Deep Learning with Unsupervised Data Labeling for Weed Detection in Line Crops in UAV Images
Abstract
1. Introduction
2. Related Work
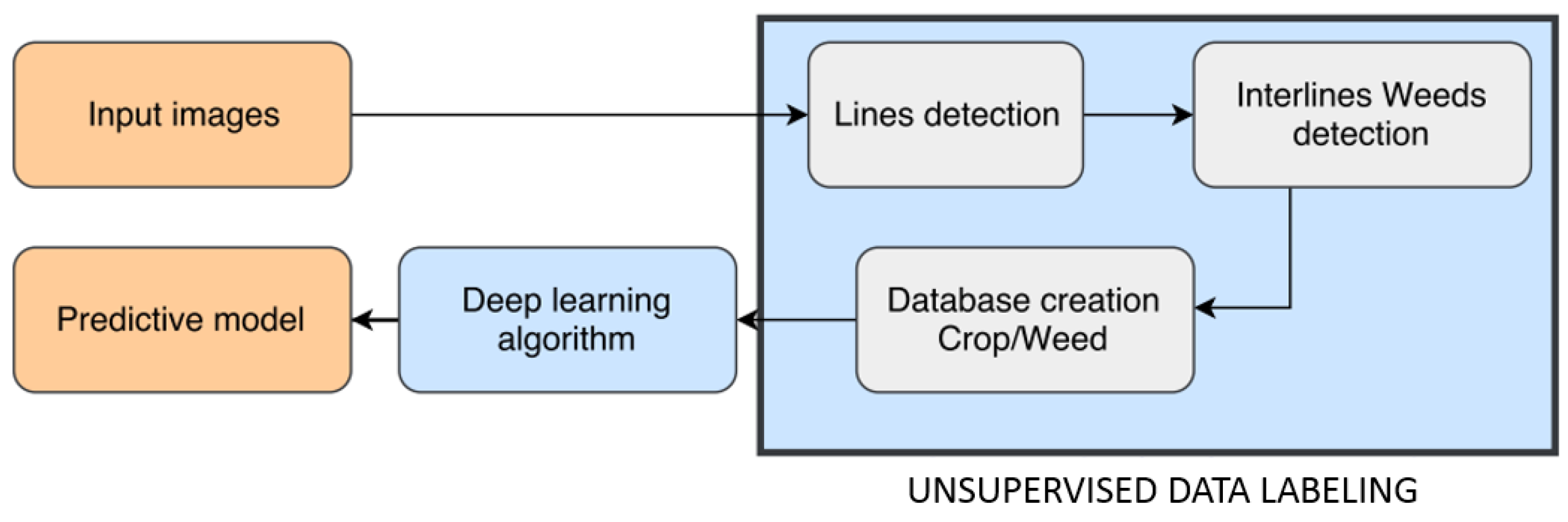
3. Proposed Method
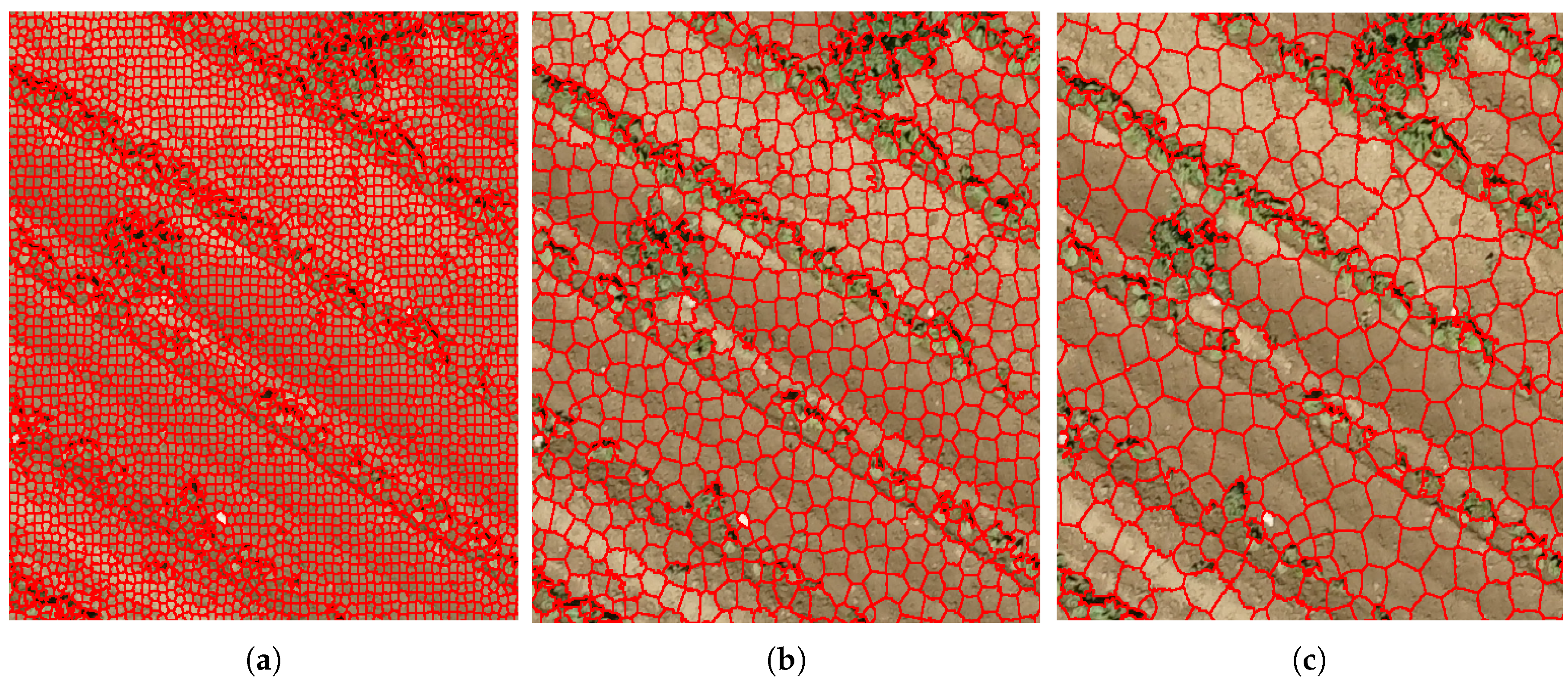
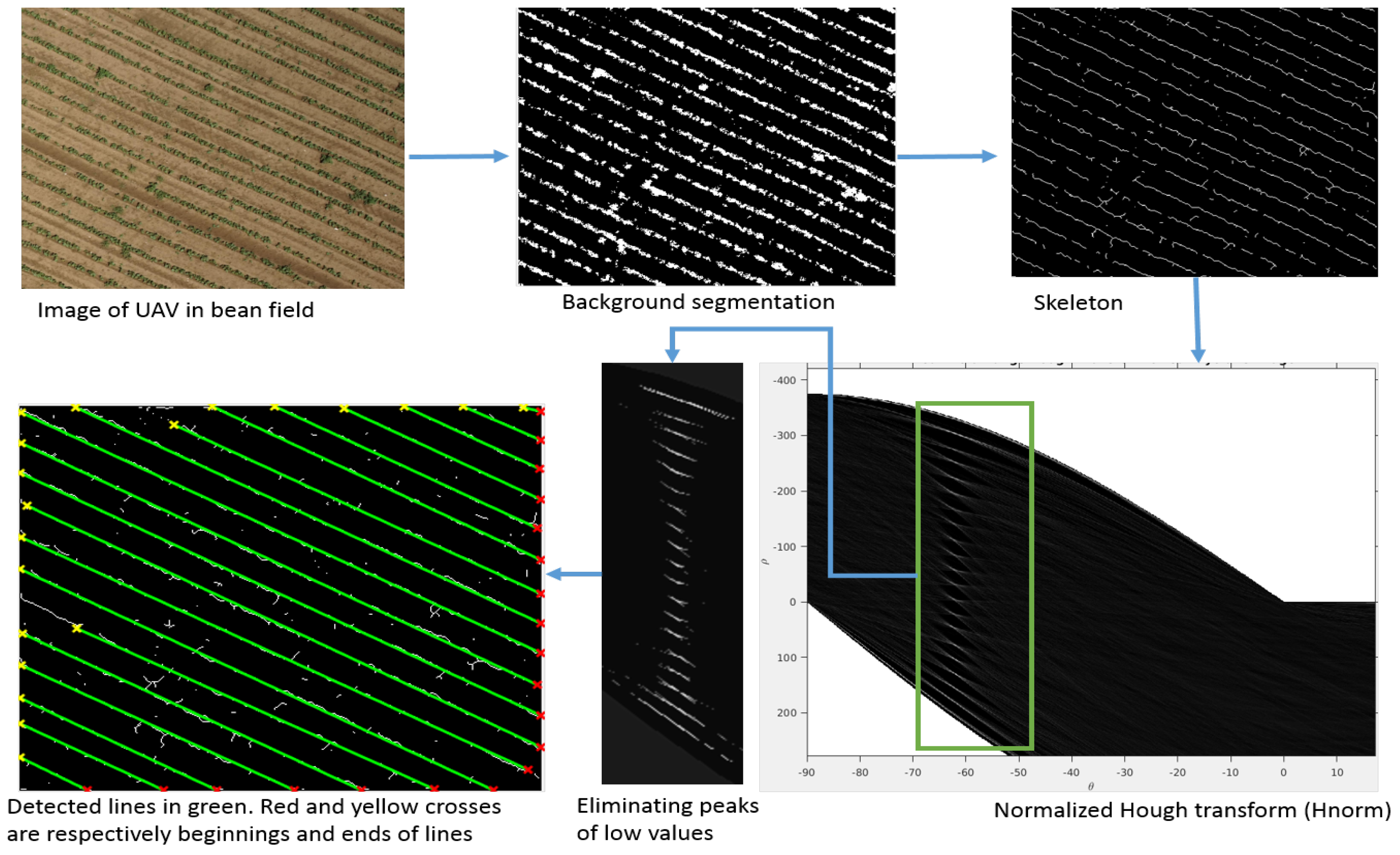
3.1. Detection of Crop Lines
| Algorithm 1: Crop line detection. |
 |
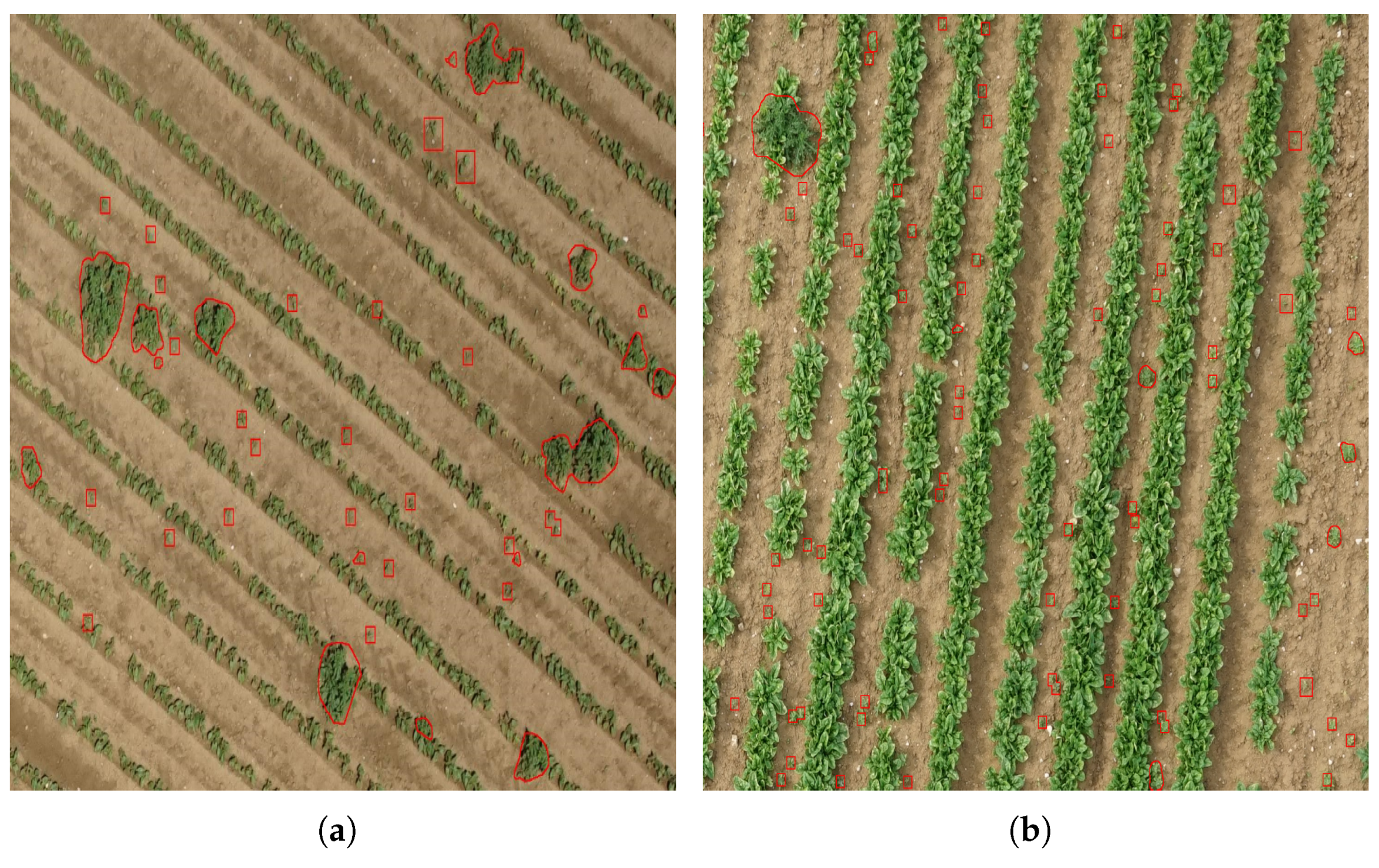
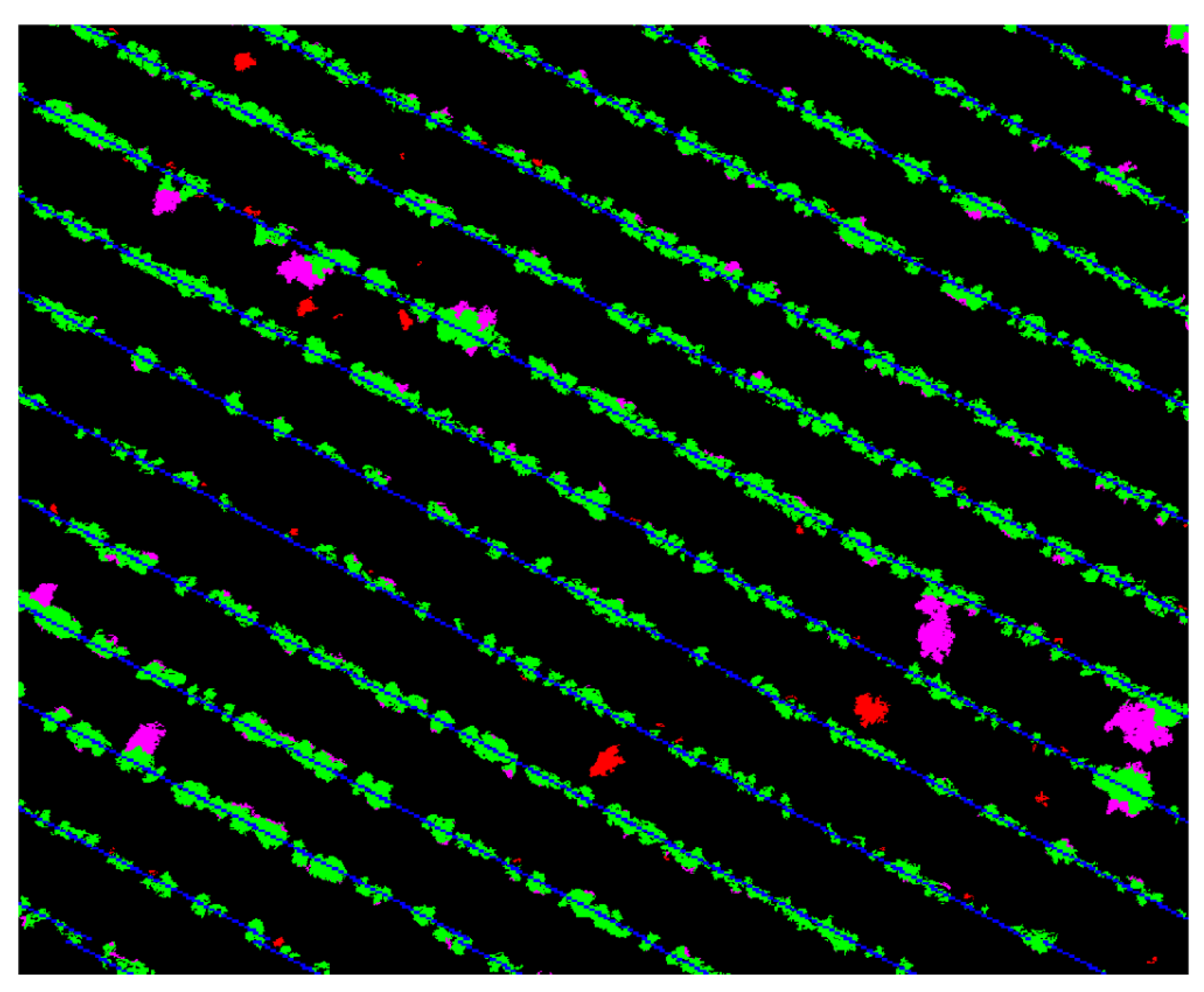
3.2. Unsupervised Training Data Labeling
3.3. Crop/Weed Classification Using Convolutional Neural Networks
3.4. Feature Extraction
3.4.1. Color Features
3.4.2. Geometric Shape Features
3.4.3. Edge density
3.4.4. Histogram of Oriented Gradients (HOG)
3.4.5. Haralick Texture
3.4.6. Gabor Wavelets
3.5. SVM or Support Vector Machine
3.6. Random Forest (RF)
4. Results and Discussion
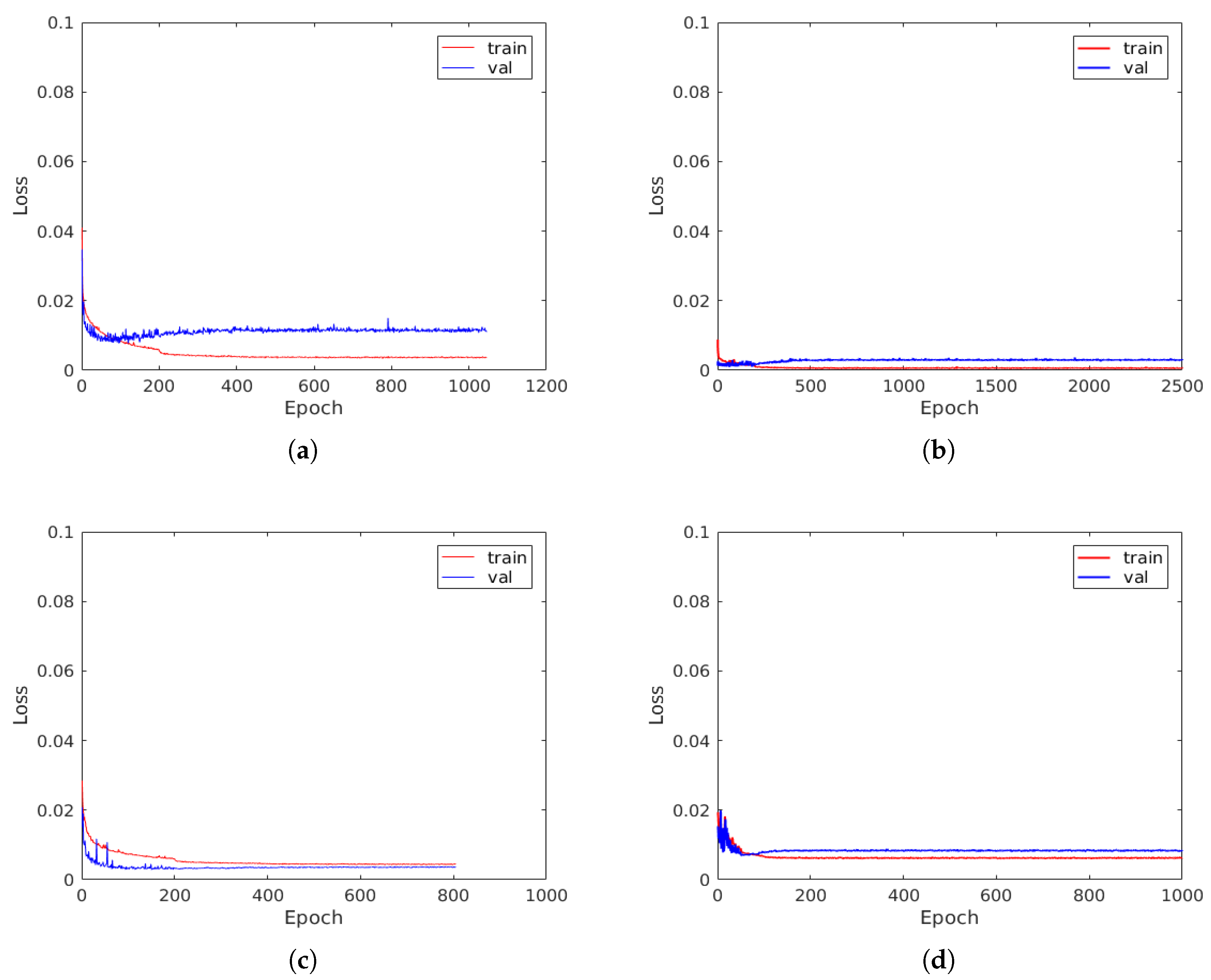
4.1. Results and Discussion
4.2. ResNet vs. Feature Extraction with SVM and RF
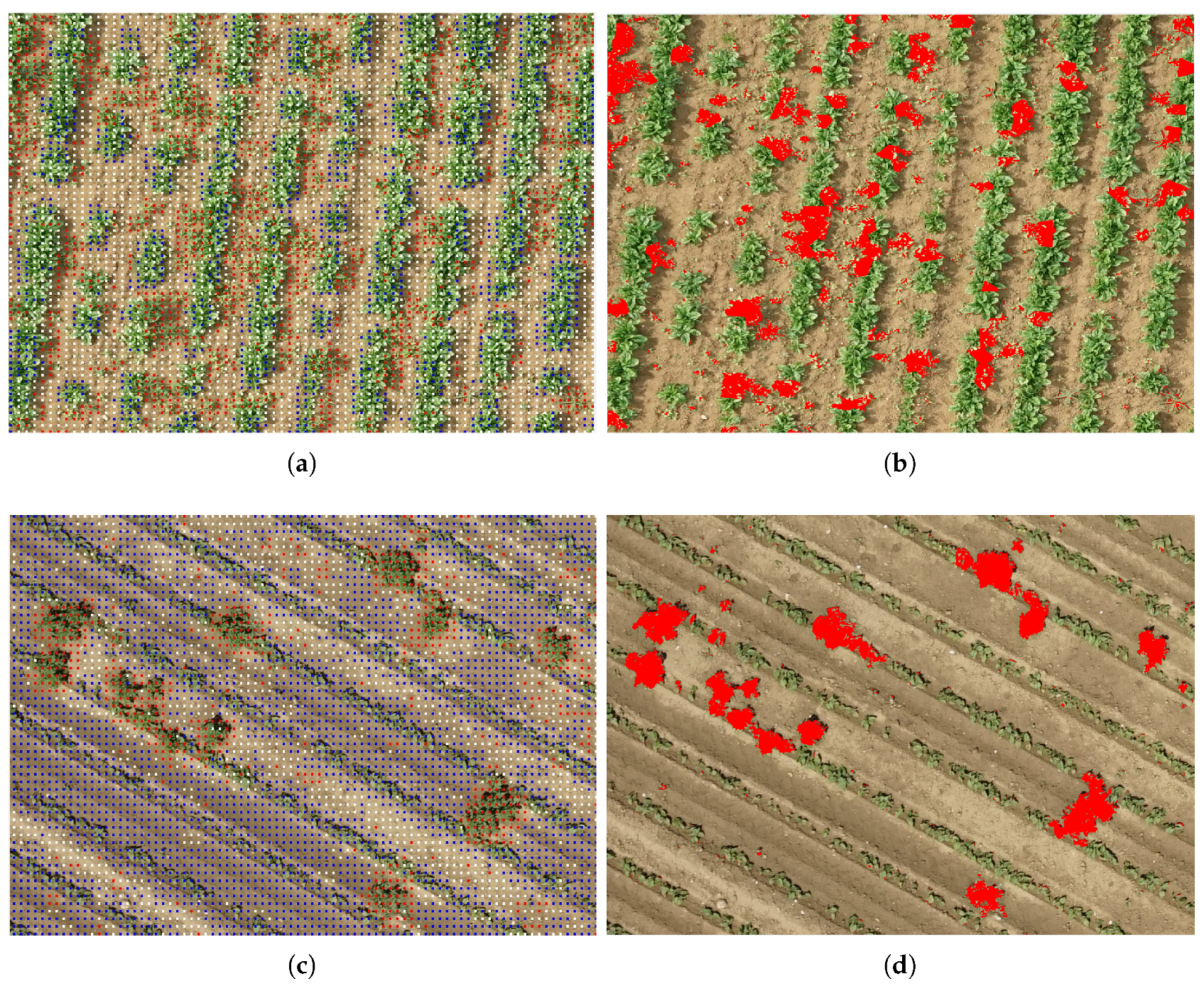
4.3. Weed Detection
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- European Crop Protection. With or Without Pesticides? ECPA: Brussels, Belgium, 2017. [Google Scholar]
- Oerke, E.C. Crop losses to pests. J. Agric. Sci. 2006, 144, 31. [Google Scholar] [CrossRef]
- Pierce, F.J.; Nowak, P. Aspects of Precision Agriculture. Adv. Agron. 1999, 67, 1–85. [Google Scholar] [CrossRef]
- McBratney, A.; Whelan, B.; Ancev, T.; Bouma, J. Future Directions of Precision Agriculture. Precis. Agric. 2005, 6, 7–23. [Google Scholar] [CrossRef]
- Mulla, D.J. Twenty five years of remote sensing in precision agriculture: Key advances and remaining knowledge gaps. Biosyst. Eng. 2013, 114, 358–371. [Google Scholar] [CrossRef]
- Weis, M.; Gutjahr, C.; Ayala, V.R.; Gerhards, R.; Ritter, C.; Schölderle, F. Precision farming for weed management: Techniques. Gesunde Pflanzen 2008, 60, 171–181. [Google Scholar] [CrossRef]
- Herrmann, I.; Shapira, U.; Kinast, S.; Karnieli, A.; Bonfil, D.J. Ground-level hyperspectral imagery for detecting weeds in wheat fields. Precis. Agric. 2013, 14, 637–659. [Google Scholar] [CrossRef]
- Alchanatis, V.; Ridel, L.; Hetzroni, A.; Yaroslavsky, L. Weed detection in multi-spectral images of cotton fields. In Computers and Electronics in Agriculture; Elsevier: New York, NY, USA, 2005; Volume 47, pp. 243–260. [Google Scholar] [CrossRef]
- Torres-Sánchez, J.; López-Granados, F.; Peña, J.M. An automatic object-based method for optimal thresholding in UAV images: Application for vegetation detection in herbaceous crops. Comput. Electron. Agric. 2015, 114, 43–52. [Google Scholar] [CrossRef]
- Zhang, C.; Kovacs, J.M. The application of small unmanned aerial systems for precision agriculture: A review. Precis. Agric. 2012, 13, 693–712. [Google Scholar] [CrossRef]
- Dos Santos Ferreira, A.; Matte Freitas, D.; Gonçalves da Silva, G.; Pistori, H.; Theophilo Folhes, M. Weed detection in soybean crops using ConvNets. Comput. Electron. Agric. 2017, 143, 314–324. [Google Scholar] [CrossRef]
- Bah, M.D.; Dericquebourg, E.; Hafiane, A.; Canals, R. Deep learning based classification system for identifying weeds using high-resolution UAV imagery. In Proceedings of the 2018 Computing Conference, London, UK, July 2018. [Google Scholar]
- Hamuda, E.; Glavin, M.; Jones, E. A survey of image processing techniques for plant extraction and segmentation in the field. Comput. Electron. Agric. 2016, 125, 184–199. [Google Scholar] [CrossRef]
- Gée, C.; Bossu, J.; Jones, G.; Truchetet, F. Crop/weed discrimination in perspective agronomic images. Comput. Electron. Agric. 2008, 60, 49–59. [Google Scholar] [CrossRef]
- Woebbecke, D.; Meyer, G.; Von Bargen, K.; Mortensen, D. Color indices for weed identification under various soil, residue, and lighting conditions. Trans. ASAE 1995, 38, 259–269. [Google Scholar] [CrossRef]
- Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. 1979, 9, 62–66. [Google Scholar] [CrossRef]
- Hough, P.V.C. Method and Means for Recognizing Complex Patterns. US Patent 3,069,654, 18 December 2018. [Google Scholar] [CrossRef]
- Peña, J.M.; Torres-Sánchez, J.; Isabel De Castro, A.; Kelly, M.; López-Granados, F. Weed Mapping in Early-Season Maize Fields Using Object-Based Analysis of Unmanned Aerial Vehicle (UAV) Images. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Blaschke, T. Object based image analysis for remote sensing. ISPRS J. Photogramm. Remote Sens. 2010, 65, 2–16. [Google Scholar] [CrossRef]
- Tang, L.; Tian, L.; Steward, B.L. Classification of Broadleaf and Grass Weeds Using Gabor Wavelets and an Artificial Neural Network. Trans. Am. Soc. Agric. Eng. 2003, 46, 1247–1254. [Google Scholar] [CrossRef]
- Jeon, H.Y.; Tian, L.F.; Zhu, H. Robust crop and weed segmentation under uncontrolled outdoor illumination. Sensors 2011, 11, 6270–6283. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, F.; Al-Mamun, H.A.; Bari, A.S.M.H.; Hossain, E.; Kwan, P. Classification of crops and weeds from digital images: A support vector machine approach. Crop Protect. 2012, 40, 98–104. [Google Scholar] [CrossRef]
- Latha, M.; Poojith, A.; Reddy, A.; Kumar, V. Image Processing in Agriculture. Int. J. Innov. Res. Electr. Electron. Instrum. Control Eng. 2014, 2. [Google Scholar]
- Pérez-Ortiz, M.; Peña, J.M.; Gutiérrez, P.A.; Torres-Sánchez, J.; Hervás-Martínez, C.; López-Granados, F. Selecting patterns and features for between- and within-crop-row weed mapping using UAV-imagery. Expert Syst. Appl. 2015, 47, 85–94. [Google Scholar] [CrossRef]
- Bakhshipour, A.; Jafari, A.; Nassiri, S.M.; Zare, D. Weed segmentation using texture features extracted from wavelet sub-images. Biosyst. Eng. 2017, 157, 1–12. [Google Scholar] [CrossRef]
- Bakhshipour, A.; Jafari, A. Evaluation of support vector machine and artificial neural networks in weed detection using shape features. Comput. Electron. Agric. 2018, 145, 153–160. [Google Scholar] [CrossRef]
- Gao, J.; Liao, W.; Nuyttens, D.; Lootens, P.; Vangeyte, J.; Pižurica, A.; He, Y.; Pieters, J.G. Fusion of pixel and object-based features for weed mapping using unmanned aerial vehicle imagery. Int. J. Appl. Earth Observ. Geoinf. 2018, 67, 43–53. [Google Scholar] [CrossRef]
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2323. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. In Advances in Neural Information Processing Systems; MIT Press Ltd.: Cambridge, MA, USA, 2012; pp. 1–9. [Google Scholar] [CrossRef]
- Hung, C.; Xu, Z.; Sukkarieh, S. Feature Learning Based Approach for Weed Classification Using High Resolution Aerial Images from a Digital Camera Mounted on a UAV. Remote Sens. 2014, 6, 12037–12054. [Google Scholar] [CrossRef]
- Mortensen, A.K.; Dyrmann, M.; Karstoft, H.; Nyholm Jørgensen, R.; Gislum, R. Semantic Segmentation of Mixed Crops using Deep Convolutional Neural Network. In Proceedings of the International Conference on Agricultural Engineering, Aarhus, Denmark, 26–29 June 2016. [Google Scholar]
- Milioto, A.; Lottes, P.; Stachniss, C. Real-time blob-wise sugar beets vs. weeds classification for monitoring fields using convolutional neural networks. ISPRS Ann. Photogramm. Remote Sens. Spat. Inf. Sci. 2017, IV-2/W3, 41–48. [Google Scholar] [CrossRef]
- Di Cicco, M.; Potena, C.; Grisetti, G.; Pretto, A. Automatic model based dataset generation for fast and accurate crop and weeds detection. In Proceedings of the 2017 IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS), Vancouver, BC, Canada, 24–28 September 2017; pp. 5188–5195. [Google Scholar] [CrossRef]
- Louargant, M.; Jones, G.; Faroux, R.; Paoli, J.N.; Maillot, T.; Gée, C.; Villette, S. Unsupervised Classification Algorithm for Early Weed Detection in Row-Crops by Combining Spatial and Spectral Information. Remote Sens. 2018, 10, 761. [Google Scholar] [CrossRef]
- Jones, G.; Gée, C.; Truchetet, F. Modelling agronomic images for weed detection and comparison of crop/weed discrimination algorithm performance. Precis. Agric. 2009, 10, 1–15. [Google Scholar] [CrossRef]
- Montalvo, M.; Pajares, G.; Guerrero, J.M.; Romeo, J.; Guijarro, M.; Ribeiro, A.; Ruz, J.J.; Cruz, J.M. Automatic detection of crop rows in maize fields with high weeds pressure. Expert Syst. Appl. 2012, 39, 11889–11897. [Google Scholar] [CrossRef]
- Bah, M.D.; Hafiane, A.; Canals, R. Weeds detection in UAV imagery using SLIC and the hough transform. In Proceedings of the 2017 Seventh International Conference on Image Processing Theory, Tools and Applications (IPTA), Montreal, QC, Canada, 28 November–1 December 2017; pp. 1–6. [Google Scholar] [CrossRef]
- Achanta, R.; Shaji, A.; Smith, K.; Lucchi, A.; Fua, P.; Süsstrunk, S. SLIC Superpixels Compared to State-of-the-Art Superpixel Methods. IEEE Trans. Pattern Anal. Mach. Intell. 2011, 34, 2274–2282. [Google Scholar] [CrossRef] [PubMed]
- Kamilaris, A.; Prenafeta-Boldú, F.X. Deep learning in agriculture: A survey. Comput. Electron. Agric. 2018, 147, 70–90. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 26 June–1 July 2016; pp. 770–778. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. arXiv, 2015; arXiv:1409.1556. [Google Scholar]
- Pan, S.J.; Yang, Q. A Survey on Transfer Learning. IEEE Trans. Knowl. Data Eng. 2010, 22, 1345–1359. [Google Scholar] [CrossRef]
- Huang, Z.; Pan, Z.; Lei, B. Transfer learning with deep convolutional neural network for SAR target classification with limited labeled data. Remote Sens. 2017, 9, 907. [Google Scholar] [CrossRef]
- Lottes, P.; Khanna, R.; Pfeifer, J.; Siegwart, R.; Stachniss, C. UAV-based crop and weed classification for smart farming. In Proceedings of the 2017 IEEE International Conference on Robotics and Automation (ICRA), Sands Expo, NV, USA, 29 May–3 June 2017; pp. 3024–3031. [Google Scholar] [CrossRef]
- Dalal, N.; Triggs, W.; Triggs, B. Histograms of Oriented Gradients for Human Detection. In Proceedings of the 2005 IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR’05), San Diego, CA, USA, 20–26 June 2005. [Google Scholar] [CrossRef]
- Xiao, X.Y.; Hu, R.; Zhang, S.W.; Wang, X.F. HOG-based approach for leaf classification. Advanced Intelligent Computing Theories and Applications. In Proceedings of the 6th International Conference on Intelligent Computing, ICIC 2010, Changsha, China, 18–21 August 2010; Springer: Berlin/Heidelberg, Germany, 2010; Volume 6216, pp. 149–155. [Google Scholar] [CrossRef]
- Ibrahim, Z.; Sabri, N.; Abu Mangshor, N.N. Leaf Recognition using Texture Features for Herbal Plant Identification. Indones. J. Electr. Eng. Comput. Sci. 2018, 9, 152–156. [Google Scholar] [CrossRef]
- Haralick, R.M.; Shanmugam, K. Textural Features for Image Classification. Syst. Man Cybern. 1973, 3, 610–621. [Google Scholar] [CrossRef]
- Agrawal, K.N.; Singh, K.; Bora, G.C.; Lin, D. Weed Recognition Using Image-Processing Technique Based on Leaf Parameters. J. Agric. Sci. Technol. B J. Agric. Sci. Technol. 2012, 2, 899. [Google Scholar]
- Soh, L.K.; Tsatsoulis, C. Texture analysis of sar sea ice imagery using gray level co-occurrence matrices. IEEE Trans. Geosci. Remote Sens. 1999, 37, 780–795. [Google Scholar] [CrossRef]
- Brynolfsson, P.; Nilsson, D.; Torheim, T.; Asklund, T.; Karlsson, C.T.; Trygg, J.; Nyholm, T.; Garpebring, A. Haralick texture features from apparent diffusion coefficient (ADC) MRI images depend on imaging and pre-processing parameters. Sci. Rep. 2017, 7, 4041. [Google Scholar] [CrossRef] [PubMed]
- Bovik, A.C.; Clark, M.; Geisler, W.S. Multichannel Texture Analysis Using Localized Spatial Filters. IEEE Trans. Pattern Anal. Mach. Intell. 1990, 12, 55–73. [Google Scholar] [CrossRef]
- Naghdy, G.A. Texture analysis using Gabor wavelets. Proc. SPIE 1996, 2657, 74–85. [Google Scholar] [CrossRef]
- Cortes, C.; Vapnik, V. Support-vector networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Kazmi, W.; Garcia-Ruiz, F.; Nielsen, J.; Rasmussen, J.; Andersen, H.J. Exploiting affine invariant regions and leaf edge shapes for weed detection. Comput. Electron. Agric. 2015, 118, 290–299. [Google Scholar] [CrossRef]
- Yang, Y.; Rushmeier, H. Special object extraction from medieval books using superpixels and bag-of-features. J. Electron. Imaging 2016, 26, 011008. [Google Scholar] [CrossRef]
- Cristianini, N.; Shawe-Taylor, J. An Introduction to Support Vector Machines and Other Kernel Based Learning Methods; Cambridge University Press: Cambridge, UK, 2000; Volume 22, p. 190. [Google Scholar]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- De Castro, A.I.; Torres-Sánchez, J.; Peña, J.M.; Jiménez-Brenes, F.M.; Csillik, O.; López-Granados, F. An automatic random forest-OBIA algorithm for early weed mapping between and within crop rows using UAV imagery. Remote Sens. 2018, 10, 285. [Google Scholar] [CrossRef]
- Geurts, P.; Ernst, D.; Wehenkel, L. Extremely randomized trees. Mach. Learn. 2006, 63, 3–42. [Google Scholar] [CrossRef]
- Tucker, C.J. Red and photographic infrared linear combinations for monitoring vegetation. Remote Sens. Environ. 1979, 8, 127–150. [Google Scholar] [CrossRef]
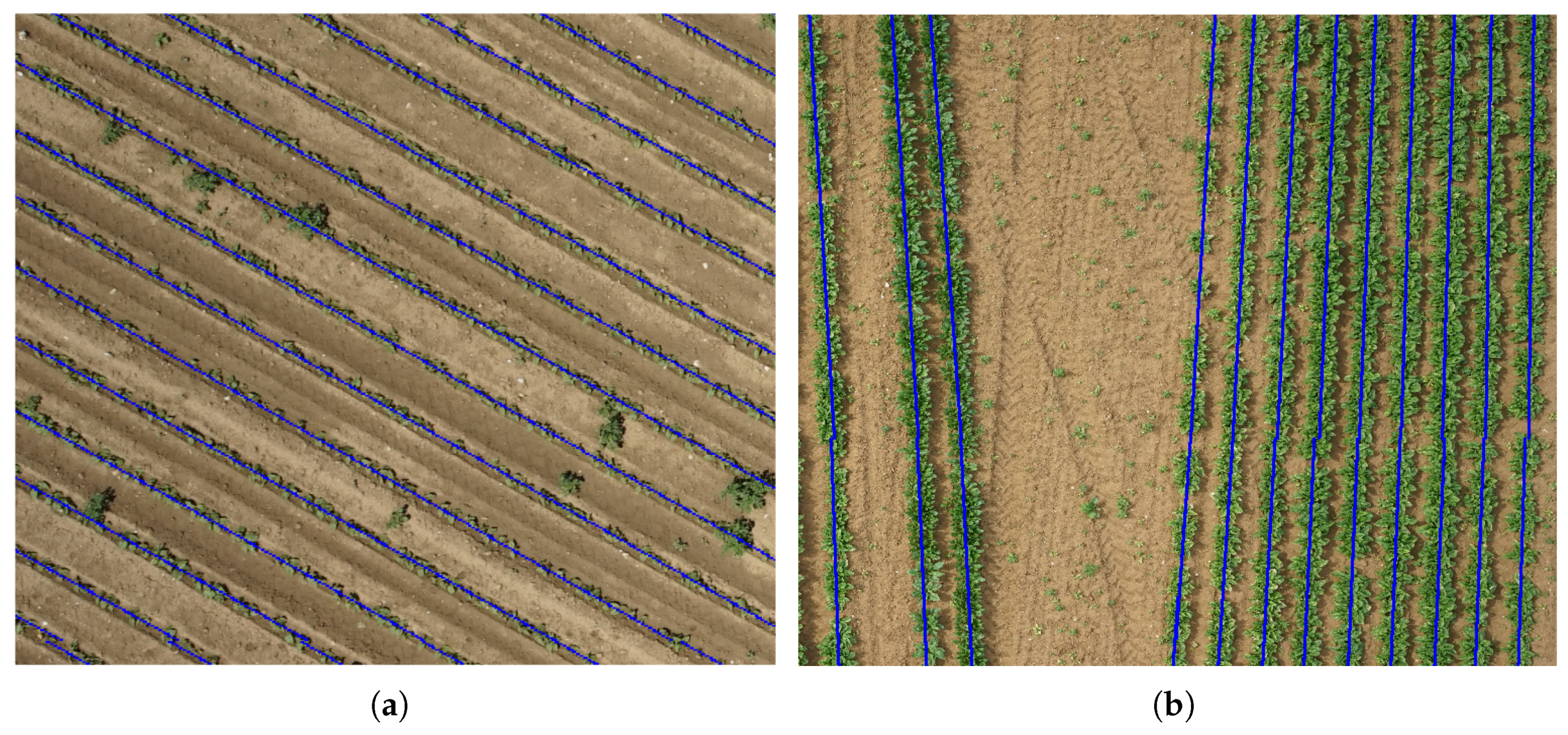
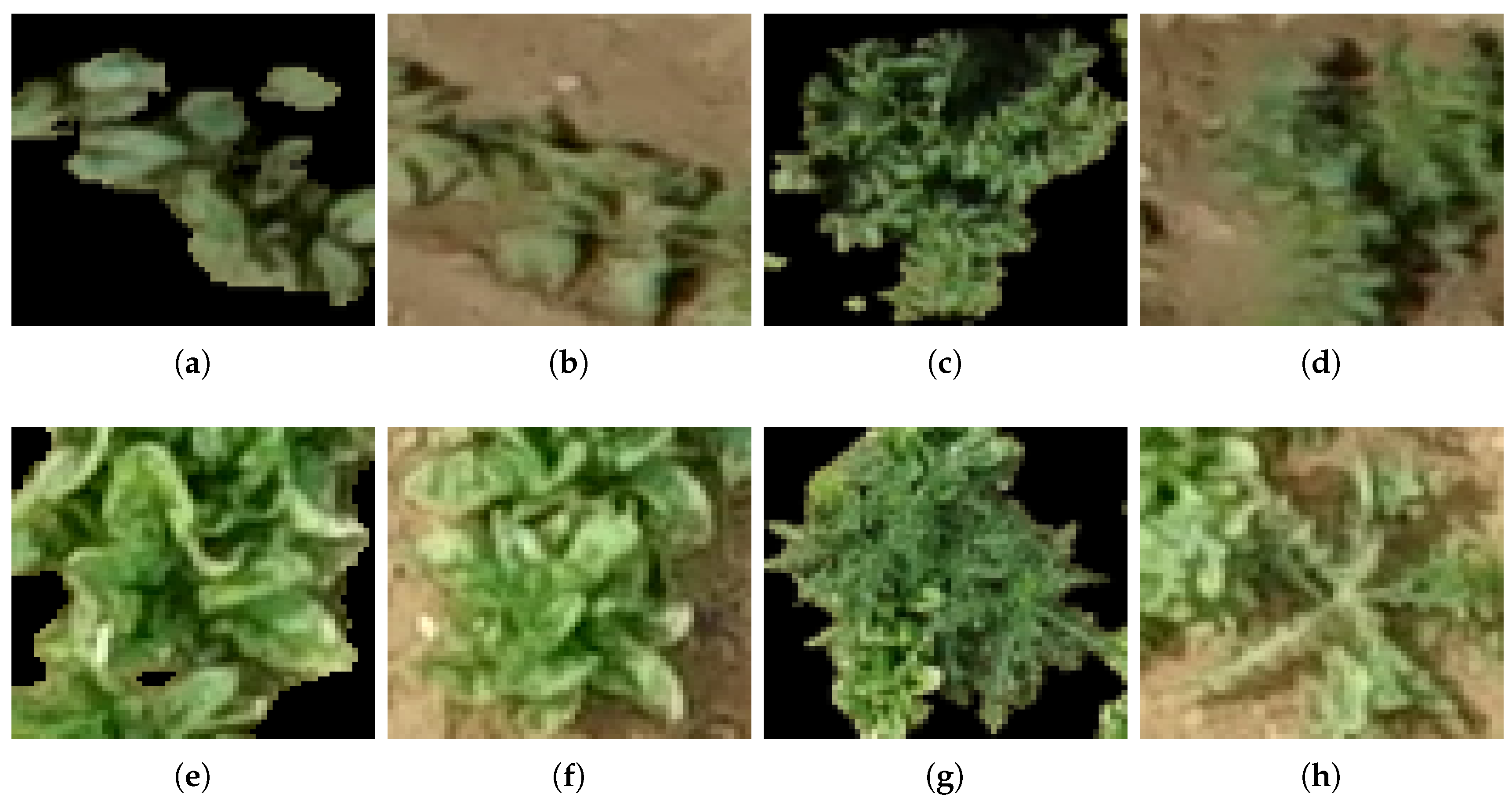






| Bean Field | ||||
| Data | Class | Training | Validation | Total |
| Supervised | Crop | 17,192 | 11,694 | 28,886 |
| labeling | Weed | 17,076 | 9060 | 16,136 |
| Total | 34,868 | 20,754 | 45,022 | |
| Unsupervised | Crop | 7688 | 1928 | 9616 |
| labeling | Weed | 5935 | 1493 | 7428 |
| Total | 13,623 | 3421 | 17,044 | |
| Spinach field | ||||
| Data | Class | Training | Validation | Total |
| Supervised | Crop | 11,350 | 2838 | 14,188 |
| labeling | Weed | 8234 | 2058 | 10,292 |
| Total | 19,584 | 4896 | 34,772 | |
| Unsupervised | Crop | 6884 | 1722 | 8606 |
| labeling | Weed | 5800 | 1452 | 7252 |
| Total | 12,684 | 3174 | 15,858 | |
| Field | Crop Samples | Weed Samples |
|---|---|---|
| Bean | 2139 | 1852 |
| Spinach | 1523 | 1825 |
| SVM (AUC%) | RF (AUC%) | ResNet18 (AUC%) | ||||
|---|---|---|---|---|---|---|
| Best Features | Sup | Unsup | Sup | Unsup | Sup | Unsup |
| Labeling | Labeling | Labeling | Labeling | Labeling | Labeling | |
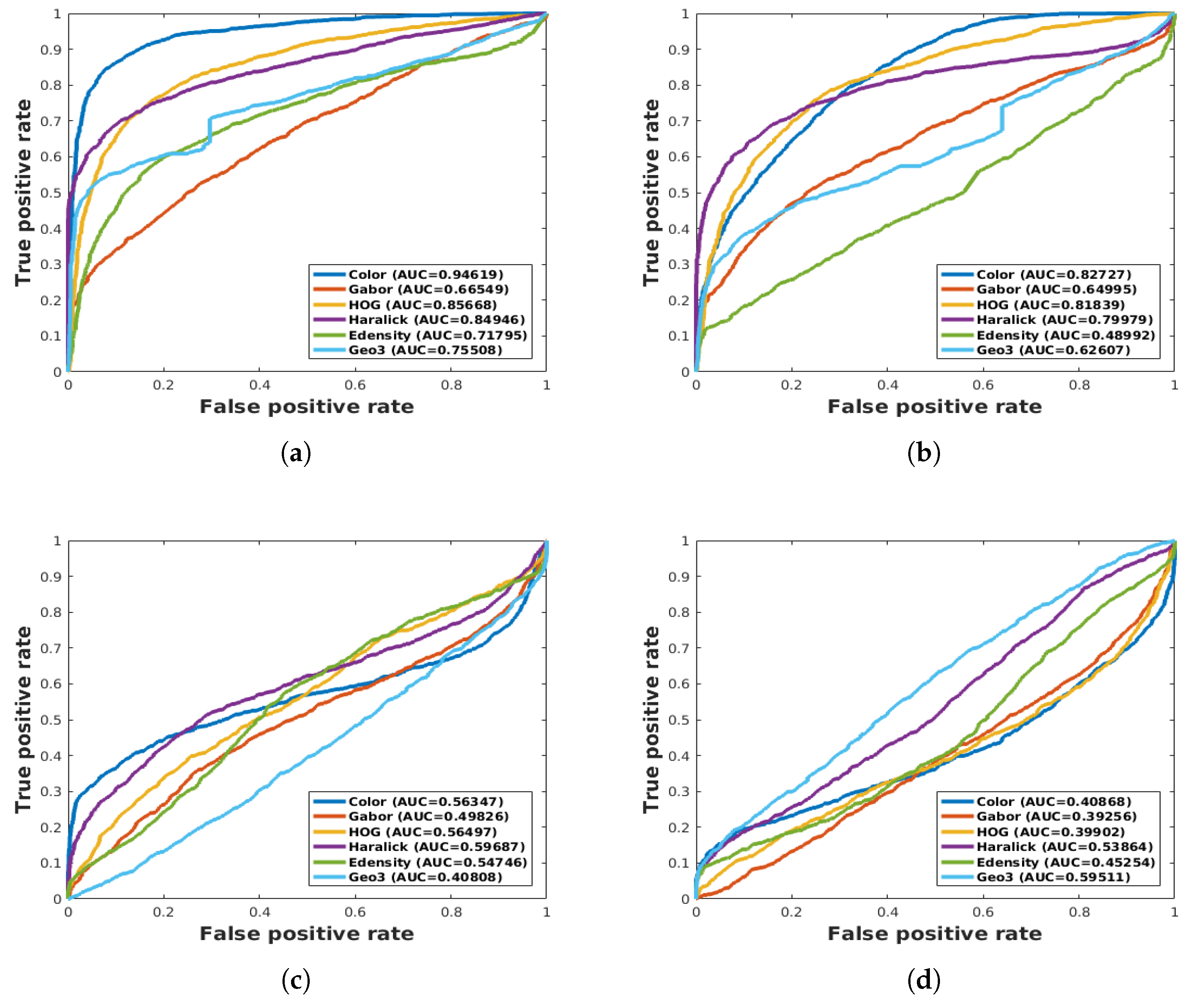
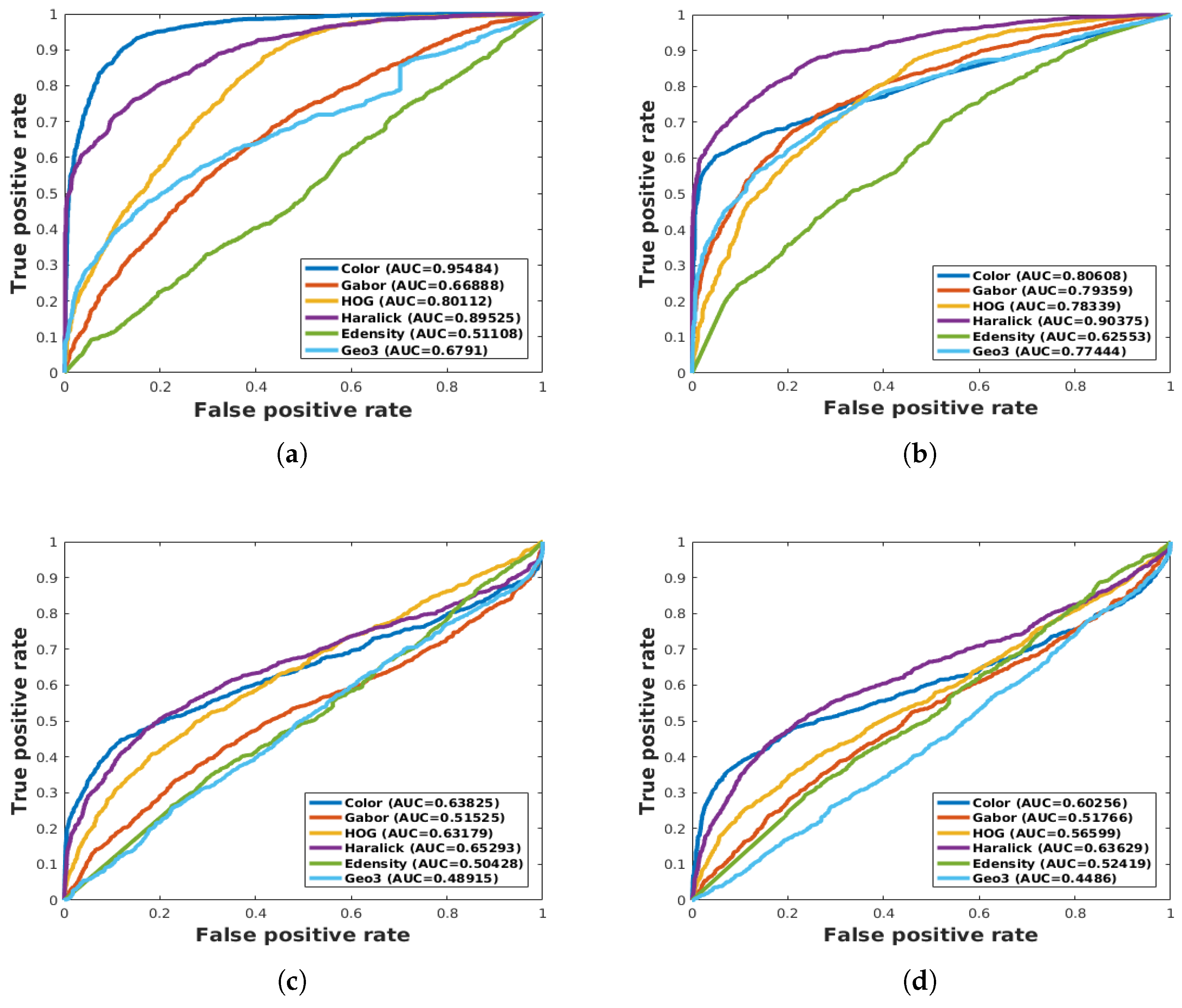
| ALL features | 60.60 | 44.76 | 70.16 | 63.95 | - | - |
| Geo3 | 40.80 | 59.51 | 48.91 | 44.86 | - | - |
| Haralick, Color | 59.78 | 40.46 | 68.15 | 65.40 | - | - |
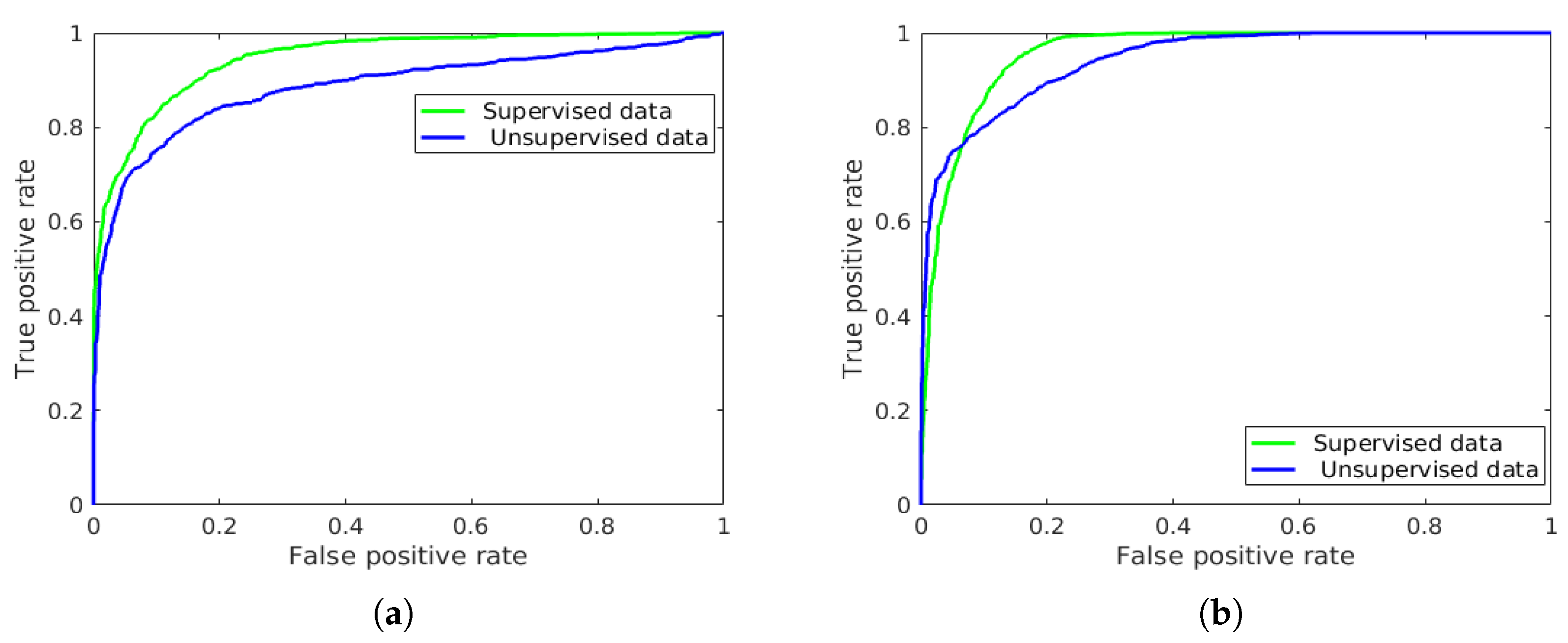
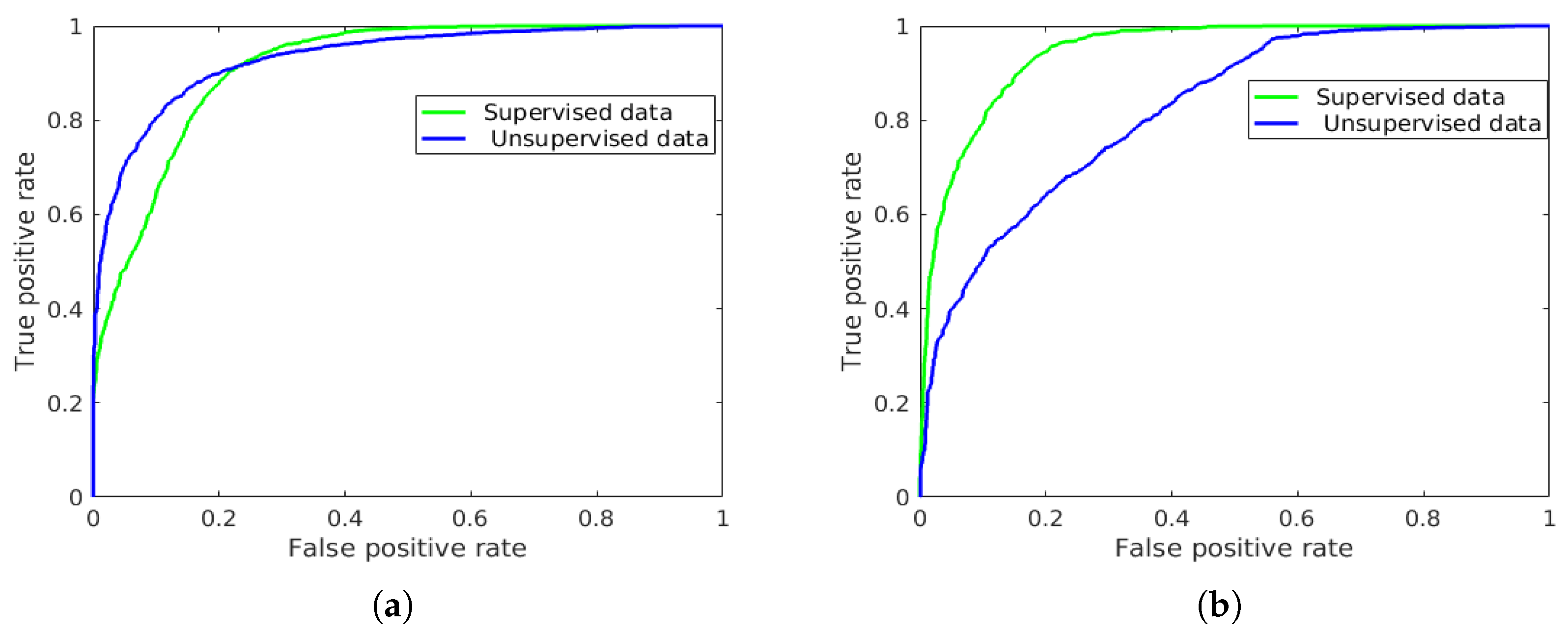
| - | - | - | - | - | 94.84 | 88.73 |
| SVM (AUC%) | RF (AUC%) | ResNet18 (AUC%) | ||||
|---|---|---|---|---|---|---|
| Best Features | Sup | Unsup | Sup | Unsup | Sup | Unsup |
| Labeling | Labeling | Labeling | Labeling | Labeling | Labeling | |
| Color, HOG, Gabor | 95.94 | 87.38 | 93.50 | 95.131 | - | - |
| Haralick, Color, HOG, Gabor | 93.93 | 90.77 | 95.464 | 96.177 | - | - |
| All features | 93.352 | 90.70 | 96.99 | 95.162 | - | - |
| - | - | - | - | - | 95.70 | 94.34 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bah, M.D.; Hafiane, A.; Canals, R. Deep Learning with Unsupervised Data Labeling for Weed Detection in Line Crops in UAV Images. Remote Sens. 2018, 10, 1690. https://doi.org/10.3390/rs10111690
Bah MD, Hafiane A, Canals R. Deep Learning with Unsupervised Data Labeling for Weed Detection in Line Crops in UAV Images. Remote Sensing. 2018; 10(11):1690. https://doi.org/10.3390/rs10111690
Chicago/Turabian StyleBah, M Dian, Adel Hafiane, and Raphael Canals. 2018. "Deep Learning with Unsupervised Data Labeling for Weed Detection in Line Crops in UAV Images" Remote Sensing 10, no. 11: 1690. https://doi.org/10.3390/rs10111690
APA StyleBah, M. D., Hafiane, A., & Canals, R. (2018). Deep Learning with Unsupervised Data Labeling for Weed Detection in Line Crops in UAV Images. Remote Sensing, 10(11), 1690. https://doi.org/10.3390/rs10111690





