The Post-COVID-19 Era: Interdisciplinary Demands of Contagion Surveillance Mass Spectrometry for Future Pandemics
Abstract
:1. Introduction
2. Possible Pathogen Detection Using Benchtop MS with Machine Learning
3. Need for Viral Degradation Study Using MS to Understand the Range of Infectivity
4. A Potential Way for Detecting Viral Infectivity in Aerosols, Droplets and Fomites Using MS with Van Krevelen Diagram
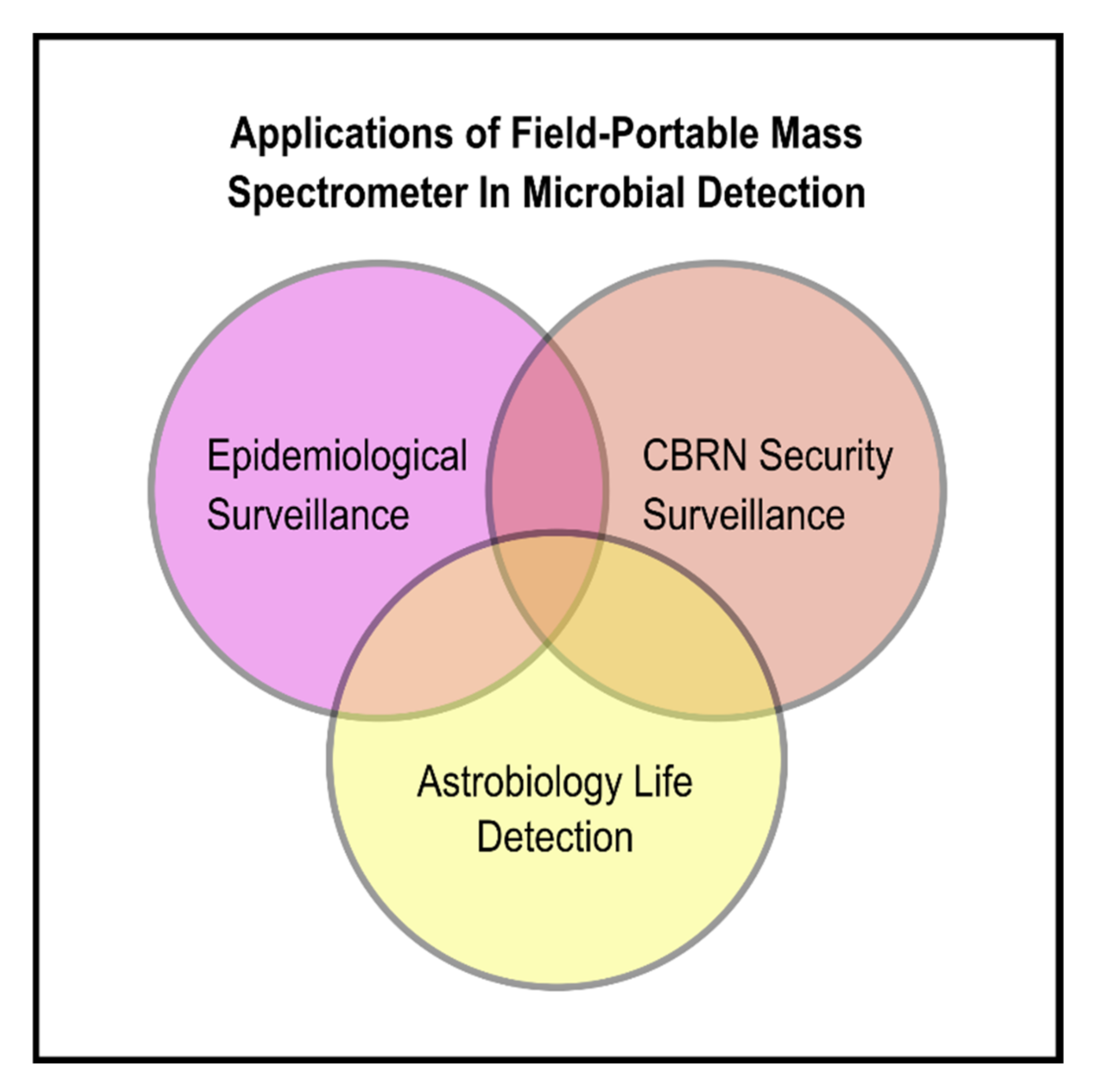
5. Sharing Engineering Attributes of MS used in Astrobiology and CBRN Monitoring with Future MS for In Situ Contagion-Laden ADF Surveillance
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Durzyńska, J.; Goździcka-Józefiak, A. Viruses and Cells Intertwined since the Dawn of Evolution. Virol. J. 2015, 12, 169. [Google Scholar] [CrossRef] [Green Version]
- Hegde, N.R.; Maddur, M.S.; Kaveri, S.V.; Bayry, J. Reasons to Include Viruses in the Tree of Life. Nat. Rev. Microbiol. 2009, 7, 615. [Google Scholar] [CrossRef]
- Moreira, D.; López-García, P. Ten Reasons to Exclude Viruses from the Tree of Life. Nat. Rev. Microbiol. 2009, 7, 306–311. [Google Scholar] [CrossRef]
- Sorensen, G. The Role of the Virus-Phytoplankton System in Marine Biogeochemical Cycling: Possible Impacts of Climate Change; Univerity of Plymouth: Plymouth, UK, 2009. [Google Scholar]
- Rohwer, F.; Prangishvili, D.; Lindell, D. Roles of Viruses in the Environment. Environ. Microbiol. 2009, 11, 2771–2774. [Google Scholar] [CrossRef]
- WHO Announces COVID-19 Outbreak a Pandemic. 2020. Available online: https://www.euro.who.int/en/health-topics/health-emergencies/coronavirus-covid-19/news/news/2020/3/who-announces-covid-19-outbreak-a-pandemic (accessed on 12 September 2020).
- Al Huraimel, K.; Alhosani, M.; Kunhabdulla, S.; Stietiya, M.H. SARS-CoV-2 in the Environment: Modes of Transmission, Early Detection and Potential Role of Pollutions. Sci. Total Environ. 2020, 744, 140946. [Google Scholar] [CrossRef]
- Liu, Y.; Ning, Z.; Chen, Y.; Guo, M.; Liu, Y.; Gali, N.K.; Sun, L.; Duan, Y.; Cai, J.; Westerdahl, D.; et al. Aerodynamic Analysis of SARS-CoV-2 in Two Wuhan Hospitals. Nature 2020. [Google Scholar] [CrossRef] [PubMed]
- Santarpia, J.L.; Rivera, D.N.; Herrera, V.; Morwitzer, M.J.; Creager, H.; Santarpia, G.W.; Crown, K.K.; Brett-Major, D.; Schnaubelt, E.; Broadhurst, M.J.; et al. Aerosol and surface contamination of SARS-CoV-2 observed in quarantine and isolation care. Sci. Rep. 2020, 10, 12732. [Google Scholar] [CrossRef] [PubMed]
- Chia, P.Y.; Coleman, K.K.; Tan, Y.K.; Ong, S.W.X.; Gum, M.; Lau, S.K.; Lim, X.F.; Lim, A.S.; Sutjipto, S.; Lee, P.H.; et al. Detection of Air and Surface Contamination by SARS-CoV-2 in Hospital Rooms of Infected Patients. Nat. Commun. 2020, 11, 2800. [Google Scholar] [CrossRef]
- Aydogdu, M.O.; Altun, E.; Chung, E.; Ren, G.; Homer-Vanniasinkam, S.; Chen, B.; Edirisinghe, M. Surface Interactions and Viability of Coronaviruses. J. R. Soc. Interface 2021, 18, 20200798. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical Features of Patients Infected with 2019 Novel Coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef] [Green Version]
- Moriarty, L.F.; Plucinski, M.M.; Marston, B.J.; Kurbatova, E.V.; Knust, B.; Murray, E.L.; Pesik, N.; Rose, D.; Fitter, D.; Kobayashi, M.; et al. Public Health Responses to COVID-19 Outbreaks on Cruise Ships—Worldwide, February-March 2020. MMWR Morb. Mortal. Wkly. Rep. 2020, 69, 347–352. [Google Scholar] [CrossRef]
- Nor, N.S.; Yip, C.W.; Ibrahim, N.; Jaafar, M.H.; Rashid, Z.Z.; Mustafa, N.; Hamid, H.H.A.; Chandru, K.; Latif, M.T.; Saw, P.E.; et al. Particulate Matter (PM2.5) as a Potential SARS-CoV-2 Carrier. Sci. Rep. 2021, 11, 2508. [Google Scholar] [CrossRef]
- Van Doremalen, N.; Bushmaker, T.; Morris, D.H.; Holbrook, M.G.; Gamble, A.; Williamson, B.N.; Tamin, A.; Harcourt, J.L.; Thornburg, N.J.; Gerber, S.I.; et al. Aerosol and Surface Stability of SARS-CoV-2 as Compared with SARS-CoV-1. N. Engl. J. Med. 2020, 382, 1564–1567. [Google Scholar] [CrossRef] [PubMed]
- Tellier, R.; Li, Y.; Cowling, B.J.; Tang, J.W. Recognition of Aerosol Transmission of Infectious Agents: A Commentary. BMC Infect. Dis. 2019, 19, 101. [Google Scholar] [CrossRef]
- Wilson, N.M.; Norton, A.; Young, F.P.; Collins, D.W. Airborne Transmission of Severe Acute Respiratory Syndrome Coronavirus-2 to Healthcare Workers: A Narrative Review. Anaesthesia 2020. [Google Scholar] [CrossRef]
- Wei, J.; Li, Y. Airborne Spread of Infectious Agents in the Indoor Environment. Am. J. Infect. Control. 2016, 44, S102–S108. [Google Scholar] [CrossRef]
- Xie, X.; Li, Y.; Chwang, A.T.Y.; Ho, P.L.; Seto, W.H. How Far Droplets Can Move in Indoor Environments—Revisiting the Wells Evaporation-Falling Curve. Indoor Air 2007, 17, 211–225. [Google Scholar] [CrossRef]
- Stadnytskyi, V.; Bax, C.E.; Bax, A.; Anfinrud, P. The Airborne Lifetime of Small Speech Droplets and Their Potential Importance in SARS-CoV-2 Transmission. Proc. Natl. Acad. Sci. USA 2020. [Google Scholar] [CrossRef] [PubMed]
- Saw, L.H.; Leo, B.F.; Nor, N.S.M.; Yip, C.W.; Ibrahim, N.; Hamid, H.H.A.; Latif, M.T.; Lin, C.Y.; Nadzir, M.S.M. Modeling Aerosol Transmission of SARS-CoV-2 from Human-Exhaled Particles in a Hospital Ward. Environ. Sci. Pollut. Res. Int. 2021. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, M.; Lee, S.J.; Epstein, M. Modeling Aerosol Transmission of SARS-CoV-2 in Multi-Room Facility. J. Loss Prev. Process. Indust. 2021, 69, 104336. [Google Scholar] [CrossRef]
- Azimi, P.; Keshavarz, Z.; Cedeno Laurent, J.G.; Stephens, B.; Allen, J.G. Mechanistic Transmission Modeling of COVID-19 on the Diamond Princess Cruise Ship Demonstrates the Importance of Aerosol Transmission. Proc. Natl. Acad. Sci. USA 2021, 118. [Google Scholar] [CrossRef]
- Shereen, M.A.; Khan, S.; Kazmi, A.; Bashir, N.; Siddique, R. COVID-19 Infection: Origin, Transmission, and Characteristics of Human Coronaviruses. J. Advert. Res. 2020, 24, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; et al. Structure of the SARS-CoV-2 Spike Receptor-Binding Domain Bound to the ACE2 Receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huffman, J.A.; Alex Huffman, J.; Perring, A.E.; Savage, N.J.; Clot, B.; Crouzy, B.; Tummon, F.; Shoshanim, O.; Damit, B.; Schneider, J.; et al. Real-Time Sensing of Bioaerosols: Review and Current Perspectives. Aerosol Sci. Technol. 2020, 54, 465–495. [Google Scholar] [CrossRef] [Green Version]
- Tobias, H.J.; Schafer, M.P.; Pitesky, M.; Fergenson, D.P.; Horn, J.; Frank, M.; Gard, E.E. Bioaerosol Mass Spectrometry for Rapid Detection of Individual Airborne Mycobacterium Tuberculosis H37Ra Particles. Appl. Environ. Microbiol. 2005, 71, 6086–6095. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fergenson, D.P.; Pitesky, M.E.; Tobias, H.J.; Steele, P.T.; Czerwieniec, G.A.; Russell, S.C.; Lebrilla, C.B.; Horn, J.M.; Coffee, K.R.; Srivastava, A.; et al. Reagentless Detection and Classification of Individual Bioaerosol Particles in Seconds. Anal. Chem. 2004, 76, 373–378. [Google Scholar] [CrossRef]
- Phares, D.J.; Rhoads, K.P.; Wexler, A.S.; Kane, D.B.; Johnston, M.V. Application of the ART-2a Algorithm to Laser Ablation Aerosol Mass Spectrometry of Particle Standards. Anal. Chem. 2001, 73, 2338–2344. [Google Scholar] [CrossRef]
- Song, X.-H.; Hopke, P.K.; Fergenson, D.P.; Prather, K.A. Classification of Single Particles Analyzed by ATOFMS Using an Artificial Neural Network, ART-2A. Anal. Chem. 1999, 71, 860–865. [Google Scholar] [CrossRef]
- Stein, S.E.; Scott, D.R. Optimization and Testing of Mass Spectral Library Search Algorithms for Compound Identification. J. Am. Soc. Mass Spectrom. 1994, 5, 859–866. [Google Scholar] [CrossRef] [Green Version]
- Werther, W.; Lohninger, H.; Stancl, F.; Varmuza, K. Classification of Mass Spectra: A Comparison of Yes/No Classification Methods for the Recognition of Simple Structural Properties. Chemom. Intellig. Lab. Syst. 1994, 22, 63–76. [Google Scholar] [CrossRef]
- Varmuza, K.; Werther, W. Mass Spectral Classifiers for Supporting Systematic Structure Elucidation. J. Chem. Inf. Comput. Sci. 1996, 36, 323–333. [Google Scholar] [CrossRef]
- Kerber, A.; Laue, R.; Meringer, M.; Rücker, C.; Schymanski, E. Mathematical Chemistry and Chemoinformatics: Structure Generation, Elucidation and Quantitative Structure-Property Relationships; Walter de Gruyter: Berlin, Germany, 2013; ISBN 9783110254075. [Google Scholar]
- Yan, L.; Yi, J.; Huang, C.; Zhang, J.; Fu, S.; Li, Z.; Lyu, Q.; Xu, Y.; Wang, K.; Yang, H.; et al. Rapid Detection of COVID-19 Using MALDI-TOF-Based Serum Peptidome Profiling. Anal. Chem. 2021, 93, 4782–4787. [Google Scholar] [CrossRef]
- Delafiori, J.; Navarro, L.C.; Siciliano, R.F.; de Melo, G.C.; Busanello, E.N.B.; Nicolau, J.C.; Sales, G.M.; de Oliveira, A.N.; Val, F.F.A.; de Oliveira, D.N.; et al. Covid-19 Automated Diagnosis and Risk Assessment through Metabolomics and Machine Learning. Anal. Chem. 2021, 93, 2471–2479. [Google Scholar] [CrossRef]
- Kampf, G.; Todt, D.; Pfaender, S.; Steinmann, E. Persistence of Coronaviruses on Inanimate Surfaces and Their Inactivation with Biocidal Agents. J. Hosp. Infect. 2020, 104, 246–251. [Google Scholar] [CrossRef] [Green Version]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 Novel Coronavirus (2019-nCoV) by Real-Time RT-PCR. Euro Surveill. 2020, 25. [Google Scholar] [CrossRef] [Green Version]
- Yamagishi, T.; Ohnishi, M.; Matsunaga, N.; Kakimoto, K.; Kamiya, H.; Okamoto, K.; Suzuki, M.; Gu, Y.; Sakaguchi, M.; Tajima, T.; et al. Environmental Sampling for Severe Acute Respiratory Syndrome Coronavirus 2 during a COVID-19 Outbreak on the Diamond Princess Cruise Ship. J. Infect. Dis. 2020, 222, 1098–1102. [Google Scholar] [CrossRef] [PubMed]
- Shang, J.; Wan, Y.; Luo, C.; Ye, G.; Geng, Q.; Auerbach, A.; Li, F. Cell Entry Mechanisms of SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2020, 117, 11727–11734. [Google Scholar] [CrossRef] [PubMed]
- Walker, C.M.; Ko, G. Effect of Ultraviolet Germicidal Irradiation on Viral Aerosols. Environ. Sci. Technol. 2007, 41, 5460–5465. [Google Scholar] [CrossRef]
- Duan, S.-M.; Zhao, X.-S.; Wen, R.-F.; Huang, J.-J.; Pi, G.-H.; Zhang, S.-X.; Han, J.; Bi, S.-L.; Ruan, L.; Dong, X.-P.; et al. Stability of SARS Coronavirus in Human Specimens and Environment and Its Sensitivity to Heating and UV Irradiation. Biomed. Environ. Sci. 2003, 16, 246–255. [Google Scholar] [PubMed]
- Casanova, L.M.; Jeon, S.; Rutala, W.A.; Weber, D.J.; Sobsey, M.D. Effects of Air Temperature and Relative Humidity on Coronavirus Survival on Surfaces. Appl. Environ. Microbiol. 2010, 76, 2712–2717. [Google Scholar] [CrossRef] [Green Version]
- Garcia de Abajo, F.J.; Hernández, R.J.; Kaminer, I.; Meyerhans, A.; Rosell-Llompart, J.; Sanchez-Elsner, T. Back to Normal: An Old Physics Route to Reduce SARS-CoV-2 Transmission in Indoor Spaces. ACS Nano 2020, 14, 7704–7713. [Google Scholar] [CrossRef]
- Abad, F.X.; Pintó, R.M.; Bosch, A. Survival of Enteric Viruses on Environmental Fomites. Appl. Environ. Microbiol. 1994, 60, 3704–3710. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Casadevall, A. The Pathogenic Potential of a Microbe. mSphere 2017, 2. [Google Scholar] [CrossRef] [Green Version]
- Overmyer, K.A.; Shishkova, E.; Miller, I.J.; Balnis, J.; Bernstein, M.N.; Peters-Clarke, T.M.; Meyer, J.G.; Quan, Q.; Muehlbauer, L.K.; Trujillo, E.A.; et al. Large-Scale Multi-Omic Analysis of COVID-19 Severity. Cell Syst. 2021, 12, 23–40.e7. [Google Scholar] [CrossRef] [PubMed]
- Mahmud, I.; Garrett, T.J. Mass Spectrometry Techniques in Emerging Pathogens Studies: COVID-19 Perspectives. J. Am. Soc. Mass Spectrom. 2020, 31, 2013–2024. [Google Scholar] [CrossRef] [PubMed]
- Chia, J.W.F.; Sawai, O.; Nunoura, T. Reaction Pathway of Poly(ethylene) Terephthalate Carbonization: Decomposition Behavior Based on Carbonized Product. Waste Manag. 2020, 108, 62–69. [Google Scholar] [CrossRef]
- Medeiros, P.M.; Seidel, M.; Powers, L.C.; Dittmar, T.; Hansell, D.A.; Miller, W.L. Dissolved Organic Matter Composition and Photochemical Transformations in the Northern North Pacific Ocean. Geophys. Res. Lett. 2015, 42, 863–870. [Google Scholar] [CrossRef]
- Heinrichs, M.E.; Tebbe, D.A.; Wemheuer, B.; Niggemann, J.; Engelen, B. Impact of Viral Lysis on the Composition of Bacterial Communities and Dissolved Organic Matter in Deep-Sea Sediments. Viruses 2020, 12, 922. [Google Scholar] [CrossRef]
- Miller, R.L.; Plagemann, P.G. Effect of Ultraviolet Light on Mengovirus: Formation of Uracil Dimers, Instability and Degradation of Capsid, and Covalent Linkage of Protein to Viral RNA. J. Virol. 1974, 13, 729–739. [Google Scholar] [CrossRef] [Green Version]
- Lytle, C.D.; Sagripanti, J.-L. Predicted Inactivation of Viruses of Relevance to Biodefense by Solar Radiation. J. Virol. 2005, 79, 14244–14252. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ong, S.W.X.; Tan, Y.K.; Coleman, K.K.; Tan, B.H.; Leo, Y.-S.; Wang, D.L.; Ng, C.G.; Ng, O.-T.; Wong, M.S.Y.; Marimuthu, K. Lack of Viable Severe Acute Respiratory Coronavirus Virus 2 (SARS-CoV-2) among PCR-Positive Air Samples from Hospital Rooms and Community Isolation Facilities; Cambridge University Press: Cambridge, UK, 2021. [Google Scholar]
- Rodríguez, R.A.; Pepper, I.L.; Gerba, C.P. Application of PCR-Based Methods to Assess the Infectivity of Enteric Viruses in Environmental Samples. Appl. Environ. Microbiol. 2009, 75, 297–307. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aubrey, A.D.; Cleaves, H.J.; Bada, J.L. The Role of Submarine Hydrothermal Systems in the Synthesis of Amino Acids. Orig. Life Evol. Biosph. 2009, 39, 91–108. [Google Scholar] [CrossRef] [PubMed]
- Chandru, K.; Imai, E.; Kaneko, T.; Obayashi, Y.; Kobayashi, K. Survivability and Abiotic Reactions of Selected Amino Acids in Different Hydrothermal System Simulators. Orig. Life Evol. Biosph. 2013, 43, 99–108. [Google Scholar] [CrossRef] [PubMed]
- Saladino, R.; Neri, V.; Crestini, C.; Costanzo, G.; Graciotti, M.; Di Mauro, E. Synthesis and Degradation of Nucleic Acid Components by Formamide and Iron Sulfur Minerals. J. Am. Chem. Soc. 2008, 130, 15512–15518. [Google Scholar] [CrossRef]
- Johnson, P.V.; Hodyss, R.; Tang, K.; Brinckerhoff, W.B.; Smith, R.D. The Laser Ablation Ion Funnel: Sampling for in Situ Mass Spectrometry on Mars. Planet. Space Sci. 2011, 59, 387–393. [Google Scholar] [CrossRef]
- Goesmann, F.; Rosenbauer, H.; Bredehöft, J.H.; Cabane, M.; Ehrenfreund, P.; Gautier, T.; Giri, C.; Krüger, H.; Le Roy, L.; MacDermott, A.J.; et al. COMETARY SCIENCE. Organic Compounds on Comet 67P/Churyumov-Gerasimenko Revealed by COSAC Mass Spectrometry. Science 2015, 349, aab0689. [Google Scholar] [CrossRef] [PubMed]
- Briois, C.; Thissen, R.; Thirkell, L.; Aradj, K.; Bouabdellah, A.; Boukrara, A.; Carrasco, N.; Chalumeau, G.; Chapelon, O.; Colin, F.; et al. Orbitrap Mass Analyser for in Situ Characterisation of Planetary Environments: Performance Evaluation of a Laboratory Prototype. Planet. Space Sci. 2016, 131, 33–45. [Google Scholar] [CrossRef]
- Goesmann, F.; Brinckerhoff, W.B.; Raulin, F.; Goetz, W.; Danell, R.M.; Getty, S.A.; Siljeström, S.; Mißbach, H.; Steininger, H.; Arevalo, R.D., Jr.; et al. The Mars Organic Molecule Analyzer (MOMA) Instrument: Characterization of Organic Material in Martian Sediments. Astrobiology 2017, 17, 655–685. [Google Scholar] [CrossRef] [PubMed]
- Vazquez, T.; Vuppala, S.; Ayodeji, I.; Song, L.; Grimes, N.; Evans-Nguyen, T. IN SITU MASS SPECTROMETERS FOR APPLICATIONS IN SPACE. Mass Spectrom. Rev. 2020. [Google Scholar] [CrossRef] [PubMed]
- National Academies of Sciences, Engineering, and Medicine; Division on Engineering and Physical Sciences; Space Studies Board; Committee on Astrobiology Science Strategy for the Search for Life in the Universe. An Astrobiology Strategy for the Search for Life in the Universe; National Academies Press: Washington, DC, USA, 2019; ISBN 9780309484169. [Google Scholar]
- Mahaffy, P.R.; Webster, C.R.; Cabane, M.; Conrad, P.G.; Coll, P.; Atreya, S.K.; Arvey, R.; Barciniak, M.; Benna, M.; Bleacher, L.; et al. The Sample Analysis at Mars Investigation and Instrument Suite. Space Sci. Rev. 2012, 170, 401–478. [Google Scholar] [CrossRef]
- Meringer, M.; Giri, C.; Cleaves, H.J. Fitting Cometary Sampling and Composition Mass Spectral Results Using Non-Negative Least Squares: Reducing Detection Ambiguity for In Situ Solar System Organic Compound Measurements. ACS Earth Space Chem. 2018, 2, 1256–1261. [Google Scholar] [CrossRef]
- Waite, J.H., Jr.; Niemann, H.; Yelle, R.V.; Kasprzak, W.T.; Cravens, T.E.; Luhmann, J.G.; McNutt, R.L.; Ip, W.-H.; Gell, D.; De La Haye, V.; et al. Ion Neutral Mass Spectrometer Results from the First Flyby of Titan. Science 2005, 308, 982–986. [Google Scholar] [CrossRef] [Green Version]
- Ali, A.; Sittler, E.C., Jr.; Chornay, D.; Rowe, B.R.; Puzzarini, C. Organic Chemistry in Titan’ S Upper Atmosphere and Its Astrobiological Consequences: I. Views towards Cassini Plasma Spectrometer (CAPS) and Ion Neutral Mass Spectrometer (INMS) Experiments in Space. Planet. Space Sci. 2015, 109, 46–63. [Google Scholar] [CrossRef]
- Janjic, A. The Need for Including Virus Detection Methods in Future Mars Missions. Astrobiology 2018, 18, 1611–1614. [Google Scholar] [CrossRef]
- Demirev, P.A.; Fenselau, C. Mass Spectrometry in Biodefense. J. Mass Spectrom. 2008, 43, 1441–1457. [Google Scholar] [CrossRef] [PubMed]
- Trapp, R. Living with Chemicals: How to Prevent Their Use for Hostile Purposes and Mitigate Chemical Risks. In Cyber and Chemical, Biological, Radiological, Nuclear, Explosives Challenges: Threats and Counter Efforts; Martellini, M., Malizia, A., Eds.; Springer International Publishing: Cham, Switzerland, 2017; pp. 357–383. ISBN 9783319621081. [Google Scholar]
- Zhang, X.; Zhang, H.; Yu, K.; Li, N.; Liu, Y.; Liu, X.; Zhang, H.; Yang, B.; Wu, W.; Gao, J.; et al. Rapid Monitoring Approach for Microplastics Using Portable Pyrolysis-Mass Spectrometry. Anal. Chem. 2020, 92, 4656–4662. [Google Scholar] [CrossRef]
- Fatigante, W.L.; Mukta, S.; Lawton, Z.E.; Bruno, A.M.; Traub, A.; Gasa, A.J.; Stelmack, A.R.; Wilson-Frank, C.R.; Mulligan, C.C. Filter Cone Spray Ionization Coupled to a Portable MS System: Application to On-Site Forensic Evidence and Environmental Sample Analysis. J. Am. Soc. Mass Spectrom. 2020, 31, 336–346. [Google Scholar] [CrossRef] [PubMed]
- La Nasa, J.; Nardella, F.; Modugno, F.; Colombini, M.P.; Ribechini, E.; Degano, I. SIFT-Ing Archaeological Artifacts: Selected Ion Flow Tube-Mass Spectrometry as a New Tool in Archaeometry. Talanta 2020, 207, 120323. [Google Scholar] [CrossRef]
- Tulej, M.; Riedo, A.; Neuland, M.B.; Meyer, S.; Wurz, P.; Thomas, N.; Grimaudo, V.; Moreno-García, P.; Broekmann, P.; Neubeck, A.; et al. CAMAM: A Miniature Laser Ablation Ionisation Mass Spectrometer and Microscope-Camera System forIn SituInvestigation of the Composition and Morphology of Extraterrestrial Materials. Geostand. Geoanal. Res. 2014, 38, 441–466. [Google Scholar] [CrossRef]
- Snyder, D.T.; Pulliam, C.J.; Ouyang, Z.; Cooks, R.G. Miniature and Fieldable Mass Spectrometers: Recent Advances. Anal. Chem. 2016, 88, 2–29. [Google Scholar] [CrossRef] [Green Version]
- Walker, H.J.; Burrell, M.M. Could Breath Analysis by MS Could Be a Solution to Rapid, Non-Invasive Testing for COVID-19? Bioanalysis 2020, 12, 1213–1217. [Google Scholar] [CrossRef] [PubMed]
- Gould, O.; Ratcliffe, N.; Król, E.; de Lacy Costello, B. Breath Analysis for Detection of Viral Infection, the Current Position of the Field. J. Breath Res. 2020, 14, 041001. [Google Scholar] [CrossRef] [PubMed]
- Ruszkiewicz, D.M.; Sanders, D.; O’Brien, R.; Hempel, F.; Reed, M.J.; Riepe, A.C.; Bailie, K.; Brodrick, E.; Darnley, K.; Ellerkmann, R.; et al. Diagnosis of COVID-19 by Analysis of Breath with Gas Chromatography-Ion Mobility Spectrometry—A Feasibility Study. EClinicalMedicine 2020, 29, 100609. [Google Scholar] [CrossRef]
- Cardozo, K.H.M.; Lebkuchen, A.; Okai, G.G.; Schuch, R.A.; Viana, L.G.; Olive, A.N.; Lazari, C.D.S.; Fraga, A.M.; Granato, C.F.H.; Pintão, M.C.T.; et al. Establishing a Mass Spectrometry-Based System for Rapid Detection of SARS-CoV-2 in Large Clinical Sample Cohorts. Nat. Commun. 2020, 11, 6201. [Google Scholar] [CrossRef] [PubMed]

| Portable Mass Spectrometer for Astrobiology Applications | Portable Mass Spectrometer for Contagion Surveillance Applications |
|---|---|
| High resolution and mass/charge range for intact biosignature detection (10–1000 daltons) (e.g., amino acids, nucleotides, nucleobases etc.) | High resolution and mass/charge range for intact microbial signature detection (100 megadaltons—1 giga-daltons) (e.g., lipid, RNA and proteins) |
| Low payload mass for space payload needs | Low payload mass for hand-held devices aerial drone and autonomous ground vehicles |
| Low energy consumption with provisions for energy storage backups | Low energy consumption with provisions for intermittent charging and plug-in options |
| Miniaturization of 2–4 GHz magnets (ion/electron cyclotron) to match low payload requirements | Miniaturization of 2–4 GHz magnets (ion/electron cyclotron) to match with ergonomics and portability requirements |
| Use of nano-mechanical resonators/aerodynamic lenses for providing greater m/z resolution and higher signal-to-noise ratio | Use of nano-mechanical resonators/aerodynamic lenses for greater resolution along with sample nebulization |
| Ability to detect biosignature molecules in atmospheric aerosols, geological rocks, regolith, brine and clays, and liquid samples | Ability to detect infectious microbial signatures and estimate contagion infectivity in bioaerosols and fomites in closed or public spaces |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Giri, C.; Cleaves, H.J., II; Meringer, M.; Chandru, K. The Post-COVID-19 Era: Interdisciplinary Demands of Contagion Surveillance Mass Spectrometry for Future Pandemics. Sustainability 2021, 13, 7614. https://doi.org/10.3390/su13147614
Giri C, Cleaves HJ II, Meringer M, Chandru K. The Post-COVID-19 Era: Interdisciplinary Demands of Contagion Surveillance Mass Spectrometry for Future Pandemics. Sustainability. 2021; 13(14):7614. https://doi.org/10.3390/su13147614
Chicago/Turabian StyleGiri, Chaitanya, Henderson James Cleaves, II, Markus Meringer, and Kuhan Chandru. 2021. "The Post-COVID-19 Era: Interdisciplinary Demands of Contagion Surveillance Mass Spectrometry for Future Pandemics" Sustainability 13, no. 14: 7614. https://doi.org/10.3390/su13147614
APA StyleGiri, C., Cleaves, H. J., II, Meringer, M., & Chandru, K. (2021). The Post-COVID-19 Era: Interdisciplinary Demands of Contagion Surveillance Mass Spectrometry for Future Pandemics. Sustainability, 13(14), 7614. https://doi.org/10.3390/su13147614








