDAR data to segment individual tree crowns in managed pine forests. Int. J. Remote Sens. 2010, 31, 117–139. [Google Scholar] [CrossRef]
- Bienert, A.; Queck, R.; Schmidt, A.; Bernhofer, C.; Maas, H. Voxel space analysis of terrestrial laser scans in forests for wind field modelling. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2010, 38, 92–97. [Google Scholar]
- Reitberger, J.; Schnörr, C.; Krzystek, P.; Stilla, U. 3D segmentation of single trees exploiting full waveform LIDAR data. ISPRS J. Photogramm. Remote Sens. 2009, 64, 561–574. [Google Scholar] [CrossRef]
- Duncanson, L.; Cook, B.; Hurtt, G.; Dubayah, R. An efficient, multi-layered crown delineation algorithm for mapping individual tree structure across multiple ecosystems. Remote Sens. Environ. 2014, 154, 378–386. [Google Scholar] [CrossRef]
- Wang, Y.; Hyyppa, J.; Liang, X.; Kaartinen, H.; Yu, X.; Lindberg, E.; Holmgren, J.; Qin, Y.; Mallet, C.; Ferraz, A.; et al. International Benchmarking of the Individual Tree Detection Methods for Modeling 3-D Canopy Structure for Silviculture and Forest Ecology Using Airborne Laser Scanning. IEEE Trans. Geosci. Remote Sens. 2016, 54, 5011–5027. [Google Scholar] [CrossRef]
- Li, W.; Guo, Q.; Jakubowski, M.K.; Kelly, M. A new method for segmenting individual trees from the lidar point cloud. Photogramm. Eng. Remote Sens. 2012, 78, 75–84. [Google Scholar] [CrossRef]
- Lu, X.; Guo, Q.; Li, W.; Flanagan, J. A bottom-up approach to segment individual deciduous trees using leaf-off lidar point cloud data. ISPRS J. Photogramm. Remote Sens. 2014, 94, 1–12. [Google Scholar] [CrossRef]
- Heurich, M.; Persson, Å.; Holmgren, J.; Kennel, E. Detecting and measuring individual trees with laser scanning in mixed mountain forest of central Europe using an algorithm developed for Swedish boreal forest conditions. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2004, 36, 307–312. [Google Scholar]
- Weinacker, H.; Koch, B.; Heyder, U.; Weinacker, R. Development of filtering, segmentation and modelling modules for lidar and multispectral data as a fundament of an automatic forest inventory system. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2004, 36, W2. [Google Scholar]
- Xia, S.; Wang, C.; Pan, F.; Xi, X.; Zeng, H.; Liu, H. Detecting stems in dense and homogeneous forest using single-scan TLS. Forests 2015, 6, 3923–3945. [Google Scholar] [CrossRef]
- Bienert, A.; Schneider, D. Range image segmentation for tree detection in forest scans. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2013, II-5/W2, 49–54. [Google Scholar] [CrossRef]
- Zhong, L.; Cheng, L.; Xu, H.; Wu, Y.; Chen, Y.; Li, M. Segmentation of Individual Trees From TLS and MLS Data. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2017, 10, 774–787. [Google Scholar] [CrossRef]
- Zou, X.; Cheng, M.; Wang, C.; Xia, Y.; Li, J. Tree Classification in Complex Forest Point Clouds Based on Deep Learning. IEEE Geosci. Remote Sens. Lett. 2017, 14, 2360–2364. [Google Scholar] [CrossRef]
- Rutzinger, M.; Pratihast, A.; Oude Elberink, S.; Vosselman, G. Detection and modelling of 3D trees from mobile laser scanning data. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2010, 38, 520–525. [Google Scholar]
- Lin, Y.; Hyyppä, J.; Jaakkola, A.; Yu, X. Three-level frame and RD-schematic algorithm for automatic detection of individual trees from MLS point clouds. Int. J. Remote Sens. 2012, 33, 1701–1716. [Google Scholar] [CrossRef]
- Wu, B.; Yu, B.; Yue, W.; Shu, S.; Tan, W.; Hu, C.; Huang, Y.; Wu, J.; Liu, H. A Voxel-Based Method for Automated Identification and Morphological Parameters Estimation of Individual Street Trees from Mobile Laser Scanning Data. Remote Sens. 2013, 5, 584–611. [Google Scholar] [CrossRef]
- Von Luxburg, U. A tutorial on spectral clustering. Stat. Comput. 2007, 17, 395–416. [Google Scholar] [CrossRef]
- Shi, J.; Malik, J. Normalized cuts and image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2000, 22, 888–905. [Google Scholar]
- Lee, J.; Cai, X.; Lellmann, J.; Dalponte, M.; Malhi, Y.; Butt, N.; Morecroft, M.; Schonlieb, C.B.; Coomes, D.A. Individual Tree Species Classification From Airborne Multisensor Imagery Using Robust PCA. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2016, 9, 2554–2567. [Google Scholar] [CrossRef]
- Hu, H.; Feng, J.; Yu, C.; Zhou, J. Multi-class constrained normalized cut with hard, soft, unary and pairwise priors and its applications to object segmentation. IEEE Trans. Image Process. 2013, 22, 4328–4340. [Google Scholar] [CrossRef] [PubMed]
- Klein, D.; Kamvar, S.D.; Manning, C.D. From instance-level constraints to space-level constraints: Making the most of prior knowledge in data clustering. In Proceedings of the Nineteenth International Conference on Machine Learning, Sydney, Australia, 8–12 July 2002; pp. 307–314. [Google Scholar]
- Peluffo-Ordóñez, D.H.; Castro-Ospina, A.E.; Chavez-Chamorro, D.; Acosta-Medina, C.D.; Castellanos-Domínguez, G. Normalized cuts clustering with prior knowledge and a pre-clustering stage. In Proceedings of the European Symposium on Artificial Neural Networks, Bruges, Belgium, 24–26 April 2013; pp. 309–314. [Google Scholar]
- Yao, T.; Yang, X.; Zhao, F.; Wang, Z.; Zhang, Q.; Jupp, D.; Lovell, J.; Culvenor, D.; Newnham, G.; Ni-Meister, W.; et al. Measuring forest structure and biomass in New England forest stands using Echidna ground-based lidar. Remote Sens. Environ. 2011, 115, 2965–2974. [Google Scholar] [CrossRef]
- Zande, D.V.d.; Jonckheere, I.; Stuckens, J.; Verstraeten, W.W.; Coppin, P. Sampling design of ground-based lidar measurements of forest canopy structure and its effect on shadowing. Can. J. Remote Sens. 2008, 34, 526–538. [Google Scholar] [CrossRef]
- FARO Technologies Inc. FARO SCENE 5.1 Users Manual; FARO: Lake Mary, FL, USA, 2014. [Google Scholar]
- Ayachit, U. The ParaView Guide; Kitware Inc.: New York, NY, USA, 2016; p. 237. [Google Scholar]
- Heinzel, J.; Huber, M. Detecting Tree Stems from Volumetric TLS Data in Forest Environments with Rich Understory. Remote Sens. 2017, 9, 9. [Google Scholar] [CrossRef]
- Soille, P. Morphological Image Analysis: Principles and Applications; Springer: Berlin/Heidelberg, Germany; New York, NY, USA, 2004; p. 302. [Google Scholar]
- Komodakis, N.; Tziritas, G. Approximate Labeling via Graph Cuts Based on Linear Programming. IEEE Trans. Pattern Anal. Mach. Intell. 2007, 29, 1436–1453. [Google Scholar] [CrossRef] [PubMed]
- Kamvar, K.; Sepandar, S.; Klein, K.; Dan, D.; Manning, M.; Christopher, C. Spectral learning. In Proceedings of the International Joint Conference of Artificial Intelligence, Acapulco, Mexico, 9–15 August 2003. [Google Scholar]
- Komodakis, N.; Tziritas, G.; Paragios, N. Performance vs computational efficiency for optimizing single and dynamic MRFs: Setting the state of the art with primal-dual strategies. Comput. Vis. Image Underst. 2008, 112, 14–29. [Google Scholar] [CrossRef]
- Congalton, R.G. A review of assessing the accuracy of classifications of remotely sensed data. Remote Sens. Environ. 1991, 37, 35–46. [Google Scholar] [CrossRef]
- Brolly, G.; Király, G.; Czimber, K. Mapping Forest Regeneration from Terrestrial Laser Scans. Acta Silv. Lign. Hung. 2013, 9, 135–146. [Google Scholar] [CrossRef]
- Liang, X.; Hyyppä, J. Automatic Stem Mapping by Merging Several Terrestrial Laser Scans at the Feature and Decision Levels. Sensors 2013, 13, 1614–1634. [Google Scholar] [CrossRef] [PubMed]
| Plot № | Forest Type and Species | Number of Mature Trees | Regeneration Canopy Coverage | General Description of Layers and Structure | |
|---|---|---|---|---|---|
| Tree Height m | Tree Height m and DBH ≤ 12 cm | ||||
| 1 | Mixed (fir, pine, spruce, beech) | 7 DBH 12–36 cm 4 DBH > 36 cm | Fir 6–15% | Fir 16–25% Spruce 6–15% Beech 6–15% | Lower and upper layer predominant. Lower layer contains small trees in dense and patchy clusters (height < 130 cm). |
| 2 | Mixed (pine, spruce, beech, ash, elm) | 8 DBH 12–36 cm 9 DBH > 36 cm | Beech 16–25% Others 6–15% | Beech 26–50% Elm 6–15% | Lower layer dominated by very small trees (< 130 cm), mature trees of relatively low height (∼17 m). |
| 3 | Mixed (pine, spruce, rowan, sorb, alder) | 9 DBH 12–36 cm 2 DBH > 36 cm | All spec. ≤ 5% | Spruce 6–15% Sorb 6–15% Alder 6–15% | Mid layer interlocking with upper layer trees. Upper layer of relatively low height (∼16 m). |
| 4 | Mixed (fir, pine, spruce, beech) | 10 DBH 12–36 cm 7 DBH > 36 cm | All spec. ≤ 5% | Fir 16–25% Spruce 26–50% Beech 16–25% | Continuous height transition between mid and upper layers. Strongly interlocking crowns but good stem visibility. |
| 5 | Mixed (pine, spruce, larch, beech, ash, maple) | 15 DBH 12–36 cm 3 DBH > 36 cm | Ash 51–75% Maple 6–15% | Ash 6–15% Beech 16–25% | All layers equally represented. Appears visually unstructured with strongly interlocking crowns. |
| 6 | Mixed (pine, spruce, larch, beech, oak, poplar) | 6 DBH 12–36 cm 7 DBH > 36 cm | Beech 6–15% | Beech 51–75% Poplar 26–59% | Lower an mid layers predominant. Upper layer with irregular distribution of trees. |
| 7 | Coniferous (spruce, larch), deciduous lower layer (rowan) | 9 DBH 12–36 cm | All spec. ≤ 5% | Larch 6–15% | Predominant upper layer with spruce mature trees. Sparse lower an mid layers. |
| 8 | Coniferous (spruce), deciduous lower layer (rowan) | 3 DBH 12–36 cm 3 DBH > 36 cm | All spec. ≤ 5% | Spruce 16–25% | Equal ratio of lower, mid and upper layer trees. Clustered distribution. |
| 9 | Coniferous (spruce), deciduous lower and mid layers (rowan) | 12 DBH > 36 cm | Rowan 16–25% | Spruce 6–15% Rowan 16–25% | Predominant upper and mid layers with spruce trees. Sparse lower layer. |
| Plot № | Mature Trees | Regeneration Trees | |
|---|---|---|---|
| DBH ≥ 12 cm | DBH ≥ 36 cm | DBH < 12 cm | |
| Auto/Ref | Auto/Ref | Auto/Ref | |
| 1 | 7/7 | 4/4 | 8/9 |
| 2 | 10/11 | 6/6 | 1/2 |
| 3 | 10/10 | 1/1 | 0/1 |
| 4 | 13/13 | 4/4 | 6/7 |
| 5 | 15/15 | 3/3 | 3/5 |
| 6 | 10/10 | 3/3 | 9/12 |
| 7 | 4/5 | 3/4 | 4/4 |
| 8 | 4/4 | 2/2 | 9/10 |
| 9 | 2/2 | 10/10 | 15/15 |
| Rate [%] | 97.40 | 97.30 | 84.62 |
| Q1 of Plot № | Producer’s Accuracy [%] | User’s Accuracy [%] | ||
|---|---|---|---|---|
| All Clusters | Incorrect Clusters | All Clusters | Incorrect Clusters | |
| 1 | 99.45 | 89.34 | 98.06 | 95.51 |
| 4 | 89.02 | 65.84 | 87.74 | 85.70 |
| 5 | 92.50 | 64.66 | 96.37 | 87.39 |
| Average | 93.66 | 73.28 | 94.06 | 89.53 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Heinzel, J.; Huber, M.O. Constrained Spectral Clustering of Individual Trees in Dense Forest Using Terrestrial Laser Scanning Data. Remote Sens. 2018, 10, 1056. https://doi.org/10.3390/rs10071056
Heinzel J, Huber MO. Constrained Spectral Clustering of Individual Trees in Dense Forest Using Terrestrial Laser Scanning Data. Remote Sensing. 2018; 10(7):1056. https://doi.org/10.3390/rs10071056
Chicago/Turabian StyleHeinzel, Johannes, and Markus O. Huber. 2018. "Constrained Spectral Clustering of Individual Trees in Dense Forest Using Terrestrial Laser Scanning Data" Remote Sensing 10, no. 7: 1056. https://doi.org/10.3390/rs10071056
APA StyleHeinzel, J., & Huber, M. O. (2018). Constrained Spectral Clustering of Individual Trees in Dense Forest Using Terrestrial Laser Scanning Data. Remote Sensing, 10(7), 1056. https://doi.org/10.3390/rs10071056





