Microbial Cell Factories: Biodiversity, Pathway Construction, Robustness, and Industrial Applicability
Abstract
1. Introduction
2. Biodiversity of Microbial Cell Factories
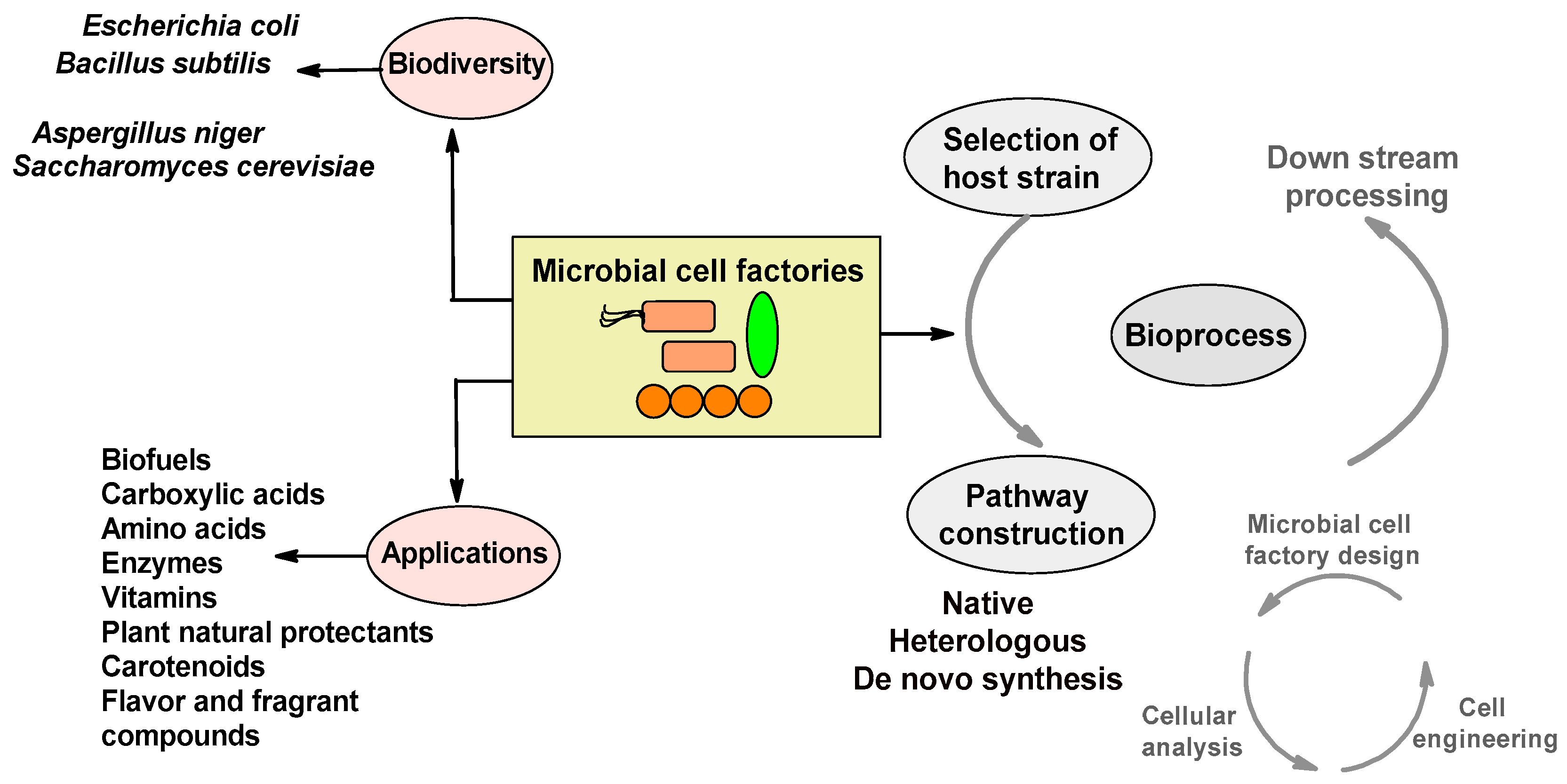
3. Designing Microbial Cell Factories
3.1. Microbial Strain Selection
3.2. Pathway Construction: Native, Heterologous, and Artificial De Novo
3.2.1. Native Pathway
3.2.2. Heterologous Pathway
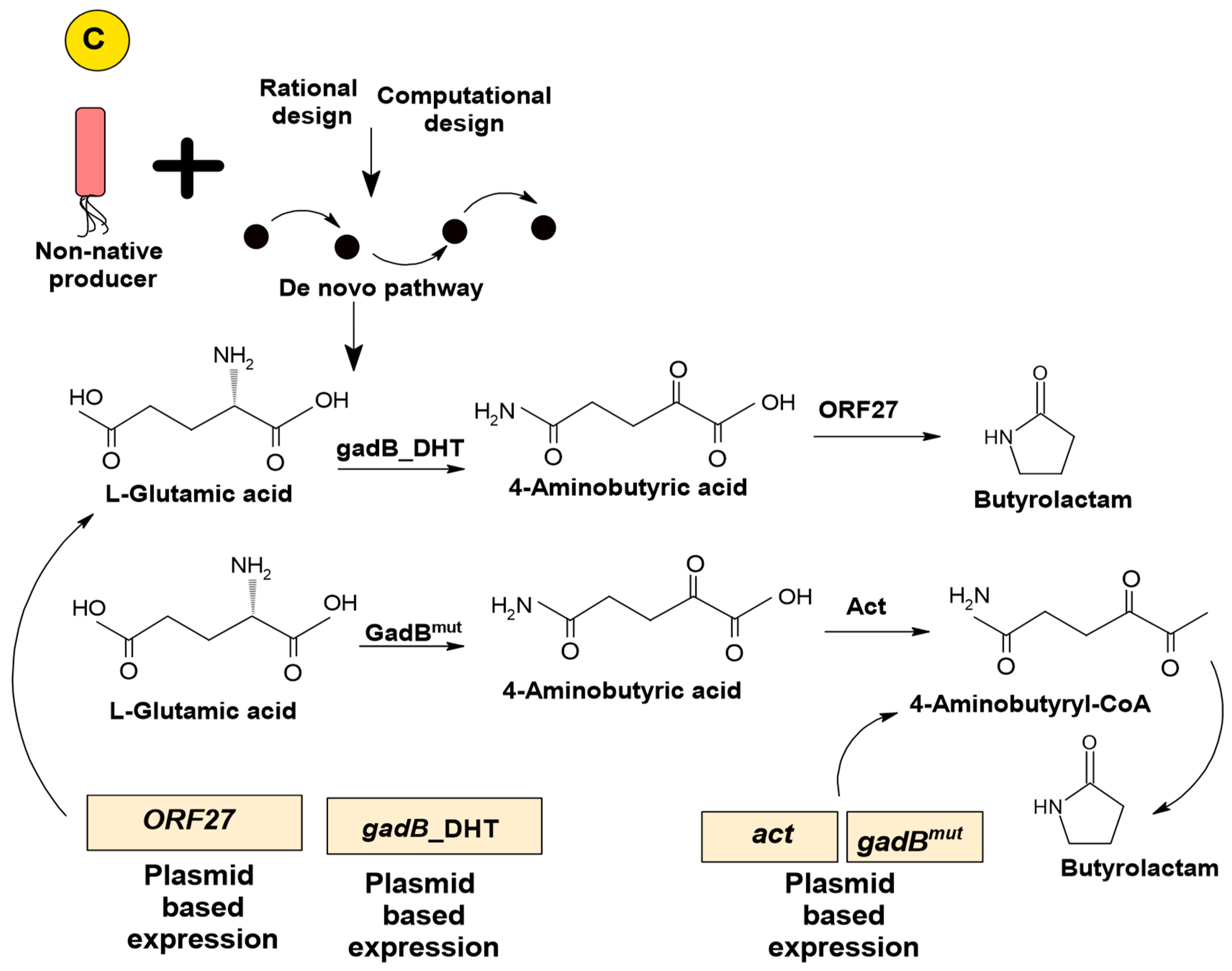
3.2.3. Artificial De Novo Pathway
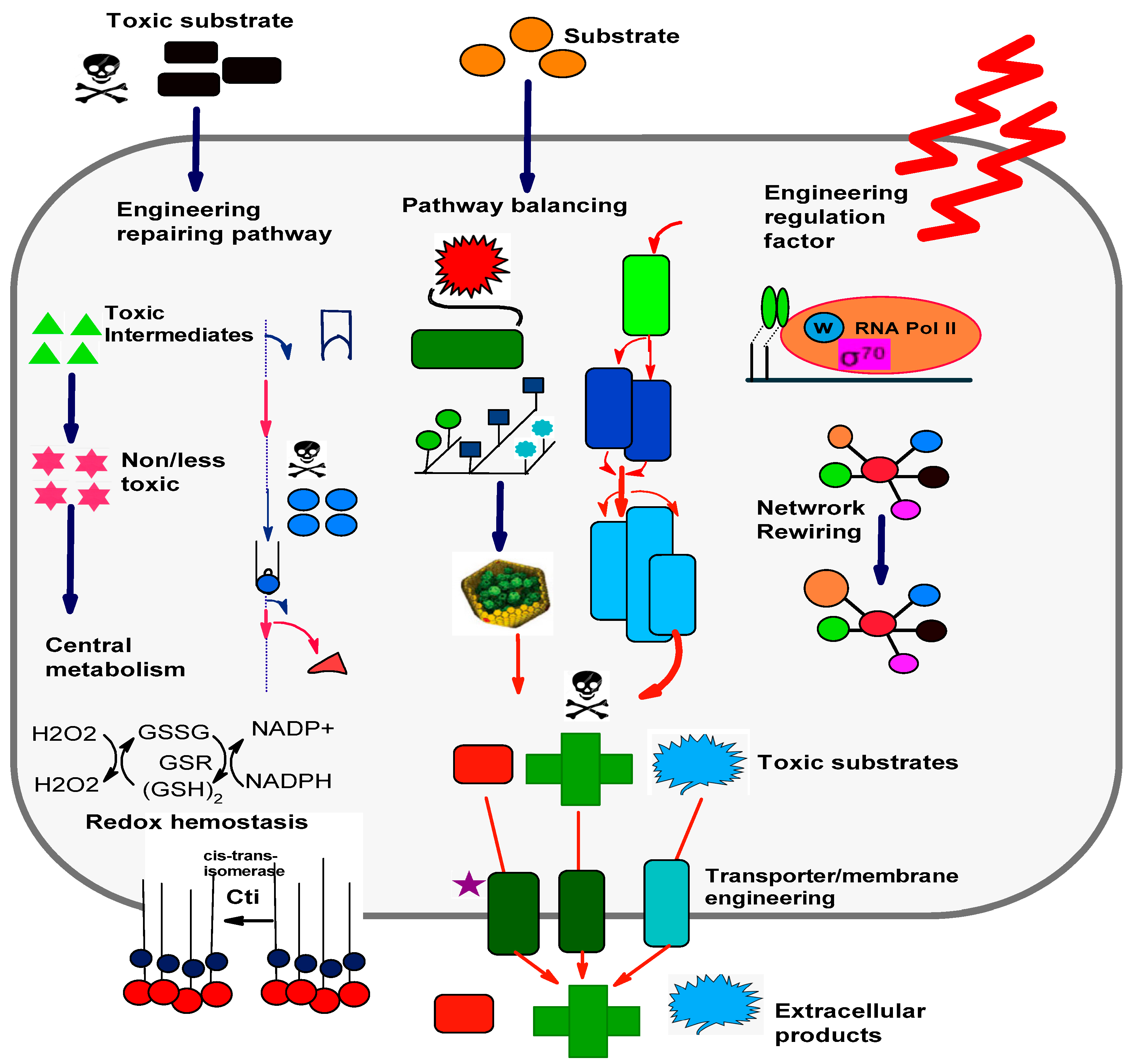
4. Robustness of Microbial Cell Factories
5. Application of Microbial Cell Factories
5.1. Carboxylic Acids
5.2. Vitamins
5.3. Amino Acids
5.4. Plant Natural Products
5.5. Carotenoids
5.6. Flavors and Fragrances
5.7. Bioenergy
6. Industrial Progress, Commercial Limitations, and Future Prospects
Author Contributions
Funding
Conflicts of Interest
References
- Wilbanks, T.J.; Fernandez, S. Climate Change and Infrastructure, Urban Systems, and Vulnerabilities: Technical Report for the US Department of Energy in Support of the National Climate Assessment; Island Press: Washington, DC, USA, 2014. [Google Scholar]
- Shi, T.Q.; Darvishi, F.; Cao, M.; Ji, B.; Ji, X.J. Design and construction of microbial cell factories for the production of fuels and chemicals. Front. Bioeng. Biotechnol. 2023, 11, 1198317. [Google Scholar] [CrossRef] [PubMed]
- Becker, J.; Wittmann, C. Advanced biotechnology: Metabolically engineered cells for the bio-based production of chemicals and fuels, materials, and health-care products. Angew. Chem. Int. Ed. 2015, 54, 3328–3350. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.L.; Zhou, J.J.; Quan, C.S.; Xiu, Z.L. Advances in industrial microbiome based on microbial consortium for biorefinery. Bioresour. Biproc. 2017, 4, 11. [Google Scholar] [CrossRef]
- Zuroff, T.R.; Curtis, W.R. Developing symbiotic consortia for lignocellulosic biofuel production. Appl. Microbiol. Biotechnol. 2012, 93, 1423–1435. [Google Scholar] [CrossRef]
- He, Y.; Xie, K.; Xu, P.; Huang, X.; Gu, W.; Zhang, F.; Tang, S. Evolution of microbial community diversity and enzymatic activity during composting. Res. Microbiol. 2013, 164, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Demain, A.L. Importance of microbial natural products and the need to revitalize their discovery. J. Ind. Microbiol. Biotechnol. 2014, 41, 185–201. [Google Scholar] [CrossRef]
- Abedi, E.; Hashemi, S.M.B. Lactic acid production–producing microorganisms and substrates sources-state of art. Heliyon 2020, 6, 04974. [Google Scholar] [CrossRef]
- Piwowarek, K.; Lipińska, E.; Hać-Szymańczuk, E.; Kieliszek, M.; Ścibisz, I. Propionibacterium spp.—Source of propionic acid, vitamin B12, and other metabolites important for the industry. Appl. Microbiol. Biotechnol. 2018, 102, 515–538. [Google Scholar] [CrossRef]
- Chubukov, V.; Mukhopadhyay, A.; Petzold, C.J.; Keasling, J.D.; Martín, H.G. Synthetic and systems biology for microbial production of commodity chemicals. NPJ Syst. Biol. Appl. 2016, 2, 16009. [Google Scholar] [CrossRef]
- Fisher, A.K.; Freedman, B.G.; Bevan, D.R.; Senger, R.S. A review of metabolic and enzymatic engineering strategies for designing and optimizing performance of microbial cell factories. Comput. Struct. Biotechnol. J. 2014, 11, 91–99. [Google Scholar] [CrossRef]
- Novo Nordisk Global. Available online: https://www.novonordisk.com/ (accessed on 4 January 2024).
- DSM. Available online: https://www.dsm.com/corporate/home.html (accessed on 13 December 2023).
- Cargill. Available online: https://www.cargill.com/home (accessed on 11 January 2024).
- Navarrete, C.; Jacobsen, I.H.; Martínez, J.L.; Procentese, A. Cell factories for industrial production processes: Current issues and emerging solutions. Processes 2020, 8, 768. [Google Scholar] [CrossRef]
- Locey, K.J.; Lennon, J.T. Scaling laws predict global microbial diversity. Proc. Natl. Acad. Sci. USA 2016, 113, 5970–5975. [Google Scholar] [CrossRef]
- Petrova, P.; Petrov, K. Lactic acid fermentation of cereals and pseudocereals: Ancient nutritional biotechnologies with modern applications. Nutrients 2020, 12, 1118. [Google Scholar] [CrossRef]
- Wang, Y.; Wu, J.; Lv, M.; Shao, Z.; Hungwe, M.; Wang, J.; Bai, X.; Xie, J.; Wang, Y.; Geng, W. Metabolism characteristics of lactic acid bacteria and the expanding applications in food industry. Front. Bioeng. Biotechnol. 2021, 9, 612285. [Google Scholar] [CrossRef] [PubMed]
- Meruvu, H.; Harsa, S.T. Lactic acid bacteria: Isolation–characterization approaches and industrial applications. Crit. Rev. Food Sci. Nutr. 2022, 63, 8337–8356. [Google Scholar] [CrossRef] [PubMed]
- Satyanarayana, T.; Kunze, G. Yeast Biotechnology: Diversity and Applications; Springer Science: Berlin/Heidelberg, Germany, 2009; Volume 78. [Google Scholar]
- Abdel-Azeem, A.M.; Abdel-Azeem, M.A.; Abdul-Hadi, S.Y.; Darwish, A.G. Aspergillus: Biodiversity, ecological significances, and industrial applications. In Recent Advancement in White Biotechnology through Fungi; Springer: Cham, Switzerland, 2019; pp. 121–179. [Google Scholar]
- Baeshen, N.A.; Baeshen, M.N.; Sheikh, A.; Bora, R.S.; Ahmed, M.M.M.; Ramadan, H.A.; Saini, K.S.; Redwan, E.M. Cell factories for insulin production. Microb. Cell Factories 2014, 13, 141. [Google Scholar] [CrossRef]
- Bomgardner, M. Food Ingredients Cargill, DSM start up stevia factory. Chem. Eng. News. 2019, 97, 14–15. [Google Scholar]
- Tolieng, V.; Tanaka, N.; Shiwa, Y.; Thitiprasert, S.; Kanchanasin, P.; Phongsopitanun, W.; Booncharoen, A.; Thongchul, N.; Tanasupawat, S. Weizmannia acidilactici sp. nov., a lactic acid producing bacterium isolated from soils. Syst. Appl. Microbiol. 2023, 46, 126389. [Google Scholar] [CrossRef]
- Romero-Mota, D.I.; Estrada-García, J.; Sales-Pérez, R.E.; Méndez-Contreras, J.M. Valorization of Bovine Manure and Molasses by the Production of Lactic Acid and Biomass through Probiotic Anaerobic Cofermentation with Lactobacillus acidophilus, Lactobacillus fermentum, and Bacillus subtilis. J. Environ. Eng. 2024, 150, 04023100. [Google Scholar] [CrossRef]
- Sansatchanon, K.; Sudying, P.; Promdonkoy, P.; Kingcha, Y.; Visessanguan, W.; Tanapongpipat, S.; Runguphan, W.; Kocharin, K. Development of a Novel D-Lactic Acid Production Platform Based on Lactobacillus saerimneri TBRC 5746. J. Microbiol. 2023, 61, 853–863. [Google Scholar] [CrossRef]
- Tong, Z.; Tong, Y.; Wang, D.; Shi, Y.C. Whole maize flour and isolated maize starch for production of citric acid by Aspergillus niger: A review. Starch-Stärke 2023, 75, 2000014. [Google Scholar] [CrossRef]
- Diamantopoulou, P.; Sarris, D.; Tchakouteu, S.S.; Xenopoulos, E.; Papanikolaou, S. Growth Response of Non-Conventional Yeasts on Sugar-Rich Media: Part 1: High Production of Lipid by Lipomyces starkeyi and Citric Acid by Yarrowia lipolytica. Microorganisms 2023, 11, 1863. [Google Scholar] [CrossRef] [PubMed]
- Behera, B.C.; Mishra, R.; Mohapatra, S. Microbial citric acid: Production, properties, application, and future perspectives. Food Front. 2021, 2, 62–76. [Google Scholar] [CrossRef]
- Saichana, N.; Matsushita, K.; Adachi, O.; Frébort, I.; Frebortova, J. Acetic acid bacteria: A group of bacteria with versatile biotechnological applications. Biotechnol. Adv. 2015, 33, 1260–1271. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Chang, Y.; Xie, S.; Song, J.; Wang, M. Impacts of bioprocess engineering on product formation by Acetobacter pasteurianus. Appl. Microbiol. Biotechnol. 2018, 102, 2535–2541. [Google Scholar] [CrossRef]
- Kuenz, A.; Krull, S. Biotechnological production of itaconic acid—Things you have to know. Appl. Microbiol. Biotechnol. 2018, 102, 3901–3914. [Google Scholar] [CrossRef]
- Salgado, J.M.; Abrunhosa, L.; Venâncio, A.; Domínguez, J.M.; Belo, I. Integrated use of residues from olive mill and winery for lipase production by solid state fermentation with Aspergillus sp. Appl. Biochem. Biotechnol. 2014, 172, 1832–1845. [Google Scholar] [CrossRef]
- Tanyildizi, M.S.; Özer, D.; Elibol, M. Production of bacterial α-amylase by B. amyloliquefaciens under solid substrate fermentation. Biochem. Eng. J. 2007, 37, 294–297. [Google Scholar] [CrossRef]
- Vijayaraghavan, P.; Lazarus, S.; Vincent, S.G.P. De-hairing protease production by an isolated Bacillus cereus strain AT under solid-state fermentation using cow dung: Biosynthesis and properties. Saudi J. Biol. Sci. 2014, 21, 27–34. [Google Scholar] [CrossRef]
- Rajkumar, R.; Jayappriyan, K.R.; Rengasamy, R. Purification and characterization of a protease produced by Bacillus megaterium RRM2: Application in detergent and dehairing industries. J. Basic Microbiol. 2011, 51, 614–624. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, W.A.; Yusof, N.Z.; Nordin, N.; Zakaria, Z.A.; Rezali, M.F. Production and characterization of violacein by locally isolated Chromobacterium violaceum grown in agricultural wastes. Appl. Biochem. Biotechnol. 2012, 167, 1220–1234. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Jiang, P.; Lu, Y.; Ruan, Z.; Jiang, R.; Xing, X.H.; Wei, D. Optimization of culture conditions for violacein production by a new strain of Duganella sp. B2. Biochem. Eng. J. 2009, 44, 119–124. [Google Scholar] [CrossRef]
- Chen, J.; Zhao, L.; Xu, J.; Yang, R.; He, S.; Yan, X. Determination of oxidized scytonemin in Nostoc commune Vauch cultured on different conditions by high performance liquid chromatography coupled with triple quadrupole mass spectrometry. J. Appl. Phycol. 2013, 25, 1001–1007. [Google Scholar] [CrossRef]
- Kurbanoglu, E.B.; Ozdal, M.; Ozdal, O.G.; Algur, O.F. Enhanced production of prodigiosin by Serratia marcescens MO-1 using ram horn peptone. Braz. J. Microbiol. 2015, 46, 631–637. [Google Scholar] [CrossRef]
- Borić, M.; Danevčič, T.; Stopar, D. Prodigiosin from Vibrio sp. DSM 14379; a new UV-protective pigment. Microb. Ecol. 2011, 62, 528–536. [Google Scholar] [CrossRef]
- Zhao, J.; Li, Q.; Sun, T.; Zhu, X.; Xu, H.; Tang, J.; Zhang, X.; Ma, Y. Engineering central metabolic modules of Escherichia coli for improving β-carotene production. Metabol Eng. 2013, 17, 42–50. [Google Scholar] [CrossRef]
- Larroude, M.; Celinska, E.; Back, A.; Thomas, S.; Nicaud, J.M.; Ledesma-Amaro, R. A synthetic biology approach to transform Yarrowia lipolytica into a competitive biotechnological producer of β-carotene. Biotechnol. Bioeng. 2018, 115, 464–472. [Google Scholar] [CrossRef]
- Tokui, M.; Kubodera, T.; Gomi, K.; Yamashita, N.; Nishimura, A. Construction of a thiamine pyrophosphate high-producing strain of Aspergillus oryzae by overexpression of three genes involved in thiamine biosynthesis. J. Biosci. Bioeng. 2011, 111, 388–390. [Google Scholar] [CrossRef]
- Schwechheimer, S.K.; Park, E.Y.; Revuelta, J.L.; Becker, J.; Wittmann, C. Biotechnology of riboflavin. Appl. Microbiol. Biotechnol. 2016, 100, 2107–2119. [Google Scholar] [CrossRef]
- Wang, Y.; Liu, L.; Jin, Z.; Zhang, D. Microbial Cell Factories for Green Production of Vitamins. Front. Bioeng. Biotechnol. 2021, 9, 473. [Google Scholar] [CrossRef] [PubMed]
- Dmytruk, K.V.; Ruchala, J.; Fayura, L.R.; Chrzanowski, G.; Dmytruk, O.V.; Tsyrulnyk, A.O.; Andreieva, Y.A.; Fedorovych, D.V.; Motyka, O.I.; Mattanovich, D.; et al. Efficient production of bacterial antibiotics aminoriboflavin and roseoflavin in eukaryotic microorganisms, yeasts. Microb. Cell Factories 2023, 22, 132. [Google Scholar] [CrossRef] [PubMed]
- Chandra, N.; Kumar, S. Antibiotics Producing Soil Microorganisms. In Antibiotics and Antibiotics Resistance Genes in Soils; Hashim, M.Z., Strezov, V., Varma, A., Eds.; Springer International Publishing AG: Berlin/Heidelberg, Germany, 2017; Volume 51, pp. 1–18. [Google Scholar]
- Bhosale, P.; Larson, A.J.; Bernstein, P.S. (Factorial analysis of tricarboxylic acid cycle intermediates for optimization of zeaxanthin production from Flavobacterium multivorum. J. Appl. Microbiol. 2004, 96, 623–629. [Google Scholar] [CrossRef] [PubMed]
- Shahina, M.; Hameed, A.; Lin, S.Y.; Lee, R.J.; Lee, M.R.; Young, C.C. Gramella planctonica sp. nov., a zeaxanthin-producing bacterium isolated from surface seawater, and emended descriptions of Gramella aestuarii and Gramella echinicola. Anton. Van Leeu. 2014, 105, 771–779. [Google Scholar] [CrossRef]
- Sowmya, R.; Sachindra, N.M. Carotenoid production by Formosa sp. KMW, a marine bacteria of Flavobacteriaceae family: Influence of culture conditions and nutrient composition. Biocatal. Agri Biotechnol. 2015, 4, 559–567. [Google Scholar] [CrossRef]
- Joshi, C.; Singhal, R.S. Modelling and optimization of zeaxanthin production by Paracoccus zeaxanthinifaciens ATCC 21588 using hybrid genetic algorithm techniques. Biocatal. Agri Biotechnol. 2016, 8, 228–235. [Google Scholar] [CrossRef]
- Barredo, J.L.; García-Estrada, C.; Kosalkova, K.; Barreiro, C. Biosynthesis of astaxanthin as a main carotenoid in the heterobasidiomycetous yeast Xanthophyllomyces dendrorhous. J. Fungi 2017, 3, 44. [Google Scholar] [CrossRef]
- Krupa, D.; Nakkeeran, E.; Kumaresan, N.; Vijayalakshmi, G.; Subramanian, R. Extraction, purification and concentration of partially saturated canthaxanthin from Aspergillus carbonarius. Bioresour. Technol. 2010, 101, 7598–7604. [Google Scholar] [CrossRef] [PubMed]
- Hojjati, M.; Razavi, S.H.; Rezaei, K.; Gilani, K. Stabilization of canthaxanthin produced by Dietzia natronolimnaea HS-1 with spray drying microencapsulation. J. Food Sci. Technol. 2014, 51, 2134–2140. [Google Scholar] [CrossRef]
- Market Research Future. Available online: https://www.marketresearchfuture.com/reports/bio-based-chemicals-market-5706 (accessed on 17 January 2024).
- Ko, Y.S.; Kim, J.W.; Lee, J.A.; Han, T.; Kim, G.B.; Park, J.E.; Lee, S.Y. Tools and strategies of systems metabolic engineering for the development of microbial cell factories for chemical production. Chem. Soc. Rev. 2020, 49, 4615–4636. [Google Scholar] [CrossRef]
- Runguphan, W.; Keasling, J.D. Metabolic engineering of Saccharomyces cerevisiae for production of fatty acid-derived biofuels and chemicals. Metab. Eng. 2014, 21, 103–113. [Google Scholar] [CrossRef]
- Patra, P.; Das, M.; Kundu, P.; Ghosh, A. Recent advances in systems and synthetic biology approaches for developing novel cell-factories in non-conventional yeasts. Biotechnol. Adv. 2021, 47, 107695. [Google Scholar] [CrossRef] [PubMed]
- Goel, A.; Wortel, M.T.; Molenaar, D.; Teusink, B. Metabolic shifts: A fitness perspective for microbial cell factories. Biotechnol. Lett. 2012, 34, 2147–2160. [Google Scholar] [CrossRef] [PubMed]
- Archer, D.B. Filamentous fungi as microbial cell factories for food use. Curr. Opin. Biotechnol. 2000, 11, 478–483. [Google Scholar] [CrossRef]
- Woolston, B.M.; Edgar, S.; Stephanopoulos, G. Metabolic engineering: Past and future. Annu. Rev. Chem. Biomol. Eng. 2013, 4, 259–288. [Google Scholar] [CrossRef] [PubMed]
- Ling, H.; Teo, W.; Chen, B.; Leong, S.S.J.; Chang, M.W. Microbial tolerance engineering toward biochemical production: From lignocellulose to products. Curr. Opin. Biotechnol. 2014, 29, 99–106. [Google Scholar] [CrossRef]
- Lee, S.Y.; Kim, H.U.; Chae, T.U.; Cho, J.S.; Kim, J.W.; Shin, J.H.; Kim, D.I.; Ko, Y.S.; Jang, W.D.; Jang, Y.S. A comprehensive metabolic map for production of bio-based chemicals. Nat. Catal. 2019, 2, 18–33. [Google Scholar] [CrossRef]
- Rangel, A.E.; Gomez Ramirez, J.M.; Gonzalez Barrios, A.F. From industrial by-products to value-added compounds: The design of efficient microbial cell factories by coupling systems metabolic engineering and bioprocesses. Biofuels. Bioprod. Biorefin. 2020, 14, 1228–1238. [Google Scholar] [CrossRef]
- Li, Y.; Lin, Z.; Huang, C.; Zhang, Y.; Wang, Z.; Tang, Y.J.; Chen, T.; Zhao, X. Metabolic engineering of Escherichia coli using CRISPR–Cas9 meditated genome editing. Metab. Eng. 2015, 31, 13–21. [Google Scholar] [CrossRef]
- Tong, Y.; Charusanti, P.; Zhang, L.; Weber, T.; Lee, S.Y. CRISPR-Cas9 based engineering of actinomycetal genomes. ACS Synth. Biol. 2015, 4, 1020–1029. [Google Scholar] [CrossRef]
- Blin, K.; Pedersen, L.E.; Weber, T.; Lee, S.Y. CRISPy-web: An online resource to design sgRNAs for CRISPR applications. Synth. Syst. Biotechnol. 2016, 1, 118–121. [Google Scholar] [CrossRef]
- Choi, S.; Song, H.; Lim, S.W.; Kim, T.Y.; Ahn, J.H.; Lee, J.W.; Lee, M.H.; Lee, S.Y. Highly selective production of succinic acid by metabolically engineered Mannheimia succiniciproducens and its efficient purification. Biotechnol. Bioeng. 2016, 113, 2168–2177. [Google Scholar] [CrossRef]
- Zhang, X.; Tervo, C.J.; Reed, J.L. Metabolic assessment of E. coli as a biofactory for commercial products. Metab. Eng. 2016, 35, 64–74. [Google Scholar] [CrossRef]
- Otero, J.M.; Cimini, D.; Patil, K.R.; Poulsen, S.G.; Olsson, L.; Nielsen, J. Industrial systems biology of Saccharomyces cerevisiae enables novel succinic acid cell factory. PLoS ONE 2013, 8, e54144. [Google Scholar] [CrossRef]
- Huang, H.; Liu, H.; Gan, Y.R. Genetic modification of critical enzymes and involved genes in butanol biosynthesis from biomass. Biotechnol. Adv. 2010, 28, 651–657. [Google Scholar] [CrossRef] [PubMed]
- Gopinath, V.; Nampoothiri, K.M. Corynebacterium glutamicum. In Encyclopedia of Food Microbiology, 1st ed.; Batt, C.A., Tortorello, M.L., Eds.; Elsevier Ltd, Academic Press: Amsterdam, Netherlands, 2007; Volume 1, pp. 504–517. [Google Scholar]
- Kim, H.T.; Baritugo, K.A.; Oh, Y.H.; Hyun, S.M.; Khang, T.U.; Kang, K.H.; Jung, S.H.; Song, B.K.; Park, K.; Kim, K.; et al. Metabolic engineering of Corynebacterium glutamicum for the high-level production of cadaverine that can be used for the synthesis of biopolyamide 510. ACS Sustain. Chem. Eng. 2018, 6, 5296–5305. [Google Scholar] [CrossRef]
- Paddon, C.J.; Westfall, P.J.; Pitera, D.J.; Benjamin, K.; Fisher, K.; McPhee, D.; Leavell, M.D.; Tai, A.; Main, A.; Eng, D.; et al. High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 2013, 496, 528–532. [Google Scholar] [CrossRef] [PubMed]
- Choi, O.; Wu, C.Z.; Kang, S.Y.; Ahn, J.S.; Uhm, T.B.; Hong, Y.S. Biosynthesis of plant-specific phenylpropanoids by construction of an artificial biosynthetic pathway in Escherichia coli. J. Ind. Microbiol. Biotechnol. 2011, 38, 1657–1665. [Google Scholar] [CrossRef] [PubMed]
- Park, S.Y.; Binkley, R.M.; Kim, W.J.; Lee, M.H.; Lee, S.Y. Metabolic engineering of Escherichia coli for high-level astaxanthin production with high productivity. Metabol Eng. 2018, 49, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Chae, T.U.; Ko, Y.S.; Hwang, K.S.; Lee, S.Y. Metabolic engineering of Escherichia coli for the production of four-, five-and six-carbon lactams. Metabol Eng. 2017, 41, 82–91. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Kao, E.; Wang, G.; Baidoo, E.E.; Chen, M.; Keasling, J.D. Metabolic engineering of Escherichia coli for the biosynthesis of 2-pyrrolidone. Metabol Eng. Commun. 2016, 3, 1–7. [Google Scholar] [CrossRef]
- Choi, O.; Lee, J.K.; Kang, S.Y.; Pandey, R.P.; Sohng, J.K.; Ahn, J.S.; Hong, Y.S. Construction of artificial biosynthetic pathways for resveratrol glucoside derivatives. J. Microbiol. Biotechnol. 2014, 24, 614–618. [Google Scholar] [CrossRef] [PubMed]
- Yim, H.; Haselbeck, R.; Niu, W.; Pujol-Baxley, C.; Burgard, A.; Boldt, J.; Khandurina, J.; Trawick, J.D.; Osterhout, R.E.; Stephen, R.; et al. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol. Nat. Chem. Biol. 2011, 7, 445–452. [Google Scholar] [CrossRef] [PubMed]
- Gong, Z.; Nielsen, J.; Zhou, Y.J. Engineering robustness of microbial cell factories. Biotechnol. J. 2017, 12, 1700014. [Google Scholar] [CrossRef]
- Cho, J.S.; Kim, G.B.; Eun, H.; Moon, C.W.; Lee, S.Y. Designing microbial cell factories for the production of chemicals. JACS Au 2022, 2, 1781–1799. [Google Scholar] [CrossRef]
- Choi, K.R.; Jang, W.D.; Yang, D.; Cho, J.S.; Park, D.; Lee, S.Y. Systems metabolic engineering strategies: Integrating systems and synthetic biology with metabolic engineering. Trends Biotechnol. 2019, 37, 817–837. [Google Scholar] [CrossRef]
- Thomford, N.E.; Senthebane, D.A.; Rowe, A.; Munro, D.; Seele, P.; Maroyi, A.; Dzobo, K. Natural products for drug discovery in the 21st century: Innovations for novel drug discovery. Int. J. Mol. Sci. 2018, 19, 1578. [Google Scholar] [CrossRef]
- Li, J.; Rong, L.; Zhao, Y.; Li, S.; Zhang, C.; Xiao, D.; Foo, J.L.; Yu, A. Next-generation metabolic engineering of non-conventional microbial cell factories for carboxylic acid platform chemicals. Biotechnol. Adv. 2020, 43, 107605. [Google Scholar] [CrossRef] [PubMed]
- Gomes, R.J.; de Fatima Borges, M.; de Freitas Rosa, M.; Castro-Gómez, R.J.H.; Spinosa, W.A. Acetic acid bacteria in the food industry: Systematics, characteristics and applications. Food Technol. Biotechnol. 2018, 56, 139. [Google Scholar] [CrossRef]
- Song, H.; Lee, S.Y. Production of succinic acid by bacterial fermentation. Enzym. Microb. Technol. 2006, 39, 352–361. [Google Scholar] [CrossRef]
- Dwidar, M.; Park, J.Y.; Mitchell, R.J.; Sang, B.I. The future of butyric acid in industry. Sci. World. J. 2012, 2012, 471417. [Google Scholar] [CrossRef]
- Jiang, L.; Cui, H.; Zhu, L.; Hu, Y.; Xu, X.; Li, S.; Huang, H. Enhanced propionic acid production from whey lactose with immobilized Propionibacterium acidipropionici and the role of trehalose synthesis in acid tolerance. Green Chem. 2015, 17, 250–259. [Google Scholar] [CrossRef]
- Wang, Z.; Ammar, E.M.; Zhang, A.; Wang, L.; Lin, M.; Yang, S.T. Engineering Propionibacterium freudenreichii subsp. Shermanii for enhanced propionic acid fermentation: Effects of overexpressing propionyl-CoA: Succinate CoA transferase. Metabol Eng. 2015, 27, 46–56. [Google Scholar] [CrossRef]
- Guan, N.; Li, J.; Shin, H.D.; Du, G.; Chen, J.; Liu, L. Metabolic engineering of acid resistance elements to improve acid resistance and propionic acid production of Propionibacterium jensenii. Biotechnol. Bioeng. 2016, 113, 1294–1304. [Google Scholar] [CrossRef] [PubMed]
- Rabah, H.; Carmo, F.L.R.D.; Jan, G. Dairy propionibacteria: Versatile probiotics. Microorganisms. 2017, 5, 24. [Google Scholar] [CrossRef] [PubMed]
- Upadhyaya, B.P.; DeVeaux, L.C.; Christopher, L.P. Metabolic engineering as a tool for enhanced lactic acid production. Trends Biotechnol. 2014, 32, 637–644. [Google Scholar] [CrossRef] [PubMed]
- Tamakawa, H.; Ikushima, S.; Yoshida, S. Efficient production of L-lactic acid from xylose by a recombinant Candida utilis strain. J. Biosci. Bioeng. 2012, 113, 73–75. [Google Scholar] [CrossRef] [PubMed]
- Papagianni, M.; Avramidis, N. Lactococcus lactis as a cell factory: A twofold increase in phosphofructokinase activity results in a proportional increase in specific rates of glucose uptake and lactate formation. Enzym. Microbiol. Technol. 2011, 49, 197–202. [Google Scholar] [CrossRef] [PubMed]
- Dhillon, G.S.; Brar, S.K.; Kaur, S.; Verma, M. Bioproduction and extraction optimization of citric acid from Aspergillus niger by rotating drum type solid-state bioreactor. Ind. Crop. Prod. 2013, 41, 78–84. [Google Scholar] [CrossRef]
- McKinlay, J.B.; Laivenieks, M.; Schindler, B.D.; McKinlay, A.A.; Siddaramappa, S.; Challacombe, J.F.; Lowry, S.R.; Clum, A.; Lapidus, A.L.; Burkhart, K.B.; et al. genomic perspective on the potential of Actinobacillus succinogenes for industrial succinate production. BMC Genom. 2010, 11, 680. [Google Scholar] [CrossRef]
- Tsuji, A.; Okada, S.; Hols, P.; Satoh, E. Metabolic engineering of Lactobacillus plantarum for succinic acid production through activation of the reductive branch of the tricarboxylic acid cycle. Enzym. Microb. Technol. 2013, 53, 97–103. [Google Scholar] [CrossRef]
- Uchikura, H.; Ninomiya, K.; Takahashi, K.; Tsuge, Y. Requirement of de novo synthesis of pyruvate carboxylase in long-term succinic acid production in Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 2020, 104, 4313–4320. [Google Scholar] [CrossRef]
- Lan, E.I.; Wei, C.T. Metabolic engineering of cyanobacteria for the photosynthetic production of succinate. Metabol Eng. 2016, 38, 483–493. [Google Scholar] [CrossRef]
- Cok, B.; Tsiropoulos, I.; Roes, A.L.; Patel, M.K. Succinic acid production derived from carbohydrates: An energy and greenhouse gas assessment of a platform chemical toward a bio-based economy. Biofuel Bioprod. Biorefin. 2016, 8, 16–29. [Google Scholar] [CrossRef]
- Cardinale, S.; Tueros, F.G.; Sommer, M.O.A. Genetic-metabolic coupling for targeted metabolic engineering. Cell Rep. 2017, 20, 1029–1037. [Google Scholar] [CrossRef]
- Sych, J.M.; Lacroix, C.; Stevens, M.J.A. Vitamin B12 : Physiology, production and application. In Industrial Biotechnology of Vitamins, Biopigments, and Antioxidants; Vandamme, E.J., Revuelta, J.L., Eds.; Wiley-VCH: Hoboken, NJ, USA, 2016; pp. 129–159. [Google Scholar]
- Xia, W.; Chen, W.; Peng, W.F.; Li, K.T. Industrial vitamin B12 production by Pseudomonas denitrificans using maltose syrup and corn steep liquor as the cost-effective fermentation substrates. Bioprocess Biosyst. Eng. 2015, 38, 1065–1073. [Google Scholar] [CrossRef]
- Cai, Y.; Xia, M.; Dong, H.; Qian, Y.; Zhang, T.; Zhu, B.; Wu, J.; Zhang, D. Engineering a vitamin B12 high-throughput screening system by riboswitch sensor in Sinorhizobium meliloti. BMC Biotechnol. 2018, 18, 27. [Google Scholar] [CrossRef]
- Wang, H.; Sun, X.; Wang, L.; Wu, H.; Zhao, G.; Liu, H.; Wang, P.; Zheng, Z. Coproduction of menaquinone-7 and nattokinase by Bacillus subtilis using soybean curd residue as a renewable substrate combined with a dissolved oxygen control strategy. Ann. Microbiol. 2018, 68, 655–665. [Google Scholar] [CrossRef]
- Georgi, T.; Rittmann, D.; Wendisch, V.F. Lysine and glutamate production by Corynebacterium glutamicum on glucose, fructose and sucrose: Roles of malic enzyme and fructose-1, 6-bisphosphatase. Metabol Eng. 2005, 7, 291–301. [Google Scholar] [CrossRef] [PubMed]
- Bommareddy, R.R.; Chen, Z.; Rappert, S.; Zeng, A.P. A de novo NADPH generation pathway for improving lysine production of Corynebacterium glutamicum by rational design of the coenzyme specificity of glyceraldehyde 3-phosphate dehydrogenase. Metabol Eng. 2014, 25, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Hirasawa, T.; Shimizu, H. Recent advances in amino acid production by microbial cells. Curr. Opin. Biotechnol. 2016, 42, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Hajnal, I. Meeting Report: Cold Spring Harbor Asia Synthetic Biology Meeting. Biotechnol. J. 2016, 11, 197–198. [Google Scholar] [CrossRef] [PubMed]
- Heider, S.A.; Wendisch, V.F. Engineering microbial cell factories: Metabolic engineering of Corynebacterium glutamicum with a focus on non-natural products. Biotechnol. J. 2015, 10, 1170–1184. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, S.; Suda, M.; Uematsu, K.; Natsuma, Y.; Hiraga, K.; Jojima, T.; Inui, M.; Yukawa, H. Engineering of Corynebacterium glutamicum for high-yield L-valine production under oxygen deprivation conditions. Appl. Environ. Microbiol. 2013, 79, 1250–1257. [Google Scholar] [CrossRef] [PubMed]
- Park, J.H.; Jang, Y.S.; Lee, J.W.; Lee, S.Y. Escherichia coli W as a new platform strain for the enhanced production of L-Valine by systems metabolic engineering. Biotechnol. Bioeng. 2011, 108, 1140–1147. [Google Scholar] [CrossRef] [PubMed]
- Nagai, Y.; Ito, H.; Yasueda, H. Amino acid production: L-Lysine. In Encyclopedia of Industrial Biotechnology: Bioprocess, Bioseparation, and Cell Technology, 1st ed.; Flickinger, M.C., Ed.; Wiley Online Library: Hoboken, NJ, USA, 2009; Volume 7, pp. 1–10. [Google Scholar]
- Xu, J.M.; Li, J.Q.; Zhang, B.; Liu, Z.Q.; Zheng, Y.G. Fermentative production of the unnatural amino acid L-2-aminobutyric acid based on metabolic engineering. Microb. Cell Factories 2019, 18, 43. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Chen, X.; Wan, F.; Zhang, B.; Chen, J.; Xiong, Y. Effect of Tween 40 and DtsR1 on l-arginine overproduction in Corynebacterium crenatum. Microb. Cell Factories 2015, 14, 119. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Yu, M.; Zhou, Y.; Li, Y.; Ye, B.C. Systematic pathway engineering of Corynebacterium glutamicum S9114 for L-ornithine production. Microb. Cell Factories 2017, 16, 158. [Google Scholar] [CrossRef] [PubMed]
- Qin, J.; Zhou, Y.J.; Krivoruchko, A.; Huang, M.; Liu, L.; Khoomrung, S.; Nielsen, J. Modular pathway rewiring of Saccharomyces cerevisiae enables high-level production of L-ornithine. Nat. Commun. 2015, 6, 8224. [Google Scholar] [CrossRef]
- Yang, B.; Feng, X.; Li, C. Microbial Cell Factory for Efficiently Synthesizing Plant Natural Products via Optimizing the Location and Adaptation of Pathway on Genome Scale. Front. Bioeng. Biotechnol. 2020, 8, 969. [Google Scholar] [CrossRef]
- Liu, X.; Ding, W.; Jiang, H. Engineering microbial cell factories for the production of plant natural products: From design principles to industrial-scale production. Microb. Cell Factories 2017, 16, 125. [Google Scholar] [CrossRef]
- Beyraghdar Kashkooli, A.; Farmanpour-Kalalagh, K.; Babaei, A. Yeast Synthetic Biology for Production of Artemisinin as an Antimalarial Drug. In Synthetic Biology of Yeasts: Tools and Applications; Springer International Publishing: Cham, Switzerland, 2022; pp. 157–180. [Google Scholar]
- Wu, C.-F.; Yang, J.-Y.; Wang, F.; Wang, X.-X. Resveratrol: Botanical origin, pharmacological activity and applications. Chin. J. Nat. Med. 2013, 11, 1–15. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, S.; Yu, O. Metabolic engineering of flavonoids in plants and microorganisms. Appl. Microbiol. Biotechnol. 2011, 91, 949–956. [Google Scholar] [CrossRef]
- Beekwilder, J.; Wolswinkel, R.; Jonker, H.; Hall, R.; de Vos, C.R.; Bovy, A. Production of resveratrol in recombinant microorganisms. Appl. Environ. Microbiol. 2006, 72, 5670–5672. [Google Scholar] [CrossRef]
- Lim, C.G.; Fowler, Z.L.; Hueller, T.; Schaffer, S.; Koffas, M.A. High-yield resveratrol production in engineered Escherichia coli. Appl. Environ. Microbiol. 2011, 77, 3451–3460. [Google Scholar] [CrossRef]
- Marienhagen, J.; Bott, M. Metabolic engineering of microorganisms for the synthesis of plant natural products. J. Biotechnol. 2013, 163, 166–178. [Google Scholar] [CrossRef] [PubMed]
- Mata-Gómez, L.C.; Montañez, J.C.; Méndez-Zavala, A.; Aguilar, C.N. Biotechnological production of carotenoids by yeasts: An overview. Microb. Cell Factories 2014, 13, 12. [Google Scholar] [CrossRef] [PubMed]
- Global Carotenoids Market. Available online: https://www.bccresearch.com/partners/verified-market-research/global-carotenoids-market.html (accessed on 17 January 2024).
- Sarnaik, A.; Sawant, K.; Khadilkar, J.; Pillai, G.; Pandit, R.; Lali, A. Cyanobacterial Cell Factories for Improved Carotenoid Biosynthesis through a Synthetic Biology Approach. In Next Generation Biomanufacturing Technologies; American Chemical Society: Washington, DC, USA, 2019; pp. 23–39. [Google Scholar]
- Kuzina, V.; Cerdá-Olmedo, E. Ubiquinone and carotene production in the Mucorales Blakeslea and Phycomyces. Appl. Microbiol. Biotechnol. 2007, 76, 991–999. [Google Scholar] [CrossRef] [PubMed]
- Das, A.; Yoon, S.H.; Lee, S.H.; Kim, J.Y.; Oh, D.K.; Kim, S.W. An update on microbial carotenoid production: Application of recent metabolic engineering tools. Appl. Microbiol. Biotechnol. 2007, 77, 505–512. [Google Scholar] [CrossRef]
- Sarnaik, A.; Nambissan, V.; Pandit, R.; Lali, A. Recombinant Synechococcus elongatus PCC 7942 for improved zeaxanthin production under natural light conditions. Algal Res. 2018, 36, 139–151. [Google Scholar] [CrossRef]
- Kim, J.H.; Kang, H.J.; Yu, B.J.; Kim, S.C.; Lee, P.C. Planococcus faecalis sp. nov., a carotenoid-producing species isolated from stools of Antarctic penguins. Int. J. Syst. Evol. Microbiol. 2015, 65, 3373–3378. [Google Scholar] [CrossRef] [PubMed]
- Köcher, S.; Breitenbach, J.; Müller, V.; Sandmann, G. Structure, function and biosynthesis of carotenoids in the moderately halophilic bacterium Halobacillus halophilus. Arch. Microbiol. 2009, 191, 95–104. [Google Scholar] [CrossRef]
- Steiger, S.; Perez-Fons, L.; Cutting, S.M.; Fraser, P.D.; Sandmann, G. Annotation and functional assignment of the genes for the C30 carotenoid pathways from the genomes of two bacteria: Bacillus indicus and Bacillus firmus. Microbiology 2015, 161, 194–202. [Google Scholar] [CrossRef] [PubMed]
- Hameed, A.; Shahina, M.; Lin, S.Y.; Cho, J.C.; Lai, W.A.; Young, C.C. Kordiaaquimaris sp. nov., a zeaxanthin-producing member of the family Flavobacteriaceae isolated from surface seawater, and emended description of the genus Kordia. Int. J. Syst. Evol. Microbiol. 2013, 63, 4790–4796. [Google Scholar] [CrossRef] [PubMed]
- Hameed, A.; Shahina, M.; Lin, S.Y.; Lai, W.A.; Hsu, Y.H.; Liu, Y.C.; Young, C.C. Aquibacter zeaxanthinifaciens gen. nov., sp. nov., a zeaxanthin-producing bacterium of the family Flavobacteriaceae isolated from surface seawater, and emended descriptions of the genera Aestuariibaculum and Gaetbulibacter. Int. J. Syst. Evol. Microbiol. 2014, 64, 138–145. [Google Scholar] [CrossRef] [PubMed]
- Hameed, A.; Shahina, M.; Lin, S.Y.; Liu, Y.C.; Lai, W.A.; Young, C.C. Gramella oceani sp. nov., a zeaxanthin-producing bacterium of the family Flavobacteriaceae isolated from marine sediment. Int. J. Syst. Evol. Microbiol. 2014, 64, 2675–2681. [Google Scholar] [CrossRef] [PubMed]
- Manikandan, R.; Thiagarajan, R.; Goutham, G.; Arumugam, M.; Beulaja, M.; Rastrelli, L.; Woźniak, K.S.; Habtemariam, S.; Orhan, I.E.; Nabavi, S.F.; et al. Zeaxanthin and ocular health, from bench to bedside. Fitoterapia 2016, 109, 58–66. [Google Scholar] [CrossRef] [PubMed]
- Ambati, R.R.; Phang, S.M.; Ravi, S.; Aswathanarayana, R.G. Astaxanthin: Sources, extraction, stability, biological activities and its commercial applications—A review. Mar. Drugs 2014, 12, 128–152. [Google Scholar] [CrossRef] [PubMed]
- Tao, L.; Rouvière, P.E.; Cheng, Q. A carotenoid synthesis gene cluster from a non-marine Brevundimonas that synthesizes hydroxylated astaxanthin. Gene 2006, 379, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Kim, Y.S.; Choi, T.J.; Lee, W.J.; Kim, Y.T. Paracoccus haeundaensis sp. nov., a Gram-negative, halophilic, astaxanthin-producing bacterium. Int. J. Syst. Evol. Microbiol 2004, 54, 1699–1702. [Google Scholar] [CrossRef]
- Asker, D.; Beppu, T.; Ueda, K. Sphingomonas astaxanthinifaciens sp. nov., a novel astaxanthin-producing bacterium of the family Sphingomonadaceae isolated from Misasa, Tottori, Japan. FEMS Microbiol. Lett. 2007, 273, 140–148. [Google Scholar] [CrossRef]
- Li, C.; Swofford, C.A.; Sinskey, A.J. Modular engineering for microbial production of carotenoids. Metab. Eng. Commun. 2020, 10, e00118. [Google Scholar] [CrossRef]
- Jin, J.; Wang, Y.; Yao, M.; Gu, X.; Li, B.; Liu, H.; Ding, M.; Xiao, W.; Yuan, Y. Astaxanthin overproduction in yeast by strain engineering and new gene target uncovering. Biotechnol. Biofuels 2018, 11, 230. [Google Scholar] [CrossRef] [PubMed]
- Basiony, M.; Ouyang, L.; Wang, D.; Yu, J.; Zhou, L.; Zhu, M.; Wang, X.; Feng, J.; Dai, J.; Shen, Y.; et al. Optimization of microbial cell factories for astaxanthin production: Biosynthesis and regulations, engineering strategies and fermentation optimization strategies. Synth. Syst. Biotechnol. 2022, 7, 689–704. [Google Scholar] [CrossRef] [PubMed]
- Global Premium Fragrances Market to Reach $31.4 Billion by 2027. Report Linker. Available online: https://www.reportlinker.com/p06032650/?utm_source=GNW (accessed on 11 January 2024).
- Flavors and Fragrance Market by Type (Flavors and Fragrance), Nature (Natural and Synthetic), and Application (Food & Beverages, Cosmetics & Personal Care, Home Care and Fabric Care): Global Opportunity Analysis and Industry Forecast, 2021–2027. Available online: https://www.alliedmarketresearch.com/flavors-and-fragrances-market (accessed on 14 January 2024).
- Cosmetics Market Size, Share & Trends Analysis Report By Product (Skin Care, Hair Care, Makeup, Fragrance), by End-user (Men, Women), by Distribution Channel, by Region, and Segment Forecasts, 2023–2030. Research and Markets. Available online: https://www.researchandmarkets.com/reports/5648046/cosmetics-market-size-share-and-trends-analysis (accessed on 7 January 2024).
- Van Wyk, N.; Kroukamp, H.; Pretorius, I.S. The smell of synthetic biology: Engineering strategies for aroma compound production in yeast. Fermentation 2018, 4, 54. [Google Scholar] [CrossRef]
- Barden, T.C. Indoles: Industrial, agricultural and over-the-counter uses. Heterocyclic Scaffolds II. ChemInform 2014, 42, 31–46. [Google Scholar]
- Mindt, M.; Beyraghdar Kashkooli, A.; Suarez-Diez, M.; Ferrer, L.; Jilg, T.; Bosch, D.; Santos, V.M.; Wendisch, V.F.; Cankar, K. Production of indole by Corynebacterium glutamicum microbial cell factories for flavor and fragrance applications. Microb. Cell Factories 2022, 21, 45. [Google Scholar] [CrossRef] [PubMed]
- Czajka, J.J.; Nathenson, J.A.; Benites, V.T.; Baidoo, E.E.; Cheng, Q.; Wang, Y.; Tang, Y.J. Engineering the oleaginous yeast Yarrowia lipolytica to produce the aroma compound β-ionone. Microb. Cell Factories 2018, 17, 136. [Google Scholar] [CrossRef]
- Mi, J.; Becher, D.; Lubuta, P.; Dany, S.; Tusch, K.; Schewe, H.; Buchhaupt, M.; Schrader, J. De novo production of the monoterpenoid geranic acid by metabolically engineered Pseudomonas putida. Microb. Cell Factories 2014, 13, 170. [Google Scholar] [CrossRef]
- Barghini, P.; Di Gioia, D.; Fava, F.; Ruzzi, M. Vanillin production using metabolically engineered Escherichia coli under non-growing conditions. Microb. Cell Factories 2007, 6, 13. [Google Scholar] [CrossRef]
- Liu, Z.; Zhang, X.; Lei, D.; Qiao, B.; Zhao, G.R. Metabolic engineering of Escherichia coli for de novo production of 3-phenylpropanol via retrobiosynthesis approach. Microb. Cell Factories 2021, 20, 121. [Google Scholar] [CrossRef]
- Tao, X.Y.; Lin, Y.C.; Wang, F.Q.; Liu, Q.H.; Ma, Y.S.; Liu, M.; Wei, D.Z. Production of sesquiterpene patchoulol in mitochondrion-engineered Saccharomyces cerevisiae. Biotechnol. Lett. 2022, 44, 571–580. [Google Scholar] [CrossRef]
- Kampranis, S.C.; Makris, A.M. Developing a yeast cell factory for the production of terpenoids. Comput. Struct. Biotechnol. J. 2012, 3, e201210006. [Google Scholar] [CrossRef]
- Tilman, D.; Socolow, R.; Foley, J.A.; Hill, J.; Larson, E.; Lynd, L.; Pacala, S.; Reilly, J.; Searchinger, T.; Somerville, C.; et al. Beneficial biofuels—The food, energy, and environment trilemma. Science 2009, 325, 270–271. [Google Scholar] [CrossRef]
- Lan, E.I.; Liao, J.C. Metabolic engineering of cyanobacteria for 1-butanol production from carbon dioxide. Metab. Eng. 2011, 13, 353–363. [Google Scholar] [CrossRef]
- Silva, J.P.A.; Mussatto, S.I.; Roberto, I.C.; Teixeira, J.A. Ethanol production from xylose by Pichia stipitis NRRL Y-7124 in a stirred tank bioreactor. Braz. J. Chem. Eng. 2011, 28, 151–156. [Google Scholar]
- Aruna, A.; Nagavalli, M.; Girijashankar, V.; Ponamgi, S.P.D.; Swathisree, V.; Venkateswar, R.L. Direct bioethanol production by amylolytic yeast C andida albicans. Lett. Appl. Microbiol. 2015, 60, 229–236. [Google Scholar] [CrossRef]
- Kim, J.H.; Ryu, J.; Huh, I.Y.; Hong, S.K.; Kang, H.A.; Chang, Y.K. Ethanol production from galactose by a newly isolated Saccharomyces cerevisiae KL17. Bioprocess Biosyst. Eng. 2014, 37, 1871–1878. [Google Scholar] [CrossRef]
- Mauerhofer, L.M.; Zwirtmayr, S.; Pappenreiter, P.; Bernacchi, S.; Seifert, A.H.; Reischl, B.; Schmider, T.; Taubner, R.S.; Paulik, C.; Rittmann, S.K.M. Hyperthermophilic methanogenic archaea act as high-pressure CH4 cell factories. Commun. Biol. 2021, 4, 289. [Google Scholar] [CrossRef] [PubMed]
- Seifert, A.H.; Rittmann, S.; Bernacchi, S.; Herwig, C. Method for assessing the impact of emission gasses on physiology and productivity in biological methanogenesis. Bioresour. Technol. 2013, 136, 747–751. [Google Scholar] [CrossRef] [PubMed]
- Simon, K.M.R.; Seifert, A.H.; Bernacchi, S. Kinetics, multivariate statistical modelling, and physiology of CO2-based biological methane production. Appl. Energy 2018, 216, 751–760. [Google Scholar]
- Willquist, K.; Zeidan, A.A.; van Niel, E.W. Physiological characteristics of the extreme thermophile Caldicellulosiruptor saccharolyticus: An efficient hydrogen cell factory. Microb. Cell Factories 2010, 9, 89. [Google Scholar] [CrossRef] [PubMed]
- Gabrielyan, L.; Sargsyan, H.; Trchounian, A. Novel properties of photofermentative biohydrogen production by purple bacteria Rhodobacter sphaeroides: Effects of protonophores and inhibitors of responsible enzymes. Microb. Cell Factories 2015, 14, 131. [Google Scholar] [CrossRef] [PubMed]
- Callegari, A.; Bolognesi, S.; Cecconet, D.; Capodaglio, A.G. Production technologies, current role, and future prospects of biofuels feedstocks: A state-of-the-art review. Crit. Rev. Environ. Sci. Technol. 2020, 50, 384–436. [Google Scholar] [CrossRef]
- Singh, V.; Mani, I.; Chaudhary, D.K.; Dhar, P.K. Metabolic engineering of biosynthetic pathway for production of renewable biofuels. Appl. Biochem. Biotechnol. Appl. 2014, 172, 1158–1171. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.R.; Jiao, S.; Lee, S.Y. Metabolic engineering strategies toward production of biofuels. Curr. Opin. Chem. Biol. 2020, 59, 1–14. [Google Scholar] [CrossRef]
- Eng, T.; Demling, P.; Herbert, R.A.; Chen, Y.; Benites, V.; Martin, J.; Lipzen, A.; Baidoo, E.E.; Blank, L.M.; Petzold, C.J.; et al. Restoration of biofuel production levels and increased tolerance under ionic liquid stress is enabled by a mutation in the essential Escherichia coli gene cydC. Microb. Cell Factories 2018, 17, 159. [Google Scholar] [CrossRef]
- Cao, Y.; Mu, H.; Liu, W.; Zhang, R.; Guo, J.; Xian, M.; Liu, H. Electricigens in the anode of microbial fuel cells: Pure cultures versus mixed communities. Microb. Cell Factories 2019, 18, 39. [Google Scholar] [CrossRef]
- Cho, Y.K.; Donohue, T.J.; Tejedor, I.; Anderson, M.A.; McMahon, K.D.; Noguera, D.R. Development of a solar-powered microbial fuel cell. J. Appl. Microbiol. 2008, 104, 640–650. [Google Scholar] [CrossRef]
- Gomez, M.V.; Mai, G.; Greenwood, T.; Mullins, J.P. The development and maximization of a novel photosynthetic microbial fuel cell using Rhodospirillum rubrum. J. Emerg. Invest. 2014, 3, 1–7. [Google Scholar] [CrossRef]
- Borole, A.P.; O’Neill, H.; Tsouris, C.; Cesar, S. A microbial fuel cell operating at low pH using the acidophile Acidiphilium cryptum. Biotechnol. Lett. 2008, 30, 1367–1372. [Google Scholar] [CrossRef]
- Ren, Z.; Ward, T.E.; Regan, J.M. Electricity production from cellulose in a microbial fuel cell using a defined binary culture. Environ. Sci. Tech. 2007, 41, 4781–4786. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.A.; Ethiraj, B.; Cheng, C.K.; Yousuf, A.; Thiruvenkadam, S.; Prasad, R.; Rahman Khan, M.M. Enhanced current generation using mutualistic interaction of yeast-bacterial coculture in dual chamber microbial fuel cell. Ind. Eng. Chem. Res. 2018, 57, 813–821. [Google Scholar] [CrossRef]
- Islam, M.A.; Ong, H.R.; Ethiraj, B.; Cheng, C.K.; Khan, M.M.R. Optimization of co-culture inoculated microbial fuel cell performance using response surface methodology. J. Environ. Manag. 2018, 225, 242–251. [Google Scholar] [CrossRef] [PubMed]
- Recombinant Insulin. Novo Nordisk Pharmatech A/S. Available online: https://novonordiskpharmatech.com/products/recombinant-insulin/ (accessed on 23 November 2023).
- Quorn Mycoprotein. Available online: https://www.quorn.co.uk/mycoprotein (accessed on 4 January 2024).
- Sunsonzymes. Available online: https://www.sunsonzyme.com/ (accessed on 4 January 2024).
- Biosuccinium™ Reverdia. Available online: https://www.reverdia.com/biosuccinium/ (accessed on 7 January 2024).
- BioAstin® Cyanotech. Available online: https://www.cyanotech.com/astaxanthin/bioastin-water-dispersible-astaxanthin/ (accessed on 11 January 2024).
- Astapure® Algatech. Available online: https://www.algatech.com/algatech-product/astapure-natural-astaxanthin/ (accessed on 7 December 2023).
- Fucovital™ AlgaTech. Available online: https://www.algatech.com/algatech-product/fucovital/ (accessed on 13 January 2024).
- Purecane™ Amyris. Available online: https://amyris.com/2427/purecane-launches-in-kroger-stores-in-time-for-healthy-holiday-baking (accessed on 15 January 2024).
- Aji-No-Moto®. Available online: https://www.ajinomoto.com/msg/what-is-aji-no-moto-and-how-is-it-made#cf_block_1_cf_heading_txt (accessed on 31 December 2023).
- Nootkatone by Evolva™. Available online: https://evolva.com/nootkatone/ (accessed on 21 January 2024).
- Geno BDO™. Genomatica. Available online: https://www.genomatica.com/products/ (accessed on 21 January 2024).
- AvelaTM. Genomatica. Available online: https://www.genomatica.com/products/ (accessed on 18 December 2023).
- CaroCare® DSM. Available online: https://www.dsm.com/food-beverage/en_US/ingredients/beverages-and-brewing/beverage/carocare.html# (accessed on 7 January 2024).
- Biolys® Evonik. Available online: https://animal-nutrition.evonik.com/en/products-and-solutions/amino-acids/l-lysine/biolys-131967.html (accessed on 17 January 2024).
- Threamino® Evonik. Available online: https://animal-nutrition.evonik.com/en/products-and-solutions/amino-acids/l-threonine/threamino-131965.html (accessed on 17 January 2024).
- Song, C.W.; Park, J.M.; Chung, S.C.; Lee, S.Y.; Song, H. Microbial production of 2, 3-butanediol for industrial applications. J. Ind. Microbiol. 2019, 46, 1583–1601. [Google Scholar] [CrossRef]

| Bio Product | Microbes | References |
|---|---|---|
| Carboxylic/Organic Acids | ||
| Lactic acid | Lactobacillus acidophilus Lactobacillus fermentum Bacillus subtilis Weizmannia acidilactici Lactobacillus saerimneri | [24,25,26] |
| Citric acid | Aspergillus niger Aspergillus aculeatus Yarrowia lipolytica Bacillus licheniformis Arthrobacter paraffinens | [27,28,29] |
| Gluconic acid | Gluconobacter oxydans | [30] |
| Acetic acid | Acetobacter pasteurianus Acetobacter aceti | [31] |
| Itaconic acid | Aspergillus terreus | [32] |
| Enzymes | ||
| Lipase | Aspergillus ibericus Aspergillus niger Aspergillus uvarum | [33] |
| Amylase | Bacillus amyloliquefaciens | [34] |
| Protease | Bacillus cereus Bacillus megaterium | [35,36] |
| Indole Derivatives | ||
| Violacein | Chromobacterium violaceum Duganella sp. B2 | [37,38] |
| Scytonemin | Nostoc commune | [39] |
| Prodigiosin | Saccharomyces marcescens Vibrio sp. | [40,41] |
| Vitamins | ||
| Vitamin A | Yarrowia lipolytica Escherichia coli | [42,43] |
| Vitamin B | Aspergillus oryzae Bacillus subtilis | [44,45] |
| Vitamin C | Gluconobacter oxydans Bacillus megaterium | [46] |
| Vitamin D | Rhodococcus sp. Streptomyces sp. Pseudonocardia sp. Mycobacterium sp. | [46] |
| Antibiotics | ||
| Roseoflavin | Streptomyces davaonensis Streptomyces cinnabarinus Komagataella phaffii | [47] |
| Penicillin | Penicillium notatum | [48] |
| Rifamycin | Amycolatopsis mediterrane | [48] |
| Tetracyclines | Streptomyces aureofaciens | [48] |
| Streptomycin | Streptomyces griseus | [48] |
| Carotenoids | ||
| Zeaxanthin | Flavobacterium multivorum Mesoflavibacter zeaxanthinifaciens Gramella planctonica Formosa sp. KMW Paracoccus zeaxanthinifaciens | [49,50,51,52] |
| Astaxanthin | Xanthophyllomyces dendrorhous | [53] |
| Canthaxanthin | Aspergillus carbonarius Dietzia natronolimnaea | [54,55] |
| Metabolic Engineering Approaches | Tools |
|---|---|
| Molecular tools |
|
| Enzyme engineering |
|
| Genome-scale metabolic models (GEMs) |
|
| Random mutagenesis of chassis strain |
|
| Tolerance and transporter engineering |
|
| |
| |
| |
| Lipid and membrane morphology engineering |
|
| Antibiotic-free systems |
|
| |
|
| Biocompound/Biochemical | Product Name | Company | Reference |
|---|---|---|---|
| Succinic acid | Biosuccinium™ | Reverdia | [184] |
| Astaxanthin | BioAstin® | Cyanotech | [185] |
| Astaxanthin | AstaPure® | AlgaTech | [186] |
| Fucoxanthin | Fucovital™ | AlgaTech | [187] |
| Erythritol (flavour/sweetener) | Purecane™ | Amyris | [188] |
| Monosodium glutamate (MSG) (flavor enhancer) | Aji-No-Moto® | Ajinomoto | [189] |
| Sesquiterpenoid (fragrance compound) | Nootkatone™ | Evolva | [190] |
| Bio-1,4-butanediol | Geno BDO™ | Genomatica | [191] |
| Natural (R)-1,3 Butanediol | Avela TM | Genomatica | [192] |
| β-Carotene | CaroCare® | DSM | [193] |
| L-lysine | Biolys® | Evonik | [194] |
| L-threonine | Threamino® | Evonik | [195] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chaudhary, R.; Nawaz, A.; Fouillaud, M.; Dufossé, L.; Haq, I.u.; Mukhtar, H. Microbial Cell Factories: Biodiversity, Pathway Construction, Robustness, and Industrial Applicability. Microbiol. Res. 2024, 15, 247-272. https://doi.org/10.3390/microbiolres15010018
Chaudhary R, Nawaz A, Fouillaud M, Dufossé L, Haq Iu, Mukhtar H. Microbial Cell Factories: Biodiversity, Pathway Construction, Robustness, and Industrial Applicability. Microbiology Research. 2024; 15(1):247-272. https://doi.org/10.3390/microbiolres15010018
Chicago/Turabian StyleChaudhary, Rida, Ali Nawaz, Mireille Fouillaud, Laurent Dufossé, Ikram ul Haq, and Hamid Mukhtar. 2024. "Microbial Cell Factories: Biodiversity, Pathway Construction, Robustness, and Industrial Applicability" Microbiology Research 15, no. 1: 247-272. https://doi.org/10.3390/microbiolres15010018
APA StyleChaudhary, R., Nawaz, A., Fouillaud, M., Dufossé, L., Haq, I. u., & Mukhtar, H. (2024). Microbial Cell Factories: Biodiversity, Pathway Construction, Robustness, and Industrial Applicability. Microbiology Research, 15(1), 247-272. https://doi.org/10.3390/microbiolres15010018











