Pb(II) Bio-Removal, Viability, and Population Distribution of an Industrial Microbial Consortium: The Effect of Pb(II) and Nutrient Concentrations
Abstract
1. Introduction
2. Materials and Methods
2.1. Project Overview
2.2. Materials
2.3. Microbial Culture
2.4. Experimental
2.5. Sampling
2.6. Batch Reactor Analysis
2.7. Microbial Plate Preparation for Characterization
2.8. Microbial Characterization
2.9. SEM Analysis
Sample Preparation and Analysis
3. Results
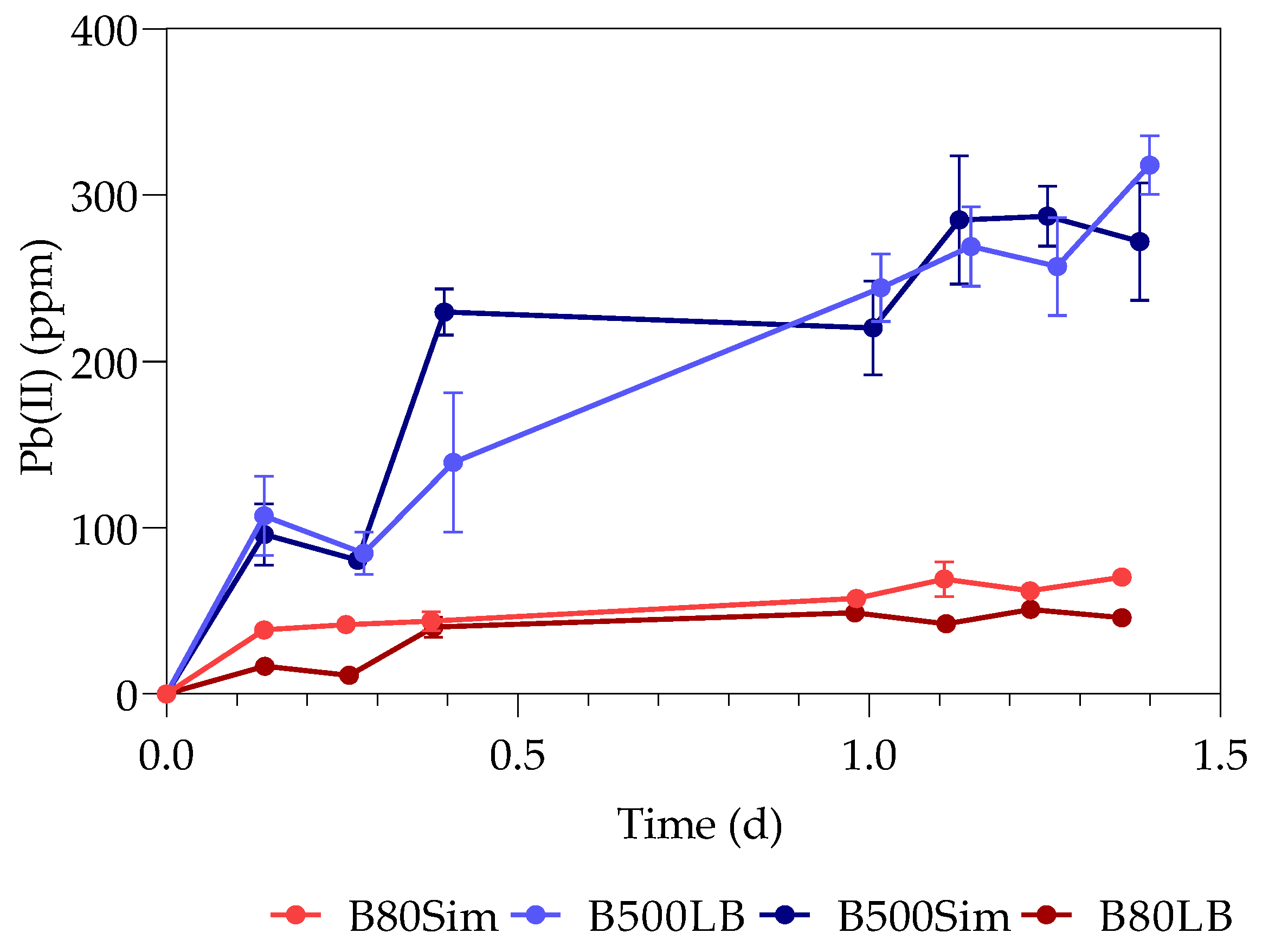
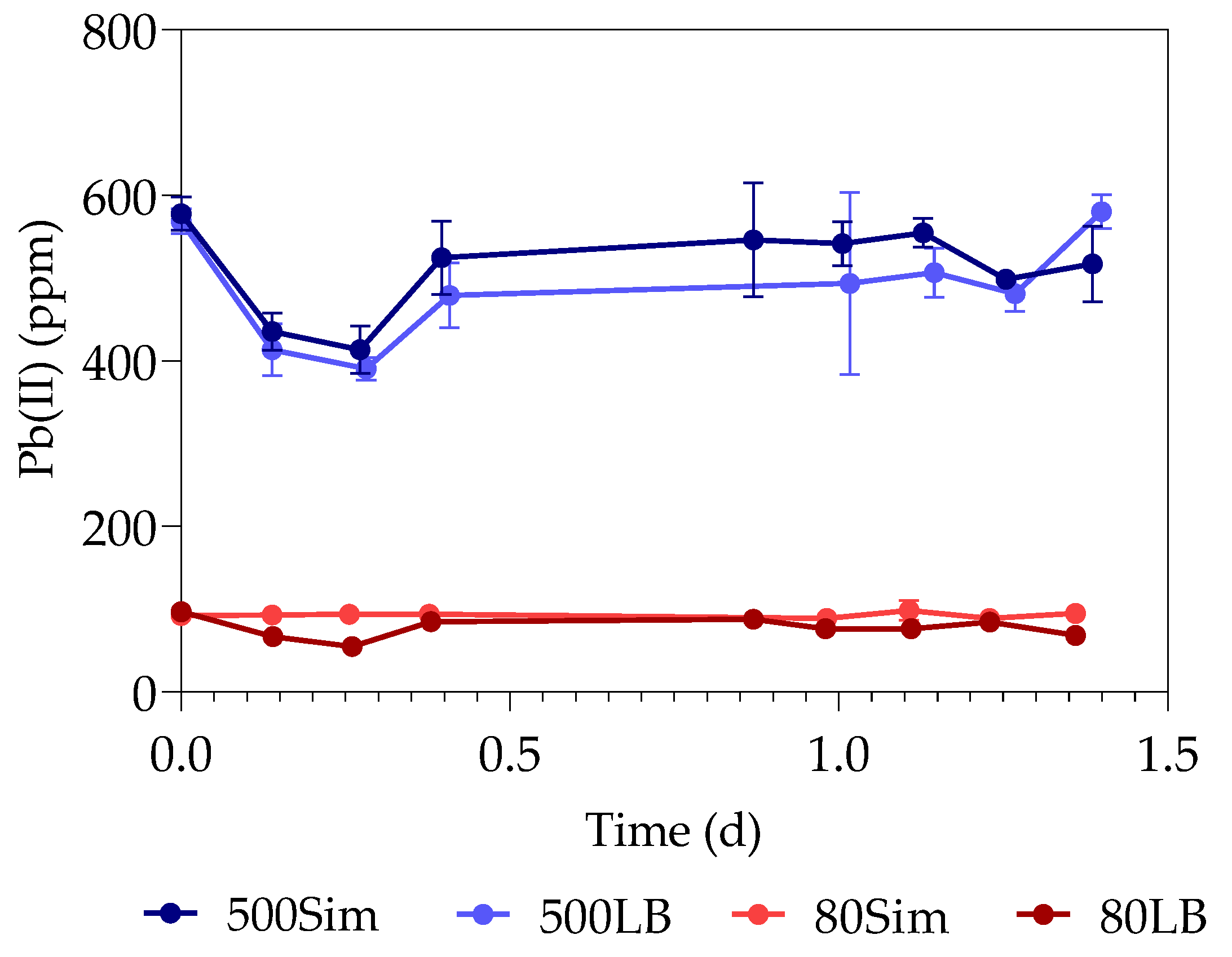
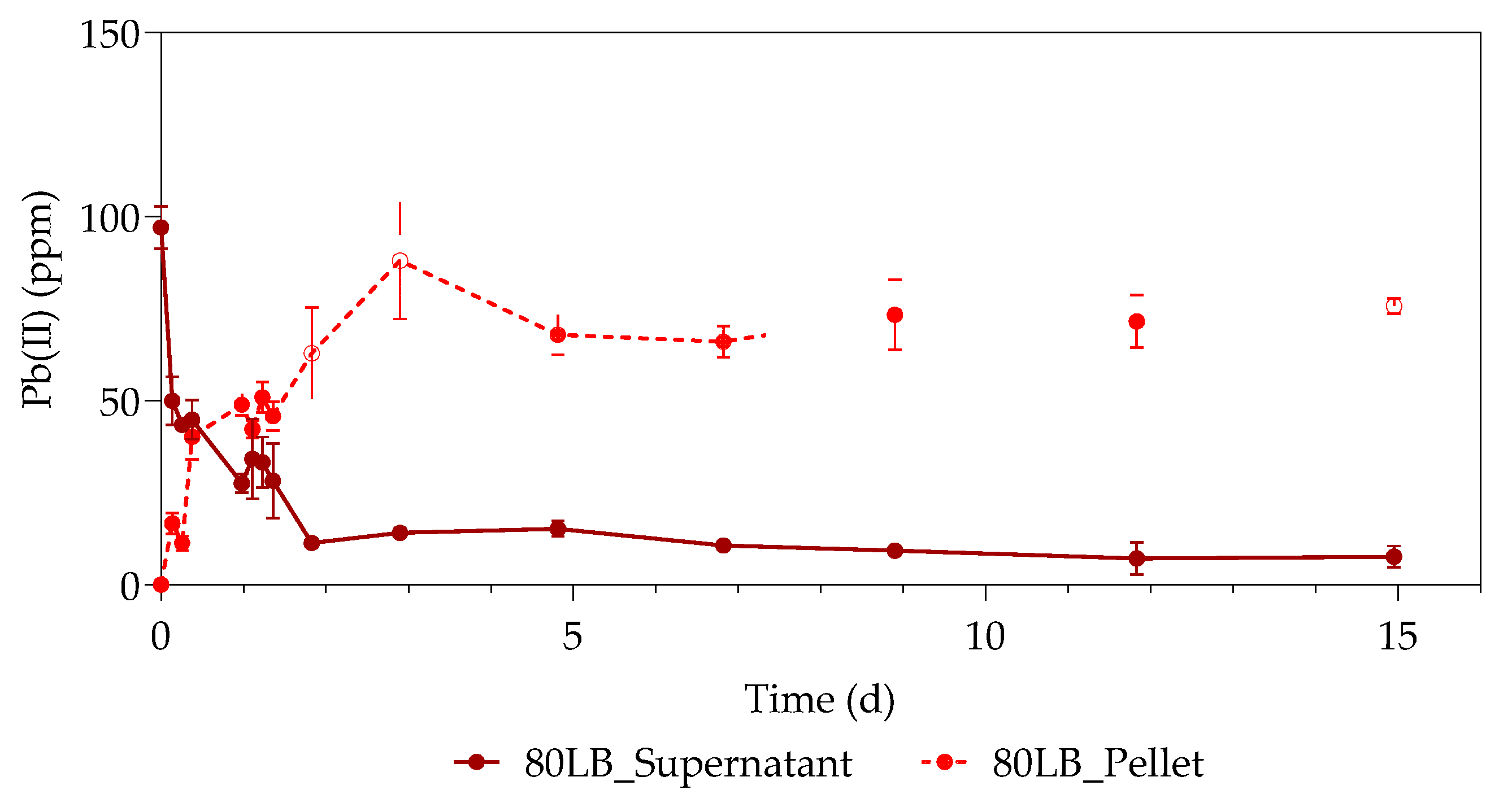
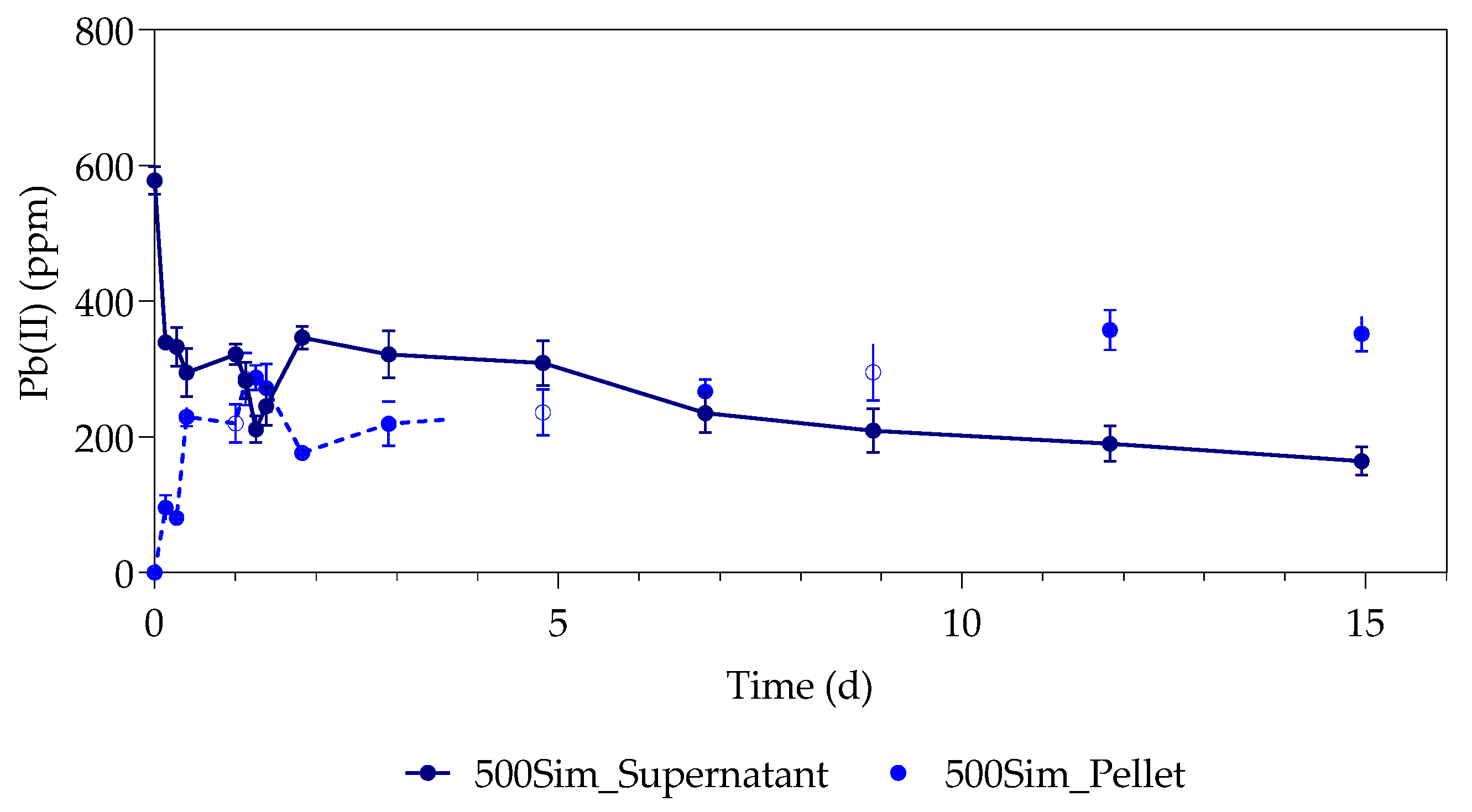
3.1. Pb(II) Removal
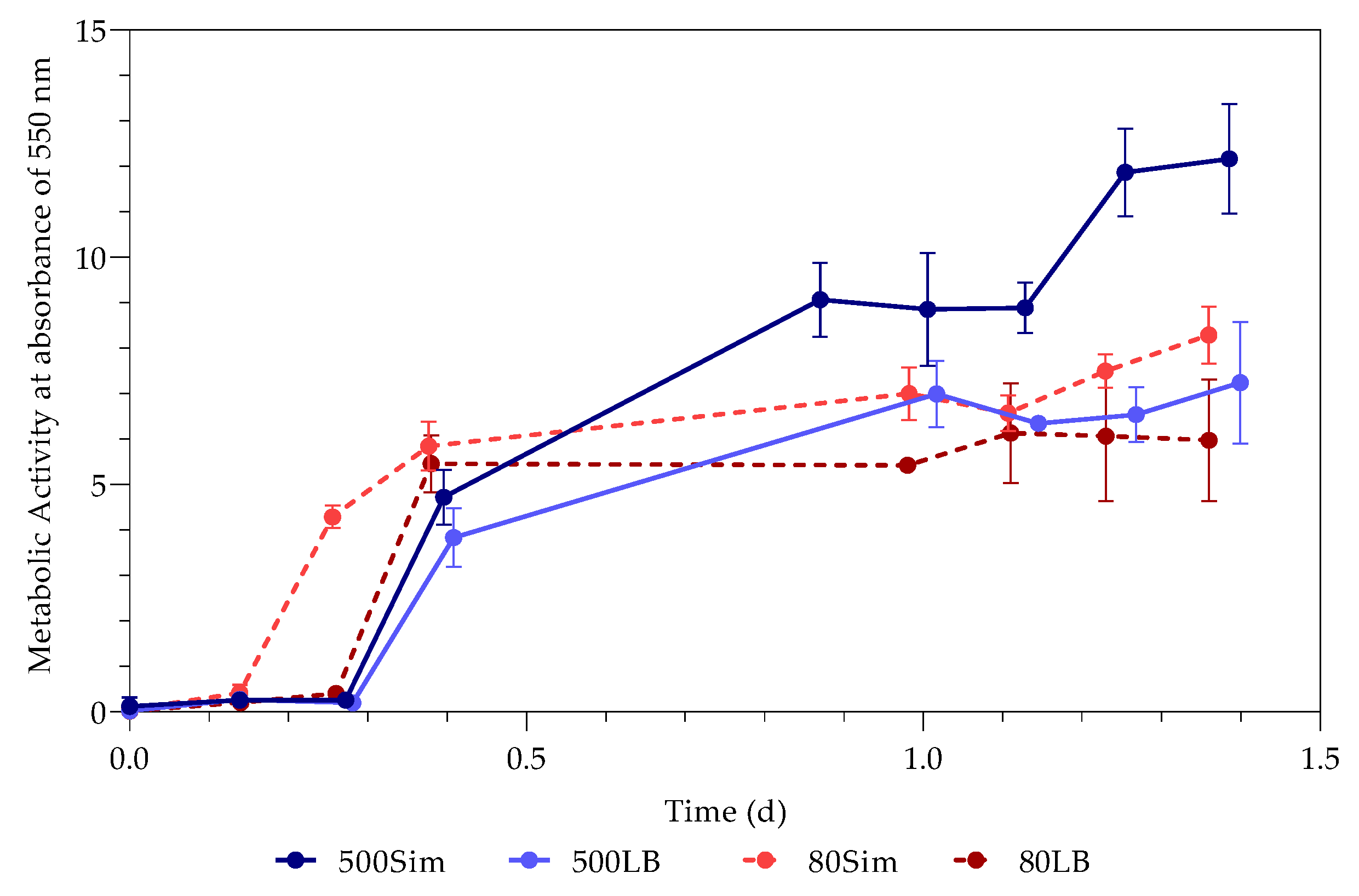
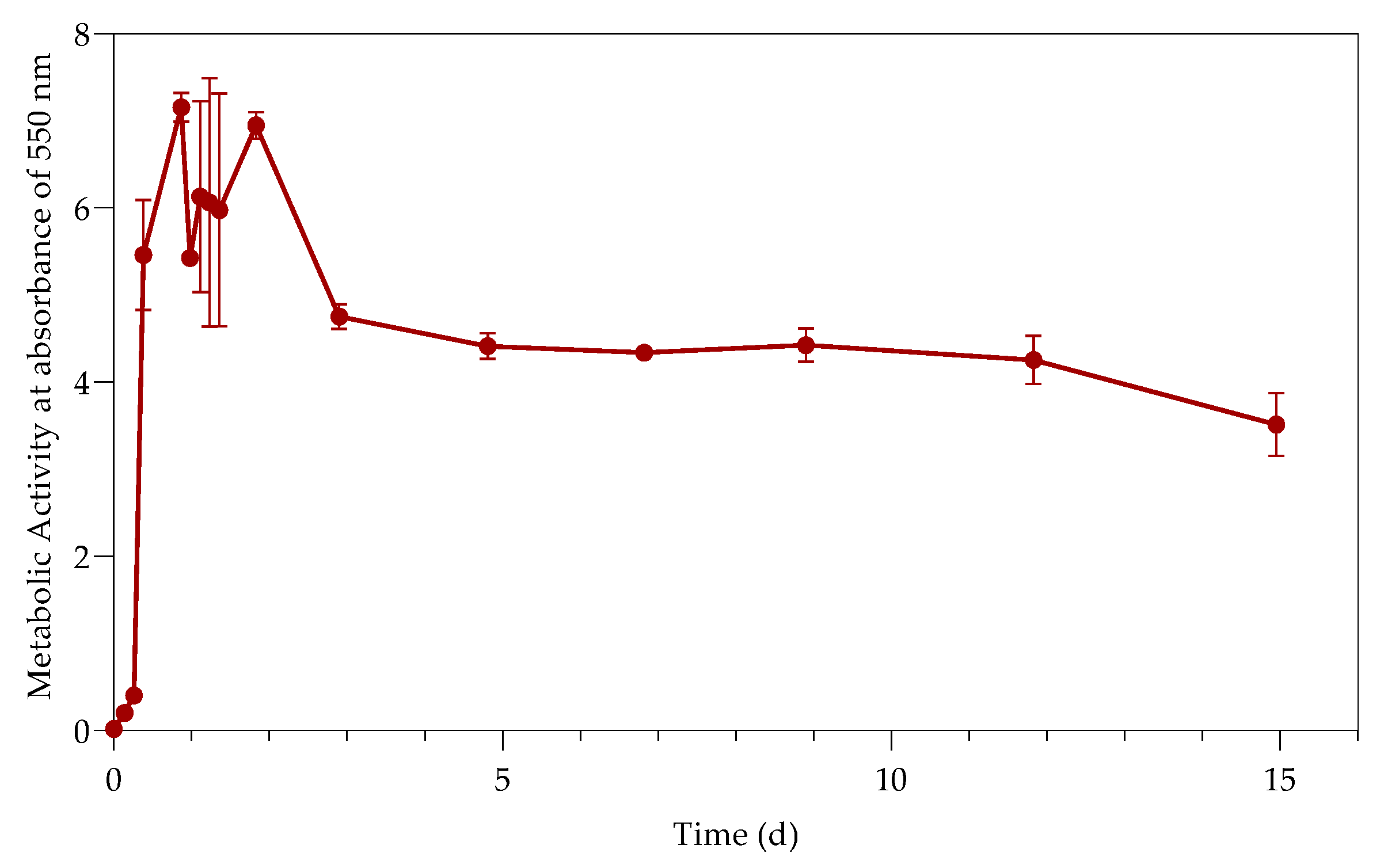
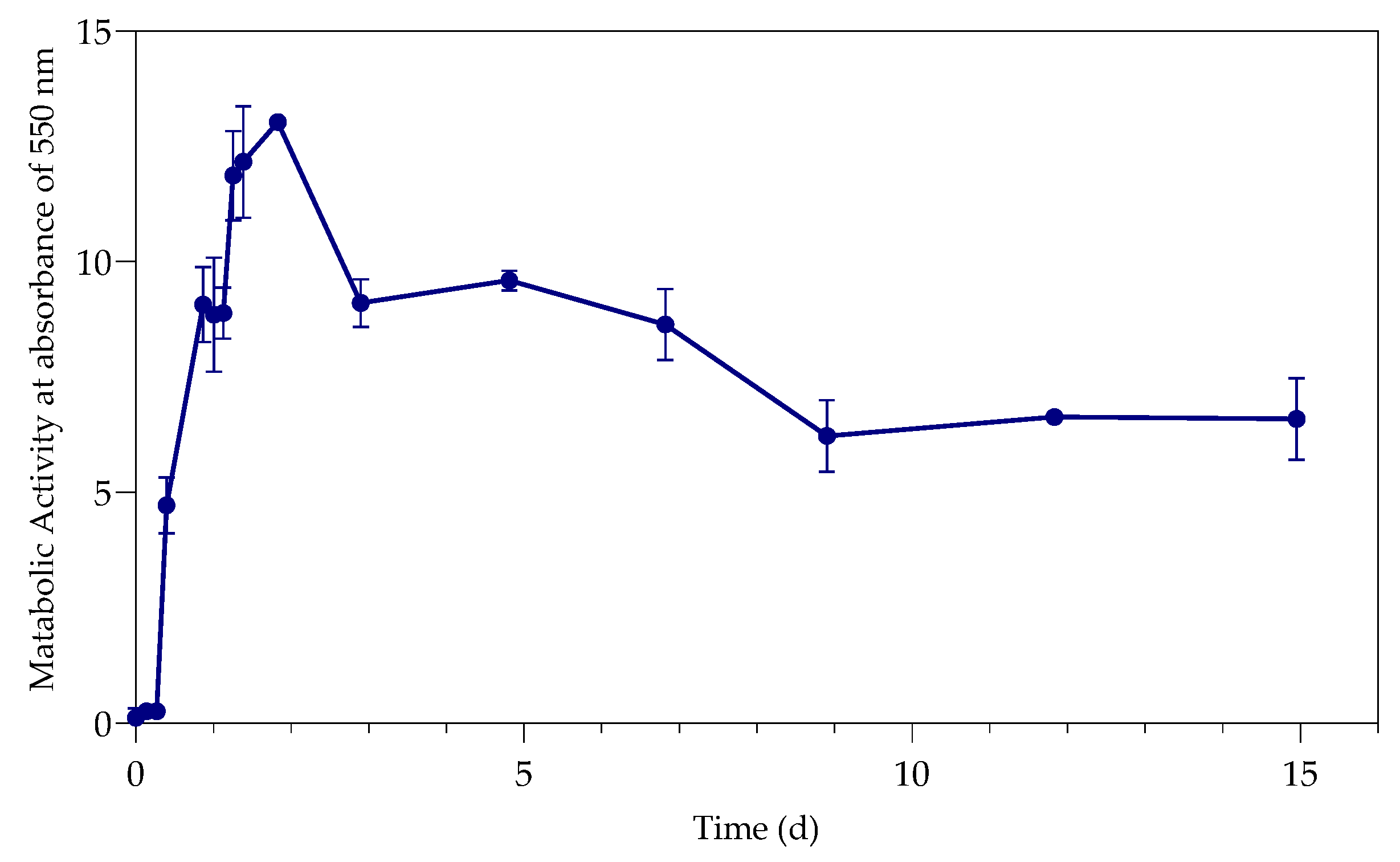
3.2. Metabolic Activity (Growth)
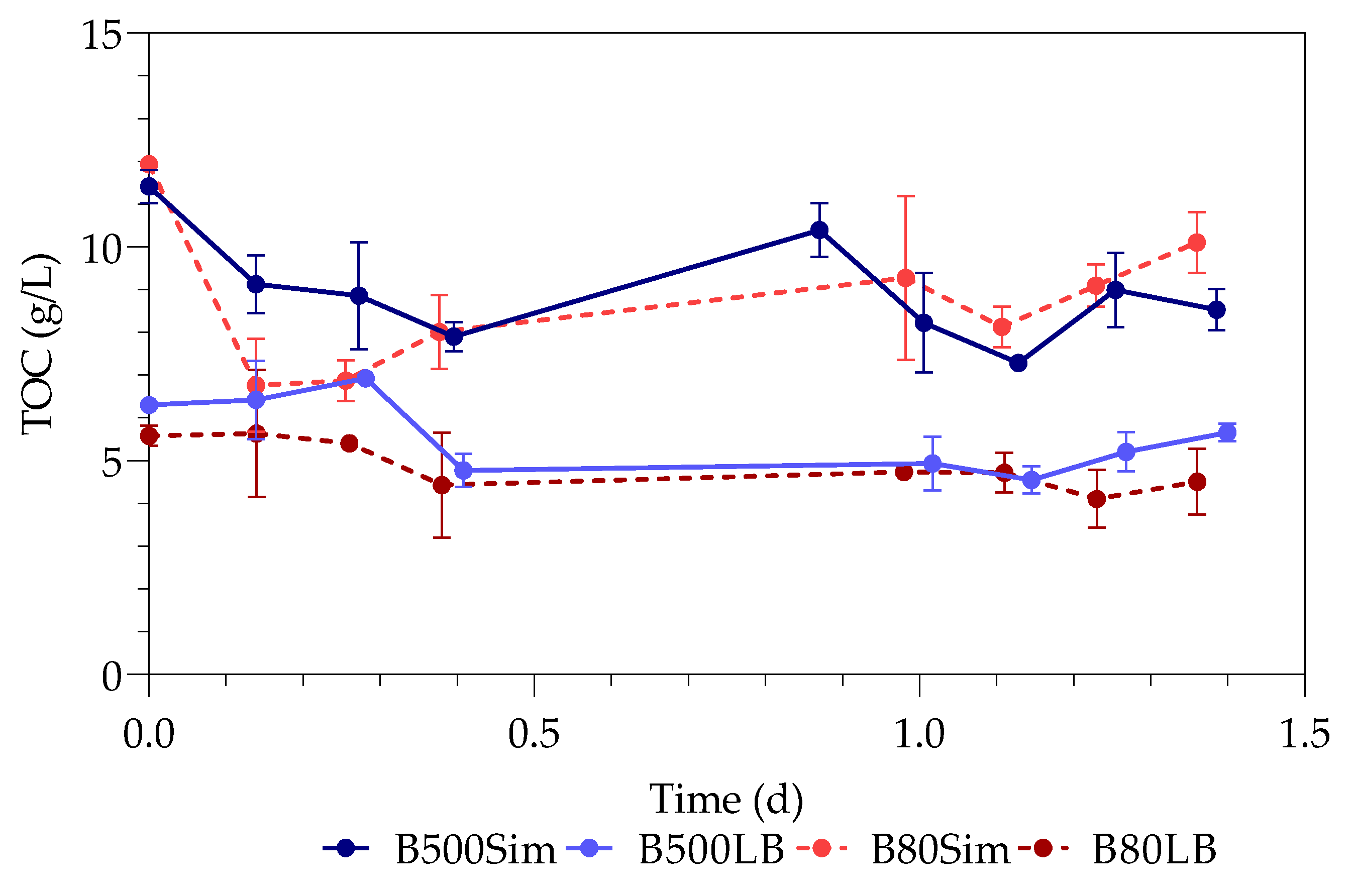
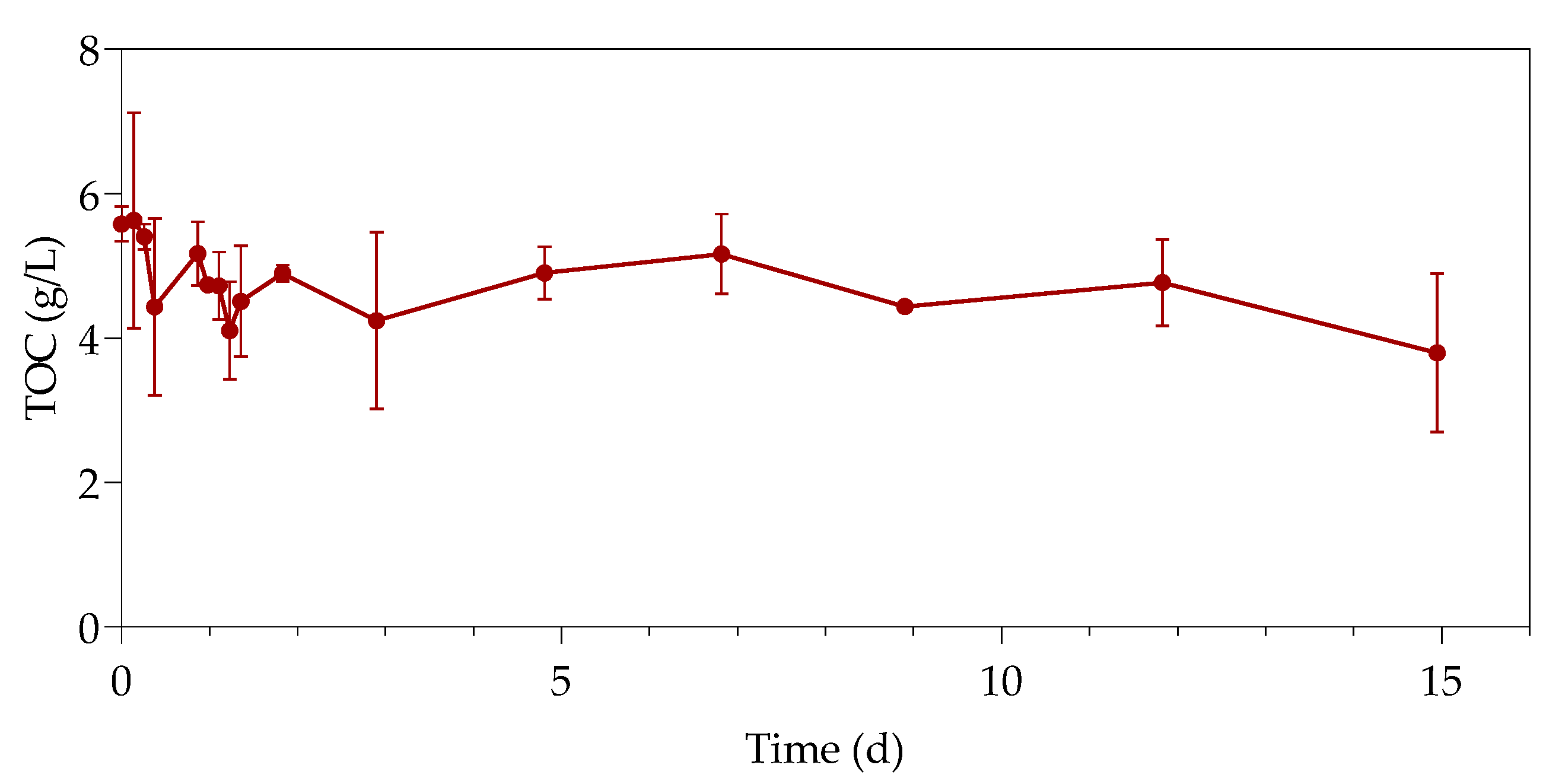
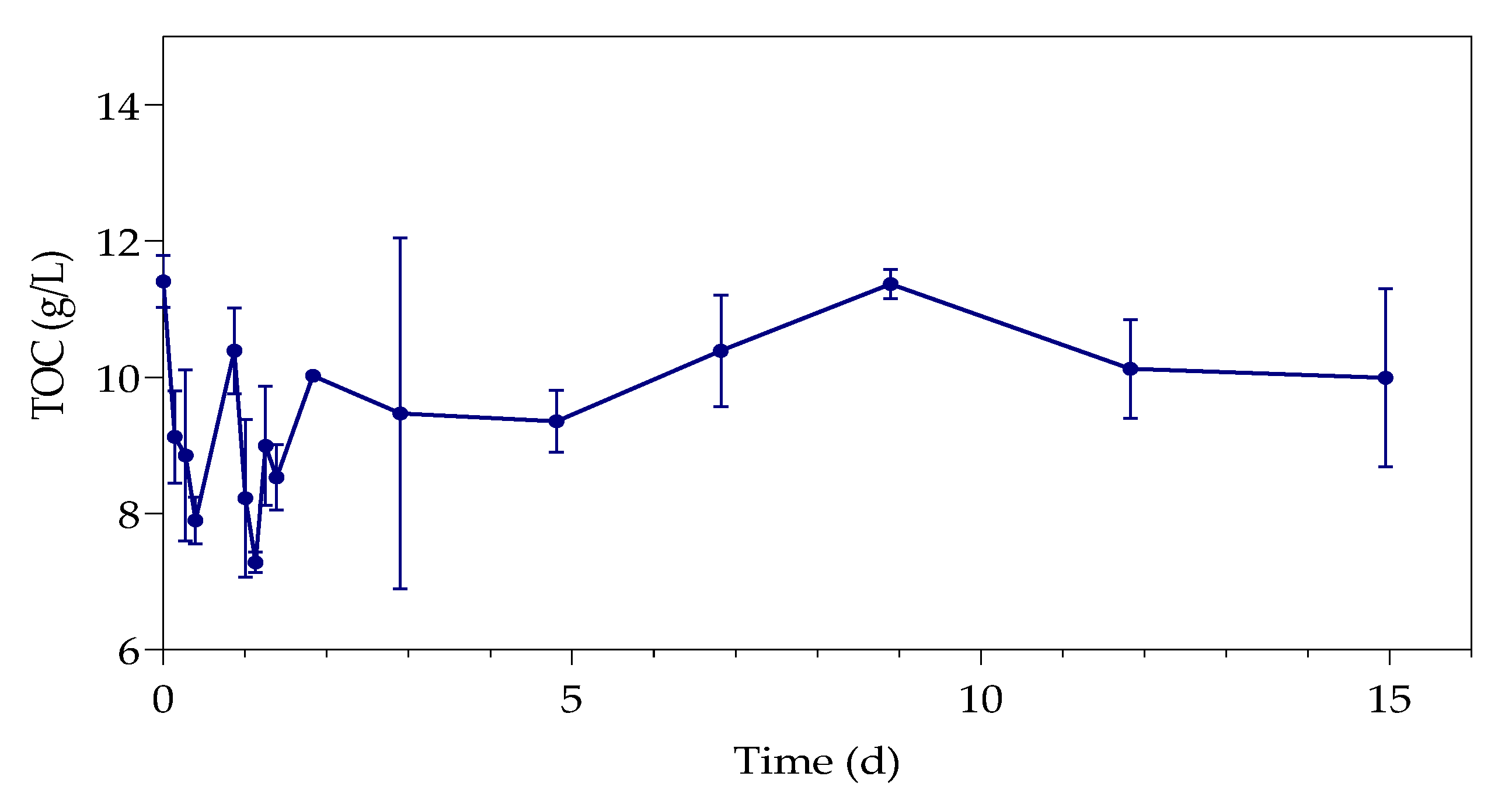
3.3. Total Organic Carbon
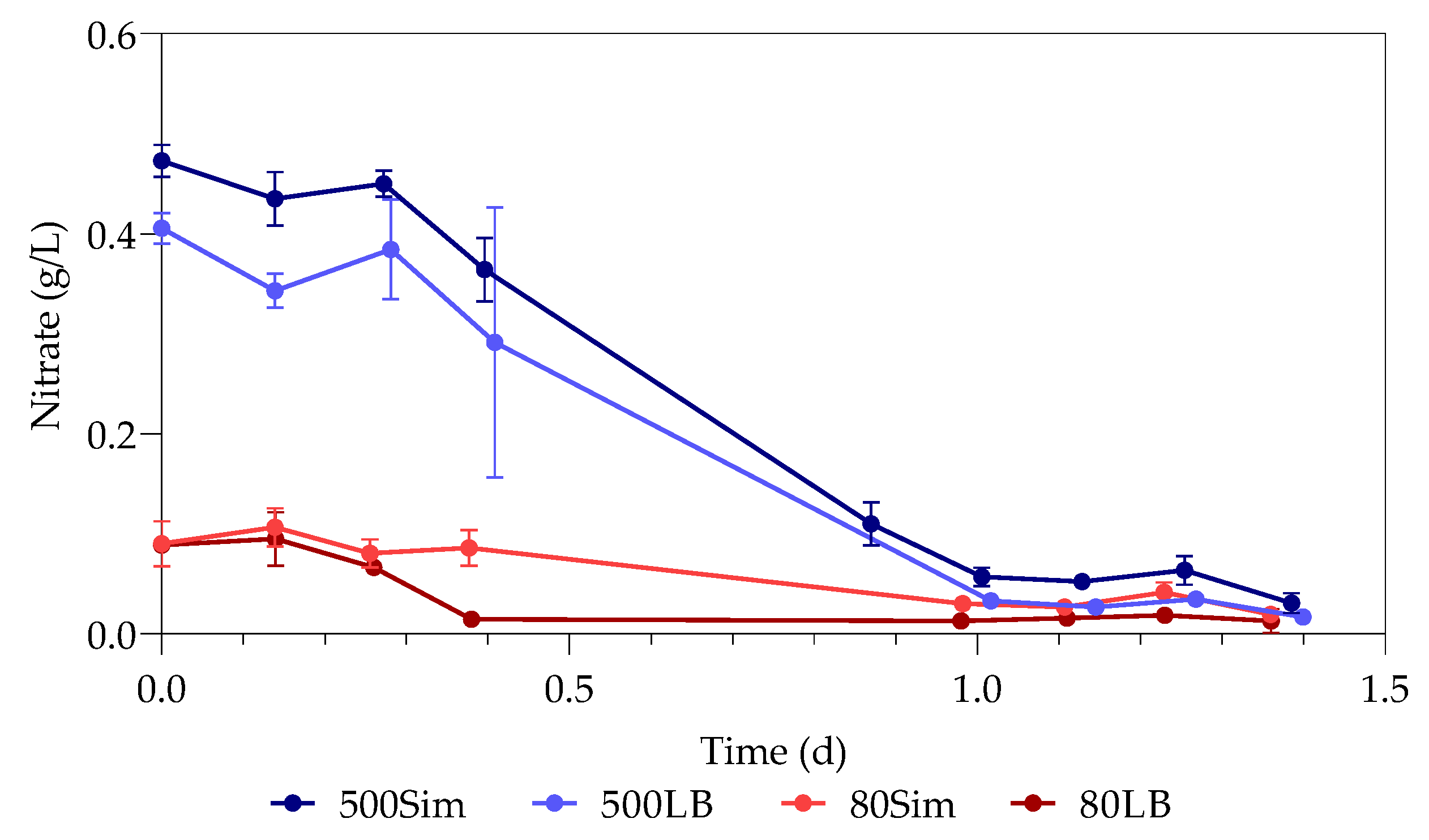
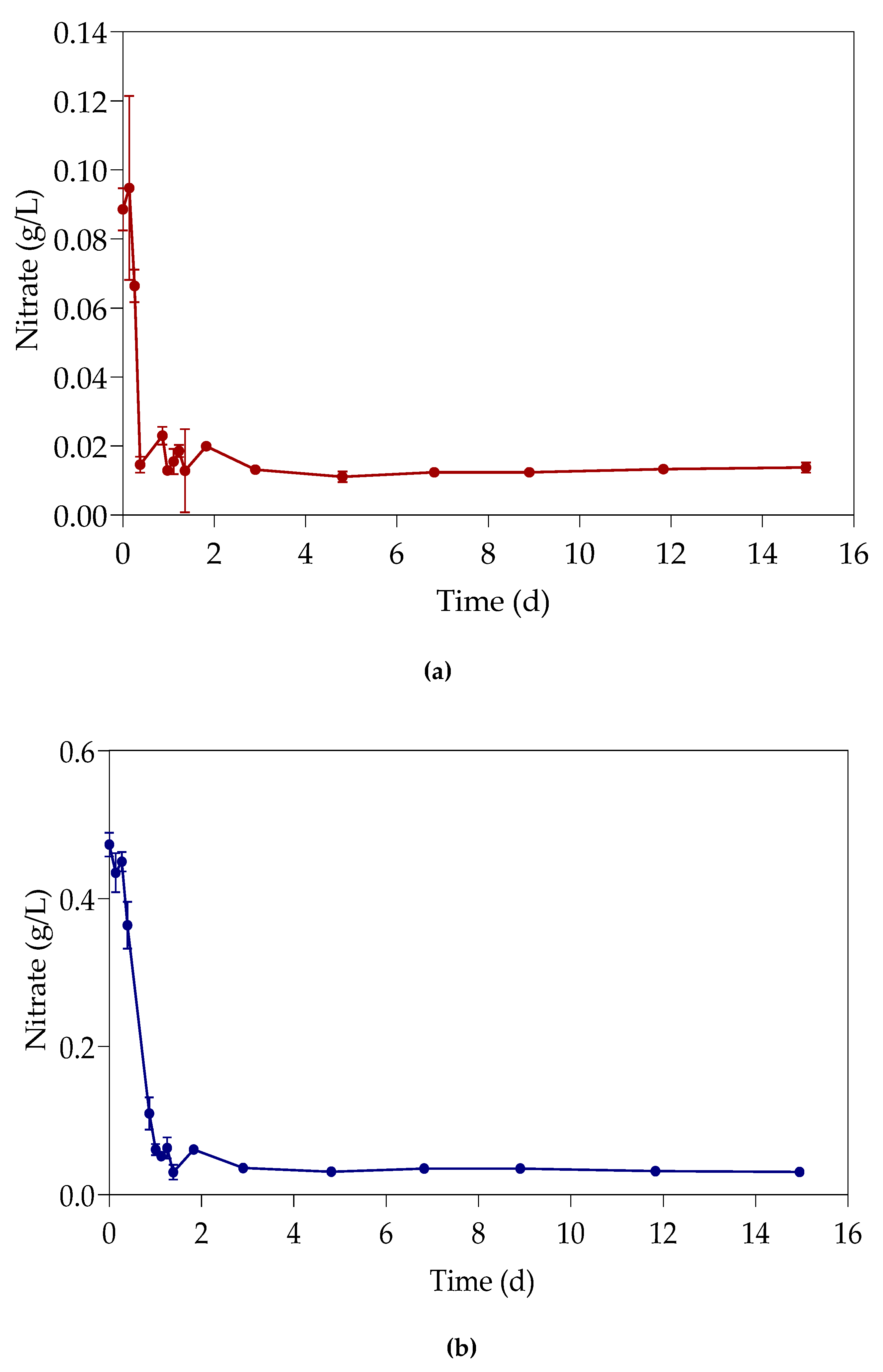
3.4. Nitrates
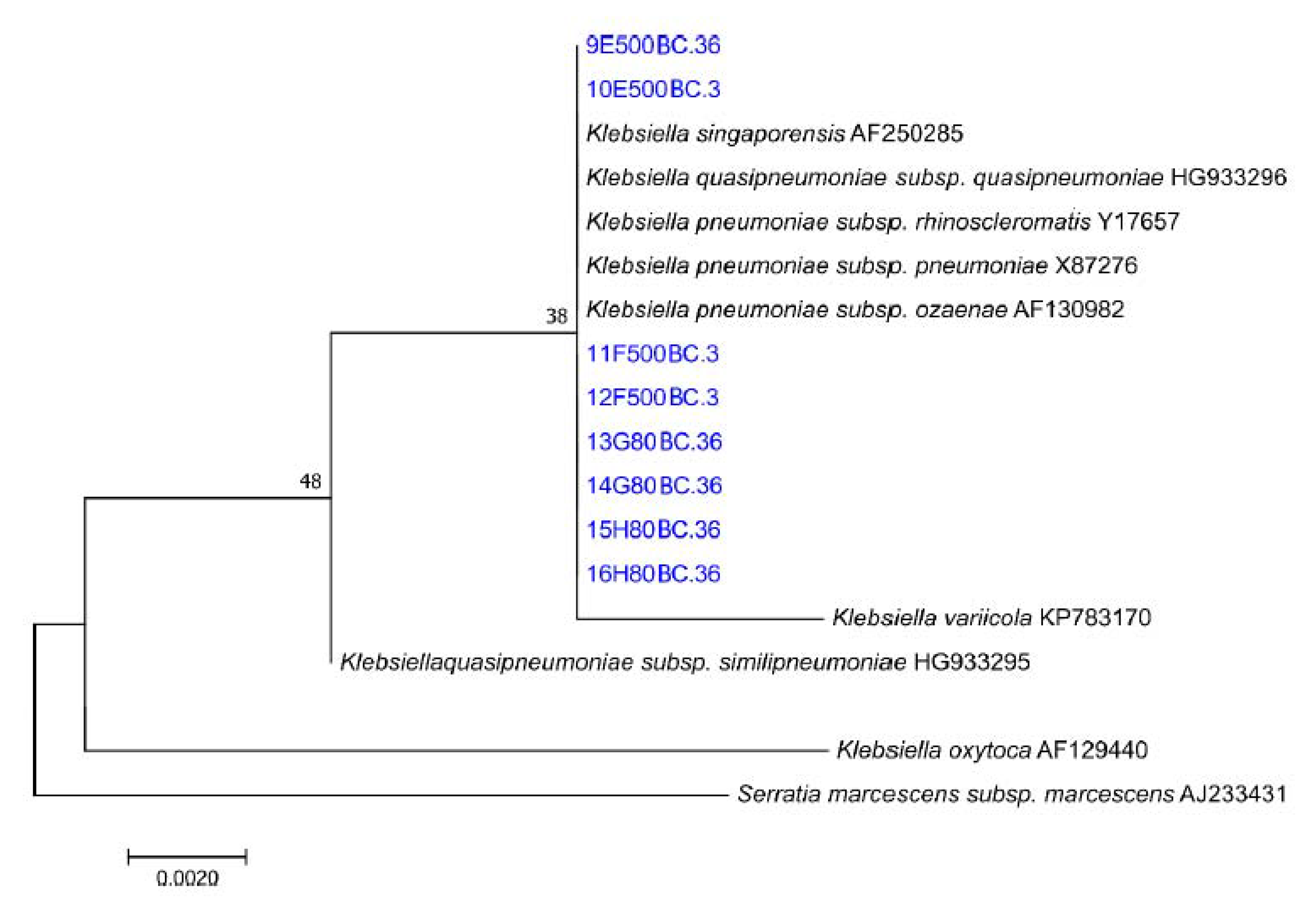
3.5. Microbial Characterisation
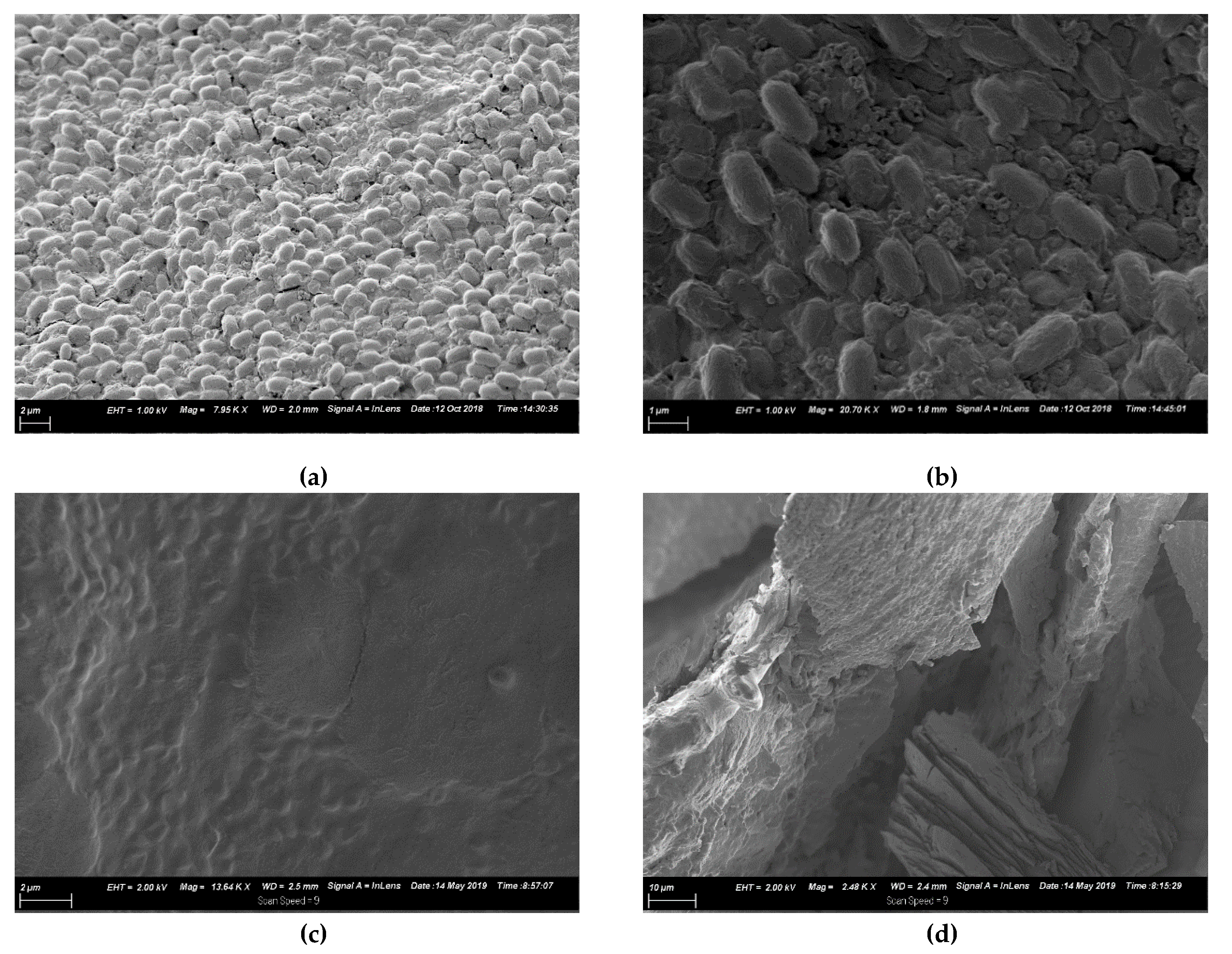
3.6. SEM Morphology
3.7. Specific Growth Rate
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Duruibe, J.O.; Ogwuegbu, M.O.C.; Egwurugwu, J.N. Heavy metal pollution and human biotoxic effects. Int. J. Phys. Sci. 2007, 2, 112–118. [Google Scholar]
- Substances Agency for Toxic and Disease Registry. Toxicological Profile for Lead. Available online: https://www.atsdr.cdc.gov/toxprofiles/tp.asp?id=96&tid=22 (accessed on 24 July 2019).
- U.S. Geological Survey. Mineral Commodity Summaries 2019. Available online: https://pubs.usgs.gov/periodicals/mcs2020/mcs2020.pdf (accessed on 5 January 2020).
- International Lead Association. Lead Production & Statistics. Available online: https://www.ila-lead.org/lead-facts/lead-production--statistics (accessed on 6 May 2019).
- Pan, X.; Chen, Z.; Li, L.; Rao, W.; Xu, Z.; Guan, X. Microbial strategy for potential lead remediation: A review study. World J. Microbiol. Biotechnol. 2017, 33, 1–7. [Google Scholar] [CrossRef]
- Hee, J.; Bolan, N. Lead immobilization and bioavailability in microbial and root interface. J. Hazard. Mater. 2013, 261, 777–783. [Google Scholar] [CrossRef] [PubMed]
- Aiking, H.; Govers, H.; Van’t Riet, J. Detoxification of mercury, cadmium, and lead in Klebsiella aerogenes NCTC 418 growing in continuous culture. Appl. Environ. Microbiol. 1985, 50, 1262–1267. [Google Scholar] [CrossRef] [PubMed]
- Bowman, N.; Patel, D.; Sanchez, A.; Xu, W.; Alsaffar, A.; Tiquia-arashiro, S.M. Lead-resistant bacteria from Saint Clair River sediments and Pb removal in aqueous solutions. Appl. Microbiol. Biotechnol. 2018, 102, 2391–2398. [Google Scholar] [CrossRef] [PubMed]
- Brink, H.G.; Hörstmann, C.; Peens, J. Microbial Pb(II)-precipitation: The influence of oxygen on Pb(II)-removal from aqueous environment and the resulting precipitate identity. Int. J. Environ. Sci. Technol. 2019, 17, 409–420. [Google Scholar] [CrossRef]
- Brink, H.G.; Mahlangu, Z. Microbial Lead(II) precipitation: The influence of growth substrate. Chem. Eng. Trans. 2018, 64, 439–444. [Google Scholar]
- Brink, H.G.; Hörstmann, C.; Feucht, C.B. Microbial Pb (II) Precipitation: Minimum Inhibitory Concentration and Precipitate Identity. Chem. Eng. Trans. 2018, 74, 1453–1458. [Google Scholar]
- Peens, J.; Wu, Y.W.; Brink, H.G. Microbial Pb (II) Precipitation: The Influence of Elevated Pb (II) Concentrations. Chem. Eng. Trans. 2018, 64, 1453–1458. [Google Scholar]
- Wang, H.; Cheng, H.; Wang, F.; Wei, D.; Wang, X. An improved 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT) reduction assay for evaluating the viability of Escherichia coli cells. J. Microbiol. Methods 2010, 82, 330–333. [Google Scholar] [CrossRef] [PubMed]
- Perkin-Elmer. Atomic Spectroscopy a Guide to Selecting the Appropriate Technique and System. Available online: www.perkinelmer.com/atomicspectroscopy (accessed on 17 August 2019).
- Kikuchi, S.; Ishimoto, M. Nitrate respiration of Klebsiella pneumoniae on amino acids, especially on serine. Z. Allg. Mikrobiol. 1980, 20, 405–413. [Google Scholar] [CrossRef] [PubMed]
- Tiquia-arashiro, S.M. Lead absorption mechanisms in bacteria as strategies for lead bioremediation. Appl. Microbiol. Biotechnol. 2018, 102, 5437–5444. [Google Scholar] [CrossRef] [PubMed]
- Pugazhendhi, A.; Manogari, G. New insight into effective biosorption of lead from aqueous solution using Ralstonia solanacearum: Characterization and mechanism studies. J. Clean. Prod. 2018, 174, 1234–1239. [Google Scholar] [CrossRef]
- Kirchman, D.L. Calculating microbial growth rates from data on production and standing stocks. Mar. Ecol. Prog. Ser. 2002, 233, 303–306. [Google Scholar] [CrossRef]
- Bionumbers. Available online: https://bionumbers.hms.harvard.edu/search.aspx (accessed on 25 February 2020).
- Kim, C.; Ainala, S.K.; Oh, Y.; Jeon, B.; Park, S.; Kim, J.R. Metabolic Flux Change in Klebsiella pneumoniae L17 by Anaerobic Respiration in Microbial Fuel Cell. Biotechnol. Bioproc. Eng. 2016, 260, 250–260. [Google Scholar] [CrossRef]
- Cerqueira, N.M.; Gonzalez, P.J.; Brondino, C.D.; Romão, C.C.; Moura, I.; Mouraet, J.J. The Effect of the Sixth Sulfur Ligand in the Catalytic Mechanism of Periplasmic Nitrate Reductase. J. Comput. Chem. 2009, 30, 2466–2484. [Google Scholar] [CrossRef] [PubMed]


| Bacteria | Pb Species | Mechanism | Description | Product | Reference |
|---|---|---|---|---|---|
| Enterobacter cloacae | Pb-contaminated soils | Precipitation | Mobilizing insoluble Phosphoric compounds to precipitate Pb(II) | Pb10(PO4)6(OH)2 | Hee and Bolan [6] |
| Klebsiella aerogenes NCTC 418 | PbCl2 | Precipitation | Releasing Sulfide from insoluble Sulfur compounds to precipitate Pb(II) | PbS | Aiking et al. [7] |
| Klebsiella michiganensis | Pb(NO3)2 | Biosorption | Biosorption of Pb(II) to extracellular polymeric substances (EPS) | Not mentioned | Bowman et al. [8] |
| Consortium (this research team) | Pb(NO3)2 | Dissimilatory precipitation | Sulfide released from insoluble Sulfur compounds to bind with Pb(II), oxidation-reduction to produce elemental Pb and PbO | PbS, PbO and elemental Pb | Brink et al. [9] |
| Independent Variables | ||||
| Pb(II) | 80 ppm | 500 ppm | ||
| Nutrients | Standard LB broth
| Simulated LB broth
| Standard LB broth
| Simulated LB broth
|
| Nitrates | Pb(NO3)2 | Pb(NO3)2 | Pb(NO3)2 | Pb(NO3)2 |
| Dependent Variables | ||||
| ||||
| Time (h) | B80LB | B80Sim | B500LB | B500Sim |
|---|---|---|---|---|
| 0 |  |  |  |  |
| 6 |  |  |  |  |
| 9 |  |  |  |  |
| 24 |  |  |  |  |
| 27 |  |  |  |  |
| 33 |  |  |  |  |
| Control | ||||
| 0 |  | |||
| 72 |  | |||
| 80LB | 500Sim | ||
|---|---|---|---|
| BLAST Hit | % | BLAST Hit | % |
| (Clostridium) bifermentans | 25.74 | Ralstonia solanacearum | 33.54 |
| Klebsiella pneumoniae | 21.52 | (Clostridium) bifermentans | 25.41 |
| Listeria monocytogenes | 20.36 | Klebsiella pneumoniae | 23.81 |
| HIV-1 isolate | 15.82 | Clostridium bifermentans | 6.26 |
| Uncultured bacterium | 6.12 | Uncultured bacterium | 4.57 |
| Uncultured organism | 2.43 | Clostridium sporogenes | 3.76 |
| No hits | 2.22 | Uncultured klebsiella | 1.44 |
| Clostridium bifermentans | 1.58 | Uncultured clostridium | 0.41 |
| Human immunodeficiency | 1.48 | Clostridium sp. | 0.26 |
| Bacillus sp. | 0.74 | HIV-1 isolate | 0.22 |
| Listeria phage | 0.32 | Uncultured organism | 0.09 |
| Pediococcus pentosaceus | 0.32 | Enterococcus casseliflavus | 0.08 |
| Uncultured klebsiella | 0.21 | Uncultured enterobacteriales | 0.06 |
| Experiment | ||
|---|---|---|
| B80Sim | 23.8 | 0.029 d = 0.55 h = 33 min 12 sec |
| B80LB | 23.6 | 0.029 d = 0.55 h = 33 min 12 sec |
| B500Sim | 13.1 | 0.053 d = 1 h 16 min = 76.2 min |
| B500LB | 13.2 | 0.053 d = 1 h 16 min = 76.2 min |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hörstmann, C.; Brink, H.G.; Chirwa, E.M.N. Pb(II) Bio-Removal, Viability, and Population Distribution of an Industrial Microbial Consortium: The Effect of Pb(II) and Nutrient Concentrations. Sustainability 2020, 12, 2511. https://doi.org/10.3390/su12062511
Hörstmann C, Brink HG, Chirwa EMN. Pb(II) Bio-Removal, Viability, and Population Distribution of an Industrial Microbial Consortium: The Effect of Pb(II) and Nutrient Concentrations. Sustainability. 2020; 12(6):2511. https://doi.org/10.3390/su12062511
Chicago/Turabian StyleHörstmann, Carla, Hendrik G. Brink, and Evans M.N. Chirwa. 2020. "Pb(II) Bio-Removal, Viability, and Population Distribution of an Industrial Microbial Consortium: The Effect of Pb(II) and Nutrient Concentrations" Sustainability 12, no. 6: 2511. https://doi.org/10.3390/su12062511
APA StyleHörstmann, C., Brink, H. G., & Chirwa, E. M. N. (2020). Pb(II) Bio-Removal, Viability, and Population Distribution of an Industrial Microbial Consortium: The Effect of Pb(II) and Nutrient Concentrations. Sustainability, 12(6), 2511. https://doi.org/10.3390/su12062511







