Effect of the C/N Ratio on Biodegradation of Ciprofloxacin and Denitrification from Low C/N Wastewater as Assessed by a Novel 3D-BER System
Abstract
1. Introduction
2. Materials and Methods
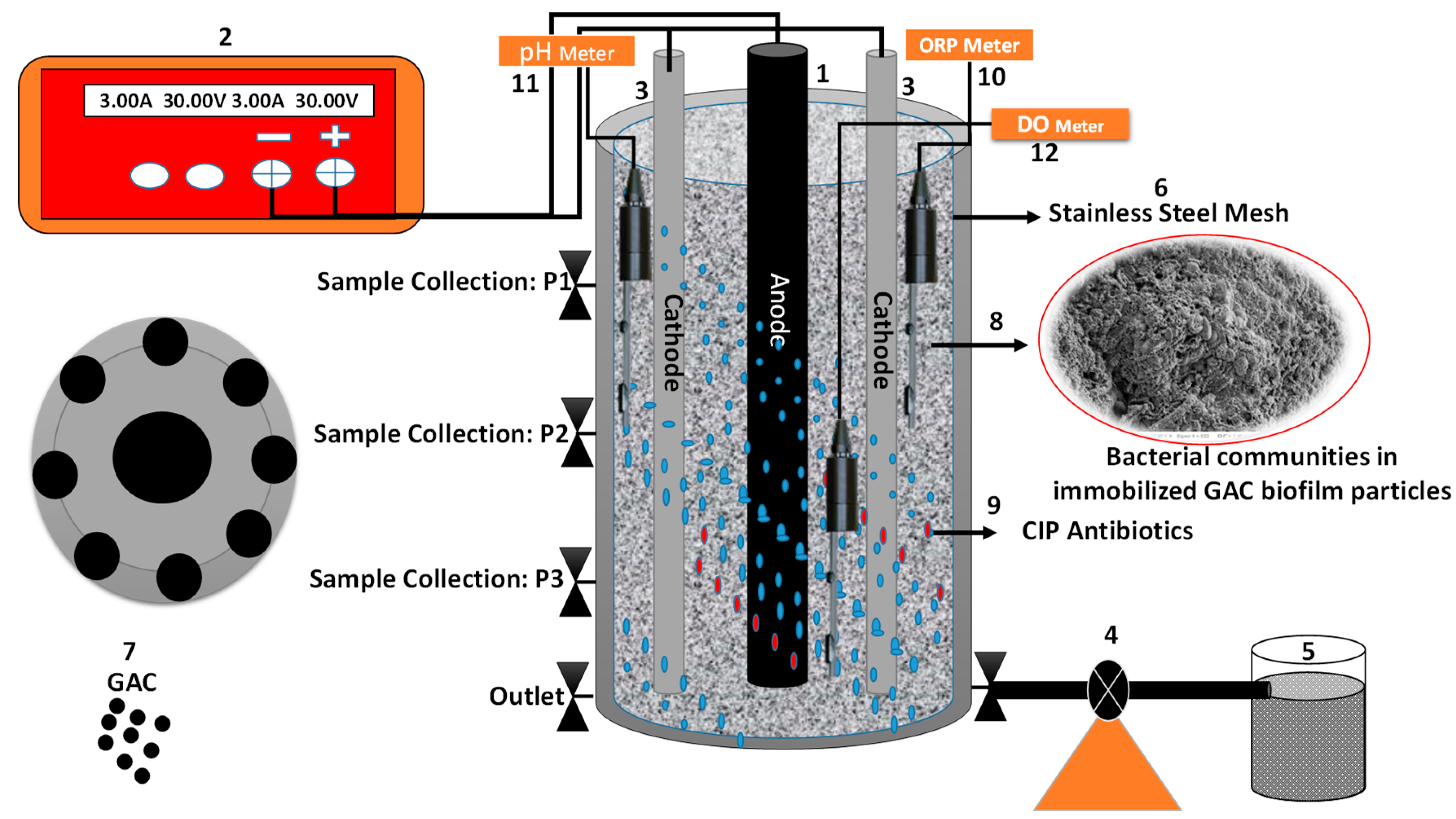
2.1. Configuration of the 3D-BER System
2.2. Chemicals and Reagents
2.3. Experimental Procedure
2.3.1. Bacterial Adaptation Phase
2.3.2. Immobilized GAC Biofilm at Various C/N Ratios
2.3.3. Analytical Methods and Calculations
2.4. Ciprofloxacin Concentrations Measurement and By-Products Identification
2.5. DNA Extraction, PCR Amplification, and High-Throughput Sequencing
2.6. Biofilms GAC Samples Analysis by SEM
2.7. Statistical Data Analysis
3. Results and Discussion
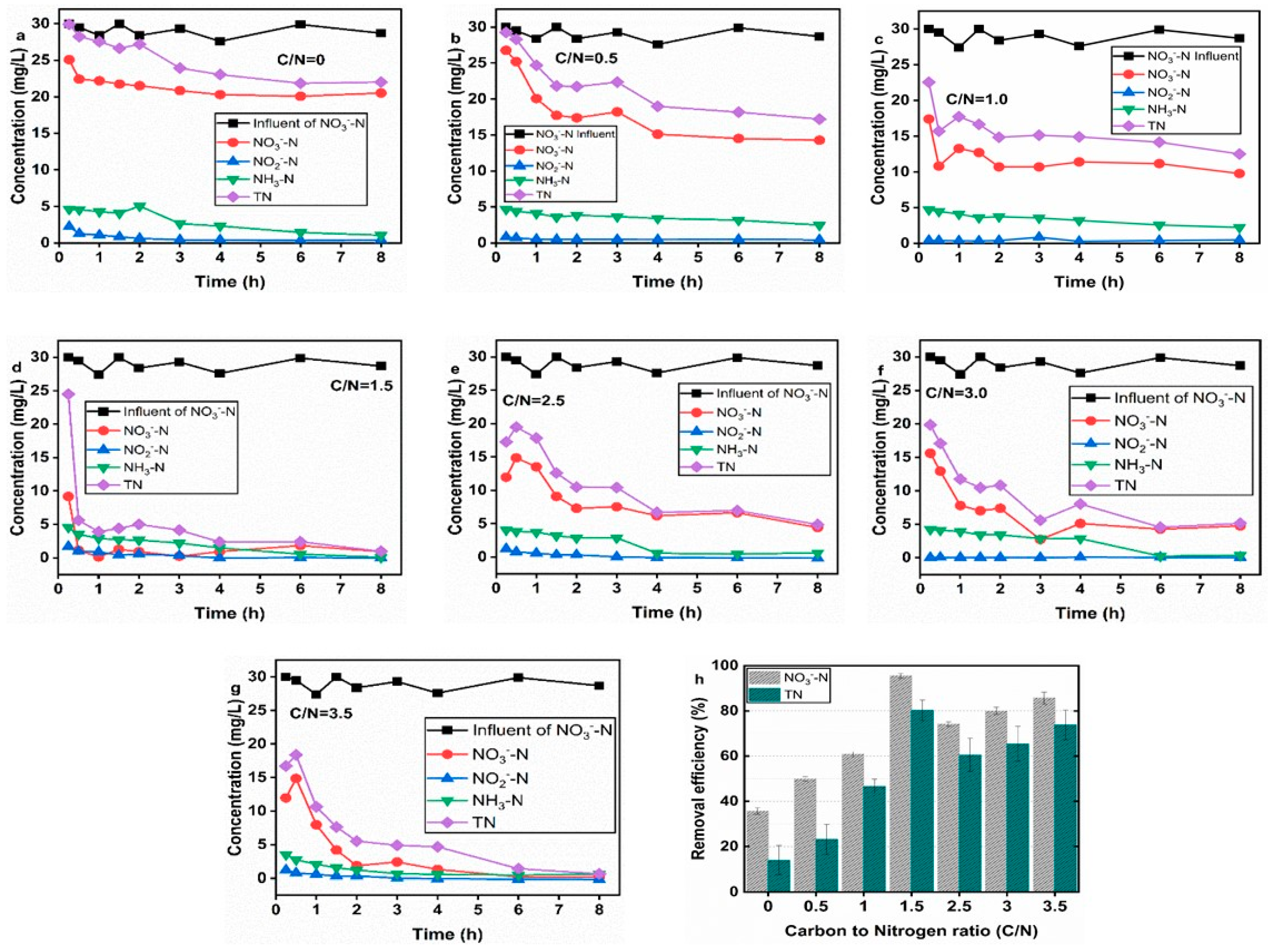
3.1. Removal Performance and the Effects of C/N Ratios
3.2. Effects of pH on Antibiotics CIP, Nitrate and Total Nitrogen Removal
3.3. Effect of pH on Biodegradation Mechanism of the Antibiotic Ciprofloxacin
3.4. Identification of Degradation Pathways by LC-MS/MS
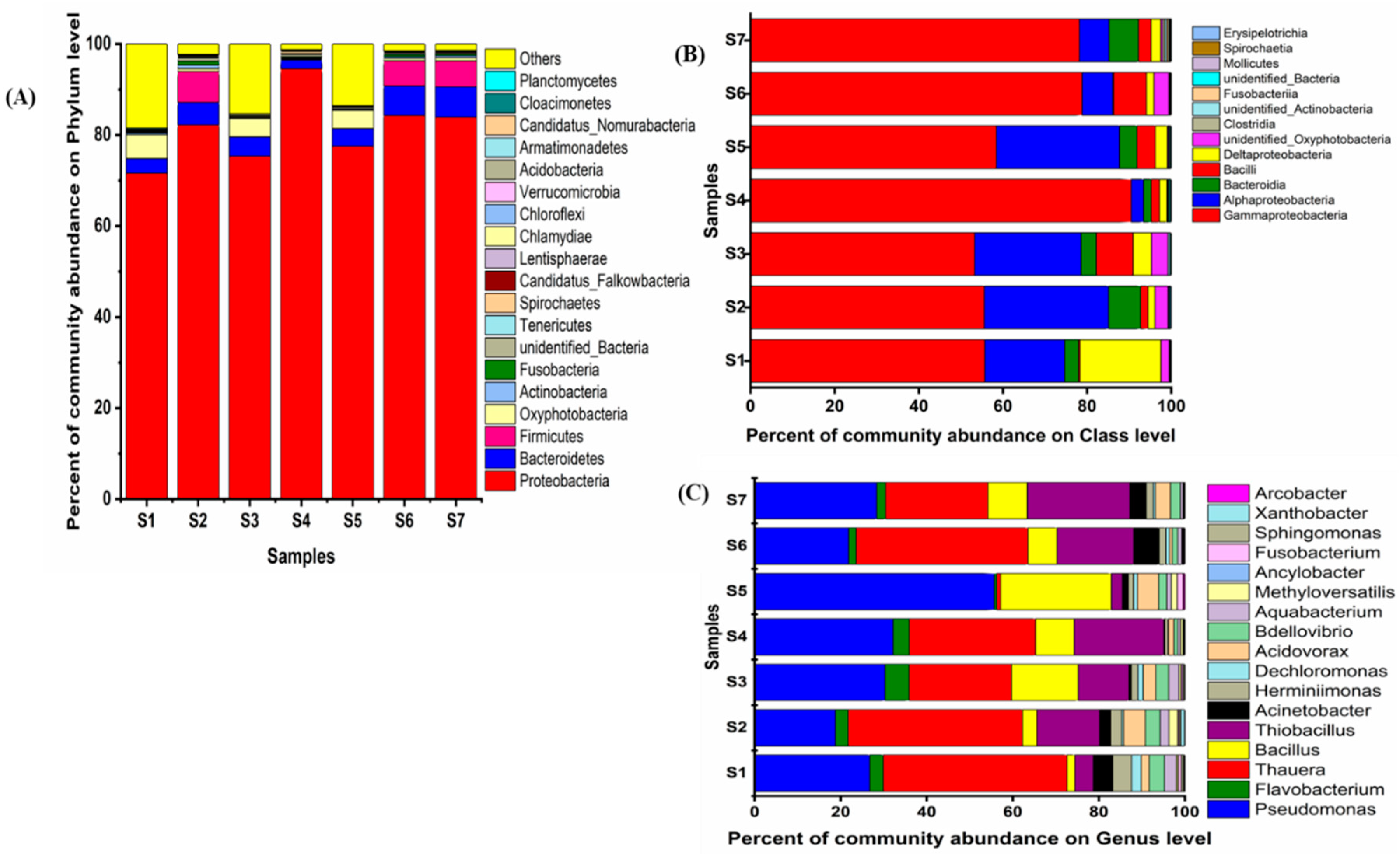
3.5. Bacterial Diversity and Community Composition
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Pan, L.-J.; Li, J.; Li, C.-X.; Tang, X.-D.; Yu, G.-W.; Wang, Y. Study of ciprofloxacin biodegradation by a Thermus sp. isolated from pharmaceutical sludge. J. Hazard. Mater. 2018, 343, 59–67. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; Gao, G.; Yang, T.-T.; Chen, X.; Chao, Y.; Na, G.; Ge, L.; Huang, L.-N. Effects of fluoroquinolone antibiotics on reactor performance and microbial community structure of a membrane bioreactor. Chem. Eng. J. 2015, 280, 448–458. [Google Scholar] [CrossRef]
- Nguyen, T.-T.; Bui, X.-T.; Dang, B.-T.; Ngo, H.-H.; Jahng, D.; Fujioka, T.; Chen, S.-S.; Dinh, Q.-T.; Nguyen, C.-N.; Nguyen, T.-T. Effect of ciprofloxacin dosages on the performance of sponge membrane bioreactor treating hospital wastewater. Bioresour. Technol. 2019, 273, 573–580. [Google Scholar] [CrossRef] [PubMed]
- Liao, X.; Li, B.; Zou, R.; Dai, Y.; Xie, S.; Yuan, B. Biodegradation of antibiotic ciprofloxacin: Pathways, influential factors, and bacterial community structure. Environ. Sci. Pollut. Res. 2016, 23, 7911–7918. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Hong, H.; Liao, L.; Ackley, C.J.; Schulz, L.A.; Macdonald, R.A.; Mihelich, A.L.; Emard, S.M. A mechanistic study of ciprofloxacin removal by kaolinite. Colloids Surf. B Biointerfaces 2011, 88, 339–344. [Google Scholar] [CrossRef]
- Liu, Z.; Sun, P.; Pavlostathis, S.G.; Zhou, X.; Zhang, Y. Inhibitory effects and biotransformation potential of ciprofloxacin under anoxic/anaerobic conditions. Bioresour. Technol. 2013, 150, 28–35. [Google Scholar] [CrossRef]
- Sahlin, S.; Joakim Larsson, D.; Ågerstrand, M. Ciprofloxacin: EQS Data Overview. In The Department of Environmental Science and Analytical Chemistry (ACES) Report; Stockholms Universitet: Stockholm, Sweden, 2018; p. 15. [Google Scholar]
- Mougin, C.; Cheviron, N.; Repincay, C.; Hedde, M.; Hernandez-Raquet, G. Earthworms highly increase ciprofloxacin mineralization in soils. Environ. Chem. Lett. 2013, 11, 127–133. [Google Scholar] [CrossRef]
- Liu, Z.; Sun, P.; Pavlostathis, S.G.; Zhou, X.; Zhang, Y. Adsorption, inhibition, and biotransformation of ciprofloxacin under aerobic conditions. Bioresour. Technol. 2013, 144, 644–651. [Google Scholar] [CrossRef]
- Lima, V.B.; Goulart, L.A.; Rocha, R.S.; Steter, J.R.; Lanza, M.R. Degradation of antibiotic ciprofloxacin by different AOP systems using electrochemically generated hydrogen peroxide. Chemosphere 2020, 247, 125807. [Google Scholar] [CrossRef]
- Li, J.; Cheng, W.; Xu, L.; Strong, P.; Chen, H. Antibiotic-resistant genes and antibiotic-resistant bacteria in the effluent of urban residential areas, hospitals, and a municipal wastewater treatment plant system. Environ. Sci. Pollut. Res. 2015, 22, 4587–4596. [Google Scholar] [CrossRef]
- Polesel, F.; Lehnberg, K.; Dott, W.; Trapp, S.; Thomas, K.V.; Plósz, B.G. Factors influencing sorption of ciprofloxacin onto activated sludge: Experimental assessment and modelling implications. Chemosphere 2015, 119, 105–111. [Google Scholar] [CrossRef] [PubMed]
- Shokoohi, R.; Ghobadi, N.; Godini, K.; Hadi, M.; Atashzaban, Z. Antibiotic detection in a hospital wastewater and comparison of their removal rate by activated sludge and earthworm-based vermifilteration: Environmental risk assessment. Process Saf. Environ. Prot. 2020, 134, 169–177. [Google Scholar] [CrossRef]
- Sukul, P.; Spiteller, M. Fluoroquinolone Antibiotics in the Environment. Reviews of Environmental Contamination and Toxicology; Springer: Berlin/Heidelberg, Germany, 2007; pp. 131–162. [Google Scholar]
- Alexandrino, D.A.; Mucha, A.P.; Almeida, C.M.R.; Gao, W.; Jia, Z.; Carvalho, M.F. Biodegradation of the veterinary antibiotics enrofloxacin and ceftiofur and associated microbial community dynamics. Sci. Total. Environ. 2017, 581, 359–368. [Google Scholar] [CrossRef] [PubMed]
- Amorim, C.L.; Moreira, I.S.; Maia, A.S.; Tiritan, M.E.; Castro, P.M. Biodegradation of ofloxacin, norfloxacin, and ciprofloxacin as single and mixed substrates by Labrys portucalensis F. Appl. Microbiol. Biotechnol. 2014, 98, 3181–3190. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Yang, Y.; Dai, Y.; Xie, S. Anaerobic biodegradation of nonylphenol in river sediment under nitrate-or sulfate-reducing conditions and associated bacterial community. J. Hazard. Mater. 2015, 286, 306–314. [Google Scholar] [CrossRef]
- Hao, R.; Li, S.; Li, J.; Meng, C. Denitrification of simulated municipal wastewater treatment plant effluent using a three-dimensional biofilm-electrode reactor: Operating performance and bacterial community. Bioresour. Technol. 2013, 143, 178–186. [Google Scholar] [CrossRef]
- Tao, M.; Guan, L.; Jing, Z.; Tao, Z.; Wang, Y.; Luo, H.; Wang, Y. Enhanced denitrification and power generation of municipal wastewater treatment plants (WWTPs) effluents with biomass in microbial fuel cell coupled with constructed wetland. Sci. Total. Environ. 2020, 709, 136159. [Google Scholar] [CrossRef]
- Wang, S.-Y.; Yang, X.-Y.; Meng, H.-S.; Zhang, Y.-C.; Li, X.-Y.; Xu, J. Enhanced denitrification by nano ɑ-Fe2O3 induced self-assembled hybrid biofilm on particle electrodes of three-dimensional biofilm electrode reactors. Environ. Int. 2019, 125, 142–151. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhang, B.; Feng, C.; Huang, F.; Zhang, P.; Zhang, Z.; Yang, Y.; Sugiura, N. Behavior of autotrophic denitrification and heterotrophic denitrification in an intensified biofilm-electrode reactor for nitrate-contaminated drinking water treatment. Bioresour. Technol. 2012, 107, 159–165. [Google Scholar] [CrossRef]
- Zhou, M.; Fu, W.; Gu, H.; Lei, L. Nitrate removal from groundwater by a novel three-dimensional electrode biofilm reactor. Electrochim. Acta 2007, 52, 6052–6059. [Google Scholar] [CrossRef]
- Gros, H. Biological denitrification process with hydrogen-oxidizing bacteria for drinking water treatment. Water Supply 1988, 6, 193–198. [Google Scholar]
- Kurt, M.; Dunn, I.J.; Bourne, J.R. Biological denitrification of drinking water using autotrophic organisms with H2 in a fluidized-bed biofilm reactor. Biotechnol. Bioeng. 1987, 29, 493–501. [Google Scholar] [CrossRef] [PubMed]
- Tong, Y.; He, Z. Nitrate removal from groundwater driven by electricity generation and heterotrophic denitrification in a bioelectrochemical system. J. Hazard. Mater. 2013, 262, 614–619. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zhang, T. Biodegradation and adsorption of antibiotics in the activated sludge process. Environ. Sci. Technol. 2010, 44, 3468–3473. [Google Scholar] [CrossRef] [PubMed]
- Sobieszuk, P.; Szewczyk, K. Estimation of (C/N) ratio for microbial denitrification. Environ Technol. 2006, 27, 103–108. [Google Scholar] [CrossRef] [PubMed]
- Park, H.I.; Kim, J.S.; Kim, D.K.; Choi, Y.-J.; Pak, D. Nitrate-reducing bacterial community in a biofilm-electrode reactor. Enzym. Microb. Technol. 2006, 39, 453–458. [Google Scholar] [CrossRef]
- Percheron, G.; Bernet, N.; Moletta, R. Interactions between methanogenic and nitrate reducing bacteria during the anaerobic digestion of an industrial sulfate rich wastewater. FEMS Microbiol. Ecol. 1999, 29, 341–350. [Google Scholar] [CrossRef]
- Wei, X.; Chen, J.; Xie, Q.; Zhang, S.; Ge, L.; Qiao, X. Distinct photolytic mechanisms and products for different dissociation species of ciprofloxacin. Environ. Sci. Technol. 2013, 47, 4284–4290. [Google Scholar] [CrossRef]
- Ji, Y.; Ferronato, C.; Salvador, A.; Yang, X.; Chovelon, J.-M. Degradation of ciprofloxacin and sulfamethoxazole by ferrous-activated persulfate: Implications for remediation of groundwater contaminated by antibiotics. Sci. Total. Environ. 2014, 472, 800–808. [Google Scholar] [CrossRef]
- Hamilton, E. Laboratory manual for the examination of water, waste water and soil. Sci. Total Environ. 1990, 92, 289. [Google Scholar] [CrossRef]
- Guo, H.; Jiang, N.; Wang, H.; Shang, K.; Lu, N.; Li, J.; Wu, Y. Pulsed discharge plasma induced WO3 catalysis for synergetic degradation of ciprofloxacin in water: Synergetic mechanism and degradation pathway. Chemosphere 2019, 230, 190–200. [Google Scholar] [CrossRef] [PubMed]
- Hijosa-Valsero, M.; Fink, G.; Schlüsener, M.P.; Sidrach-Cardona, R.; Martín-Villacorta, J.; Ternes, T.; Becares, E. Removal of antibiotics from urban wastewater by constructed wetland optimization. Chemosphere 2011, 83, 713–719. [Google Scholar] [CrossRef] [PubMed]
- Dorival-García, N.; Zafra-Gómez, A.; Navalón, A.; González, J.; Vilchez, J. Removal of quinolone antibiotics from wastewaters by sorption and biological degradation in laboratory-scale membrane bioreactors. Sci. Total Environ. 2013, 442, 317–328. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Wang, H.; Yang, K.; Ma, F. Performance and microbial communities in a combined bioelectrochemical and sulfur autotrophic denitrification system at low temperature. Chemosphere 2018, 193, 337–342. [Google Scholar] [CrossRef]
- Zhang, B.; Liu, Y.; Tong, S.; Zheng, M.; Zhao, Y.; Tian, C.; Liu, H.; Feng, C. Enhancement of bacterial denitrification for nitrate removal in groundwater with electrical stimulation from microbial fuel cells. J. Power Sources 2014, 268, 423–429. [Google Scholar] [CrossRef]
- Fry, D.E.; Garrison, R.N.; Trachtenberg, L.; Polk, H.C. Bacterial inoculum and the activity of antimicrobial agents. Surg. Gynecol. Obstet. 1985, 160, 105–108. [Google Scholar]
- Chen, C.; Ho, K.-L.; Liu, F.-C.; Ho, M.; Wang, A.-J.; Ren, N.; Lee, D.-J. Autotrophic and heterotrophic denitrification by a newly isolated strain Pseudomonas sp. C27. Bioresour. Technol. 2013, 145, 351–356. [Google Scholar] [CrossRef]
- Srivastava, P.; Yadav, A.K.; Abbassi, R.; Garaniya, V.; Lewis, T.W. Denitrification in a low carbon environment of a constructed wetland incorporating a microbial electrolysis cell. J. Environ. Chem. Eng. 2018, 6, 5602–5607. [Google Scholar] [CrossRef]
- Mousavi, S.A.; Ibrahim, S.; Aroua, M.; Ghafari, S. Development of nitrate elimination by autohydrogenotrophic bacteria in bio-electrochemical reactors—A review. Biochem. Eng. J. 2012, 67, 251–264. [Google Scholar] [CrossRef]
- Wu, G.; Li, Z.; Huang, Y.; Zan, F.; Dai, J.; Yao, J.; Yang, B.; Chen, G.; Lei, L. Electrochemically assisted sulfate reduction autotrophic denitrification nitrification integrated (e-SANI®) process for high-strength ammonium industrial wastewater treatment. Chem. Eng. J. 2020, 381, 122707. [Google Scholar] [CrossRef]
- Bertanza, G.; Menoni, L.; Capoferri, G.U.; Pedrazzani, R. Promoting biological phosphorus removal in a full scale pre-denitrification wastewater treatment plant. J. Environ. Manag. 2019, 254, 109803. [Google Scholar] [CrossRef] [PubMed]
- Ge, S.; Ma, J.; Liu, L.; Yuan, Z.-M. The Impact of Exogenous Aerobic Bacteria on Sustainable Methane Production Associated with Municipal Solid Waste Biodegradation: Revealed by High-Throughput Sequencing. Sustainability 2020, 12, 1815. [Google Scholar] [CrossRef]
- Wan, D.; Liu, H.; Qu, J.; Lei, P.; Xiao, S.; Hou, Y. Using the combined bioelectrochemical and sulfur autotrophic denitrification system for groundwater denitrification. Bioresour. Technol. 2009, 100, 142–148. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Nguyen, L.N.; Oh, S. Ecological impact of the antibiotic ciprofloxacin on microbial community of aerobic activated sludge. Environ. Geochem. Heal. 2019, 2019, 1–11. [Google Scholar] [CrossRef]
- Zhang, L.; Yue, Q.; Yang, K.; Zhao, P.; Gao, B. Analysis of extracellular polymeric substances (EPS) and ciprofloxacin-degrading microbial community in the combined Fe-C micro-electrolysis-UBAF process for the elimination of high-level ciprofloxacin. Chemosphere 2018, 193, 645–654. [Google Scholar] [CrossRef]
- Zhao, Y.; Feng, C.; Wang, Q.; Yang, Y.; Zhang, Z.; Sugiura, N. Nitrate removal from groundwater by cooperating heterotrophic with autotrophic denitrification in a biofilm–electrode reactor. J. Hazard. Mater. 2011, 192, 1033–1039. [Google Scholar] [CrossRef] [PubMed]
- Song, P.; Huang, G.; Hong, Y.; An, C.; Xin, X.; Zhang, P. A biophysiological perspective on enhanced nitrate removal from decentralized domestic sewage using gravitational-flow multi-soil-layering systems. Chemosphere 2020, 240, 124868. [Google Scholar] [CrossRef]
- Si, Z.; Song, X.; Wang, Y.; Cao, X.; Zhao, Y.; Wang, B.; Chen, Y.; Arefe, A.; Wang, Y. Intensified heterotrophic denitrification in constructed wetlands using four solid carbon sources: Denitrification efficiency and bacterial community structure. Bioresour. Technol. 2018, 267, 416–425. [Google Scholar] [CrossRef]
- Chiu, Y.-C.; Chung, M.-S. Determination of optimal COD/nitrate ratio for biological denitrification. Int. Biodeterior. Biodegrad. 2003, 51, 43–49. [Google Scholar] [CrossRef]
- Yi, K.; Wang, D.; Yang, Q.; Li, X.; Chen, H.; Sun, J.; An, H.; Wang, L.; Deng, Y.; Liu, J.; et al. Effect of ciprofloxacin on biological nitrogen and phosphorus removal from wastewater. Sci. Total. Environ. 2017, 605, 368–375. [Google Scholar] [CrossRef]
- Zhang, H.; Jia, Y.; Khanal, S.K.; Lu, H.; Fang, H.; Zhao, Q. Understanding the Role of Extracellular Polymeric Substances on Ciprofloxacin Adsorption in Aerobic Sludge, Anaerobic Sludge, and Sulfate-Reducing Bacteria Sludge Systems. Environ. Sci. Technol. 2018, 52, 6476–6486. [Google Scholar] [CrossRef] [PubMed]
- Hermosilla, D.; Han, C.; Nadagouda, M.N.; Machala, L.; Gascó, A.; Campo, P.; Dionysiou, D.D. Environmentally friendly synthesized and magnetically recoverable designed ferrite photo-catalysts for wastewater treatment applications. J. Hazard. Mater. 2019, 381, 121200. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Gao, D.; Xu, J.; Cai, L.; Cheng, J.; Yu, Z.; Hu, Z.; Yu, J. Interaction of ciprofloxacin with the activated sludge of the sewage treatment plant. Environ. Sci. Pollut. Res. 2018, 25, 35064–35073. [Google Scholar] [CrossRef]
- Kim, D.G.; Choi, D.; Cheon, S.; Ko, S.-O.; Kang, S.; Oh, S. Addition of biochar into activated sludge improves removal of antibiotic ciprofloxacin. J. Water Process. Eng. 2020, 33. [Google Scholar] [CrossRef]
- Jiang, M.; Zheng, X.; Chen, Y. Enhancement of denitrification performance with reduction of nitrite accumulation and N2O emission by Shewanella oneidensis MR-1 in microbial denitrifying process. Water Res. 2020, 169, 115242. [Google Scholar] [CrossRef] [PubMed]
- Jia, Y.; Khanal, S.K.; Shu, H.; Zhang, H.; Chen, G.-H.; Lu, H. Ciprofloxacin degradation in anaerobic sulfate-reducing bacteria (SRB) sludge system: Mechanism and pathways. Water Res. 2018, 136, 64–74. [Google Scholar] [CrossRef] [PubMed]
- Girardi, C.; Greve, J.; Lamshöft, M.; Fetzer, I.; Miltner, A.; Schäffer, A.; Kästner, M. Biodegradation of ciprofloxacin in water and soil and its effects on the microbial communities. J. Hazard. Mater. 2011, 198, 22–30. [Google Scholar] [CrossRef]
- Hosono, T.; Alvarez, K.; Lin, I.-T.; Shimada, J. Nitrogen, carbon, and sulfur isotopic change during heterotrophic (Pseudomonas aureofaciens) and autotrophic (Thiobacillus denitrificans) denitrification reactions. J. Contam. Hydrol. 2015, 183, 72–81. [Google Scholar] [CrossRef]
- Sakakibara, Y.; Araki, K.; Watanabe, T.; Kuroda, M. The denitrification and neutralization performance of an electrochemically activated biofilm reactor used to treat nitrate-contaminated groundwater. Water Sci. Technol. 1997, 36, 61. [Google Scholar] [CrossRef]
- Huo, S.; Kong, M.; Zhu, F.; Qian, J.; Huang, D.; Chen, P.; Ruan, R. Co-culture of Chlorella and wastewater-borne bacteria in vinegar production wastewater: Enhancement of nutrients removal and influence of algal biomass generation. Algal Res. 2020, 45, 101744. [Google Scholar] [CrossRef]
- Stuckey, D.C.; Oh, S. Effect of ciprofloxacin on methane production and anaerobic microbial community. Bioresour. Technol. 2018, 261, 240–248. [Google Scholar]
- Gonzalez-Martinez, A.; Rodriguez-Sanchez, A.; Martinez-Toledo, M.; García-Mesa, J.J.; Hontoria, E.; Osorio-Robles, F.; Gonzalez–Lopez, J. Effect of ciprofloxacin antibiotic on the partial-nitritation process and bacterial community structure of a submerged biofilter. Sci. Total. Environ. 2014, 476, 276–287. [Google Scholar] [CrossRef] [PubMed]




| Name | Properties | Molecular Formula | Structure |
|---|---|---|---|
| Ciprofloxacin | C17H18FN3O3 | 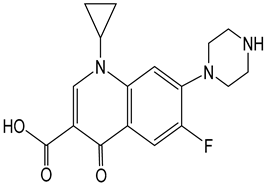 | |
| CAS number | 85721-33-1 | ||
| * pKa | 10.58 ± 0.30 | ||
| 8.71 ± 0.089 | |||
| 6.14 ± 0.13 | |||
| 3.01 ± 0.30 | |||
| * Log Kowb | 0.40 | ||
| Molecular weight | 331.35 g/mol | ||
| Parameters (mg·L−1) | S1 | S2 | S3 | S4 | S5 | S6 | S7 |
|---|---|---|---|---|---|---|---|
| Electric Current (mA) | 60 | 60 | 60 | 60 | 60 | 60 | 60 |
| COD | 0 | 15 | 30 | 45 | 75 | 90 | 105 |
| NO3−-N | 30 | 30 | 30 | 30 | 30 | 30 | 30 |
| C/N | 0 | 0.5 | 1.0 | 1.5 | 2.5 | 3.0 | 3.5 |
| Reactor Type | Mode | H/A-D | Treatment Source | E.V (L) | Initial (NO3-N) (mg/L) | C/N Source | Power Source | Input Energy | D.R.R (%) | CIP Removal (%) | Country | References |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Three-dimensional bioelectrochemical reactor system (3D-BERS) | Batch | H&A | Synthetic municipal wastewater | 0.738 | 30 mg/L | CH3COONa | DC | 60 mA | 95.53 | 94.20 | China | This Study |
| Three-dimensional biofilm-electrode reactor (3D-BER) | Continuous | H&A | Synthetic municipal wastewater | 3.4 | 30 mg/L | CH3CooNa | DC | 40 mA | 98.3 | N/A | China | [18] |
| Three-dimensional biofilm electrode reactors (3D-BERs) | Batch | A | Synthetic wastewater | 2 | 50–100 mg/L | Hydrogen & α-Fe2O3 | DC | 20–80 mA | 95 to 98.9 | N/A | China | [20] |
| A three-dimensional BER | Continuous | H&A | Synthetic groundwater | 0.5 | 20 mg/L | C2H5OH | DC | 15mA | 100 | N/A | China | [22] |
| Bioelectrochemical System (BES) | Batch | H | Synthetic groundwater | 1.0 | 250 g/m3 day | CH3CooNa | DC | N/A | 14.6 ± 0.2 36.2 ± 5.0 | N/A | USA | [25] |
| A combined single-chamberMFC and BER system | Continuous | H&A | Synthetic groundwater | 0.480 | 364 mg/L | CH3OH | MFC | 500–700 mV | 30 | N/A | China | [37] |
| A combined BES and sulfur autotrophic denitrification system (CBSAD) | Continuous | A | Synthetic groundwater | 33.47 | 20.9–22.0 mg NO3-N/L | Hydrogen | DC | 1000 mA | 95 | N/A | China | [45] |
| Bioreactor | Continuous | H&A | Synthetic municipal wastewater | 0.9 | 30 mg/L | C6H12O6 | -------- | -------- | N/A | 20–22 | Singapore | [46] |
| Fe-C micro-electrolysis reactor and up-flow biological aerated filter (UBAF) | Continuous | H&A | Synthetic municipal wastewater | 15 | 60–100 mg/L | Fe-C micro-electrolysis | N/A | N/A | N/A | 90 | China | [47] |
| Condition (C/N) | Sample ID | No. of Reads | No.of OTUs | ACE | Shannon | Simpson | Chao1 | Coverage |
|---|---|---|---|---|---|---|---|---|
| 0 | S1 | 39,322 | 182 | 254.321 | 2.642 | 0.321 | 254.119 | 0.993 |
| 0.5 | S2 | 40,215 | 189 | 259.583 | 2.997 | 0.224 | 245.337 | 0.995 |
| 1.0 | S3 | 41,234 | 254 | 288.981 | 3.456 | 0.423 | 288.327 | 0.996 |
| 1.5 | S4 | 73,382 | 382 | 390.705 | 6.367 | 0.838 | 399.229 | 0.999 |
| 2.5 | S5 | 54,633 | 267 | 297.149 | 3.994 | 0.455 | 345.657 | 0.994 |
| 3.0 | S6 | 59,236 | 263 | 311.597 | 4.214 | 0.754 | 358.279 | 0.995 |
| 3.5 | S7 | 63,324 | 353 | 363.642 | 5.247 | 0.936 | 361.571 | 0.999 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hassan, M.; Zhu, G.; Yang, Z.; Lu, Y.; Lang, Y.; Gong, L.; Shan, H. Effect of the C/N Ratio on Biodegradation of Ciprofloxacin and Denitrification from Low C/N Wastewater as Assessed by a Novel 3D-BER System. Sustainability 2020, 12, 7611. https://doi.org/10.3390/su12187611
Hassan M, Zhu G, Yang Z, Lu Y, Lang Y, Gong L, Shan H. Effect of the C/N Ratio on Biodegradation of Ciprofloxacin and Denitrification from Low C/N Wastewater as Assessed by a Novel 3D-BER System. Sustainability. 2020; 12(18):7611. https://doi.org/10.3390/su12187611
Chicago/Turabian StyleHassan, Mahdi, Guangcan Zhu, Zhonglian Yang, Yongze Lu, Yan Lang, Liying Gong, and Huang Shan. 2020. "Effect of the C/N Ratio on Biodegradation of Ciprofloxacin and Denitrification from Low C/N Wastewater as Assessed by a Novel 3D-BER System" Sustainability 12, no. 18: 7611. https://doi.org/10.3390/su12187611
APA StyleHassan, M., Zhu, G., Yang, Z., Lu, Y., Lang, Y., Gong, L., & Shan, H. (2020). Effect of the C/N Ratio on Biodegradation of Ciprofloxacin and Denitrification from Low C/N Wastewater as Assessed by a Novel 3D-BER System. Sustainability, 12(18), 7611. https://doi.org/10.3390/su12187611








