1. Introduction
Healthcare has drawn considerable attention in recent years, and increasing numbers of patients are engaging in online health communities (OHCs) for health information exchange [
1,
2,
3]. Online health communities are becoming an essential channel for users to search for health information and share their experiences of medical treatments [
4]. According to Health Information National Trends Survey 2017, about 80 percent of adults in the U.S. search for health-related information online [
5]. In China, there were around 195 million people using online medical services by the end of 2016 [
6]. With the rapid growth of healthcare service delivery, a number of new models have been developed recently, including online health consultations [
7]. Patients not only interact with their peers, but also consult doctors about their diseases through online communities [
8], which forms a new communication channel between patients and doctors. This new form of online patient–doctor communication has greatly changed the traditional delivery model of healthcare service. Online communication between patients and physicians can potentially alleviate the medical resource shortage problem and eliminate geographic barriers and time constraints to some extent [
9].
Online health consultations generate large amounts of valuable health-related information [
10]. The wide spread of periodic general health examinations [
11] also contributes to the fast growth of the medical datasets available. The vast development of information and communication technologies dramatically improves the deposition and exchange of health-related data, which facilitates healthcare Big Data analytics. As one of the Sustainable Development Goals (SDGs), sustainable healthcare is dedicated to ensuring healthy lives and promoting well-being for all people. Medical-related entity extraction from online health consultations can contribute to the sustainability of visualized healthcare in the following aspects. First, the extracted entities in online health consultations can facilitate the procedure of online patient–doctor communication by automatically recognizing and classifying the critical health concepts in patient- and doctor-generated text. The high efficiency of online health consultation helps improve the convenience and flexibility, saving costs and time for healthcare service delivery [
7,
12]. It can potentially support users to manage their health conditions electronically and thus attain more promising health outcomes and reduce future health risks [
13]. OHCs can also benefit from entity extraction by attracting more participants to engage in the information exchange platforms. Second, medical entity recognition is an essential task in clinical information extraction and medical knowledge discovery [
14], and can facilitate a number of healthcare-related applications such as disease surveillance [
15] and adverse drug reaction detection [
16]. Early detection of disease activity can reduce the impact of certain diseases such as seasonal influenza with a rapid response [
17]. Adverse drug reactions are among the top causes of morbidity and mortality and have been drawing considerable public attention [
18]. Disease surveillance and adverse drug reaction detection using social media data can enhance public health monitoring and ensure a healthier life [
19].
In this study, we aim to recognize several types of medical entities, namely, medical problems, medical tests, and treatments [
20], which are critical health concepts in medical knowledge discovery. Medical problem entity recognition aims to identify the states of diseases or symptoms in text to extract the health conditions of a patient, such as “breast cancer” and “fever”. The medical test entity recognition task seeks to find the medical examinations mentioned in text, including laboratory tests and physical examinations such as “blood test” and “CT scan”. Treatment entity recognition attempts to extract the mentions of therapy in medical text including drug names and surgery procedures, such as “glucose” and “heart transplantation”. For example, in the post “My right face was slightly swollen and accompanied by fever. … I didn’t feel better after taking glucocorticoid. After a blood test and other thorough examination, it was diagnosed as a facial lymphoma and now I’m ready for chemotherapy”. In this post, “slightly swollen”, “fever”, and “facial lymphoma” are medical problem entities; “blood test” is a medical test entity; and “glucocorticoid”, “chemotherapy” are treatment entities.
Extensive studies have been conducted to extract medical-related entities. Lexical-based methods recognize an entity by matching to the most similar or identical terms in a dictionary [
21], which makes lexical-based approaches particularly useful for practical information extraction [
22]. In the medical field, the most widely used controlled terminology dictionaries include UMLS (Unified Medical Language System) [
23], ICD (International Classification of Diseases) [
24], and SNOMED CT (Systematized Nomenclature of Medicine–Clinical Terms) [
25]. However, the short terms in the dictionary would result in false positives and significantly degrade the overall accuracy, and spelling variations which are quite common in the social media context make lexical-based approaches less usable. Machine learning approaches have been widely adopted for entity recognition because of their excellent environmental adaptability. The commonly used algorithms in entity recognition tasks include Maximum Entropy (ME) [
26], Support Vector Machine (SVM) [
27], Hidden Markov Model (HMM) [
28], and Conditional Random Fields (CRF) [
16,
29,
30]. Despite their excellent performance in some studies, the machine-learning-based models usually need tough feature engineering work. In recent years, the rapid improvement of deep learning techniques has brought new opportunities for natural language processing (NLP) studies including entity extraction [
31,
32,
33], and has significantly contributed to overcoming the above problem. Due to their capacity for automatically learning effective features from word embedding, deep neural network (DNN)-based models such as recurrent neural networks (RNNs) have been employed in state-of-the-art models [
31,
34]. As a unique RNN architecture, Long Short-Term Memory (LSTM) and its variant bidirectional LSTM (BiLSTM) have been utilized in entity extraction tasks and have shown encouraging performance [
31,
34,
35].
Although medical entity extraction has been widely studied, existing approaches have several limitations when applied to the Chinese social media context. First, most traditional machine learning approaches require complicated feature engineering work [
16,
23,
27,
28,
29,
30]. Feature engineering relies on handcrafted rules and language-specific knowledge, which is inherently tough and time-consuming [
34]. Second, most current works are designed for the English language context and ignore the uniqueness of Chinese. Compared with English, there is no blank space between words, and seldom do morphological changes exist in Chinese, making existing entity extraction approaches challenging in the context of Chinese. Third, unlike clinical notes that are described by healthcare professionals, the content in social media could be extensively informal, with features such as lexical variants, internet slang, typos, and grammatical errors. Previous approaches that used clinical notes as a data resource may fail to recognize out-of-vocabulary (OOV) terms, resulting in unsatisfying entity extraction performance [
36].
Recognizing the significance of medical entity extraction and the limitations of existing related works, this study proposes a novel DNN-based approach to extracting medical entities in the context of Chinese social media that can overcome the aforementioned problems. This study intends to enhance the sustainability of online healthcare services and public health monitoring by improving the performance of health concept extraction in online health consultations. Recent developments in DNN have achieved great success in many areas, providing new opportunities for natural language processing (NLP) research [
31,
33]. Specifically, we aimed to design a model that can automatically capture the context features of text to avoid tough feature engineering work and is effective in medical entity recognition in Chinese social media text. Considering the uniqueness of Chinese, we also evaluate the effect of recognition granularity on the performance of the entity extraction.
The rest of the paper is organized as follows. In
Section 2, we introduce the proposed medical entity extraction model, followed by the evaluation procedure in
Section 3. The experimental results are presented in
Section 4.
Section 5 discusses the evaluation results and reviews the practical implications of our model for the healthcare system. Lastly, we conclude our major research findings and research limitations in
Section 6.
2. Method
This study proposes a novel DNN-based model named CNMER (Chinese Medical Entity Recognition) to extract medical entities from Chinese OHCs.
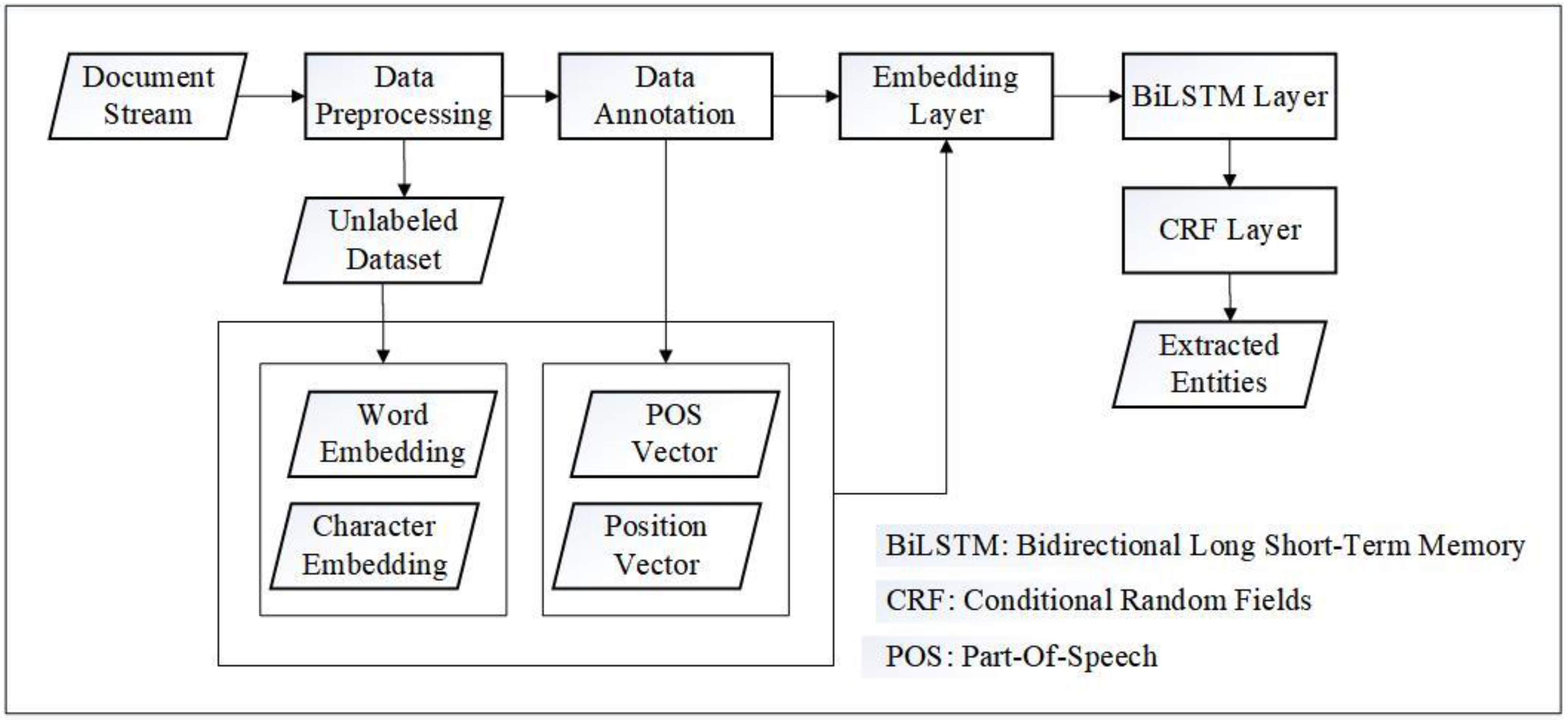
Figure 1 depicts an overview of our approach. After data collection, preprocessing was performed, and a subset of the processed data was randomly selected for data annotation. The remaining unlabeled dataset was utilized as the text corpus for unsupervised training on the domain word and character embeddings. With the part-of-speech (POS) feature and position feature, the trained embeddings were then used to formulate the character representation as the input for the BiLSTM-CRF.
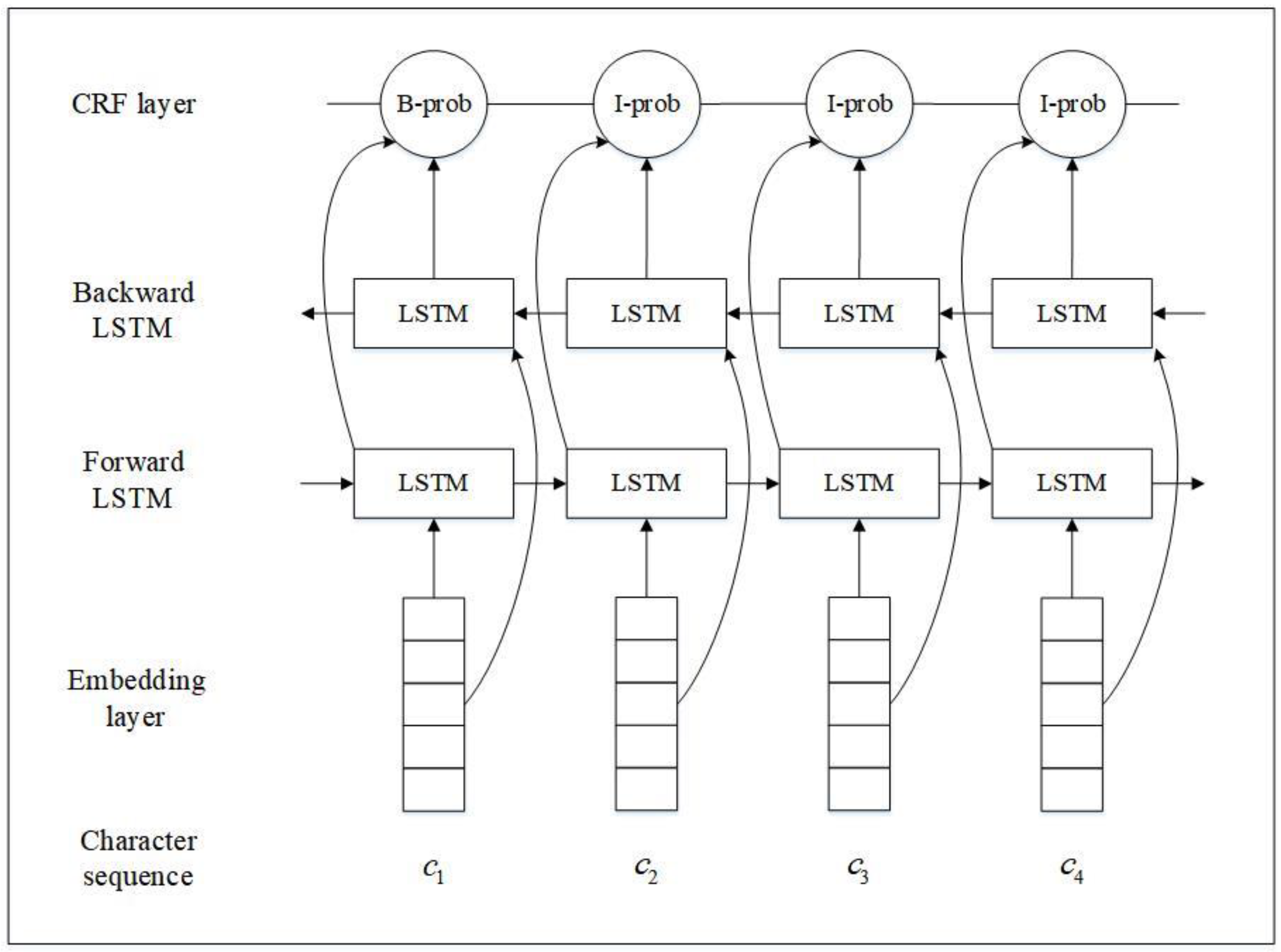
As shown in
Figure 2, the BiLSTM-CRF architecture consists of an embedding layer, a BiLSTM layer, and a CRF layer. The embedding layer maps each character in a sentence using the predefined numerical representation vector. The BiLSTM layer includes forward LSTM and backward LSTM, and takes the representation vectors of the character sequence as input and returns another sequence by considering both left and right context information. The CRF layer makes final tagging decisions based on the output of the BiLSTM layer using the CRF model.
2.1. Data Preprocessing and Annotation
The communications between physicians and patients in OHCs generate abundant health-related text. In this study, we exploited the online consultation text as the data resource. First, data preprocessing was performed to remove irrelevant contents such as private information, html tags, and other invalid characters. We also filtered out the consultations that were shorter than five characters. Unlike English text, words are not separated by blank spaces in Chinese sentences; thus, word segmentation was conducted to split each word in a sentence. We utilized Jieba, an open source NLP application in the Python language to segment sentences into words and perform POS tagging, for which a total of 40 types of POS tags were predefined. In this study, we employed Chinese Unified Medical Language System (CUMLS) [
37], a repository of biomedical terminologies developed by Chinese Academy of Medical Sciences to help improve the performance of Chinese word segmentation for the health-related corpus. CUMLS integrates more than ten biomedical sources such as biomedical thesauri, classifications, and text words of biomedical literature. Specifically, CUMLS includes 100,000 medical terms. Using CUMLS as the supplementary dictionary, terms in consultations that are matched with the repository can be extracted and segmented as a single word automatically.
After data preprocessing, we randomly selected a small subset from the obtained corpus as the source of data annotation. An annotation protocol was developed before annotation. To obtain the annotation dataset, two expert annotators were recruited to independently label the entity boundaries and types in sentences. Another expert annotator was asked to check any disagreements and make the final judgement. In this study, we labeled entities using the “BIO” tagging formalism, where the “B” category represents the beginning of an entity, the “I” category represents the continuity of an entity, and “O” denotes all other characters. As an illustration, for a medical problem entity which consists of four characters in total, namely , the annotators are supposed to tag the character sequence as “B-prob, I-prob, I-prob, I-prob”.
2.2. Embedding Layer
Conventional machine learning approaches lack the ability to process natural data in their raw form and require careful engineering and designing work to extract effective features from raw data such as plain text [
38]. The input of machine learning approaches is usually represented in the form of a fixed-length feature vector. For text input, bag-of-words is one of the most common used features. Although they have been widely used, bag-of-words features have certain disadvantages: they fail to capture the order of words in text and they miss the semantic information of words. For example, the word “sickness”, “illness”, and “hospital” are represented with the same distance by bag-of-words, although “sickness” should be closer to “illness” than to “hospital”, semantically.
Distributed representations of words in the form of a vector space can group similar words and facilitate many natural language processing tasks toward better performance. Representation learning approaches can automatically detect the information needed and represent it at a higher and more abstract level. A word embedding maps a word to a numerical vector in a low-dimensional vector space which can capture semantic or syntactic properties of the word; semantically similar words are expected to be assigned similar vectors [
34]. The learned word representations explicitly encode many linguistic regularities and patterns, and many of these patterns can be represented as linear translations [
39]. For example, the result of calculation vec(“Beijing”) − vec(“China”) + vec(“Japan”) is closer to vec(“Tokyo”) than to any other learned word vectors, where “vec” represents the learned embedding vector of a word. This study uses the skip-gram method for both word- and character-level embedding training [
39], which predicts the words that are most likely to appear around the focused word. Given a sequence of training words
, the model is trained by maximizing the average log probability
where
is the size of training context, and
are the words surrounding the focused
. The basic skip-gram formulation defines
using the Softmax function
where
and
denote the “input” and “output” vector representations of
, respectively, and
represents the total number of words in the vocabulary [
39]. We use word2vec, an open source tool developed by Google, to train the character and word embedding [
39]. We trained a 100-dimensional embedding for both characters and words based on the unlabeled dataset [
33].
For Chinese online health-related text, word segmentation is a challenging task [
40] which could result in unsatisfying performance for word-based entity extraction methods. To address this issue, character-based entity recognition was proposed [
41]. The character representation has been recognized as an important factor that impacts the entity recognition performance [
32,
33]. However, the semantic information of a character varies according to context, while the same character in different contexts is usually represented by the same embedding. Therefore, the direct use of character-level embedding for the various contexts will lead to inaccurate character feature representation [
33]. In this study, we propose to combine character embedding with the context word embedding as a part of the character embedding vector. Thus, the character representation incorporates not only the features of the focal character, but also the context information of the related word.
The POS feature and the position of a character in the context word [
42] were also incorporated into our model as they carry critical context information for the focused character. According to the tagging scheme in Jieba, we predefined a list of POS tags and mapped each tag to a 40-dimensional one-hot vector to represent the POS feature of the context word. To represent the position feature of a character, we used a 4-dimensional one-hot vector to present the positional information of a character in the context word: single-character word, the beginning of a word, the middle of a word, or the end of a word. All the embeddings and vectors were then concatenated together as a single vector, and finally we obtained a 244-dimensional numerical representation for each character as input for the BiLSTM network.
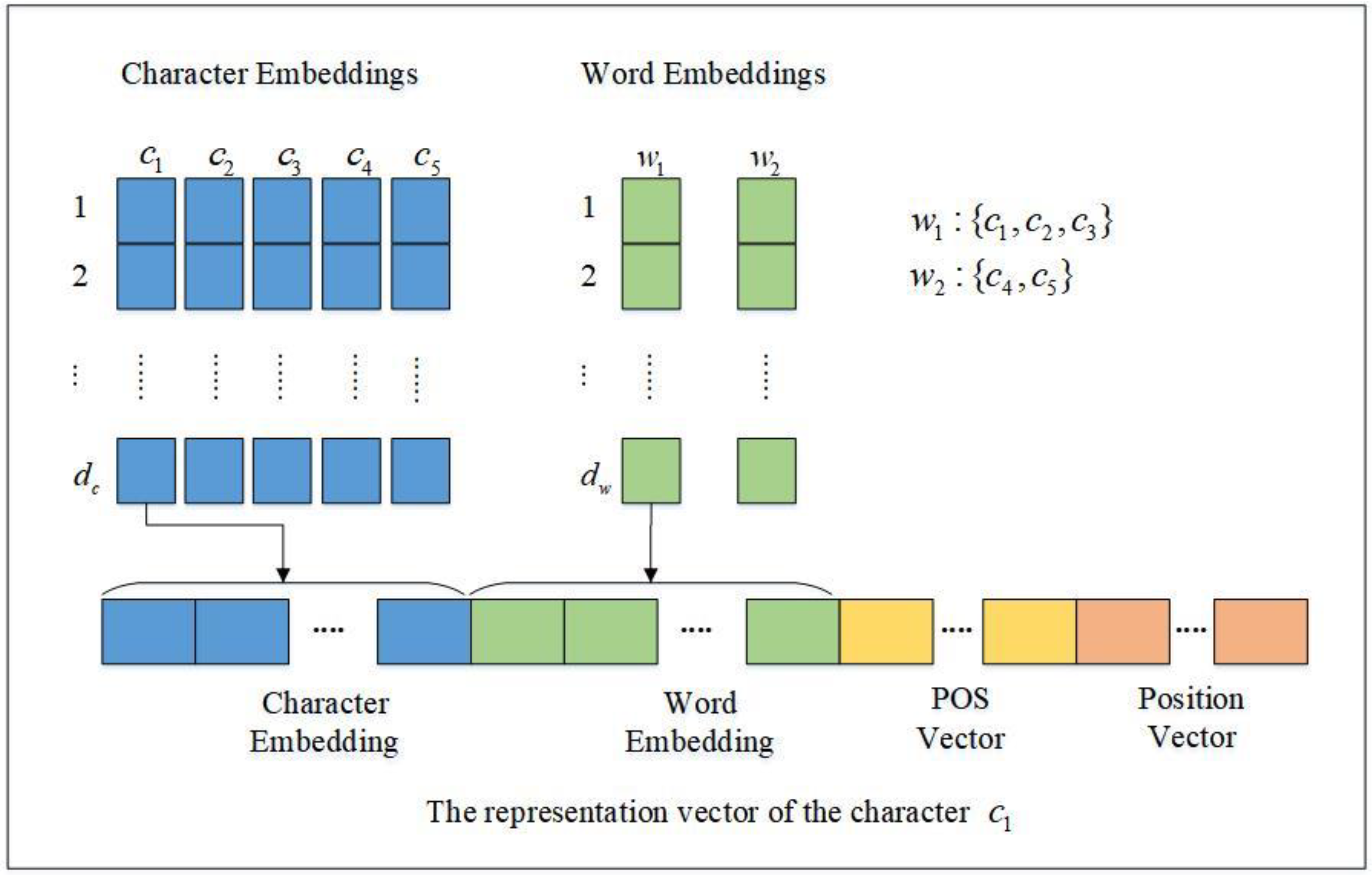
Figure 3 illustrates an example of character representation used in our model, where
represents the dimension of the character embedding and
indicates the dimension of the word embedding. In the example, the entity is divided into two parts during word segmentation, namely,
which consists of
, and
, which consists of
. Therefore, the representation vector of the character
consists of four parts, which are the character embedding of
, the word embedding of
, the POS feature vector of the word
, and the position feature (i.e., “the beginning of a word”) vector of the character
in the word
.
2.3. BiLSTM Layer
A typical neural network contains a set of input units, multiple hidden layers that contain hidden units, a set of output units that stands for tags, and the connections between those units [
43]. The model is trained using an algorithm named “back-propagation” to adjust the weights of connections between units, so that any input tends to generate the corresponding output. The relationship between inputs and outputs that a neural network learns can be regarded as a mapping, and neural networks with multiple hidden layers are believed to be good at learning mappings.
Deep neural networks are neural networks with a large number of hidden layers. A deep neural network system is usually regarded as a classification system that decides what category (e.g., entity type) a given input (e.g., word) is mapped to. Theoretically, given infinite data, a deep learning system is capable of representing any deterministic mapping for any given inputs and corresponding outputs [
43]. However, due to the finite amount of data available in real-world applications, deep learning systems have to generalize beyond the training data.
Compared with human beings, deep learning systems lack the ability to learn abstractions from explicit and verbal definitions. Instead, they rely on the large amount of training examples to learn these rules. In the context of entity recognition, given the definition of a medical entity, humans can easily tell whether a word is a medical entity and what type the entity is. However, deep learning models have to learn this “definition” through large numbers of annotated examples. In a DNN, the final tagging result of a given input character in terms of medical recognition depends on many features, such as the POS information, the positional information, and the context words. The hidden layers in a DNN are considered as complex feature transformations in the networks and produce the most abstract features for the final output layer; this is a critical process in learning the implicit rules embedded in the training set.
The RNN is an extension of the traditional feedforward neural network, and can handle variable-length input sequences. An RNN contains a recurrent hidden state, and the activation of the hidden state depends on that of the previous time. Nevertheless, RNNs fail to capture long-term dependencies as the gradient tends to either vanish or explode during training.
The LSTM is a special kind of RNN that is designed to avoid the long-term dependency issue by joining with a gated memory cell [
44]. Typically, an LSTM unit consists of an input gate
, an output gate
, a forget gate
, a memory cell
, and a hidden state
. The LSTM incorporates these structures called gates to optionally remove or add information; they contain a sigmoid neural net layer and a pointwise multiplication operation. The sigmoid layer outputs values between 0 and 1 to indicate how much of each component should be reserved, in which a value of 0 denotes “let nothing through” and 1 denotes “let everything through”. The LSTM computes the output by iterating the following equations:
where
means the sigmoid function;
denotes pointwise multiplication;
,
,
, and
(with subscripts
,
, and
) are the weight matrices for input
, hidden state
, memory cell
, and output
, respectively; and
,
,
, and
denote the bias vectors. The BiLSTM is composed of a forward LSTM and a backward LSTM to capture both past and future information, which are two separated networks with different parameters.
The entity extraction task can be modeled by deep learning methods as a sequence labeling task. In OHCs, there are many long sentences in patient-contributed content, and the semantic meaning of a focused character can be shaped by the characters before and after it over a long distance. In the text sequence of online consultations, users report their health conditions in detail and the mentions of each medical entity could rely on long-distance information in the text. Based on these intuitions, we utilized BiLSTM to extract medical named entities, as BiLSTM can learn long-distance dependencies and the bidirectional information of a character at the same time.
2.4. CRF Layer
When it comes to the context of entity recognition in text, it is always beneficial to consider the correlations between the sequential labels as there are many tagging constraints in natural language sentences. However, the widely used Softmax method predicts the final labels independently, and using Softmax as the top inference layer to extract medical entities will probably break these constraints.
CRF is the most successful model that can control the structure prediction of tagging results. Therefore, CRF was employed to predict the final label sequence in the proposed model. CRF is a probabilistic framework and is usually adopted for sequential data including text [
45]. The basic idea of CRF is to use a series of potential functions to estimate the conditional probability of the output label sequence given the input sequence. More specifically, CRF uses an undirected graphical model to calculate the conditional probability
of a label sequence
given an input sequence
, where
denotes the parameters in the model.
denotes the feature vector and
is the cumulative sum of
over all the possible
:
The model is trained over a given training set
, by maximizing the conditional likelihood:
For the input sequence
and the trained parameters
, the final prediction of a trained CRF is the label sequence
that maximizes the model:
CRF predicts the optimal sequence of labels using a Viterbi algorithm for the input sequence. In our model, the final output of the entity recognition task imposes several hard constraints; for example, “I-cure” cannot follow “B-prob”. The CRF layer considers the interactions between successive labels and can automatically learn these constraints from training data to ensure the validity of the final entity tagging results.
5. Discussion
In spite of the comparatively weak performance in terms of precision, the experimental results reveal the substantial ability of our proposed model over existing approaches in medical entity extraction, indicating the advantage of our designed model. The further character representation evaluation implies that using pretrained embeddings based on the domain corpus can dramatically improve the performance of medical entity recognition over randomly initialized ones, and incorporating position and POS features can further improve the overall performance. The evaluation also suggests the advantage of character-based methods over word-based methods in the Chinese social media context. By the inclusion of context word embedding with character embedding as the representation of the text input, our model can effectively extract medical-related entities in Chinese OHCs without complex feature engineering.
The Bidirectional LSTM architecture is capable of learning long-term dependencies from both forward and backward directions to capture further context features, which results in better performance of the DNN model over traditional machine learning models in terms of overall recall and F-measure. The inclusion of context word embedding with the character embedding can partly capture the context information and avoid use of the same character embedding vector in varied contexts. Therefore, a character with the same location tag might be assigned a different representation vector, while a character in a different context would be represented with the same embedding in CWME; this could be the reason why CNMER generally outperforms the CWME approach. The overall higher recall over traditional machine learning approaches and better F-measure over the three baseline models demonstrates that our model is more appropriate for medical entity extraction in online medical consultations.
In the setting of online consultations, physicians need to process abundant unstructured text information, among which medical entities are the most critical part for efficient health assistance. The rapid development of information and communication technology in recent years has greatly changed the manner of health service delivery in modern society. Compared with real-world face-to-face visits, e-mediated patient–doctor communication has certain unique characteristics that touch the critical components of the relationship between patients and physicians [
56] and would potentially affect the sustainability and effectiveness of online communities.
Confirmation bias means that one is more inclined to the evidence that supports their existing beliefs, expectations, and hypothesis in hand [
57]. In online health consultations, users are anonymous to physicians and the user-generated content is the only cue for physicians to infer the health conditions of patients. With the limited information available in online consultations, physicians need to evaluate the health conditions of patients and make professional medical suggestions. However, adequate and accurate evidence are required in medical services. During this process, confirmation bias could occur in two possible ways. First, before reporting their conditions to healthcare professionals online, some patients may have their own prior judgement for the medical problem and thus unconsciously describe their conditions with bias. Second, due to the limited information available, even the most seasoned healthcare practitioner can be prejudiced occasionally and be led to misdiagnose a problem by confirmation bias [
58]. CNMER has been proven effective in extracting health-related concepts from Chinese OHCs, and these medical concepts are essential components for health professionals to provide feedback. In the context of online medical consultations, the principal contents submitted by users are highlighted with the extracted medical entities. Users can check and edit what they have written, and physicians can effectively examine the posts without ignoring the critical information in the text, which could potentially help to alleviate the effect of confirmation bias.
Trust is another critical concern in e-mediated patient–doctor interaction. The first and foremost function of trust is to reduce complexity [
59]. Trust has been shown to affect a host of behaviors including patients’ willingness to seek care, reveal sensitive information, and remain with a physician [
60]. In the context of e-mediated communication, patients are anonymous to healthcare service providers, which further highlights the importance of trust. Patients’ trust in their doctor and doctors’ trust in their patients during online consultations play an essential role in dealing with the health issues of patients. For patients that seek online medical support, trust in their doctors can help to sustain well-being when coping with health risks. The continuous trust between patients and physicians in online heath consultations is one of the key critical elements that ensure the sustainability of online healthcare service delivery. The higher level and status of a healthcare system have proven to be associated with more trust [
61]. The extracted medical concepts facilitate efficient information processing and boost the information exchange in online consultations, improving the relationship between patient and doctor. Patient–doctor communication is more than transferring information about medical conditions from patient to doctor and medical knowledge from doctor to patient: it is about releasing the patient’s feelings of stress, anxiety, and risk in health issues [
56]. The significant positive relationship between trust and the perceived value of social interaction has been reported in a previous study [
62]. An efficient, intelligent healthcare system employing an online platform can thus improve the trust between patients and doctors as the social exchange is perceived to be beneficial.
Despite the wide use of health insurance and other related programs, economic or time cost is usually inevitable for most healthcare consumers when dealing with their health problems. Healthcare consumers tend to reduce the cost without impairing the quality of care; they evaluate the return and corresponding cost of different healthcare services and make decisions according to their related knowledge and experience. Chronic diseases such as diabetes, cancer, cardiovascular disease, and chronic respiratory diseases have been a substantial economic burden for patients due to expenditure on long-term medical care, especially for those from low-income and middle-income countries such as China [
63]. The introduction of online healthcare services provides users with alternative options to cope with these health concerns. OHCs have been reported as powerful platforms for chronic disease patients to tackle some of the challenges, with certain advantages including the exchange of medical knowledge, supporting self-management, and improving patient-centered care [
64]. Online healthcare platforms not only provide modern patients an open communication channel with their physicians, but also facilitate patients gaining control over their lives and improving the quality of care by self-management [
64]. While the sensitivity to cost of healthcare services varies [
65], individuals can seek medical support with minimal time and cost restrictions in OHCs.
Health information technology has been widely adopted in recent years due to its capacity to improve the cost savings, efficiency, quality, and safety of medical service delivery. Among all the components, cost remains the primary barrier that impedes the adoption of health information technology [
66], and cost–benefit analysis of healthcare system adoption is meaningful. For the proposed health system designed for OHCs, online platforms can employ the system on their websites, and both patients and doctors can utilize the system to enhance the healthcare service delivery. As stated before, the engagement of the intelligent system can benefit the OHCs by attracting more doctors and patients to participant in healthcare information exchanges due to its advantages including diminishing confirmation bias, gaining trust, and reducing cost. Despite the potentially high cost of DNN systems at the moment, the rapid development of deep learning technologies and the booming of related web services make the system more applicable. It is economically feasible for online healthcare platforms to employ the system as further considerable benefits are expected.
The utilization of DNN in our model achieved more promising performance over conventional machine learning methods. From the perspective of practical implementation, DNN systems are known for their lack of transparency, and the prediction results are tough to explain. Consequently, there are concerns regarding the safety issue of employing the system as DNN-based models remain mysterious to their users. However, as our system are designed for extracting medical entities to facilitate information processing rather than providing health-related professional suggestions, the transparency of our model and the explanation of results are actually dispensable in real-world applications.
The sustainable employment of the proposed DNN-based healthcare system by online health platforms relies on the continuous benefits that are obtained from the system. The powerful capacity to extract medical concepts from the healthcare system can moderately improve the quality of information transmission between patient and doctors in OHCs, which reduces economic and time cost and enhances quality of life [
67,
68]. According to the medical advice provided by health professionals, users can cope with their health issues more properly and thus decrease their medical expenditures. The effective health-related information seeking powered by the proposed model can also minimize patients’ future health risks by reducing medical uncertainty. The sustainable development of OHCs depends on the participation of health professionals, and doctors can gain social and economic returns by participating in OHCs [
69]. For healthcare service providers, the system can assist them to improve the efficiency of medical information processing with higher accuracy. As increasing numbers of participants engage with and benefit from the system, OHCs can gain more profit and thus invest more in the development of intelligent healthcare systems, which in turn attracts more participants to the platforms. Medical concept discovery is the basis of healthcare knowledge discovery strategies such as disease surveillance and adverse drug reaction detection. Healthcare knowledge discovery from social media has been validated as viable in previous works [
15,
16], and can contribute to the sustainability of public health. Therefore, the adoption of the proposed system can directly or indirectly benefit various participants including health consumers, health service providers, and online healthcare platforms, contributing to the sustainability of the virtualized healthcare industry.
6. Conclusions
Our study contributes to the literature mainly in terms of the following points. Firstly, this work designs an effective DNN model that can automatically learn context features of text to replace complex and time-consuming handcrafted feature engineering work. The evaluation results demonstrate that the proposed model considerably outperforms traditional machine learning approaches and a strong DNN baseline model. Second, this paper investigates the medical entity extraction task in the context of Chinese social media, while prior research primarily focused on the English language context. Considering the uniqueness of health-related Chinese social media text, this study proposes concatenating character embedding with context word embedding, together with position and a POS features vector, to enhance the feature representation of characters in Chinese online medical text. As far as we know, this research is among the first works to focus on medical-related entity recognition in Chinese social media. Third, based on a large domain text corpus collected from a well-known Chinese OHC, this work builds a word embedding dataset and a character embedding dataset in the context of Chinese medical-related social media, which are available to the public online [
48]. The learned distributed representations of words and characters capture both syntactic and semantic features, and can facilitate learning algorithms to achieve more promising performance in many NLP-related tasks, including sentiment analysis [
70], text classification [
71], and recommendation [
72].
Previous studies have certain limitations when applied to the context of Chinese health-related social media. This study designed a BiLSTM-CRF-based model named CNMER to extract medical-related entities from Chinese OHCs. The model utilizes character embedding, word embedding, position, and POS feature vectors as the character representation and avoids tough feature engineering work. Despite the relatively unsatisfying results in terms of precision compared with the CRF-based methods, the proposed CNMER approach attained statistically better performance in terms of recall and F-measure over all three baseline models including a strong DNN model, which indicates that our model is more effective in extracting health-related entities from Chinese OHCs. The advantages of using characters as the basic tag units are also validated in this study. The proposed medical entity extraction system contributes to the sustainable development of virtualized healthcare as it benefits many stakeholders including health consumers, health service providers, and online healthcare platforms.
Besides the above achievements, the designed model has certain limitations. First, we only considered the recognition of three main types of medical-related concepts; other entity types such as body part, medical department, and time which are also essential for medical decision support were not investigated in this study. Second, only the focal character and word were considered when constructing a representation vector, while further context characters and words could also contribute to additional performance improvement; this was not explored in our study. Lastly, although the evaluation results indicate that our model outperforms the baseline approaches, the performance is still not satisfying enough for real-world applications. Medical entity extraction in Chinese social media remains a challenging task and deserves further investigation.