Tissue-Based Genomic Testing in Prostate Cancer: 10-Year Analysis of National Trends on the Use of Prolaris, Decipher, ProMark, and Oncotype DX
Abstract
1. Introduction
2. Materials and Methods
2.1. Dataset
2.2. Study Population Statistical Analyses
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hamdy, F.C.; Donovan, J.L.; Lane, J.A.; Metcalfe, C.; Davis, M.; Turner, E.L.; Martin, R.M.; Young, G.J.; Walsh, E.I.; Bryant, R.J.; et al. Fifteen-Year Outcomes after Monitoring, Surgery, or Radiotherapy for Prostate Cancer. N. Engl. J. Med. 2023, 388, 1547–1558. [Google Scholar] [CrossRef]
- Tohi, Y.; Kato, T.; Sugimoto, M. Aggressive Prostate Cancer in Patients Treated with Active Surveillance. Cancers 2023, 15, 4270. [Google Scholar] [CrossRef] [PubMed]
- Schaeffer, E.M.; Srinivas, S.; Adra, N.; An, Y.; Barocas, D.; Bitting, R.; Bryce, A.; Chapin, B.; Cheng, H.H.; D’amico, A.V.; et al. Prostate Cancer, Version 4.2023, NCCN Clinical Practice Guidelines in Oncology. J. Natl. Compr. Cancer Netw. 2023, 21, 1067–1096. [Google Scholar] [CrossRef] [PubMed]
- Mottet, N.; van den Bergh, R.C.N.; Briers, E.; van den Broeck, T.; Cumberbatch, M.G.; De Santis, M.; Fanti, S.; Fossati, N.; Gandaglia, G.; Gillessen, S.; et al. EAU-EANM-ESTRO-ESUR-SIOG Guidelines on Prostate Cancer—2020 Update. Part 1: Screening, Diagnosis, and Local Treatment with Curative Intent. Eur. Urol. 2020, 79, 243–262. [Google Scholar] [CrossRef] [PubMed]
- Cooperberg, M.R.; Meeks, W.; Fang, R.; Gaylis, F.D.; Catalona, W.J.; Makarov, D.V. Time Trends and Variation in the Use of Active Surveillance for Management of Low-risk Prostate Cancer in the US. JAMA Netw. Open 2023, 6, e231439. [Google Scholar] [CrossRef] [PubMed]
- Bolla, M.; van Poppel, H.; Tombal, B.; Vekemans, K.; Da Pozzo, L.; de Reijke, T.M.; Verbaeys, A.; Bosset, J.-F.; van Velthoven, R.; Colombel, M.; et al. Postoperative radiotherapy after radical prostatectomy for high-risk prostate cancer: Long-term results of a randomised controlled trial (EORTC trial 22911). Lancet 2012, 380, 2018–2027. [Google Scholar] [CrossRef] [PubMed]
- Thompson, I.M.; Tangen, C.M.; Paradelo, J.; Lucia, M.S.; Miller, G.; Troyer, D.; Messing, E.; Forman, J.; Chin, J.; Swanson, G.; et al. Adjuvant Radiotherapy for Pathological T3N0M0 Prostate Cancer Significantly Reduces Risk of Metastases and Improves Survival: Long-Term Followup of a Randomized Clinical Trial. J. Urol. 2009, 181, 956–962. [Google Scholar] [CrossRef]
- Kneebone, A.; Fraser-Browne, C.; Duchesne, G.M.; Fisher, R.; Frydenberg, M.; Herschtal, A.; Williams, S.G.; Brown, C.; Delprado, W.; Haworth, A.; et al. Adjuvant radiotherapy versus early salvage radiotherapy following radical prostatectomy (TROG 08.03/ANZUP RAVES): A randomised, controlled, phase 3, non-inferiority trial. Lancet Oncol. 2020, 21, 1331–1340. [Google Scholar] [CrossRef]
- Del Giudice, F.; Huang, J.; Li, S.; Sorensen, S.; Enemchukwu, E.; Maggi, M.; Salciccia, S.; Ferro, M.; Crocetto, F.; Pandolfo, S.D.; et al. Contemporary trends in the surgical management of urinary incontinence after radical prostatectomy in the United States. Prostate Cancer Prostatic Dis. 2022, 26, 367–373. [Google Scholar] [CrossRef]
- Franco, A.; Autorino, R. ExoDx test for prostate cancer: The future is liquid—Editorial Comment. Prostate Cancer Prostatic Dis. 2023, 26, 443–444. [Google Scholar] [CrossRef]
- Di Lorenzo, G.; Autorino, R.; D’Armiento, F.; Mignogna, C.; De Laurentiis, M.; De Sio, M.; D’Armiento, M.; Damiano, R.; Vecchio, G.; De Placido, S. Expression of proto-oncogene c-kit in high risk prostate cancer. Eur. J. Surg. Oncol. (EJSO) 2004, 30, 987–992. [Google Scholar] [CrossRef] [PubMed]
- Moreno, C.S.; Winham, C.L.; Alemozaffar, M.; Klein, E.R.; Lawal, I.O.; Abiodun-Ojo, O.A.; Patil, D.; Barwick, B.G.; Huang, Y.; Schuster, D.M.; et al. Integrated Genomic Analysis of Primary Prostate Tumor Foci and Corresponding Lymph Node Metastases Identifies Mutations and Pathways Associated with Metastasis. Cancers 2023, 15, 5671. [Google Scholar] [CrossRef] [PubMed]
- Klein, E.A.; Cooperberg, M.R.; Magi-Galluzzi, C.; Simko, J.P.; Falzarano, S.M.; Maddala, T.; Chan, J.M.; Li, J.; Cowan, J.E.; Tsiatis, A.C.; et al. A 17-gene Assay to Predict Prostate Cancer Aggressiveness in the Context of Gleason Grade Heterogeneity, Tumor Multifocality, and Biopsy Undersampling. Eur. Urol. 2014, 66, 550–560. [Google Scholar] [CrossRef] [PubMed]
- Erho, N.; Crisan, A.; Vergara, I.A.; Mitra, A.P.; Ghadessi, M.; Buerki, C.; Bergstralh, E.J.; Kollmeyer, T.; Fink, S.; Haddad, Z.; et al. Discovery and Validation of a Prostate Cancer Genomic Classifier that Predicts Early Metastasis Following Radical Prostatectomy. PLoS ONE 2013, 8, e66855. [Google Scholar] [CrossRef] [PubMed]
- Cuzick, J.; Swanson, G.P.; Fisher, G.; Brothman, A.R.; Berney, D.M.; Reid, J.E.; Mesher, D.; Speights, V.O.; Stankiewicz, E.; Foster, C.S.; et al. Prognostic value of an RNA expression signature derived from cell cycle proliferation genes in patients with prostate cancer: A retrospective study. Lancet Oncol. 2011, 12, 245–255. [Google Scholar] [CrossRef] [PubMed]
- Blume-Jensen, P.; Berman, D.M.; Rimm, D.L.; Shipitsin, M.; Putzi, M.; Nifong, T.P.; Small, C.; Choudhury, S.; Capela, T.; Coupal, L.; et al. Development and Clinical Validation of an In Situ Biopsy-Based Multimarker Assay for Risk Stratification in Prostate Cancer. Clin. Cancer Res. 2015, 21, 2591–2600. [Google Scholar] [CrossRef]
- Shipitsin, M.; E Small, C.; Choudhury, S.; Giladi, E.; Friedlander, S.F.; Nardone, J.; Hussain, S.; Hurley, A.D.; Ernst, C.; Huang, Y.E.; et al. Identification of proteomic biomarkers predicting prostate cancer aggressiveness and lethality despite biopsy-sampling error. Br. J. Cancer 2014, 111, 1201–1212. [Google Scholar] [CrossRef]
- Research Capabilities. Available online: https://pearldiverinc.com/researchinfo.html (accessed on 30 July 2023).
- Alluri, R.K.; Leland, H.; Heckmann, N. Surgical research using national databases. Ann. Transl. Med. 2016, 4, 393. [Google Scholar] [CrossRef]
- Schaeffer, E.M.; Srinivas, S.; Adra, N.; An, Y.; Barocas, D.; Bitting, R.; Bryce, A.; Chapin, B.; Cheng, H.H.; D’Amico, A.V.; et al. Prostate Cancer, Version 1.2023 Featured Updates to the NCCN Guidelines. Natl. Compr. Cancer Netw. 2022, 20, 1288–1298. [Google Scholar] [CrossRef]
- Chung, J.-S.; Morgan, T.M.; Hong, S.K. Clinical implications of genomic evaluations for prostate cancer risk stratification, screening, and treatment: A narrative review. Prostate Int. 2020, 8, 99–106. [Google Scholar] [CrossRef]
- Peabody, J.W.; DeMaria, L.M.; Tamondong-Lachica, D.; Florentino, J.; Acelajado, M.C.; Ouenes, O.; Richie, J.P.; Burgon, T. Impact of a protein-based assay that predicts prostate cancer aggressiveness on urologists’ recommendations for active treatment or active surveillance: A randomized clinical utility trial. BMC Urol. 2017, 17, 51. [Google Scholar] [CrossRef] [PubMed]
- Kornberg, Z.; Cooperberg, M.R.; Cowan, J.E.; Chan, J.M.; Shinohara, K.; Simko, J.P.; Tenggara, I.; Carroll, P.R. A 17-Gene Genomic Prostate Score as a Predictor of Adverse Pathology in Men on Active Surveillance. J. Urol. 2019, 202, 702–709. [Google Scholar] [CrossRef] [PubMed]
- Lin, D.W.; Zheng, Y.; McKenney, J.K.; Brown, M.D.; Lu, R.; Crager, M.; Boyer, H.; Tretiakova, M.; Brooks, J.D.; Dash, A.; et al. 17-Gene Genomic Prostate Score Test Results in the Canary Prostate Active Surveillance Study (PASS) Cohort. J. Clin. Oncol. 2020, 38, 1549–1557. [Google Scholar] [CrossRef]
- Eggener, S.; Karsh, L.I.; Richardson, T.; Shindel, A.W.; Lu, R.; Rosenberg, S.; Goldfischer, E.; Korman, H.; Bennett, J.; Newmark, J.; et al. A 17-gene Panel for Prediction of Adverse Prostate Cancer Pathologic Features: Prospective Clinical Validation and Utility. Urology 2019, 126, 76–82. [Google Scholar] [CrossRef]
- Murphy, A.B.; Abern, M.R.; Liu, L.; Wang, H.; Hollowell, C.M.P.; Sharifi, R.; Vidal, P.; Kajdacsy-Balla, A.; Sekosan, M.; Ferrer, K.; et al. Impact of a Genomic Test on Treatment Decision in a Predominantly African American Population With Favorable-Risk Prostate Cancer: A Randomized Trial. J. Clin. Oncol. 2021, 39, 1660–1670. [Google Scholar] [CrossRef] [PubMed]
- Eymech, O.; Brunckhorst, O.; Fox, L.; Jawaid, A.; Van Hemelrijck, M.; Stewart, R.; Dasgupta, P.; Ahmed, K. An exploration of wellbeing in men diagnosed with prostate cancer undergoing active surveillance: A qualitative study. Support. Care Cancer 2022, 30, 5459–5468. [Google Scholar] [CrossRef] [PubMed]
- Eure, G.; Germany, R.; Given, R.; Lu, R.; Shindel, A.W.; Rothney, M.; Glowacki, R.; Henderson, J.; Richardson, T.; Goldfischer, E.; et al. Use of a 17-Gene Prognostic Assay in Contemporary Urologic Practice: Results of an Interim Analysis in an Observational Cohort. Urology 2017, 107, 67–75. [Google Scholar] [CrossRef]
- Badani, K.K.; Kemeter, M.J.; Febbo, P.G.; Lawrence, H.J.; Denes, B.S.; Rothney, M.P.; Rothberg, M.B.; Brown, G.A. The Impact of a Biopsy Based 17-Gene Genomic Prostate Score on Treatment Recommendations in Men with Newly Diagnosed Clinically Prostate Cancer Who are Candidates for Active Surveillance. Urol. Pract. 2015, 2, 181–189. [Google Scholar] [CrossRef]
- Herlemann, A.; Huang, H.-C.; Alam, R.; Tosoian, J.J.; Kim, H.L.; Klein, E.A.; Simko, J.P.; Chan, J.M.; Lane, B.R.; Davis, J.W.; et al. Decipher identifies men with otherwise clinically favorable-intermediate risk disease who may not be good candidates for active surveillance. Prostate Cancer Prostatic Dis. 2019, 23, 136–143. [Google Scholar] [CrossRef]
- Tosoian, J.J.; Chappidi, M.R.; Bishoff, J.T.; Freedland, S.J.; Reid, J.; Brawer, M.; Stone, S.; Schlomm, T.; Ross, A.E. Prognostic utility of biopsy-derived cell cycle progression score in patients with National Comprehensive Cancer Network low-risk prostate cancer undergoing radical prostatectomy: Implications for treatment guidance. BJU Int. 2017, 120, 808–814. [Google Scholar] [CrossRef]
- Jo, J.K.; Hong, S.K.; Byun, S.S.; Zargar, H.; Autorino, R.; Lee, S.E. Positive surgical margin in robot-assisted radical prostatec-tomy: Correlation with pathology findings and risk of biochemical recurrence. Minerva Urol. Nephrol. 2017, 69, 493–500. [Google Scholar] [CrossRef] [PubMed]
- Parker, C.C.; Clarke, N.W.; Cook, A.D.; Kynaston, H.G.; Petersen, P.M.; Catton, C.; Cross, W.; Logue, J.; Parulekar, W.; Payne, H.; et al. Timing of radiotherapy after radical prostatectomy (RADICALS-RT): A randomised, controlled phase 3 trial. Lancet 2020, 396, 1413–1421. [Google Scholar] [CrossRef] [PubMed]
- Morgan, T.M.; Hawken, S.R.; Ghani, K.R.; Miller, D.C.; Feng, F.Y.; Linsell, S.M.; A Salisz, J.; Gao, Y.; E Montie, J.; Cher, M.L.; et al. Variation in the use of postoperative radiotherapy among high-risk patients following radical prostatectomy. Prostate Cancer Prostatic Dis. 2016, 19, 216–221. [Google Scholar] [CrossRef] [PubMed]
- Suardi, N.; Gallina, A.; Lista, G.; Gandaglia, G.; Abdollah, F.; Capitanio, U.; Dell’oglio, P.; Nini, A.; Salonia, A.; Montorsi, F.; et al. Impact of Adjuvant Radiation Therapy on Urinary Continence Recovery After Radical Prostatectomy. Eur. Urol. 2013, 65, 546–551. [Google Scholar] [CrossRef]
- Pra, A.D.; Ghadjar, P.; Hayoz, S.; Liu, V.; Spratt, D.; Thompson, D.; Davicioni, E.; Huang, H.-C.; Zhao, X.; Liu, Y.; et al. Validation of the Decipher genomic classifier in patients receiving salvage radiotherapy without hormone therapy after radical prostatectomy—An ancillary study of the SAKK 09/10 randomized clinical trial. Ann. Oncol. 2022, 33, 950–958. [Google Scholar] [CrossRef]
- Spratt, D.E.; Yousefi, K.; Deheshi, S.; Ross, A.E.; Den, R.B.; Schaeffer, E.M.; Trock, B.J.; Zhang, J.; Glass, A.G.; Dicker, A.P.; et al. Individual Patient-Level Meta-Analysis of the Performance of the Decipher Genomic Classifier in High-Risk Men After Prostatectomy to Predict Development of Metastatic Disease. J. Clin. Oncol. 2017, 35, 1991–1998. [Google Scholar] [CrossRef] [PubMed]
- Dalela, D.; Santiago-Jiménez, M.; Yousefi, K.; Karnes, R.J.; Ross, A.E.; Den, R.B.; Freedland, S.J.; Schaeffer, E.M.; Dicker, A.P.; Menon, M.; et al. Genomic Classifier Augments the Role of Pathological Features in Identifying Optimal Candidates for Adjuvant Radiation Therapy in Patients With Prostate Cancer: Development and Internal Validation of a Multivariable Prognostic Model. J. Clin. Oncol. 2017, 35, 1982–1990. [Google Scholar] [CrossRef]
- NRG Oncology. Parallel Phase III Randomized Trials of Genomic-Risk Stratified Unfavorable Intermediate Risk Prostate Cancer: De-Intensification and Intensification Clinical Trial Evaluation (Guidance). Available online: https://www.nrgoncology.org/Clinical-Trials/Protocol/nrg-gu010-1?filter=nrg-gu010-1 (accessed on 6 January 2024).
- NRG Oncology. Parallel Phase III Randomized Trials for High Risk Prostate Cancer Evaluating De-Intensification for Lower Genomic Risk and Intensification of Concurrent Therapy for Higher Genomic Risk with Radiation (Predict-RT*). Available online: https://www.nrgoncology.org/Clinical-Trials/Protocol/nrg-gu009-1?filter=nrg-gu009-1 (accessed on 6 January 2024).
- Cuzick, J.; Berney, D.M.; Fisher, G.; Mesher, D.; Møller, H.; Reid, J.E.; Perry, M.; Park, J.; Younus, A.; Gutin, A.; et al. Prognostic value of a cell cycle progression signature for prostate cancer death in a conservatively managed needle biopsy cohort. Br. J. Cancer 2012, 106, 1095–1099. [Google Scholar] [CrossRef]
- Bishoff, J.T.; Freedland, S.J.; Gerber, L.; Tennstedt, P.; Reid, J.; Welbourn, W.; Graefen, M.; Sangale, Z.; Tikishvili, E.; Park, J.; et al. Prognostic Utility of the Cell Cycle Progression Score Generated from Biopsy in Men Treated with Prostatectomy. J. Urol. 2014, 192, 409–414. [Google Scholar] [CrossRef]
- Saad, F.; Latour, M.; Lattouf, J.-B.; Widmer, H.; Zorn, K.C.; Mes-Masson, A.-M.; Ouellet, V.; Saad, G.; Prakash, A.; Choudhury, S.; et al. Biopsy Based Proteomic Assay Predicts Risk of Biochemical Recurrence after Radical Prostatectomy. J. Urol. 2017, 197, 1034–1040. [Google Scholar] [CrossRef]
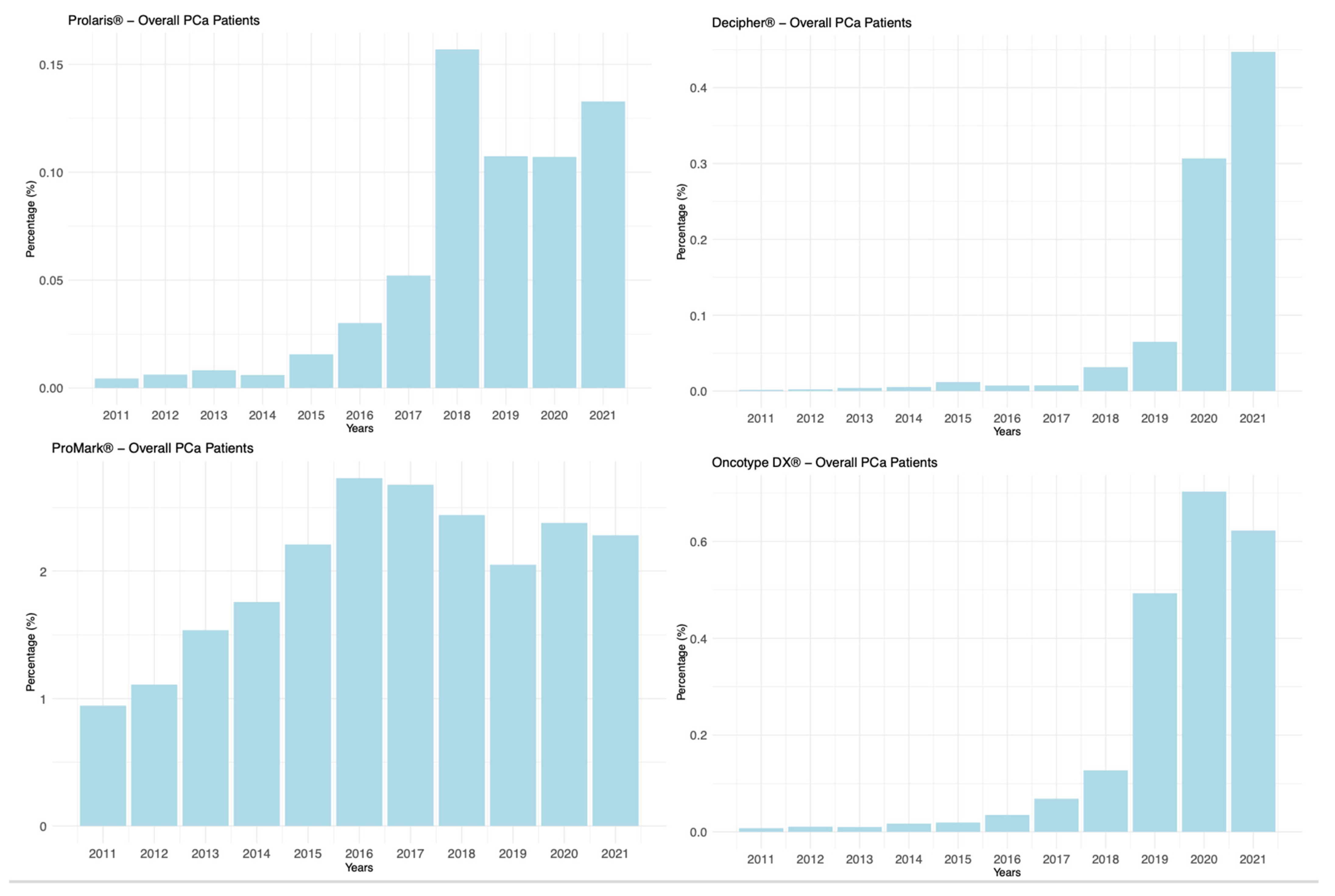
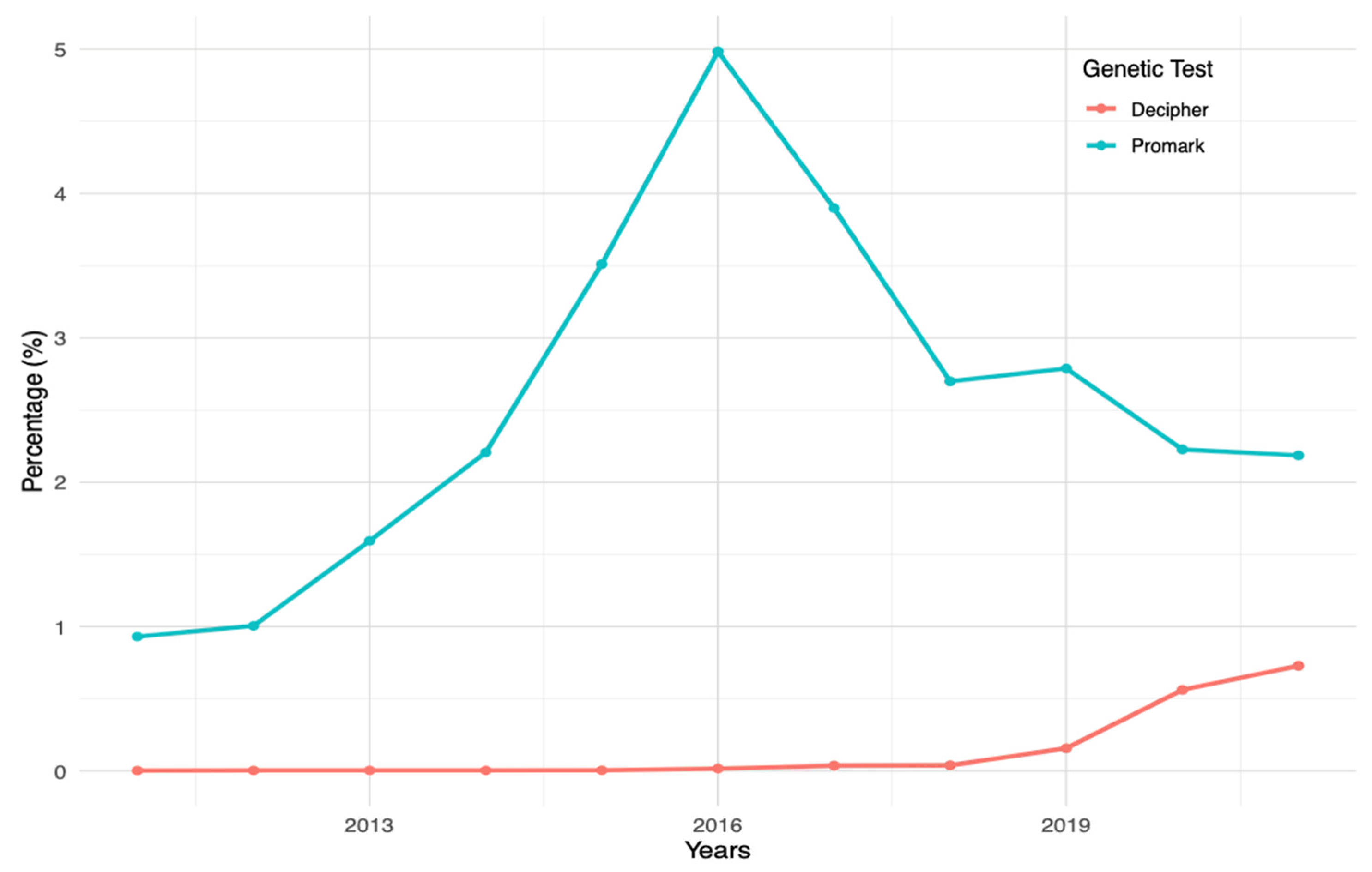
| Variable | Prostate Biopsy Patients (n = 1,561,203) | RP Patients (n = 241,445) |
|---|---|---|
| Age, years, Mean ± SD | 68.51 ± 8.33 | 64.39 ± 7.60 |
| Region, n (%) | ||
| Midwest | 352,299 (22.5) | 63,632 (26.3) |
| Northeast | 362,326 (23.2) | 49,380 (20.4) |
| South | 620,196 (39.7) | 92,368 (38.3) |
| West | 220,271 (14.2) | 35,127 (14.5) |
| Unknown | 6111 (0.4) | 938 (0.5) |
| Charlson Comorbidity Index, Mean ± SD | 2.79 ± 2.64 | 2.85 ± 2.57 |
| Obesity, n (%) | 473,656 (30.3) | 82,082 (33.9) |
| Diabetes, n (%) | 411,863 (26.4) | 54,763 (22.7) |
| Social Determinants of Health (SDOH), n (%) | ||
| Lack of Education Access and Quality | 460 (0.03) | 62 (0.03) |
| Inadequate Health Care Access and Quality | 241 (0.02) | 22 (0.01) |
| Poor Neighborhood and Built Environment | 4305 (0.28) | 747 (0.31) |
| Negative Social and Community Context | 10,209 (0.65) | 1503 (0.62) |
| Economic instability | 5070 (0.32) | 604 (0.25) |
| Overall | 19,451 (1.25) | 2836 (1.17) |
| Use of Tissue-based genomic Testing, n (%) | 20,748 (1.32) | 3076 (1.27) |
| Genetic Test | Regression Coefficient (Slope) | Model Fit (Adjusted R2) | p-Value |
|---|---|---|---|
| Prolaris® | 0.0152 | 0.7397 | 0.0004 |
| Oncotype DX® | 0.0687 | 0.6677 | 0.0013 |
| Decipher® | 0.0334 | 0.4888 | 0.01 |
| Promark® | 0.1375 | 0.5248 | 0.007 |
| Genetic Test | Clinical Indication | Testing Method | Assessed Parameters | Scoring | Clinical Implications | Other Characteristics |
|---|---|---|---|---|---|---|
| Prolaris® | After biopsy: NCCN very low low, favorable intermediate-risk localized prostate cancer After RP: patients who may benefit from aggressive intervention/at high risk of recurrence | Reverse transcriptase PCR | Gene activity related to cell cycle: 46 genes (31 Cell Cycle Progression + 15 housekeeping genes) | Cell Cycle Progression (CCP) score between 0 and 10 Higher scores indicative of more aggressive disease | Provides risk assessment to aid treatment choice between AS, single modal or multi-modal treatment Provides:
| Result combined with patient’s clinical data (CAPRA score and NCCN) |
| Decipher® | After biopsy: all GS, all PSA values, all Stages After RP: patients with adverse pathology, all PSA values (including undetectable, rising, and persistently elevated PSA) | Microarray genomic testing | Expression of 22 coding and noncoding RNAs | Genomic Risk (GR) Score between 0 and 1 Higher scores indicative of more aggressive disease | After biopsy: High risk (>0.6): patients may benefit from treatment intensification with multimodal therapy Low risk (<0.45): patients can be candidates for AS Provides:
High risk (>0.6): patients may benefit from RT with concurrent ADT; patients may benefit from earlier, more intense, or multimodality therapy, and may consider clinical trials of novel therapies Provides:
| Result not combined with other clinical or pathologic parameters Additional information: After biopsy: Personalized risk of metastasis if combined with patient’s NCCN risk category After RP:
|
| ProMark® | NCCN very low, low and intermediate risk localized prostate cancer | Proteomic analysis | Quantify the values of 8 tumor progression-related biomarker proteins | ProMark Score between 0 and 100 Higher scores indicative of more aggressive disease | Provides risk assessment to aid treatment choice between AS and active treatment Predicts BCR in patients after RP Provides:
| Result not combined with other clinical or diagnostic data (NCCN, CAPRA, D’Amico) Additional information: Likelihood (%) of Adverse Pathology at RP Personalized risk of aggressive disease if combined with patient’s NCCN risk category |
| Oncotype DX® | NCCN low, intermediate, and high-risk localized prostate | Reverse transcriptase PCR | Expression of 17 genes (12 cancer-related and 5 reference genes) | Genomic Prostate Score (GPS) between 0 and 100 Higher scores indicative of more aggressive disease | Low risk patients: help inform the AS decision High Risk patients: help inform the treatment intensity decision Provides: Low risk patients: Likelihood (%) of Adverse Pathology at RP High Risk patients: Likelihood (lower\higher) of disease progression | Result combined to NCCN risk group Additional information:
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bologna, E.; Ditonno, F.; Licari, L.C.; Franco, A.; Manfredi, C.; Mossack, S.; Pandolfo, S.D.; De Nunzio, C.; Simone, G.; Leonardo, C.; et al. Tissue-Based Genomic Testing in Prostate Cancer: 10-Year Analysis of National Trends on the Use of Prolaris, Decipher, ProMark, and Oncotype DX. Clin. Pract. 2024, 14, 508-520. https://doi.org/10.3390/clinpract14020039
Bologna E, Ditonno F, Licari LC, Franco A, Manfredi C, Mossack S, Pandolfo SD, De Nunzio C, Simone G, Leonardo C, et al. Tissue-Based Genomic Testing in Prostate Cancer: 10-Year Analysis of National Trends on the Use of Prolaris, Decipher, ProMark, and Oncotype DX. Clinics and Practice. 2024; 14(2):508-520. https://doi.org/10.3390/clinpract14020039
Chicago/Turabian StyleBologna, Eugenio, Francesco Ditonno, Leslie Claire Licari, Antonio Franco, Celeste Manfredi, Spencer Mossack, Savio Domenico Pandolfo, Cosimo De Nunzio, Giuseppe Simone, Costantino Leonardo, and et al. 2024. "Tissue-Based Genomic Testing in Prostate Cancer: 10-Year Analysis of National Trends on the Use of Prolaris, Decipher, ProMark, and Oncotype DX" Clinics and Practice 14, no. 2: 508-520. https://doi.org/10.3390/clinpract14020039
APA StyleBologna, E., Ditonno, F., Licari, L. C., Franco, A., Manfredi, C., Mossack, S., Pandolfo, S. D., De Nunzio, C., Simone, G., Leonardo, C., & Franco, G. (2024). Tissue-Based Genomic Testing in Prostate Cancer: 10-Year Analysis of National Trends on the Use of Prolaris, Decipher, ProMark, and Oncotype DX. Clinics and Practice, 14(2), 508-520. https://doi.org/10.3390/clinpract14020039









